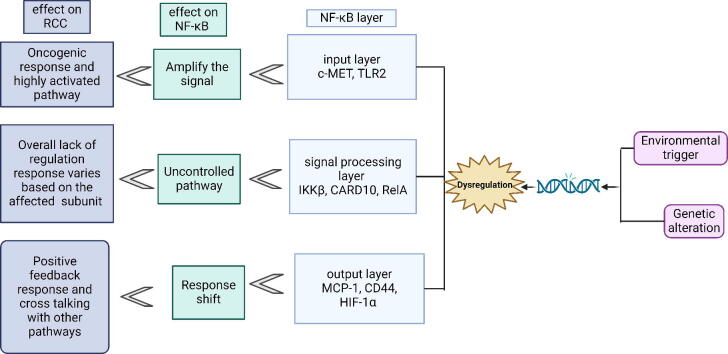
Graphical abstract
Keywords: Renal cell carcinoma, Genetic modification, NF-κB dysregulation, Molecular mechanism
Highlights
-
•
NF-κB is a cellular orchestrator that allow the shift of cellular response based on the environmental trigger.
-
•
Dysregulated NF-κB genes have been implicated in different cancers, including RCC.
-
•
Dysregulation of NF-κB would affect different layers of the pathway to sustain different cancer hallmarks.
-
•
Understanding the lack of regulation of NF-κB in different RCC subtypes impedes disease progression and treatment resistance.
Abstract
Background
The nuclear factor kappa B (NF-κB) is a critical pathway that regulates various cellular functions, including immune response, proliferation, growth, and apoptosis. Furthermore, this pathway is tightly regulated to ensure stability in the presence of immunogenic triggers or genotoxic stimuli. The lack of control of the NF-κB pathway can lead to the initiation of different diseases, mainly autoimmune diseases and cancer, including Renal cell carcinoma (RCC). RCC is the most common type of kidney cancer and is characterized by complex genetic composition and elusive molecular mechanisms.
Aim of review
The current review summarizes the mechanism of NF-κB dysregulation in different subtypes of RCC and its impact on pathogenesis.
Key scientific concept of review
This review highlights the prominent role of NF-κB in RCC development and progression by driving the expression of multiple genes and interplaying with different pathways, including the phosphatidylinositol 3-kinase (PI3K)/protein kinase B (Akt) pathway. In silico analysis of RCC cohorts and molecular studies have revealed that multiple NF-κB members and target genes are dysregulated. The dysregulation includes receptors such as TLR2, signal-transmitting members including RelA, and target genes, for instance, HIF-1α. The lack of effective regulatory mechanisms results in a constitutively active NF-κB pathway, which promotes cancer growth, migration, and survival. In this review, we comprehensively summarize the role of dysregulated NF-κB-related genes in the most common subtypes of RCC, including clear cell RCC (ccRCC), chromophobe RCC (chRCC), and papillary RCC (PRCC).
Introduction
In terms of global incidence, kidney cancer is one of the ten most common cancers among men and the 14th most common cancer among women. The incidence of kidney cancer is increasing every year, with a 1 % annual increase rate recorded between 2010 and 2019 in the United States [1]. In 2020, more than 400,000 cases of kidney cancer were registered, and almost 180,000 deaths were reported [2]. The most common type of kidney cancer is renal cell carcinoma (RCC) (accounting for >90 % of the cases).
The interplay between inflammation and cancer has been investigated extensively in many cancer types since the first link was proposed in 1863 by Rudolf Virchow [3]. Inflammation represents an instant and unspecific innate response to the stimulus, including pathogens, physical injury, or dust particles. In some cases, immune cells and pro-inflammatory mediators persist in the absence of external stimuli, leading to chronic inflammation. Besides, chronic inflammation has been stated as a major cancer risk factor [4]. The histological and molecular changes induced by chronic inflammation were demonstrated to enhance cell proliferation [5].
The nuclear factor kappa B (NF-κB) pathway is a key pathway that regulates inflammation along with other cellular functions such as growth, apoptosis, and immune response [6]. Dysregulation of the NF-κB pathway has been associated with multiple conditions, such as inflammatory and neurodegenerative diseases as well as cancer [7]. NF-κB pathway can be activated by different stimuli and various receptors, enabling it to switch between cellular functions. The innate immune response due to microbes or cell damage trigger instant and expeditious NF-κB activation, inducing immune cells recruitment and pro-inflammatory cytokines release. Conversely, during cell stress, NF-κB activation could enhance the expression of anti-apoptotic genes to promote cell survival [8].
NF-κB pathway: An overview and its dysregulation
The NF-κB signaling pathway is a crucial regulator of body responses discovered at the end of the 20th century. Initially, NF-κB was identified as an immunoglobulin (Ig) binding protein [9]. Activation of NF-κB can be initiated by a range of receptors that transduce the cell signals to generate a range of cellular responses [10]. The first layer of NF-κB is the input layer, which is composed of the different receptors and ligands that provoke canonical and non-canonical NF-κB activation. Correspondingly, a signal is then transmitted to a signal processing layer that consists of various mediators and adaptors linked to different receptors [11].
The NF-κB pathway constitutes different transcription factors that can directly affect gene transcription by binding to the promoter. The family of NF-κB transcription factors encompasses five transcription factors, which are RelA (p65), RelB, cRel, NF-κB1 (p50), and NF-κB2 (p52). Transcription factors shuttle between the signal processing and execution layers to bind to the target gene promoter. The execution layer is the last layer of NF-κB that represents the genes under the control of NF-κB transcription factors [12].
In the absence of stimuli, NF-κB transcription factors’ nuclear localization signals are sequestered in the cytoplasm by binding to the inhibitors: IκBα, IκBβ or IκBε. The IκB family is a group of NF-κB inhibitors composed of five different members that inhibit the NF-κB pathway: IκBα, IκBβ, IκBε, IκBζ, and BCL3. Upon activation of the NF-κB pathway, one of the NF-κB inducible genes IκBα will be expressed to form a negative feedback response and bind to NF-κB transcription factors to ensure that the NF-κB pathway will remain firmly regulated after producing the target response [13]. Moreover, upon stimulation of NF-κB, IκBε is expressed but in delayed onset in comparison to IκBα, forming a second negative feedback loop [14].
A better understanding of this mechanism may provide a deeper understanding of how the NF-κB pathway consistently produces the intended response and ensures that the activation will cease to maintain balance and homeostasis after achieving the required response.
BCL3 is another member of the NF-κB inhibitor protein family. Under basal conditions, BCL3 is not highly expressed, but upon activation of NF-κB, it binds to the Bcl-3 gene and induces its expression [15]. Moreover, BCL3 can either suppress or promote NF-κB transcriptional activity based on the stimulus and cell type [16].
A major class of NF-κB are the regulators, which are the enzymes or protein subunits that modulate NF-κB pathway activity in response to various triggers. A20, encoded by tumor necrosis factor alpha induced protein 3 (TNFAIP3), is an NF-κB negative regulator. A20 could act as an oncoprotein or a tumor suppressor based on cancer type [17]. TNFAIP3 has a conserved site for RelB-p52, suggesting that it will be expressed after activation of the NF-κB pathway to form a negative feedback response [18]. Also, cylindromatosis (CYLD) is a deubiquitinase enzyme that regulates the canonical NF-κB pathway by different mechanisms [19] (Fig. 1).
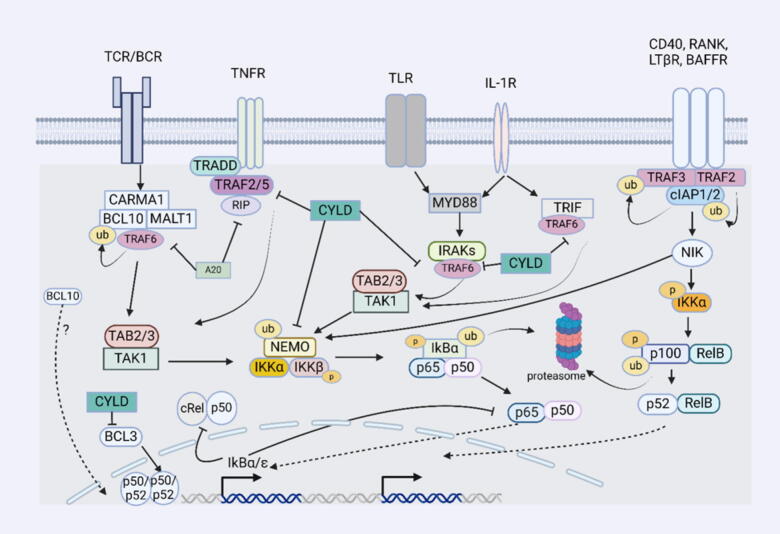
Fig. 1.
Canonical and non-canonical NF-κB and the different mechanisms of regulations. The input layer of canonical NF-κB comprises different types of receptors, including BCR, TCR, TLR, IL-1R, and TNFR. These are activated in response to different triggers, resulting in canonical NF-κB pathway activation. Under basal conditions, IκBα masks the nuclear localization signal of p65 and p50, hence hindering NF-κB transcription factors from translocating to the nucleus. Stimulation of the receptors in the input layer transmits the signal to the processing layer through recruiting various adapters, resulting in β complex activation in the canonical NF-κB pathway. Subsequently, IKK phosphorylates IκBα to release p65-p50 and/or c-REL-p50 heterodimer that translocates to the nucleus and binds to the target gene promotor. Conversely, the non-canonical pathway is regulated by the TRAF-cIAP complex. Under basal conditions, NIK is instantly bound to TRAF3 and recruited to the E3 ubiquitin ligase complex consisting of TRAF2 and cIAP1/2. Thus, NIK will be ubiquitinated and degraded. Upon stimulation of the non-canonical NF-κB pathway by triggering lymphotoxin β, CD40, RANK, or BAFFR, NIK phosphorylates and activates IKKα that in turn phosphorylates p100, hence mediates its proteasomal processing. p52-RelB will translocate to the nucleus and stimulate gene transcription, producing the output response. NF-κB activity is regulated by several enzymes and proteins that keep NF-κB balanced. Mainly, IκBα/ε is induced in response to NF-κB activation to produce a negative feedback loop. In contrast, CYLD detaches the K63-ubiquitin chain and deactivates different subunits, including TRAF2, TRAF6, NEMO, and BCL3, therefore inactivating IKK kinase complex and inhibiting canonical NF-κB pathway activation. A20 regulates NF-κB through physically binding to TRAF2 to impede NF-κB activation, interfere with TRAF6 ubiquitination, and target RIP-1 for degradation. Also, A20 competes for the binding of ubiquitin chain on NEMO added by LUBAC, hence impeding IKKβ recognition by TAK-1deubiquitinating TRAF6 and RIP1 as well as interfering with NEMO ubiquitination. Moreover, BCL3 regulates NF-κB and affects its transcriptional activity through binding to p50 and p52 homodimers. This figure was generated using Biorender. BAFFR, B-cell activating factor receptor; BCR, B-cell receptor; cIAP1/2, cellular inhibitor of apoptosis 1 and 2; CYLD, cylindromatosis; IL-1R, interleukin-1 receptor; IKK, inhibitor of nuclear factor-κB kinase; IκB, inhibitor of nuclear factor-κB; LTβR, lymphotoxin β receptor; MyD88, myeloid differentiation primary response gene 88; NEMO, NF-κB essential modulator; TRIF, TIR domain-containing adaptor inducing interferon-beta; RIP1, receptor-interacting protein 1; NIK, NF-κB-inducing kinase; TAK1, TGF-β-activated protein kinase 1; TCR, T-cell receptor; TLR, toll-like receptor; TNFR, Tumor necrosis factor-α receptor; TRADD, TNF-R-associated death domain; TRAF2/5/6, TNF-R-associated factor 2, 5 and 6.
This diversity in the NF-κB regulators explains that there is always a compensatory mechanism to keep NF-κB under control, so if any of the regulators got affected by genetic alterations or epigenetic modification, others will compensate [20]. This point has been confirmed in a knock-in mouse model where they replaced IκBα with the IκBβ gene. The mouse model exhibits no visible abnormalities in contrast to the IκBα-knockout model, which died very early due to multiple deficiencies [21]. Notably, Failure of regulatory mechanisms produces immense dysregulation of the NF-κB pathway, leading to autoimmune diseases or cancer [22].
Cells are subjected to environmental triggers at diverse intensities, which can lead to the constitutive activation of NF-κB. Some of these triggers such as smoking or ultraviolet light exposure could introduce DNA alterations and histone modification. But mostly these lesions or mutations could be repaired after removal of the trigger to restore balanced gene expression and keep critical pathways including NF-κB regulated [23], [24], [25].
NF-κB regulators can work in harmony to restrain the balance and impede the formation of uncontrollable responses by forming negative feedback responses [13]. For instance, IκBα can translocate to the nucleus by associating with proteins that possess nuclear localization signal (NLS) to enhance the dissociation of NF-κB from DNA. Still, these proteins are yet to be identified [26]. These findings suggest that not all the mechanisms of NF-κB regulation are still known.
One more level of regulation was observed in NF-κB, where some NF-κB subunits could play a dual role as pro- and anti-inflammatory, such as p50 and IKKβ [27]. Therefore, the same subunit known to activate NF-κB early during infection plays a regulatory role in the last phase of inflammation. As a result, mutations that affect these subunits could lead to aberrant NF-κB pathways. Based on these points, it can be hypothesized that the mutation frequency that may cause uncontrolled NF-κB pathway varies between individuals based on the exhibited level of NF-κB regulation.
