Abstract
The G1 cyclin Cln2 negatively regulates the mating-factor pathway. In a genetic screen to identify factors required for this regulation, we identified an allele of CDC28 (cdc28-csr1) that blocked this function of Cln2. Cln2 immunoprecipitated from cdc28-csr1 cells was completely defective in histone H1 kinase activity, due to defects in Cdc28 binding and activation by Cln2. In contrast, Clb2-associated H1 kinase and Cdc28 binding was normal in immunoprecipitates from these cells. cdc28-csr1 was significantly deficient in other aspects of genetic interaction with Cln2. The cdc28-csr1 mutation was determined to be Q188P, in the T loop distal to most of the probable Cdk-cyclin interaction regions. We performed random mutagenesis of CDC28 to identify additional alleles incapable of causing CLN2-dependent mating-factor resistance but capable of complementing cdc28 temperature-sensitive and null alleles. Two such mutants had highly defective Cln2-associated kinase, but, surprisingly, two other mutants had levels of Cln2-associated kinase near to wild-type levels. We performed a complementary screen for CDC28 mutants that could cause efficient Cln2-dependent mating-factor resistance but not complement a cdc28 null allele. Most such mutants were found to alter residues essential for kinase activity; the proteins had little or no associated kinase activity in bulk or in association with Cln2. Several of these mutants also functioned in another assay for CLN2-dependent function not involving the mating-factor pathway, complementing the temperature sensitivity of a cln1 cln3 cdc28-csr1 strain. These results could indicate that Cln2-Cdc28 kinase activity is not directly relevant to some CLN2-mediated functions. Mutants of this sort should be useful in differentiating the function of Cdc28 complexed with different cyclin regulatory subunits.
Cyclin-dependent kinases (Cdks) regulate the occurrence of diverse cell cycle events in eukaryotic cells. There is considerable diversity of the kinases and still more diversity of the cyclins, even in a single organism. In the budding yeast, Saccharomyces cerevisiae, Cdc28 is the major Cdk involved in cell cycle regulation, and it is activated by nine different cyclins: the three G1 cyclins Cln1, Cln2, and Cln3, involved in regulating the Start event; and the six B-type cyclins (Clbs) involved in regulating S phase and mitosis (6, 23). Although activation of a Cdk by multiple cyclins is commonly observed in other organisms, the situation in budding yeast is unusual in that cyclins of divergent sequence classes all activate the same Cdk. B-type, CLN1,2-type, and CLN3 cyclins have only ∼25% identity in the most highly conserved cyclin box region, which comprises the Cdk binding interface. The high efficiency of functional activation of Cdc28 by these very divergent cyclins poses an interesting problem in molecular recognition.
Cln3-Cdc28 probably functions in vivo primarily as a transcriptional activator of CLN1 and CLN2 as well as other genes (10, 18, 33, 35). Cln1 and Cln2 are thought to directly drive the Start event, consisting of bud emergence, negative regulation of the mating-factor response pathway, and activation of the Clb-Cdc28 kinases by stabilizing Clb proteins and by inducing degradation of the Sic1 inhibitor of Clb-Cdc28 kinase activity (6). Clb-Cdc28 complexes are required for later cell cycle events such as DNA replication and mitosis (23). In most cases, these different regulatory specificities of the cyclin-Cdc28 complexes can be shown to be due to the identity of the cyclin, although significant functional overlap between different cyclins is observed.
The activity of the mating-factor signalling pathway is cell cycle regulated. Both basal and pheromone-induced transcription of pheromone-responsive genes are reduced as cells enter S phase (25, 26, 37, 40). It was observed that deletion of CLN1 and CLN2 abolishes the cell cycle-regulated inducibility of the pheromone-responsive gene FUS1 while overexpression of CLN2 leads to both strong repression of pheromone-induced FUS1 transcription and resistance to mating-pheromone arrest (25). Expression of the G1 cyclin genes CLN1 and CLN2 as cells enter S phase is therefore required for repression of the mating-factor-induced signalling (25). Inhibition of the mating-factor pathway by Cln1 and Cln2 appears to be highly specific to these homologous cyclins, since neither Cln3 nor Clb cyclins have a detectable ability to inhibit the mating-factor pathway (25). The mechanism by which Cln1 or Cln2 inhibits the mating-factor pathway is unknown, but the step in the pathway that is inhibited has been deduced to be postreceptor and probably post-G-protein activation (24, 25, 37). To study the mechanism by which Cln2 inhibits the mating-factor pathway, we initiated a genetic screen for mutations that block this inhibition. From this screen, we isolated a mutant allele of CDC28 that was defective in binding and activation by Cln2. This mutant Cdc28 functioned well with Clb cyclins and complemented a cdc28 null allele. We screened for additional cdc28 mutants with similar phenotypes and also for cdc28 mutants with the reverse phenotype (i.e., the ability to confer Cln2-dependent mating-factor pathway inhibition combined with a failure to complement a cdc28 null allele). Both screens produced cdc28 mutants with diverse biochemical defects in cyclin binding and kinase activation.
MATERIALS AND METHODS
Plasmids.
The plasmids used in this study are described in Table 1. pKL038-1 (cdc28::HIS3/TRP1) was constructed as follows. pRD47 (CDC28/TRP1/Ampr) (9) was digested with AflII and ClaI, releasing a 568-bp fragment of the CDC28 coding region. The 3′ recessed ends were blunted with Klenow. pJA50-Delta (1), a plasmid containing a HIS3/Kanr cassette, was digested with SmaI, releasing the HIS3/Kanr fragment. The blunted pRD47 fragments were ligated to the SmaI-digested pJA50-Delta fragments and transformed into Escherichia coli. E. coli carrying the HIS3/Kanr-disrupted plasmids were selected on Luria-Bertani (LB) medium containing ampicillin and kanamycin, and correct constructs were confirmed by KpnI digests. pKL049-1 (cdc28::Camr/TRP1) and pKL050-1 (cdc28-HA::Camr/TRP1) were constructed as follows. The chloramphenicol resistance gene, Camr, contained within the pBCKS(+) plasmid was amplified by PCR, with the incorporation of AflII and ClaI sites at the 5′ and 3′ ends of the gene, respectively. The resulting PCR product was digested with AflII and ClaI, ligated to the AflII-ClaI-digested and gel-purified pRD58 (cdc28T169A/TRP1) or pSF19 (CDC28-HA/TRP1) vector fragment, and transformed into E. coli. E. coli carrying the Camr-disrupted plasmids were selected on Luria-Bertani medium containing ampicillin and chloramphenicol, and the correct constructs were confirmed by digestion. Both hemagglutinin (HA)-tagged and untagged versions of the cdc28-csr and cdc28-rsc plasmids (pKL045-KL048 and pKL054-KL066) were constructed by in vivo gap repair (described below). pKL039-1 (cdc28-csr1/URA3) was constructed by switching the TRP1 marker of pKL054 to URA3 with pTU10, as described previously (7). Similarly, pKL051 (CLN2-3P/URA3), pKL052 (cln2K129A-3P/URA3), and pKL053 (cln2E183A-3P/URA3) were constructed by switching the TRP1 markers of pT411, pKL003, and pKL004 (18), respectively, to URA3 with pTU10 (7). pKL019 (cdc28T169A-HA/TRP1) was constructed by digesting pRD58 (cdc28T169A/TRP1) with AflII and ClaI and subcloning the AflII-ClaI fragment into an AflII-ClaI-digested pSF19 (CDC28-HA/TRP1). Plasmids containing the single-mutant alleles cdc28-L61S and cdc28-K187E were constructed by the splice overlap extension method described previously (13), with both wild-type CDC28 and cdc28-csr19 or cdc28-csr41, respectively, as the templates. Following PCR amplification of the single-mutant fragments, pKL075 and pKL077 (cdc28-L61S) and pKL081 and pKL082 (cdc28-K187E) were constructed by in vivo gap repair (described below) with the AflII-ClI-digested untagged vector pKL049-1; KL078 and KL080 (cdc28-L61S-HA), and KL084 and KL085 (cdc28-K187E-HA) were constructed with the AflII-ClI-digested HA-tagged vector pKL050-1. Each of the single-mutant alleles was fully sequenced to ensure the correct construction.
TABLE 1.
Plasmids used in this study
| Plasmid | Relevant genes | Source |
|---|---|---|
| 143 | GAL1::CLB2-HA URA3 AmprCEN | R. J. Deshaies |
| BCKS(+) | Camr | Stratagene |
| JA50-Delta | HIS3 Kanr | 1 |
| KH100 | GAL1::CLN2-HA TRP1 AmprCEN | 18 |
| KL002 | GAL1::CLN3-HA TRP1 AmprCEN | 18 |
| KL038-1 | cdc28::HIS3 TRP1 Ampr KanrCEN | This study |
| KL039-1 | cdc28-csr1 URA3 Ampr KanrCEN | This study |
| KL045 | cdc28-csr3 TRP1 AmprCEN | This study |
| KL046 | cdc28-csr19 TRP1 AmprCEN | This study |
| KL047 | cdc28-csr35 TRP1 AmprCEN | This study |
| KL048 | cdc28-csr41 TRP1 AmprCEN | This study |
| KL049-1 | cdc28::CamrTRP1 AmprCEN | This study |
| KL050-1 | cdc28-HA::CamrTRP1 AmprCEN | This study |
| KL051 | CLN2-3P URA3 Ampr KanrCEN | This study |
| KL052 | cln2-K129A-3P URA3 Ampr KanrCEN | This study |
| KL053 | cln2-E183A-3P URA3 Ampr KanrCEN | This study |
| KL054 | cdc28-csr1 TRP1 AmprCEN | This study |
| KL055 | cdc28-csr3-HA TRP1 AmprCEN | This study |
| KL056 | cdc28-csr19-HA TRP1 AmprCEN | This study |
| KL057 | cdc28-csr35-HA TRP1 AmprCEN | This study |
| KL058 | cdc28-csr41-HA TRP1 AmprCEN | This study |
| KL059 | cdc28-rsc1-HA TRP1 AmprCEN | This study |
| KL060 | cdc28-rsc5-HA TRP1 AmprCEN | This study |
| KL061 | cdc28-rsc13-HA TRP1 AmprCEN | This study |
| KL062 | cdc28-rsc15-HA TRP1 AmprCEN | This study |
| KL063 | cdc28-rsc1 TRP1 AmprCEN | This study |
| KL064 | cdc28-rsc5 TRP1 AmprCEN | This study |
| KL065 | cdc28-rsc13 TRP1 AmprCEN | This study |
| KL066 | cdc28-rsc15 TRP1 AmprCEN | This study |
| KL075 | cdc28-L61S TRP1 AmprCEN | This study |
| KL077 | cdc28-L61S TRP1 AmprCEN | This study |
| KL078 | cdc28-L61S-HA TRP1 AmprCEN | This study |
| KL080 | cdc28-L61S-HA TRP1 AmprCEN | This study |
| KL081 | cdc28-K187E TRP1 AmprCEN | This study |
| KL082 | cdc28-K187E TRP1 AmprCEN | This study |
| KL084 | cdc28-K187E-HA TRP1 AmprCEN | This study |
| KL085 | cdc28-K187E-HA TRP1 AmprCEN | This study |
| RD47 | CDC28 TRP1 AmprCEN | 9 |
| RD58 | cdc28-T169A TRP1 AmprCEN | P. Sorger |
| KL019 | cdc28-T169A-HA TRP1 AmprCEN | This study |
| RS414 | TRP1 AmprCEN | 31 |
| RS416 | URA3 AmprCEN | 31 |
| SF19 | CDC28-HA TRP1 AmprCEN | 9 |
| TU10 | trp1::URA3/Kanr | 7 |
Yeast strains.
All the strains used in this study are described in Table 2. All strains were congenic with BF264-15D (MATa trp1 leu2 ura3 ade1 his2) (29). The strains were constructed and analyzed by standard genetic methods. DNA transformation was by the lithium acetate method. The cdc28::HIS3 disruption in strain 2198-3A-2a was constructed as follows. pKL038-1 (cdc28::HIS3/TRP1) was digested with XhoI and SacII, releasing a fragment containing the disrupted cdc28::HIS3 gene, and the digest was used to transform strain BOY760 containing pKL039-1. Yeast clones carrying the disrupted cdc28::HIS3 gene were identified by their His+ Trp− FOA− phenotype. The FUS1::HIS3, far1::URA3, leu2::LEU2::GAL1::CLN2, and leu2::LEU2::GAL1::CLN3 gene constructs, cln gene deletion alleles, and cdc28-13 mutant allele have been described previously (4, 5, 28, 32).
TABLE 2.
Strains used in this study
| Strain | Genotype |
|---|---|
| BOY1395 | MATa bar1 far1::URA3 HIS3 |
| BOY1396 | MATa bar1 far1::URA3 GAL1::CLN2 HIS3 |
| BOY916 | MATa bar1 far1::URA3 FUS1::HIS3 |
| BOY915 | MATa bar1 far1::URA3 FUS1::HIS3 leu2::LEU2::GAL1::CLN2 |
| 2198-3A | MATa bar1 TRP1::GAL1::CLN2 |
| 2180-1B | MATa bar1 csr1-1 LEU2::GAL1::CLN2 |
| 1607-2D | MATa bar1 cln1-del cln2-del cln3-del leu2::LEU2::GAL1::CLN3 |
| 1706-2B | cln1-del cln2-del cln3-del cdc28-csr1 leu2::LEU2::GAL1::CLN3 |
| 1255-5C | MATa bar1 CLN1 CLN2 CLN3 |
| 2195-13B | MATa bar1 CLN1 CLN2 CLN3 cdc28-csr1 |
| 2192-2B | MATa bar1 CLN1 CLN2 CLN3 leu2::LEU2::GAL1::CLN2 FUS1::HIS3 HIS2 |
| 2192-2C | MATa bar1 CLN1 CLN2 CLN3 cdc28-csr1 leu2::LEU2::GAL1::CLN2 FUS1::HIS3 HIS2 |
| 2112-10C | MATa bar1 cln1 cln2 CLN3 cdc28-13 leu2::LEU2::GAL1::CLN2 |
| 1227-3C | MATa cln1 CLN2 cln3 cdc28-13 pGAL1::CLN3/URA3 |
| BOY760 | MATa CLN1 CLN2 CLN3 leu2::LEU2::GAL1::CLN2 |
| 2198-3A-2a | MATa CLN1 CLN2 CLN3 leu2::LEU2::GAL1::CLN2 cdc28::HIS3 pKL039-1 |
| 2195-3A | MATa bar1 CLN1 CLN2 CLN3 cdc28-csr1 leu2::LEU2::GAL1::CLN2-HA |
| 2180-14A-1a | MATa bar1 CLN1 CLN2 CLN3 cdc28-csr1 far1::URA3 leu2::LEU2::GAL1::CLN2 |
Growth conditions.
