Abstract
We present a genome assembly from an individual male specimen of Crioceris asparagi (Asparagus Beetle; Arthropoda; Insecta; Coleoptera; Chrysomelidae). The genome sequence has a total length of 639.30 megabases. Most of the assembly (93.04%) is scaffolded into 9 chromosomal pseudomolecules, including the X and Y sex chromosomes. The mitochondrial genome has also been assembled and is 15.76 kilobases in length. Gene annotation of this assembly on Ensembl identified 26,673 protein-coding genes.
Keywords: Crioceris asparagi, Asparagus Beetle, genome sequence, chromosomal, Coleoptera
Species taxonomy
Eukaryota; Opisthokonta; Metazoa; Eumetazoa; Bilateria; Protostomia; Ecdysozoa; Panarthropoda; Arthropoda; Mandibulata; Pancrustacea; Hexapoda; Insecta; Dicondylia; Pterygota; Neoptera; Endopterygota; Coleoptera; Polyphaga; Cucujiformia; Chrysomeloidea; Chrysomelidae; Criocerinae; Crioceris; Crioceris asparagi (Linnaeus, 1758) (NCBI:txid131627)
Background
Crioceris asparagi (Linnaeus, 1758), commonly known as the “Asparagus Beetle” is one of about 250 species of the leaf beetle family (Chrysomelidae) found on the British Isles ( Duff, 2018). Chrysomelidae are one of the largest families of Coleoptera, with over 40,000 currently valid species. Crioceris Geoffroy, 1762 is the type genus of the subfamily Criocerinae, represented by seven species in Britain ( Duff, 2018) and about 1500 worldwide ( Monrós, 1960; Vencl & Leschen, 2014). Crioceris is native to the Palaearctic, Oriental and Afrotropical regions, counting about 30 described species ( Bezděk & Sekerka, 2024; Heinze, 1962; Kimoto & Gressitt, 1979; Monrós, 1960). C. asparagi is currently the only Crioceris on the British checklist, though occasional records of two more species, C. duodecimpunctata (Linnaeus, 1758) and C. macilenta Weise, 1881, exist. The latter two are considered non-established accidental introductions ( Duff, 2018).
Adults and larvae of C. asparagi feed externally and conspicuously on the leaves and stems of wild, and more often cultivated, asparagus ( Asparagus spp.: Asparagaceae) and can be a pest ( Rheinheimer & Hassler, 2018). Pupation takes place in the soil at the base of the plant. Immature stages are described by Steinhausen ( 1994; 2001).
The native distribution of C. asparagi includes almost all of Europe except the far North, the Middle East, Central Asia and Siberia ( Bezděk & Sekerka, 2024). It has been an important non-native pest of asparagus in the USA and Canada for more than a century ( Chittenden, 1917).
Cox (2007) shows it to be widespread in England, but with only scarce records towards the North and West. He mentions a few scattered records in Wales, but considers it absent from Scotland and Northern Ireland. He also observes that its British distribution much more closely matches the range of cultivated asparagus, than the more coastal distribution of the uncommon wild asparagus. NBN Atlas Partnership (2024) shows that C. asparagi has apparently increased both its range and abundance, now also occurring in a few places in Scotland, but remains very scarce in Wales. Whether it is indigenous to the UK is still a matter of speculation, but Fowler (1890) believed it to be, so if it is an introduction, it is a long-standing one.
C. asparagi cannot easily be confused with any other British leaf beetle. Its cylindrical shape is somewhat atypical for Chrysomelidae, but not unusual for a Criocerinae. The elegant blue metallic or shiny black ladder-like elytral pattern on top of a whitish background, with reddish elytral margins and pronotum is very much unique. Variations with reduced or expanded elytral pattern exist ( Warchałowski, 2010), but those individuals are still easily recognisable among other Criocerinae. Specimens with a dark spot in the middle of the red pronotum seem to be unique to the eastern Mediterranean and Middle East and are sometimes classified as subspecies maculipes (Germar, 1834). Specimens with more coarsely punctate pronotum from Central Asia have previously been assigned to another subspecies, turkestanica Medvedev, 1955 (a synonym according to Bezděk & Sekerka, 2024). In the British fauna, only the nominate subspecies has been recorded ( Duff, 2016; Warchałowski, 2010).
The genome of the Asparagus Beetle, Crioceris asparagi, was sequenced as part of the Darwin Tree of Life Project, a collaborative effort to sequence all named eukaryotic species in the Atlantic Archipelago of Britain and Ireland. Here we present a chromosomally complete genome sequence for Crioceris asparagi, based on a male specimen collected from cultivated asparagus at an allotment at Winsford Gardens, Penge, England, United Kingdom ( Figure 1). This appears to be the first genome for a member of the subfamily Criocerinae to be made available.