Conversely, constant exposure to the trigger for a long time causes the accumulation of mutations that eventually lead to irreversible genetic mutations and irritable NF-κB response [28]. This dialogue occurs in cases where the repair and compensatory mechanisms are deficient, the damage will be major to cause dysregulated NF-κB pathway and producing autoimmune diseases or cancer. During cell transformation, aberrant NF-κB shifts the response from immunogenic to sustaining cancer hallmarks. This can explain why patients with very early cancer exhibit high levels of various inflammatory mediators, suggesting a high immunogenic response to cell dysplasia or necrosis. Despite this, most inflammatory mediators stabilize and return to normal levels after cancer progression [29].
Such response alteration occurs mainly because of two points. Notably, genetic alterations in NF-κB-target genes lead to constitutively active NF-κB pathway because of the positive feedback loop. Also, it is frequently accompanied by the activation of other oncogenic pathways such as MAPK or PI3K/Akt [30]. Several cancers have been linked to NF-κB dysregulation. However, the most prominent role has been reported in mucosa-associated lymphoid tissue (MALT) lymphoma. Gastric MALT lymphoma can be classified into two subtypes: translocation-positive and translocation-negative. Translocation-negative MALT lymphoma mainly activates immune response by upregulating co-stimulatory molecules. However, translocation of B-cell lymphoma/leukemia 10 (BCL10) to IgH locus is associated with upregulation and nuclear translocation of BCL10, inducing constant activation of the NF-κB pathway and sustaining different cancer hallmarks [30].
The dysregulation of NF-κB can happen at the three layers of the pathway. The input layer can be affected by upregulation or mutation of NF-κB activating receptor or ligand upregulation and, in some cases, both. The second type of dysregulation affects the signal processing layer; this layer represents the algorithmic and analytical part of this pathway. As a final layer, the execution layer is responsible for producing genes engaged in the NF-κB pathway and producing positive feedback loops, as well as activating other pathways [6], [30]. The impact of dysregulated NF-κB genes in different health conditions is summarized in Table 1.
Table 1.
Dysregulated genes related to different layers of NF-κB are implied in different disorders.
| NF-κB layer | Dysregulated gene | Condition | The Effect of dysregulation | References |
|---|---|---|---|---|
| Input layer | c-MET | – Head and neck squamous cell carcinoma – Non-small cell lung cancer – Papillary renal cell carcinoma |
Growth and invasion | [31], [32], [33] |
| BAFF | – Rheumatoid arthritis – Systamic lupus erythematosus |
B-lymphocyte proliferation and auto-antibodies formation | [34], [35] | |
| TLR9 | – Hepatocellular carcinoma | Proliferation | [36] | |
| Processing layer | RELA | Psoriasis | Auto-immune reaction | [37] |
| BCL10 | – MALT lymphoma – pancreatic cancer – Extranodal NK-cell lymphoma |
[30], [38], [39] | ||
| IKKβ | Pneumonia | Defected immune response | [40] | |
| IKKβ | Sepsis | Enhanced pro-inflammatory cytokines | [41] | |
| IκBα | Multiple sclerosis | Defected negative feedback response and enhanced inflammation | [42] | |
| MyD88 | colorectal cancer | Signal amplification and driving proliferation and metastasis | [43] | |
| Execution layer | TNF-α | Rheumatoid arthritis | Produce immunogenic costimulatory signals in lymphocytes | [44] |
| MCP-1 | Renal cell carcinoma | Self-sufficiency growth signals | [45] | |
| BCL2 | Gastric MALT lymphoma | Survival | [30] | |
| TLR2 and TLR4 | Various cancer types and Tumor microenvironment | Anti-apoptotic signals, immune escape and chemotherapy resistance | [46], [47], [48], [49] | |
| CCR2 and TLR6 | Gastric MALT lymphoma | Positive feedback response, activate oncogenic pathway leading to survival and proliferation |
Various methods of inhibiting NF-κB are used to treat different cancers and immune diseases. For instance, corticosteroids alleviate the symptoms of almost all immune diseases by targeting the NF-κB pathway along with other immune-related pathways. Targeting NF-κB in immune diseases develops tolerance while blocking the immune triggers [29]. Besides, in cancer, NF-κB blockers inhibit the transcription of various oncogenes that will sustain cancer hallmarks. For this purpose, NF-κB inhibitors have been developed [50]. NF-κB inhibitors can be classified into different classes based on molecular target. BAY 11 is an irreversible selective NF-κB inhibitor that showed anti-cancer effect in several types of cancer. BAY 11 selectively interferes with TNF-α-induced NF-κB and stabilizes IκBα, hence BAY 11 suppresses proliferation and causes cell apoptosis [51]. In addition, multiple proteasome inhibitors are employed to target the NF-κB pathway including MG-115 and MG-132, however, proteasome inhibitors are non-specific to NF-κB [52]. Another class of NF-κB inhibitors is IKK complex inhibitors such as IKK-16 [53]. Each of these inhibitors targets a specific part of this pathway, however, the NF-κB pathway is composed of intricate layers and factors that would be very challenging to control and restore the balance. Each of the conditions caused by irritable NF-κB response could employ different mediators and factors. Thereby, identifying the molecular dynamic of NF-κB-linked diseases is a major milestone in controlling NF-κB and restoring a balanced pathway.
Role of NF-κB in different cancer hallmarks
Dysregulation of the NF-κB pathway may enhance cancer initiation, promotion, and spread [8]. NF-κB performs its tumorigenic role by sustaining different cancer hallmarks specifically, invasion and metastasis [54]. NF-κB induces the expression of SNAIL, TWIST1, SLUG, matrix metalloproteinase-9 (MMP-9), and C-X-C motif chemokine ligand 12 (CXCL12) which enhance cancer migration and result in epithelial-mesenchymal transition (EMT) [55]. Moreover, inhibiting NF-κB can effectively reverse EMT [56], confirming the critical role of NF-κB in metastasis.
Furthermore, stimulating angiogenesis is another hallmark of cancer. NF-κB as a part of an inflammation-induced pathway, could induce angiogenesis directly and indirectly. NF-κB directly induces the expression of hypoxia-inducible factor-1α (HIF-1α) and vascular endothelial growth factor (VEGF) during hypoxia. In response to platelet factor-4 (PF4) treatment, NF-κB binds to E-selectin to induce its expression. In contrast, the indirect effect is through NF-κB-activated macrophages and fibroblasts that in turn produce VEGF in the tumor microenvironment [57]. Besides, the indirect mechanism includes several pro-inflammatory cytokines such as IL-1 and TNF-α secreted in response to NF-κB activation which can trigger the expression of adhesion molecules such as intracellular adhesion molecule-1/2 (ICAM-1/2), E-selectin and VCAM-1 [58]. Besides, NF-κB has a critical role in evading apoptosis, where cancer cells tend to highly activate the NF-κB pathway, resulting in high expression of anti-apoptotic genes such as A1/Bfl-1 and BCL-xL and cIAP2 [59]. Furthermore, BCL-2 family members such as BCL-xL and A1/Bfl-1 inhibit apoptosis by suppressing mitochondrial depolarization and cytochrome c release. However, cIAP2 acts as an anti-apoptotic protein by mediating the ubiquitin degradation of receptor-interacting protein kinase 1 (RIPK1) involved in the TNFR signaling pathway and by directly binding to caspase 3, 7, and 9 and inhibits their activity [60]. NF-κB provides cancer cells with sustained proliferation ability. In detail, NF-κB regulates the collection of genes that control cell cycle and proliferation, such as cyclin D, and cyclin A. Numerous NF-κB members such as c-Rel, p65, and p50 interact with the cyclin E/cyclin-dependent kinase (CDK)2 and regulate the activity of this complex [61]. Moreover, NF-κB can positively regulate mouse double minute 2 (MDM2), thus inducing p53 instability and cell growth [62]. In addition, various pro-inflammatory cytokines induced by NF-κB such as IL-6, TNFα, and IL-1β, encourage cancer cell proliferation. Furthermore, constitutive activation of NF-κB maintains high levels of granulocyte colony-stimulating factor (G-CSF) which positively regulates growth and survival [6].
One of the crucial steps of cancer development is immune escape. Transformed cells can be detected by the immune system and eliminated. However, cancer cells develop various strategies to avoid immune response. The role of NF-κB in immune surveillance is major, thus multiple NF-κB transcription factors are included in immune cells regulation. Specifically, p65 maintains mature peripheral regulatory T cells (Tregs), whereas c-Rel contributes to the development of thymic Tregs. Hence, interference with c-Rel either by deletion or inhibition, diminishes Treg immunosuppressive effect and enhances the response to immune checkpoint blockade [63], hence the filtration of Tregs in the tumor site is linked to poor prognosis in many cancers such as kidney cancer [64]. Moreover, the p50 subunit of NF-κB contributes to shifting macrophage polarization toward M2 macrophage, thereby enhancing immunosuppression and tumor promotion [65]. Therefore, as a result, NF-κB could attribute to all stages of cancer by mediating cancer cells to proliferate and invade without getting stopped by the host immune system. In summary, several cancers have been linked to NF-κB dysregulation. However, the most prominent role has been reported in MALT lymphoma [30].
The role of NF-κB subunits in different cancer hallmarks is summarized in Table 2.
Table 2.
NF-κB induces the expression of various genes that are implicated in cancer hallmarks including invasion, migration, and proliferation.
| NF-κB subunit/s | NF-κB-responsive genes | Cancer hallmarks | References |
|---|---|---|---|
| p65 | SNAIL | Migration and invasion | [66] |
| Canonical NF-κB pathway specifically IKKβ and p65 | TWIST1 | Invasion and migration | [67] |
| p65 | MMP-9 | Invasion and metastasis | [68] |
| Non-canonical NF-κB pathway subunits p52 and RELB | CXCL12 | Migration and angiogenesis | [69] |
| P65, p50 and NF-κB2 | VCAM-1 | Migration and angiogenesis | [70] |
| All NF-κB subunits, especially p50 | HIF-1α | Angiogenesis | [71] |
| c-REL and p65 | GM-CSF | Growth and migration | [72] |
| Canonical NF-κB pathway subunits p65 and p50 | VEGF | Angiogenesis, migration, and invasion | [73] |
| Canonical NF-κB pathway subunits p65 and p50 | Indirect expression of ICAM1/2, E-selectin and VCAM1 through TNF-α and IL-1 | Angiogenesis | [58] |
| p50 | M-CSF | Immunosuppression | [74] |
| c-REL | IL-2 | Immunosuppression | [6] |
| Canonical NF-κB pathway | Foxp3 | Immunosuppression | [75] |
| p65 and -REL | ICAM-1 | Proliferation and migration | [76] |
| p65 and p50 | G-CSF | Proliferation | [77] |
| p65 and p50 | Cyclin D | Proliferation | [61] |
| p65 | Cyclin A | Proliferation | [61] |
| p65 | MDM2 | Anti-apoptotic | [78] |
| p65 and p50 | BCL-2 | Anti-apoptotic | [59] |
A highly activated NF-κB is implicated in renal inflammation in response to various triggers. NF-κB is initially activated to resolve the trigger and produce an immune response. However, the persistence of the cause or a failure in NF-kB regulatory mechanisms produces the activation of the immune system for a prolonged period, even after the resolution of the trigger, which leads to chronic inflammation. The constant activation of NF-κB incites a positive feedback response that will outweigh the inhibition regulatory mechanism imposed by the negative feedback loop. The resultant shift from chronic inflammation to renal cell carcinoma is usually sustained by the anti-apoptotic effect of the NF-κB pathway, which will be further amplified by cross-talking with other inflammatory pathways, including Janus kinase/signal transducer and activator of transcription (JAK/STAT), to enhance cell proliferation [79]. Exposure to inflammatory conditions for a long time usually triggers genetic alteration by different mechanisms, affecting the regulation of key pathways, mainly NF-kB, and shifting the response from immune to proliferation, survival, and metastasis [80].
Methodology
The current study employs data from different databases including COSMIC, TCGA, GEO, and GTex. The mutation frequency was obtained from the COSMIC database based on multiple projects like TCGA, Pan-Cancer Analysis of Whole Genomes project (PCAWG), and AACR Project GENIE. The box plot figures were generated using Tumor, Normal, and Metastatic (TNM) plot tools that utilize data from different databases such as TARGET, GEO, and TCGA. The data of the TNM plot involves 277 Normal, 556 tumors, and 58 metastatic tissue samples obtained from the kidney.
NF-κB in kidney cancer
Kidney cancer is one of the least studied cancers due to the complexity of its genetic composition and the elusive molecular mechanisms. RCC, which is the major type of kidney cancer, can be subclassified into three major subtypes based on the kidney region where the cancer originates. The most common subtype of RCC is clear cell renal cell carcinoma (ccRCC), which accounts for more than 70 % of RCC. Therefore, it’s the most studied subtype. Other subtypes represent a lower frequency of RCC, including papillary renal cell carcinoma (PRCC) and chromophobe renal cell carcinoma (chRCC) [81]. NF-κB role in RCC development and progression has been investigated in diverse studies [82], [83], [84].
