Abstract
Breaching the vascular barrier is a critical step in pancreatic ductal adenocarcinoma (PDAC) metastasis, yet the mechanisms enabling this process remain incompletely understood. Transforming growth factor beta (TGFβ) receptors have been extensively studied in many cancer types. However, activin receptor-like kinase 7 (ALK7), one of the TGFβ receptors, is under-investigated, and its roles in PDAC metastasis have been unclear. This study identifies two distinct but interconnected ALK7-driven non-canonical pathways that promote PDAC dissemination. The ALK7–β-catenin–EMT axis enhances intrinsic tumor cell motility, driving epithelial-mesenchymal transition (EMT). In parallel, the ALK7–β-catenin–MMP axis facilitates metastatic invasion by upregulating MMP production, leading to ECM degradation and invadosome formation, which promote vascular barrier breakdown and intravasation. An orthotopic PDAC metastasis model reveals that both pharmacological and genetic ALK7 inhibition suppresses metastasis. 3D microfluidic vessel-on-chip platforms further demonstrate that ALK7 inhibition preserves basement membrane (BM) integrity, limiting intravasation. While MMP inhibition effectively blocks BM breakdown and intravasation, extravasation remains unaffected, highlighting distinct molecular requirements for different metastatic stages. These findings establish ALK7 as a dual-function pro-metastatic regulator that orchestrates both tumor cell plasticity and ECM remodeling, positioning ALK7 inhibition as a promising strategy to target early metastatic dissemination in PDAC.
Supplementary Information
The online version contains supplementary material available at 10.1186/s12943-025-02384-w.
Keywords: Pancreatic cancer, ALK7, Matrix metalloproteinases, Organ-on-a-chip, Intravasation
Introduction
Pancreatic ductal adenocarcinoma (PDAC) is among the most lethal malignancies, defined by its exceptional resistance to therapy, frequent recurrence, and high metastatic potential [1]. Despite decades of research, survival rates remain dismal, with fewer than 10% of patients exceeding five years post-diagnosis and over 80% succumbing to metastatic progression within two years [1, 2]. The uniquely hostile tumor microenvironment of PDAC, characterized by a dense desmoplastic stroma, not only acts as a barrier to drug delivery but also imposes significant physical constraints on tumor cell movement [3]. For most cancers, including PDAC, blood vessels are the critical mediators of metastasis [4]. However, unlike many other cancers, where metastasis depends on extravasation, PDAC metastasis is highly dependent on successful intravasation, as tumor cells must first overcome the fibrotic stroma and breach the vascular barrier to enter circulation [5, 6]. This makes intravasation a rate-limiting step in the dissemination of PDAC. Once in the bloodstream, circulating tumor cells (CTCs) can travel to distant organs, where extravasation occurs under less restrictive conditions. Given these unique constraints, targeting the mechanisms that regulate tumor-stroma interactions and vascular invasion represents a crucial strategy for limiting PDAC metastasis.
Transforming growth factor beta (TGFβ) receptors have been extensively studied in many types of cancers [7]. However, activin receptor-like kinase 7 (ALK7), one of the TGFβ receptors, is under-investigated, and its roles in cancer metastasis have been controversial. Some studies have reported that ALK7 is anti-metastatic [8–10]; by contrast, some other studies have reported that ALK7 is pro-metastatic [11–14]. Those controversies, delaying the translation of ALK7 inhibitors in development, could stem from researchers studying different types of cancers; but more importantly, those discrepancies came from what metastatic steps researchers focused on and mainly investigated using different experimental models. The TGFβ receptor signaling is a well-known regulator of anoikis, a specialized form of apoptosis that occurs when cells detach from the extracellular matrix (ECM) [15, 16]. While TGFβ can promote anoikis in normal and early-stage cancer cells, advanced cancer cells often exploit TGFβ signaling to resist anoikis by activating epithelial-mesenchymal transition (EMT) and pro-survival pathways [15, 16]. This anoikis resistance facilitates cancer cell survival in circulation, allowing efficient metastatic dissemination. Despite these findings, the roles of ALK7 in PDAC metastasis still remain to be elucidated.
In this study, we investigate the roles of ALK7 in PDAC metastasis. Using orthotopic PDAC spontaneous metastasis models in vivo and three-dimensional (3D) microfluidic platforms in vitro, we identify and report two distinct but interconnected ALK7-driven non-canonical pathways that promote PDAC dissemination. We further reveal the different contributions of ALK7 to tumor extravasation and intravasation, revealing that ALK7 inhibition can delay metastasis by primarily targeting intravasation.