Figure 1. Photographs of the Crioceris asparagi (icCriAspa1) specimen used for genome sequencing.
Genome sequence report
The genome of Crioceris asparagi ( Figure 1) was sequenced using Pacific Biosciences single-molecule HiFi long reads, generating a total of 25.24 Gb (gigabases) from 2.11 million reads, providing an estimated 36-fold coverage. Primary assembly contigs were scaffolded with chromosome conformation Hi-C data, which produced 164.92 Gb from 1,092.17 million reads. Specimen and sequencing details are summarised in Table 1.
Table 1. Specimen and sequencing data for Crioceris asparagi.
| Project information | |||
|---|---|---|---|
| Study title | Crioceris asparagi (common asparagus beetle) | ||
| Umbrella BioProject | PRJEB59077 | ||
| Species | Crioceris asparagi | ||
| BioSample | SAMEA111458050 | ||
| NCBI taxonomy ID | 131627 | ||
| Specimen information | |||
| Technology | ToLID |
BioSample
accession |
Organism part |
| PacBio long read sequencing | icCriAspa1 | SAMEA111458352 | abdomen |
| Hi-C sequencing | icCriAspa1 | SAMEA111458147 | Head and thorax |
| Sequencing information | |||
| Platform | Run accession | Read count | Base count (Gb) |
| Hi-C Illumina NovaSeq 6000 | ERR10802453 | 1.09e+09 | 164.92 |
| PacBio Sequel IIe | ERR10802387 | 2.11e+06 | 25.24 |
Assembly errors were corrected by manual curation, including 12 missing joins or mis-joins. This reduced the scaffold number by 1.25% and increased the scaffold N50 by 106.78%. The final assembly has a total length of 639.30 Mb in 709 sequence scaffolds, with 161 gaps, and a scaffold N50 of 90.6 Mb ( Table 2).
Table 2. Genome assembly data for Crioceris asparagi, icCriAspa1.1.
| Genome assembly | ||
|---|---|---|
| Assembly name | icCriAspa1.1 | |
| Assembly accession | GCA_958507055.1 | |
| Accession of alternate haplotype | GCA_958501965.1 | |
| Span (Mb) | 639.30 | |
| Number of contigs | 871 | |
| Number of scaffolds | 709 | |
| Longest scaffold (Mb) | 131.42 | |
| Assembly metrics * | Benchmark | |
| Contig N50 length (Mb) | 5.5 | ≥ 1 Mb |
| Scaffold N50 length (Mb) | 90.6 | = chromosome N50 |
| Consensus quality (QV) | 58.8 | ≥ 40 |
| k-mer completeness | primary: 90.19%; alternate:
61.69%; combined: 98.64% |
≥ 95% |
| BUSCO ** | C:98.4%[S:97.6%,D:0.8%],
F:0.2%,M:1.4%,n:2,124 |
S > 90%, D < 5% |
| Percentage of assembly mapped
to chromosomes |
93.04% | ≥ 90% |
| Sex chromosomes | XY | localised homologous pairs |
| Organelles | Mitochondrial genome: 15.76 kb | complete single alleles |
| Genome annotation of assembly GCA_958507055.1 at Ensembl | ||
| Number of protein-coding genes | 26,673 | |
| Number of gene transcripts | 26,898 | |
* Assembly metric benchmarks are adapted from Rhie et al. (2021) and the Earth BioGenome Project Report on Assembly Standards September 2024.
** BUSCO scores based on the endopterygota_odb10 BUSCO set using version 5.3.2. C = complete [S = single copy, D = duplicated], F = fragmented, M = missing, n = number of orthologues in comparison. A full set of BUSCO scores is available at https://blobtoolkit.genomehubs.org/view/icCriAspa1_1/dataset/icCriAspa1_1/busco.
The snail plot in Figure 2 provides a summary of the assembly statistics, indicating the distribution of scaffold lengths and other assembly metrics. Figure 3 shows the distribution of scaffolds by GC proportion and coverage. Figure 4 presents a cumulative assembly plot, with separate curves representing different scaffold subsets assigned to various phyla, illustrating the completeness of the assembly.
Figure 2. Genome assembly of Crioceris asparagi, icCriAspa1.1: metrics.
The BlobToolKit snail plot provides an overview of assembly metrics and BUSCO gene completeness. The circumference represents the length of the whole genome sequence, and the main plot is divided into 1,000 equal-sized bins around the circumference. The outermost blue tracks display the distribution of GC, AT, and N percentages across the bins. Scaffolds are arranged clockwise from longest to shortest and are depicted in dark grey. The longest scaffold is indicated by the red arc, and the deeper orange and pale orange arcs represent the N50 and N90 lengths. A light grey spiral at the centre shows the cumulative scaffold count on a logarithmic scale. A summary of complete, fragmented, duplicated, and missing BUSCO genes in the endopterygota_odb10 set is presented at the top right. An interactive version of this figure is available at https://blobtoolkit.genomehubs.org/view/icCriAspa1_1/dataset/icCriAspa1_1/snail.
