Abstract
Epstein-Barr virus Nuclear Antigen Leader Protein (EBNA-LP) plays a pivotal role in the transformation of B cells by Epstein-Barr virus (EBV), functioning independently of EBNA2 to regulate chromatin architecture and gene expression. Our study reveals that EBNA-LP binds to chromatin regions distinct from EBNA2 and facilitates the formation of long-distance chromatin loops by interacting with the cellular factor YY1. This interaction reconfigures the three-dimensional structure of the host genome, enhancing the integrity of topologically associating domains (TADs) and promoting the interaction between enhancers and promoters within these domains. In EBV-infected B cells, EBNA-LP strengthens YY1-mediated chromatin loops within TADs, which helps maintain stable regulatory programs essential for B cell transformation. Notably, EBNA-LP is crucial for establishing EBV-induced enhancers, yet it is not required for their maintenance once formed. Additionally, our data suggest a compensatory increase in CTCF binding in the absence of EBNA-LP, leading to more promiscuous chromatin interactions between TADs and a reduced TAD insulation at their boundaries. These findings provide new insights into the molecular mechanisms by which EBV reshapes the host genome chromatin architecture to support B cell transformation and highlight potential therapeutic targets for disrupting EBV-driven oncogenesis.
Graphical Abstract
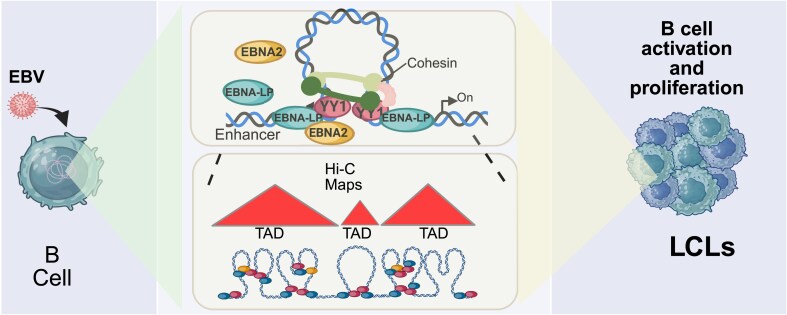
Graphical Abstract.
Introduction
Epstein-Barr virus (EBV) is the first known human DNA tumor virus which is responsible for around 200 000 cancer cases annually [1]. EBV establishes a lifelong latent infection in human B-cells by maintaining its genomes as circular chromatinized episomes, which replicate in sync with the host genome through the action of both viral and cellular factors [2]. Because episomes do not integrate into the host genome, it is essential for the virus to acquire a chromatin conformation that allows expression of genes required for transcription and replication while maintaining genomic stability. During latency, EBV expresses only a limited number of viral genes in at least four different transcriptional patterns that are referred to as latency types. The switch between these is dynamic and the different latency types reflect different stages of B cell development and are reliant on the maintenance of a stable 3D viral chromatin structure [3–5]. The most transcriptionally active state (Latency III) is characterized by the expression of viral noncoding RNAs, such as BART and EBER small non-coding RNAs and the viral protein called EBV nuclear antigen 1 (EBNA1), which functions as a transcription factor (TF) and facilitates episome replication and segregation during host cell division [2] as well as the other viral nuclear antigens EBNA2, EBNA-3A, -3B, -3C, and EBNA-leader protein (EBNA-LP) [2]. After EBV infection, the first two viral proteins expressed are EBNA2 and EBNA-LP. Genome-wide studies using techniques like chromatin immunoprecipitation followed by sequencing (ChIP-seq) have shown that both host and EBV TFs colocalize on both the viral and host genomes [6–12]: EBNA2 is essential for driving B-cell proliferation as it activates the expression of both host genes required for activation of the infected B cell, and the LMP1 viral gene. Moreover it modulates the binding of host TFs – such as RBPJ and EBF1– to the viral genome. EBNA-LP has been proposed to act mostly as a coactivator of EBNA2, by removing transcription repressors and activating EP300 [13–16]. However during infection, EBNA2 is entirely capable of activating cell genes in the absence of EBNA-LP, suggesting that the role of EBNA-LP in gene expression during virus infection is more nuanced or complex than simply boosting EBNA2 activity [17].
Type II latency [2], which expresses EBNA1, noncoding RNAs, and latency membrane proteins (LMP1, LMP2A and LMP2B), has been observed in germinal center B cells in healthy individuals, and most resembles the transcriptional state found in cancers such as nasopharyngeal carcinoma, Hodgkin disease, T/natural killer-cell lymphomas, and EBV-associated Gastric Carcinoma (EBVaGC) [2, 18, 19].
Latency I is the least transcriptionally active form and is observed in healthy adults’ resting memory B-cells [2]. It is characterized by expression of EBNA1 and the non-cording RNAs. Burkitt lymphoma, is associated with type I EBV latency [18].
The full repertoire of latency proteins is essential for the virus-driven transformation of human B-cells into continuously proliferating lymphoblastoid cell lines (LCLs) in vitro which serve as a valuable model to study some of the EBV-associated malignances [20].
The host genome is intricately packaged within the small confines of the nucleus in highly complex, yet organized patterns [21]. This packaging allows remote enhancers and their target genes to communicate efficiently by looping out large sections of DNA [22]. 3D genome interactions can be measured using chromatin conformation capture followed by deep sequencing (Hi-C), which reveals interaction frequencies between different genomic loci [23, 24]. This technique has rendered possible the identification of large, megabase-sized local chromatin interaction domains, termed as topologically associating domains (TADs). These domains correlate with regions of the genome that constrain the spread of heterochromatin and are stable across different cell types [25]. Most of the DNA looping is mediated by architectural proteins such as CCCTC-binding factor (CTCF) [26] and Yin-Yang 1 (YY1) [27, 28] TF together with the Cohesin complex [29]. HiChIP combines chromatin immunoprecipitation with Hi-C allowing to further elucidate the mechanisms by which these cellular factors orchestrate chromatin looping [30].
Infection of B-cells with EBV determines a global rearrangement of host chromatin mostly through the ability of some of the viral nuclear factors to interact and alter host TFs occupancy across cellular genome [6, 14–16, 31, 32].
Here we report the role of the viral nuclear factor EBNA-LP in reshaping host chromatin architecture through its colocalization with the host TF YY1 at both cellular and viral gene promoters essential for transformation. By using ChIP-seq, HiChIP, and HiC and employing EBNA-LP KO LCLs, we found that EBNA-LP colocalizes with YY1 at YY1-mediated loops and that these interactions are stabilized in latently infected cells. EBNA-LP KO cells exhibited not only altered cellular chromatin architecture at YY1 binding sites but an overall loss in TAD identity, changes in chromatin accessibility and gene expression. These data highlight key role for EBNA-LP in how EBV reconfigures the cell and partly the viral genomes to promote B-cell transformation.
Material and methods
Cell culture and treatment
Cell lines were maintained in a humidified incubator containing 5% CO2 at 37°C. LCL cells, EBNA-LP WT and KO cell lines were cultured in RPMI 1640 supplemented with Fetal Bovine Serum at a concentration of 15%, supplemented with 1% penicillin-streptomycin and plasmocin at a concentration 1:500 (Invivogen ant-mpp). Mutu1 cell lines were cultured under the same conditions except for the concentration of FBS supplemented to the media (10%).
B cells were negatively selected from PBMCs using EasySepTM Human B Cell Isolation Kit (StemCell Technologies 100–0971) as per manufacturer’s protocol. Cells were then counted and used for experiments carried out before infection. The remaining cells were spun down 600 g 3 min and resuspended in RPMI 1640 supplemented with HyCloneTM Fetal Bovine Serum at a concentration of 15% (Cytiva SH30071.03) and infected with EBV (B95.8 strain).
Nuclear extract and western blot
EBNA-LP WT/KO cells were cultured and prepared for nuclear extract as previously described [33]. Briefly, roughly 100 × 106 cells were washed twice with ice cold PBS before resuspension in buffer A (10 mM Tris pH 7.9, 1.5 mM MgCl2, 10 mM KCl, 0.5 mM DTT, 1 mg/ml each of protease inhibitors), and incubated at 4°C for 10 min. After spinning down at 2000 rpm for 10 min, the pellet was resuspended in buffer A and subject to dounce homogenization (with B pestle) and spun down again. The supernatant was kept as cytoplasmic extract. The pellet was resuspended in buffer C (20 mM Tris pH 8.0, 1.5 mM MgCl2, 0.42M NaCl, 25% glycerol, 0.2 mM EDTA, 0.5 mM DTT, protease inhibitors) and dounce homogenized (with B pestle), followed by incubation at 4°C for 30 min. The resulting lysate was spun down at 12 000 rpm for 30 min. The supernatant was kept as nuclear extract. Both extracts were dialyzed overnight in BC80 (20 mM Tris pH 8.0, 80 mM KCl, 0.2 mM EDTA, 10% glycerol, 1 mM β-mercaptoethanol, 0.2 mM phenylmethylsulfonyl fluoride (PMSF)), cleared at 20 000 g for 10 min, and stored at −80°C.
Protein concentration measurement and western blotting were carried out as previously described [4]. Membranes were incubated overnight at 4°C with primary antibodies against SMC3 (Bethyl Laboratories A300-060A), CTCF (Active Motif 61 311), EBNA-LP (Cosmo Bio #CAC-EBV-01), YY1 (Active Motif 61 779) and β-Actin (Cell Signaling Technology 12262S) as per manufacturer recommendation.
Quantification was performed using iBright Analysis Software version 5.3.0 (ThermoFisher).
RNA extraction and RNA-seq
Total RNA from EBNA-LP WT and KO cell lines was isolated from 2 × 106 cells using a Direct-zol RNA Kit (Zymo Research, cat. No. R2050) according to the manufacturer's protocol. RNA samples were either used for downstream RT-qPCR or submitted to the Wistar Institute genomics core facility for RNA quality control and sequencing library preparation using the SENSE mRNA-Seq Library Prep Kit V2 (Lexogen) to generate Illumina-compatible sequencing libraries according to the manufacturer's instructions. Paired-end reads of 150 bp were obtained using an Illumina HiSeq 2500 sequencer.
RNAseq analysis
RNA-seq data was aligned using STAR [34] against hg19 version of the Human genome and all unaligned reads were then aligned against NC_007605.1 version of EBV genome and featureCounts software [35] was used to estimate raw read counts for Human and EBV genes. DESeq2 [36] was used to perform differential analysis.
deepTools [37] was used for data visualization. To compare different conditions, alignment files were normalized using the reciprocal of the size factors calculated in DESeq2 [36] and inputted as the –scaleFactor argument of the bamCoverage function of deepTools [37]. For visualization purposes, all replicates for a same condition were merged.
Ingenuity Pathway Analysis (IPA) [38] was used to perform gene ontology and to identify the upstream regulators of differentially expressed genes.
CIBERSORTx [39] was used with default options to identify cell identity in EBNA-LP WT/KO conditions and GraphPad was used to asses statistical significance.
Heatmaps for differentially expressed genes were generated using pheatmap R package [40].
Datasets are available in Gene Expression Omnibus (see Data Availability section for accession number).
Chromatin immunoprecipitation assays
Chromatin immunoprecipitation sequencing (ChIP-seq)
Chromatin immunoprecipitation with next-generation sequencing (ChIP-seq) was performed as previously described [3]. Briefly, 50 × 106 cells per immunoprecipitation were collected and fixed with 1% formaldehyde for 15 min and then quenched with 0.25 M glycine for 5 min on ice. After three washes with 1X PBS, pellets were resuspended in 10 mL each of a series of two lysis buffers and resuspended in 1 mL of the third lysis buffer before fragmentation in Covaris ME220 Ultrasonicator (peak power 75, duty factor 25, cycles/burst 1000, average power 18.8, time 720 s) to generate chromatin fragments roughly 200–500 bp in size. Chromatin was centrifuged to clear debris and a 1:20 of this cleared chromatin was kept as standard input for comparison against immunoprecipitations. Chromatin was incubated rotating at 4°C 1 h with 50 μg YY1 (Active Motif 61 779) and 25 μg H3K27ac (Active Motif 39 133), then chromatin–antibody complexes were precipitated using 250 μL of Dynabeads Protein A (ThermoFisher, product No. 10001D) incubated rotating at 4°C overnight. DNA was purified using Zymo ChIP DNA Clean & Concentrator Kit (Zymo Research, product No. D5205). Libraries for sequencing were made using NEBNext Ultra II DNA Library Prep Kit (New England Biolabs, product No. E7103) and sequenced on the Illumina HiSeq 2500.
RNA pol II ChIP-seq
Chromatins were prepared as described above. Immunoprecipitations were carried out using total Pol II antibody (a gift from Alessandro Gardini at The Wistar Institute). RNAP2 antibody was incubated with Dynabeads Protein A for 1 h at 4°C, then beads were washed twice using the IP Dilution Buffer and added to the chromatin and incubated o/n at 4°C with rotation. All the subsequent steps were performed as described above.
ChIP-seq analysis
Reads were mapped against the hg19 version of the human genome and the human gammaherpesvirus 4 (HHV4) NC_007605.1 genome assembly using Burrows-Wheeler Aligner (BWA) [41]. We used MACS2 [42, 43] software package to call peaks using input samples as control. deepTools [37] was used for data visualization.
For comparisons, alignment files were read-depth normalized and replicates were merged for visualization purposes.
For TF-binding motif analysis, HOMER [44] findMotifsGenome.pl function was used using default options. HOMER annotatePeaks.pl function was used to annotate peaks. DESeq2 [36] was used to do differential analysis of ChIP-seq peaks. Average plots were obtained using seqMINER [45] and plotted using ggplot2 R package [46]. IPA [38] was used to perform gene ontology on ChIP-seq peaks.
ChIP-seq data were deposited for public access at Gene Expression Omnibus (see Data Availability section for accession number).