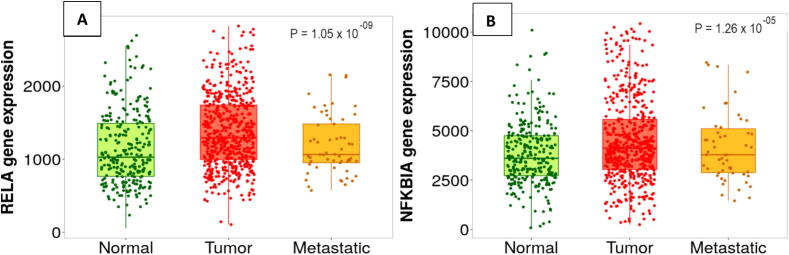
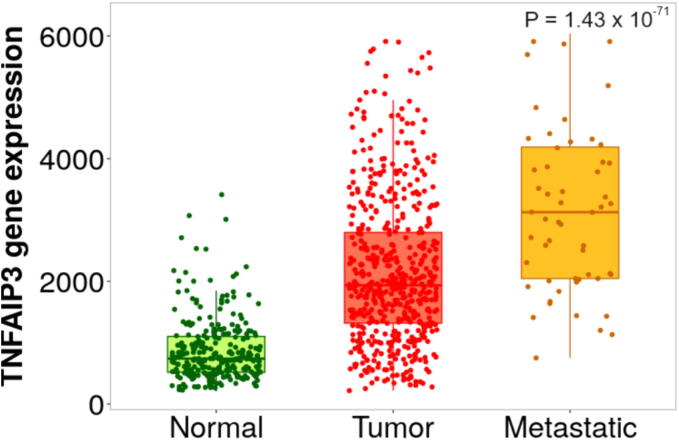
When the cells are subjected to growth-stimulating triggers, guard mechanisms will be activated to induce cell cycle arrest or apoptosis to protect the cells from mutations and transformation. However, in RCC, multiple NF-κB-related genes are genetically altered; hence, the NF-κB-mediated survival mechanisms are utilized by cancer cells to promote tumor growth [85]. Based on data from TNM plot (tnmplot.com [86] cohort study that involved 277 Normal, 556 tumors, and 58 metastatic kidney cancer samples, RELA is highly expressed in tumor samples in contrast to healthy control and metastatic kidney cancer patients (p = 1.05 × 10−9) (Fig. 2A) [87]. No significant difference was detected in the expression of RELB and NF-κB2 between normal, tumor, and metastatic renal carcinoma, suggesting the substantial role of the canonical NF-κB pathway in driving tumor development in renal carcinoma. Moreover, the NFKBIA gene that encodes for IκBα was found to be highly expressed in tumor samples of kidney cancer compared with healthy control and metastatic kidney cancer with a p-value equal to 1.26 × 10−5 in TNM plot cohort [15] (Fig. 2B). The high expression of RELA in tumor samples, along with the elevated levels of IκBα as negative feedback, suggests the activation of the canonical NF-κB pathway upon exposure to harmful stimuli, which can progress to cancer with constant exposure to the stimuli and the lack of regulation possessed by the decreasing of NFKBIA seen in metastatic kidney cancer (Fig. 2B). This evidence proposes the prominent role of the canonical NF-κB in cancer initiation and the requirement of high levels of NF-κB-responsive genes in the initial stages of cancer. Importantly, the transformation is accompanied by other mutations that activate cancer-related pathways to shift the response from immune response to proliferation and survival [88]. A similar result was observed in the phosphorylated status of p65 and IκBα in RCC tissues obtained from fresh specimens. In detail, specimens were obtained from three patients' tumor and normal corresponding tissues, and the expression of p-p65 and p-IκBα were assessed. Notably, the p-p65 and p-IκBα levels were significantly increased in RCC tissues compared with normal tissues, confirming the constant activation of NF-κB in RCC [89]. Accordingly, the dysregulation of NF-κB of RCC occurs at the transcriptional and post-translational levels. However, more studies must be done to detect the downstream target of NF-κB, which will provide insight into the molecular mechanism and dynamic NF-κB pathway in RCC. Additionally, A cohort study in the TNM plot database with 277 Normal, 556 tumors, and 58 metastatic kidney cancer participants found that TNFAIP3 was most highly expressed among metastatic renal cancer samples in comparison to tumor and control samples, with a p = 1.43 × 10−71, suggesting that high expression of A20 in correlated to metastasis in RCC (Fig. 3). Moreover, consistent with the previous study, the TNFAIP3 was found to be over-expressed in 3.93 % (21/534) in ccRCC based on a cohort study conducted by the COSMIC database, suggesting a tumor-promoting and anti-apoptotic role of A20 in kidney cancer [90]. However, the mechanism of the A20 tumor-promoting effect has not yet been studied in kidney cancer. Molecular analysis of A20 may reveal how the genetic and epigenetic alterations of TNFAIP3 promote kidney cancer progression.
Fig. 2.
Canonical NF-κB-related genes are elevated in kidney tumor tissues. A TNM box plot of RELA gene expression in kidney normal, tumor, and metastatic samples. B TNM box plot of NFKBIA gene expression in kidney 277 normal, 556 tumors, and 58 metastatic samples (N = 891).
Fig. 3.
TNFAIP3 expression is correlated positively with the tumor stage. TNM box plot of TNFIAP3 gene in kidney 277 normal, 556 tumors, and 58 metastatic samples (N = 891).
In addition, caspase recruitment domain family member10 (CARD10)-mediated NF-κB activation in response to epidermal growth factor receptor (EGFR) has been implied in RCC. Activating CARD10 using EGF effectively activates NF-κB in renal tubular epithelial cells; hence, silencing CARD10 significantly reduces the migration and proliferation of RCC cell lines. This evidence confirms the role of the CARD10-mediated NF-κB pathway in enhancing the progression of RCC. Consistent with the previous study, CARD10 was found to be over-expressed in 7 % among 534 ccRCC patients based on a COSMIC cohort study, which confirms its role in augmenting NF-κB activity and providing a shift toward survival and cancer progression in response to growth factors such as EGF [91]. Moreover, HIF-1α, which is highly expressed in ccRCC, triggers various cells in the tumor microenvironment to release growth factors, including EGF, to promote intravasation and migration. Accordingly, CARD10-mediated NF-κB activation by EGF tends to be activated in the later stage of cancer in response to other oncogenic pathways [92].
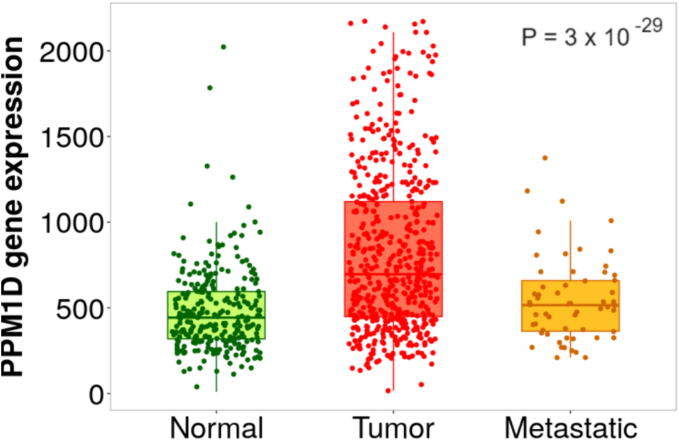
Also, another NF-κB inducible gene that has a conserved site for p65. PPM1D encodes Wild-type P53-induced phosphatase 1 (Wip1), which acts as a serine/threonine phosphatase [93]. Many papers have demonstrated Wip1′s role as an oncoprotein. Wip1 was correlated to poor prognosis, higher TNM stage, and less differentiated tumor in kidney cancer. In a cohort study done in 277 Normal, 556 tumors, and 58 metastatic kidney cancer samples in the TNM plot database, tumor samples have the highest expression of PPM1D in comparison to healthy control and metastatic renal cancer with p = 3 × 10−29, supporting the evidence that PPM1D has a crucial role in cancer development rather than invasion and progression (Fig. 4). Also, referring to the COSMIC database, PPM1D is overexpressed in 2 % of 534 ccRCC samples [94]. Upregulation of PPM1D suggests the shift toward survival in response to triggers that can activate DNA damage response, such as radiation or chemotherapy instead of apoptosis or cell cycle arrest. Accordingly, this is consistent with the growing evidence that PPM1D plays a role in resistance to cancer treatments [95].
Fig. 4.
PPM1D NF-κB-responsive gene is highly expressed in kidney tumor tissues. TNM box plot of PPM1D gene in 277 normal, 556 tumors, and 58 metastatic cancer kidney samples (N = 891).
Cancer cells’ genetic profiles vary; as a result, each of the subtypes exhibits a unique genetic mutation landscape. However, all RCC subtypes have a common gene mutation that drives carcinogenesis, mainly von Hippel-Lindau (VHL). VHL is a tumor suppressor gene that acts as an E3 ubiquitin ligase and induces HIF-1α degradation [96]. Moreover, different renal cancer subtypes exhibit different rates of VHL missense mutations that frequently lead to VHL inactivation; however, the highest rate of VHL mutations was found in ccRCC.
Role of NF-κB in ccRCC
The most frequently mutated gene in ccRCC, based on a cohort study done by the COSMIC database, is VHL. Notably, 52 % of the VHL mutations were reported among 6232 ccRCC cases. Loss of the VHL gene in sporadic and familial ccRCC is correlated to the elevated NF-κB-signature genes, suggesting the inverse crosstalk between VHL and NF-κB in ccRCC. Moreover, VHL mutations or loss could drive NF-κB expression by accumulating HIF-α and transforming growth factor-alpha (TGF-α), resulting in PI3K/Akt/NF-κB pathway activation [97]. Referring to the COSMIC database, a cohort study that included 534 ccRCC patients found that 3.2 % (17/534) patients were overexpressing HIF1A, and 4.9 % (26/534) patients exhibited overexpression in the TGFA gene, suggesting the prominent role of this pathway in NF-κB activation in ccRCC. In addition, the increased expression of HIF1A allows the metabolic shift that occurs in ccRCC toward glycolysis and promotes angiogenesis and migration in hypoxic stressful conditions [92], whereas TGF-α overexpression sustains cell proliferation [98]. Similarly, immunofluorescence staining of ccRCC cells shows nuclear and cytoplasmic expression of HIF-1α, which is correlated to nuclear p50 expression. In contrast, restoring the ZIP1 tumor suppressor gene, significantly abrogated HIF-1α levels and p50 nuclear localization [99]. Thus, the pivotal role of NF-κB in regulating and interacting with a vast number of oncogenic pathways and proteins provides a continuous loop of sustaining RCC progression and migration.
Another previous study has reported that VHL presence does not affect the expression or the phosphorylation status of p65 and IκBα. Interestingly, tested VHL-wildtype or VHL-deficient ccRCC cell lines exhibited an activated NF-κB pathway, regardless of VHL status [89]. This evidence provides proof of concept that the NF-κB pathway is activated in RCC through interplaying with different pathways mainly, PI3K/Akt pathway. subsequently, not a single genetic alteration would drive NF-κB dysregulation.
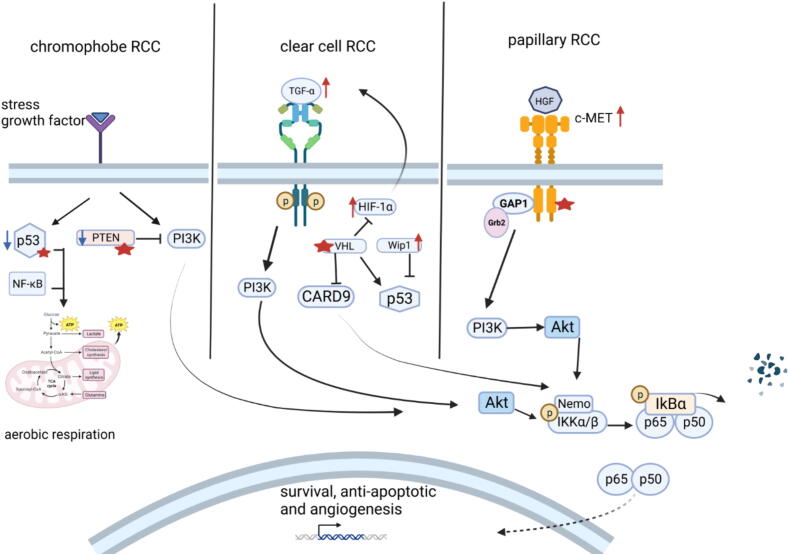
ccRCC exhibits a total of 7 % mutation in TP53 in a total of 2316 samples, which will augment PI3K/NF-κB by the absence of one more tumor suppressor gene, allowing this pathway to be constitutively active. Moreover, this, in turn, will suppress the activity of p53 in ccRCC cells that harbor wild-type p53, rendering it inactive. As a result, the NF-κB pathway is not suppressed by the main regulators VHL and p53 (Fig. 5) [100]. Moreover, NF-κB activity has been assessed in a study that included 42 ccRCC and 3 PRCC samples. Two out of the three PRCC samples had activated the NF-κB pathway. Whereas, 5/7 (71 %) grade three ccRCC samples had activated NF-κB, which is the highest percentage compared to grades 1 and 2. Moreover, all three metastatic RCC cases exhibit highly activated NF-κB. 15/45 RCC samples (33 %) exhibit over 200 % increased NF-κB activity [82]. This evidence shows that despite the employing of different oncogenic pathways in the two different RCC subtypes, NF-κB is highly activated in both subtypes. NF-κB might be activated in different stages and by unique genetic alterations in each RCC subtype, however, its constant activation is a critical step in cancer development and progression.
Fig. 5.