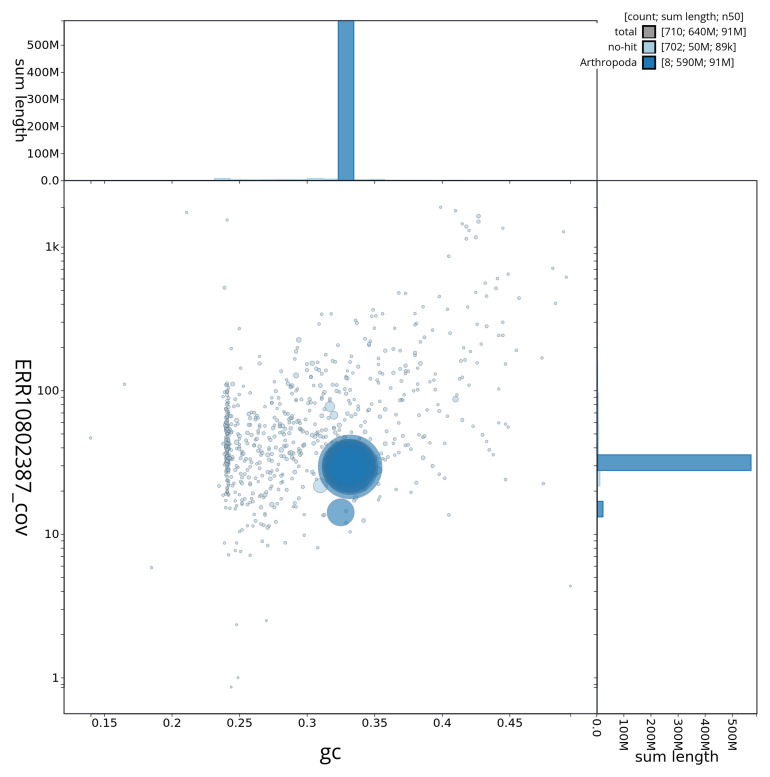
Figure 3. Genome assembly of Crioceris asparagi, icCriAspa1.1: BlobToolKit GC-coverage plot.
BlobToolKit GC-coverage plot showing sequence coverage (vertical axis) and GC content (horizontal axis). The circles represent scaffolds, with the size proportional to scaffold length and the colour representing phylum membership. The histograms along the axes display the total length of sequences distributed across different levels of coverage and GC content. An interactive version of this figure is available at https://blobtoolkit.genomehubs.org/view/icCriAspa1_1/dataset/icCriAspa1_1/blob.
Figure 4. Genome assembly of Crioceris asparagi icCriAspa1.1: BlobToolKit cumulative sequence plot.
The grey line shows cumulative length for all sequences. Coloured lines show cumulative lengths of sequences assigned to each phylum using the buscogenes taxrule. An interactive version of this figure is available at https://blobtoolkit.genomehubs.org/view/icCriAspa1_1/dataset/icCriAspa1_1/cumulative.
Most of the assembly sequence (93.04%) was assigned to 9 chromosomal-level scaffolds, representing 7 autosomes and the X and Y sex chromosomes. These chromosome-level scaffolds, confirmed by the Hi-C data, are named in order of size ( Figure 5; Table 3). During manual curation it was noted that Chromosomes X and Y were assigned based on read coverage statistics. While not fully phased, the assembly deposited is of one haplotype. Contigs corresponding to the second haplotype have also been deposited. The mitochondrial genome was also assembled and can be found as a contig within the multifasta file of the genome submission, and as a separate fasta file with accession OY293835.1.
Figure 5. Genome assembly of Crioceris asparagi icCriAspa1.1: Hi-C contact map of the icCriAspa1.1 assembly, visualised using HiGlass.
Chromosomes are shown in order of size from left to right and top to bottom. An interactive version of this figure may be viewed at https://genome-note-higlass.tol.sanger.ac.uk/l/?d=OtktpgA8SYaTECusDNb9xg.