CUT&RUN
CUT&RUN was performed as per EpiCypher protocol. Briefly, 500 000 cells per condition were crosslinked using 0.1% formaldehyde for 1 min, then crosslinking was quenched using 0.125M glycine. Cells were spun down 600 g 3 min, washed twice with XL Wash Buffer (20 mM HEPES-KOH pH 7.9, 150 mM NaCl, 1% Triton X-100, 0.05% SDS, 1X Protease Inhibitor without EDTA, 0.5 mM Spermidine) at RT. Activated ConA beads (Bead Activation Buffer: 20 mM HEPES pH 7.9, 10 mM KCl, 1 mM CaCl2, 1 mM MnCl2) were added to the cells and incubated on nutator for 10 min. Beads were then collected on a magnet and 50 uL/reaction of fresh ice-cold Antibody Buffer (20 mM HEPES-KOH pH 7.9, 150 mM NaCl, 1% Triton X-100, 0.05% SDS, 1X Protease Inhibitor without EDTA, 0.5 mM Spermidine, 0.01% Digitonin, 2 mM EDTA) was added to the beads. 1 ug of H3K27ac antibody (Active Motif 39 133), 1 ug of YY1 antibody (Active Motif 61 779), 1 ug of CTCF antibody (Active Motif 61 311) or 1 ug of IgG antibody (CST 66362S) was added to the reaction and incubated o/n on a nutator at 4°C with cap elevated.
Next day beads were collected on a magnet, washed twice with 200 uL of ice-cold Digitonin Buffer (20 mM HEPES-KOH pH 7.9, 150 mM NaCl, 1% Triton X-100, 0.05% SDS, 1X Protease Inhibitor without EDTA, 0.5 mM Spermidine, 0.01% Digitonin), resuspended in 50 uL of the same buffer and 2.5 uL of pAG-MNase was added to each reaction. Samples were incubated for 10 min at RT on nutator with caps elevated. Beads were washed again twice with ice-cold Digitonin Buffer and resuspended in 50 uL of the same buffer. Around 1 uL of 100 mM CaCl2 was added to each sample on ice, and samples were incubated for 2 h on nutator at 4°C with caps elevated. Chromatin digestion with MNase was stopped using Stop Buffer (340 mM NaCl, 20 mM EDTA, 4 mM EGTA, 50 μg/mL RNase A, 50 μg/mL Glycogen, 1:100 Yeast Spike-in DNA) at 37°C for 10 min in a thermocycler. Supernatant was incubated with 0.8 uL of SDS and 1 uL of Proteinase K o/n at 55°C in a thermocycler to revert crosslinking. DNA was then purified using 1.1X AMPure XP Beads (Beckman-Coulter A63881), quantified using Qubit 1X dsDNA HS quantification kit (ThermoFisher Q33230) and libraries were prepared using NEBNext Ultra II DNA Library Prep Kit (New England Biolabs, product No. E7103) and sequenced on the Illumina HiSeq 2500.
CUT&RUN analysis
Reads were mapped against the hg19 version of the human genome, the human gammaherpesvirus 4 (HHV4) NC_007605.1 genome assembly and the Saccharomyces cerevisiae R64 assembly using BWA [41]. We used SEACR [47] to call peaks with default options. HOMER [44] annotatePeaks.pl function was used to annotate peaks. deepTools [37] was used for data visualization.
For comparisons, alignment files for YY1 and H3K27ac experiments in B cells and LCLs were read-depth normalized and replicates were merged for visualization purposes.
For CTCF CUT&RUN on EBNA-LP WT and KO cells, alignment files were normalized on spike-in DNA read counts and replicates were merged for visualization purposes. MACS2 [42, 43] was used to call peaks.
For TF-binding motif analysis, HOMER [44] findMotifsGenome.pl function was used using default options.
CUT&RUN data were deposited for public access at Gene Expression Omnibus (see Data Availability section for accession number).
Assay for transposase-accessible chromatin using sequencing (ATACseq)
ATACseq was performed as described by Grandi et al. [48]. Briefly, 100 000 cells were spun down at 500 g for 5 min at 4°C. Cell pellet was resuspended in 50 μL of fresh ice-cold ATAC-seq Lysis Buffer by pipetting up and down three times and incubated on ice for 3 min. Then, 1 mL of fresh ice-cold ATAC-seq Wash Buffer was added to dilute the lysis reagents and tubes inverted five times to mix. Nuclei were spun down at 500 g for 10 min at 4°C. Nuclei pellets were then resuspended in 50 μL of freshly made Transposition Mix by pipetting up and down six times and reactions were incubated at 37°C for 30 min in a thermomixer with 1000 rpm mixing. The transposition reaction was terminated by adding 250 μL (five volumes) of DNA Binding Buffer from the DNA Clean and Concentrator-5 Kit and mixed well by inversion. Reactions were kept o/n at −20°C. The next day DNA was purified using the Zymo DNA Clean and Concentrator-5 Kit as per protocol.
Each sample in the study was assigned to a unique combination of the Adapter 1 (Ad1) and Adapter 2 (Ad2) sequences. Each cleaned-up transposed DNA sample was transferred to a 200 μL PCR tube and mixed with 25 μL of NEBNext Ultra II Q5 2X Master Mix, 2.5 μL of the corresponding Ad1 and 2.5 μL of the corresponding Ad2.
ATACseq analysis
Reads were mapped against the hg19 version of the human genome and the human gammaherpesvirus 4 (HHV4) NC_007605.1 genome assembly using BWA [41]. MACS2 [42] software package was used to call peaks. deepTools [37] was used for data visualization.
For comparisons, alignment files were read-depth normalized and replicates were merged for visualization purposes using samtools [49].
DESeq2 [36] was used for differential analysis of ATACseq peaks.
Chromatin conformation capture assays
HiC assay
Hi-C assay was performed as previously described [3]. Briefly, 5 × 106 cells per condition were collected for in-situ Hi-C. Libraries of total ligation products were produced using Ultralow Library Systems V2 (Tecan Genomics, part no. 0344NB-32) as per manufacturer’s protocol.
Libraries were sequenced using the Illumina HiSeq 2500 sequencing platform with paired-end 75bp read length. HiC data was preprocessed using HiC-Pro v2.10.0 pipeline [50] with default settings using hg19 human genome. HiC matrices for the same condition were combined to get 600 M reads per condition and HiCcompare [51] was used to estimate significance of differential contact based on raw count matrix files at 150 kb resolution.
GENOVA R package [52] was used to estimate the relative contact probability, insulation score at CTCF peaks, to generate aggregate peak analysis (APA), to perform Aggregate TAD Analysis (ATA), to perform TAD + N analysis and to generate saddle plots for A/B compartments.
plotgardener R package [53] was used to generate HiC matrix plots.
HOMER [44] was used to detect significant A/B compartment switching and ggplot2 R package [46] was used to generate a scatterplot for A/B compartments.
To detect stripes from HiC Kight-Ruitz normalized matrices at a resolution of 10 kb stripenn [54] software was used. stripenn compute with the option –maxpixel 0.93,0.94,0.95,0.96,0.97,0.98,0.99 was used to detect stripes, while stripenn score was used to compute stripiness for every detected stripe in both conditions. The function stripenn seeimage was used to plot images of the stripes.
The detailed protocol with all minor alterations will be happily supplied by corresponding author per request. Datasets are available in GEO (see Data Availability section for the accession number).
HiChIP assay
HiChIP assay was performed as per protocol [30] with minor changes as previously described [4].
Briefly, 25 × 106 cells per condition were fixed with 1% formaldehyde for 10 min and then quenched with 0.125 M glycine for 5 min at RT. Ligation products were immunoprecipitated with 50 μg YY1 antibody (Active Motif 61 779). Around 5 μL of Streptavidin C-1 beads were used for biotin pull-down. After the last wash with Tween Wash Buffer, bead-bound DNA was used to prepare libraries with Ultralow Library Systems V2 (Tecan Genomics, part no. 0344NB-32) as per manufacturer’s protocol.
Libraries were then deep sequenced using the Illumina HiSeq 2500 sequencing platform.
Data were preprocessed using HiC-Pro v2.10.0 pipeline [50] with default settings using a combined version of hg19 and EBV NC_007605.1 genomes at 5kb resolution and hichipper software [55] was used to perform restriction site bias modeling and interaction identification using YY1 ChIP-seq peaks.
Only statistically significant interactions (FDR < 5%) were kept for downstream analysis.
diffloop Bioconductor R package [56] was used to merge anchors between samples, interactions with less than 5 reads in each condition were discarded and log2(KO counts/WT counts) was calculated for comparison between HiChIP experiments performed in EBNA-LP WT/KO cells. Interactions were then plotted using plotgardener R package [53].
YY1 HiChIP anchors were annotated and based on the annotation interactions were divided in Enhancer-Enhancer, Enhancer-Promoter, or Promoter-Promoter.
The detailed protocol with all minor alterations will be happily supplied by corresponding author per request.
Statistical analysis
All experiments presented were conducted at least in duplicate to ensure reproducibility of results. The Prism statistical software package (GraphPad) was used to identify statistically significant differences between experimental conditions and control samples, using statistical tests as indicated in the figure legends.
Datasets
The following publicly available datasets have been used: EBNA2 ChIP-seq GSE148396 (GEO Series), source 10.1101/gr.264705.120; EBNA-LP ChIP-seq GSE49338 (GEO Series), source 10.1073/pnas.1317608110; for H3K4me1 ChIP-seq in GM12878, ENCLB695AMJ (library), source ENCODE; for H3K4me1 ChIP-seq B-cells, ENCLB970ANP, ENCLB269HCT, ENCLB107GYS (library), source ENCODE; Time course B-cell RNAseq, GSE125974, source doi.org/10.1128/JVI.00226–19. Technical information regarding datasets generated are provided in Supplementary Table S1.
Results
YY1 Occupancy changes between EBV latency programs
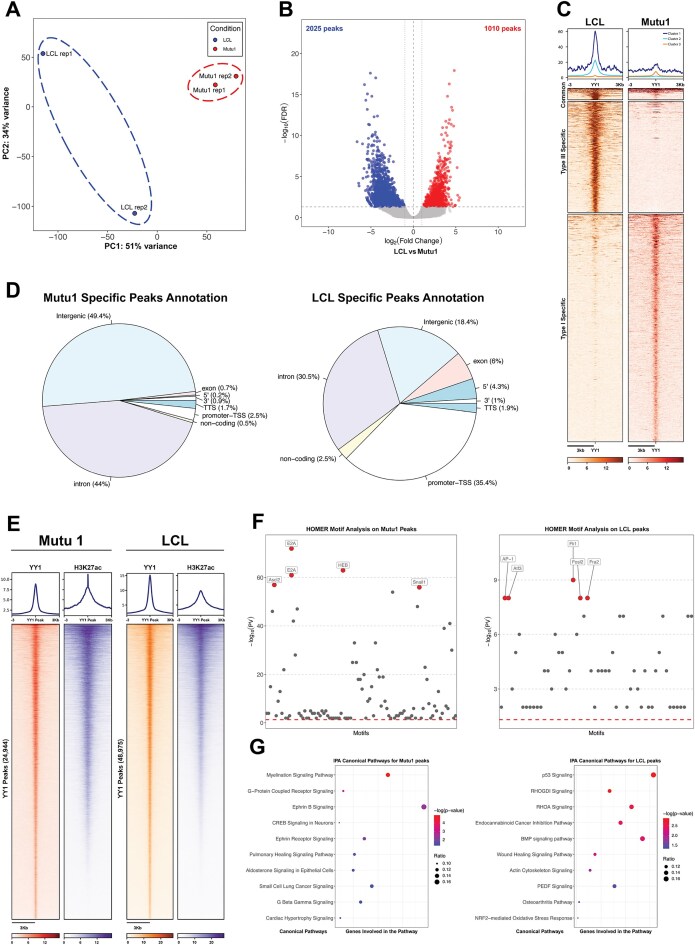
Our previous work suggested that viral chromatin remodeling is an essential feature of EBV latency programs. Recent evidence [57] unveiled the ability of EBV to reshape the host genome architecture to achieve cell transformation. To gain a better understanding of the mechanism regulating EBV-mediated host genome reorganization we decided to investigate the role of host factors involved in chromatin remodeling. In particular, we focused on YY1, a chromatin remodeling protein that is important for B cells [58] and that interacts with CTCF [27, 28]. We used ChIP-seq to assess YY1 occupancy in EBV + latency III LCL cells, and EBV + latency I Mutu1 cells. Our PCA analysis of ChIP-seq data revealed that YY1 binding distinctly separates EBV + B cells according to the viral latency program (Fig. 1A). When we compared YY1 occupancy in type III LCL cells to type I Mutu1 cells, we detected 3035 sites differentially occupied by YY1 between the two cell types (Fig. 1B). Of these differentially occupied YY1 loci, 2025 (∼67%) are regions where YY1 occupancy is lost in type III LCL cells compared to Mutu1 cells, and 1010 (∼33%) are regions where YY1 occupancy is gained in type III EBV cells compared to type I cells (Fig. 1B). Further analysis of YY1 ChIP-seq peaks in type I and type III EBV cells revealed three distinct clusters: sites commonly bound in both, sites specific to type III LCLs, and sites specific to type I Mutu cells (Fig. 1C). We observed that very few loci (<10%) were occupied by YY1 in both cell lines, suggesting that YY1 binding is influenced by both cell type and EBV latency.
Figure 1.
YY1 binding differs between latency types. (A) Principal Component Analysis (PCA) of YY1 ChIP-seq experiments in LCL (blue) and Mutu1 (red) cell lines. (B) Volcano plot of differentially bound regions on the human genome by YY1 between LCL and Mutu1 cell lines. (C) Heatmap showing the regions plotted as a volcano plot in B. (D) Annotation of the differentially bound regions of YY1. (E) Heatmaps showing the binding of YY1 and the active enhancer histone mark H3K27ac in Mutu1 (left) and LCL (right) cell lines. (F) HOMER TF-motif analysis of the differentially bound regions of YY1 in Mutu1 (left) and LCL (right) cell lines. (G) IPA of the differential YY1 peaks in Mutu1 (left) and LCL (right) cell lines.