Cells were grown in YEP medium (1% Difco yeast extract, 2% Difco Bacto Peptone) containing 2% glucose (YEPDex) or 3% galactose (YEPGal) as the carbon source. In experiments where the plasmids were retained, synthetic dropout medium (SC) containing 2% glucose (SCDex) or 3% galactose (SCGal) was used. The SC was prepared as described previously (25). For growth on solid media, 2% Difco agar was added. In experiments designed to select against the functional URA3 gene, 5-fluoroorotic acid (FOA) plates containing 6.7 g of Difco yeast nitrogen base complete (with amino acids and ammonium sulfate) per liter, 1 g of FOA powder per liter, 2% Difco agar, 12 mg of uracil per liter, and the remaining amino acids in the concentrations described by Ausubel et al. (2) were used. As a carbon source, 2% glucose (FOADex) or 3% galactose (FOAGal) was added.
Mutagenesis.
Random mutagenesis of CDC28 was performed by PCR with Taq polymerase, as described previously (17). In all cases, all four deoxynucleoside triphosphates were present at equal molar amounts of 1 mM each. In some cases, MnCl2, at a final concentration of either 0.25 or 0.5 mM, was added to increase the mutation frequency.
Gap repair to introduce mutant CDC28 into plasmids; untagging CDC28-HA.
A previously described (22) method of in vivo recombination, or gap repair, was used to introduce the original cdc28-csr1 allele into a plasmid, to generate the library of mutant CDC28 plasmids used in the screens, and to untag the HA-tagged CDC28 alleles. To introduce the cdc28-csr1 allele into a plasmid (pKL054), the cdc28-csr1 coding sequence was amplified by PCR with a genomic DNA preparation from the 2195-13B strain as the template. The amplification used the seq#1-5′CDC28 primer (about 60 bp 5′ of the start of the coding sequence and about 140 bp 5′ of the AflII site within the CDC28 coding sequence), and the seq#4-3′CDC28 primer (about 60 bp 3′ of the end of the coding sequence and about 300 bp 3′ of the ClaI site within the CDC28 coding sequence). The resulting 1,022-bp fragment was cotransformed with AflII-ClaI-digested (or gapped) and gel-purified pRD47 (CDC28/TRP1) into strain 2112-10C. Transformants were selected on defined medium lacking tryptophan and screened for the appropriate Csr mutant phenotype. The plasmids were isolated from the transformants, and two independently isolated plasmids were sequenced, each containing the single Q188P mutation. To generate the library of mutant CDC28 plasmids, the wild-type CDC28 allele was amplified by error-prone PCR (17). The primers used in the amplification reactions were at least 100 bp 5′ or 3′ of the AflII and ClaI sites in the CDC28 coding region, respectively. Then the amplified fragments were cotransformed with AflII-ClaI-digested and gel-purified pSF19 (CDC28-HA/TRP1) into strain 2112-10C (cdc28-csr screen) or 2198-3A-2a (cdc28-rsc screen). Transformants were selected on defined medium lacking tryptophan, and screened for appropriate phenotypes. To untag the pSF19-based HA-tagged CDC28 plasmids recovered in these screens, the HA-tagged plasmid constructs were digested with KpnI to release the mutant CDC28 coding sequence; KpnI cuts once upstream of the AflII site (within the polylinker of the plasmid) and once approximately 130 bp 3′ of the ClaI site (within the coding sequence of the gene). For all mutant alleles, all the mutations found within the HA-tagged constructs were included within this KpnI fragment. For the cdc28-csr allele untagging, mutant KpnI fragments were cotransformed with AflII-ClaI-digested and gel-purified pRD58 (cdc28-T169A/TRP1) containing an untagged and mutated version of the CDC28 gene (used to reduce the functional background). For the cdc28-rsc allele untagging, AflII-ClaI-digested pKL049-1, containing a Camr-disrupted version of pRD58, was used instead of pRD58 to further reduce the background. For all mutant alleles, except for cdc28-rsc15, which contained two mutations downstream of the ClaI site, all the mutations were included within the AflII-ClaI gap region. Transformants were selected on defined medium lacking tryptophan and screened for the appropriate mutant phenotype. Plasmids were isolated and tested by digestion and retransformation. The untagged allele of cdc28-rsc15 was sequenced to ensure the presence of mutations downstream of the ClaI site.
Immunoprecipitation, immunoblot analysis, and protein kinase assay.
The immunoprecipitation, immunoblot analysis, and protein kinase assay procedures were adapted from those of Cross and Blake (8) and Tyers et al. (35, 36) and have been described previously (18). For experiments analyzing Cln2-HA or Clb2-HA, the protocol used was described previously (18). For experiments analyzing Cdc28-HA, or bulk kinase, the protocol was as described, except that in the experiments analyzing the bulk kinase of the cdc28-csr alleles, the cells were grown overnight at 38 instead of 30°C and kinase assays were performed at 38 instead of 30°C. For the experiment analyzing Cln3-HA, the following modifications were made to optimize Cln3-Cdc28 association: following filtration, the cells were resuspended in 250 μl of LSHNN extraction buffer (10 mM HEPES [pH 7.5], 50 mM NaCl, 0.1% Nonidet P-40, 10% glycerol) instead of TNN extraction buffer; following incubation with protein A-agarose (Repligen), the immunoprecipitates were washed with LSHNN extraction buffer three times for 1 min each and washed with HNN extraction buffer (10 mM HEPES [pH 7.5], 250 mM NaCl, 0.1% Nonidet P-40, 10% glycerol) once for 2 min.
RESULTS
Isolation of a mutation blocking Cln2 regulation of the mating-factor pathway.
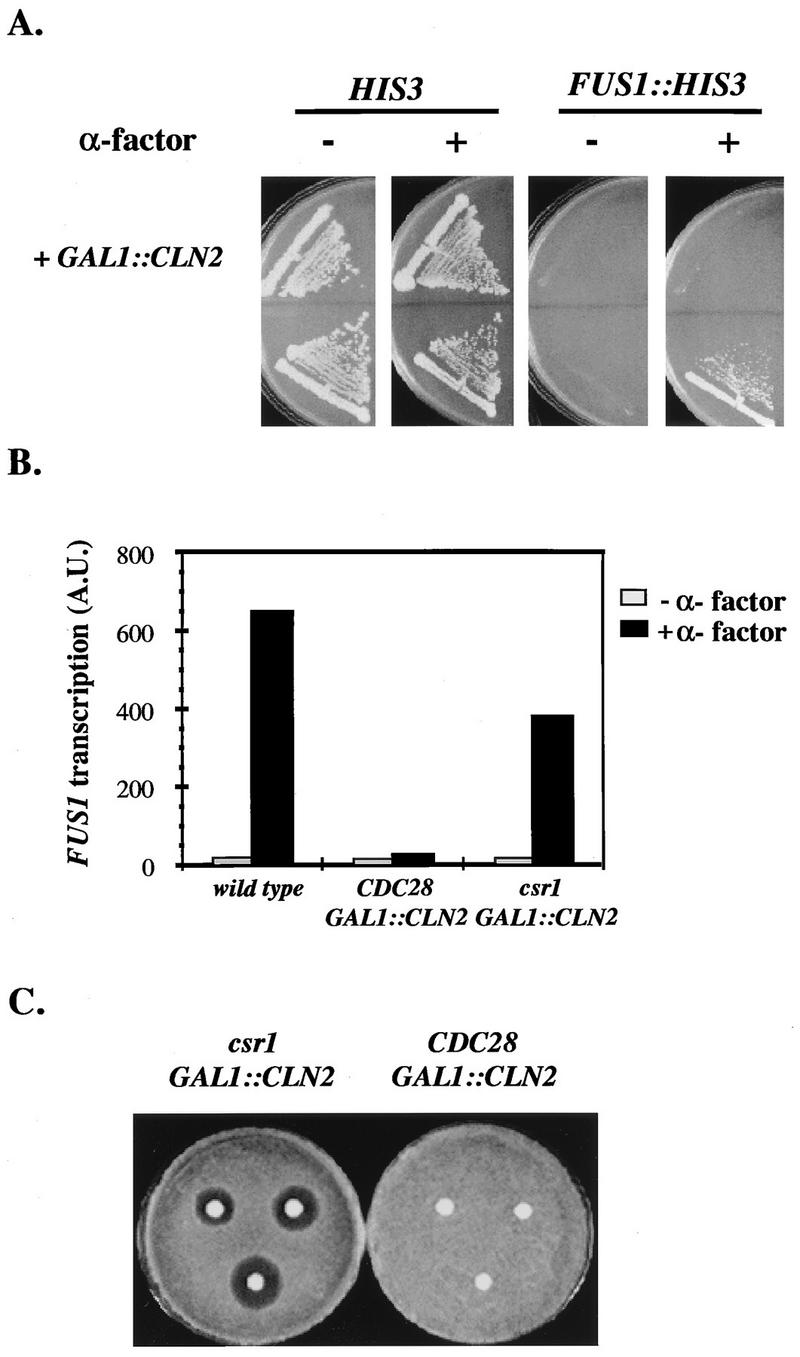
Overexpression of CLN2 from the GAL1 promoter results in repression of mating-pheromone signal transduction and resistance to mating-pheromone-induced cell cycle arrest (25). We devised a genetic screen to identify factors involved in the Cln2-mediated inhibition of the mating-pheromone pathway. We constructed a strain of the genotype far1 FUS1::HIS3 MATa bar1 (Fig. 1A). The FUS1 promoter is strongly induced by the mating-pheromone pathway (21, 34), and the FUS1::HIS3 fusion is regulated similarly (32). Thus, on medium lacking histidine and containing aminotriazole, the growth of this strain is mating-pheromone dependent (Fig. 1A, bottom right panel). This strain is able to form colonies on mating factor because of the far1 mutation. Introduction of GAL1::CLN2 into this strain eliminated colony formation on galactose-histidine plates containing mating factor because GAL1::CLN2 inhibited the mating-pheromone pathway, and thus FUS1::HIS3 transcription, making the strain phenotypically His− (Fig. 1A, top right panel). To identify mutants that were defective in GAL1::CLN2-mediated inhibition of the mating-pheromone pathway, we screened for mutant clones that were His+ when grown on galactose plates containing mating factor. Most of the His+ mutant clones that were obtained were Leu−, indicative of loss of the LEU2::GAL1::CLN2 cassette. The Leu+ mutant clones were analyzed by Northern blot analysis for both CLN2 expression and FUS1 transcriptional induction. Most of the Leu+ derivatives expressed elevated levels of endogenous FUS1 transcript constitutively (data not shown). These probably represented clones with constitutive activation of the mating-pheromone pathway similar to the ones described by Stevenson et al. (32). We did not analyze these mutants further. One mutant exhibited essentially wild-type activation of FUS1 transcription by mating-factor treatment, despite continued high expression of the CLN2 transcript from the GAL1 promoter (Fig. 1B). This mutant also suppressed the hyperpolarization phenotype, or elongated buds, normally found in cells overexpressing CLN2 (19). The mutant strain was crossed to a MATα FAR1 FUS1::HIS3 LEU2::GAL1::CLN2 strain and analyzed by tetrad analysis. The mutation, as determined by its hyperpolarization phenotype, segregated 2:2 (i.e., as a single gene) that recombined freely with the LEU2::GAL1::CLN2 cassette. In FAR1 LEU2::GAL1::CLN2 segregants from the cross, the presence of the mutation restored the sensitivity to mating factor of cells overexpressing CLN2, as measured by the halo assay (Fig. 1C). We named the mutation csr1-1, for cyclin-dependent signalling repression.
FIG. 1.
Isolation of the csr1-1 mutant; failure of repression by GAL1::CLN2 expression and restoration of mating-factor resistance to GAL1::CLN2 cells. (A) Construction of a strain in which CLN2 overexpression results in death due to inactivation of the mating-factor pathway. MATa bar1 far1 strains with and without a LEU2::GAL1::CLN2 construct and with and without an FUS1::HIS3 construct were grown in the absence and presence of mating factor on SCGal-His plates containing 50 mM aminotriazole. The strains used were BOY1396 (GAL1::CLN2 HIS3), BOY1395 (HIS3), BOY916 (GAL1::CLN2 FUS1::HIS3), and BOY915 (FUS1::HIS3). All the strains were MATa bar1 far1::URA3. (B) The csr1-1 mutation restores FUS1 inducibility by mating factor to GAL1::CLN2 cells. Cultures were grown on YEPGal medium, and part of the culture was treated with mating factor for 20 min. Samples were analyzed by Northern analysis for FUS1 and TCM1 transcript levels. The transcripts were quantitated with a PhosphorImager, and the FUS1 levels indicated were normalized for loading in the different lanes, as deduced from the TCM1 transcript levels. Values indicated in the histogram are arbitrary units (A.U.). The strains used were 1255-5C (wild type), 2198-3A (TRP1::GAL1::CLN2), and 2180-1B (csr1-1 LEU2::GAL1::CLN2). (C) The csr1-1 mutation restores mating-factor sensitivity to GAL1::CLN2 cells. Cultures were grown to saturation on YEPGal medium and spread on YEPGal plates. Sterile discs containing 15 μl of a 0.025, 0.05, or 0.1 mM alpha-factor dilution were placed on each lawn, clockwise from the top left. The plates were incubated for 2 days at 30°C. The strains used were 2198-3A (LEU2::GAL1::CLN2) and 2180-1B (csr1-1 LEU2::GAL1::CLN2).
csr1-1 is an allele of CDC28.