Table 3. Chromosomal pseudomolecules in the genome assembly of Crioceris asparagi, icCriAspa1.
| INSDC
accession |
Name | Length
(Mb) |
GC% |
|---|---|---|---|
| OY293826.1 | 1 | 131.42 | 33.0 |
| OY293827.1 | 2 | 97.98 | 33.0 |
| OY293828.1 | 3 | 90.62 | 33.0 |
| OY293829.1 | 4 | 82.3 | 33.0 |
| OY293830.1 | 5 | 69.17 | 33.0 |
| OY293831.1 | 6 | 48.75 | 33.0 |
| OY293832.1 | 7 | 47.7 | 33.0 |
| OY293833.1 | X | 21.83 | 32.5 |
| OY293834.1 | Y | 5.05 | 31.0 |
| OY293835.1 | MT | 0.02 | 24.0 |
The final assembly has a Quality Value (QV) of 58.8 and k-mer completeness of 98.64% for the combined assembly. BUSCO (v5.4.3) analysis using the endopterygota_odb10 reference set ( n = 2,124) indicated a completeness score of 98.4% (single = 97.6%, duplicated = 0.8%). The assembly achieves the EBP reference standard of 6.C.59. Other quality metrics are given in Table 2.
Genome annotation report
The Crioceris asparagi genome assembly (GCA_958507055.1) was annotated at the European Bioinformatics Institute (EBI) on Ensembl Rapid Release. The resulting annotation includes 26,898 transcribed mRNAs from 26,673 protein-coding genes ( Table 2; https://rapid.ensembl.org/Crioceris_asparagi_GCA_958507055.1/Info/Index). The average transcript length is 4,132.90, with an average of 3.55 exons per transcript.
Methods
Sample acquisition and DNA barcoding
An adult male specimen of Crioceris asparagi (specimen ID NHMUK014443406, ToLID icCriAspa1) was collected from Winsford Gardens, Penge, England, United Kingdom (latitude 51.41, longitude –0.06) on 2021-07-03. The specimen was collected and identified by Michael Geiser (Natural History Museum) and preserved by dry freezing at –80 °C.
The initial identification was verified by an additional DNA barcoding process according to the framework developed by Twyford et al. (2024). A small sample was dissected from the specimens and stored in ethanol, while the remaining parts were shipped on dry ice to the Wellcome Sanger Institute (WSI). The tissue was lysed, the COI marker region was amplified by PCR, and amplicons were sequenced and compared to the BOLD database, confirming the species identification ( Crowley et al., 2023). Following whole genome sequence generation, the relevant DNA barcode region was also used alongside the initial barcoding data for sample tracking at the WSI ( Twyford et al., 2024). The standard operating procedures for Darwin Tree of Life barcoding have been deposited on protocols.io ( Beasley et al., 2023).
Nucleic acid extraction
The workflow for high molecular weight (HMW) DNA extraction at the Wellcome Sanger Institute (WSI) Tree of Life Core Laboratory includes a sequence of procedures: sample preparation and homogenisation, DNA extraction, fragmentation and purification. Detailed protocols are available on protocols.io ( Denton et al., 2023b).
The icCriAspa1 sample was prepared for DNA extraction by weighing and dissecting it on dry ice ( Jay et al., 2023). Tissue from the abdomen was homogenised using a PowerMasher II tissue disruptor ( Denton et al., 2023a).
HMW DNA was extracted in the WSI Scientific Operations core using the Automated MagAttract v2 protocol ( Oatley et al., 2023). The DNA was sheared into an average fragment size of 12–20 kb in a Megaruptor 3 system ( Bates et al., 2023). Sheared DNA was purified by solid-phase reversible immobilisation, using AMPure PB beads to eliminate shorter fragments and concentrate the DNA ( Strickland et al., 2023). The concentration of the sheared and purified DNA was assessed using a Nanodrop spectrophotometer and Qubit Fluorometer using the Qubit dsDNA High Sensitivity Assay kit. Fragment size distribution was evaluated by running the sample on the FemtoPulse system.
Hi-C preparation
Tissue from the head and thorax of the icCriAspa1 sample was processed at the WSI Scientific Operations core, using the Arima-HiC v2 kit. Tissue (stored at –80 °C) was fixed, and the DNA crosslinked using a TC buffer with 22% formaldehyde. After crosslinking, the tissue was homogenised using the Diagnocine Power Masher-II and BioMasher-II tubes and pestles. Following the kit manufacturer's instructions, crosslinked DNA was digested using a restriction enzyme master mix. The 5’-overhangs were then filled in and labelled with biotinylated nucleotides and proximally ligated. An overnight incubation was carried out for enzymes to digest remaining proteins and for crosslinks to reverse. A clean up was performed with SPRIselect beads prior to library preparation.
Library preparation and sequencing
Library preparation and sequencing were performed at the WSI Scientific Operations core. Pacific Biosciences HiFi circular consensus DNA sequencing libraries were prepared using the PacBio Express Template Preparation Kit v2.0 (Pacific Biosciences, California, USA) as per the manufacturer's instructions. The kit includes the reagents required for removal of single-strand overhangs, DNA damage repair, end repair/A-tailing, adapter ligation, and nuclease treatment. Library preparation also included a library purification step using AMPure PB beads (Pacific Biosciences, California, USA) and size selection step to remove templates shorter than 3 kb using AMPure PB modified SPRI. DNA concentration was quantified using the Qubit Fluorometer v2.0 and Qubit HS Assay Kit and the final library fragment size analysis was carried out using the Agilent Femto Pulse Automated Pulsed Field CE Instrument and 165kb gDNA and 55kb BAC analysis kit. Samples were sequenced using the Sequel IIe system (Pacific Biosciences, California, USA). The concentration of the library loaded onto the Sequel IIe was in the range 40–135 pM. The SMRT link software, a PacBio web-based end-to-end workflow manager, was used to set-up and monitor the run, as well as perform primary and secondary analysis of the data upon completion.