To determine whether differentially occupied loci indicated genomic regions where YY1 potentially exerts differential regulatory effects between type I and type III EBV cells, we annotated cell specific YY1 peaks. We observed that in type I cells, YY1 occupancy predominantly occurred in intronic or intergenic regions, comprising 93% of the detected peaks. In contrast, in type III latency, the majority of the YY1 peaks (35%) were localized on promoters and transcription start sites (TSS) (Fig. 1D). These data revealed that YY1 binding patterns vary significantly between Mutu1 and LCLs. Next, we performed HOMER motif analysis to identify and characterize enriched TF-binding sites within the unique DNA sequences associated with YY1 occupancy in type I and III EBV + B cells. In type I cells, YY1 peaks were enriched for cognate motifs for ASCL2, E2A, HEB, and SNAIL1 TFs, which are involved in B cell development (Fig. 1F) [59–61]. In type III cells, YY1 peaks were variously enriched for motifs recognized by TFs like AP-1, ATF3, FL1, and Fosl2 that regulate gene expression programs associated with B cell transformation (Fig. 1F) [62–64]. This functional diversity of YY1 occupancy across the host genome was further confirmed by our gene ontology pathway analysis, which showed that YY1 is associated with distinct biological pathways in different EBV latency programs (Fig. 1G). In type I Mutu cells YY1-bound genes are involved in pathways associated with B cell development and activation including Ephrin B signaling and Ephrin Receptor signaling (Fig. 1G) [65]. In type III LCL cells, YY1 occupied regulatory regions of cellular genes that are associated with proliferation and activation of B cells, including p53 signaling, RHOGDI and RHOA signaling, and BMP signaling (Fig. 1G). Overall, our data show that YY1 functions differently and regulates distinct gene sets depending on both the type of EBV-infected B cell and the form of EBV latency present. This suggests that EBV may have evolved mechanisms to modulate YY1 activity in a context-specific manner.
YY1 and EBNA-LP share common binding sites on the host genome
Previous ChIP-seq analysis and recent work from Dr. Zhao’s group suggests that EBNA2 and EBNA-LP binding may be co-occurring with YY1 binding [15]. Subsequently, recent work from our collaborator Dr. Luftig showed that YY1 and EBNA-LP physically interact and form a complex in EBV + cells [66]. To determine whether EBV evolved mechanisms to control YY1 function, we assessed whether a relationship exists between YY1 and the two viral proteins, EBNA2 and EBNA-LP, that bind the host genome and trigger lineage-specific gene expression program that are essential for B cells proliferation. Given the differences we observed in YY1 binding between type I and type III EBV + cells, we assessed whether YY1 peaks identified in our ChIP-seq experiment overlapped with available EBNA2 and EBNA-LP ChIP-seq datasets on the host genome. Our findings indicate that in type III latency, there are no loci bound by YY1 and EBNA2 together without EBNA-LP (Fig. 2A, cluster 1). On the contrary, we observed several genomic regions where EBNA-LP binding overlapped with YY1 without the presence of EBNA2 (Fig. 2A, cluster 2) or where EBNA-LP exhibited a stronger binding intensity than EBNA2 (Fig. 2A, cluster 3). This suggests that by localizing with EBNA-LP and/ or EBNA2, YY1 potentially play distinct roles in regulating host chromatin. To further prove that differences in binding between type I latency and type III latency are associated to the expression and binding of these two viral proteins we assessed YY1 binding between Mutu1 and LCLs on the previously mentioned clusters (Fig. 2B). Interestingly, cluster 1 sites exhibited YY1 binding only in the type III LCL cell line, while cluster 2 showed a reduced level of YY1 binding in type I Mutu1 cell line. Contrary to cluster 2, cluster 3 shows YY1 binding intensity in Mutu1 cell line being opposite to the one of EBNA-LP and EBNA2 in LCLs. This data suggests that the majority of the YY1 peaks co-localize with EBNA-LP (alone or together with EBNA2), indicating a shared regulatory mechanism among these factors. Gene ontology analysis showed that genomic regions bound by YY1 and EBNA-LP were enriched for cellular genes involved in RNA biology, deubiquitination and control of cell cycle (Fig. 2C). These regions were also enriched for motifs recognized by TFs belonging to the Ets family, including Elk1 and Elk4 (Fig. 2D, bottom). These factors are involved in various cellular processes, including growth and differentiation, by regulating gene expression through the MAPK/ERK signaling pathway, which is activated by EBV [67–69]. Notably, in regions co-occupied by YY1 and EBNA-LP together with EBNA2 (Cluster 1 Fig. 2A), we observed an enrichment for motifs recognized by TFs involved in cell proliferation, including PU.1 and IRF8, that are also known EBNA2 partners (Fig. 2D, top). Recent work has demonstrated that IRF8 and PU.1 are essential for transcriptionally coordinating the germinal center response and the development of late-stage B cells [70]. Overall, these data indicate that in EBV-infected cells, YY1 and EBNA-LP binding co-occurred cross the host genome, potentially regulating EBV-driven B cell proliferation through YY1 binding.
Figure 2.
YY1 co-localizes with EBNA-LP and EBNA2 in LCLs. (A) Heatmaps for EBNA-LP (pink), EBNA2 (blue) and YY1 (orange) ChIP-seq experiments. Heatmaps clusters based on the ChIP-seq signal were obtained with deepTools. (B) YY1 signal in Mutu1 and LCL cell lines using the same clusters obtained in A. (C) IPA analysis of the EBNA-LP/YY1 co-bound regions. (D) HOMER motif analysis of the EBNA-LP/EBNA2/YY1 (top) and EBNA-LP/YY1 (bottom) co-bound regions.
EBNA-LP and EBNA2 are present at YY1-mediated loop anchors
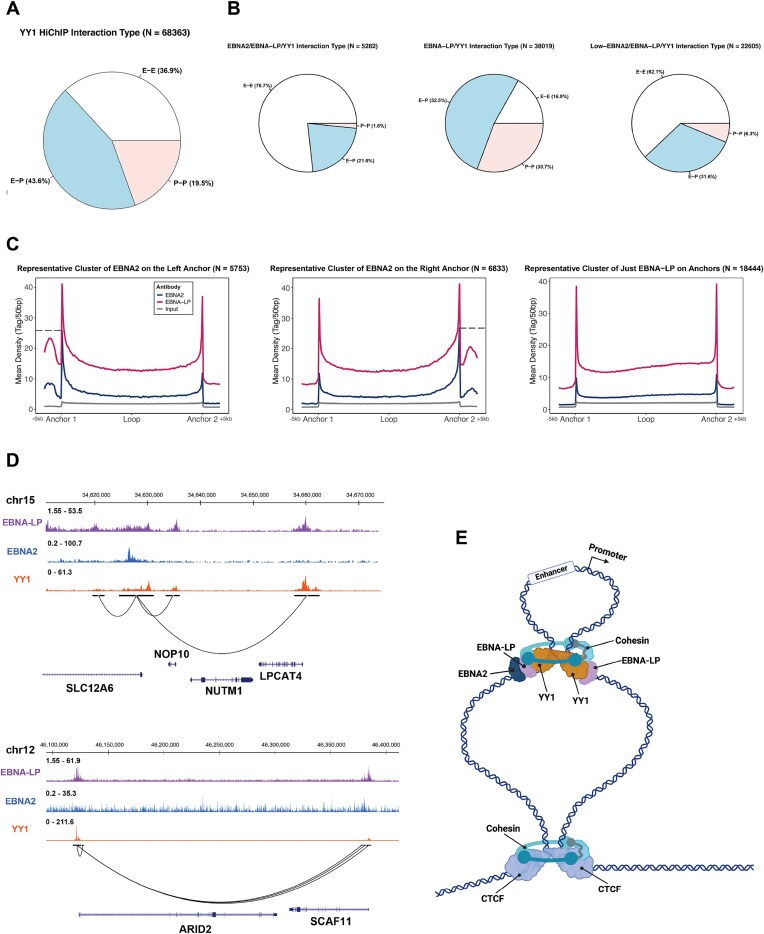
YY1 contributes to enhancer-promoter structural interactions through chromatin looping [28]. To determine whether YY1-mediated interactions are influenced by EBNA-LP and EBNA2, we used HiChIP for YY1 to identify YY1-associated chromatin loops and integrated these datasets with the EBNA2 and EBNA-LP publicly available occupancy maps generated in type III EBV + B cells. First, we confirmed that YY1-mediated interactions occurred between regulatory regions, i.e. enhancer or promoter regions, with the majority (∼44%) of the interactions mediated by YY1 connecting enhancers to promoters (Fig. 3A). Next, we integrated the structural studies with the occupancy data for EBNA2 and EBNA-LP to assess whether those viral proteins participate also in YY1-mediated chromatin loops. Interestingly, most of the YY1-mediated interactions occurred at regulatory regions occupied also by EBNA-LP (Fig. 3B, middle). Notably, only a small fraction (8%) of these structural interactions involved regions that were strongly occupied by EBNA2 (cluster 1, Fig. 2A) (Fig. 3B, left). These data indicated that most YY1-mediated interactions involved EBNA-LP associated regions, indicating a role for EBNA-LP in regulating chromatin looping. To gain a better understanding of how EBNA-LP, EBNA2, and YY1 interact to form physical connections between regulatory regions, we examined the genomic regulatory interactions mediated by YY1 (i.e. enhancer-enhancer, enhancer-promoter, and promoter-promoter) and intersected them with clusters from Fig. 2A (Figure 3B). Our analysis revealed that most EBNA-LP-YY1 interactions connect enhancer and promoter regions, whereas most EBNA2-YY1 interactions occurred between enhancers (Fig. 3B). Similarly, EBNA2-EBNA-LP-YY1 interactions primarily involved enhancers, either connecting enhancers to other enhancers or, less frequently, linking enhancers to promoters (Fig. 3B). Moreover, the distribution of EBNA-LP binding showed that EBNA-LP binds to both regions connected by YY1 (Fig. 3C and D). In contrast, EBNA2 binds to only one of the two regions physically connected by YY1 (Fig. 3C, left and middle, Fig. 3D, left and Fig. 3E), consistent with the ability of EBNA2 to interact with enhancers. These results demonstrated the ability of EBNA-LP to interact with YY1 in connecting regulatory elements, possibly ensuring the structural integrity of these interactions to facilitate B cell epigenetic reprogramming upon EBV infection.
Figure 3.
YY1 bridges enhancers and promoters together with EBNA-LP. (A) Pie charts representing the type of YY1-mediated interactions (E-E = enhancer-enhancer interactions, E-P = enhancer-promoter interactions, P-P = promoter-promoter interactions). (B) Pie charts showing the type of interactions listed in A per cluster type listed in Fig. 2A. (C) Average plots showing the ChIP-seq signal of EBNA-LP (purple) and EBNA2 (blue) on YY1 HiChIP interactions in LCLs. Left: representative cluster of EBNA-LP/EBNA2/YY1 interactions with EBNA2 on the left anchor. Middle: representative cluster of EBNA-LP/EBNA2/YY1 interactions with EBNA2 on the right anchor. Right: representative cluster of EBNA-LP/YY1 interactions. (D) UCSC Genome Browser views of EBNA-LP/EBNA2/YY1 interactions (top) and EBNA-LP/YY1 only interactions (bottom). (E) Schematic representation of YY1 HiChIP loops and YY1/EBNA-LP/EBNA2 interactions at loop anchors. Created in BioRender. Tempera, I. (2025) https://BioRender.com/vz4kxsr
EBNA-LP-YY1 loops exhibit an increased frequency upon B cell infection
To test whether YY1/EBNA-LP interactions play any role during EBV infection, we used CUT&RUN YY1 to compare binding to genetic loci in B cells before and after EBV infection. Since we observed that regions where YY1 and EBNA-LP binding co-occurred included enhancers, we also measured H3K27ac deposition in resting primary B cells and LCLs at 28 days post-infection. We integrated these datasets also with publicly available datasets for H3K4me1, a histone mark associated with primed or poised enhancers (Fig. 4). We observed that at those regions where in type III LCL cells EBNA-LP and YY1 co-bind, YY1 binding decreases in EBV infected cells compared to matching uninfected B cells (Fig. 4A). Moreover, at these same regions, the decrease in YY1 binding coincides with an increase in H3K27ac and a decrease of H3K4me1 deposition (Fig. 4A and B). The combined increase in H3K27ac and decrease in H3K4me1 typically indicates that these EBNA-LP-YY1 co-bound regions shifted, on average, from a poised or primed enhancer state in resting B cells to a fully active state in LCLs (Fig. 4B). Activated enhancers are associated with the formation of chromatin loops that bring them into close contact with their target promoters. Thus, to determine whether these EBNA-LP-YY1 regions establish structural connections after EBV infection, we performed in situ Hi-C assay in matching B cells before and after EBV infection (Supplementary Fig. S1). Consistent with previous reports, our PCA analysis revealed that the activation, differentiation, and proliferation of B cells driven by EBV significantly reshapes the global network of structural chromatin interactions within infected B cells (Supplementary Fig. S1A). Specifically, EBV infection leads to a notable reorganization of transcriptionally permissive (A compartment) and repressive (B compartment) chromatin compartments, with a significant number of these compartment switching from one type into the other after EBV infection (Supplementary Fig. S1B). However, EBNA-LP-YY1 co-bound regions in LCLs remain localized in A (active) compartments both before and after EBV infection (Fig. 4C). To determine whether in B cells EBNA-LP-YY1 forms chromatin loops after EBV infection, we used APA [23] to examine the composite signal of EBNA-LP-YY1 associated loops genome-wide. Detailed analysis of the APA plots focusing on YY1 and EBNA-LP binding sites highlighted an enrichment of loops involving these proteins after infection of B cells with EBV (Fig. 4D, Supplementary Fig. S1D, bottom). Similarly, for the regions occupied by all three factors (EBNA2-EBNA-LP-YY1), the APA plots showed an increased frequency of chromatin loops after EBV infection (Supplementary Fig. S1C and D, top). Overall, these results indicate that upon infection EBV promotes the formation of structural chromatin interactions mediated by YY1 through EBNA-LP binding. These data suggest that EBNA-LP could either be necessary for the formation of a subsets of YY1-associated chromatin loops or be recruited at these loops.
