Abstract
Clinical testing and public health surveillance can be significantly improved by incorporating sequencing-based molecular detection and subtyping for real-time monitoring of virus evolution. With phylogenetic analysis used for speciation and variant subtyping, target analyte specificity can be relaxed well beyond typical parameters acceptable in PCR-based diagnostics. Hybrid capture is a promising way to enrich large numbers of sequences with maximal flexibility, using standard molecular biology laboratory equipment and small benchtop sequencers. Here, we report the development and bench validation of a hybrid capture based next-generation sequencing diagnostic panel for all known viral tick-borne pathogens, TITAN-RNA. Based on systematic testing with simulated novel viruses and field samples, we determined a 10% tolerance for evenly distributed mutations or 27% tolerance for naturally occurring viral divergence. The TITAN-RNA extrapolated limit of detection in blood is 19.1 genome copies by complementary log-log analysis, and linearity performance (R2 ≥ 0.99) is amenable for its use as a quantitative assay. As proof of principle for public health surveillance and evolutionary studies, we report two putatively novel segmented Flavi-like viruses in New York State, USA, identified from the invasive Haemaphysalis longicornis tick.
1. Introduction
There are more than 50,000 cases of vector-borne diseases reported annually in the USA, and 95% of cases are caused by pathogens spread by ticks [1]. Of the three families of ticks, Ixodidae and Argasidae are associated with tick-borne diseases (TBDs). Hard ticks (ixodid) are the most important vectors in the United States. However, soft ticks belonging to the genus Ornithodoros can be vectors of public health importance [1]. Two Ixodidae species of interest in the Eastern US are the blacklegged tick (Ixodes scapularis) and the Asian longhorned tick (Haemaphysalis longicornis). Both I. scapularis and H. longicornis are spreading in the USA likely due to environmental warming and longer seasonal periods of high humidity [2]. H. longicornis is tolerant to cold climates, which is the likely reason for its proliferation in the Northeast and Midwest United States after being first discovered in 2017 [3, 4].
Tick-borne viruses of the Flaviviridae family belong to one of two genomic classes: unsegmented or segmented. Unsegmented flavivirus genomes are positive-sense single stranded (ss), RNA viruses, with genomes that are approximately 9.2 to 11 kb long. The RNA translates into a polyprotein encoding seven non-structural proteins (NS1, NS2A, NS2B, NS3, NS4A, NS4B and NS5) and three structural proteins: capsid (C), pre-membrane (prM), membrane (M), and envelope (E). Tick-borne encephalitis virus (TBEV), Louping ill virus (LIV), Asian Omsk hemorrhagic fever virus (AOHFV), Kyasanur Forest disease (KFD), and Powassan virus (POWV) are examples of unsegmented flavivirus genomes. Segmented jingmenvirus genomes, from a more recently described genus in this family, contain four or five segments of positive sense ssRNA. One segment encodes either NS2B/NS3-like proteins, another segment encodes the NS5-like RNA-dependent RNA Polymerase (NSP1), and two or three other unique segments encode VP1-, VP2-, and VP3-like structural proteins which include the capsid protein and the membrane protein. In the case of the emerging Jingmen-tick virus (JMTV), the segments are putatively characterized as segment 1 encoding NSP1 (NS5-like), segment 2 encoding VP1/1ab (glycoprotein), segment 3 encoding NSP2 (NS2B/NS3-like), and segment 4 encoding VP2 and VP3 (capsid and membrane proteins). Alongshan virus (ALSV), Mogiana tick virus (MGTV), and Guaico culex virus (GCXV) represent other segmented jingmenviruses [5–7]. The unsegmented Tamana bat virus is an unclassified member of the family Flaviviridae [8].
Tick-borne viruses (TBV’s) belonging to the Flaviviridae family are widely reported around the world and lead to more than 10,000 hospitalizations annually. TBEV, causes encephalitis, non-specific viremia, neurological, and hemorrhagic symptoms in the infected patients [9–11]. The primary known vectors for TBEV are I. ricinus and I. persulcatus [12, 13]. Notably, TBEV incidence has been increasing in Europe and the Eurasian region likely due to low vaccination uptake, landscape changes, and climate change [14–24]. The AOH, LIV, and KFD fever viruses have been detected in Europe, Russia, China, Japan, India, Southeast Asia, and the Middle east and are slow to expand beyond their localized regions. The tick vectors for these viruses are ticks belonging species in Ixodes, Dermacentor, and Haemaphysalis genera. POWV is categorized as lineages I or II. POWV lineage II is steadily expanding in the Northeast and Midwest USA and is now considered an emerging pathogen with localized transmission foci [25, 26]. It has also been detected in Canada and Russia with the primary tick vector being I. scapularis in the Americas. POWV lineage I is vectored by I. cookei and I. marxi which are widely distributed in the regions which they populate. POWV illness is more severe than other TBV’s, with a fatality rate as high as 19% in adults [27]. POWV illness symptoms including encephalitis, meningitis, loss of coordination, speech difficulties, high fever, and headaches. One in ten cases of POWV are fatal, and 5 in 10 cause severe long-term neurological damage [28–31]. JMTV has been detected in patients with fatal Crimean-Congo Hemorrhagic Fever (CCHF) in Kosovo [7] and associated with mild to severe illness in patients infected in China [5]. The first report of a segmented TBV detection in the USA was in 2020 where rodents in Pennsylvania were identified with sequences with 70% amino acid identity with the JMTV polymerase, provisionally named Flavi-like segmented virus US (FLSV-US) [32].
Current molecular diagnostic methods for tick-borne pathogen surveillance.
Polymerase chain reaction (PCR) is the most used detection method in surveillance and is currently and accepted as the standard. PCR is limited as it can only identify closely related pathogens, and when targeting highly conserved regions it provides limited genomic information. This means that further analysis between closely related pathogens is required to determine taxonomical relationships [33]. Hybrid-capture Next Generation Sequencing (HCNGS) is an alternative enrichment method that addresses limitations of current metagenomic and amplicon-based sequencing methods. It requires biotinylated nucleic acid baits to complementarily bind to target DNA (or in vitro reverse-transcribed RNA) in prepared libraries during hybridization. Unlike PCR-based modalities, HCNGS panels allow for semi-targeted enrichment of target genomic regions in large quantities. A single HCNGS panel can hold hundreds to thousands of complementary baits and thus can facilitate the enrichment of many target sequences at once. Baits are generally 75 bp to 140 bp long and engineered with sequence degeneracy for variation tolerance. Flexibility can be increased further with degenerate nucleotides in the bait sequence. While the duration and cost of HCNGS is still an area of development, it does not require any specialized equipment other than a benchtop Illumina sequencer, which is now increasingly common in public health and hospital laboratories [34–36].
Tick-borne pathogens are a public health threat. This study observes linearity, limit of detection, and divergent sequence tolerance to develop, and bench validate hybrid capture based next-generation sequencing technology as a diagnostic or surveillance tool for viral tick-borne pathogens and other viral febrile illness agents. Field sample testing with this panel serves as proof of principle for detection of endemic and emerging tick-borne viruses.
2. Methods
2.1. Synthetic nucleic acids
Naturally occurring sequences.
Heartland virus (HRTV) L, Dabie bandavirus (DBV) L, and POWV NS5 synthetic high-fidelity DNA (Supplemental Table 1) were purchased from Integrated DNA Technologies (Newark, NJ, USA), with an initial specification concentration of 106 copies per μL. Each lyophilized sample was normalized at 1000 ng and resuspended in nuclease-free water at 10 ng per μL. Quantitative Synthetic POWV lineage II RNA was purchased from American Type Culture Collection (Manassas, Virginia, USA) (ATCC catalog number #VR-3275SD), with an initial specification range ≥ 105 to 106 copies per μL. Concentrations of synthetic POWV DNA or RNA were confirmed with digital droplet PCR (ddPCR) using the ddPCR Supermix for Probes or the 1-Step RT-ddPCR Advanced Kit for Probes (BioRad; Hercules, California, USA). YUAN and probe are listed in Table 1. The manufacturer’s standard methods were followed for the ddPCR using the Droplet Generator Oil for Probes (BioRad), the QX200 Droplet Generator (BioRad), and the QX200 Droplet Reader (BioRad). The DNA or RNA concentration was measured using the QX Manager software (BioRad).
Table 1.
Primer and Probe sequence information.
| Primer or Probe | Sequence (5’ to 3’) | Annealing temperature (°C) | Notes |
|---|---|---|---|
| FLAVI-1 (F) FLAVI-2 (R) |
AATGTACGCTGATGACACAGCTGGCTGGGACAC TCCAGACCTTCAGCATGTCTTCTGTTGTCATCCA |
50 | SuperScript III One-step Flavivirus NS5 [5, 38] |
| YUAN-F YUAN-R YUAN-P |
AGAATGGCCATGACAGACACAA AGCCAGTCACTCACHGCTCTCAT GCCCAAGAGCCRCAGCCAGG |
50 | TaqMan Fast Virus 1-step POWV NS5 [39] |
| FLSV-US-F1 FLSV-US-R1 |
GGWGCYATGGGYTACCAGAT TCCARGGTGAGTARTCCTTTCG |
Round 1 56 (+ 0.5°C per cycle) (10 cycles) Round 2 54 (30 cycles) |
Superscript IV One-step FLSV-US NS5-like [32] |
| FLSV-US-F2 FLSV-US-R2 |
GGWGCYATGGGYTACCAGATGGA CCARGGTGAGTARTCCTTTCGARATC |
Round 1 60 (10 cycles) Round 2 58 (30 cycles) |
Platinum II Hot Start FLSV-US NS5-like [32] |
Artificially divergent sequences.