Overexpressed Cln2 requires Cdc28 to inactivate the mating-factor pathway (25). Thus, one explanation for the csr1-1 mutant phenotype would be that the mutation rendered Cln2-Cdc28 kinase defective. Indeed, Cln2 immunoprecipitates from csr1-1 mutant cells, compared to wild-type cells, had extremely low levels of associated kinase activity and Cdc28 protein (see below). We therefore tested whether the mutation could be in CDC28. In a cross to a cdc28-13 strain, the csr1-1 mutation segregated 2:2 in opposition to cdc28-13. The csr1-1 allele was complemented by CDC28 on a low-copy-number plasmid, providing further evidence that csr1-1 was an allele of CDC28. To confirm this, we amplified the CDC28 gene from a csr1-1 strain by using high-fidelity Vent polymerase to avoid the accumulation of errors and introduced the recovered CDC28 gene into a low-copy-number plasmid by gap repair (see Materials and Methods). The plasmids recovered by using the PCR product from the csr1-1 strain complemented cdc28-13 at 38°C but failed to complement csr1-1 for mating-factor resistance upon CLN2 overexpression. A plasmid containing the wild-type CDC28 gene complemented both cdc28-13 and csr1-1. These results mapped the csr1-1 mutation to CDC28, and we renamed the allele cdc28-csr1. The sequence of the cdc28-csr1 allele revealed only one nucleotide change: the Q188P substitution (Table 3). This position is predicted to be in the activation loop, distal to most regions involved in cyclin binding, based on the cyclin A-Cdk2 crystal structure (15).
TABLE 3.
cdc28-csr mutants used in this study
| Name | Mutation(s) | Bulk kinasea | Cln2 kinasea | Cln2 bindinga | Notes |
|---|---|---|---|---|---|
| CDC28 | None | 1.0 | 1.0 | 1.0 | |
| csr1 | Q188P | 0.8 | 0.003 | 0.02 | |
| csr3 | R43K, G101V, D147G, L165S | 0.9 | 0.002 | 0.1 | |
| csr19 | P30S, K41R, L61S | 1.0 | 0.3 | 1.0 | |
| L61S | L61S | 1.0 | 0.4 | 1.0 | |
| csr35 | I56T | 0.9 | 0.7 | 0.7 | I49 of CDK2; contacts cyclin A |
| csr41 | K187E, G192S, R208G | 1.0 | 0.002 | 0.1 | |
| K187E | K187E | 0.9 | 0.03 | 0.4 |
Average values relative to those for CDC28.
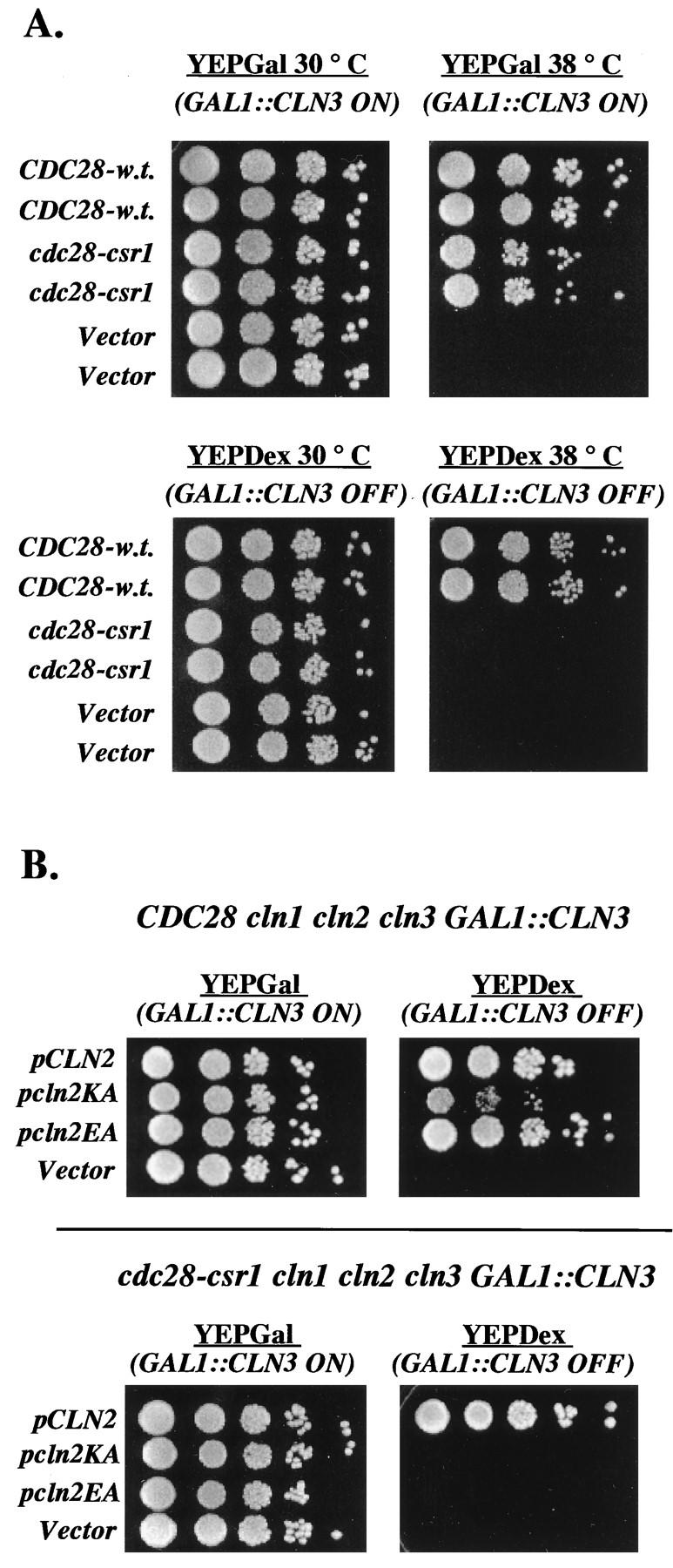
We tested cdc28-csr1 for additional phenotypes indicative of defective cyclin interaction. CLN1 CLN2 CLN3 cdc28-csr1 strains were about 35% larger in cell volume than were controls, suggesting a moderate in vivo defect in this aspect of G1 cyclin function. cln1 cln2 CLN3 cdc28-csr1 and cln1 CLN2 cln3 cdc28-csr1 strains were viable; however, the latter strains but not the former were temperature sensitive, suggesting that Cln2/Cdc28-csr1 function was less efficient than Cln3/Cdc28-csr1 function. This is surprising because in most assays Cln2 is much more potent than Cln3 (18), and it suggested specificity in the csr1 defect. The temperature sensitivity of cln1 CLN2 cln3 cdc28-csr1 strains was further analyzed in a cln1 CLN2 cln3 cdc28-13 GAL1::CLN3 strain, carrying plasmid copies of either CDC28 or cdc28-csr1 (Fig. 2A). Cells carrying the wild-type copy of CDC28 were viable at both 30 and 38°C (the permissive and restrictive temperatures for the cdc28-13 allele, respectively), independent of the expression of GAL1::CLN3. In contrast, cells carrying the cdc28-csr1 plasmid were viable at 38°C only when GAL1::CLN3 was expressed (Fig. 2A). cdc28-csr1 failed to show significant genetic interaction with disruption of the major mitotic B-type cyclin clb2: clb2 cdc28-csr1 double mutants were fully viable and temperature resistant and had only a moderately greater cell size than did clb2 single mutants (data not shown).
FIG. 2.
cdc28-csr1 is defective in genetic interaction with CLN2. (A) Temperature sensitivity of the cln1 CLN2 cln3 cdc28-csr1-1 strain. Strain 1227-3C (cln1 CLN2 cln3 cdc28-13 pURA3/GAL1::CLN3) was transformed with plasmids RD47 (CDC28), KL054 (cdc28-csr1), or RS414 (Vector). For each transformed strain, 10-fold serial dilutions were prepared for two independently isolated transformants, and 5 μl of each dilution was plated onto both a YEPGal plate (GAL1::CLN3 on) and a YEPDex plate (GAL1::CLN3 off) at both 30 and 38°C. The YEPGal plates were incubated for 3 days, and the YEPDex plates were incubated for 2 days. (B) Weakened alleles of CLN2 fail to support the viability of a cln1 cln2 cln3 cdc28-csr1 strain. Strain 1607-2B (CDC28 cln1 cln2 cln3 GAL1::CLN3) and strain 1706-2B (cdc28-csr1 cln1 cln2 cln3 GAL1::CLN3) were each transformed with plasmid KL051 (CLN2-3P), KL052 (cln2-K129A-3P), KL053 (cln2-E183A-3P), or RS416 (Vector). For each transformed strain, 10-fold serial dilutions were prepared, and 5 μl of each dilution was plated onto both a YEPGal plate (GAL1::CLN3 on) and a YEPDex plate (GAL1::CLN3 off). The plates were incubated at 30°C for 3 days.
We recently described CLN2 mutants with reduced function due to an inability to interact efficiently with Cdc28 (18). These mutants, cln2-K129A-3P and cln2-E183A-3P (each driven by the CLN3 promoter), were able to complement a cln1 cln2 cln3 strain; however, they were unable to complement a cln1 cln2 cln3 cdc28-csr1 strain (although both CLN3 and a wild-type CLN2 gene driven by the CLN3 promoter, CLN2-3P, were able to do so) (Fig. 2B and data not shown).
Thus, overall, these genetic results suggested a strong and relatively Cln2-specific functional defect associated with the csr1 mutant Cdc28.
Biochemical characterization of Cdc28-csr1-cyclin interaction.
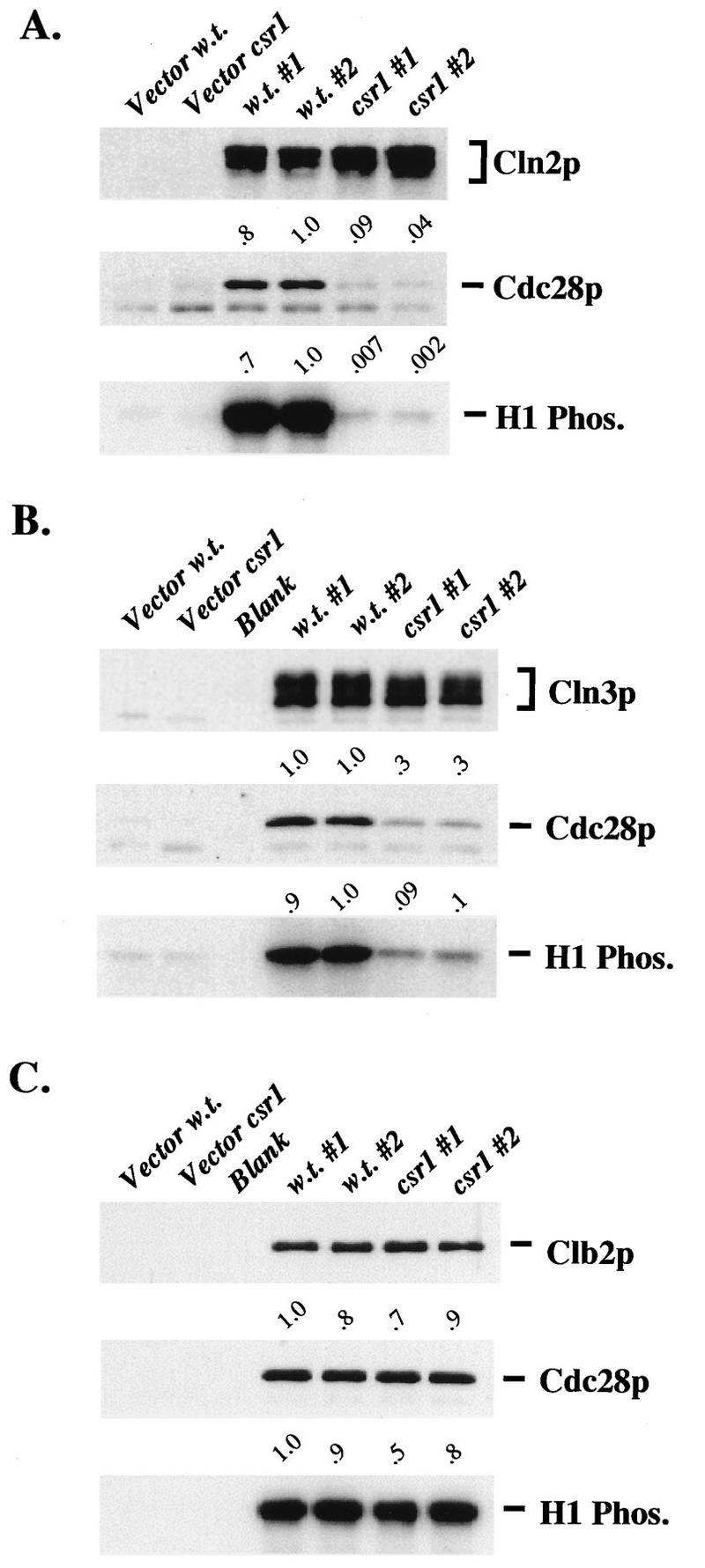
Since the genetic assays suggested specific defects in Cln2-Cdc28 function, we wanted to further analyze these defects biochemically. We immunoprecipitated HA-tagged Cln2 from the cdc28-csr1 strain and from the CDC28 strain. We detected, on average, about 400-fold less Cln2-associated histone H1 kinase activity from the cdc28-csr1 strain than from the CDC28 strain (Fig. 3A). This was correlated with about a 40-fold defect in coimmunoprecipitation of Cdc28-csr1; the small amount of Cdc28 coimmunoprecipitated appeared to have an additional defect in activation as a histone H1 kinase. We observed no defect in Cdc28-csr1 protein abundance relative to the wild type following purification of Cdc28 by immunoprecipitation or by p13 affinity chromatography (data not shown); thus, the defect in Cln2-Cdc28 interaction is not due to low Cdc28-csr1 protein levels. When we carried out a similar experiment with HA-tagged Cln3, we also observed a significant defect in Cdc28 binding and activity from the cdc28-csr1 strain, although the effect was reproducibly smaller than the effect on Cln2 (Fig. 3B). In sharp contrast, Clb2-associated Cdc28 and kinase activity, as well as bulk Cdc28-HA kinase activity, was near normal in immunoprecipitates from the cdc28-csr1 strain (Fig. 3C and data not shown).