Figure 4.
During EBV-driven transformation of B cells YY1-mediated interactions are strengthened and YY1 peaks gain more H3K27ac histone mark. (A) Heatmap showing YY1, H3K27ac and H3K4me1 CUT&RUN and ChIP-seq signals between B cells before EBV infection and after 28 days post infection. Clusters are the same as in Fig. 2A. (B) Boxplots showing the difference in YY1 and H3K27ac CUT&RUN signals and the respective P-value (calculated using Wilcoxon t test). (C) Barplot representing the percentage of EBNA-LP/YY1 co-bound regions overlapping with HiC compartments. HiC compartments were divided in A to A, A to B, B to A, and B to B based on HiC eigenvectors calculated with HOMER. (D) APA plots measuring the contact frequency ± 50Kb from EBNA-LP/YY1 co-bound loops in B cells (top square) and LCLs (middle square).
EBNA-LP is essential for chromatin loop formation
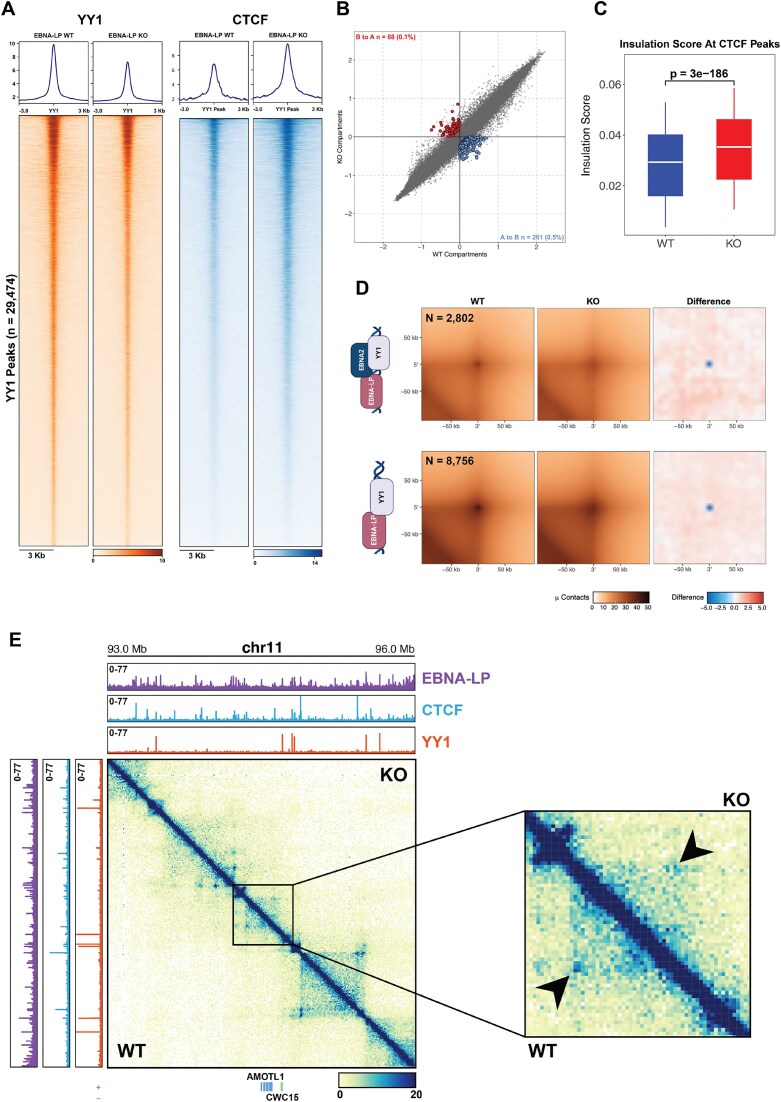
To establish a role for EBNA-LP in chromatin loop formation, we conducted structural chromatin studies in type III EBNA-LP KO LCLs, generated as described previously [17, 66] using an EBV virus with a disrupted EBNA-LP ORF [17]. The matched pairs of EBNA-LP KO and WT LCL cells generated from two different donors exhibited minimal or no effect on the expression of either YY1 or other proteins essential for chromatin looping, such as CTCF and the Cohesin subunit SMC3 (Supplementary Fig. S2A and B). First, we determined by ChIP-seq in these LCLs whether EBNA-LP caused any changes in YY1 binding across the viral genome. We observed in LP-KO cells a decrease in YY1 occupancy across the genome compared to WT cells, suggesting an effect of EBNA-LP in stabilizing YY1 binding on chromatin (Fig. 5A). YY1 cooperates with CTCF in establishing structural chromatin interactions, therefore we assessed whether EBNA-LP KO also affects CTCF binding at YY1 bound regions. Using CUT&RUN for CTCF, we observed a significant enrichment of CTCF binding at YY1 bound regions in LP-KO cells compared to WT (Fig. 5A). Notably, these results indicate that in EBNA-LP KO cells the reduction in YY1 occupancy correlates to an increase in CTCF binding at the same regions, suggesting an interplay between these two factors. Both CTCF and YY1 are integral in the organization of the three-dimensional structure of the genome. Thus, we used Hi-C to determine whether EBNA-LP KO changes the structural interactions of the host genome in EBV infected cells. We observed compartmentalization switches between LP-WT and LP-KO cells, with most of these involving an A-to-B (active-to-inactive chromatin) compartment switching (Fig. 5B). These data indicate that compartmentalization is altered in EBNA-LP KO, with most of these changes occurring in the B compartment. Next, we determined whether EBNA-LP KO alters interaction patterns between compartments by assessing compartment strength using saddle plot analysis (Supplementary Fig. S3A). The saddle plot analysis indicates a slight increase in A-A interactions and conversely a reduction in B-B interactions in LP-KO cells compared to WT controls (Supplementary Fig. S3A). LP-KO cells also had more interactions between A and B compartments than WT LCLs, indicating a significant reorganization of chromatin compartmentalization in EBNA-LP KO. Compartmentalization is achieved through the preserved integrity of topologically associated domains (TADs) within compartments. Thus, we assess the effect of EBNA-LP KO on TAD organization. ATA showed that intra-TADs interactions were reduced in LP-KO cells compared to WT (Supplementary Fig. S2B), indicating weakened TAD integrity and less distinct boundaries.
Figure 5.
EBNA-LP KO determines global changes in YY1 and CTCF binding and in chromatin architecture. (A) Heatmaps showing YY1 ChIP-seq signal and CTCF CUT&RUN signal on YY1 peaks identified in EBNA-LP WT and KO cell lines. (B) Scatterplot of A/B HiC compartments obtained using HOMER in EBNA-LP WT and KO LCLs. Compartment switching from B to A (red) and A to B (blue) that pass the p-value < 0.05 filter are colored. (C) Boxplot representing the insulation score calculated on CTCF peaks between EBNA-LP WT (blue) and KO (red) cell lines with respective p-value (calculated using Wilcoxon t test). (D) APA plots showing contact frequency of EBNA-LP/EBNA2/YY1 (top) and EBNA-LP/YY1 only (bottom) co-bound regions. (E) HiC Knight-Ruiz-balanced matrices showing differences in contact frequency on chromosome 11 between EBNA-LP WT (bottom half) and KO (top half) LCLs. EBNA-LP (purple), CTCF (light blue) and YY1 (orange) WT (left) and KO (top) signals are plotted.
Typically, the integrity of TADs is maintained by CTCF binding, which insulates adjacent TADs and preserves their boundaries. To determine whether EBNA-LP affects CTCF insulating ability, we calculated the insulation score for CTCF CUT&RUN binding sites in LP-KO and WT cells. Consistent with weak and less distinct TADs, LP-KO cells showed a higher insulation score for CTCF bound regions compared to WT (Fig. 5C), which suggest a reduction in insulation ability of CTCF. Loss of TAD boundaries may lead to the formation of promiscuous interactions between different TADs (inter-TADs) or compartments. To test this, we plotted the contact probability as a function of genomic distance for both LP-WT and LP-KO cells and compared the two curves. Consistent with loss of insulation between TADs, EBNA-LP KO cells showed higher contact probability for long-range interactions occurring at the levels of TADs and compartments than WT cells (Supplementary Fig. S3C). Indeed, the number of inter-TAD interactions was higher in EBNA-LP KO cells than WT ones (Supplementary Fig. S3D), consistent with weakening of boundaries in absence of EBNA-LP. Overall, these data indicated that EBNA-LP increases TAD boundary integrity and compartmentalization.
EBNA-LP regulates the formation of YY1-mediated chromatin loops
The results on inter-TADs indicate that EBNA-LP contributes to regulating long-rage chromatin interactions. Next, we explored whether EBNA-LP is also involved in short-range interactions mediated by chromatin loops (interactions between sites < 100 kbp apart). Differential analysis of loops showed examples of genomic regions with both increased (upregulated) and decreased (downregulated) interaction frequencies in EBNA-LP KO cells compared to WT cells (Supplementary Fig S4A and B). Based on our observations on the loss of TAD integrity and the increased formation of promiscuous interactions (Supplementary Fig. S3), we hypothesized that EBNA-LP KO would exhibit an increase in in chromatin loop length given the weakened robustness of TAD boundaries. Indeed, our data show that, compared to wild-type (WT) cells, the loops gained in EBNA-LP KO cells are on average longer than the loops that have been lost (Supplementary Fig. S4C). The binding profile of YY1 at the loop anchor points of both upregulated and downregulated loops showed a reduced binding of YY1 at both anchors in LP-KO cells compared to WT (Supplementary Fig. S4D). CTCF showed strictly differences in binding at loop anchors. In upregulated loops, CTCF binding increased at the start anchor point while decreased at the end anchors in LP-KO cells compared to LP-WT (Supplementary Fig. S4D). In downregulated loops, CTCF binding occurred predominantly at the start anchor point and its signal was reduced in LP-KO compared to LP-WT (Supplementary Fig. S4D). Analysis of chromatin accessibility using transposase-accessible chromatin sequencing (ATAC-seq) revealed that, compared to WT cells, EBNA-LP KO cells exhibited decreased chromatin accessibility at regions adjacent to both anchor points. However, there were no changes in chromatin accessibility within the intervening regions of the loops (Supplementary Fig. S4D). Consistent with a role for EBNA-LP and, sometimes alongside EBNA2, in the formation of these loops, on either up- or down-regulated loops both viral proteins are present at the loop anchors (Supplementary Fig. S4E). Given the interaction between EBNA-LP and YY1 at anchor points of YY1-mediated loops, we explored further YY1 occupancy at loop anchors in EBNA-LP KO cell lines. In KO cells, we observed the disruption of chromatin loops formed at EBNA-LP-YY1 occupied regions (Fig. 5D and E); notably we observed similar disruption of chromatin loops at those regions where also EBNA2 was part of the complex (Fig. 5D), indicating that EBNA-LP could influence CTCF and YY1 occupancy at YY1-bound chromatin loops. Overall, these results indicate that EBNA-LP in B cells helps solidify TADs and regulates the formation or disruption of chromatin loops.
In the absence of EBNA-LP LCLs exhibit disrupted stripes
HiC data revealed regions of high chromatin interaction which appears in contact matrices as stripes. Stripes consist of a super-enhancer (SE), generally present at the anchor of the stripe, which forms multiple interactions with genes present in a gene-dense locus. Stripes are frequently associated with active regulatory elements such as enhancers, promoters, and insulators. We observed that EBNA-LP deregulated 10% of all the stripes we were able to identify with our Hi-C assay resolution (Supplementary Fig. S5A). EBNA-LP KO induced the formation of novel stripes (n = 10) and the disruption of existing ones (n = 20) (Supplementary Fig. S5A). For example, on chromosome 4, we observed a stripe extending from a region enriched for YY1, EBNA-LP, and CTCF binding, that is visible in the Hi-C contact map of LP-WT cells but is disrupted in KO cells: the gap in the stripe coincides with a site of increased CTCF binding and loss of YY1 binding in KO cells (Supplementary Fig. S5B). Similarly, in KO cells, we observed a disruption of a stripe by gap formation in a region on chromosome 20 (Supplementary Fig. S5D), which also correlated with increased CTCF binding in KO cells. An example of a newly formed stripe is on chromosome 10 (Supplementary Fig. S5C) where the loops forming that stripe appear to be more frequent. The changes in stripe formation in LP-KO cells, coinciding with increased CTCF binding and loss of YY1 binding, suggests that EBNA-LP plays a crucial role in maintaining chromatin organization of these regulatory regions.
EBNA-LP loss determines changes in chromatin accessibility and gene expression
The previous results indicated that EBNA-LP normally participates in the reconfiguration of the host genome, and its absence leads to architectural chromatin changes. To investigate whether these altered chromatin structures also affect the chromatin state, we analyzed chromatin accessibility using transposase-accessible chromatin sequencing (ATAC-seq) in LP-WT and KO type III EBV + cells.
PCA analysis revealed that LP-WT and LP-KO cells can be separated based on their chromatin accessibility (Fig. 6A). However, the donor origin also has a noticeable influence on the chromatin state of the samples (Fig. 6A).