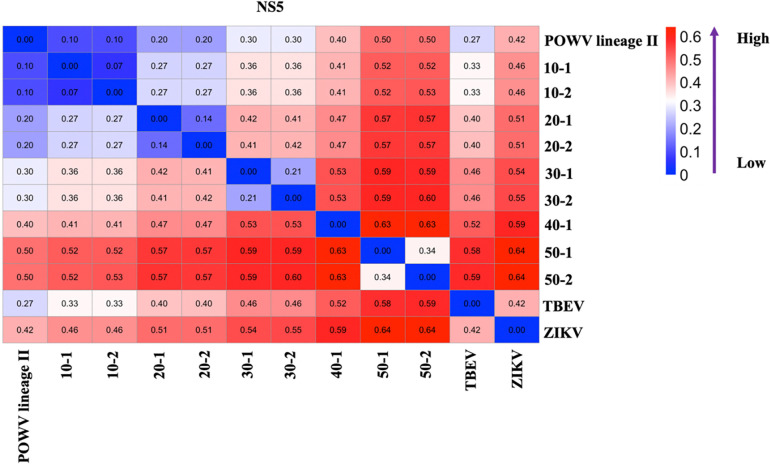
The sequence of POWV NS5 (NCBI # OL704303.1), which is the RNA-dependent RNA polymerase (RdRP) was used as a baseline (0%) to design modified sequences that were 10%, 20%, 30%, 40%, and 50% variable using a custom Python code. At total of sixteen sequences were designed: five (10%), four (20), three (30%), two (40%), and two (50%) independent replicate randomly varied sequences respectively (Supplemental Table 1). To ensure that the sequences were appropriately diverged from the baseline “ancestral” sequence, code was written to determine the number of mutations needed based on the original sequence length. POWV NS5 is 2709 nucleotides long so a 10% different sequence will have approximately 271 mutations. The code was written to create mutations that were random, evenly distributed, and with no more than 10% overlap between variable sequences. The python code used to generate the sequence can be found at (https://gitlab.com/goodmanlab/). The percent dissimilarity between the baseline and variant NS5 sequences is shown in Fig. 1. Artificial simulated variants, TBEV NS5 (NCBI# NC_001672.1), and ZIKV NS5 (NCBI# NC_012532.1) was ordered as high-fidelity DNA from IDT (2709 bp). Each lyophilized sample resuspended in nuclease-free water at 10 ng per μL. The concentration of each resuspended sample was measured using fluorimetry (Qubit Flex, Thermo Fisher Scientific; Liverpool, NY, USA).
Figure 1. Divergent Sequences.
Heatmap showing the pairwise-distance relationship between experimental and naturally occurring sequences. The bluest shade represents lowest dissimilarity, and the reddest shade represents the highest dissimilarity.
2.2. Field samples
Questing tick samples were collected and identified in June 2022, 2023, and 2024 as part of the New York State Tick Blitz citizen science project, as previously described [37]. Briefly, ticks were collected one time per day to reduce the likelihood of transporting ticks to other locations, and they were collected along a 300-m linear transect, either continuous or divided into shorter parcels in multiple sites across New York State. Visible ticks were collected with forceps and placed in plastic vials. Larvae were collected with lint rollers and then placed into plastics bags. Ticks were refrigerated in plastic sealable bags with moistened paper towels. Live ticks were mailed to Cornell University for identification as previously described [37]. Haemaphysalis longicornis ticks were homogenized using a 2.0-mm hollow brass bead in 800 uL of PBS and the BeadBlaster D2400 (Benchmark Scientific; Sayreville, NJ, USA) at 2.3.4 meters per second for 5 minutes. Homogenized samples were then centrifuged at 14,000 rpm for 1 minute and incubated for 1 hour with Proteinase K. Samples were centrifuged again for 3 minutes at 14,000 rpm, and 300 uL supernatant was used for the subsequent DNA/RNA extraction on the KingFisher Apex (Thermo Fisher Scientific) using the Quick-DNA/RNA Pathogen MagBead extraction kit (Zymo Research, catalog number # R2145 or R2146). Nucleic acid samples eluted from tick homogenates were frozen at −80°C.
2.3. Panel Curation and Design
Target curation.
Conserved regions of gene segments that were specific to each targeted pathogen were identified from alignments of nucleotide sequences of strains available from the GenBank nucleotide sequence database (https://www.ncbi.nlm.nih.gov/nucleotide/). Alignments of multiple sequences were performed using Clustal Omega on Geneious Version 2022.2.2. or MUSCLE v5 [40]. Accuracy of the alignment results was confirmed by querying sequences using Basic Local Alignment Search Tool (BLAST, https://blast.ncbi.nlm.nih.gov/Blast.cgi, [41]) and comparing the query sequences and results. The accuracy of consensus sequences in finding the desired agent was examined on BLAST. Consensus sequences of conserved gene regions with pairwise identity percentages of 90% and above, and identical site coverage of 85% and above were selected. Consensus sequences that met the requirements from the last step were used as query sequences to search for similar biological sequences in GenBank using BLASTN. Qualified consensus sequences that had more than 50% non-specific similarities, those with query sequence coverage being less than 100%, and those with queried sequences’ identity percentage of less than 98% were removed.
Panel design and synthesis.
Targeted Identification of Tick-borne Analytes by hybrid-capture NGS for RNA (TITAN-RNA) bait panels were designed and synthesized by Twist Biosciences (South San Francisco, California). Baits in each panel consist of 120 bp biotinylated DNA segments matching genes identified by literature and database review. Target sequences longer than 120bp were divided into 120bp fragments (baits) with overlapping segments at both ends. For validation purposes, the panel was split into TBD_Virus (Table 2) and TBD_viral_discovery (Table 3). TBD_Virus primarily targets known domestic and endemic tick-borne viruses, and TBD_viral_discovery primarily targets foreign tick-borne viruses. Both panels also target other vector-borne febrile illness-associated viruses as the intent is for syndromic use in diagnosing fevers of unknown origin. Together, these panels contain 8,805 baits targeting a total of 159 target regions and over 100 viruses.
Table 2. TBD_virus.
Target pathogen descriptions, target genes, and length of baits in TBD_Virus (v. 1–3)
| Virus | Accession | Description | Target gene | Target size (bp) |
|---|---|---|---|---|
| Alkhurma hemorrhagic fever virus | MN509835 | NS5 | 2708 | |
| Alongshan virus | MN107160 | NS5 | 2708 | |
| Bandavirus dabieense | MN509835 | Clade I consensus | L | 6210 (Deletion at 3' and 5' ends) |
| Bandavirus dabieense | Clade I consensus | 2853 | ||
| Bandavirus dabieense | Clade II consensus | 2852 | ||
| Bandavirus dabieense | Clade II consensus | 6210 | ||
| Bandavirus dabieense | Clade III consensus | 2853 | ||
| Bandavirus dabieense | Clade IV consensus | 2853 | ||
| Bandavirus dabieense | Clade V consensus | 2853 | ||
| Bandavirus dabieense | Clade VI consensus | 6210 | ||
| Bandavirus dabieense | Clade VII consensus | 2853 | ||
| Bandavirus heartlandense | KJ740148 | L | 2800 | |
| Bandavirus heartlandense | NC_024494 | M | 3427 | |
| Bandavirus heartlandense | NC_024496 | S | 1772 | |
| Bandavirus heartlandense | NC_078062 | NSs and N | 1753 | |
| Bandavirus heartlandense | NC_078063 | M | 3403 | |
| Bhanja virus | JX961616 (Group 1) | L | 2800 | |
| Bhanja virus | JX961619 | L | 2800 | |
| Bhanja virus | Consensus of all NCBI sequences | L | 2800 | |
| Bhanja virus | NC 027141 | M | ||
| Bhanja virus | NC_027142 | S | ||
| Bourbon virus | Consensus of all NCBI sequences | PB1 | 2148 | |
| Bourbon virus | Consensus of all NCBI sequences | NP | 1381 | |
| Chikungunya | MG649980 | ECSA_MASA | E1 | 1321 |
| Chikungunya | JF274082 | ECSA_IOL | E1 | 1321 |
| Chikungunya | KY680405 | AUL_Cbn | E1 | 1321 |
| Chikungunya | KY703937 | AUL_America | E1 | 1321 |
| Chikungunya | HM045796 | AUL_Asia | E1 | 1321 |
| Chikungunya | AY726732 | WA | E1 | 1321 |
| Colorado tick fever virus | Consensus of all NCBI sequences | Segment 1 | 3000 | |
| Colorado tick fever virus | Consensus of all NCBI sequences | Segment 9 | 1361 | |
| Crimean Congo Hemorrhagic virus | KR011838 | Clade V consensus | S | 1,550 |
| Crimean Congo Hemorrhagic virus | AJ538198 | Clade IV consensus | S | 1493 |
| Crimean Congo Hemorrhagic virus | OP345194 | Clade III consensus | S | 1600 |
| Crimean Congo Hemorrhagic virus | HQ849545 | Clade II consensus | S | 1528 |
| Crimean Congo Hemorrhagic virus | Clade I consensus | S | 1686 | |
| Gadgets gully virus | NC_033723.1 | NS5 | 2708 | |
| Kyasanur Forest disease virus | NC_039218.1 | NS5 | 2708 | |
| Langat virus | NC 003690 | NS5 | 2708 | |
| Louping ill virus | Clade I consensus | E | 1488 | |
| Louping ill virus | Clade II consensus | E | 1493 | |
| Louping ill virus | Clade III consensus | E | 1457 | |
| Omsk hemorrhagic fever virus | NC_005062.1 | NS5 | 2708 | |
| Powassan virus | Lineage I consensus | NS5 | 2709 | |
| Powassan virus | Lineage II consensus | NS5 | 2709 | |
| Powassan virus | Lineage I & II consensus | NS5 | 2709 | |
| Powassan virus | Lineage I consensus | Whole genome | 10.8 kb | |
| Powassan virus | Lineage II Consensus | Whole genome | 10.8 kb | |
| Royal Farm virus | NC_039219.1 | NS5 | 2708 | |
| Taï Forest reovirus | Consensus of all NCBI sequences | Segment 1 | 3000 | |
| Tarumizu tick virus | Consensus of all NCBI sequences | Segment 1 | 3000 | |
| Tick-borne encephalitis virus | Clade I consensus | E | 1488 | |
| Tick-borne encephalitis virus | Clade II consensus | E | 1206 | |
| Tick-borne encephalitis virus | Consensus of all NCBI sequences | Whole genome | 10.8 kb | |
Table 3. TBD_viral_discovery.