FIG. 3.
Biochemical characterization of cyclin-dependent activation of Cdc28-csr1. (A) Comparison of Cln2-associated kinase activity, associated Cdc28 protein, and associated Cdc28 specific kinase activity in CDC28 wild-type and cdc28-csr1 cells. A CDC28 wild-type strain (1255-5C) and a cdc28-csr1 strain (2195-13B) were each transformed with plasmids RS414 (Vector) and KH100 (GAL1::CLN2-HA). Cultures of each transformant strain were grown overnight at 30°C in SCGal-Trp. Before extraction, log-phase cultures of each of the pKH100 transformants were split in half and processed in duplicate. Cellular protein was extracted with TNN buffer, HA-tagged Cln2 protein was isolated, and kinase assays were performed. Cln2-HA protein and associated Cdc28 protein were analyzed by immunoblotting. All the protocols are described in Materials and Methods. The anti-HA immunoblot, the anti-Cdc28 immunoblot, and the H1 kinase blot are shown. The lanes of each blot correspond (from left to right) to transformed strains 1255-5C pRS414, 2195-13B pRS414, 1255-5C pKH100, 1255-5C pKH100, 2195-13B pKH100, and 2195-13B pKH100. Values indicated above the anti-Cdc28 immunoblot were obtained by standardizing the associated Cdc28 protein levels to Cln2 protein levels. Values indicated above the H1 kinase blot were obtained by standardizing the histone H1 radioactivity to Cln2 protein levels. (B) Comparison of Cln3-associated kinase activity, associated Cdc28 protein, and associated Cdc28 specific kinase activity in CDC28 wild-type and cdc28-csr1 cells. This experiment is identical to the one described for panel A, with the following exceptions: strains 1255-5C and 2195-13B were transformed with plasmids RS414 (Vector) and KL002 (GAL1::CLN3-HA), cellular protein was extracted with LSHNN extraction buffer (see Materials and Methods), and Cln3-HA protein was analyzed instead of Cln2-HA protein. The lanes of each blot correspond (from left to right) to transformed strains 1255-5C pRS414, 2195-13B pRS414, blank lane, 1255-5C pKL002, 1255-5C pKL002, 2195-13B pKL002, and 2195-13B pKL002. (C) Comparison of Clb2-associated kinase activity, associated Cdc28 protein, and associated Cdc28 specific kinase activity in CDC28 wild-type and cdc28-csr1 cells. This experiment is identical to the one described for panel A, with the following exceptions: strains 1255-5C and 2195-13B were transformed with plasmids RS416 (Vector) and 143 (GAL1::CLB2-HA), each of the plasmids carries the URA3 gene, the cultures were grown overnight in SCGal-Ura, and Clb2-HA protein was analyzed instead of Cln2-HA protein. The lanes of each blot correspond (from left to right) to transformed strains 1255-5C pRS416, 2195-13B pRS416, blank lane, 1255-5C p143, 1255-5C p143, 2195-13B p143, and 2195-13B p143.
The striking defects in Cln2-associated Cdc28 and H1 kinase activity were observed under both the relatively stringent immunoprecipitation conditions of Tyers et al. (35) and the gentler conditions designed to optimize Cln3-Cdc28 binding and kinase activity (8, 12); using these conditions, we detected about 100-fold less Cln2-associated histone H1 kinase activity from the cdc28-csr1 strain than from the CDC28 strain (data not shown). It appears likely that the in vitro defect is more extreme than the in vivo defect, although the specificity (worst interaction with Cln2, better with Cln3, no defect with Clb2) is in agreement with that deduced from the genetic studies described above.
Isolation of additional cdc28 alleles with the Csr phenotype.
Identification of additional alleles with similar phenotypes to cdc28-csr1 could give information about the Cdc28-Cln2 binding interface or the residues important for Cdc28-Cln2-dependent functions. Therefore, we developed a method for random mutagenesis of CDC28 and recovery of mutated cdc28 genes with the same phenotype as cdc28-csr1: that is, complementation of CDC28 function for viability but not for Cln2-dependent mating-factor resistance. To do this, we mutagenized CDC28 by error-prone PCR and recovered the mutated genes on plasmids by cotransforming the mutant fragments together with gapped vector (see Materials and Methods) into a cln1 cln2 CLN3 GAL1::CLN2 cdc28-13 strain. We then screened for viability on glucose at 38°C (requiring complementation of cdc28-13 even in the absence of the major G1 cyclins CLN1 and CLN2), combined with tight sensitivity to mating factor on galactose medium (the Csr phenotype), indicating an inability to interact productively with overexpressed Cln2 to cause mating-factor resistance. We wanted to eliminate simple low-functioning alleles of CDC28; therefore, as a secondary screen, we tested histone H1 kinase activity from asynchronously cycling cultures (the CDC28 gene used for mutant isolation was tagged with the HA epitope, making this determination simple). This bulk Cdc28 histone H1 kinase activity is probably largely dependent on a mixture of Clb cyclins expressed in asynchronous culture (11). We selected four mutant CDC28 genes that showed vigorous growth on glucose medium at 38°C, tight alpha-factor sensitivity on galactose medium at 30 or 38°C, and >80% of wild-type bulk Cdc28 H1 kinase activity (Fig. 4A and 5A and data not shown).
FIG. 4.
New cdc28-csr alleles are defective in their genetic interaction with CLN2. (A) Halo assays for new csr alleles. The alleles were tested in a cdc28-13 cln1 cln2 GAL-CLN2 background; the viability of the lawn at 38°C indicates complementation of cdc28-13 cln1 cln2; mating-factor sensitivity indicates failure of effective CLN2-dependent function. Strain 2112-10C (cln1 cln2 CLN3 cdc28-13 GAL1::CLN2) was transformed with plasmids RS414 (Vector), SF19 (CDC28-HA), KL054 (cdc28-csr1), KL055 (cdc28-csr3-HA), KL056 (cdc28-csr19-HA), KL057 (cdc28-csr35-HA), and KL058 (cdc28-csr41-HA). Representative transformant strains were grown up overnight in SCGal-Trp medium. A 200-μl volume of culture was spread onto YEPGal plates and allowed to incubate at 30 or 38°C for about 4 h. Following incubation, sterile disks containing 15 μl of a 0.05, 0.1, or 0.2 mM alpha-factor dilution were placed on each lawn, clockwise from the top left. The plates were allowed to incubate at 30°C for 2 days or 38°C for 3 days. (B) New cdc28-csr alleles are defective in their interactions with weakened alleles of CLN2. Strain 1706-2B (cdc28-csr1 cln1 cln2 cln3 GAL1::CLN3) was first transformed with plasmids KL051 (CLN2-3P), KL052 (cln2KA), KL053 (cln2EA), or RS416 (Vector). Each of these transformed strains (described in the legend to Fig. 2B), was additionally transformed with plasmids SF19 (CDC28-HA), KL054 (cdc28-csr1), KL055 (cdc28-csr3-HA), KL056 (cdc28-csr19-HA), KL057 (cdc28-csr35-HA), KL058 (cdc28-csr41-HA), or RS414 (Vector). For each cotransformed strain, 10-fold serial dilutions were prepared, and 5 μl of each dilution was plated onto both a YEPGal plate (GAL1::CLN3 on) and a YEPDex plate (GAL1::CLN3 off). The plates were incubated at 30°C for 3 days.
FIG. 5.
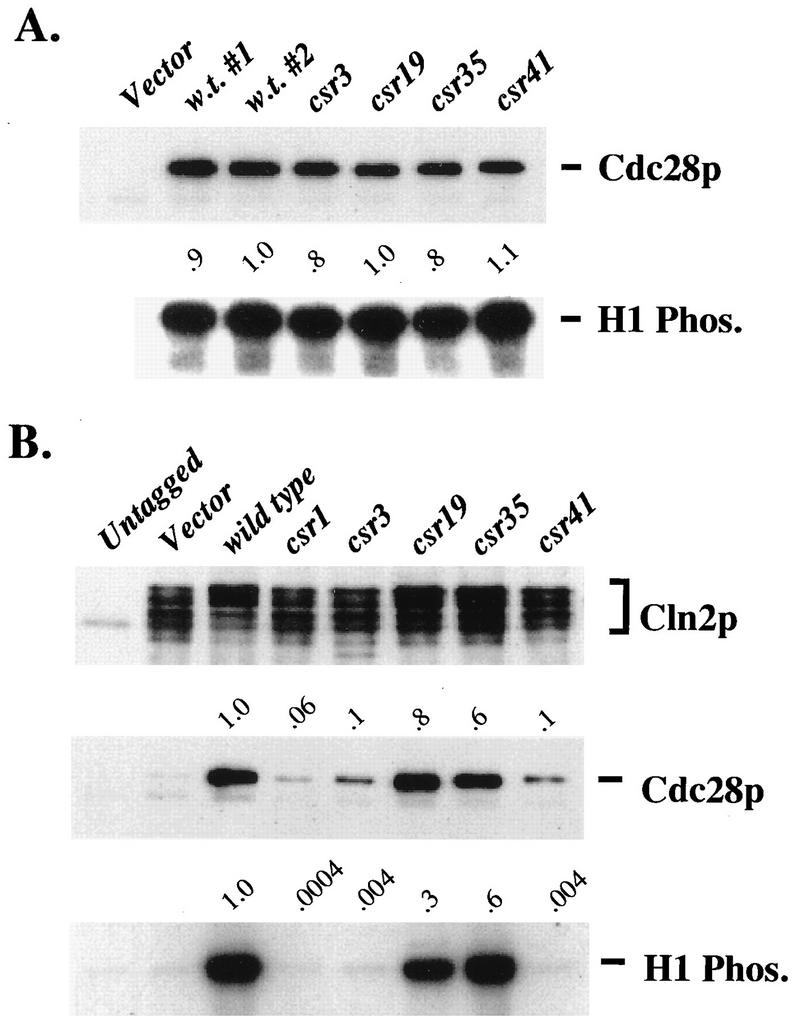
Total and Cln2-associated kinase assays and Cdc28 binding for new cdc28-csr alleles. (A) Comparison of total Cdc28 kinase activities for new cdc28-csr alleles. The wild-type strain 1255-5C was transformed with plasmids RS414 (Vector), SF19 (CDC28-HA), KL055 (cdc28-csr3-HA), KL056 (cdc28-csr19-HA), KL057 (cdc28-csr35-HA), or KL058 (cdc28-csr41-HA). Cultures of each transformant strain were grown overnight at 38°C in SCDex-Trp. HA-tagged Cdc28 protein was isolated, and kinase assays were performed at 38°C. Cdc28 protein levels were analyzed by immunoblotting. All the procedures are described in Materials and Methods. The anti-HA immunoblot and H1 kinase blot are shown. The lanes in each blot correspond (from left to right) to transformed strains 1255-5C pRS414, 1255-5C pSF19, 1255-5C pSF19, 1255-5C pKL055, 1255-5C pKL056, 1255-5C pKL057, and 1255-5C pKL058. Values indicated above the H1 kinase blot are the total H1 kinase activities relative to wild-type values, not standardized to Cdc28 protein levels. (B) Comparison of Cln2-associated kinase activity, associated Cdc28 protein, and associated Cdc28 specific kinase activity for new cdc28-csr alleles. Strain 2195-3A (cdc28-csr1 GAL1::CLN2-HA) was transformed with plasmids RS414 (Vector), RD47 (CDC28), KL054 (cdc28-csr1), KL045 (cdc28-csr3), KL046 (cdc28-csr19), KL047 (cdc28-csr35), or KL048 (cdc28-csr41). Strain 2193-13B (cdc28-csr1) was transformed with pRS414. Cultures of each transformant strain were grown overnight at 30°C in SCGal-Trp medium. HA-tagged Cln2 protein was isolated, and kinase assays were performed. Cln2-HA protein and associated Cdc28 protein were analyzed by immunoblotting. All the methods are described in Materials and Methods. The anti-HA immunoblot, anti-Cdc28 immunoblot, and H1 kinase blot are shown. The lanes of each blot correspond (from left to right) to transformed strains 2195-13B pRS414, 2195-3A pRS414, 2195-3A pRD47, 2195-3A pKL054, 2195-3A pKL045, 2195-3A pKL046, 2195-3A pKL047, and 2195-3A pKL048. Values indicated above the anti-Cdc28 immunoblot were obtained by standardizing associated Cdc28 protein levels to Cln2 protein levels. Levels from the untagged culture and vector control (representing the level of Cln2-associated Cdc28-csr1 protein) have been subtracted. Values indicated above the H1 kinase blot were obtained by standardizing histone H1 radioactivity to Cln2 protein levels. Levels from the untagged culture and vector control (representing the level of H1 phosphorylation by the Cdc28-csr1 in the strain) have been subtracted.
These mutant cdc28 genes resulted in tight mating-factor sensitivity, essentially comparable to that observed with cdc28-csr1 (Fig. 4A). Additionally, each of the mutant cdc28 genes demonstrated defective genetic interactions with the weakened alleles of CLN2 (Fig. 4B). However, the new cdc28-csr alleles were clearly less defective than cdc28-csr1. As shown in Fig. 4B, cdc28-csr alleles 19, 35, and 41 were less defective than cdc28-csr1 in their genetic interactions with the weakened alleles of CLN2. Additionally, each of the new cdc28-csr alleles, but not cdc28-csr1, was competent at rescue of a cln1 CLN2 cln3 cdc28-13 strain at 38°C (i.e., they could function for Start with CLN2 as their sole G1 cyclin) (data not shown). Furthermore, each of the new cdc28-csr alleles, but not cdc28-csr1, could confer mating-factor resistance in a different GAL1::CLN2 strain additionally containing endogenous CLN1 and CLN2 (data not shown).