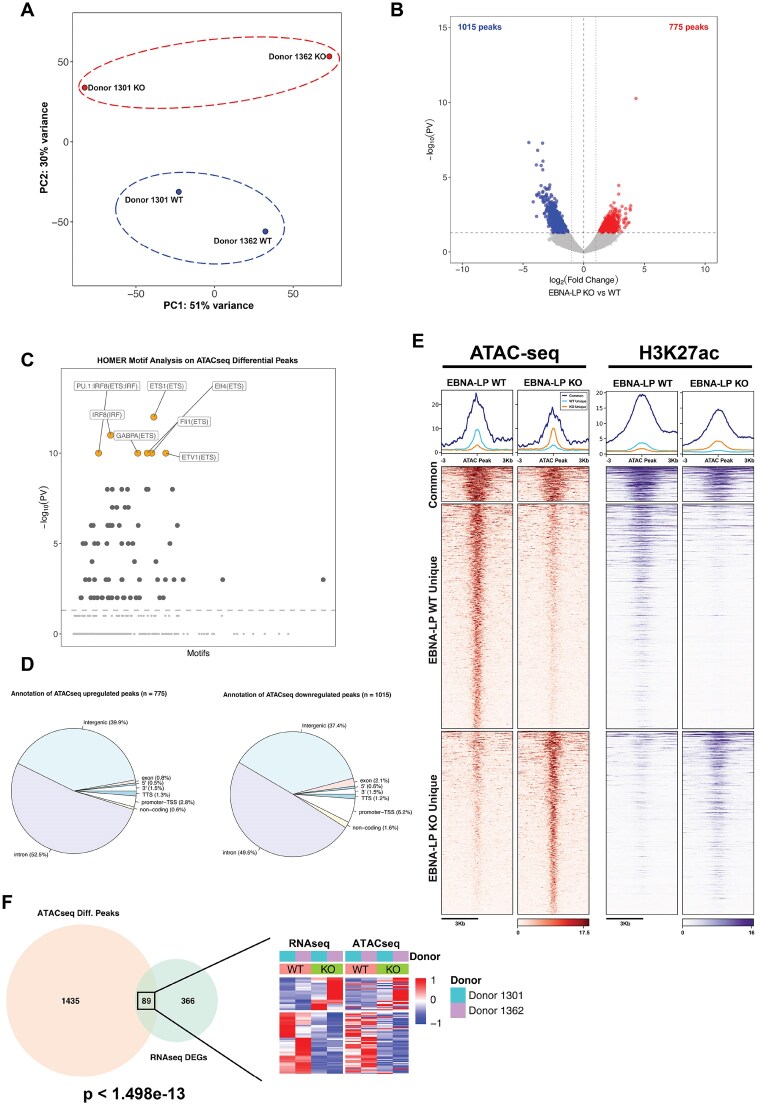
Figure 6.
Chromatin accessibility is altered by EBNA-LP KO. (A) PCA analysis of ATAC-seq peaks between EBNA-LP WT (blue) and KO (red) cells. (B) Volcano plot showing differentially accessible regions of the chromatin between EBNA-LP WT and KO cell lines. (C) HOMER motif analysis of the differentially accessible regions. (D) Annotation of the differential ATAC-seq peaks. (E) Heatmaps showing ATAC-seq and H3K27ac ChIP-seq signals on the differential ATAC-seq peaks. (F) Venn diagram showing the overlap between ATAC-seq differential peaks and differentially expressed genes between EBNA-LP WT and KO cell lines. Of the overlapping genes the respective heatmaps showing the normalized read counts of both experiments are shown.
EBNA-LP KO induces changes in chromatin accessibility of 1790 genomic regions, with a slight but noticeable tendency (56% of the differential peaks) towards a loss of chromatin accessibility (Fig. 6B). Changes in chromatin accessibility in EBNA-LP KO cell lines occurred at genomic regions containing motifs recognized by TFs critical for regulating lineage-specific transcriptional programs, including PU.1 and ETS-1 (Fig. 6C). Notably, PU.1 and ETS motifs were also enriched among the genomic regions co-occupied by EBNA-LP and YY1 in type III LCL cells (Fig. 2D), supporting a role for EBNA-LP in regulating chromatin accessibility for these TFs.
However, chromatin regions with both increased (upregulated) and decreased (downregulated) accessibility after EBNA-LP KO occurred mostly at intergenic and intronic regions (Fig. 6D). Consistent with changes in chromatin accessibility, we observed gain or loss of the active histone mark H3K27ac at upregulated (EBNA-LP KO Unique) and downregulated (EBNA-LP WT Unique and Common) ATAC-seq peaks in LP-KO cells compared to WT cells (Fig. 6E).
To determine whether changes in chromatin accessibility were directly linked to changes in gene transcription in LP-KO cells, we assessed global changes in gene expression in LP-KO cells compared to WT cells by RNA-seq (Fig. 7). By overlapping ATAC-seq changes with RNA-seq data, we found that 89 cellular genes (20% of all differentially expressed genes) showed changes in transcription that correlated with changes in chromatin accessibility (Fig. 6F). As expected, upregulation of gene expression coincided with an increased open chromatin state, while downregulation of gene expression corresponded with decreased chromatin accessibility (Fig. 6F). These results suggest that EBNA-LP is essential for sustaining the profile of chromatin structure and accessibility found in LP-WT LCLs.
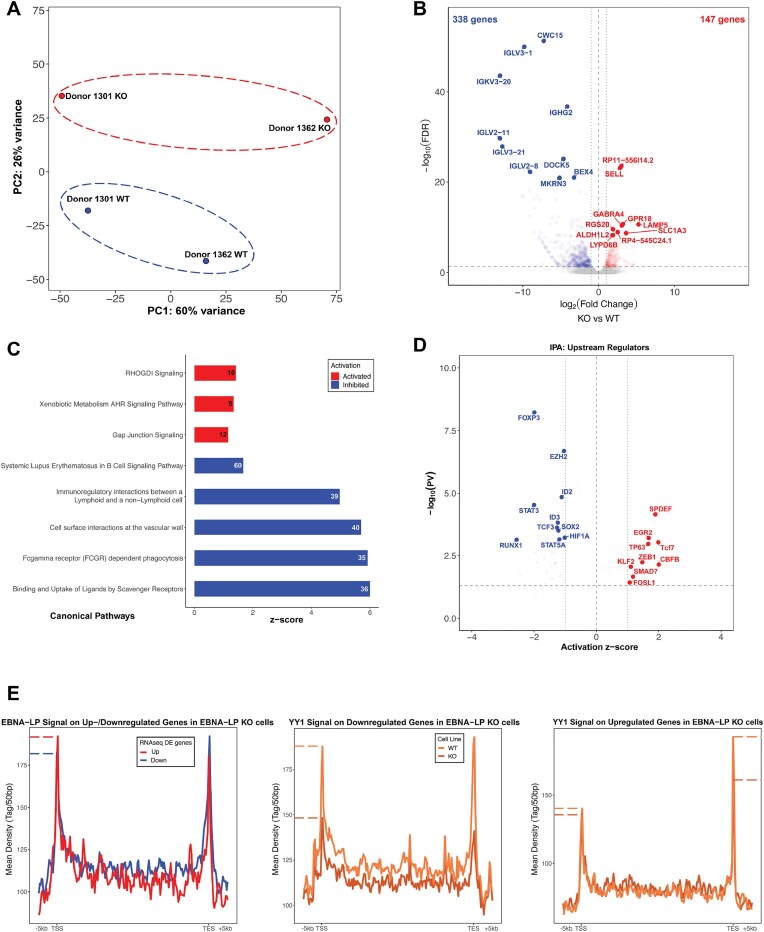
Figure 7.
A global downregulation of gene expression is determined by EBNA-LP KO. (A) PCA analysis of the RNA-seq experiment between EBNA-LP WT and KO cell lines. (B) Volcano plot showing the differentially expressed genes in the two conditions. The top ten up- and downregulated genes are reported on top of their respective dot. (C) IPA analysis of the differentially expressed genes showing the top five up- and downregulated canonical pathways. (D) Scatterplot showing the up- and downregulated upstream regulators. The top ten for each condition are reported. (E) Average plots showing EBNA-LP (left, red = upregulated, blue = downregulated), YY1 (middle and right, orange = WT, dark orange = KO) ChIP-seq signals on up- and downregulated genes.
We further examined which genes were deregulated upon depletion of EBNA-LP. PCA analysis of the RNA-seq data showed that 26% of the variance is attributable to the presence of EBNA-LP (Fig. 7A). We identified 485 differentially regulated (DE) genes in LP-KO cells compared to WT cells (Fig. 7B). We observed both upregulated and downregulated genes in absence of EBNA-LP, although DE genes were skewed toward downregulation (338 out of 485; 70% of DEG) (Fig. 7B), consistent with a role for EBNA-LP in reorganizing chromatin to alter host gene expression. To assess if changes in transcription correlate with epigenetic changes, we chose two of the top downregulated genes AMOTL1 and CWC15 (Supplementary Fig. S6A) that are located within regions exhibiting altered chromatin structures in LP-KO cells (Fig. 5E). We confirmed that changes in transcription correlate to changes in deposition of the active histone chromatin mark H3K27ac across the body of DE genes between LP-WT and KO cells (Supplementary Fig. S7F).
To further determine what cellular pathway EBNA-LP regulates, we performed gene ontology analysis of DE genes. Genes involved in B cell signaling were enriched among upregulated genes in KO cells. Among the upregulated pathway, we found RHOGDI signaling, which contribute through B cell receptor (BCR) signaling to activation and differentiation of B cells into antibody-producing plasma cells or memory B cells (Fig. 7C). CIBERSORTx [71] analysis of RNA-seq data confirmed that compared to LP-WT, KO cells exhibit a global transcriptional profile indicative of differentiation into memory B-cells, while WT cells were transcriptionally similar to activated immune cells which is consistent with the preference of EBNA-LP KO virus to infect memory B-cells (Supplementary Fig. S6B). Consistent with the importance of EBNA-LP for induction of transcriptional programs for B cells activation, downregulated genes in KO cells were enriched for targets of TFs essential for EBV-mediated activation and proliferation of B cells, including RUNX1, STAT3, EZH2, and HIF1A (Fig. 7D). Among the downregulated genes in LP-KO cells, we observed a significant enrichment for targets regulated by TFs involved in B cell differentiation, including FOSL1, TP63, CBFB, KLF2 and SMAD7 (Fig. 7D). Next, we assessed whether EBNA-LP and YY1 bind the EBNA-LP KO DE genes. EBNA-LP binding occurred at the TSS or TES region of both upregulated and downregulated genes (Fig. 7E). Moreover, in downregulated genes YY1 binding significantly decrease across the gene body of EBNA-LP targets (Fig. 7E), while in upregulated genes, we observed a loss of YY1 binding preferentially at the TES in EBNA-LP KO cells compared to WT (Fig. 7E).
Our RNA-seq data suggests that EBNA-LP is necessary for the expression of cellular genes for the establishment of a latent EBV infection in B cells partly through its interaction with YY1. To test whether EBNA-LP-deregulated genes that we found in our cell lines changed in expression upon EBV infection, we assessed how EBNA-LP DE genes behave during primary infection of B cells with EBV. We used available RNAseq datasets [72] at different timepoints after EBV infection and correlated those with our RNAseq data (Supplementary Fig. S6C). We speculated that genes that are upregulated in EBNA-LP KO LCLs should be repressed by EBNA-LP in wild-type EBV-infected cells compared to uninfected ones, and conversely, genes downregulated in EBNA-LP KO LCLs should be upregulated by EBNA-LP expression in EBV infected cells. Notably, while we did not observe this reciprocal relationship for genes elevated in LP-KO LCLs, for genes downregulated in LP-KO cells we observed increasing levels of upregulation over time after EBV infection of B cells (Supplementary Fig. S6C with most of the EBNA-LP targets becoming significantly upregulated from 7 and 28 days post-EBV infection, consistent with expression of EBNA-LP in infected cells after 2 d.p.i. These data indicate that EBNA-LP is necessary for the expression of cellular genes involved in EBV-mediated cell transformation.
EBNA-LP KO influences H3K27ac deposition and SE looping
Given the differences in chromatin accessibility and gene expression upon EBNA-LP KO we decided to investigate how the well-known open chromatin associated histone mark, H3K27ac, changes in LP-KO cells. We performed ChIP-seq for H3K27ac in LP-WT and KO cells and performed a differential analysis between the two conditions using DESeq2. We observed that 30% of the variance could be explained by the absence of EBNA-LP (Supplementary Fig. S7A). We identified 5137 H3K27ac peaks that significantly change upon EBNA-LP KO (Fig. 8B). There was no substantial difference in localization of these peaks, most of which are localized at intergenic/intron regions (Supplementary Fig. S7G). Moreover, differences in H3K27ac deposition positively correlate with differences in gene expression and RNA Pol II occupancy across the gene body (Supplementary Fig. S7E and F). We then performed a motif analysis on the H3K27ac differential peaks and observed again ETS-motif binding TFs mainly present at gained peaks, while NFkB and bZIP-motif binding TFs in lost peaks (Fig. 8F).
Figure 8.
H3K27ac and RNA Pol II binding change upon B cell infection with an EBNA-LP Ko virus. (A) Heatmaps showing H3K27ac and RNA Pol II ChIP-seq signals in EBNA-LP WT and KO cell lines on YY1 peaks. Clusters are the same as in Fig. 2A. (B) Volcano plot representing differentially H3K27ac-rich regions between EBNA-LP WT and KO LCLs. (C) YY1 ChIP-seq signal on EBNA-LP/YY1-bound genomic regions (cluster 2 of the heatmap in (A)). (D) Violin plots showing H3K27ac (left) and RNA Pol II (right) coverage per replicate for all clusters in (A) with respective p-value (calculated using Wilcoxon t test). (E) UCSC Genome Browser view of one of the top ten downregulated genes, AMOTL1. (F) HOMER motif analysis on the differentially enriched regions for H3K27ac in EBNA-LP WT and KO LCLs. (G) Number of loops that originate from an SE in EBNA-LP WT and KO cell lines. (H) Violin plots representing H3K27ac signal in upregulated (left) and downregulated (right) SEs in EBNA-LP WT and KO cells per replicate. P-values are reported on top of the graphs and calculated using Wilcoxon t test.