The activation of the NF-κB pathway by various upstream receptors and triggers in different subtypes of RCC. NF-κB is activated in RCC different subtypes through different dysregulations that occur upstream, mainly in the PI3K/Akt pathway. In chromophobe RCC, mutations, and deletion of p53 and PTEN allow the overactivation of PI3K/Akt by growth factors and hormones that will, in turn, activate the NF-κB pathway and support aerobic oxidation mechanisms. Chromophobe RCC cancer cells exhibit increased expression of several Kreb’s cycle enzymes including citrate synthase and acotinase2. Also, NF-κB could enhance p53 suppression by inducing the expression of MDM2, providing a positive feedback response and shifting the response to survival and proliferation. In clear cell RCC, VHL mutation causes the accumulation and activation of HIF-1α and NF-κB, respectively. HIF-1α accumulation induces the expression of TGF-α that binds to the epidermal growth factor receptor and activates NF-κB in an Akt-mediated manner. Moreover, NF-κB provides a positive feedback loop by inducing the expression of multiple genes, including HIF-1α. p53 activity is suppressed in clear cell RCC through VHL loss. Additionally, Wip1 is another inhibitor of p53 that will suppress its function by dephosphorylating ser15 which will further potentiate NF-κB pathway. Besides, TGF-α activates PI3K providing an indirect NF-κB pathway activation. Papillary RCC is characterized by mutations in the c-MET receptor that will constantly activate its tyrosine kinase domain. Upon the dimerization of the receptor and phosphorylation and activation of the tyrosine kinase domain, GAP1 and Grb2 are recruited and bind to the multifactional docking site of c-MET, resulting in PI3K/Akt activation. Consequently, Akt phosphorylates IKK and activates the canonical NF-κB pathway. Accordingly, the p65-p50 heterodimer binds to multiple genes, which sustains different cancer hallmarks such as survival and migration. This figure was designed using Biorender. CARD9, caspase recruitment domain family member 9; c-MET, mesenchymal-epithelial transition factor; GAP1, general amino acid permease; Grb2, growth factor receptor-bound protein 2; HGF, hepatocyte growth factor, HIF-1α, hypoxia-inducible factor 1-α; IKK, inhibitor of nuclear factor-κB kinase; IκB, inhibitor of nuclear factor-κB; NEMO, NF-κB essential modulator; NF-κB, nuclear factor kappa light chain enhancer of activated B cells; PI3K, phosphatidylinositol 3-kinase; PTEN, phosphatase and tensin homolog; RCC; renal cell carcinoma TGF-α, Tissue growth factor-α; VHL, von Hippel_Lindau; Wip1, wild-type p53-induced phosphatase 1Different subsets of RCC exploit intricate pathways to sustain various cancer hallmarks. In fact, different genetic alteration patterns are observed within the same RCC subtype. Curiously, a biopsy from clear cell carcinoma patients revealed intratumor heterogeneity, where different regions in the same tumor mass exhibit unique mutational landscapes, and only 34 % of the mutations are shared between these regions [124]. This evidence suggests that RCC comprises a heterogeneous group of cancer cells with distinct molecular profiles. However, overall, each subtype of RCC is characterized by some biomarkers or gene signature. While VHL mutation or deletion is detected in more than 70 % of ccRCC [125], another hereditary condition, Birt–Hogg–Dubé (BHD) syndrome, is associated with chRCC. Around 30 % of BHD patients develop renal tumors, and 34 % of the developed BHD-related renal tumors are chRCC [126]. BHD syndrome is associated with FLCN gene mutation, which encodes folliculin. In detail, folliculin executes its tumor-suppressing action by regulating multiple energy metabolic pathways, including mTOR and MAPK. Loss of function of FLCN allows AMPK and mTOR pathways to be constitutively active, which aligns with the augmented mitochondrial biogenesis observed in chRCC [127]. Lastly, heredity mutation in the MET gene results in Hereditary papillary renal carcinoma (HPRC), which significantly increases the risk of developing type 1 PRCC (which accounts for >80 % of PRCC) [128]. In light of the cohort and translational studies of different subtypes of RCC, each subtype employs specific NF-κB genes based on the cancer hallmarks required. Moreover, several factors contribute to somatic mutations, including immune surveillance, cell-specific mutations, and environmental factors that patients may be exposed to, including ultraviolet (UV) light [129]. Based on the above observations, ccRCC is associated with increased expression and dysregulation of several NF-κB genes, which are utilized to drive ccRCC development and progression. In contrast, chRCC and PRCC mainly drive NF-κB activation indirectly, hence specific genes of NF-κB pathway will be dysregulated.
The expression of p50 NF-κB subunit has been linked to different apoptosis and angiogenesis markers in ccRCC. In detail, 40 ccRCC samples were measured to express the p50 subunit of NF-κB, VEGF, EGFR, p53, and BCL-2. A significant correlation was detected between NF-κB level and EGFR (p = 0.004), VEGF (p = 0.001), BCL-2 (p = 0.01), and p53 (p-value = 0.p037) [83]. Consistent with this finding, 786–0 RCC cell lines with p50 knockout showed decreased expression of crucial angiogenic markers, primarily IL-6 [101]. Accordingly, p50 could act as a novel therapeutic target and prognostic marker in ccRCC. However, more research is needed to study the molecular complexes and mechanisms that employ the p50 subunit and its distinct role in ccRCC pathogenesis.
Y-box binding protein 1 (YBX1) is a transcription factor that exhibits oncogenic properties in several types of cancer. In ccRCC, YBX1 is associated with a higher stage and grade by activating the NF-κB pathway In detail, YBX1 interacts with RCC-related proteins including secreted phosphoprotein 1 (SPP1) and Ras-GTPase activating protein SH3 domain binding proteins 1 (G3BP1), which will activate NF-κB and enhance the phosphorylation and nuclear translocation of p65 in RCC cell lines. Additionally, an indirect interaction between NF-κB and glycogen synthase 1 (GYS1) has been observed in ccRCC. In ccRCC tissues, GYS1 levels are significantly increased, which is associated with tumor growth The accumulation of GYS1 indirectly activates the canonical NF-κB pathway and enhances glycogen levels. In contrast, silencing GYS1 causes metabolic perturbations and synthetic lethality of ccRCC cells. Accordingly, these findings indicate that NF-κB plays an important role in enhancing proliferation and migration of RCC, by interacting directly and indirectly with different oncoproteins. Moreover, consistent with previous findings NF-κB plays its tumor-promoting role by shifting the metabolic machinery in cancer cells toward glycolysis.
In addition, a meta-analysis done using data of ccRCC from gene expression omnibus (GEO) and ArrayExpress revealed a prominent alteration of NF-κB-responsive genes and regulators. Multiple genes were upregulated, including IKBKB (IKKβ), MMP9, SOD2, and PSMB9. Hence, these genes mediate the survival and pro-inflammatory role of the NF-κB pathway. The expression of these genes was correlated to poor prognosis and reduced survival [85]. As p50 is a major transcription factor of canonical NF-κB, this evidence highlights the critical role of this pathway in driving ccRCC aggressiveness.
An in vivo model of ccRCC was established by exposing the mice to a carcinogen known as ferric nitrilotriacetate (FeNTA) to simulate the histological features of ccRCC, and the levels of p65, IκBα, and EGFR were assessed. Notably, tumor tissue derived from the experimental mouse model exhibited elevated levels of p65 and EGFR, whereas IκBα levels were decreased. Conversely, after one month from the exposure to FeNTA, IkBα levels increased, while EGFR and p65 expression remained elevated [102]. This study is consistent with the TNM plot data, which shows that IκBα is increased as negative feedback when patients are exposed to a trigger and an immune response is activated. Moreover, the persistence of the trigger shifts the response from immune to different cancer hallmarks based on the dysregulated gene. In this case, NF-κB activation induces the expression of proliferation-promoting genes such as cyclin D, shifting the response to cell growth and proliferation. Subsequently, the increased IκBα level because of the negative feedback loop can no longer counteract or suppress the consistently activated NF-κB pathway and restore the balance, especially since other oncogenic pathways will also be activated [103].
Cezanne (encoded by Otud7b) is another regulator of the canonical NF-κB pathway in the kidney, specifically in glomerular endothelial cells [104]. Notably, cezanne is under-expressed in 3 % of 534 cases of ccRCC [105], suggesting an uncontrolled constantly activated canonical NF-κB pathway in ccRCC and the shift toward cell growth.
Bcl-3 is a known proto-oncogene that enhances tumor progression [106]. In the context of ccRCC, Bcl-3 was found to be overexpressed by 3 % across 534 samples in a cohort study established by the COSMIC database [105]. In response to DNA damage, BCL3 was found to be upregulated, induce the expression of MDM2, and inhibit p53-mediated apoptosis; as a result, oncogenic pathways such as NF-κB will shift the cellular response toward survival, resulting in the propagation of cells with mutation and generation of transformed cells [107]. Another study revealed the presence of BCL3 in the nucleus of metastatic RCC. Particularly, co-immunoprecipitation demonstrated that nuclear extracts from metastatic lung tissues contain p50/BCL3 complexes. These studies provide insight into the role of NF-κB subunits in RCC by forming complexes and binding to the promoter of different genes to mediate cancer migration [108]. Conducting research on the molecular dynamic and relative expression of NF-κB members will enhance the understating of the specific mechanism employed.
Based on these findings we can suggest that the most dysregulated NF-κB-related factors and regulators in ccRCC are involved in the canonical NF-κB pathway. Altered genes were found at the regulatory, signal-transmitting, transcription factors, and inducible gene levels of the canonical pathway.
Role of NF-κB in chRCC
One of the most dysregulated genes in cancer is TP53. TP53 has a prominent role in RCC development, especially in chRCC. According to a cohort study done by the COSMIC database, chRCC has the highest rate of TP53 mutation RCC subtypes; notably, TP53 was found to be under-expressed in 34.4 % (28/66) chRCC patients, and it was mutated in 26 % out of 203 patients. p53 and NF-κB regulate each other negatively by different mechanisms. At the transcription level, NF-κB can bind to the promoter of MDM2 and indue its transcription, hence promoting p53 degradation. p53 and NF-κB compete for the limited pool of p300 and CREB-binding protein (p300/CBP), affecting the transcription of each other’s genes [109]. In addition, p53 interferes with the non-canonical NF-κB pathway by decreasing NIK and inhibiting the NF-κB dependent gene transcription [110]. NF-κB was shown to interfere with p53 at the protein level through IKK-mediated p53 phosphorylation, which results in its ubiquitination and degradation [111]. Another mechanism where NF-κB can interfere with P53 is through some NF-κB-responsive genes, such as macrophage migration inhibitory factor (MIF) and IL-6, which can inhibit p53-mediated transcription. As a result, TP53-negative cancer cells are characterized by constant activation of the NF-κB pathway and enhanced tumor progression.
Also, it was demonstrated that even in p53-wild-type cancer cells that exhibit an over-activated NF-κB pathway, p53 will not be fully functional because NF-κB will shut the p53-mediated apoptotic role [112]. Based on COSMIC and TCGA databases, the second most frequent mutated gene in chRCC is Phosphatase and tensin homolog (PTEN). Specifically, PTEN was found to be mutated in 7 % in a study composed of 194 patients, and under expressed in 39.4 % of 66 patients [113]. PTEN loss or inactivating mutations allow Akt to be constitutively active so that it turns to phosphorylate MDM2 and stabilizes it, rendering p53 inactive. At the same time, the PI3K/Akt pathway activates NF-κB [114]. The formerly mentioned mechanism clarifies the role of Akt and NF-κB in inactivating p53; hence, in chRCC, this mechanism suppresses p53 even in patients expressing wild-type p53. In normal cases, p53 is activated in response to stress, inflammation, or DNA damage. However, deletion or suppression of p53 will shift the mechanism to favor the activation and upregulation of NF-κB, hence initiating the transformation process in chRCC. Moreover, by building a conclusion from these studies, we can say that Akt/NF-κB/p53 is the dominant pathway that induces cell transformation in chRCC (Fig. 5). Furthermore, 4.3 % (10/233) mutations were detected in the VHL gene among chRCC cases in a cohort study performed by the COSMIC database, suggesting the absence of another NF-κB regulator, switching NF-κB activity on.
CD44 is a gene regulated by the NF-κB pathway. CD44 acts as an adhesion receptor, and upon activation, it supports cancer migration and invasion in many cancers. Moreover, CD44 is activated and upregulated in response to T lymphocyte activation, whereby, after resolving the trigger, CD44 levels decrease. Notably, among a chRCC cohort study reported by the COSMIC database, 15.2 % (10/66) patients are overexpressing CD44 [114], indicating a highly active NF-κB, producing a positive feedback loop and shifting the response from immune activation to migration and invasion. This evidence provides insight into how NF-κB can be constantly active in chRCC, sustaining the release of factors needed for cancer development and progression.