Biochemical and molecular characterization of cdc28-csr alleles.
We tested the new csr alleles for Cln2 binding and kinase activation. To do this, we first untagged each of the cdc28-csr alleles (see Materials and Methods) and then introduced the untagged alleles into plasmids into a cdc28-csr1 GAL1::CLN2-HA strain. In extracts from this strain, Cln2 could be immunoprecipitated with the HA tag, but no significant Cdc28 protein or kinase activity coprecipitated due to the csr1 mutation. Therefore, we could attribute all Cdc28 binding and kinase activity to the introduced plasmid cdc28-csr gene. For two of the alleles, we observed results similar to those for cdc28-csr1: low levels of Cdc28 protein and kinase activity coimmunoprecipitated with overexpressed Cln2 (Fig. 5B). For two of the alleles, in contrast, we observed high levels of Cdc28 protein and kinase activity coimmunoprecipitated with Cln2 (Fig. 5B). These four mutants did not differ significantly in the tightness of their mating-factor sensitivity phenotype, although cdc28-csr19 and cdc28-csr35 were less defective than cdc28-csr3 and cdc28-csr41 in their genetic interactions with the weakened alleles of cln2 (Fig. 4B).
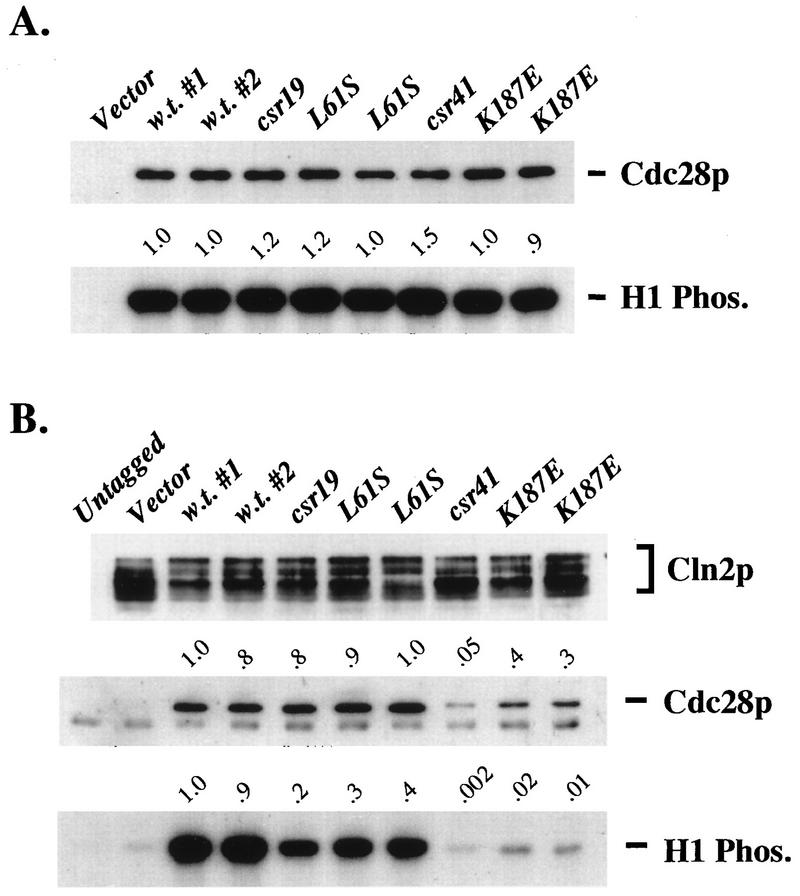
We sequenced these mutant cdc28 genes (Table 3). Three of them had multiple amino acid substitutions. One of the mutations in cdc28-csr41, K187E, was at a neighboring position to the cdc28-csr1 amino acid substitution, Q188P, and these mutant Cdc28 proteins shared the property of defective interaction with Cln2. We therefore constructed K187E as a single mutant and found that this single mutation largely (although not completely) reconstituted the genetic and biochemical behavior of cdc28-csr41 (Fig. 6 and 7). Therefore, this region of the Cdc28 T-loop may be important for the Cdc28-Cln2 biochemical interaction. One mutant, cdc28-csr35, contained only the single mutation I56T. This residue is conserved in Cdk2. It is the “I” in the so-called PSTAIRE helix. In the cyclin A-Cdk2 crystal structure, this hydrophobic isoleucine side chain inserts into a hydrophobic pocket in cyclin A (15). Surprisingly, this mutant exhibits near-wild-type Cln2-associated and “bulk” (probably mainly Clb-associated) histone H1 kinase activity. Because cdc28-csr19, with similar genetic and biochemical behavior to cdc28-csr35, had as one of its mutations L61S, near to I56 (both predicted to be in the PSTAIRE helix), we constructed the L61S single mutant and found that it largely reconstituted the cdc28-csr19 phenotype (Fig. 6 and 7). Therefore, the correct sequence or conformation of the PSTAIRE helix may be important for the function of the Cdc28-Cln2 complex although not for effective biochemical interaction (binding and kinase activation).
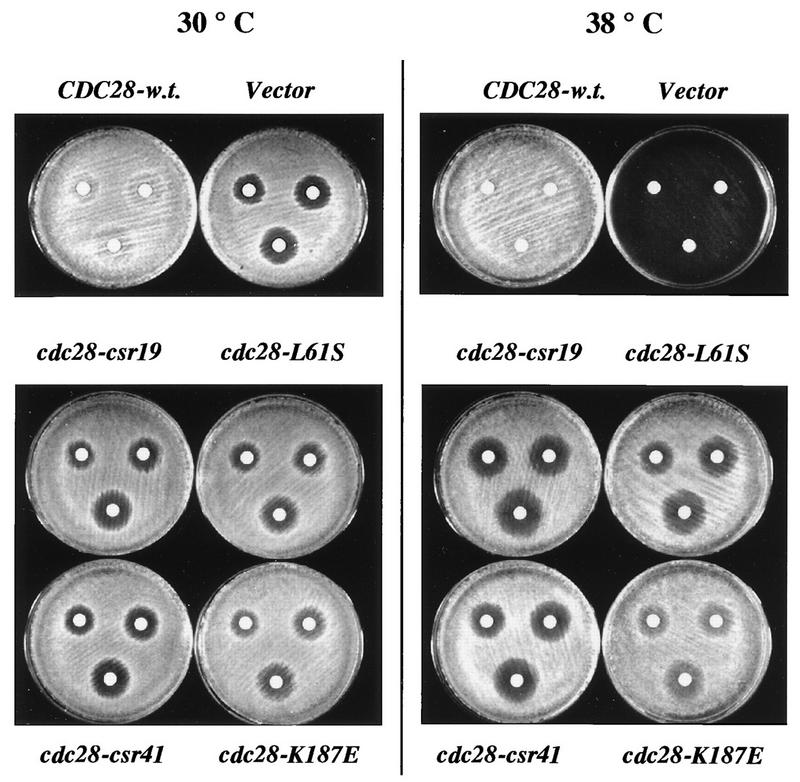
FIG. 6.
cdc28-L61S and cdc28-K187E single-mutant alleles are defective in their genetic interaction with CLN2. Halo assays for single-mutant alleles, tested in a cdc28-13 cln1 cln2 GAL-CLN2 background, are shown. Strain 2112-10C was transformed with plasmids SF19 (CDC28-HA), RS414 (Vector), KL056 (cdc28-csr19-HA), KL078 (cdc28-L61S-HA), KL058 (cdc28-csr41-HA), and KL084 (cdc28-K187E-HA). Pooled transformants were grown overnight in SCGal-Trp medium, and halo assays were prepared as described in the legend to Fig. 4. The plates were allowed to incubate for 1 day.
FIG. 7.
Total and Cln2-associated kinase assays and Cdc28 binding for cdc28-L61S and cdc28-K187E alleles. (A) Comparison of total Cdc28 kinase activities. The wild-type strain 1255-5C was transformed with plasmids RS414 (Vector), SF19 (CDC28-HA), KL056 (cdc28-csr19-HA), KL078 (cdc28-L61S-HA), KL080 (cdc28-L61S-HA), KL058 (cdc28-csr41-HA), KL084 (cdc28-K187E-HA), or KL085 (cdc28-K187E-HA). Samples were prepared as described in the legend to Fig. 5A and in Materials and Methods. The anti-HA immunoblot and H1 kinase blot are shown. The lanes on each blot correspond to transformed strains 1255-5C pRS414, 1255-5C pSF19, 1255-5C pSF19, 1255-5C pKL056, 1255-5C pKL078, 1255-5C pKL080, 1255-5C pKL058, 1255-5C pKL084, and 1255-5C pKL085. The values indicated above the H1 kinase blot are the total H1 kinase activities relative to wild-type values, not standardized to Cdc28 protein levels. (B) Comparison of Cln2-associated kinase activity, associated Cdc28 protein, and associated Cdc28 specific kinase activity for new cdc28-csr alleles. Strain 2195-3A (cdc28-csr1 GAL1::CLN2-HA) was transformed with plasmids RS414 (Vector), RD47 (CDC28), KL046 (cdc28-csr19), KL075 (cdc28-L61S), KL077 (cdc28-L61S), KL048 (cdc28-csr41), KL081 (cdc28-K187E), or KL082 (cdc28-K187E). Strain 2193-13B (cdc28-csr1) was transformed with pRS414. Samples were prepared as described in the legend to Fig. 5B and in Materials and Methods. The anti-HA immunoblot, anti-Cdc28 immunoblot, and H1 kinase blot are shown. The lanes of each blot correspond to transformed strains 2195-13B pRS414, 2195-3A pRS414, 2195-3A pRD47, 2195-3A pRD47, 2195-3A pKL046, 2195-3A pKL075, 2195-3A pKL077, 2195-3A pKL048, 2195-3A pKL081, and 2195-3A pKL082. Values indicated above the anti-Cdc28 immunoblot and the H1 kinase blot were obtained as described in the legend to Fig. 5B.
Screen for cdc28 mutants defective for viability but not for Cln2-dependent mating-factor resistance (Rsc phenotype).