YY1 binding and chromatin loops have been linked to SE functions [28, 73]. Both EBNA2 and EBNA-LP are bound together to SEs, and while the function of EBNA2 has been characterized at these regions, the role of EBNA-LP remains unknown [74, 75]. We used ChIP-seq to first assess deposition of the active chromatin mark H3K27ac at YY1 peaks between LP-WT and KO cells and integrate this information with RNA Pol II localization by ChIP-seq in the same samples (Fig. 8A). We observed that H3K27ac deposition coincides with RNA Pol II localization at YY1 binding sites in both WT and LP-KO cells (Fig. 8A). However, in EBNA-LP KO cells RNA Pol II occupancy significantly decreased at YY1 peaks that would normally bind EBNA-LP (Fig. 8A and 8D, right), while H3K27ac deposition seems unaffected on cluster 2, where YY1 and EBNA-LP bind together, while decreases at cluster 1 where EBNA2 is strongly bound together with YY1 and EBNA-LP (Fig. 8A and 8D, left) suggesting that in the absence of EBNA-LP, RNA Pol II is less likely to be recruited to active gene promoters. At YY1 peaks previously seen to be occupied by EBNA-LP but not EBNA2 (cluster 2), we observed a noticeably lower YY1 occupancy in EBNA-LP KO LCLs (Fig. 8C). These data suggest that EBNA-LP might stabilize YY1 binding which in turn affects RNA Pol II occupancy at these genomic regions. Next, we used HOMER to identify and compare SEs in LP-WT and KO cells. All detected SEs are in common to both LP-WT and KO cells but H3K27ac signal changed between cell types (Fig. 8H, Supplementary Fig. S7B). We were able to identify 24 SEs that exhibited a high H3K27ac signal in the LP-KO cells and 30 SEs that exhibited a higher signal in the WT cells. All these SEs are characterized by a high occupancy of EBNA-LP and a lower binding of EBNA2 compared to input signal (Supplementary Fig. S7C). Interestingly, those SEs that are preferentially enriched for H3K27ac histone mark in the LP-KO cells show a lower H3K27ac signal in transformed B cells, while those with a higher H3K27ac signal in LP-WT LCLs exhibit a higher H3K27ac enrichment after B cell transformation by EBV (Supplementary Fig. S7D). These data suggest that EBV-driven SE formation is impaired in EBNA-LP knockout LCLs, while SEs normally present in uninfected B cells still acquire the H3K27ac activation mark upon infection with the EBNA-LP KO virus.
SEs are associated with the formation of extensive chromatin loops. Therefore we interrogated our Hi-C dataset to determine the number of chromatin interactions originated by SEs in WT and KO cells. In LP-KO LCLs, SEs participated in more distinct chromatin loops than in WT cells (Fig. 8G) which is consistent with our observation that the absence of EBNA-LP increases promiscuous long-range interactions between TADs (Fig. 5C and Supplementary Fig. S3D).
These results suggest that EBNA-LP plays a crucial role in the regulation of chromatin architecture and the functional state of SEs, with a subset of SEs existing in B cells that are directly influenced by EBNA-LP.
EBNA-LP absence influences gene expression through chromatin looping
Our analysis indicates that EBNA-LP together with YY1 contributes to redesign B cell chromatin structure. To determine whether these changes in chromatin structure contribute to changes in transcription due to EBNA-LP expression we overlapped changing chromatin loops (Hi-C data) with differentially expressed genes (RNA-seq data) to identify genes that are regulated by EBNA-LP through chromatin looping (Supplementary Fig. S8). We identified 172 deregulated genes that laid within chromatin loops deregulated in LP-KO cells compared to WT cells (Supplementary Fig. S8). Our analysis revealed a significant correlation between changes in gene expression and the formation of chromatin loops. Using stringent conditions, we found 82 genes where increased transcription was linked to loop formation, or decreased transcription coincided with loop disruption (Supplementary Fig. S8A). These genes were notably enriched in functions related to immune response and B cell activity (Supplementary Fig. S8A).
In addition, we identified a separate group of 90 genes that showed the opposite correlation, where increased transcription was associated with loop disruption, and vice versa (Supplementary Fig. S8B). Our gene ontology analysis indicated that these genes were enriched for hormone response, and cytokine-mediated response, supporting the role of EBNA-LP in regulating B cell activation in EBV infected cells (Supplementary Fig. S8B) and suggesting that EBV, through EBNA-LP, reshapes the biological responses of infected cells to cytokines. Overall, these results indicate that EBNA-LP-mediated chromatin loops are closely associated with gene expression changes, where the formation and disruption of loops selectively influence genes involved in immune responses, B cell functions, and cell differentiation processes.
EBNA-LP and YY1 are essential in shaping EBV genome 3D organization.
EBV genome is epigenetically regulated by cellular factors that control viral chromatin and contribute to form structural interactions within the viral epigenome. Since we found that EBNA-LP, together with YY1, is necessary for the formation of chromatin loops across the host genome, we determine whether similarly EBNA-LP and YY1 participate in modulating the 3D architecture of the EBV genome. ChIP-seq experiments showed that EBNA-LP and YY1 colocalize at two regions of the viral genome: the RPMS1 promoter and the LMP1/LMP2B promoter, adjacent to the TR locus (Fig. 9A). By HiChIP assay we confirmed that the RPMS1-LMPs loop is occupied by YY1. Moreover, we found that in WT cells these regions form a chromatin loop, that is lost in KO cells, indicating that EBNA-LP is necessary for this chromatin loop formation (Fig. 9A). Next, we assessed how the loss of the YY1-mediated interaction affected the chromatin composition of these loci. Our ChIP-seq data showed a decrease in YY1 binding, and H3K27ac deposition at both loci in LP-KO cells compared to WT (Fig. 9C and D). These changes in YY1 binding correlated with a slightly increased CTCF occupancy at the RPMS1 locus but no changes at the LMPs promoter (Fig. 9C). Consistent with the observed decrease in YY1 binding, we observed a substantial decrease of RNA Pol II signal across the RPMS1 region (Fig. 9C and D) and to a lesser extent across the LMP2A coding regions (Fig. 9C). To determine whether these changes in chromatin organization correlated with changes in transcription, we analyzed our RNA-seq for changes in viral transcripts between WT cells and EBNA-LP KO cells. We found three viral transcripts that were significantly downregulated in KO cells compared to WT cells (Fig. 9B). Notably, all the three downregulated transcripts belong to the BARTs transcript family, which is located within the YY1-chromatin loop previously described as lost in LPKO cells. These results are consistent with the epigenetic data showing loss of open chromatin and loss of chromatin looping in LP-KO cells compared to WT cells. In particular, our data suggest that RPMS1 loop is important to facilitate BART transcription. Overall these results indicate that EBNA-LP contributes together with YY1 at organizing the three-dimensional structure of the EBV genome and to modulating the expression of viral genes important for persistent viral infection according to the model we have proposed in Fig. 10.
Figure 9.
The absence of EBNA-LP affects also EBV genome architecture and gene expression. (A) YY1 HiChIP iced-normalized matrices for the EBV genome in WT (top half) and KO (bottom half) showing the region of the viral genome between 100 Kb and ∼172 Kb. On the left side and on top of the matrices are represented EBNA2 (blue), EBNA-LP (purple), and YY1 (orange) ChIP-seq tracks. The bottom of the matrices indicated the loops originating from the YY1 ChIP-seq peak on the RPMS1 promoter. The color of the ribbons represents the log2 ratio of read depth normalized counts between EBNA-LP WT and KO LCLs. (B) Volcano plot showing differentially expressed viral genes (blue = downregulated, red = upregulated, grey = unchanged). (C) UCSC Genome Browser view of ATAC-seq (red), CTCF CUT&RUN (light blue), H3K27ac (light purple), RNA Pol II (dark purple), YY1 (orange), EBNA2 (blue), EBNA-LP (purple) ChIP-seq and RNA-seq (bottom four lines), and YY1 HiChIP tracks on the viral genome focused on the RPMS1 gene. (D) Average plots showing RNA Pol II (top) and YY1 (bottom) ChIP-seq signals on the RPMS1 promoter bbewteen EBNA-LP WT and KO cell lines.
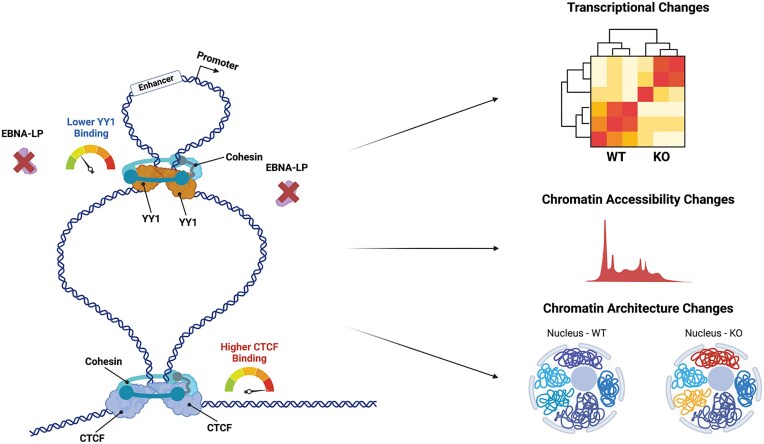
Figure 10.
EBNA-LP is essential in organizing the 3D structure of the genome in EBV-transformed cell lines. Schematic of working model for EBNA-LP KO. The KO of EBNA-LP induces a reduction in YY1 binding and increase in CTCF binding and global changes in transcription, chromatin accessibility and architecture. Created in BioRender. Tempera, I. (2025) https://BioRender.com/vz4kxsr.
Discussion
EBNA-LP plays a crucial role in B cell transformation caused by EBV by activating EBNA2 [7, 13, 76–78]. Although it was previously believed that transcriptional regulation by EBNA-LP only worked alongside EBNA2, recent research using ChIP-seq to map EBNA-LP occupancy in B cells, has revealed that only 29% of EBNA-LP binding overlaps with EBNA2 [15]. Additionally, genes whose EBNA2-mediated activation was previously identified as boosted by EBNA-LP activity did not exhibit reduced expression during LP-KO infection [17]. This suggests that EBNA-LP regulates B cell transformation independently of EBNA2-induced gene activation. Our current study supports this idea, showing that EBNA-LP influences the three-dimensional structure of the host chromatin by connecting regulatory elements such as enhancers and promoters. This function of EBNA-LP is mostly separate from the one EBNA2 exerts by binding to the genome and involves the ability of EBNA-LP to interact with the cellular factor YY1 [66].
YY1 plays a crucial role in B cell biology and differentiating B cells in the germinal center [79–83]. Our functional genomic studies have revealed that in EBV + B cells, EBNA-LP and YY1 combine to stabilize a network of long-distance chromatin interactions that connect regulatory regions with enhancers. By comparing the 3D structure of the host genome in B cells before and after EBV infection, we discovered that EBNA-LP boosts YY1 binding either stabilizing or disrupting functional chromatin interactions that are vital for driving cell proliferation and activation.
Our data suggest that YY1 binds chromatin regardless of EBNA-LP presence, but the addition of EBNA-LP allows YY1 binding sites to loop more stably with genomic regions that have epigenetic features of active enhancers, thus reshaping the 3D organization of the host genome. Our data indicate that the YY1-mediated chromatin loops promoted by EBNA-LP occur within TADs, suggesting that EBNA-LP expression may strengthen TAD identity. Consistent with this hypothesis, compared to KO LCL cells, EBNA-LP WT cells show an increase in intra-TADs interaction frequency and more robust boundaries, which indicate that in B cells infected with EBV and expressing EBNA-LP, the chromatin structure becomes more rigidly compartmentalized and less promiscuous in its interactions. The increased TAD integrity mediated by EBNA-LP can help maintain stable regulatory programs in infected B cells, as regulatory elements (such as enhancers and promoters) are more likely to interact with their intended targets within the same TAD. It is well known that TADs are maintained by the TF CTCF and their boundaries are enriched in arrays of CTCF motifs [84]. Our findings show that the decrease in YY1 binding observed upon EBNA-LP KO is accompanied by an increase in CTCF binding on the same regions suggesting an interplay between the two architectural proteins, which has previously been described [27, 85]. Moreover, the more promiscuous interactions observed between TADs is a result of a reduction of the insulation score at CTCF sites, even though the binding of CTCF is increased at these regions. Genes within a well-defined TAD are more likely to be coregulated, as they are more insulated from the influence of regulatory elements outside their TAD. This can result in a more controlled and specific expression pattern [86]. In this regard, we observed that IgG genes that cluster together on chromosomes 2, 14, and 22 are among the most down-regulated genes in EBNA-LP KO cells and exhibit an altered chromatin conformation respect to WT cells. In line with the concept that EBNA-LP stabilizes YY1 binding to promote the formation of chromatin loops within TADs, we noticed changes in stripes in EBNA-LP KO cells. Stripes are linear patterns of increased interaction frequency in Hi-C data, which extend from a specific genomic locus and indicate regions where an enhancer interacts with multiple promoters across a region, facilitating coordinated gene regulation. Overall, our data suggest that EBNA-LP could fine-tune and coregulate the transcription of several genes by regulating the 3D structure of the host genome.