Metabolic machinery in ccRCC versus chRCC
Cancer cells favor the shift to glycolysis even in the presence of oxygen because, along with ATP, glycolysis produces multiple metabolites that enter various biosynthetic pathways. For instance, glucose 6-phosphate produced by glycolysis enters the pentose phosphate pathway, generating nucleotides needed for DNA synthesis of highly replicating cancerous cells. p53-NF-κB crosstalk has a major impact on glucose metabolism in cancer cells. Consequently, different metabolic pathways are implicated in various renal carcinoma subtypes based on the TCGA database. In which mitochondrial genes in chRCC were found to be highly replicated, overexpressed, and mutated. Moreover, most Krebs cycle enzymes were upregulated, and at least one gene in each electron transport chain complex (ETC) was also upregulated [114]. Conversely, in ccRCC, genes for enzymes involved in the Krebs cycle were suppressed. However, enzymes that play a central role in glycolysis, the pentose phosphate pathway, and fatty acid synthesis were enriched [115], [116]. Some studies demonstrated that NF-κB encourages aerobic respiration in cancer cells that harbor wild-type p53 by increasing the expression of some enzymes, such as the synthesis of cytochrome c oxidase 2 (SCO2). In contrast, in p53-deficient cells, NF-κB increases the tendency to glycolysis and the Warburg effect by enhancing the level of GLUT3 [117], [118]. However, the dialogue in ccRCC and chRCC contradicts the previous study. Based on scientific findings, in some conditions, NF-κB exerts its pro-tumorigenic and metabolic functions through mutated p53, which is the case in chRCC, where NF-κB can switch various mitochondrial respiration machinery even p53 is mutated. Whereas several studies have reported that while p53 is not mutated in high frequency in ccRCC, it’s functionally inhibited by the upregulations of multiple p53 suppressors [118].
Role of NF-κB in PRCC
PRCC represents another rare subtype of RCC; hence, it is characterized by a distinct genetic background [116]. According to a cohort study done by the COSMIC database, the most mutated gene in PRCC is the MET gene, which was found to be mutated in 9 % across 709 patients, whereas 5 % of 286 patients exhibited an increase in gene copy number. Moreover, most cancers tend to mutate and/or upregulate MET, rendering it constitutively active to drive survival, angiogenesis, and migration through cross-talking with different signaling pathways, including NF-κB pathway [119]. Upon activation of c-MET receptor by HGF, one of the main downstream that is activated in response to c-MET is the PI3K/Akt pathway. Akt mediates NF-κB pathway activation through phosphorylation and activating the IKK complex, which subsequently activates the p65-p50 dimer through phosphorylating the inhibitory protein IκBα [120]. Even though MET is mutated in multiple cancers, the first mutations were detected in PRCC, indicating that this receptor plays a critical role in PRCC development and progression. Moreover, the missense mutations in MET frequently affect the tyrosine kinase domain, resulting in oncogenic signals. A constantly active c-MET receptor amplifies the input signal and activates NF-κB to drive cancer survival and proliferation and overcome the apoptotic signals.
TLR2 is one of the NF-κB-inducible genes that will further activate NF-κB upon expression. Hence, TLR2 was mutated in 0.6 %/338 patients in the COSMIC cohort among PRCC. This evidence provides a clue of how the mutations in any of the NF-κB layers could produce countless cycles of NF-κB that will sustain different cancer hallmarks (Fig. 5) [121].
Another gene related to NF-κB and PRCC is the I kappa B kinase interacting protein (IKIP). IKIP was shown to bind to IKKα/β and block its interaction with NEMO, hence negatively regulating the NF-κB pathway. While it acts as a pro-apoptotic molecule and is activated by p53, it was found that many cancers have an increased level of IKIP. According to TIMER and GEPIA databases, IKIP is highly expressed in PRCC, and its expression is negatively correlated to survival while positively linked to higher TNM stage. The reduced expression of IKIP indicates highly activated NF-κB in PRCC, which will induce the expression of oncogenes to sustain survival and migration [122].
Also, 2 % of 645 PRCC samples had VHL mutations in a COSMIC database-based cohort study, suggesting that VHL mutations have a weaker role in PRCC for switching the NF-κB activity. However, the upregulation of the c-MET receptor is the critical player in this cancer subtype to drive NF-κB constant activity [123].
Targeting NF-κB in RCC
The tumor-promoting role of NF-κB encourages scientists to use diverse methods to interfere with NF-κB in RCC. Notably, this includes pharmacological inhibitors as well as genetic interference. Knocking down the p50 subunit in RCC mouse model significantly decreases tumorigenicity. Besides, knocking down p50 in the RCC mouse cell line suppresses cell proliferation and causes late apoptosis [130]. This evidence supports the tumorigenic role of p50 observed in cohort studies done in RCC patients. Besides, Transfecting NOAT113026 long non-coding RNA in RCC cell lines suppresses cell invasion, colony formation, and proliferation by interplay with the p50 NF-κB subunit to interfere with its level [131].
A number of compounds isolated from natural herbs that exhibit anti-inflammatory and anti-cancer properties were found to inhibit NF-κB in RCC. Triptolidenol is a drug used in chronic nephritis, however, its role in ccRCC has been studied. In addition, Triptolidenol inhibited cell migration and induced cell cycle arrest and apoptosis in ccRCC cells by interfering with IKKβ kinase activity. Subsequently, suppressing the transactivation of NF-κB provides a mechanism to hinder RCC progression [132]. Physalin F is a plant extract with an anti-tumor effect. Physalin F showed an anti-cancer effect in RCC cell lines by targeting the NF-κB pathway. The cytotoxic role of Physalin F was mediated by decreased p65 and p50 nuclear translocation and gene transcription including Bcl-2 and Bcl-xL, hence inducing cell apoptosis [133]. Piperlongumine is a natural alkaloid that ablates c-Met expression at RCC cells' gene and protein levels. Furthermore, Piperlongumine was shown to suppress downstream signaling pathways including NF-κB pathway, hence it acts on metastatic RCC [134]. Liguistium wallichii is a natural herb used for medical purposes. The anti-cancer effect of Liguistium wallichii has been evaluated in experimental rat models of RCC. Liguistium wallichii showed a significant decrease in nuclear division and cancer RCC progression by targeting NF-κB activation and p65 nuclear localization. Thus, it is evident that NF-κB plays a major role in DNA replication and gene transcription, resulting in cancer development and progression [135].
These findings suggest that the anti-cancer effect exhibited by natural compounds is mostly through targeting NF-κB. In RCC, especially the ccRCC subtype, the immune mediators are elevated. subsequently, these compounds due to their anti-inflammatory effect were potent in mediating apoptosis by interfering with NF-κB-mediated gene transcription.
The expression of NF-κB has been linked to multiple drug resistance in ccRCC. Notably, the sensitivity of ccRCC to metformin, sorafenib, and gefitinib was better in the group with downregulated NF-κB than in the group with upregulated NF-κB [115]. Also, the NF-κB role has been implied in sunitinib resistance. One of the main tyrosine kinase inhibitors used in advanced ccRCC is sunitinib. However, over 20 % of ccRCC patients develop resistance, and even most patients who showed response in the initial period will develop resistance after months of treatment. Two pathways emanated by a stress response in the endoplasmic reticulum (ER) were shown to mediate sunitinib resistance in ccRCC. One of these two pathways is NF-κB, where sunitinib was shown to activate NF-κB by stabilizing inositol-requiring enzyme 1 (IRE1) in response to ER stress which further recruits and activates TRAF2 leading to canonical NF-κB pathway activation. Notably, targeting NF-κB effectively re-sensitizes RCC to sunitinib while blocking the other resistance mediating pathway, PERK, only suppress the release of pro-inflammatory mediators such as TNF-α and IL-6 [136]. This evidence suggests the main role of NF-κB pathway in mediating RCC resistance to tyrosine kinase inhibitors by promoting several cancer hallmarks.
In a study that used ccRCC cell lines and an RCC mouse model, the effect of BAY-11 was studied. BAY-11 NF-κB inhibitor effectively induced cell apoptosis in cultured cells in a concentration-dependent pattern. Importantly, this effect was accompanied by a decrease in p-p65, indicating that the anti-cancer effect observed was due to NF-κB inhibition. In addition, BAY 11 significantly reduced tumor growth in xenograft nude mice, while it had no effect on cell proliferation [89]. This study proves that NF-κB has a major role in mediating RCC survival and using a selective and irreversible NF-κB inhibitor which is BAY 11 deprives RCC cells of the genes that are needed to sustain survival they committed programmed death.
Most of the tested inhibitors except for BAY 11 are non-specific for the NF-κB pathway, hence the anti-cancer effect observed can’t be exploited to detect the molecular mechanism. Therefore, more studies involving RCC cell lines or mouse models are required to investigate the role of specific NF-κB inhibitors in RCC.
Conclusion and future perspectives
NF-κB pathway orchestrates cellular responses, including proliferation, apoptosis, and immune response. Extensive research has delved into the role of NF-κB in cancer, yet a major gap in the degree of NF-κB dysregulation that shifts the response remains to be discovered. An array of different inhibitors and subunits tightly controls the regulation of NF-κB. Major dysregulation in NF-κB can lead to chronic inflammation due to irritable NF-κB pathway that is unsusceptible to negative feedback response. Chronic inflammation is usually associated with extensive tissue damage, cell type changes, and genetic alterations. Genetic dysregulations of NF-κB multiple subunits could favor the shift of NF-κB from immune response to proliferation survival, and growth leading to a major imbalance in cellular guard mechanisms. In cancer, NF-κB tends to be constitutively active in an autonomous pattern, producing self-sufficient growth signals and promoting RCC cell survival.
Restoring NF-κB regulation could be a novel mechanism to control various auto-immune diseases as well as cancer. NF-κB inhibitors could have a limited role in restoring controlled NF-κB activity. Investigating new mechanisms to modulate the NF-κB pathway could provide a novel method to treat multiple types of cancer, especially RCC. Various molecular studies should be conducted to specify NF-κB subunits that are linked to RCC pathogenesis. Identifying the molecular dynamic of RCC subtypes is essential to overcome RCC’s bad prognosis and current limited treatment options. Each subtype of RCC is characterized by some biomarkers or gene signature. Relative expression levels of different NF-κB subunits could have a major role in impacting its activity and shuttling to the nucleus, hence affecting NF-κB pathway activation and RCC progression. Therefore, NF-κB subunit involved in different RCC subtypes development could be used as a diagnostic and therapeutic biomarker.
Most pharmacological inhibitors of the NF-κB pathway target specific subunits of this pathway while this pathway is composed of tens of receptors, signal transduction complexes, and transcription factors. As a result, other subunits will compensate, and the overall activation will not be modulated. Accordingly, the detailed genetic composition of RCC is required to characterize the genetic landscape of each subtype and target it.
Funding
R.H. is funded by University of Sharjah (grant no: 22010902103) and ASPIRE, the technology program management pillar of Abu Dhabi’s Advanced Technology Research Council (ATRC), via the ASPIRE Precision Medicine Research Institute Abu Dhabi (VRI-20–10).
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Biographies

Nour Abu Jayab MSc, PharmB, 3rd year PhD student in the molecular medicine and translational research program conducted in college of Medicine_university of Sharjah. Graduated from college of pharmacy, university of Sharjah, UAE. She did her master’s in molecular medicine and translational research, college of medicine, university of Sharjah, UAE. A researcher worked in cancer research and currently taking part in molecular genetics research group, Research institute for Medical and Health Sciences. Her masters project focused on genetic mutations in colorectal cancer and melanoma with their impacts on carcinogenesis. She is interested in complex molecular genetics and discovering novel cancer genetic biomarkers. Nour Abu Jayab contributed to multiple research projects in the cancer field with an author h-index of 2. She is working mainly in liver, colorectal, and kidney cancer. She contributed to Conceptualization, Investigation, Writing-Original draft preparation, Writing- Review and editing.

Alaa Abed PD, PhD, MSc, and BDS is a postdoctoral research fellowship in Biomedically-Informed artificial Intelligence laboratory at the Center of Excellence for Precision Medicine, Research institute for Medical and Health Sciences, University of Sharjah, Sharjah, UAE. Alaa Altaie has a dental and medical background and integrative molecular microbiology and immunology. In addition, she is working in Molecular biology, Molecular Microbiology, Metabolomics, Immunology, Bioinformatics, Epidemiology, Systematic Review, Meta-analysis, Oral Microbiology, Cancer research including breast, renal, colorectal, and oral cancer. Her published articles were cited 74 with an author h-index of 6. Alaa Abed contributed as a reviewer in Plos One, Scientific Reports, and Journal of Advanced Research. She contributed to Writing – original draft, Writing-Reviewing and editing.

Iman Talaat M.D, PhD, MSc, MBBCh, graduated from the Faculty of Medicine, Alexandria University, Egypt. in Basic Medical Sciences in Pathology. She is a Full Professor of Pathology at the Clinical Sciences Department, College of Medicine, University of Sharjah, UAE, and a Full Professor of Pathology in the Pathology Department, Faculty of Medicine, Alexandria University, Egypt. She is also a consultant anatomic pathologist. She is known for her research work with an author h-index of 21. Her research work in the field of cancer, focusing mainly on breast, colorectal, prostate, and renal cancers, as well as other solid tumors. In addition, she has good experience in immunohistochemistry (IHC), in-situ hybridization (ISH), and fluorescence in situ hybridization (FISH) techniques. She has more than 70 publications in high-impact peer-reviewed journals and is the PI and co-PI of various internal and external grants. Additionally, she is the coordinator of the immuno-oncology research group at the Research Institute of Medical and Health Sciences (RIMHS), University of Sharjah, Sharjah, UAE. She contributed to Writing-Reviewing and editing.