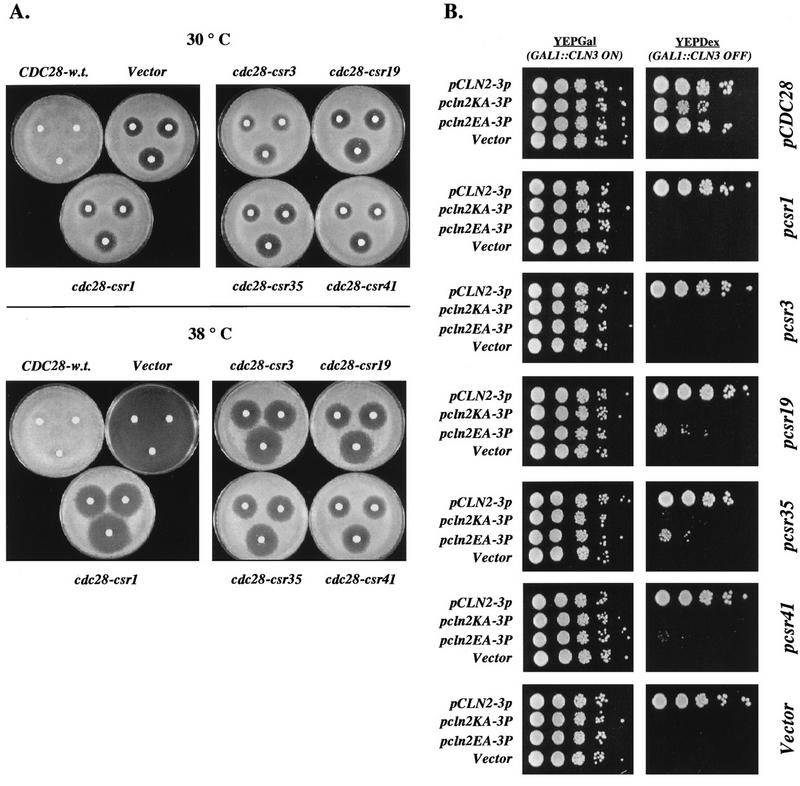
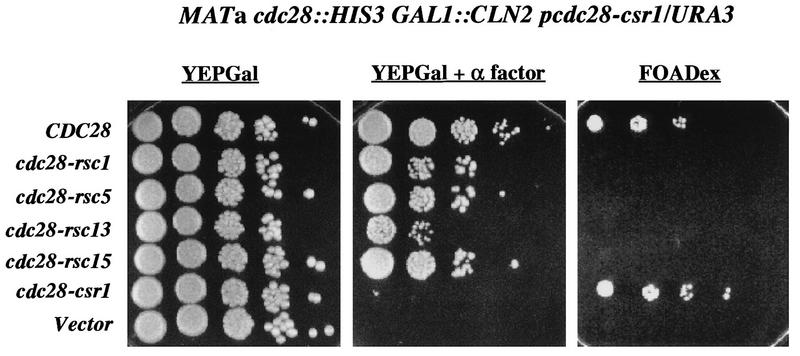
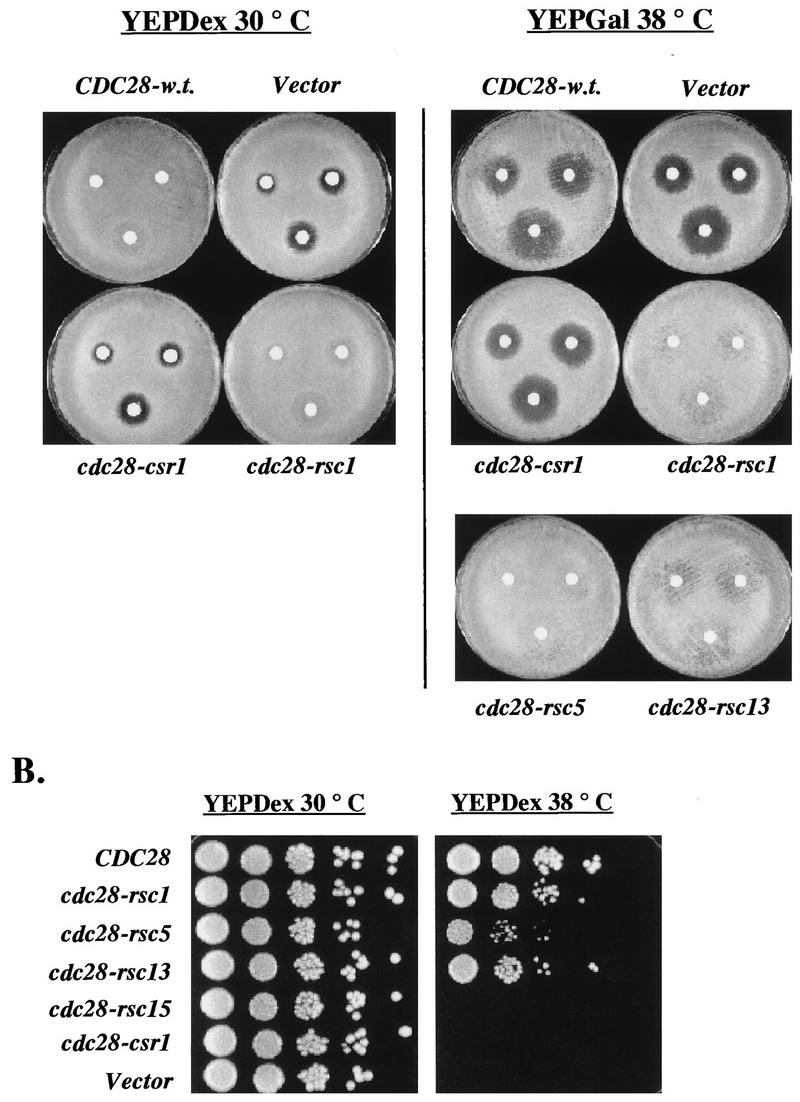
The Csr screen described above identified cdc28 mutants that were fully competent for viability (complementation of cdc28-13 at 38°C) but did not support Cln2-dependent inhibition of the mating-factor pathway. We were concerned that this genetic assay might simply reflect greater mutational or intrinsic sensitivity of the mating-factor pathway assay and/or Cln2 association, so that subtle defects in Cdc28 function overall might be reflected in the apparently highly specific Csr phenotype. In an attempt to address this question, we performed the converse screen, looking for cdc28 mutants that could support Cln2-dependent inhibition of the mating-factor pathway but could not complement cdc28 for viability (resistant to mating factor, still cdc28-). To do this, we constructed a strain of genotype GAL1::CLN2 cdc28::HIS3 pcdc28-csr1/URA3 ura3. This strain was viable due to complementation of the cdc28::HIS3 disruption by cdc28-csr1 on the URA3-containing plasmid. However, because of the Csr phenotype of the cdc28-csr1 allele, this strain was alpha-factor sensitive on galactose medium. Additionally, because this strain was dependent upon the cdc28-csr1/URA3 plasmid for viability, it was sensitive to FOA, a compound lethal to URA3 strains (3). When this strain was transformed with wild-type CDC28 on a TRP1 plasmid, it became alpha-factor resistant on galactose medium and FOA resistant as expected; when transformed with cdc28-csr1, it remained alpha-factor sensitive on galactose medium yet became FOA resistant, as expected (Fig. 8). We transformed the strain with the mutated CDC28 library and screened for clones that were alpha-factor resistant on galactose medium (indicating an effective Cln2-Cdc28 interaction) yet FOA sensitive (perhaps indicating a defective Clb-Cdc28 interaction). We identified four mutant plasmids that conferred this phenotype (Fig. 8). These mutants were characterized quantitatively by serial dilution on control YEPGal medium, on YEPGal medium containing alpha-factor, and on FOA medium. cdc28-rsc mutants 1, 5, and 15 gave alpha-factor resistance with about a 10-fold-lower colony-forming efficiency than that of wild-type CDC28; the colony-forming efficiency of cdc28-rsc13 was reduced by 100-fold, and the colonies were smaller. Vector and cdc28-csr1 on a plasmid were completely negative. Neither the wild-type CDC28 nor mutant cdc28-rsc alleles allowed growth on YEPDex medium containing alpha-factor (data not shown), suggesting that the mating-pheromone resistance conferred by these alleles is dependent on the overexpression of CLN2. On FOA, the cdc28-rsc alleles gave no viable colonies whereas both CDC28 and cdc28-csr1 were equally effective. We transferred the plasmids to a GAL1::CDC28 strain (38) to confirm by a different assay that they failed to complement a cdc28 null allele. Transformants were streaked on glucose medium to inactivate GAL1::CDC28. While transformants with wild-type CDC28 were fully viable on glucose medium, transformants with these mutant cdc28 genes were inviable on glucose medium (data not shown), confirming the conclusion reached on the basis of their FOA-sensitive phenotype (Fig. 8).
FIG. 8.
cdc28-rsc alleles complement cdc28-csr1 for mating-factor resistance but do not complement a cdc28 null for viability. (A) Serial dilution assays on alpha-factor and FOA. Strain 2198-3A-2a (cdc28::HIS3 pURA3/cdc28-csr1) was transformed with plasmids SF19 (CDC28-HA), KL059 (cdc28-rsc1-HA), KL060 (cdc28-rsc5-HA), KL061 (cdc28-rsc13-HA), KL062 (cdc28-rsc15-HA), KL054 (cdc28-csr1), or RS414 (Vector). Transformants were grown on SCDex-Trp-Ura plates. For each transformed strain, 10-fold serial dilutions were prepared, and 5 μl of each dilution was plated onto a YEPGal plate, a YEPGal plate containing 0.3 mM alpha-factor, and an FOADex plate (3). The plates were allowed to incubate at 30°C for 3 days. A representative set of transformants is shown.
We determined the mutations in the cdc28-rsc mutants (Table 4). Several of these mutations disrupt residues likely to be essential for protein kinase catalytic activity. In cdc28-rsc13, Glu 58 is mutated to Gly. This residue (Glu 51 of CDK2) is conserved in all eukaryotic protein kinases and belongs to a triad of catalytic-site residues. These residues (Lys 33, Glu 51, and Asp 145 of CDK2) are involved in ATP phosphate orientation and magnesium coordination and are likely to be critical for catalysis (15, 41). In cdc28-rsc1, Arg 159 is mutated to Gly. This residue (Arg 150 of CDK2), together with two additional arginine residues (Arg 50 and Arg126 of CDK2), forms part of the basic pocket that binds the phosphorylated threonine residue. The arginine-phosphate interactions are thought to stabilize the region of the phosphorylation site (30). A positive residue in this position is highly conserved in kinases with phosphorylated activation loops (16), including all cyclin-dependent kinases and protein kinase A. In cdc28-rsc5, Arg 178 in the P+1 loop is mutated to Ser. This arginine (Arg 169 of CDK2) is conserved in proline-directed kinases (Cdks and mitogen-activated protein kinase) and is speculated to interact with residues of the substrate in the vicinity of the proline C-terminal to the Ser/Thr phosphoacceptor residue (30). We were surprised to detect biological activity by the mating-factor resistance assay in CDC28 genes mutated at these presumably critical residues. cdc28-rsc15 was multiply mutant.
TABLE 4.
cdc28-rsc mutants used in this study
| Name | Mutation(s) | Bulk kinasea | Cln2 kinasea | Cln2 bindinga | Notes |
|---|---|---|---|---|---|
| CDC28 | None | 1.0 | 1.0 | 1.0 | |
| rsc1 | R159G | 0.002 | 0.02 | 0.08 | R150 of CDK2; one of three R’s that neutralize phosphate group; contacts cyclin A |
| rsc5 | R178S | 0.1 | 0.009 | 0.5 | R169 of CDK2; likely to H-bond to substrate near P of S/TP consensus |
| rsc13 | E58G | 0.01 | 0.001 | 0.6 | E51 of CDK2; residue of catalytic triad |
| rsc15 | D92N, P164T, S216R, V227A, A234T | 0.4 | 0.007 | 0.07 |
Average values relative to those for CDC28.
Biochemical characterization of cdc28-rsc kinase activity: total and Cln2-associated.
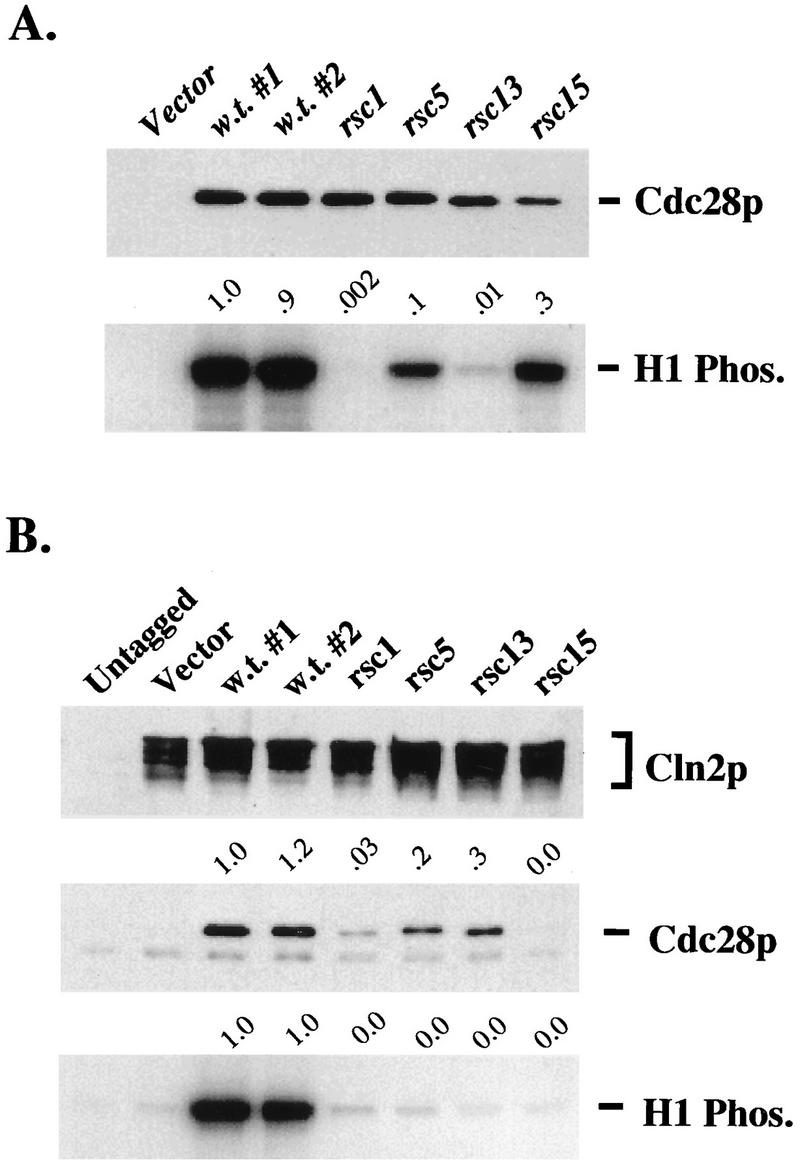
We tested the total (bulk) kinase activity associated with the Cdc28-rsc proteins in asynchronous culture, taking advantage of the HA epitope on the mutant Cdc28 protein to avoid interference from Cdc28-csr1. cdc28-rsc1 (R159G) exhibited essentially undetectable H1 kinase activity, and cdc28-rsc13 (E58G) exhibited only about 1% of wild-type levels (Fig. 9A). cdc28-rsc5 (R178S) had about 10% of the wild-type activity but was still clearly reduced from the wild type cdc28-rsc15 (multiply mutant) had about 40% of the wild-type activity, but Cdc28 protein abundance was somewhat reduced. Next, we tested the binding and kinase activity of these Cdc28-rsc proteins complexed to Cln2. As for the cdc28-csr alleles, we first untagged the cdc28-rsc alleles (see Materials and Methods) and then introduced the untagged cdc28-rsc alleles into plasmids into a cdc28-csr1 GAL1::CLN2-HA strain. As described above, in extracts from this strain, Cln2 could be immunoprecipitated by using the HA tag, but no significant Cdc28 protein or kinase activity coprecipitated due to the csr1 mutation. Therefore, we could attribute almost all the Cdc28 binding and kinase activity to the introduced plasmid cdc28-rsc allele. The cdc28-rsc mutants had almost no detectable kinase coprecipitated with Cln2-HA (<2% of the wild-type level). Cdc28 coimmunoprecipitation was variable, being at least 50% of the wild-type level for cdc28-rsc mutants 5 and 13 and roughly 10% of the wild-type level for cdc28-rsc mutants 1 and 15 (Fig. 9B).
FIG. 9.