Target pathogens, descriptions, target genes, and length of baits in TBD_viral_discovery (v. 1–3).
| Genus | Virus species | Accession | Target gene | Target size (bp) |
|---|---|---|---|---|
| Orthonairovirus | Huangpi Tick Virus 1 | NC_031135.1 | L | 947 |
| Pacific coast tick nairovirus | KU933934.1 | L | 947 | |
| Tamdy virus | MT815992.1 | L | 947 | |
| Tacheng Tick Virus 1 | NC_031284.1 | L | 947 | |
| Burana virus | NC_043439.1 | L | 947 | |
| Shanxi tick virus 2 | MZ244235.1 | L | 947 | |
| Wenzhou Tick Virus | NC_031291.1 | L | 947 | |
| Songling virus | ON408079.1 | L | 947 | |
| Thiafora virus | NC_039220.1 | L | 947 | |
| Artashat virus | NC_043440.1 | L | 947 | |
| Avalon virus | NC_040460.1 | L | 947 | |
| Crimean Congo hemorrhagic fever virus 2 | DQ211612.1 | L | 947 | |
| Dugbe virus | NC_004159.1 | L | 947 | |
| Nairobi sheep disease virus | NC_034387.1 | L | 947 | |
| Tofla virus | NC_029124.1 | L | 947 | |
| Hazara virus | NC_038709.1 | L | 947 | |
| Meihua Mountain virus | ON184084.1 | L | 947 | |
| Estero Real orthobunyavirus | NC_055217.1 | L | 947 | |
| Great Saltee virus | NC_040512.1 | L | 947 | |
| IssykKul virus | LC495734.1 | L | 947 | |
| Keterrah virus | NC_034392.1 | L | 947 | |
| Uzun Agach virus | KP792741.1 | L | 947 | |
| Kasokero virus | NC_036636.1 | L | 947 | |
| Yogue virus | NC_029931.1 | L | 947 | |
| Chim virus | NC_043434.1 | L | 947 | |
| Qalyub virus | NC_034511.1 | L | 947 | |
| Geran virus | KP792714.1 | L | 947 | |
| Phlebovirus | Murre virus | NC_055353.1 | L | 716 |
| Alenquer virus | NC_055330.1 | L | 716 | |
| Echarte virus | NC_055339.1 | L | 716 | |
| Maldonado virus | NC_055344.1 | L | 716 | |
| Gabek Forest virus | NC_055319.1 | L | 716 | |
| Anhanga virus | NC_033836.1 | L | 716 | |
| Chagres virus | NC_055327.1 | L | 716 | |
| Laurel Lake virus | NC_043679.1 | L | 965 | |
| Lone Star virus | NC_021242.1 | L | 965 | |
| Forecariah virus | JX961625.2 | L | 965 | |
| Razdan virus | NC_022630.1 | L | 965 | |
| Razdan virus | KC335496.1 | L | 965 | |
| Palma virus | JQ956379.1 | L | 965 | |
| Hunter Island virus | KF848980.1 | L | 965 | |
| Guertu virus | NC_043611.1 | L | 965 | |
| Zwiesel bat banyangvirus | MN823639.1 | L | 965 | |
| Blacklegged tick phlebovirus | NC_055432.1 | L | 965 | |
| Mukawa virus | NC_043510.1 | L | 965 | |
| Candiru virus | NC_015374.1 | L | 965 | |
| Toscana virus | NC_006319.1 | L | 965 | |
| Sandfly fever | HM566172.1 | L | 965 | |
| Lihan Tick Virus | NC_055423.1 | L | 965 | |
| Tacheng Tick Virus 2 | NC_055425.1 | L | 965 | |
| Yongjia Tick Virus 1 | NC_055427.1 | L | 965 | |
| Silverwater virus | NC_055370.1 | L | 965 | |
| Huangpi Tick Virus 2 | NC_031138.1 | L | 965 | |
| Kaisodi virus | NC_040494.1 | L | 965 | |
| Kabuto mountain virus | NC_036604.1 | L | 965 | |
| Rukutama virus | NC_055368.1 | L | 965 | |
| Grand Arbaud virus | NC_055333.1 | L | 965 | |
| Precarious point virus | NC_055337.1 | L | 965 | |
| Zaliv Terpenia virus | NC_055356.1 | L | 965 | |
| Unclassified | Soft tick bunyavirus | LC027465.1 | L | 882 |
| Wanowrie virus | MK896438.1 | L | 882 | |
| Ji'an nariovirus | ON408088.1 | L | 882 | |
| Henan tick virus | MZ244224.1 | L | 882 | |
| Peribunyaviridae sp | MW721847.1 | L | 882 | |
| Nairovirus | Eyach virus | NC_003696.1 | Segment 1 | 863 |
| Kundal virus | NC_055248.1 | Segment 1 | 863 | |
| O ’hara headland virus | OL452245.1 | Segment 2 | 798 | |
| Thogoto virus | NC_006495.1 | Segment 2 | 798 | |
| Oz virus | NC_040731.1 | Segment 2 | 798 | |
| Batken virus | MN98725.1 | Segment 2 | 798 | |
| Uhori virus | NC_034263.1 | Segment 2 | 798 | |
| Jingmenvirus | Jingmen Tick Virus | NC_024113.1 | segment 1 | 795 |
| Mogiana tick virus | NC_034222.1 | segment 1 | 795 | |
| Flavivirus | Yokose virus, complete genome | NC_005039.1 | NS5 | 579 |
| Zika virus | NC_012532.1 | NS5 | 579 | |
| Zika virus | KY766069.1 | NS5 | 579 | |
| Zika virus | KX377337.1 | NS5 | 579 | |
| Zika virus | KJ776791.2 | NS5 | 579 | |
| Zika virus | NC_035889.1 | NS5 | 579 | |
| Sepik virus | NC_008719.1 | NS5 | 579 | |
| Banzi virus | NC_043110.1 | NS5 | 579 | |
| Saumarez Reef virus | NC_033726.1 | NS5 | 579 | |
| Kama virus strain | NC_023439.1 | NS5 | 579 | |
| Meaban virus | NC_033721.1 | NS5 | 579 | |
| Kadam virus | NC_033724.1 | NS5 | 579 | |
| Karshi virus | AY863002.1 | NS5 | 579 | |
| Alkhurma virus | AF331718.1 | NS5 | 579 | |
| Spanish goat encephalitis virus | NC_027709.1 | NS5 | 579 | |
| Negishi virus | KT224355.1 | NS5 | 579 | |
| Hepatitis GB virus | NC_001655.1 | NS5 | 579 | |
| Yokose virus | NC_005039.1 | NS5 | 579 | |
| Zika virus | NC_012532.1 | NS5 | 579 | |
| Zika virus isolate | KY766069.1 | NS5 | 579 |
2.4. TITAN-RNA Sequencing Workflow
RNA was converted to cDNA using SuperScript IV VILO Master Mix with ezDNase (Thermo Fischer Scientific, cat# 11766050) and converted to dsDNA using DNA Polymerase I, Large (Klenow) (NEB, Ipswich, MA, USA; cat# M0210L). Libraries were prepared using the TWIST Bioscience Library Preparation EF 2.0 with Enzymatic Fragmentation and the Illumina TruSeq-compatible Twist CD Index Adapter Set 1–96 (Twist Biosciences). Per the standard protocol, the target DNA concentration for each library preparation was set to less than or equal to 50 ng in 40 μL of nuclease-free water, and the DNA fragment target was approximately 300 bp. The final concentrations of the libraries were quantified with Qubit 1X dsDNA using Broad Range or High Sensitivity assay kits (Invitrogen; Waltham, MA, USA), and DNA concentrations measured using the Qubit Flex Fluorometer. Eight libraries were pooled at a time for multiplexing to prepare for hybrid-capture. DNA in pooled libraries was ‘captured’ according to the Twist Target Enrichment Standard Hybridization v2 protocol. The custom bait panel was used to hybridize each pool for 15 to 17 hours and sequenced on the MiSeq Sequencing System (Illumina) using the MiSeq Reagent Kit v3 (Illumina) and a read length of 151 or 251 base pairs. After sequencing, the raw reads were initially screened with the Chan Zuckerberg ID (CZID) cloud-based platform v3.0.2 (https://czid.org/; [42]). To verify the CZID positives, raw reads were trimmed with Trimmomatic v.0.39 [43] and mapped to the reference sequences using BWA-MEM2 v.2.2.1 [44, 45], on the Galaxy cloud informatics platform [46]. Reference-free assemblies were built to confirm the closest sequence match.