Our analysis of enhancers and SEs in the context of EBNA-LP expression and its interaction with YY1 and EBNA2 suggests that EBNA-LP-mediated chromatin looping is necessary to bring promoters close to enhancer regions. Moreover, EBNA-LP seems essential in the formation of SEs during EBV-driven transformation. By comparing EBNA-LP WT and KO LCLs we were able to identify two sets of SEs that have altered H3K27ac mark that is directly correlated to its deposition during B cell transformation. It is worth noticing that those SEs that show a decreased H3K27ac histone mark in LP-KO cells are those that gain the same mark during transformation, consistent with previous data [15, 57, 87]. On the contrary, those SE regions that gain H3K27ac mark in LP-KO cells appear to be enriched for H3K27ac in B cell rather than LCLs. This suggests that EBNA-LP, together with EBNA2, is essential in the formation of a subset of EBV-associated SEs and disruption of B cell-specific SEs. Interestingly, the overall number of interactions originating from the SEs identified in both LP-WT and KO cell lines are more in LP-KO LCLs, again corroborating the evidence that LP-KO causes a reduction in TAD identity and therefore an increase in interaction frequency from active open chromatin regions like SEs. As EBV infection of B cells mimics the epigenetic remodeling of antigen-activated B cells through the germinal center, it is tempting to speculate that similarly memory B cells that emerged from the germinal center may acquire and maintain this similar set of enhancers and SEs, which, on the contrary, naïve B cells lack entirely. This model could, therefore, explain why EBNA-LP is essential for the immortalization of naïve but not memory B cells [17, 88, 89]. In fact, EBV infection of naïve B cells may rely on EBNA-LP to assist in the formation and stabilization of these enhancer and SE regions across the host. In contrast, EBV infection of memory B cells may not require EBNA-LP expression as certain essential structural changes in enhancer configurations are already present in mature differentiated B cells. This model could also explain the importance of EBNA-LP to interact with YY1 and bring YY1 regions in contact with these EBV-associated enhancers, as naïve B cells require YY1 to differentiate into germinal center B cells and plasma cells properly [55, 79–89].
How EBNA-LP, by interacting with it, can manipulate YY1 to form particular configurations of loops remains unknown. EBNA-LP can disrupt PML nuclear bodies by interacting with and displacing Sp100 [90, 91]. Sp100 is a major component of PML nuclear bodies and interacts with proteins involved in heterochromatin formation. Sp100 can also regulate ETS1 [92, 93] a TF that controls lymphoid cell differentiation, survival, and proliferation. Interestingly, genomic regions in which the open chromatin structure is perturbed in EBNA-LP KO cells are enriched for ETS1 binding motif. It is tempting to speculate that through YY1 binding and the displacement of Sp100, EBNA-LP could bring YY1-binding regions close to ETS1-associated promoters, modulating the expression of ETS1-regulated genes.
EBNA-LP enhances EBNA2-driven transcription from the Cp and LMP1 promoters in transient transfection assays using reporter constructs [7, 13]. Additionally, it boosts EBNA2 in up-regulating LMP1 expression in type I EBV-infected BL cells. Our data show that EBNA-LP and YY1 co-bind at several viral genomic regions. We confirmed previous observations showing that YY1 binds upstream of BZLF1 [94], although our mapping suggests that YY1 binding occurs precisely at the TSS for the BZLF1 gene rather than upstream of the promoter; this difference with previous data could be due to the ability of ChIP-seq to map more precisely binding sites compared to quantitative ChIP. As observed across the host genome, our data indicate that YY1 and EBNA-LP co-binding occur at viral genomic regions we have recently identified to possess the epigenetic signature associated with enhancers, including the LMP1/TR region and the region upstream of the RPMS1 promoter [4]. Previous work suggested that EBNA-LP coregulates LMP1; however, our RNA-seq data show no differences in LMP1 expression in LCL cells established through infection with EBNA-LP KO or wild-type EBV. Instead, we observed that all the BART transcripts expression levels significantly decreased in KO cells. Our HiChIP data showed that YY1 promotes a chromatin loop formation between RPMS1 and the enhancer TR region, and in LCL EBNA-LP KO cells, YY1 binding significantly decreases at the RPMS1 region engaged in the chromatin loop. This data reveals that EBNA-LP could be a coregulator of the BART transcripts, which has been reported to interact with the CBF1 repressor complex and negatively regulate EBNA2 [95]. Moreover, the RPMS1 locus also expresses an EBV circular RNA (circRNA) [96], whose precise function remains largely unknown, but it may interact with OriLyt during EBV lytic replication [97], suggesting a potential role for YY1-EBNA-LP chromatin loops in regulating EBV viral cycle.
The reshaping of the chromatin interaction landscape by EBNA-LP interaction with YY1 underscores the ability of EBV to stabilize host cell gene expression through extensive chromatin remodeling. The formation of new enhancer-promoter loops and the reorganization of chromatin compartments are likely mechanisms by which EBNA-LP exerts its oncogenic effects. These findings highlight potential targets for therapeutic intervention aimed at disrupting the chromatin interactions essential for EBV-driven transformation and proliferation of B cells.
Supplementary Material
Acknowledgements
Graphical abstract created in BioRender. Tempera, I. (2025) https://BioRender.com/4r8z6jp..
Author contributions: D.M. (Conceptualization, data curation, investigation, formal analysis, validation, visualization, writing – original draft, and writing – review & editing); L.B.C. (Formal analysis, investigation, and validation); J.M.C. (Investigation); R.S. (Investigation); S.P.A. (Investigation); R.E.W. (Resources and writing – review & editing); M.A.L. (Resources and writing – review & editing); I.T. (Conceptualization, funding acquisition, project administration, supervision, visualization, writing – original draft, and writing – review & editing).
Contributor Information
Davide Maestri, The Wistar Institute, Genome Regulation and Cell Signaling Program, 3601 Spruce Street Philadelphia PA 19104, United States.
Lisa B Caruso, The Wistar Institute, Genome Regulation and Cell Signaling Program, 3601 Spruce Street Philadelphia PA 19104, United States.
Jana M Cable, Department of Molecular Genetics and Microbiology, Duke University School of Medicine, Durham, NC 27710, United States.
Rachel Sklutuis, The Wistar Institute, Genome Regulation and Cell Signaling Program, 3601 Spruce Street Philadelphia PA 19104, United States.
Sarah Preston-Alp, The Wistar Institute, Genome Regulation and Cell Signaling Program, 3601 Spruce Street Philadelphia PA 19104, United States.
Robert E White, Section of Virology, Department of Infectious Disease, Imperial College London, London, SW7 2AZ, United Kingdom.
Micah A Luftig, Department of Molecular Genetics and Microbiology, Duke University School of Medicine, Durham, NC 27710, United States.
Italo Tempera, The Wistar Institute, Genome Regulation and Cell Signaling Program, 3601 Spruce Street Philadelphia PA 19104, United States.
Supplementary data
Supplementary data is available at NAR online.
Conflict of interest
None declared.
Funding
Wistar Institute core facilities was supported by Cancer Center Support Grant [P30 CA010815]; I.T. is supported by NIH R01AI130209, R01AI182056, R01AI153508, P01 CA269043, and P30 CA010815; M.L. and R.W. are supported by NIH R01CA140337; S.P.A. is supported by T32CA09171, R.S. is supported by T32CA288356. Funding to pay the Open Access publication charges for this article was provided by NIH grant funding.
Data availability
The datasets generated for this article are available in the GEO repository (GSE276491).
References
- 1. Cohen JI, Fauci AS, Varmus H et al. Epstein-Barr Virus: an important vaccine target for cancer prevention. Sci Transl Med. 2011; 3:107fs7. 10.1126/scitranslmed.3002878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Price AM, Luftig MA To be or not IIb: a multi-step process for Epstein-Barr Virus latency establishment and consequences for B cell tumorigenesis. PLoS Pathog. 2015; 11:e1004656. 10.1371/journal.ppat.1004656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Morgan SM, Tanizawa H, Caruso LB et al. The three-dimensional structure of Epstein-Barr virus genome varies by latency type and is regulated by PARP1 enzymatic activity. Nat Commun. 2022; 13:187. 10.1038/s41467-021-27894-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Maestri D, Napoletani G, Kossenkov A et al. The three-dimensional structure of the EBV genome plays a crucial role in regulating viral gene expression in EBVaGC. Nucleic Acids Res. 2023; 51:12092–110. 10.1093/nar/gkad936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Caruso LB, Maestri D, Tempera I Three-dimensional chromatin structure of the EBV genome: a crucial factor in viral infection. Viruses. 2023; 15:1088. 10.3390/v15051088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Zhao B, Zou J, Wang H et al. Epstein-Barr virus exploits intrinsic B-lymphocyte transcription programs to achieve immortal cell growth. Proc Natl Acad Sci USA. 2011; 108:14902–7. 10.1073/pnas.1108892108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Alfieri C, Birkenbach M, Kieff E Early events in Epstein-Barr virus infection of human B lymphocytes. Virology. 1991; 181:595–608. 10.1016/0042-6822(91)90893-G. [DOI] [PubMed] [Google Scholar]
- 8. Wood CD, Veenstra H, Khasnis S et al. MYC activation and BCL2L11 silencing by a tumour virus through the large-scale reconfiguration of enhancer-promoter hubs. eLife. 2016; 5:e18270. 10.7554/eLife.18270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Kaiser C, Laux G, Eick D et al. The proto-oncogene c-myc is a direct target gene of Epstein-Barr virus nuclear antigen 2. J Virol. 1999; 73:4481–4. 10.1128/JVI.73.5.4481-4484.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Zhao B, Maruo S, Cooper A et al. RNAs induced by Epstein-Barr virus nuclear antigen 2 in lymphoblastoid cell lines. Proc Natl Acad Sci USA. 2006; 103:1900–5. 10.1073/pnas.0510612103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Jiang S, Zhou H, Liang J et al. The Epstein-Barr virus regulome in lymphoblastoid cells. Cell Host Microbe. 2017; 22:561–573. 10.1016/j.chom.2017.09.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Lu F, Chen H-S, Kossenkov AV et al. EBNA2 drives formation of new chromosome binding sites and target genes for B-cell master regulatory transcription factors RBP-jκ and EBF1. PLoS Pathog. 2016; 12:e1005339. 10.1371/journal.ppat.1005339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Harada S, Kieff E Epstein-Barr virus nuclear protein LP stimulates EBNA-2 acidic domain-mediated transcriptional activation. J Virol. 1997; 71:6611–8. 10.1128/jvi.71.9.6611-6618.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Portal D, Zhao B, Calderwood MA et al. EBV nuclear antigen EBNALP dismisses transcription repressors NCoR and RBPJ from enhancers and EBNA2 increases NCoR-deficient RBPJ DNA binding. Proc Natl Acad Sci USA. 2011; 108:7808–13. 10.1073/pnas.1104991108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Portal D, Zhou H, Zhao B et al. Epstein-Barr virus nuclear antigen leader protein localizes to promoters and enhancers with cell transcription factors and EBNA2. Proc Natl Acad Sci USA. 2013; 110:18537–42. 10.1073/pnas.1317608110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Wang C, Zhou H, Xue Y et al. Epstein-Barr virus nuclear antigen leader protein coactivates EP300. J Virol. 2018; 92:e02155–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Szymula A, Palermo RD, Bayoumy A et al. Correction: epstein-Barr virus nuclear antigen EBNA-LP is essential for transforming naïve B cells, and facilitates recruitment of transcription factors to the viral genome. PLoS Pathog. 2019; 15:e1007403. 10.1371/journal.ppat.1007403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Hsu JL, Glaser SL Epstein–Barr virus-associated malignancies: epidemiologic patterns and etiologic implications. Crit Rev Oncol Hematol. 2000; 34:27–53. 10.1016/S1040-8428(00)00046-9. [DOI] [PubMed] [Google Scholar]
- 19. Thorley-Lawson DA, Gross A Persistence of the Epstein–Barr Virus and the origins of associated lymphomas. N Engl J Med. 2004; 350:1328–37. 10.1056/NEJMra032015. [DOI] [PubMed] [Google Scholar]
- 20. Lieberman PM Virology. Epstein-Barr virus turns 50. Science. 2014; 343:1323–5. 10.1126/science.1252786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Dekker J, Mirny L The 3D genome as moderator of chromosomal communication. Cell. 2016; 164:1110–21. 10.1016/j.cell.2016.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Schoenfelder S, Fraser P Long-range enhancer–promoter contacts in gene expression control. Nat Rev Genet. 2019; 20:437–55. 10.1038/s41576-019-0128-0. [DOI] [PubMed] [Google Scholar]
- 23. Rao SSP, Huntley MH, Durand NC et al. A 3D map of the Human genome at kilobase resolution reveals principles of chromatin looping. Cell. 2014; 159:1665–80. 10.1016/j.cell.2014.11.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Lieberman-Aiden E, van Berkum NL, Williams L et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science. 2009; 326:289–93. 10.1126/science.1181369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Dixon JR, Selvaraj S, Yue F et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature. 2012; 485:376–80. 10.1038/nature11082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Kim S, Yu N-K, Kaang B-K CTCF as a multifunctional protein in genome regulation and gene expression. Exp Mol Med. 2015; 47:e166–. 10.1038/emm.2015.33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Beagan JA, Duong MT, Titus KR et al. YY1 and CTCF orchestrate a 3D chromatin looping switch during early neural lineage commitment. Genome Res. 2017; 27:1139–52. 10.1101/gr.215160.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Weintraub AS, Li CH, Zamudio AV et al. YY1 Is a structural regulator of enhancer-promoter loops. Cell. 2017; 171:1573–1588. 10.1016/j.cell.2017.11.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Mach P, Kos PI, Zhan Y et al. Cohesin and CTCF control the dynamics of chromosome folding. Nat Genet. 2022; 54:1907–18. 10.1038/s41588-022-01232-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Mumbach MR, Rubin AJ, Flynn RA et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat Methods. 2016; 13:919–22. 10.1038/nmeth.3999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Beer S, Wange LE, Zhang X et al. EBNA2-EBF1 complexes promote MYC expression and metabolic processes driving S-phase progression of Epstein-Barr virus–infected B cells. Proc Natl Acad Sci USA. 2022; 119:e2200512119. 10.1073/pnas.2200512119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Yang Y, Ye X, Dai R et al. Phase separation of Epstein-Barr virus EBNA2 protein reorganizes chromatin topology for epigenetic regulation. Commun Biol. 2021; 4:967. 10.1038/s42003-021-02501-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Vervoort SJ, Welsh SA, Devlin JR et al. The PP2A-Integrator-CDK9 axis fine-tunes transcription and can be targeted therapeutically in cancer. Cell. 2021; 184:3143–62. 10.1016/j.cell.2021.04.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Dobin A, Davis CA, Schlesinger F et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013; 29:15–21. 10.1093/bioinformatics/bts635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Liao Y, Smyth GK, Shi W featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014; 30:923–30. 10.1093/bioinformatics/btt656. [DOI] [PubMed] [Google Scholar]