Rifat Hamoudi Ph.D, C.Eng, Eur Eng, C.Sci, FHEA and MIEEE is Professor in Molecular, Cellular and Computational Medicine within the College of Medicine, University of Sharjah, UAE and he is also an honorary Associate Professor in Molecular and Computational Pathology at University College London, UK. He worked in cancer genetics, molecular and cellular pathology and software engineering projects and was involved in the design and analysis of more than 200 scientific research studies including phase I clinical trials in high impact journals including PNAS, Nature, Nature Genetics, Lancet, Science and Cell spanning the fields of pathology, computer science, engineering and molecular medicine with an H index 59. His main field of research interest is modelling immune regulation, deciphering the role of immune response in disease using engineering and mathematics principles and applying them to the acquisition and analysis of data obtained from cellular and molecular biomedical experiments. He also has patents for the development of non-invasive biomarkers from saliva for esophageal cancer and asthma. He is on the editorial board of many scientific journals. He contributed to Supervision, Conceptualization, investigation, Writing- Reviewing and Editing.
Contributor Information
Iman M. Talaat, Email: italaat@sharjah.ac.ae.
Rifat Hamoudi, Email: rhamoudi@sharjah.ac.ae.
References
- 1.Siegel R.L., Giaquinto A.N., Jemal A. Cancer statistics, 2024. CA Cancer J Clin. 2024;74(1):12–49. doi: 10.3322/caac.21820. [DOI] [PubMed] [Google Scholar]
- 2.Hu M., Fan J.Y., Zhou X., Cao G.W., Tan X. Global incidence and mortality of renal cell carcinoma in 2020. Zhonghua Liu Xing Bing Xue Za Zhi. 2023;44(4):575–580. doi: 10.3760/cma.j.cn112338-20220624-00558. [DOI] [PubMed] [Google Scholar]
- 3.Karin M. Nuclear factor-kappaB in cancer development and progression. Nature. 2006;441(7092):431–436. doi: 10.1038/nature04870. [DOI] [PubMed] [Google Scholar]
- 4.Chen L., Deng H., Cui H., Fang J., Zuo Z., Deng J., et al. Inflammatory responses and inflammation-associated diseases in organs. Oncotarget. 2018;9(6):7204–7218. doi: 10.18632/oncotarget.23208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Singh N., Baby D., Rajguru J.P., Patil P.B., Thakkannavar S.S., Pujari V.B. Inflammation and cancer. Ann Afr Med. 2019;18(3):121–126. doi: 10.4103/aam.aam_56_18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Liu T., Zhang L., Joo D., Sun S.C. NF-κB signaling in inflammation. Signal Transduct Target Ther. 2017;2:17023 -. doi: 10.1038/sigtrans.2017.23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Guan C., Zhou X., Li H., Ma X., Zhuang J. NF-κB inhibitors gifted by nature: The anticancer promise of polyphenol compounds. Biomed Pharmacother. 2022;156 doi: 10.1016/j.biopha.2022.113951. [DOI] [PubMed] [Google Scholar]
- 8.Taniguchi K., Karin M. NF-κB, inflammation, immunity and cancer: coming of age. Nat Rev Immunol. 2018;18(5):309–324. doi: 10.1038/nri.2017.142. [DOI] [PubMed] [Google Scholar]
- 9.Sen R., Baltimore D. Multiple nuclear factors interact with the immunoglobulin enhancer sequences. Cell. 1986;46(5):705–716. doi: 10.1016/0092-8674(86)90346-6. [DOI] [PubMed] [Google Scholar]
- 10.Karin M., Greten F.R. NF-kappaB: linking inflammation and immunity to cancer development and progression. Nat Rev Immunol. 2005;5(10):749–759. doi: 10.1038/nri1703. [DOI] [PubMed] [Google Scholar]
- 11.Yamamoto M., Gohda J., Akiyama T., Inoue J.I. TNF receptor-associated factor 6 (TRAF6) plays crucial roles in multiple biological systems through polyubiquitination-mediated NF-κB activation. Proc Jpn Acad Ser B Phys Biol Sci. 2021;97(4):145–160. doi: 10.2183/pjab.97.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Trares K., Ackermann J., Koch I. The canonical and non-canonical NF-κB pathways and their crosstalk: a comparative study based on Petri nets. Biosystems. 2022;211 doi: 10.1016/j.biosystems.2021.104564. [DOI] [PubMed] [Google Scholar]
- 13.Oeckinghaus A., Ghosh S. The NF-kappaB family of transcription factors and its regulation. Cold Spring Harb Perspect Biol. 2009;1(4) doi: 10.1101/cshperspect.a000034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kearns J.D., Basak S., Werner S.L., Huang C.S., Hoffmann A. IkappaBepsilon provides negative feedback to control NF-kappaB oscillations, signaling dynamics, and inflammatory gene expression. J Cell Biol. 2006;173(5):659–664. doi: 10.1083/jcb.200510155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Morotti A., Crivellaro S., Panuzzo C., Carrà G., Guerrasio A., Saglio G. IκB-α: at the crossroad between oncogenic and tumor-suppressive signals. Oncol Lett. 2017;13(2):531–534. doi: 10.3892/ol.2016.5465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Liu H., Zeng L., Yang Y., Guo C., Wang H. Bcl-3: a double-edged sword in immune cells and inflammation. Front Immunol. 2022;13 doi: 10.3389/fimmu.2022.847699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Shi Y., Wang X., Wang J., Wang X., Zhou H., Zhang L. The dual roles of A20 in cancer. Cancer Lett. 2021;511:26–35. doi: 10.1016/j.canlet.2021.04.017. [DOI] [PubMed] [Google Scholar]
- 18.Du M.Q. MALT lymphoma: a paradigm of NF-κB dysregulation. Semin Cancer Biol. 2016;39:49–60. doi: 10.1016/j.semcancer.2016.07.003. [DOI] [PubMed] [Google Scholar]
- 19.Sun S.C. CYLD: a tumor suppressor deubiquitinase regulating NF-kappaB activation and diverse biological processes. Cell Death Differ. 2010;17(1):25–34. doi: 10.1038/cdd.2009.43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Zhang L., Xiao X., Arnold P.R., Li X.C. Transcriptional and epigenetic regulation of immune tolerance: roles of the NF-κB family members. Cell Mol Immunol. 2019;16(4):315–323. doi: 10.1038/s41423-019-0202-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Cheng J.D., Ryseck R.P., Attar R.M., Dambach D., Bravo R. Functional redundancy of the nuclear factor kappa B inhibitors I kappa B alpha and I kappa B beta. J Exp Med. 1998;188(6):1055–1062. doi: 10.1084/jem.188.6.1055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Begalli F., Bennett J., Capece D., Verzella D., D'Andrea D., Tornatore L., et al. Unlocking the NF-κB conundrum: embracing complexity to achieve specificity. Biomedicines. 2017;5(3) doi: 10.3390/biomedicines5030050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Zong D., Liu X., Li J., Ouyang R., Chen P. The role of cigarette smoke-induced epigenetic alterations in inflammation. Epigen Chrom. 2019;12(1):65. doi: 10.1186/s13072-019-0311-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Rastogi R.P., Richa K.A., Tyagi M.B., Sinha R.P. Molecular mechanisms of ultraviolet radiation-induced DNA damage and repair. J Nucl Acids. 2010; 2010: doi: 10.4061/2010/592980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Zhang C., Qin S., Qin L., Liu L., Sun W., Li X., et al. Cigarette smoke extract-induced p120-mediated NF-κB activation in human epithelial cells is dependent on the RhoA/ROCK pathway. Sci Rep. 2016;6:23131. doi: 10.1038/srep23131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Wang X., Peng H., Huang Y., Kong W., Cui Q., Du J., et al. Post-translational modifications of IκBα: the state of the art. Front Cell Dev Biol. 2020;8 doi: 10.3389/fcell.2020.574706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Lawrence T. The nuclear factor NF-kappaB pathway in inflammation. Cold Spring Harb Perspect Biol. 2009;1(6) doi: 10.1101/cshperspect.a001651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.de Jesus A.A., Canna S.W., Liu Y., Goldbach-Mansky R. Molecular mechanisms in genetically defined autoinflammatory diseases: disorders of amplified danger signaling. Annu Rev Immunol. 2015;33:823–874. doi: 10.1146/annurev-immunol-032414-112227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Goodnow C.C. Pathways for self-tolerance and the treatment of autoimmune diseases. Lancet. 2001;357(9274):2115–2121. doi: 10.1016/s0140-6736(00)05185-0. [DOI] [PubMed] [Google Scholar]
- 30.Hamoudi R.A., Appert A., Ye H., Ruskone-Fourmestraux A., Streubel B., Chott A., et al. Differential expression of NF-kappaB target genes in MALT lymphoma with and without chromosome translocation: insights into molecular mechanism. Leukemia. 2010;24(8):1487–1497. doi: 10.1038/leu.2010.118. [DOI] [PubMed] [Google Scholar]
- 31.Kong-Beltran M., Seshagiri S., Zha J., Zhu W., Bhawe K., Mendoza N., et al. Somatic mutations lead to an oncogenic deletion of met in lung cancer. Cancer Res. 2006;66(1):283–289. doi: 10.1158/0008-5472.Can-05-2749. [DOI] [PubMed] [Google Scholar]
- 32.Di Renzo M.F., Olivero M., Martone T., Maffe A., Maggiora P., Stefani A.D., et al. Somatic mutations of the MET oncogene are selected during metastatic spread of human HNSC carcinomas. Oncogene. 2000;19(12):1547–1555. doi: 10.1038/sj.onc.1203455. [DOI] [PubMed] [Google Scholar]
- 33.Lubensky I.A., Schmidt L., Zhuang Z., Weirich G., Pack S., Zambrano N., et al. Hereditary and sporadic papillary renal carcinomas with c-met mutations share a distinct morphological phenotype. Am J Pathol. 1999;155(2):517–526. doi: 10.1016/s0002-9440(10)65147-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Wei F., Chang Y., Wei W. The role of BAFF in the progression of rheumatoid arthritis. Cytokine. 2015;76(2):537–544. doi: 10.1016/j.cyto.2015.07.014. [DOI] [PubMed] [Google Scholar]
- 35.Stohl W., Metyas S., Tan S.M., Cheema G.S., Oamar B., Xu D., et al. B lymphocyte stimulator overexpression in patients with systemic lupus erythematosus: longitudinal observations. Arthritis Rheum. 2003;48(12):3475–3486. doi: 10.1002/art.11354. [DOI] [PubMed] [Google Scholar]
- 36.Liu Y., Yan W., Tohme S., Chen M., Fu Y., Tian D., et al. Hypoxia induced HMGB1 and mitochondrial DNA interactions mediate tumor growth in hepatocellular carcinoma through Toll-like receptor 9. J Hepatol. 2015;63(1):114–121. doi: 10.1016/j.jhep.2015.02.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Lizzul P.F., Aphale A., Malaviya R., Sun Y., Masud S., Dombrovskiy V., et al. Differential expression of phosphorylated NF-kappaB/RelA in normal and psoriatic epidermis and downregulation of NF-kappaB in response to treatment with etanercept. J Invest Dermatol. 2005;124(6):1275–1283. doi: 10.1111/j.0022-202X.2005.23735.x. [DOI] [PubMed] [Google Scholar]
- 38.Kuo S.H., Yang S.H., Wei M.F., Lee H.W., Tien Y.W., Cheng A.L., et al. Contribution of nuclear BCL10 expression to tumor progression and poor prognosis of advanced and/or metastatic pancreatic ductal adenocarcinoma by activating NF-κB-related signaling. Cancer Cell Int. 2021;21(1):436. doi: 10.1186/s12935-021-02143-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Chan K.K., Shen L., Au W.Y., Yuen H.F., Wong K.Y., Guo T., et al. Interleukin-2 induces NF-kappaB activation through BCL10 and affects its subcellular localization in natural killer lymphoma cells. J Pathol. 2010;221(2):164–174. doi: 10.1002/path.2699. [DOI] [PubMed] [Google Scholar]
- 40.Gordon S., Taylor P.R. Monocyte and macrophage heterogeneity. Nat Rev Immunol. 2005;5(12):953–964. doi: 10.1038/nri1733. [DOI] [PubMed] [Google Scholar]
- 41.Greten F.R., Arkan M.C., Bollrath J., Hsu L.C., Goode J., Miething C., et al. NF-kappaB is a negative regulator of IL-1beta secretion as revealed by genetic and pharmacological inhibition of IKKbeta. Cell. 2007;130(5):918–931. doi: 10.1016/j.cell.2007.07.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Yan J., McCombe P.A., Pender M.P., Greer J.M. Reduced IκB-α protein levels in peripheral blood cells of patients with multiple sclerosis-a possible cause of constitutive NF-κB activation. J Clin Med. 2020;9(8) doi: 10.3390/jcm9082534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Wang E.L., Qian Z.R., Nakasono M., Tanahashi T., Yoshimoto K., Bando Y., et al. High expression of Toll-like receptor 4/myeloid differentiation factor 88 signals correlates with poor prognosis in colorectal cancer. Br J Cancer. 2010;102(5):908–915. doi: 10.1038/sj.bjc.6605558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Brennan F.M., Maini R.N., Feldmann M. Role of pro-inflammatory cytokines in rheumatoid arthritis. Springer Semin Immunopathol. 1998;20(1–2):133–147. doi: 10.1007/bf00832003. [DOI] [PubMed] [Google Scholar]
- 45.Küper C., Beck F.X., Neuhofer W. Autocrine MCP-1/CCR2 signaling stimulates proliferation and migration of renal carcinoma cells. Oncol Lett. 2016;12(3):2201–2209. doi: 10.3892/ol.2016.4875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Janeckova L., Pospichalova V., Fafilek B., Vojtechova M., Tureckova J., Dobes J., et al. HIC1 tumor suppressor loss potentiates TLR2/NF-κB signaling and promotes tissue damage-associated tumorigenesis. Mol Cancer Res. 2015;13(7):1139–1148. doi: 10.1158/1541-7786.Mcr-15-0033. [DOI] [PubMed] [Google Scholar]
- 47.Shi S., Xu C., Fang X., Zhang Y., Li H., Wen W., et al. Expression profile of Toll-like receptors in human breast cancer. Mol Med Rep. 2020;21(2):786–794. doi: 10.3892/mmr.2019.10853. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Song I.J., Yang Y.M., Inokuchi-Shimizu S., Roh Y.S., Yang L., Seki E. The contribution of toll-like receptor signaling to the development of liver fibrosis and cancer in hepatocyte-specific TAK1-deleted mice. Int J Cancer. 2018;142(1):81–91. doi: 10.1002/ijc.31029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Garner H., de Visser K.E. Immune crosstalk in cancer progression and metastatic spread: a complex conversation. Nat Rev Immunol. 2020;20(8):483–497. doi: 10.1038/s41577-019-0271-z. [DOI] [PubMed] [Google Scholar]
- 50.Rasmi R.R., Sakthivel K.M., Guruvayoorappan C. NF-κB inhibitors in treatment and prevention of lung cancer. Biomed Pharmacother. 2020;130 doi: 10.1016/j.biopha.2020.110569. [DOI] [PubMed] [Google Scholar]