2.5. Analytic Validation
Linearity.
To assess the relationship between the amount of POWV RNA input versus POWV RNA detected, the, TBD_Virus portion of the TITAN-RNA panel (Table 2) was used to detect POWV lineage II RNA in 10-fold serial dilutions from 1×102 copies per microliter to 1×10−4 copies per μL, Input volumes of each dilution sample for library preparations were normalized, and so was the final pooling volume of each sample’s libraries. The input volume was calculated based on that of the 1×102 copies/μl sample. Linearity experiment replicates were performed by different personnel, on different days.
Limit of Detection (LOD).
Limit of detection testing was performed to compare the low-level detection capabilities of the TITAN-RNA (TBD_Virus) panel system in comparison to qRT-PCR, in a variety of sample matrices (water, blood, and skin biopsy) and nucleic acid spikes (dsDNA or RNA). POWV NS5 double-stranded DNA (dsDNA) was serially diluted to 0.0005, 0.00025, 0.000125, 0.0000625, 0.00003125, 0.000015625, 7.8125E-06 copies per microliter in nuclease free water. POWV NS5 double-stranded DNA (dsDNA) was serially diluted to 0.0017, 0.00085, 0.000425, 0.000213, 0.000106, 5.313×10–5, and 2.65625×10–5 copies/μl in nuclease-free water. POWV lineage II RNA (ATCC VR-3275SD) was serially diluted to 5, 2.5, 1.25, 0.8, 0.625, 0.4, 0.3125, 0.2, 0.15625, 0.078125, 0.05, 0.025 copies per microliter in nuclease free water. POWV lineage II RNA (ATCC VR-3275SD) was serially diluted to 0.225, 0.1125, 0.05625, 0.028125, 0.0140625, 0.00703125, and 0.003515625 copies per μl in nuclease free water. HRTV L and DBV L dsDNA were serially diluted to 0.0017, 0.00085, 0.000425, 0.000213, 0.000106, 5.313×10−5, and 2.65625×10−5 copies per μl in nuclease-free water.
RNA for LOD experiments was either converted to dsDNA (for hybrid capture) or used directly with TaqMan Fast Virus 1-Step Master Mix for qPCR (Thermo Fischer Scientific, cat# 4444432). Four μl of each DNA or RNA dilution was mixed with 9 μl nuclease-free water, 5 μl TaqMan Fast Virus 1-step Master Mix (4X), forward primers (10 μM), reverse primer (10 μM), and probe (10 μM). Negative controls were performed by adding 4 μl water to the PCR mixture in place of the DNA template. qRT-PCR was run on the QuantStudio™ platform (Thermo Fisher Scientific) Amplification thresholds were set at 10% of highest amplification in each run. A negative control was run with every group. POWV positive samples were identified and verified for each concentration in a total of five replicates.
Blood and ear skin notch biopsy samples from specific-pathogen-free laboratory mice (FVB/NJ, JAX #001800) that were being euthanized for an unrelated study approved by the Cornell University Institutional Animal Care and Use Committee (IACUC, Protocol 2007–0115) were used in spiking experiments to mimic the complexity of clinical samples. ATCC POWV lineage II RNA was spiked into mouse blood at 0.225, 0.1125, 0.05625, 0.028125, 0.0140625, 0.00703125, and 0.003515625 copies per μl, and mouse ear notches at 0.45, 0.225, 0.1125, 0.05625, 0.028125, 0.0140625, 0.00703125, and 0.003515625 copies per microliter. Extractions were performed using the MagMAX CORE Nucleic Acid Purification Kit (Applied Biosystems; Waltham, MA, USA) for blood samples, or the Quick DNA/RNA Pathogens MagBead Kit (Zymo Research) for the ear notch samples. The ratios of positive to negative samples were used to determine the fraction-positive value for each replicate and plotted using ggplot2 v.3.5.2 [47] in RStudio v.4.4.3 [48] or Quodata PROLab POD (Dresden, Saxony, Germany) [49]. A Sigmoidal Fitting model was fit with the x-axis as Log10 of POWV total copies and the y-axis as fraction-positive samples. The LOD was calculated as the concentration at which the fraction-positive samples decreased below 95%. A complementary log-log model was used to extrapolate the limit in blood [50, 51].
2.6. Divergent sequence tolerance.
To measure the sequence variability tolerance of the TITAN-RNA (TBD_Virus) panel system, nine of the randomly altered NS5 “variant” sequences described in 2.1 (10seq1, 10seq2, 20seq1, 20seq2, 30seq1, 30seq2, 40seq1, 50seq1, 50seq2), and NS5 of TBEV (NCBI# NC_001672.1) and ZIKV (NCBI# NC_012532.1), were resuspended to a concentration of 10 ng/μL in nuclease-free water. The final concentration of each sequence stock was measured using a Qubit Fluorometer High Sensitivity Kit, and total copies per microliter were calculated based on the sequence length. Ten-thousand total copies of each sequence were spiked into individual aliquots of 40 μL water with 50 ng of clean mouse ear notch DNA. Hybrid-capture was performed and analyzed as described in 2.4. The average coverage depth was calculated as follows:
2.7. Field discovery.
Field-based results were confirmed with both untargeted and targeted methods. Untargeted deep RNA sequencing was performed in separate laboratory (Cornell University Transcriptional Regulation and Expression Facility; TREx RRID: SCR_022532). After DNAse treatment and rRNA depletion with the RiboZero HMR kit (Illumina; San Diego, California, USA), RNA sequencing libraries were generated using the NEBNext Ultra II Directional Library Prep kit (New England Biolabs, Ipswich, MA, USA), quality checked, and 100M reads were targeted per sample using 2 × 150 bp paired-end sequencing on the Illumina NovaSeq 6000 platform with the S4 flow cell. Raw reads were trimmed, assembled, and mapped to references from the TITAN-RNA (TBD_Virus) panel using GalaxyTrakr’s Trimmomatic, rnaviralSPADES v3.15.5 [52–54], and BWA-MEM2 tools. Contigs were also screened for other viral homologies using VirBot [55]. Conventional RT-PCR or nested-RT-PCR and long-read amplicon sequencing was used to further characterize viral sequences. RT-PCR or nested-RT-PCR was performed with the SuperScript III or IV One-Step RT-PCR System (Invitrogen) with FLSV-US primers and probe listed in Table 1. Ten microliters of PCR product were run on 1% agarose gels, imaged with a ChemiDoc MP Imaging System (BioRad). Gels were extracted with QIAquick Gel Extraction Kit (Qiagen, catalog# 28704), or RT-PCR products were directly cleaned with QIAquick PCR Purification Kit (Qiagen, catalog# 28104). PCR products were quantified using Qubit 1X dsDNA High Sensitivity assay kits and concentrations measured by fluorimetry (Qubit Flex). Amplicons were barcoded using the Native Barcoding Kit 96 V14 (Oxford Nanopore; Oxford, UK; cat# SQK-NBD114.96I), prepared libraries were loaded onto a MinION R10.4.1 (FLO-MIN114) flow cells and sequenced with a MinION Mk1b Oxford Nanopore Portable Sequencing device. Raw reads were trimmed, assembled, and mapped to references from the TITAN-RNA (TBD_Virus) panel using GalaxyTrakr’s Porechop v.0.2.4 [56], Flye v.2.9.5 [57, 58], and BWA-MEM2 tools, respectively. Sanger sequencing performed by Cornell Biotechnology Resource Center. Hybrid assemblies from TITAN-RNA, Sanger, and Nanopore-based sequencing reads were built with Geneious Prime de novo assembler. Sequences were analyzed with Prokka v.1.14.6 [59, 60], and maximum likelihood phylogenetic trees made using GalaxyTrakr’s IQtree v.2.4.0 [61–67], and visualized with iTOL v.7.2 [68]. Codon Adaptation Index was calculated using the E-CAI server (http://genomes.urv.es/CAIcal/E-CAI/; [69]) using the Homo sapiens codon usage table (https://www.kazusa.or.jp/codon/cgi-bin/showcodon.cgi?species=9606).