For Hi-C library preparation, DNA was fragmented to a size of 400 to 600 bp using a Covaris E220 sonicator. The DNA was then enriched, barcoded, and amplified using the NEBNext Ultra II DNA Library Prep Kit following manufacturers’ instructions. The Hi-C sequencing was performed using paired-end sequencing with a read length of 150 bp on an Illumina NovaSeq 6000 instrument.
Genome assembly, curation and evaluation
Assembly
The HiFi reads were first assembled using Hifiasm ( Cheng et al., 2021) with the --primary option. Haplotypic duplications were identified and removed using purge_dups ( Guan et al., 2020). The Hi-C reads were mapped to the primary contigs using bwa-mem2 ( Vasimuddin et al., 2019). The contigs were further scaffolded using the provided Hi-C data ( Rao et al., 2014) in YaHS ( Zhou et al., 2023) using the --break option for handling potential misassemblies. The scaffolded assemblies were evaluated using Gfastats ( Formenti et al., 2022), BUSCO ( Manni et al., 2021) and MERQURY.FK ( Rhie et al., 2020).
The mitochondrial genome was assembled using MitoHiFi ( Uliano-Silva et al., 2023), which runs MitoFinder ( Allio et al., 2020) and uses these annotations to select the final mitochondrial contig and to ensure the general quality of the sequence.
Assembly curation
The assembly was decontaminated using the Assembly Screen for Cobionts and Contaminants (ASCC) pipeline (article in preparation). Flat files and maps used in curation were generated in TreeVal ( Pointon et al., 2023). Manual curation was primarily conducted using PretextView ( Harry, 2022), with additional insights provided by JBrowse2 ( Diesh et al., 2023) and HiGlass ( Kerpedjiev et al., 2018). Scaffolds were visually inspected and corrected as described by Howe et al. (2021). Any identified contamination, missed joins, and mis-joins were corrected, and duplicate sequences were tagged and removed. The curation process is documented at https://gitlab.com/wtsi-grit/rapid-curation (article in preparation).
Evaluation of the final assembly
A Hi-C map for the final assembly was produced using bwa-mem2 ( Vasimuddin et al., 2019) in the Cooler file format ( Abdennur & Mirny, 2020). To assess the assembly metrics, the k-mer completeness and QV consensus quality values were calculated in Merqury ( Rhie et al., 2020). This work was done using Nextflow ( Di Tommaso et al., 2017) DSL2 pipelines “sanger-tol/readmapping” ( Surana et al., 2023a) and “sanger-tol/genomenote” ( Surana et al., 2023b). The genome was analysed within the BlobToolKit environment ( Challis et al., 2020) and BUSCO scores ( Manni et al., 2021) were calculated.
The genome assembly and evaluation pipelines were developed using nf-core tooling ( Ewels et al., 2020) and MultiQC ( Ewels et al., 2016), relying on the Conda package manager, the Bioconda initiative ( Grüning et al., 2018), the Biocontainers infrastructure ( da Veiga Leprevost et al., 2017), as well as the Docker ( Merkel, 2014) and Singularity ( Kurtzer et al., 2017) containerisation solutions.
Table 4 contains a list of relevant software tool versions and sources.
Table 4. Software tools: versions and sources.
Genome annotation
The BRAKER2 pipeline ( Brůna et al., 2021) was used in the default protein mode to generate annotation for the Crioceris asparagi assembly (GCA_958507055.1) in Ensembl Rapid Release at the EBI.
Wellcome Sanger Institute – Legal and Governance
The materials that have contributed to this genome note have been supplied by a Darwin Tree of Life Partner. The submission of materials by a Darwin Tree of Life Partner is subject to the ‘Darwin Tree of Life Project Sampling Code of Practice’, which can be found in full on the Darwin Tree of Life website here. By agreeing with and signing up to the Sampling Code of Practice, the Darwin Tree of Life Partner agrees they will meet the legal and ethical requirements and standards set out within this document in respect of all samples acquired for, and supplied to, the Darwin Tree of Life Project.