- 51.Xie K., Zhou D., Fang C., Pu R., Zhu Z. Inhibition of NF-κB activation by BAY 11–7821 suppresses the proliferation and inflammation of glioma cells through inducing autophagy. Transl Cancer Res. 2022;11(2):403–413. doi: 10.21037/tcr-21-2914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Yamamoto Y., Gaynor R.B. Therapeutic potential of inhibition of the NF-kappaB pathway in the treatment of inflammation and cancer. J Clin Invest. 2001;107(2):135–142. doi: 10.1172/jci11914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Yi Y.W., You K.S., Han S., Ha I.J., Park J.S., Lee S.G., et al. Inhibition of IκB kinase is a potential therapeutic strategy to circumvent resistance to epidermal growth factor receptor inhibition in triple-negative breast cancer cells. Cancers (Basel) 2022;14(21) doi: 10.3390/cancers14215215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Hanahan D., Weinberg R.A. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–674. doi: 10.1016/j.cell.2011.02.013. [DOI] [PubMed] [Google Scholar]
- 55.Oh A., Pardo M., Rodriguez A., Yu C., Nguyen L., Liang O., et al. NF-κB signaling in neoplastic transition from epithelial to mesenchymal phenotype. Cell Commun Signal. 2023;21(1):291. doi: 10.1186/s12964-023-01207-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Huber M.A., Azoitei N., Baumann B., Grünert S., Sommer A., Pehamberger H., et al. NF-kappaB is essential for epithelial-mesenchymal transition and metastasis in a model of breast cancer progression. J Clin Invest. 2004;114(4):569–581. doi: 10.1172/jci21358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Rastogi S., Aldosary S., Saeedan A.S., Ansari M.N., Singh M., Kaithwas G. NF-κB mediated regulation of tumor cell proliferation in hypoxic microenvironment. Front Pharmacol. 2023;14:1108915. doi: 10.3389/fphar.2023.1108915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Tabruyn S.P., Griffioen A.W. NF-kappa B: a new player in angiostatic therapy. Angiogenesis. 2008;11(1):101–106. doi: 10.1007/s10456-008-9094-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Verzella D., Pescatore A., Capece D., Vecchiotti D., Ursini M.V., Franzoso G., et al. Life, death, and autophagy in cancer: NF-κB turns up everywhere. Cell Death Dis. 2020;11(3):210. doi: 10.1038/s41419-020-2399-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Conte D., Holcik M., Lefebvre C.A., Lacasse E., Picketts D.J., Wright K.E., et al. Inhibitor of apoptosis protein cIAP2 is essential for lipopolysaccharide-induced macrophage survival. Mol Cell Biol. 2006;26(2):699–708. doi: 10.1128/mcb.26.2.699-708.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Joyce D., Albanese C., Steer J., Fu M., Bouzahzah B., Pestell R.G. NF-κB and cell-cycle regulation: the cyclin connection. Cytokine Growth Factor Rev. 2001;12(1):73–90. doi: 10.1016/s1359-6101(00)00018-6. [DOI] [PubMed] [Google Scholar]
- 62.Joyce D., Albanese C., Steer J., Fu M., Bouzahzah B., Pestell R.G. NF-kappaB and cell-cycle regulation: the cyclin connection. Cytokine Growth Factor Rev. 2001;12(1):73–90. doi: 10.1016/s1359-6101(00)00018-6. [DOI] [PubMed] [Google Scholar]
- 63.Oh H., Grinberg-Bleyer Y., Liao W., Maloney D., Wang P., Wu Z., et al. An NF-κB transcription-factor-dependent lineage-specific transcriptional program promotes regulatory T cell identity and function. Immunity. 2017;47(3) doi: 10.1016/j.immuni.2017.08.010. 450-65.e5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Shang B., Liu Y., Jiang S.J., Liu Y. Prognostic value of tumor-infiltrating FoxP3+ regulatory T cells in cancers: a systematic review and meta-analysis. Sci Rep. 2015;5:15179. doi: 10.1038/srep15179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Porta C., Rimoldi M., Raes G., Brys L., Ghezzi P., Di Liberto D., et al. Tolerance and M2 (alternative) macrophage polarization are related processes orchestrated by p50 nuclear factor kappaB. Proc Natl Acad Sci USA. 2009;106(35):14978–14983. doi: 10.1073/pnas.0809784106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Barberà M.J., Puig I., Domínguez D., Julien-Grille S., Guaita-Esteruelas S., Peiró S., et al. Regulation of Snail transcription during epithelial to mesenchymal transition of tumor cells. Oncogene. 2004;23(44):7345–7354. doi: 10.1038/sj.onc.1207990. [DOI] [PubMed] [Google Scholar]
- 67.Li C.-W., Xia W., Huo L., Lim S.-O., Wu Y., Hsu J.L., et al. Epithelial–mesenchymal transition induced by TNF-α requires NF-κB–mediated transcriptional upregulation of Twist1. Cancer Res. 2012;72(5):1290–1300. doi: 10.1158/0008-5472.CAN-11-3123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Al-Roub A., Akhter N., Al-Rashed F., Wilson A., Alzaid F., Al-Mulla F., et al. TNFα induces matrix metalloproteinase-9 expression in monocytic cells through ACSL1/JNK/ERK/NF-kB signaling pathways. Sci Rep. 2023;13(1):14351. doi: 10.1038/s41598-023-41514-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Kew R.R., Penzo M., Habiel D.M., Marcu K.B. The IKKα-dependent NF-κB p52/RelB noncanonical pathway is essential to sustain a CXCL12 autocrine loop in cells migrating in response to HMGB1. J Immunol. 2012;188(5):2380–2386. doi: 10.4049/jimmunol.1102454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Milstone D.S., Ilyama M., Chen M., O'Donnell P., Davis V.M., Plutzky J., et al. Differential role of an NF-κB transcriptional response element in endothelial versus intimal cell VCAM-1 expression. Circ Res. 2015;117(2):166–177. doi: 10.1161/circresaha.117.306666. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.van Uden P., Kenneth N.S., Rocha S. Regulation of hypoxia-inducible factor-1alpha by NF-kappaB. Biochem J. 2008;412(3):477–484. doi: 10.1042/bj20080476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Ghiorzo P., Musso M., Mantelli M., Garré C., Ravazzolo R., Bianchi-Scarrà G. c-Rel and p65 subunits bind to an upstream NF-kappaB site in human granulocyte macrophage-colony stimulating factor promoter involved in phorbol ester response in 5637 cells. FEBS Lett. 1997;418(1–2):215–218. doi: 10.1016/s0014-5793(97)01387-2. [DOI] [PubMed] [Google Scholar]
- 73.Tsai P.-W., Shiah S.-G., Lin M.-T., Wu C.-W., Kuo M.-L. Up-regulation of vascular endothelial growth factor C in breast cancer cells by heregulin-β1: a critical role of p38/nuclear factor-κB signaling pathway. J Biol Chem. 2003;278(8):5750–5759. doi: 10.1074/jbc.M204863200. [DOI] [PubMed] [Google Scholar]
- 74.Yang Y., Qin J., Lan L., Li N., Wang C., He P., et al. M-CSF cooperating with NFκB induces macrophage transformation from M1 to M2 by upregulating c-Jun. Cancer Biol Ther. 2014;15(1):99–107. doi: 10.4161/cbt.26718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Ayroldi E., Grohmann U. Exemplifying complexity of immune suppression by a “canonical” speech: a glimpse into TNFRSF-activated signaling pathways in Treg cells. Eur J Immunol. 2020;50(7):944–948. doi: 10.1002/eji.202048711. [DOI] [PubMed] [Google Scholar]
- 76.Wissink S., van de Stolpe A., Caldenhoven E., Koenderman L., van der Saag P.T. NF-kappa B/Rel family members regulating the ICAM-1 promoter in monocytic THP-1 cells. Immunobiology. 1997;198(1–3):50–64. doi: 10.1016/s0171-2985(97)80026-5. [DOI] [PubMed] [Google Scholar]
- 77.Chang S.F., Lin S.S., Yang H.C., Chou Y.Y., Gao J.I., Lu S.C. LPS-induced G-CSF expression in macrophages is mediated by ERK2, but Not ERK1. PLoS One. 2015;10(6):e0129685. doi: 10.1371/journal.pone.0129685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Busuttil V., Droin N., McCormick L., Bernassola F., Candi E., Melino G., et al. NF-κB inhibits T-cell activation-induced, p73-dependent cell death by induction of MDM2. Proc Natl Acad Sci. 2010;107(42):18061–18066. doi: 10.1073/pnas.1006163107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Ma C., Wang Y., Dong L., Li M., Cai W. Anti-inflammatory effect of resveratrol through the suppression of NF-κB and JAK/STAT signaling pathways. Acta Biochim Biophys Sin (Shanghai) 2015;47(3):207–213. doi: 10.1093/abbs/gmu135. [DOI] [PubMed] [Google Scholar]
- 80.Cordon-Cardo C., Prives C. At the crossroads of inflammation and tumorigenesis. J Exp Med. 1999;190(10):1367–1370. doi: 10.1084/jem.190.10.1367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Prasad SR, Humphrey PA, Catena JR, Narra VR, Srigley JR, Cortez AD, et al. Common and uncommon histologic subtypes of renal cell carcinoma: imaging spectrum with pathologic correlation. Radiographics. 2006;26(6):1795–806; discussion 806–10. 10.1148/rg.266065010. [DOI] [PubMed]
- 82.Oya M., Takayanagi A., Horiguchi A., Mizuno R., Ohtsubo M., Marumo K., et al. Increased nuclear factor-kappa B activation is related to the tumor development of renal cell carcinoma. Carcinogenesis. 2003;24(3):377–384. doi: 10.1093/carcin/24.3.377. [DOI] [PubMed] [Google Scholar]
- 83.Meteoglu I., Erdogdu I.H., Meydan N., Erkus M., Barutca S. NF-KappaB expression correlates with apoptosis and angiogenesis in clear cell renal cell carcinoma tissues. J Exp Clin Cancer Res. 2008;27(1):53. doi: 10.1186/1756-9966-27-53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Matušan-Ilijaš K., Damante G., Fabbro D., Dorđević G., Hadžisejdić I., Grahovac M., et al. Osteopontin expression correlates with nuclear factor-κB activation and apoptosis downregulation in clear cell renal cell carcinoma. Pathol Res Pract. 2011;207(2):104–110. doi: 10.1016/j.prp.2010.11.004. [DOI] [PubMed] [Google Scholar]
- 85.Peri S., Devarajan K., Yang D.H., Knudson A.G., Balachandran S. Meta-analysis identifies NF-κB as a therapeutic target in renal cancer. PLoS One. 2013;8(10):e76746. doi: 10.1371/journal.pone.0076746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Bartha Á., Győrffy B. TNMplot.com: a web tool for the comparison of gene expression in normal, tumor and metastatic tissues. Int J Mol Sci. 2021;22(5) doi: 10.3390/ijms22052622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Farhadi A., Namdari S., Chong P.P., Geramizadeh B., Behzad-Behbahani A., Sekawi Z., et al. Epstein-Barr virus infection is associated with the nuclear factor-kappa B p65 signaling pathway in renal cell carcinoma. BMC Urol. 2022;22(1):17. doi: 10.1186/s12894-022-00964-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Banumathy G., Cairns P. Signaling pathways in renal cell carcinoma. Cancer Biol Ther. 2010;10(7):658–664. doi: 10.4161/cbt.10.7.13247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Sourbier C., Danilin S., Lindner V., Steger J., Rothhut S., Meyer N., et al. Targeting the nuclear factor-kappaB rescue pathway has promising future in human renal cell carcinoma therapy. Cancer Res. 2007;67(24):11668–11676. doi: 10.1158/0008-5472.Can-07-0632. [DOI] [PubMed] [Google Scholar]
- 90.Troppan K., Hofer S., Wenzl K., Lassnig M., Pursche B., Steinbauer E., et al. Frequent down regulation of the tumor suppressor gene a20 in multiple myeloma. PLoS One. 2015;10(4):e0123922. doi: 10.1371/journal.pone.0123922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Peng L., He K., Cao Z., Bi L., Yu D., Wang Q., et al. CARD10 promotes the progression of renal cell carcinoma by regulating the NF-κB signaling pathway. Mol Med Rep. 2020;21(1):329–337. doi: 10.3892/mmr.2019.10840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Klatte T., Seligson D.B., Riggs S.B., Leppert J.T., Berkman M.K., Kleid M.D., et al. Hypoxia-inducible factor 1 alpha in clear cell renal cell carcinoma. Clin Cancer Res. 2007;13(24):7388–7393. doi: 10.1158/1078-0432.Ccr-07-0411. [DOI] [PubMed] [Google Scholar]
- 93.Lowe J.M., Cha H., Yang Q., Fornace A.J., Jr. Nuclear factor-kappaB (NF-kappaB) is a novel positive transcriptional regulator of the oncogenic Wip1 phosphatase. J Biol Chem. 2010;285(8):5249–5257. doi: 10.1074/jbc.M109.034579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Sun G.G., Wang Y.D., Liu Q., Hu W.N. Expression of Wip1 in kidney carcinoma and its correlation with tumor metastasis and clinical significance. Pathol Oncol Res. 2015;21(1):219–224. doi: 10.1007/s12253-014-9811-9. [DOI] [PubMed] [Google Scholar]
- 95.Deng W., Li J., Dorrah K., Jimenez-Tapia D., Arriaga B., Hao Q., et al. The role of PPM1D in cancer and advances in studies of its inhibitors. Biomed Pharmacother. 2020;125 doi: 10.1016/j.biopha.2020.109956. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Haase V.H. The VHL tumor suppressor: master regulator of HIF. Curr Pharm Des. 2009;15(33):3895–3903. doi: 10.2174/138161209789649394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Morais C., Gobe G., Johnson D.W., Healy H. The emerging role of nuclear factor kappa B in renal cell carcinoma. Int J Biochem Cell Biol. 2011;43(11):1537–1549. doi: 10.1016/j.biocel.2011.08.003. [DOI] [PubMed] [Google Scholar]
- 98.de Paulsen N., Brychzy A., Fournier M.C., Klausner R.D., Gnarra J.R., Pause A., et al. Role of transforming growth factor-alpha in von Hippel-Lindau (VHL)(-/-) clear cell renal carcinoma cell proliferation: a possible mechanism coupling VHL tumor suppressor inactivation and tumorigenesis. Proc Natl Acad Sci U S A. 2001;98(4):1387–1392. doi: 10.1073/pnas.98.4.1387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Zhan B., Dong X., Yuan Y., Gong Z., Li B. hZIP1 inhibits progression of clear cell renal cell carcinoma by suppressing NF-kB/HIF-1α pathway. Front Oncol. 2021;11 doi: 10.3389/fonc.2021.759818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Diesing K., Ribback S., Winter S., Gellert M., Oster A.M., Stühler V., et al. p53 is functionally inhibited in clear cell renal cell carcinoma (ccRCC): a mechanistic and correlative investigation into genetic and molecular characteristics. J Cancer Res Clin Oncol. 2021;147(12):3565–3576. doi: 10.1007/s00432-021-03786-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.LF ST, Bellini MH. NF-ĸΒ1 knockout reduces IL6 expression under hypoxia in renal cell carcinoma. Cell Mol Biol (Noisy-le-grand). 2023;69(6):8-14. doi:10.14715/cmb/2023.69.6.2. [DOI] [PubMed]
- 102.Pariente-Pérez T., Aguilar-Alonso F., Solano J.D., Vargas-Olvera C., Curiel-Muñiz P., Mendoza-Rodríguez C.A., et al. Differential behavior of NF-κB, IκBα and EGFR during the renal carcinogenic process in an experimental model in vivo. Oncol Lett. 2020;19(4):3153–3164. doi: 10.3892/ol.2020.11436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Sung M-K, Bae Y-J. Iron, Oxidative stress, and cancer. Cancer: oxidative stress and dietary antioxidants 2014:139–49. doi:10.1016/B978-0-12-405205-5.00013-1.