3. Results
3. 1. Analytic validation.
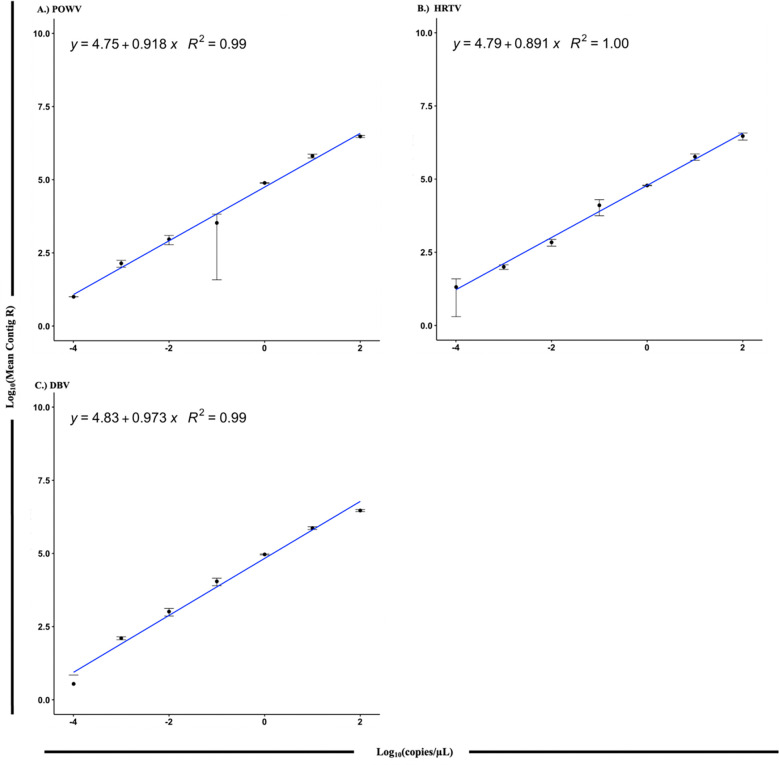
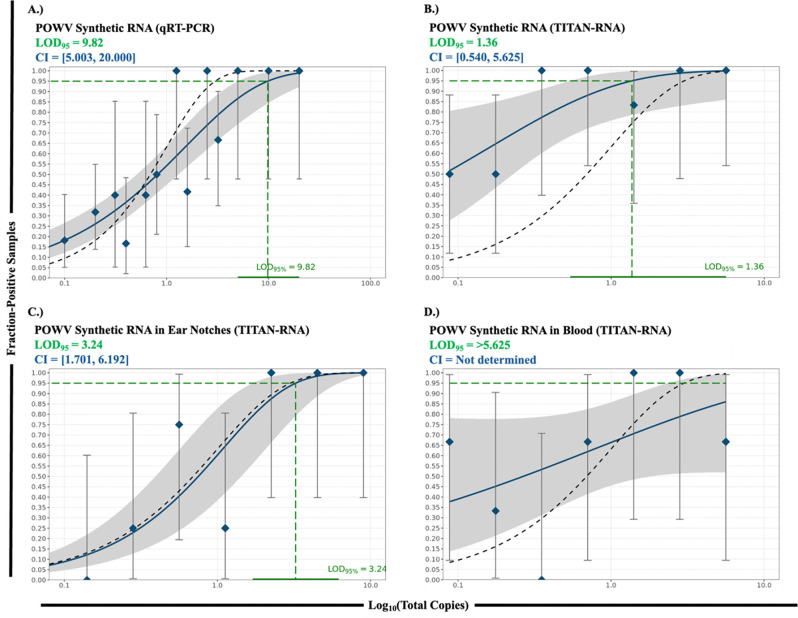
For POWV, DBV, and HRTV, a linear relationship was identified between RNA copy number and number of respective reads resulting from the hybridization capture (Fig. 2). The LODs for spikes with synthetic RNA or DNA are shown in Table 4 and Fig. 3. LOD for qRT-PCR and the TITAN-RNA (TBD_Virus) panel for synthetic POWV RNA in nuclease-free water were 9.82 and 1.36 total copies, respectively. Real-time PCR had a 7.2-fold higher LOD compared to the TITAN-RNA (TBD_Virus) panel when detecting synthetic POWV RNA. In ear notches the limit is 3.24, and in blood >5.625. Complementary log-log modelling extrapolated a limit of 19.1 in blood. The TITAN-RNA (TBD_Virus) panel further detected synthetic POWV DNA with a limit of 0.014, and qPCR detected with a limit of 0.001 total copies. The TITAN-RNA (TBD_Virus) panel detected synthetic HRTV and DBV DNA with limits of 0.023, and a limit of 0.029, respectively.
Figure 2. Linearity results.
Linearity plotted as log10 Mean Contig R (number of reads aligned per taxonomically matched contig) versus log10 Concentration (copies/μl) (n = 3). A.) POWV, B.) HRTV, C.) DBV. The X-axis represents the log10 of total copies and the Y-axis represents the log10 of mean contigs detected.
Table 4. Limit of Detection Results.
Limit of detection of POWV, HRTV, or DBV DNA by TITAN-RNA (TBD_virus).
| Sample Type | Method | LOD95 | Confidence Interval (CI) |
|---|---|---|---|
| POWV DNA | TITAN-RNA | 0.014 | 0.008, 0.024 |
| HRTV DNA | TITAN-RNA | 0.023 | 0.014, 0.034 |
| DBV DNA | TITAN-RNA | 0.029 | 0.017, 0.034 |
Figure 3. Limit of Detection Results.
Limit of detection plotted as fraction-positive samples versus total copies (n ≥ 3) A.) POWV synthetic RNA in nuclease-free water via qRT-PCR, B.) POWV synthetic RNA in nuclease-free water via TITAN-RNA (TBD_Virus), C.) POWV synthetic RNA in ear notches via TITAN-RNA (TBD_Virus), D.) POWV synthetic RNA in blood samples via TITAN RNA (TBD_Virus).
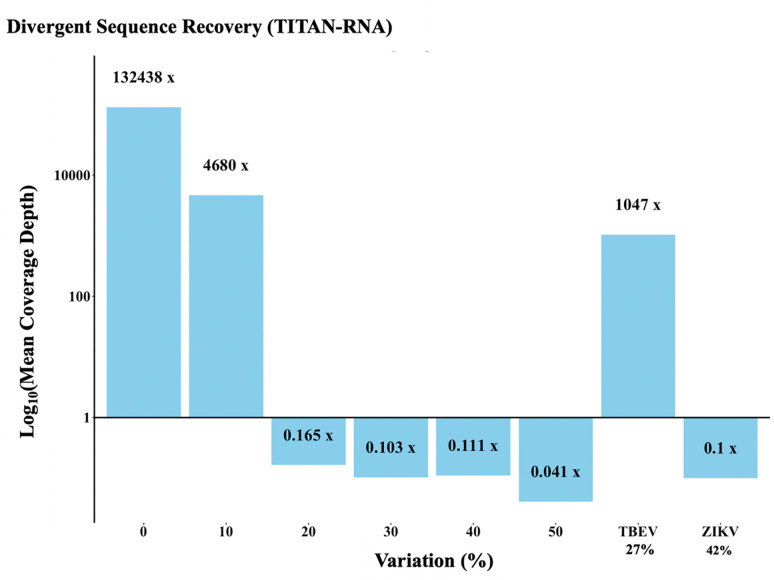
3. 2. Genetic variation Tolerance.
The TITAN-RNA (TBD_Virus) panel tolerated random sequence variability up to 10%, and natural sequence variability up to 27% (Fig. 4). With POWV NS5 as the established coverage depth control, the 10% variant had highest coverage depth among variant sequences, with TBEV NS5 (representing ~27% variance) following. The baits did not reliably capture the randomized variants with synthetic variability higher than 10% (Fig. 4). TITAN-RNA ‘captured’ an average of < 10 ZIKV reads (representing ~42% variance).
Figure 4. Mean Coverage Depth.
Log10 of mean coverage depth of NS5 using the TBD_Virus panel on random POWV sequence variants. Detection was challenged with reference sequences of two NS5 genes not included in this panel (TBEV and ZIKV) to assess the ability of POWV baits to pull down related known species.
3. 3. Viral discovery from field samples.