Further, the Wellcome Sanger Institute employs a process whereby due diligence is carried out proportionate to the nature of the materials themselves, and the circumstances under which they have been/are to be collected and provided for use. The purpose of this is to address and mitigate any potential legal and/or ethical implications of receipt and use of the materials as part of the research project, and to ensure that in doing so we align with best practice wherever possible. The overarching areas of consideration are:
• Ethical review of provenance and sourcing of the material
• Legality of collection, transfer and use (national and international)
Each transfer of samples is further undertaken according to a Research Collaboration Agreement or Material Transfer Agreement entered into by the Darwin Tree of Life Partner, Genome Research Limited (operating as the Wellcome Sanger Institute), and in some circumstances other Darwin Tree of Life collaborators.
Acknowledgements
We would like to thank to Gabriela Bittencourt-Silva and Mark Wilkinson for allowing us to collect beetles for “Darwin Tree of Life” in their allotment in 2021.
Funding Statement
This work was supported by Wellcome through core funding to the Wellcome Sanger Institute [206194, <a href=https://doi.org/10.35802/206194>https://doi.org/10.35802/206194</a>] and the Darwin Tree of Life Discretionary Award [218328, <a href=https://doi.org/10.35802/218328>https://doi.org/10.35802/218328 </a>].
The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
[version 1; peer review: 3 approved]
Data availability
European Nucleotide Archive: Crioceris asparagi (common asparagus beetle). Accession number PRJEB59077; https://identifiers.org/ena.embl/PRJEB59077. The genome sequence is released openly for reuse. The Crioceris asparagi genome sequencing initiative is part of the Darwin Tree of Life (DToL) project. All raw sequence data and the assembly have been deposited in INSDC databases. Raw data and assembly accession identifiers are reported in Table 1 and Table 2.
Metadata for specimens, BOLD barcode results, spectra estimates, sequencing runs, contaminants and pre-curation assembly statistics are given at https://links.tol.sanger.ac.uk/species/131627.
Author information
Members of the Natural History Museum Genome Acquisition Lab are listed here: https://doi.org/10.5281/zenodo.12159242.
Members of the Darwin Tree of Life Barcoding collective are listed here: https://doi.org/10.5281/zenodo.12158331.
Members of the Wellcome Sanger Institute Tree of Life Management, Samples and Laboratory team are listed here: https://doi.org/10.5281/zenodo.12162482.
Members of Wellcome Sanger Institute Scientific Operations: Sequencing Operations are listed here: https://doi.org/10.5281/zenodo.12165051.
Members of the Wellcome Sanger Institute Tree of Life Core Informatics team are listed here: https://doi.org/10.5281/zenodo.12160324.
Members of the Tree of Life Core Informatics collective are listed here: https://doi.org/10.5281/zenodo.12205391.
Members of the Darwin Tree of Life Consortium are listed here: https://doi.org/10.5281/zenodo.4783558.
References
- Abdennur N, Mirny LA: Cooler: scalable storage for Hi-C data and other genomically labeled arrays. Bioinformatics. 2020;36(1):311–316. 10.1093/bioinformatics/btz540 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Allio R, Schomaker-Bastos A, Romiguier J, et al. : MitoFinder: efficient automated large-scale extraction of mitogenomic data in target enrichment phylogenomics. Mol Ecol Resour. 2020;20(4):892–905. 10.1111/1755-0998.13160 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bates A, Clayton-Lucey I, Howard C: Sanger Tree of Life HMW DNA fragmentation: diagenode Megaruptor ®3 for LI PacBio. protocols.io. 2023. 10.17504/protocols.io.81wgbxzq3lpk/v1 [DOI] [Google Scholar]
- Beasley J, Uhl R, Forrest LL, et al. : DNA barcoding SOPs for the Darwin Tree of Life project. protocols.io. 2023; [Accessed 25 June 2024]. 10.17504/protocols.io.261ged91jv47/v1 [DOI] [Google Scholar]
- Bezděk J, Sekerka L: Catalogue of palaearctic coleoptera. Volume 6/2/1. Updated and revised second edition. Chrysomeloidea II (Orsodacnidae, Megalopodidae, Chrysomelidae).Leiden: Brill,2024. 10.1163/9789004443303 [DOI] [Google Scholar]