- 36. Love MI, Huber W, Anders S Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014; 15:550. 10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Ramírez F, Ryan DP, Grüning B et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 2016; 44:W160–5. 10.1093/nar/gkw257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Krämer A, Green J, Pollard J et al. Causal analysis approaches in ingenuity pathway analysis. Bioinformatics. 2014; 30:523–30. 10.1093/bioinformatics/btt703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Newman AM, Steen CB, Liu CL et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat Biotechnol. 2019; 37:773–82. 10.1038/s41587-019-0114-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Kolde R pheatmap: pretty Heatmaps. 2010; 10.32614/CRAN.package.pheatmap. [DOI]
- 41. Li H, Durbin R Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics. 2010; 26:589–95. 10.1093/bioinformatics/btp698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Feng J, Liu T, Qin B et al. Identifying ChIP-seq enrichment using MACS. Nat Protoc. 2012; 7:1728–40. 10.1038/nprot.2012.101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Zhang Y, Liu T, Meyer CA et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 2008; 9:R137. 10.1186/gb-2008-9-9-r137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Heinz S, Benner C, Spann N et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell. 2010; 38:576–89. 10.1016/j.molcel.2010.05.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Ye T, Krebs AR, Choukrallah M-A et al. seqMINER: an integrated ChIP-seq data interpretation platform. Nucleic Acids Res. 2011; 39:e35–. 10.1093/nar/gkq1287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Wickham H ggplot2. 2016; Cham: Springer International Publishing. [Google Scholar]
- 47. Meers MP, Tenenbaum D, Henikoff S Peak calling by sparse enrichment analysis for CUT&RUN chromatin profiling. Epigenetics Chromatin. 2019; 12:42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Grandi FC, Modi H, Kampman L et al. Chromatin accessibility profiling by ATAC-seq. Nat Protoc. 2022; 17:1518–52. 10.1038/s41596-022-00692-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Danecek P, Bonfield JK, Liddle J et al. Twelve years of SAMtools and BCFtools. GigaScience. 2021; 10:giab008. 10.1093/gigascience/giab008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Servant N, Varoquaux N, Lajoie BR et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 2015; 16:259. 10.1186/s13059-015-0831-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Stansfield JC, Cresswell KG, Vladimirov VI et al. HiCcompare: an R-package for joint normalization and comparison of HI-C datasets. BMC Bioinf. 2018; 19:279. 10.1186/s12859-018-2288-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. van der Weide RH, van den Brand T, Haarhuis JHI et al. Hi-C analyses with GENOVA: a case study with cohesin variants. NAR Genom Bioinform. 2021; 3:lqab040. 10.1093/nargab/lqab040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Kramer NE, Davis ES, Wenger CD et al. Plotgardener: cultivating precise multi-panel figures in R. Bioinformatics. 2022; 38:2042–5. 10.1093/bioinformatics/btac057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Yoon S, Chandra A, Vahedi G Stripenn detects architectural stripes from chromatin conformation data using computer vision. Nat Commun. 2022; 13:1602. 10.1038/s41467-022-29258-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Lareau C, Aryee M hichipper: a preprocessing pipeline for assessing library quality and DNA loops from HiChIP data bioinformatics. Nat Methods. 2018; 15:155–6. 10.1038/nmeth.4583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Lareau CA, Aryee MJ diffloop: a computational framework for identifying and analyzing differential DNA loops from sequencing data. Bioinformatics. 2018; 34:672–4. 10.1093/bioinformatics/btx623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Wang C, Liu X, Liang J et al. A DNA tumor virus globally reprograms host 3D genome architecture to achieve immortal growth. Nat Commun. 2023; 14:1598. 10.1038/s41467-023-37347-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Kleiman E, Jia H, Loguercio S et al. YY1 plays an essential role at all stages of B-cell differentiation. Proc Natl Acad Sci USA. 2016; 113:E3911–3920. 10.1073/pnas.1606297113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Pioli PD, Weis JH Snail transcription factors in hematopoietic cell development: a model of functional redundancy. Exp Hematol. 2014; 42:425–30. 10.1016/j.exphem.2014.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Sun L, Zhao X, Liu X et al. Transcription factor Ascl2 promotes germinal center B cell responses by directly regulating AID transcription. Cell Rep. 2021; 35:109188. 10.1016/j.celrep.2021.109188. [DOI] [PubMed] [Google Scholar]
- 61. Zhuang Y, Cheng P, Weintraub H B-lymphocyte development is regulated by the combined dosage of three basic helix-loop-helix genes, E2A, E2-2, and HEB. Mol Cell Biol. 1996; 16:2898–905. 10.1128/MCB.16.6.2898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Juilland M, Gonzalez M, Erdmann T et al. CARMA1- and MyD88-dependent activation of Jun/ATF-type AP-1 complexes is a hallmark of ABC diffuse large B-cell lymphomas. Blood. 2016; 127:1780–9. 10.1182/blood-2015-07-655647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Grötsch B, Brachs S, Lang C et al. The AP-1 transcription factor Fra1 inhibits follicular B cell differentiation into plasma cells. J Exp Med. 2014; 211:2199–212. 10.1084/jem.20130795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Zhang XK, Moussa O, LaRue A et al. The transcription factor fli-1 modulates marginal zone and follicular B cell development in mice. J Immunol. 2008; 181:1644–54. 10.4049/jimmunol.181.3.1644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Darling TK, Lamb TJ Emerging roles for eph receptors and ephrin ligands in immunity. Front Immunol. 2019; 10:1473. 10.3389/fimmu.2019.01473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Cable JM, Reinoso-Vizcaino NM, White RE et al. Epstein-Barr virus protein EBNA-LP engages YY1 through leucine-rich motifs to promote naïve B cell transformation. PLoS Pathog. 2024; 20:e1011950. 10.1371/journal.ppat.1011950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Charlot C, Dubois-Pot H, Serchov T et al. A review of post-translational modifications and subcellular localization of Ets transcription factors: possible connection with cancer and involvement in the hypoxic response. Methods Mol Biol. 2010; 647:3–30. [DOI] [PubMed] [Google Scholar]
- 68. Davis S, Vanhoutte P, Pages C et al. The MAPK/ERK cascade targets both Elk-1 and cAMP response element-binding protein to control long-term potentiation-dependent gene expression in the dentate gyrus in vivo. J Neurosci. 2000; 20:4563–72. 10.1523/JNEUROSCI.20-12-04563.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Luo Y, Liu Y, Wang C et al. Signaling pathways of EBV-induced oncogenesis. Cancer Cell Int. 2021; 21:93. 10.1186/s12935-021-01793-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70. Wang H, Jain S, Li P et al. Transcription factors IRF8 and PU.1 are required for follicular B cell development and BCL6-driven germinal center responses. Proc Natl Acad Sci USA. 2019; 116:9511–20. 10.1073/pnas.1901258116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Steen CB, Liu CL, Alizadeh AA et al.. Kidder BL Profiling cell type abundance and expression in bulk tissues with CIBERSORTx. Stem Cell Transcriptional Networks, Methods in Molecular Biology. 2020; 2117:New York, NY: Springer US; 135–57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Wang C, Li D, Zhang L et al. RNA sequencing analyses of gene expression during Epstein-Barr Virus infection of primary B lymphocytes. J Virol. 2019; 93:e00226–19. 10.1128/JVI.00226-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Wang J, Wu X, Wei C et al. YY1 Positively regulates transcription by targeting promoters and super-enhancers through the BAF complex in embryonic stem cells. Stem Cell Rep. 2018; 10:1324–39. 10.1016/j.stemcr.2018.02.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Zhou H, Schmidt SCS, Jiang S et al. Epstein-Barr virus oncoprotein super-enhancers control B cell growth. Cell Host Microbe. 2015; 17:205–16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Hong T, Parameswaran S, Donmez OA et al. Epstein–Barr virus nuclear antigen 2 extensively rewires the human chromatin landscape at autoimmune risk loci. Genome Res. 2021; 31:2185–98. 10.1101/gr.264705.120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76. Nitsche F, Bell A, Rickinson A Epstein-Barr virus leader protein enhances EBNA-2-mediated transactivation of latent membrane protein 1 expression: a role for the W1W2 repeat domain. J Virol. 1997; 71:6619–28. 10.1128/jvi.71.9.6619-6628.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Sinclair AJ, Palmero I, Peters G et al. EBNA-2 and EBNA-LP cooperate to cause G0 to G1 transition during immortalization of resting human B lymphocytes by Epstein-Barr virus. EMBO J. 1994; 13:3321–8. 10.1002/j.1460-2075.1994.tb06634.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78. Peng C-W, Xue Y, Zhao B et al. Direct interactions between Epstein-Barr virus leader protein LP and the EBNA2 acidic domain underlie coordinate transcriptional regulation. Proc Natl Acad Sci USA. 2004; 101:1033–8. 10.1073/pnas.0307808100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79. Banerjee A, Sindhava V, Vuyyuru R et al. YY1 is required for germinal center B cell development. PLoS One. 2016; 11:e0155311. 10.1371/journal.pone.0155311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Green MR, Monti S, Dalla-Favera R et al. Signatures of murine B-cell development implicate Yy1 as a regulator of the germinal center-specific program. Proc Natl Acad Sci USA. 2011; 108:2873–8. 10.1073/pnas.1019537108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81. Zaprazna K, Atchison ML YY1 controls immunoglobulin class switch recombination and nuclear activation-induced deaminase levels. Mol Cell Biol. 2012; 32:1542–54. 10.1128/MCB.05989-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Liu H, Schmidt-Supprian M, Shi Y et al. Yin Yang 1 is a critical regulator of B-cell development. Genes Dev. 2007; 21:1179–89. 10.1101/gad.1529307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83. Pan X, Papasani M, Hao Y et al. YY1 controls Igκ repertoire and B-cell development, and localizes with condensin on the Igκ locus. EMBO J. 2013; 32:1168–82. 10.1038/emboj.2013.66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84. Nanni L, Ceri S, Logie C Spatial patterns of CTCF sites define the anatomy of TADs and their boundaries. Genome Biol. 2020; 21:197. 10.1186/s13059-020-02108-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85. Schwalie PC, Ward MC, Cain CE et al. Co-binding by YY1 identifies the transcriptionally active, highly conserved set of CTCF-bound regions in primate genomes. Genome Biol. 2013; 14:R148. 10.1186/gb-2013-14-12-r148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86. Rajderkar S, Barozzi I, Zhu Y et al. Topologically associating domain boundaries are required for normal genome function. Commun Biol. 2023; 6:435. 10.1038/s42003-023-04819-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87. Liang J, Zhou H, Gerdt C et al. Epstein-Barr virus super-enhancer eRNAs are essential for MYC oncogene expression and lymphoblast proliferation. Proc Natl Acad Sci USA. 2016; 113:14121–6. 10.1073/pnas.1616697113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88. Mannick JB, Cohen JI, Birkenbach M et al. The Epstein-Barr virus nuclear protein encoded by the leader of the EBNA RNAs is important in B-lymphocyte transformation. J Virol. 1991; 65:6826–37. 10.1128/jvi.65.12.6826-6837.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89. Pich D, Mrozek-Gorska P, Bouvet M et al. First days in the life of naive human B lymphocytes infected with Epstein-Barr Virus. mBio. 2019; 10:e01723–19. 10.1128/mBio.01723-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90. Ling PD, Peng RS, Nakajima A et al. Mediation of Epstein-Barr virus EBNA-LP transcriptional coactivation by Sp100. EMBO J. 2005; 24:3565–75. 10.1038/sj.emboj.7600820. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91. Echendu CW, Ling PD Regulation of Sp100A subnuclear localization and transcriptional function by EBNA-LP and interferon. J Interferon Cytokine Res. 2008; 28:667–78. 10.1089/jir.2008.0023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92. Wasylyk C, Schlumberger SE, Criqui-Filipe P et al. Sp100 interacts with ETS-1 and stimulates its transcriptional activity. Mol Cell Biol. 2002; 22:2687–702. 10.1128/MCB.22.8.2687-2702.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93. Yordy JS, Li R, Sementchenko VI et al. SP100 expression modulates ETS1 transcriptional activity and inhibits cell invasion. Oncogene. 2004; 23:6654–65. 10.1038/sj.onc.1207891. [DOI] [PubMed] [Google Scholar]
- 94. Zalani S, Coppage A, Holley-Guthrie E et al. The cellular YY1 transcription factor binds a cis-acting, negatively regulating element in the Epstein-Barr virus BRLF1 promoter. J Virol. 1997; 71:3268–74. 10.1128/jvi.71.4.3268-3274.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95. Zhang J, Chen H, Weinmaster G et al. Epstein-Barr virus BamHi-a rightward transcript-encoded RPMS protein interacts with the CBF1-associated corepressor CIR to negatively regulate the activity of EBNA2 and NotchIC. J Virol. 2001; 75:2946–56. 10.1128/JVI.75.6.2946-2956.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96. Ungerleider N, Concha M, Lin Z et al. The Epstein Barr virus circRNAome. PLoS Pathog. 2018; 14:e1007206. 10.1371/journal.ppat.1007206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97. Zhang J-Y, Du Y, Gong L-P et al. ebv-circRPMS1 promotes the progression of EBV-associated gastric carcinoma via Sam68-dependent activation of METTL3. Cancer Lett. 2022; 535:215646. 10.1016/j.canlet.2022.215646. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets generated for this article are available in the GEO repository (GSE276491).