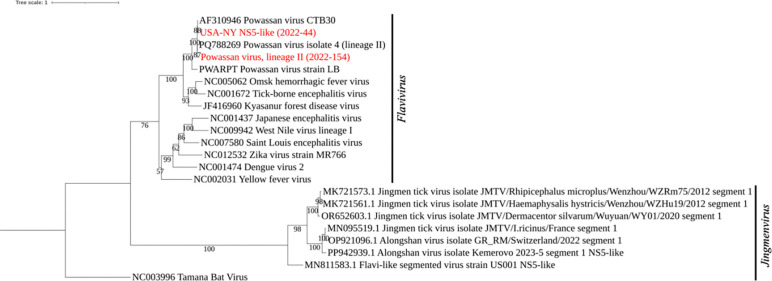
A total of 293 tick pools were screened with the TITAN-RNA panels. Ticks were pooled by species, life stage, and location collected. Fifteen I. scapularis tick pools with low levels of POWV detected by qRT-PCR were first tested with the TBD_Virus panel. Segments of the POWV genome were recovered from a pool of 5 nymphs (2022–154) from Nassau County, NY. The closest relative based on the NS5 sequence was a strain from Centre County, PA (NCBI # PQ788269) (Fig. 5). This pool had a qRT-PCR Ct value of 27.4, and 43 reads were ‘captured’ by TITAN-RNA. The other pools, with CT range 36.1–42.3, were not ‘captured’.
Figure 5. Phylogenetic analysis of tick-borne Flaviviridae RdRP (NS5 or NS5-like).
Maximum likelihood tree of segmented and unsegmented Flaviviridae RNA-dependent RNA Polymerase (NS5) homologs. Scale represents number of nucleotide substitutions. Bootstraps = 1000.
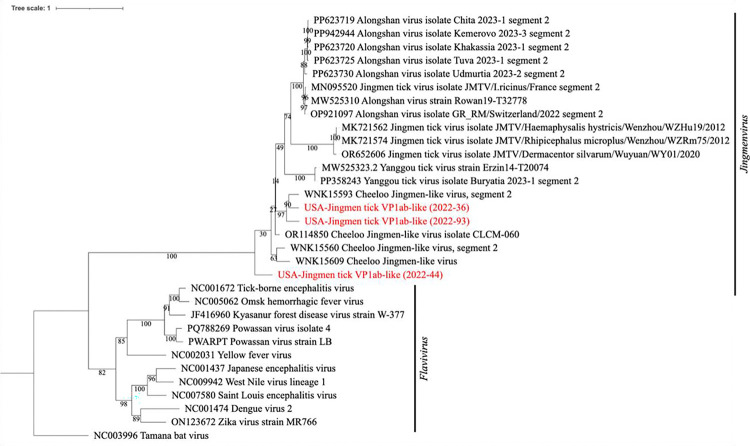
Forty-eight H. longicornis pools were then tested on the two panels combined, and select samples were validated with RNA sequencing. Three revealed VP1ab-like glycoprotein sequences of non-endemic Flavi-like Segmented Viruses (FLSV’s, Fig. 6). Alignment and phylogenetic analysis of revealed that sequences from pooled samples 2022–36 (362 bp), 2022–44 (456 bp), and 2022–93 (529 bp) clustered closest to Jingmen tick-like viruses (Fig. 6). 2022–36 contained five larvae, 2022–44 five nymphs, and 2022–93 five nymphs. Tamana bat virus is used as the outgroup [8]. The NS5 sequences recovered from sample 2022–44 (2.1 kb) by TITAN-RNA, Nanopore-based, and Sanger sequencing, however, branched alone (Fig. 5). Codon usage for these NS5 sequences was determined to be comparable to a human reference gene (Table 5), indicating likely adaptation to replication in humans. Codon adaptation for the VP1ab-like glycoprotein (segment 2) sequences was comparable to TBEV and Yellow Fever virus (YFV) reference sequences (Table 5.). From the RNA sequencing, mitochondrial sequences of dog, mouse, raccoon, and hummingbird were identified as the likely blood meal sources for reservoir investigation.
Figure 6. Phylogenetic analysis of tick-borne Flaviviridae glycoprotein (E or VP1/1ab).
Maximum likelihood tree of segmented and unsegmented Flaviviridae glycoprotein homologs. Scale represents number of nucleotide substitutions. Bootstraps = 1000.
Table 5. Codon Adaptation analysis.
Assessment of codon adaptation for replication in the human host. The expected Codon Adaptation Index (eCAI) metric at 95% confidence is shown.
| Sample or reference accession | Closest match | Gene | Average eCAI | eCAI (p<0.05) |
|---|---|---|---|---|
| 2022–154 I. scapularis | POWV | NS5 | 0.748 | 0.800 |
| 2022–93 frame 3 F H. longicornis | JMTV | VP1a-like | 0.689 | 0.747 |
| 2022–36 frame 1 R H. longicornis | 0.747 | 0.804 | ||
| 2022–44 frame 1 F H. longicornis | 0.695 | 0.751 | ||
| 2022–44 frame 1a H. longicornis | POWV | NS5-like | 0.810 | 0.865 |
| 2022–44 frame 1b H. longicornis | 0.776 | 0.828 | ||
| OP921097 | ALSV reference | VP1a | 0.724 | 0.784 |
| NC_002031 | YFV reference | full genome | 0.694 | 0.747 |
| NC_001672.1 | TBEV reference | full genome | 0.688 | 0.743 |
| NC_000006.12 | Human reference | TNF | 0.749 | 0.808 |
Finally, 230 H. longicornis pools were screened with TBD_viral_discovery_1 which probes for ZIKV NS5. One sample (2022–310) had 12 reads with the closest match having 97.6% identity to Zika virus strain MR 766 (NCBI# PP921916.1). De novo assembly revealed the longest sequence match of 120 bp with a 14–18 bp flanking the matching region with no identity to ZIKV.
4. Discussion
The TITAN-RNA (TBD_Virus) panel detected HRTV L, DBV L, and POWV NS5 with high linearity, indicating that this method is suitable for quantitative detection over a very wide dynamic range. The limit of detection results revealed that TITAN-RNA (TBD_Virus) can detect levels of spiked viral RNA lower than gold-standard qRT-PCR in nuclease-free water. Moreover, while the qRT-PCR method, limits variant detection as it utilizes target-specific primers, the TITAN-RNA could successfully detect and sequence both artificial and naturally evolved variants that were 10% and 27% divergent. The highest coverage of variant sequences was, as expected, observed in the regions with high similarity and nucleotide conservation. Coverage of less conserved sequences was attributed to both the flexible complementary binding action of the baits during hybridization and the pulldown of flanking sequences that does not depend on similarity to bait sequences.
Variation tolerance was further demonstrated using field samples of invasive ticks found to harbor a potentially novel virus species combining segmented and unsegmented flavivirus genomes, with no a prori knowledge of its existence. One of the samples (2022–44) had a 456 bp contig of RNA material for the glycoprotein of segmented Jingmen tick virus, and a highly modified RdRP from unsegmented flaviviral NS5. Hybrid-assembly of sample 2022–44 revealed a 2.2 kb contig with sequence similarity (percent identity = 90.49%, query coverage = 29%) to the NS5 of POWV (NCBI# AF310946.1) and protein similarity (percent identity = 30.30%, query coverage = 9%) to an exogenous uncharacterized protein in H. longicornis (NCBI# KAH9380833.1). The NS5 sequence was only recoverable with hybrid assembly and an extended reverse transcription, supporting its identity as part of a viral RNA genome rather than derived from the tick or a bloodmeal host genome. It is not related to the FLSV NS5 sequence discovered in mice in Pennsylvania, USA ([32], Fig. 5). Furthermore, its eCAI > 0.8 suggests strong adaptation to the human host, likely over a longer period than the glycoprotein sequence recovered from the same tick (eCAI 0.751). These initial adaptation results are based on short sequences and should be interpreted with caution.
Brown et al. (2024) recently highlighted the mystery of uncharacterized protein sequences that may code for endogenous or exogenous RdRP’s in the NCBI database and the need to annotate sequences towards higher accuracy for virus discovery [70]. The sequences identified using the TITAN-RNA panel will be added to future builds to detect additional sequence for more comprehensive evolutionary analyses. Available whole genomes of segmented and unsegmented members of the Flaviviridae family will be added, with a focus on glycoproteins (E and VP1ab) and RdRP (NS5) homologs. The TBD_Virus and TBD_virus_discovery_1 panels can be ordered through Twist Biosciences as one combined panel (TITAN-RNA). Future validation will be focused on non-flaviviral targets. Consolidating all viral baits into one panel will facilitate pooled hybrid capture sequencing with the companion bacterial/protozoal/blood meal identification panel (Wang et al., bioRxiv 2025.06.06.658355) to minimize cost and labor towards an accessible method that can be adopted in hospital and academic laboratories around the world.