- Brůna T, Hoff KJ, Lomsadze A, et al. : BRAKER2: Automatic eukaryotic genome annotation with GeneMark-EP+ and AUGUSTUS supported by a protein database. NAR Genom Bioinform. 2021;3(1): lqaa108. 10.1093/nargab/lqaa108 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Challis R, Richards E, Rajan J, et al. : BlobToolKit – interactive quality assessment of genome assemblies. G3 (Bethesda). 2020;10(4):1361–1374. 10.1534/g3.119.400908 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng H, Concepcion GT, Feng X, et al. : Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat Methods. 2021;18(2):170–175. 10.1038/s41592-020-01056-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chittenden FH: The asparagus beetles and their control.Washington, DC: United States Department of Agriculture,1917. Reference Source [Google Scholar]
- Cox ML: Atlas of the Seed and Leaf Beetles of Britain.Newbury: Pisces Publications,2007. Reference Source [Google Scholar]
- Crowley L, Allen H, Barnes I, et al. : A sampling strategy for genome sequencing the British terrestrial arthropod fauna [version 1; peer review: 2 approved]. Wellcome Open Res. 2023;8:123. 10.12688/wellcomeopenres.18925.1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- da Veiga Leprevost F, Grüning BA, Alves Aflitos S, et al. : BioContainers: an open-source and community-driven framework for software standardization. Bioinformatics. 2017;33(16):2580–2582. 10.1093/bioinformatics/btx192 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Denton A, Oatley G, Cornwell C, et al. : Sanger Tree of Life sample homogenisation: PowerMash. protocols.io. 2023a. 10.17504/protocols.io.5qpvo3r19v4o/v1 [DOI] [Google Scholar]
- Denton A, Yatsenko H, Jay J, et al. : Sanger Tree of Life wet laboratory protocol collection V.1. protocols.io. 2023b. 10.17504/protocols.io.8epv5xxy6g1b/v1 [DOI] [Google Scholar]
- Di Tommaso P, Chatzou M, Floden EW, et al. : Nextflow enables reproducible computational workflows. Nat Biotechnol. 2017;35(4):316–319. 10.1038/nbt.3820 [DOI] [PubMed] [Google Scholar]
- Diesh C, Stevens GJ, Xie P, et al. : JBrowse 2: a modular genome browser with views of synteny and structural variation. Genome Biol. 2023;24(1): 74. 10.1186/s13059-023-02914-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duff AG: Beetles of Britain and Ireland. Vol. 4: Cerambycidae to Curculionidae.West Runton: A.G. Duff publishing,2016. Reference Source [Google Scholar]
- Duff AG: Checklist of Beetles of the British Isles, 3rd Edition.Iver: Pemberley Books,2018. Reference Source [Google Scholar]
- Ewels P, Magnusson M, Lundin S, et al. : MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics. 2016;32(19):3047–3048. 10.1093/bioinformatics/btw354 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ewels PA, Peltzer A, Fillinger S, et al. : The nf-core framework for community-curated bioinformatics pipelines. Nat Biotechnol. 2020;38(3):276–278. 10.1038/s41587-020-0439-x [DOI] [PubMed] [Google Scholar]
- Formenti G, Abueg L, Brajuka A, et al. : Gfastats: conversion, evaluation and manipulation of genome sequences using assembly graphs. Bioinformatics. 2022;38(17):4214–4216. 10.1093/bioinformatics/btac460 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fowler WW: The coleoptera of the British Islands, Lamellicornia - Serricornia - Longicornia - Phytophaga.London: L. Reeve & Co,1890. [Google Scholar]
- Grüning B, Dale R, Sjödin A, et al. : Bioconda: sustainable and comprehensive software distribution for the life sciences. Nat Methods. 2018;15(7):475–476. 10.1038/s41592-018-0046-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guan D, McCarthy SA, Wood J, et al. : Identifying and removing haplotypic duplication in primary genome assemblies. Bioinformatics. 2020;36(9):2896–2898. 10.1093/bioinformatics/btaa025 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harry E: PretextView (Paired REad TEXTure Viewer): a desktop application for viewing pretext contact maps. 2022. Reference Source
- Heinze E: Die criocerinenAfrikas (Col. Chrysomelidae). EntomologischeArbeiten Aus Dem Museum G. Frey. 1962;13:156–270. [Google Scholar]
- Howe K, Chow W, Collins J, et al. : Significantly improving the quality of genome assemblies through curation. GigaScience. 2021;10(1): giaa153. 10.1093/gigascience/giaa153 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jay J, Yatsenko H, Narváez-Gómez JP, et al. : Sanger Tree of Life sample preparation: triage and dissection. protocols.io. 2023. 10.17504/protocols.io.x54v9prmqg3e/v1 [DOI] [Google Scholar]