Total and Cln2-associated kinase assays and Cdc28 binding for cdc28-rsc alleles. (A) Comparison of total Cdc28 kinase activities for cdc28-rsc alleles. Strain 2198-3A-2a (CLN1 CLN2 CLN3 cdc28::HIS3 pKL039-1) was transformed with plasmids RS414 (Vector), SF19 (CDC28-HA), KL059 (CDC28-rsc1-HA), KL060 (cdc28-rsc5-HA), KL061 (cdc28-rsc13-HA), or KL062 (cdc28-rsc15-HA). Cultures of each transformant strain were grown overnight at 30°C in SCDex-Trp. HA-tagged Cdc28 protein was isolated, and kinase assays were performed at 30°C. Cdc28 protein levels were analyzed by immunoblotting. All the procedures are described in Materials and Methods. The anti-HA immunoblot and H1 kinase blot are shown. The lanes on each blot correspond to transformed strains 2198-3A-2a pRS414, 2198-3A-2a pSF19, 2198-3A-2a pSF19, 2198-3A-2a pKL059, 2198-3A-2a pKL060, 2198-3A-2a pKL061, and 2198-3A-2a pKL062. Values indicated above the H1 kinase blot are total H1 kinase activities relative to wild-type values, not standardized to Cdc28 protein levels. Note that the levels of Cdc28-rsc15 protein were reproducibly lower than wild-type levels; when standardized to Cdc28-rsc15 protein levels, bulk Cdc28-rsc15 kinase activity for this experiment was 0.9, relative to wild-type activity. (B) Comparison of Cln2-associated kinase activity and associated Cdc28 protein for cdc28-rsc alleles. Strain 2195-3A (cdc28-csr1 GAL1::CLN2-HA) was transformed with plasmids RS414 (Vector), RD47 (CDC28), KL063 (cdc28-rsc1), KL064 (cdc28-rsc5), KL065 (cdc28-rsc13), or KL066 (cdc28-rsc15). Strain 2193-13B (cdc28-csr1) was transformed with pRS414. Cultures of each transformant strain were grown overnight at 30°C in SCGal-Trp medium. HA-tagged Cln2 protein was isolated, and kinase assays were performed at 30°C. Cln2-HA protein and associated Cdc28 protein were analyzed by immunoblot. All protocols are described in Materials and Methods. The anti-HA immunoblot, anti-Cdc28 immunoblot, and H1 kinase blot are shown. The lanes of each blot correspond to transformed strains 2195-13B pRS414, 2195-3A pRS414, 2195-3A pRD47, 2195-3A pRD47, 2195-3A pKL063, 2195-3A pKL064, 2195-3A pKL065, and 2195-3A pKL066. Values indicated above the anti-Cdc28 immunoblot and H1 kinase blot were obtained as described in the legend to Fig. 5B.
How can severely kinase-defective Cdc28 provide biological function?
The results with the csr alleles of cdc28 (see above) clearly showed that Cln2-dependent mating-factor resistance was genetically CDC28 dependent. However, the results with the rsc alleles suggested that severely kinase-defective alleles of CDC28, some of which were also significantly defective in binding to Cln2, could nevertheless support CLN2-dependent mating-factor resistance, in some cases to nearly wild-type levels (Fig. 8). The effect of the cdc28-rsc alleles on the mating-factor pathway may be indirect, working by somehow recruiting the resident Cdc28-csr1 protein, normally incompetent to interact with Cln2 to cause mating-factor resistance, into effective complexes with Cln2. However, it is clear from the measurement of Cln2-associated kinase activity that the rsc alleles did not result in the activation of Cln2-dependent Cdc28-csr1-dependent kinase activity detectable in vitro, since the activation observed in the cdc28-rsc transformants (Fig. 9B) was close to the background cdc28-csr levels (Fig. 9B, Vector lanes).
One specific model for how the cdc28-rsc alleles could complement the cdc28-csr1 defect in CLN2-dependent mating-factor resistance is that the Cdc28-rsc proteins sequester the Far1 inhibitor (4, 27). cdc28-csr1 far1 GAL1::CLN2 strains are resistant to mating factor on galactose (GAL1::CLN2 on) but not on glucose (GAL1::CLN2 off), presumably due to inefficient Cln2/Cdc28-csr1 interaction combined with the loss of Far1-dependent inhibition. (Mating-factor resistance in far1 strains is dependent on CLN2 [4].) cdc28-csr1 far1 GAL1::CLN2 strains are resistant to mating factor on galactose only at 30°C, not at 38°C, perhaps due to partial thermolability of the Cdc28-csr1 function. Therefore, we tested the cdc28-rsc alleles for the ability to complement the defect in cdc28-csr1-1 far1 GAL1::CLN2 mating-factor resistance under these two conditions: glucose at 30°C and galactose at 38°C (Fig. 10A). cdc28-rsc1 complemented mating-factor resistance under both conditions, and cdc28-rsc5 and cdc28-rsc13 complemented it on galactose at 38°C. Thus, the ability of cdc28-rsc1, cdc28-rsc5, and cdc28-rsc13 to complement mating-factor resistance in cdc28-csr1 strains (Fig. 8 and 10A) is at least not completely dependent on interaction with Far1. Furthermore, complementation by cdc28-rsc1 does not require CLN2 overexpression. In contrast, cdc28-rsc15 could not complement the defect in cdc28-csr1 far1 GAL1::CLN2 mating-factor resistance under either condition (Fig. 10A). Thus, the ability of cdc28-rsc15 to complement mating-factor resistance in the cdc28-csr1 FAR1 GAL1::CLN2 cells (Fig. 8) could be attributed to antagonizing or sequestering Far1, resulting in its inability to complement in the far1 mutant strain. Surprisingly, we found that wild-type CDC28 was unable to confer strong mating-factor resistance in this strain on galactose medium (GAL1::CLN2 expressed) at 38°C, being much weaker in this assay than the rsc alleles. In contrast, GAL1::CLN2 strains with wild-type CDC28 but lacking cdc28-csr1 are strongly mating-factor resistant at 38°C (data not shown). This suggests some dominant-negative effect of the cdc28-csr1 allele to which the rsc alleles are not subject.
FIG. 10.
Additional CLN2-dependent complementation of cdc28-csr1 by cdc28-rsc alleles. (A) Complementation of mating-factor resistance of CLN1 CLN2 CLN3 far1::URA3 cdc28-csr1 mating-factor resistance. Strain 2180-14A-1a (cdc28-csr1 far1::URA3 GAL1::CLN2) was transformed with plasmids RS414 (Vector), SF19 (CDC28-HA), KL054 (cdc28-csr1), KL059 (cdc28-rsc1-HA), KL060 (cdc28-rsc5-HA), or KL061 (cdc28-rsc13-HA). Each transformant strain was grown overnight in both SCDex-Trp and SCGal-Trp. Portions of the SCDex-Trp cultures (100 μl) and the SCGal-Trp cultures (200 μl) were spread onto a YEPDex plate and YEPGal plate, respectively. The YEPDex and YEPGal plates were allowed to incubate at 30 and 38°C, respectively, for about 4 h. Following incubation, sterile disks containing 15 μl of a 0.05, 0.1, or 0.2 mM alpha-factor dilution were placed on each plate, clockwise from the top left. YEPDex plates were allowed to incubate at 30°C for 2 days, and YEPGal plates were allowed to incubate at 38°C for 3 days. The picture on the left shows the wild-type CDC28 and cdc28-rsc1 complementation of the mating-factor resistance of this strain on YEPDex at 30°C. The pictures on the right show the cdc28-rsc1, cdc28-rsc5, and cdc28-rsc13 complementation of the mating-factor resistance of this strain on YEPGal at 38°C. Wild-type CDC28 only weakly complements in this assay. (B) Complementation of cln1 cln3 cdc28-13 pcdc28-csr1 inviability at 38°C. Strain 1227-3C (cln1 CLN2 cln3 cdc28-13 pGAL1::CLN3/URA3) was grown on YEPDex medium and screened for loss of pGAL1::CLN3/URA3. The resulting strain was cotransformed with both the KL039-1 plasmid (cdc28-csr1/URA3) and one of the TRP1 plasmids KL059 (cdc28-rsc1-HA), KL060 (cdc28-rsc5-HA), KL061 (cdc28-rsc13-HA), or KL062 (cdc28-rsc15-HA). Transformants were grown on SCDex-Trp-Ura. For each transformed strain, 10-fold serial dilutions were prepared, and 5 μl of each dilution was plated onto a YEPDex plate at both 30 and 38°C. The plates were allowed to incubate for 3 days. A representative set of dilutions is shown.
As an additional test of the CLN2-dependent function of these alleles, we tested if the cdc28-rsc alleles could complement cdc28-csr1 cln1 CLN2 cln3 temperature sensitivity (Fig. 10B). cdc28-rsc1, cdc28-rsc5, and cdc28-rsc13 were able to complement this defect; cdc28-rsc1 did so quite strongly. This extends the CLN2-dependent function of these cdc28-rsc alleles to a context in which CLN2 is not overexpressed and in which the mating-factor pathway is not involved.
Analysis of the cdc28-rsc alleles is complicated in part by their inability to complement a cdc28 null allele and the consequent requirement for the presence of an additional CDC28 allele in the cells assayed. As mentioned above, this requirement leaves open the possibility that the cdc28-rsc alleles provide CLN2-dependent functions by somehow recruiting the resident Cdc28-csr1 protein into effective Cln2-Cdc28 complexes. In an attempt to further address this possibility, we decided to ask whether the CLN2-dependent functions of the cdc28-rsc alleles were specifically dependent upon the cdc28-csr1 allele by testing for cdc28-rsc complementation of mating-pheromone sensitivity of the 2112-10C strain (cln1 cln2 CLN3 GAL1::CLN2 cdc28-13), a strain not containing the cdc28-csr1 allele, at 30°C (Fig. 4A). Of the four cdc28-rsc alleles tested, only cdc28-rsc1 and cdc28-rsc15 yielded healthy transformants; cdc28-rsc5 and cdc28-rsc13 produced only tiny transformant colonies, suggesting a dominant negative effect of these alleles in the cdc28-13 background. The cdc28-rsc1 allele was completely unable to complement the mating-pheromone sensitivity of strain 2112-10C at 30°C. Therefore, we cannot exclude the possibility that the CLN2-dependent functions of cdc28-rsc1 are dependent specifically upon the presence of cdc28-csr1. The cdc28-rsc15 allele was able to weakly complement the mating-pheromone sensitivity of strain 2112-10C at 30°C (approximately 100-fold less than wild-type CDC28 levels [data not shown]), suggesting that the mating-pheromone resistance of cdc28-rsc15 is not completely specific for the presence of the cdc28-csr1 allele.
To test whether known kinase-inactive alleles of CDC28 have similar CLN2-dependent functions in these assays, we tested the ability of cdc28-T169A (pRD58 and pKL019) to complement the mating-pheromone sensitivity of strain 2198-3A-2a (CLN1 CLN2 CLN3 GAL1::CLN2 cdc28::HIS3 pcdc28-csr1/URA3). The cdc28-T169A allele has been shown to be inactive as a kinase yet able to associate with both Cln2 and Clb2 proteins (20). cdc28-T169A was unable to rescue the pheromone sensitivity of this strain (data not shown). This result indicates that not all kinase-inactive Cdc28 proteins, even those that bind to Cln2, can provide CLN2-dependent functions.
DISCUSSION
CDC28 alleles with cyclin-specific functional defects.
Cln2 is thought to function by binding and activating the Cdc28 kinase (8, 29, 35, 36, 39) Cln2 has been found to immunoprecipitate with Cdc28 in an active kinase complex (35, 39) whose activity is dependent upon a functional CDC28 allele (39). The overexpression of CLN2 leads to repression of the mating-pheromone pathway and resistance to mating-pheromone-induced arrest (25). The overexpression of CLN2 has also been shown to cause hyperpolarization of cortical actin and elongation of buds (19). Both repression of the mating-pheromone pathway and the hyperpolarization phenotype have been shown to be dependent upon a functional CDC28 allele; at restrictive temperatures, cells containing temperature-sensitive alleles of cdc28 fail to repress the mating-pheromone pathway or to hyperpolarize in response to overexpressed CLN2 (19, 25). In these assays, cells containing temperature-sensitive alleles of cdc28 were arrested in G1. Here we have isolated and characterized an allele of CDC28, cdc28-csr1, that supports both cell cycle progression and high levels of bulk-Cdc28 and Clb2-Cdc28 kinase activity yet specifically prevents CLN2-mediated repression of the mating-pheromone pathway and hyperpolarization, as well as the formation of Cln2-Cdc28 complex and kinase activity. These observations further support the idea that these CLN2-mediated responses require Cdc28 in specific physical association with Cln2.
The cdc28-csr1 mutation appeared easy to understand, functioning poorly in both genetic and biochemical CLN2-dependent assays. Two additional cdc28-csr alleles were similar to cdc28-csr1, having both genetic and biochemical CLN2-dependent defects. However, two other cdc28-csr alleles, similarly defective in supporting CLN2-mediated mating-pheromone resistance and viability in combination with weakened alleles of CLN2, were found to promote high levels of Cln2-associated kinase activity and Cln2 binding. One possible explanation for these findings is that small quantitative defects in Cln2-associated kinase activities place these mutants below a threshold of Cln2 activity necessary to support CLN2-dependent functions. A second possible explanation for these findings is that these mutants have qualitative defects in their Cln2-associated kinase activity and, although able to phosphorylate histone H1 in vitro, are unable to phosphorylate the substrates necessary to support CLN2-dependent functions in vivo. For example, the mutants may be defective in their physical association with potential Cln2-Cdc28 substrates. A third possible explanation for these findings is that Cln2-Cdc28 kinase activity is not directly relevant to CLN2-mediated mating-pheromone resistance (see below).
The isolation of csr alleles of CDC28 that are genetically and biochemically much more defective in their interaction with Cln2 than with other cyclins could indicate that Cln2 interacts with Cdc28 in a way that is qualitatively different from other cyclins or that the binding interface between Cln2 and Cdc28 is generally more sensitive to mutation. Recent results with a large series of mutant alleles of CLN2 suggests that there are significant differences between the Cln2-Cdc28 interaction and the cyclin A-Cdk2 interaction, since some mutations in the cyclin homology region of Cln2 predicted to destroy cyclin folding based on the cyclin A crystal structure (15, 30) have relatively minor effects on Cln2 function (14). Therefore, general lability of the Cln2-Cdc28 interface may not be expected.
Kinase-inactive alleles of CDC28 with biological activity?