Ticks are responsible for nearly 77% of vector-borne disease cases in the US, and globally tick-viruses are more prevalent in other parts of the world [71, 72]. From an ecological perspective, the prevalence of tick-borne viruses may increase with land use changes, climate change and increased host populations [23, 24]. Since tick-borne pathogens primarily infect small mammals, pathogen surveillance is necessary on small synanthropic mammals as they may serve as pathogen reservoirs and increase the risk of zoonotic spillover. Unsegmented flaviviruses like TBEV, LIV, AOHFV, KFD, and POWV are harbored by ticks and well characterized genomically. Genomic references for more recently discovered segmented Flavi-like jingmenviruses are in development [73–76]. Our results highlight the need for increased monitoring of emerging disease vectors, including I. scapularis and the invasive H. longicornis species in the eastern USA, both competent vectors of POWV [25, 77–79]. Segmented and unsegmented viruses belonging to the Flaviviridae family can cause severe disease such as encephalitis, meningitis, loss of coordination, speech difficulties, high fever, and headaches [5, 28, 29]. Development of flexible enrichment methods for high-throughput sequencing technology, like the TITAN-RNA panel, will allow for improved detection and characterization of endemic and emerging viruses, potentially ahead of their spillover to humans.
Supplementary Material
Supplementary Data Captions
“Synthetic DNA/RNA Excel File” – List of all synthetic DNA or RNA used in this study with catalog and manufacturer information.
Acknowledgments
We are thankful to Dr. Lars Eisen and Dr. Joan Kenney for critical review of the manuscript. The authors also thank the de Mestre laboratory at Cornell University for assistance with ddPCR and quo data (https://www.quodata.de/) for use of their PCR analysis tool. Sequencing equipment used for this study was provided by the United States Food and Drug Administration’s Veterinary Laboratory Investigation and Response Network under Research Collaboration Agreement FY16-RCA-CVM-01-CU and the BRC Genomics Facility (RRID: SCR_022532) at the Cornell Institute of Biotechnology.
Funding
This work was supported by The Assistant Secretary of Defense for Health Affairs through the Tick-Borne Disease Research Program, endorsed by the Department of Defense under Award No. W81XWH-22-1-0891 to LBG and MD. Opinions, interpretations, conclusions and recommendations are those of the authors and are not necessarily endorsed by the Department of Defense.
Additional support was provided by the Cornell Public and Ecosystem Health Atkinson Impact Award to LBG, AIB, and LCH; National Institute Of Allergy And Infectious Diseases of the National Institutes of Health Award Number 1P20AI186093-01 to LBG; United States Geological Survey Award Number G23AC00488-00 to LBG; and Centers for Disease Control and Prevention Cooperative Agreement U50CK000633 to LCH. This content is solely the responsibility of the authors and does not necessarily represent the official views of the Centers for Disease Control and Prevention, the United States Geological Survey, or the National Institutes of Health.
Footnotes
Conflict of Interest
The authors declare no conflicts.
Data Availability
All raw sequence data from this study are deposited in NCBI BioProject PRJNA1271117. All code is deposited in https://gitlab.com/goodmanlab/.
References:
- 1.Rodino K.G., Theel E.S., and Pritt B.S., Tick-Borne Diseases in the United States. Clinical Chemistry, 2020. 66(4): p. 537–548. [DOI] [PubMed] [Google Scholar]
- 2.Suss J., et al. , What makes ticks tick? Climate change, ticks, and tick-borne diseases. J Travel Med, 2008. 15(1): p. 39–45. [DOI] [PubMed] [Google Scholar]
- 3.Raghavan R.K., et al. , Potential Spatial Distribution of the Newly Introduced Long-horned Tick, Haemaphysalis longicornis in North America. Scientific Reports, 2019. 9(1): p. 498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Eisen R.J. and Eisen L., The Blacklegged Tick, Ixodes scapularis: An Increasing Public Health Concern. Trends in Parasitology, 2018. 34(4): p. 295–309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Jia N., et al. , Emergence of human infection with Jingmen tick virus in China: A retrospective study. EBioMedicine, 2019. 43: p. 317–324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Colmant A.M.G., Charrel R.N., and Coutard B., Jingmenviruses: Ubiquitous, understudied, segmented flavi-like viruses. Frontiers in Microbiology, 2022. Volume 13 - 2022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Emmerich P., et al. , Viral metagenomics, genetic and evolutionary characteristics of Crimean-Congo hemorrhagic fever orthonairovirus in humans, Kosovo. Infection, Genetics and Evolution, 2018. 65: p. 6–11. [DOI] [PubMed] [Google Scholar]
- 8.de Lamballerie X., et al. , Genome sequence analysis of Tamana bat virus and its relationship with the genus Flavivirus. Journal of General Virology, 2002. 83(10): p. 2443–2454. [DOI] [PubMed] [Google Scholar]
- 9.Nuttall P.A. and Labuda M., Dynamics of infection in tick vectors and at the tick–host interface, in Advances in Virus Research. 2003, Academic Press. p. 233–272. [DOI] [PubMed] [Google Scholar]
- 10.Heinz F.X., et al. , Field effectiveness of vaccination against tick-borne encephalitis. Vaccine, 2007. 25(43): p. 7559–7567. [DOI] [PubMed] [Google Scholar]
- 11.Sudhindra P., Chapter 10 - Tick-Borne Infections of the Central Nervous System, in The Microbiology of Central Nervous System Infections, Kon K. and Rai M., Editors. 2018, Academic Press. p. 173–195. [Google Scholar]
- 12.Mansfield K.L., et al. , Emerging Tick-Borne Viruses in the Twenty-First Century. Frontiers in Cellular and Infection Microbiology, 2017. Volume 7 - 2017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hartemink N. and Takken W., Trends in tick population dynamics and pathogen transmission in emerging tick-borne pathogens in Europe: an introduction. Experimental and Applied Acarology, 2016. 68(3): p. 269–278. [DOI] [PubMed] [Google Scholar]
- 14.Ates F., et al. , Attitudes towards the Tick-Borne Encephalitis Vaccine among Children’s Guardians: A Cross-Sectional Survey Study in Poland. Vaccines, 2024. 12(8): p. 918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kunze M., et al. , Recommendations to Improve Tick-Borne Encephalitis Surveillance and Vaccine Uptake in Europe. Microorganisms, 2022. 10(7): p. 1283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Pilz A., Erber W., and Schmitt H.-J., Vaccine uptake in 20 countries in Europe 2020: Focus on tick-borne encephalitis (TBE). Ticks and Tick-borne Diseases, 2023. 14(1): p. 102–059. [DOI] [PubMed] [Google Scholar]
- 17.Erber W. and Schmitt H.-J., Self-reported tick-borne encephalitis (TBE) vaccination coverage in Europe: Results from a cross-sectional study. Ticks and Tick-borne Diseases, 2018. 9(4): p. 768–777. [DOI] [PubMed] [Google Scholar]
- 18.Miazga W., et al. , The long-term efficacy of tick-borne encephalitis vaccines available in Europe - a systematic review. BMC Infectious Diseases, 2023. 23(1): p. 621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Lindgren E. and Gustafson R., Tick-borne encephalitis in Sweden and climate change. The Lancet, 2001. 358(9275): p. 16–18. [DOI] [PubMed] [Google Scholar]
- 20.Kriz B., et al. , Epidemiology of Tick-Borne Encephalitis in the Czech Republic 1970–2008. Vector-Borne and Zoonotic Diseases, 2012. 12(11): p. 994–999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.KOROTKOV Y., KOZLOVA T., and KOZLOVSKAYA L., Observations on changes in abundance of questing Ixodes ricinus, castor bean tick, over a 35-year period in the eastern part of its range (Russia, Tula region). Medical and Veterinary Entomology, 2015. 29(2): p. 129–136. [DOI] [PubMed] [Google Scholar]
- 22.Tokarevich N.K., et al. , The impact of climate change on the expansion of Ixodes persulcatus habitat and the incidence of tick-borne encephalitis in the north of European Russia. Global Health Action, 2011. 4(1): p. 8448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Diuk-Wasser M.A., VanAcker M.C., and Fernandez M.P., Impact of Land Use Changes and Habitat Fragmentation on the Eco-epidemiology of Tick-Borne Diseases. Journal of Medical Entomology, 2020. 58(4): p. 1546–1564. [DOI] [PubMed] [Google Scholar]
- 24.Ogden N.H., et al. , Possible Effects of Climate Change on Ixodid Ticks and the Pathogens They Transmit: Predictions and Observations. Journal of Medical Entomology, 2020. 58(4): p. 1536–1545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Vogels C.B.F., et al. , Phylogeographic reconstruction of the emergence and spread of Powassan virus in the northeastern United States. Proc Natl Acad Sci U S A, 2023. 120(16): p. e2218012120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.CDC. Current year data 2024. May 13, 2025; Available from: https://www.cdc.gov/powassan/datahttps://www.cdc.gov/powassan/data-maps/current-year-data.html-printmaps/current-year-data.html#print.