- Kerpedjiev P, Abdennur N, Lekschas F, et al. : HiGlass: web-based visual exploration and analysis of genome interaction maps. Genome Biol. 2018;19(1): 125. 10.1186/s13059-018-1486-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kimoto S, Gressitt JL: Chrysomelidae (Coleoptera) of Thailand, Cambodia, Laos and Vietnam. I. Sagrinae, Donaciinae, Zeugophorinae, Megalopodinae and Criocerinae. Pacific Insects. 1979;20(2–3):191–256. Reference Source [Google Scholar]
- Kurtzer GM, Sochat V, Bauer MW: Singularity: scientific containers for mobility of compute. PLoS One. 2017;12(5): e0177459. 10.1371/journal.pone.0177459 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manni M, Berkeley MR, Seppey M, et al. : BUSCO update: novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Mol Biol Evol. 2021;38(10):4647–4654. 10.1093/molbev/msab199 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merkel D: Docker: lightweight Linux containers for consistent development and deployment. Linux J. 2014;2014(239): 2, [Accessed 2 April 2024]. Reference Source [Google Scholar]
- Monrós F: Los géneros de chrysomelidae (Coleoptera). Opera Lilloana. 3[1959],1960;5–337, pls. 1–3. Reference Source [Google Scholar]
- NBN Atlas Partnership: Crioceris asparagi (Linnaeus, 1758).Asparagus Beetle map on the NBN Atlas,2024. Reference Source
- Oatley G, Denton A, Howard C: Sanger Tree of Life HMW DNA extraction: automated MagAttract v.2. protocols.io. 2023. 10.17504/protocols.io.kxygx3y4dg8j/v1 [DOI] [Google Scholar]
- Pointon DL, Eagles W, Sims Y, et al. : sanger-tol/treeval v1.0.0 – Ancient Atlantis. 2023. 10.5281/zenodo.10047653 [DOI] [Google Scholar]
- Rao SSP, Huntley MH, Durand NC, et al. : A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell. 2014;159(7):1665–1680. 10.1016/j.cell.2014.11.021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rheinheimer J, Hassler M: Die blattkäfer baden-Württembergs.Karlsruhe: Kleinsteuber Books,2018. Reference Source [Google Scholar]
- Rhie A, McCarthy SA, Fedrigo O, et al. : Towards complete and error-free genome assemblies of all vertebrate species. Nature. 2021;592(7856):737–746. 10.1038/s41586-021-03451-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rhie A, Walenz BP, Koren S, et al. : Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 2020;21(1): 245. 10.1186/s13059-020-02134-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steinhausen W: 116. Familie: Chrysomelidae.In: Klausnitzer, B. (ed.) Die KäferMitteleuropas. Larven 2: Myxophaga, Polyphaga Teil 1.Krefeld: Goecke & Evers,1994;231–314. [Google Scholar]
- Steinhausen W: Die puppenmitteleuropäischerBlattkäfer - Eine vorläufigeBestimmungstabelle. 1. Teil (Coleoptera: Chrysomelidae). Mitt Munch Entomol Ges. 2001;91:35–63. Reference Source [Google Scholar]
- Strickland M, Cornwell C, Howard C: Sanger Tree of Life fragmented DNA clean up: manual SPRI. protocols.io. 2023. 10.17504/protocols.io.kxygx3y1dg8j/v1 [DOI] [Google Scholar]
- Surana P, Muffato M, Qi G: Sanger-tol/readmapping: sanger-tol/readmapping v1.1.0 - Hebridean Black (1.1.0). Zenodo. 2023a. 10.5281/zenodo.7755669 [DOI] [Google Scholar]
- Surana P, Muffato M, Sadasivan Baby C: sanger-tol/genomenote (v1.0.dev). Zenodo. 2023b. 10.5281/zenodo.6785935 [DOI] [Google Scholar]
- Twyford AD, Beasley J, Barnes I, et al. : A DNA barcoding framework for taxonomic verification in the darwin Tree of Life project [version 1; peer review: 2 approved]. Wellcome Open Res. 2024;9:339. 10.12688/wellcomeopenres.21143.1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uliano-Silva M, Ferreira JGRN, Krasheninnikova K, et al. : MitoHiFi: a python pipeline for mitochondrial genome assembly from PacBio high fidelity reads. BMC Bioinformatics. 2023;24(1): 288. 10.1186/s12859-023-05385-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vasimuddin M, Misra S, Li H, et al. : Efficient architecture-aware acceleration of BWA-MEM for multicore systems.In: 2019 IEEE International Parallel and Distributed Processing Symposium (IPDPS).IEEE,2019;314–324. 10.1109/IPDPS.2019.00041 [DOI] [Google Scholar]
- Vencl FV, Leschen RAB: 2.7.6. Criocerinae Latreille, 1807,In: Leschen, R. A. B. and Beutel, R. G. (eds.) Handbook of Zoology Arthropoda: Insecta: Coleoptera: Volume 3: Morphology and Systematics (Phytophaga).Berlin: De Gruyter,2014;237–242. Reference Source [Google Scholar]
- Warchałowski A: The palaearctic chrysomelidae. Identification keys. Vols. 1 & 2.Warszawa: Natura Optima Dux Foundation,2010.
- Zhou C, McCarthy SA, Durbin R: YaHS: yet another Hi-C scaffolding tool. Bioinformatics. 2023;39(1): btac808. 10.1093/bioinformatics/btac808 [DOI] [PMC free article] [PubMed] [Google Scholar]