When we looked for mutant CDC28 genes that would support CLN2-dependent mating-factor resistance but not complement a cdc28 null for viability, we thought that it might be possible to obtain alleles of CDC28 with close to normal levels of Cln2-associated kinase activity but with defects in Clb-associated (or bulk) kinase activity. In fact, we obtained alleles of CDC28 that were strikingly defective in Cln2-associated kinase activity, and in some cases highly defective in Cln2 binding as well, that nevertheless could complement the CLN2-dependent mating-factor resistance defect of cdc28-csr1. Some of these alleles could also complement defects of cdc28-csr1 in other CLN2-dependent functions. One possible explanation for these results is that despite the lack of detectable Cln2-associated kinase activity in our in vitro assay system, these mutants do in fact have sufficient Cln2-associated kinase activity in vivo to support CLN2-dependent functions. It is possible that these mutants, despite their inability to phosphorylate histone H1, would be fully capable of phosphorylating the substrates necessary for CLN2-dependent responses. However, given the fact that at least one of the mutants, cdc28-rsc13, has a mutation in a critical catalytic residue and that two others, cdc28-rsc1 and cdc28-rsc5, have mutations in residues that are likely to be important for kinase activity (Table 4), it appears more probable that these mutants are indeed inactive or severely reduced for Cln2-associated kinase function yet somehow function to promote CLN2-dependent functions. One explanation for kinase-inactive alleles with CLN2-dependent activities is that the kinase-inactive Cdc28 mutants are working through an indirect mechanism, somehow recruiting the resident Cdc28-csr protein into effective complexes with Cln2. For example, the Cdc28-rsc proteins could bind tightly to Cln2-Cdc28 inhibitor proteins, relieving inhibition of the Cln2/Cdc28-csr1 complex. Arguing against this idea is the observation that the kinase-inactive mutants do not restore detectable Cln2/Cdc28-csr1 binding or kinase activity in immunoprecipitates (see above). Also, we tested this possibility genetically in the case of Far1, a protein that has been reported to inhibit the activity of the Cln2-Cdc28 kinase complex (27). Our results (see above) indicate that the kinase-inactive alleles of Cdc28 do not work (or at least do not work solely) through a Far1-dependent mechanism. We cannot rule out the possibility that another (as yet unidentified) Cln2-Cdc28 inhibitor is being targeted by the kinase-inactive Cdc28 proteins.
Another possible explanation for kinase-inactive alleles of cdc28 with CLN2-dependent activities is that Cln2-associated Cdc28 kinase activity is not required for some CLN2-dependent processes and that the kinase-inactive alleles are directly capable of performing these functions. Perhaps, for example, some CLN2-dependent responses depend only upon Cln2-Cdc28 complex formation and not upon Cln2-Cdc28 kinase activity. The kinase-inactive Cdc28 complexed with Cln2 might still bind to some substrate(s), and binding might be sufficient for a biological response even without substrate phosphorylation. Along the same lines, a potentially enhanced or irreversible binding of the kinase-inactive Cln2-Cdc28 complex to some substrate(s) might bypass the need for substrate phosphorylation. For example, substrates that are usually phosphorylated and inactivated by the Cln2-Cdc28 complex might be effectively inactivated by being tightly bound by the mutant kinase complex. The suggestion that cyclins may have functions independent of their abilities to activate the kinase activities of their Cdk partners has been made for cyclin D1. Cyclin D1 has been found to activate the transcription of estrogen receptor-regulated genes, independent of complex formation to a Cdk partner (42). Our results with the cdc28-rsc alleles are similar to the cyclin D1 findings in that the Cln2 cyclin might be able to function independently of associated kinase activity. Our results differ in that the CLN2-mediated mating pheromone repression is dependent on both CLN2 overexpression and on the Cdk partner of Cln2, Cdc28.
ACKNOWLEDGMENTS
Many thanks go to Daniel Lim for his valuable contributions to the initial stages of the Csr screen. We thank Ray Deshaies and Peter Sorger for providing plasmids and Ray Deshaies for providing the anti-Cdc28 antibody.
This work was supported by NIH grant GM47238 and the Norman and Rosita Winston Foundation. K.L. is a Howard Hughes Medical Institute Predoctoral Fellow.
REFERENCES
- 1.Allen J B, Elledge S J. A family of vectors that facilitate transposon and insertional mutagenesis of cloned genes in yeast. Yeast. 1994;10:1267–1272. doi: 10.1002/yea.320101003. [DOI] [PubMed] [Google Scholar]
- 2.Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K. Current protocols in molecular biology. New York, N.Y: Wiley Interscience; 1987. [Google Scholar]
- 3.Boeke J D, LaCroute F, Fink G R. A positive selection for mutants lacking 5′ phosphate decarboxylase activity in yeast: 5-fluoro-orotic acid resistance. Mol Gen Genet. 1984;197:345–346. doi: 10.1007/BF00330984. [DOI] [PubMed] [Google Scholar]
- 4.Chang F, Herskowitz I. Identification of a gene necessary for cell cycle arrest by a negative growth factor of yeast: FAR1 is an inhibitor of a G1 cyclin, CLN2. Cell. 1990;63:999–1011. doi: 10.1016/0092-8674(90)90503-7. [DOI] [PubMed] [Google Scholar]
- 5.Cross F R. Cell cycle arrest caused by CLN gene deficiency in Saccharomyces cerevisiae resembles START-I arrest and is independent of the mating-pheromone signalling pathway. Mol Cell Biol. 1990;10:6482–6490. doi: 10.1128/mcb.10.12.6482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Cross F R. Starting the cell cycle: what’s the point? Curr Opin Cell Biol. 1995;7:790–797. doi: 10.1016/0955-0674(95)80062-x. [DOI] [PubMed] [Google Scholar]
- 7.Cross F R. Marker swap plasmids: convenient tools for yeast molecular genetics. Yeast. 1997;13:647–653. doi: 10.1002/(SICI)1097-0061(19970615)13:7<647::AID-YEA115>3.0.CO;2-#. [DOI] [PubMed] [Google Scholar]
- 8.Cross F R, Blake C M. The yeast Cln3 protein is an unstable activator of Cdc28. Mol Cell Biol. 1993;13:3266–3271. doi: 10.1128/mcb.13.6.3266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Deshaies R J, Kirschner M. G1 cyclin-dependent activation of p34CDC28 (Cdc28p) in vitro. Proc Natl Acad Sci USA. 1995;92:1182–1186. doi: 10.1073/pnas.92.4.1182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Dirick L, Böhm T, Nasmyth K. Roles and regulation of Cln-Cdc28 kinases at the start of the cell cycle of Saccharomyces cerevisiae. EMBO J. 1995;14:4803–4813. doi: 10.1002/j.1460-2075.1995.tb00162.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Grandin N, Reed S I. Differential function and expression of Saccharomyces cerevisiae B-type cyclins in mitosis and meiosis. Mol Cell Biol. 1993;13:2113–2125. doi: 10.1128/mcb.13.4.2113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hoek, M., and F. R. Cross. Unpublished results.
- 13.Horton R M, Hunt H D, Ho S N, Pullen J K, Pease L R. Engineering hybrid genes without the use of restriction enzymes: gene splicing by overlap extension. Gene. 1989;77:61–68. doi: 10.1016/0378-1119(89)90359-4. [DOI] [PubMed] [Google Scholar]
- 14.Huang K N, Odinsky S A, Cross F R. Structure-function analysis of the Saccharomyces cerevisiae G1 cyclin Cln2. Mol Cell Biol. 1997;17:4654–4666. doi: 10.1128/mcb.17.8.4654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Jeffrey P D, Russo A A, Polyak K, Gibbs E, Hurwitz J, Massague J, Pavletich N. Mechanism of CDK activation revealed by the structure of a cyclinA-CDK2 complex. Nature. 1995;376:313–320. doi: 10.1038/376313a0. [DOI] [PubMed] [Google Scholar]
- 16.Johnson L N, Noble M E M, Owen D J. Active and inactive protein kinases: structural basis for regulation. Cell. 1996;85:149–158. doi: 10.1016/s0092-8674(00)81092-2. [DOI] [PubMed] [Google Scholar]
- 17.Leung D W, Chen E, Goeddel D V. A method for random mutagenesis of a defined DNA segment using a modified polymerase chain reaction. Technique. 1989;1:11–15. [Google Scholar]
- 18.Levine K, Huang K, Cross F R. Saccharomyces cerevisiae G1 cyclins differ in their intrinsic functional specificities. Mol Cell Biol. 1996;16:6794–6803. doi: 10.1128/mcb.16.12.6794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Lew D J, Reed S I. Morphogenesis in the yeast cell cycle: regulation by Cdc28 and cyclins. J Cell Biol. 1993;120:1305–1320. doi: 10.1083/jcb.120.6.1305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Lim H H, Loy C J, Zaman S, Surana U. Dephosphorylation of threonine 169 of Cdc28 is not required for exit from mitosis but may be necessary for Start in Saccharomyces cerevisiae. Mol Cell Biol. 1996;16:4573–4583. doi: 10.1128/mcb.16.8.4573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.McCaffrey G, Clay F J, Kelsey K, Sprague G F. Identification and regulation of a gene required for cell fusion during mating of the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1987;7:2680–2690. doi: 10.1128/mcb.7.8.2680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Muhlrad D, Hunter R, Parker R. A rapid method for localized mutagenesis of yeast genes. Yeast. 1992;8:79–82. doi: 10.1002/yea.320080202. [DOI] [PubMed] [Google Scholar]
- 23.Nasmyth K. At the heart of the budding yeast cell cycle. Trends Genet. 1996;12:405. doi: 10.1016/0168-9525(96)10041-x. [DOI] [PubMed] [Google Scholar]
- 24.Oehlen, L. J. W. M., and F. R. Cross. Unpublished data.
- 25.Oehlen L J W M, Cross F R. G1 cyclins CLN1 and CLN2 repress the mating factor response pathway at Start in the yeast cell cycle. Genes Dev. 1994;8:1058–1070. doi: 10.1101/gad.8.9.1058. [DOI] [PubMed] [Google Scholar]
- 26.Oehlen L J W M, McKinney J D, Cross F R. Ste12 and Mcm1 regulate cell cycle-dependent transcription of FAR1. Mol Cell Biol. 1996;16:2830–2837. doi: 10.1128/mcb.16.6.2830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Peter M, Herskowitz I. Direct inhibition of the yeast cyclin-dependent kinase Cdc28-Cln by Far1. Science. 1994;265:1228–1231. doi: 10.1126/science.8066461. [DOI] [PubMed] [Google Scholar]
- 28.Reed S I. The selection of S. cerevisiae mutants defective in the start event of cell division. Genetics. 1980;95:561–577. doi: 10.1093/genetics/95.3.561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Richardson H E, Wittenberg C, Cross F, Reed S I. An essential G1 function for cyclin-like proteins in yeast. Cell. 1989;59:1127–1133. doi: 10.1016/0092-8674(89)90768-x. [DOI] [PubMed] [Google Scholar]
- 30.Russo A A, Jeffrey P D, Pavletich N P. Structural basis of cyclin-dependent kinase activation by phosphorylation. Nat Struct Biol. 1996;3:696–700. doi: 10.1038/nsb0896-696. [DOI] [PubMed] [Google Scholar]
- 31.Sikorski R S, Hieter P. A system of shuttle vectors and yeast strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics. 1989;122:19–27. doi: 10.1093/genetics/122.1.19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Stevenson B J, Rhodes N, Errede B, Sprague G F., Jr Constitutive mutants of the protein kinase STE11 activate the yeast pheromone response pathway in the absence of the G protein. Genes Dev. 1992;6:1293–1304. doi: 10.1101/gad.6.7.1293. [DOI] [PubMed] [Google Scholar]
- 33.Stuart D, Wittenberg C. CLN3, not positive feedback, determines the timing of CLN2 transcription in cycling cells. Genes Dev. 1995;9:2780–2794. doi: 10.1101/gad.9.22.2780. [DOI] [PubMed] [Google Scholar]
- 34.Trueheart J, Boeke J D, Fink G R. Two genes required for cell fusion during yeast conjugation: evidence for a pheromone-induced surface protein. Mol Cell Biol. 1987;7:2316–2328. doi: 10.1128/mcb.7.7.2316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Tyers M, Tokiwa G, Futcher B. Comparison of the Saccharomyces cerevisiae G1 cyclins: Cln3 may be an upstream activator of Cln1, Cln2 and other cyclins. EMBO J. 1993;12:1955–1968. doi: 10.1002/j.1460-2075.1993.tb05845.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Tyers M, Tokiwa G, Nash R, Futcher B. The Cln3-Cdc28 kinase complex of S. cerevisiae is regulated by proteolysis and phosphorylation. EMBO J. 1992;11:1773–1784. doi: 10.1002/j.1460-2075.1992.tb05229.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wassmann K, Ammerer G. Overexpression of the G1-cyclin gene CLN2 represses the mating pathway in Saccharomyces cerevisiae at the level of the MEKK Ste11. J Biol Chem. 1997;272:13180–13188. doi: 10.1074/jbc.272.20.13180. [DOI] [PubMed] [Google Scholar]
- 38.Wittenberg C, Richardson S L, Reed S I. Subcellular localization of a protein kinase required for cell cycle initiation in Saccharomyces cerevisiae: evidence for an association between the CDC28 gene product and the insoluble cytoplasmic matrix. J Cell Biol. 1987;105:1527–1538. doi: 10.1083/jcb.105.4.1527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Wittenberg C, Sugimoto K, Reed S I. G1-specific cyclins of S. cerevisiae: cell cycle periodicity, regulation by mating pheromone, and association with the p34CDC28 protein kinase. Cell. 1990;62:225–237. doi: 10.1016/0092-8674(90)90361-h. [DOI] [PubMed] [Google Scholar]
- 40.Zanolari B, Riezman H. Quantitation of α-factor internalization and response during the Saccharomyces cerevisiae cell cycle. Mol Cell Biol. 1991;11:5251–5258. doi: 10.1128/mcb.11.10.5251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Zheng J, Knighton D R, Ten Eyck L F, Karlsson R, Xuong N, Taylor S S, Sowadski J M. Crystal structure of the catalytic subunit of cAMP-dependent protein kinase complexed with MgATP and peptide inhibitor. Biochemistry. 1993;32:2154–2161. doi: 10.1021/bi00060a005. [DOI] [PubMed] [Google Scholar]
- 42.Zwijsen R M L, Wientjens E, Klompmaker R, van der Sman J, Bernards R, Michalides R J A M. CDK-independent activation of estrogen receptor by cyclin D1. Cell. 1997;88:405–415. doi: 10.1016/s0092-8674(00)81879-6. [DOI] [PubMed] [Google Scholar]