- 27.Kakoullis L., et al. , Powassan Virus Infections: A Systematic Review of Published Cases. Tropical Medicine and Infectious Disease, 2023. 8(12): p. 508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ebel G.D., Update on Powassan virus: emergence of a North American tick-borne flavivirus. Annu Rev Entomol, 2010. 55: p. 95–110. [DOI] [PubMed] [Google Scholar]
- 29.Birge J. and Sonnesyn S., Powassan virus encephalitis, Minnesota, USA. Emerg Infect Dis, 2012. 18(10): p. 1669–71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Hermance M.E. and Thangamani S., Powassan Virus: An Emerging Arbovirus of Public Health Concern in North America. Vector-Borne and Zoonotic Diseases, 2017. 17(7): p. 453–462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Krow-Lucal E.R., et al. , Powassan Virus Disease in the United States, 2006–2016. Vector-Borne and Zoonotic Diseases, 2018. 18(6): p. 286–290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Vandegrift K., et al. , Presence of Segmented Flavivirus Infections in North America. Emerging Infectious Disease journal, 2020. 26(8): p. 1810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.La Scola B., et al. , Gene-sequence-based criteria for species definition in bacteriology: the Bartonella paradigm. Trends Microbiol, 2003. 11(7): p. 318–21. [DOI] [PubMed] [Google Scholar]
- 34.Gaudin M. and Desnues C., Hybrid Capture-Based Next Generation Sequencing and Its Application to Human Infectious Diseases. Front Microbiol, 2018. 9: p. 2924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Meng Z., et al. , A novel fast hybrid capture sequencing method for high-efficiency common human coronavirus whole-genome acquisition. mSystems, 2024. 9(5): p. e01222–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Quek Z.B.R. and Ng S.H. Hybrid-Capture Target Enrichment in Human Pathogens: Identification, Evolution, Biosurveillance, and Genomic Epidemiology. Pathogens, 2024. 13, DOI: 10.3390/pathogens13040275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Foley N., et al. , New York State Tick Blitz: harnessing community-based science to understand range expansion of ticks. Journal of Medical Entomology, 2023. 60(4): p. 708–717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Ayers M., et al. , A single tube RT-PCR assay for the detection of mosquito-borne flaviviruses. Journal of Virological Methods, 2006. 135(2): p. 235–239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yuan Q., et al. , Active surveillance of pathogens from ticks collected in New York State suburban parks and schoolyards. Zoonoses and Public Health, 2020. 67(6): p. 684–696. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Edgar R.C., MUSCLE v5 enables improved estimates of phylogenetic tree confidence by ensemble bootstrapping. BioRxiv, 2021: p. 2021.06. 20.449169. [Google Scholar]
- 41.Sayers Eric W., et al. , Database resources of the National Center for Biotechnology Information in 2025. Nucleic Acids Research, 2025. 53(D1): p. D20–D29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Simmonds S.E., et al. , CZ ID: a cloud-based, no-code platform enabling advanced long read metagenomic analysis. bioRxiv, 2024: p. 2024.02.29.579666. [Google Scholar]
- 43.Bolger A.M., Lohse M., and Usadel B., Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics, 2014. 30(15): p. 2114–2120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Li H. and Durbin R., Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics, 2009. 25(14): p. 1754–1760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Li H. and Durbin R., Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics, 2010. 26(5): p. 589–595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Community T.G., The Galaxy platform for accessible, reproducible, and collaborative data analyses: 2024 update. Nucleic Acids Research, 2024. 52(W1): p. W83–W94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Wickham H., ggplot2: Elegant Graphics for Data Analysis. Springer-Verlag; New York. 2016. [Google Scholar]
- 48.RStudio, RStudio: Integrated Development Environment for R. Posit, 2025. [Google Scholar]
- 49.Uhlig S., et al. , Validation of qualitative PCR methods on the basis of mathematical–statistical modelling of the probability of detection. Accreditation and Quality Assurance, 2015. 20(2): p. 75–83. [Google Scholar]
- 50.Chen M.-H., D.D. K., and Q.-M. and Shao A New Skewed Link Model for Dichotomous Quantal Response Data. Journal of the American Statistical Association, 1999. 94(448): p. 1172–1186. [Google Scholar]
- 51.Alves J.S.B., Bazán J.L., and Arellano-Valle R.B., Flexible cloglog links for binomial regression models as an alternative for imbalanced medical data. Biometrical Journal, 2023. 65(3): p. 2100325. [DOI] [PubMed] [Google Scholar]
- 52.Antipov D., et al. , hybridSPAdes: an algorithm for hybrid assembly of short and long reads. Bioinformatics, 2015. 32(7): p. 1009–1015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Vasilinetc I., et al. , Assembling short reads from jumping libraries with large insert sizes. Bioinformatics, 2015. 31(20): p. 3262–3268. [DOI] [PubMed] [Google Scholar]
- 54.Prjibelski A.D., et al. , ExSPAnder: a universal repeat resolver for DNA fragment assembly. Bioinformatics, 2014. 30(12): p. i293–i301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Chen G., et al. , VirBot: an RNA viral contig detector for metagenomic data. Bioinformatics, 2023. 39(3). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Wick R., Porechop. Github repository, 2017. github. [Google Scholar]
- 57.Lin Y., et al. , Assembly of long error-prone reads using de Bruijn graphs. Proceedings of the National Academy of Sciences, 2016. 113(52): p. E8396–E8405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Kolmogorov M., Flye. GitHub repository, 2021. GitHub. [Google Scholar]
- 59.Cuccuru G., et al. , Orione, a web-based framework for NGS analysis in microbiology. Bioinformatics, 2014. 30(13): p. 1928–1929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Seemann T., Prokka: rapid prokaryotic genome annotation. Bioinformatics, 2014. 30(14): p. 2068–2069. [DOI] [PubMed] [Google Scholar]
- 61.Kalyaanamoorthy S., et al. , ModelFinder: fast model selection for accurate phylogenetic estimates. Nature Methods, 2017. 14(6): p. 587–589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Minh B.Q., Nguyen M.A.T., and von Haeseler A., Ultrafast Approximation for Phylogenetic Bootstrap. Molecular Biology and Evolution, 2013. 30(5): p. 1188–1195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Schrempf D., et al. , Reversible polymorphism-aware phylogenetic models and their application to tree inference. Journal of Theoretical Biology, 2016. 407: p. 362–370. [DOI] [PubMed] [Google Scholar]
- 64.Chernomor O., von Haeseler A., and Minh B.Q., Terrace Aware Data Structure for Phylogenomic Inference from Supermatrices. Systematic Biology, 2016. 65(6): p. 997–1008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Wang H.-C., et al. , Modeling Site Heterogeneity with Posterior Mean Site Frequency Profiles Accelerates Accurate Phylogenomic Estimation. Systematic Biology, 2017. 67(2): p. 216–235. [DOI] [PubMed] [Google Scholar]
- 66.Hoang D.T., et al. , UFBoot2: Improving the Ultrafast Bootstrap Approximation. Molecular Biology and Evolution, 2017. 35(2): p. 518–522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Nguyen L.-T., et al. , IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Molecular Biology and Evolution, 2014. 32(1): p. 268–274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Letunic I. and Bork P., Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Research, 2024. 52(W1): p. W78–W82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Puigbò P., Bravo I.G., and Garcia-Vallve S., CAIcal: A combined set of tools to assess codon usage adaptation. Biology Direct, 2008. 3(1): p. 38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Brown K. and Firth A.E., Uncovering hundreds of exogenous and endogenous RNA viral RdRp sequences amongst uncharacterised sequences in public protein databases. bioRxiv, 2024: p. 2024.09.25.614983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Rochlin I. and Toledo A., Emerging tick-borne pathogens of public health importance: a mini-review. Journal of Medical Microbiology, 2020. 69(6): p. 781–791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Rosenberg R., et al. , Vital Signs: Trends in Reported Vectorborne Disease Cases - United States and Territories, 2004–2016. MMWR Morb Mortal Wkly Rep, 2018. 67(17): p. 496–501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Cicculli V., et al. , First detection of Jingmen tick virus in Corsica with a new generic RTqPCR system. npj Viruses, 2024. 2(1): p. 44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Dinçer E., et al. Survey and Characterization of Jingmen Tick Virus Variants. Viruses, 2019. 11, DOI: 10.3390/v11111071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Ogola E.O., et al. Jingmen Tick Virus in Ticks from Kenya. Viruses, 2022. 14, DOI: 10.3390/v14051041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76. <.j/>Wu Z., et al. , Jingmen tick virus: an emerging arbovirus with a global threat. mSphere, 2023. 8(5): p. e00281–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Rochlin I., et al. , Rapid invasion and expansion of the Asian longhorned tick (Haemaphysalis longicornis) into a new area on Long Island, New York, USA. Ticks and Tick-borne Diseases, 2023. 14(2): p. 102088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Raney W.R., et al. , Horizontal and Vertical Transmission of Powassan Virus by the Invasive Asian Longhorned Tick, Haemaphysalis longicornis, Under Laboratory Conditions. Frontiers in Cellular and Infection Microbiology, 2022. Volume 12 - 2022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Obellianne C., et al. , Interspecies co-feeding transmission of Powassan virus between a native tick, Ixodes scapularis, and the invasive East Asian tick, Haemaphysalis longicornis. Parasites & Vectors, 2024. 17(1): p. 259. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Data Captions
“Synthetic DNA/RNA Excel File” – List of all synthetic DNA or RNA used in this study with catalog and manufacturer information.
Data Availability Statement
All raw sequence data from this study are deposited in NCBI BioProject PRJNA1271117. All code is deposited in https://gitlab.com/goodmanlab/.