- 104.Zhang H., Sun S.C. NF-κB in inflammation and renal diseases. Cell Biosci. 2015;5:63. doi: 10.1186/s13578-015-0056-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Bamford S., Dawson E., Forbes S., Clements J., Pettett R., Dogan A., et al. The COSMIC (Catalogue of Somatic Mutations in Cancer) database and website. Br J Cancer. 2004;91(2):355–358. doi: 10.1038/sj.bjc.6601894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Maldonado V., Melendez-Zajgla J. Role of Bcl-3 in solid tumors. Mol Cancer. 2011;10:152. doi: 10.1186/1476-4598-10-152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Kashatus D., Cogswell P., Baldwin A.S. Expression of the Bcl-3 proto-oncogene suppresses p53 activation. Genes Dev. 2006;20(2):225–235. doi: 10.1101/gad.1352206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.de Souza B.M., da Silva Paiva K.B., Foguer K., Barbosa Chaves K.C., de Sá L.L., Scavone C., et al. Involvement of the NF-кB/p50/Bcl-3 complex in response to antiangiogenic therapy in a mouse model of metastatic renal cell carcinoma. Biomed Pharmacother. 2014;68(7):873–879. doi: 10.1016/j.biopha.2014.07.008. [DOI] [PubMed] [Google Scholar]
- 109.Webster G.A., Perkins N.D. Transcriptional cross talk between NF-kappaB and p53. Mol Cell Biol. 1999;19(5):3485–3495. doi: 10.1128/mcb.19.5.3485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Jang H, Park S, Kim J, Kim JH, Kim SY, Cho S, et al. The tumor suppressor, p53, negatively regulates non-canonical NF-κB signaling through miRNAInduced Silencing of NF-κB-Inducing Kinase. Mol Cells 2020;43(1):23-33. 10.14348/molcells.2019.0239. [DOI] [PMC free article] [PubMed]
- 111.Xia Y., Padre R.C., De Mendoza T.H., Bottero V., Tergaonkar V.B., Verma I.M. Phosphorylation of p53 by IkappaB kinase 2 promotes its degradation by beta-TrCP. Proc Natl Acad Sci USA. 2009;106(8):2629–2634. doi: 10.1073/pnas.0812256106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Gudkov A.V., Gurova K.V., Komarova E.A. Inflammation and p53: A tale of two stresses. Genes Cancer. 2011;2(4):503–516. doi: 10.1177/1947601911409747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Roldan-Romero J.M., Santos M., Lanillos J., Caleiras E., Anguera G., Maroto P., et al. Molecular characterization of chromophobe renal cell carcinoma reveals mTOR pathway alterations in patients with poor outcome. Mod Pathol. 2020;33(12):2580–2590. doi: 10.1038/s41379-020-0607-z. [DOI] [PubMed] [Google Scholar]
- 114.Davis C.F., Ricketts C.J., Wang M., Yang L., Cherniack A.D., Shen H., et al. The somatic genomic landscape of chromophobe renal cell carcinoma. Cancer Cell. 2014;26(3):319–330. doi: 10.1016/j.ccr.2014.07.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 499(7456) (2013) 43–49. 10.1038/nature12222. [DOI] [PMC free article] [PubMed]
- 116.Zhu H., Wang X., Lu S., Ou K. Metabolic reprogramming of clear cell renal cell carcinoma. Front Endocrinol (Lausanne) 2023;14:1195500. doi: 10.3389/fendo.2023.1195500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Johnson R.F., Perkins N.D. Nuclear factor-κB, p53, and mitochondria: regulation of cellular metabolism and the Warburg effect. Trends Biochem Sci. 2012;37(8):317–324. doi: 10.1016/j.tibs.2012.04.002. [DOI] [PubMed] [Google Scholar]
- 118.Mauro C., Leow S.C., Anso E., Rocha S., Thotakura A.K., Tornatore L., et al. NF-κB controls energy homeostasis and metabolic adaptation by upregulating mitochondrial respiration. Nat Cell Biol. 2011;13(10):1272–1279. doi: 10.1038/ncb2324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Ariyawutyakorn W., Saichaemchan S., Varella-Garcia M. Understanding and targeting MET signaling in solid tumors - are we there yet? J Cancer. 2016;7(6):633–649. doi: 10.7150/jca.12663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Barzaman K., Vafaei R., Samadi M., Kazemi M.H., Hosseinzadeh A., Merikhian P., et al. Anti-cancer therapeutic strategies based on HGF/MET, EpCAM, and tumor-stromal cross talk. Cancer Cell Int. 2022;22(1):259. doi: 10.1186/s12935-022-02658-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Schmidt L., Duh F.M., Chen F., Kishida T., Glenn G., Choyke P., et al. Germline and somatic mutations in the tyrosine kinase domain of the MET proto-oncogene in papillary renal carcinomas. Nat Genet. 1997;16(1):68–73. doi: 10.1038/ng0597-68. [DOI] [PubMed] [Google Scholar]
- 122.Zhang H., Tang R., Wen X., Cai J., Huang J., Luo L., et al. Prognostic value of IKBIP in papillary renal cell carcinoma. BMC Urol. 2023;23(1):121. doi: 10.1186/s12894-023-01290-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Büscheck F., Fraune C., Simon R., Kluth M., Hube-Magg C., Möller-Koop C., et al. Prevalence and clinical significance of VHL mutations and 3p25 deletions in renal tumor subtypes. Oncotarget. 2020;11(3):237–249. doi: 10.18632/oncotarget.27428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Gerlinger M., Rowan A.J., Horswell S., Math M., Larkin J., Endesfelder D., et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med. 2012;366(10):883–892. doi: 10.1056/NEJMoa1113205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Razafinjatovo C., Bihr S., Mischo A., Vogl U., Schmidinger M., Moch H., et al. Characterization of VHL missense mutations in sporadic clear cell renal cell carcinoma: hotspots, affected binding domains, functional impact on pVHL and therapeutic relevance. BMC Cancer. 2016;16:638. doi: 10.1186/s12885-016-2688-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Hasumi H., Baba M., Hasumi Y., Furuya M., Yao M. Birt-Hogg-Dubé syndrome: clinical and molecular aspects of recently identified kidney cancer syndrome. Int J Urol. 2016;23(3):204–210. doi: 10.1111/iju.13015. [DOI] [PubMed] [Google Scholar]
- 127.Ramirez Reyes J.M.J., Cuesta R., Pause A. Folliculin: a regulator of transcription through AMPK and mTOR signaling pathways. Front Cell Dev Biol. 2021;9 doi: 10.3389/fcell.2021.667311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Sweeney P.L., Jang A., Halat S.K., Pal S.K., Barata P.C. Advanced papillary renal cell carcinoma: epidemiology, genomic drivers, current therapies, and ongoing trials. Cancer Treat Res Commun. 2022;33 doi: 10.1016/j.ctarc.2022.100639. [DOI] [PubMed] [Google Scholar]
- 129.Zhang X.B., Urlando C., Tao K.S., Heddle J.A. Factors affecting somatic mutation frequencies in vivo. Mutat Res. 1995;338(1–6):189–201. doi: 10.1016/0921-8734(95)00024-z. [DOI] [PubMed] [Google Scholar]
- 130.Ikegami A., Teixeira L.F.S., Braga M.S., Dias M., Lopes E.C., Bellini M.H. Knockdown of NF-κB1 by shRNA inhibits the growth of renal cell carcinoma in vitro and in vivo. Oncol Res. 2018;26(5):743–751. doi: 10.3727/096504017x15120379906339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Pu Y., Dong Z., Xia Y., Zhang M., Song J., Han J., et al. LncRNA NONHSAT113026 represses renal cell carcinoma tumorigenesis through interacting with NF-κB/p50 and SLUG. Biomed Pharmacother. 2019;118 doi: 10.1016/j.biopha.2019.109382. [DOI] [PubMed] [Google Scholar]
- 132.Jin J., Zhou M., Wang X., Liu M., Huang H., Yan F., et al. Triptolidenol, isolated from Tripterygium wilfordii, disrupted NF-κB/COX-2 pathway by targeting ATP-binding sites of IKKβ in clear cell renal cell carcinoma. Fitoterapia. 2021;148 doi: 10.1016/j.fitote.2020.104779. [DOI] [PubMed] [Google Scholar]
- 133.Wu S.Y., Leu Y.L., Chang Y.L., Wu T.S., Kuo P.C., Liao Y.R., et al. Physalin F induces cell apoptosis in human renal carcinoma cells by targeting NF-kappaB and generating reactive oxygen species. PLoS One. 2012;7(7):e40727. doi: 10.1371/journal.pone.0040727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Golovine K., Makhov P., Naito S., Raiyani H., Tomaszewski J., Mehrazin R., et al. Piperlongumine and its analogs down-regulate expression of c-Met in renal cell carcinoma. Cancer Biol Ther. 2015;16(5):743–749. doi: 10.1080/15384047.2015.1026511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Wu J.S., Zhang P.M., Sun L.J., Liu X.Z. Liguisticum wallichii inhibits renal carcinoma progression by downregulating UBE3A and through suppression of NF-κB signaling. Genet Mol Res. 2016;15(4) doi: 10.4238/gmr15049023. [DOI] [PubMed] [Google Scholar]
- 136.Makhov P., Naito S., Haifler M., Kutikov A., Boumber Y., Uzzo R.G., et al. The convergent roles of NF-κB and ER stress in sunitinib-mediated expression of pro-tumorigenic cytokines and refractory phenotype in renal cell carcinoma. Cell Death Dis. 2018;9(3):374. doi: 10.1038/s41419-018-0388-1. [DOI] [PMC free article] [PubMed] [Google Scholar]