Results
Pharmacological ALK7 inhibition suppresses spontaneous PDAC metastasis in an orthotopic PDAC model in vivo
To investigate the potential efficacy of ALK7 inhibition in PDAC metastasis in vivo (Fig. 1A), we established an orthotopic mouse model by injecting PDAC cells into the tail of the pancreas of 5-6-week-old immunodeficient NSG mice (NOD-scid IL2Rgammanull). The spontaneous metastasis model and treatment schedule are shown in Fig. 1B. To trace and quantify PDAC cells in live animals, PD7591 cells that express luciferase were generated and used for the orthotopic PDAC model (Supplementary Fig. S1). One week after tumor inoculation, SB431542, a widely used inhibitor of ALK family receptors, including ALK7, was administered for two weeks. At day 7, we sorted tumor-bearing mice into two cohorts with similar average tumor sizes. Next, we started to treat them with SB431542 (25 mg/kg/day, intraperitoneal (i.p.) injection), or dimethyl sulfoxide (DMSO as a vehicle) for two weeks. During the two weeks of the treatments, we assessed tumor growth weekly by injecting luciferin (150 mg/kg, i.p. injection) and measuring the bioluminescent intensity emitted from the primary tumors using the IVIS Xenogen imager. The total bioluminescent flux measured from the representative mice at 3 weeks (Fig. 1C) and the total, averaged flux from all mice during the 2-week period (Fig. 1D) indicated that there is no significant difference in primary tumor growths between the DMSO and SB431542-treated mice. We sacrificed the animals and harvested their primary tumors. Consistent with the bioluminescence-based data, we found that the overall tumor burden, as indicated by tumor weights, was not significantly different between the two groups (Fig. 1E), confirming that SB431542 treatment does not affect primary tumor growth. Next, we excised the livers (Fig. 1F, G) and lungs (Fig. 1H, I) to examine metastases in each cohort, given that the livers and lungs are one of the common distant sites of PDAC metastasis in human patients [17–20]. Noteworthily, we discovered that livers and lung metastasis was significantly reduced by the pharmacological ALK7 inhibition (Fig. 1F-I). We measured the body weight of mice receiving treatment to assess drug toxicity. No weight loss was observed in mice treated with the SB431542 inhibitor (25 mg/kg) (Supplementary Fig. S2A). Furthermore, we carried out histopathological assessments of vital organs; including the kidney, spleen, liver, and lung in non-tumor-bearing mice treated with either vehicle (DMSO) or SB431542 for two weeks. Evaluation of treatment toxicity revealed no detectable damage in these organs (Supplementary Fig. S2B). Taken together, pharmacological inhibition of ALK7 blocks spontaneous PDAC metastasis in an orthotopic in vivo model, without causing toxicity in animals. The inhibitory effects of SB431542 on aggressive PDAC phenotypes were further validated through in vitro assays. SB431542 treatment led to a dose-dependent reduction in PD7591 cell proliferation (Supplementary Fig. S3A). In transwell invasion assays, SB431542 significantly suppressed Matrigel invasion, while wound healing assays demonstrated a time-dependent impairment in cell migration (Supplementary Fig. S3B and C). In addition, SB431542 markedly reduced clonogenic capacity, as shown by colony formation assays (Supplementary Fig. S3D).
Fig. 1.
Pharmacological ALK7 blockade inhibits spontaneous PDAC metastasis in an orthotopic PDAC model in vivo. (A) A schematic of the spontaneous metastatic processes. (B) A schematic of the orthotopic PDAC spontaneous metastasis experiments with the SB431542 treatment in mice. Orthotopic PDAC tumor-bearing mice were prepared. After 1 week of tumor inoculation, DMSO or SB431542 (25 mg/kg/day, intraperitoneal (i.p.) injection) was treated for 2 weeks. (C) Representative IVIS images of PDAC tumor-bearing mice treated with DMSO or SB431542 (images at week 3). (D) Quantification of the bioluminescent signal from the luciferase-expressing tumors in PDAC tumor-bearing mice treated with DMSO or SB431542 at weeks 1, 2, and 3 after tumor inoculation (DMSO, n = 9; SB431542, n = 9). (E) Quantification of the weight of excised primary tumors obtained from the mice treated with DMSO or SB431542 (tumor excised at week 3). (F) Representative IVIS images of metastatic PDAC cells in the livers after treatment with SB431542 or DMSO (organs obtained at week 3). (G) Quantification of the bioluminescent signal from the luciferase-expressing PDAC cells metastasized to the livers of PDAC tumor-bearing mice treated with DMSO or SB431542. * p = 0.011, two-tailed unpaired Student t-test. (H) Representative IVIS images of metastatic PDAC cells in the lungs after treatment with SB431542 or DMSO (organs obtained at week 3). (I) Quantification of the bioluminescent signal from the luciferase-expressing PDAC cells metastasized to the lungs of PDAC tumor-bearing mice treated with DMSO or SB431542. * p = 0.0252, two-tailed unpaired Student t-test. Data were expressed as mean ± S.E.M. Individual data points are presented on the graphs. ‘ns’ means not significant. Representative data from one of three independent experiments is shown
Genetic ALK7 inhibition suppresses spontaneous PDAC metastasis in an orthotopic PDAC model in vivo
While these results suggested that SB431542 suppresses PDAC metastasis in vitro and in vivo, SB431542 can also block other TGFβ receptors such as ALK4 and ALK5. Also, the systemic injection of the inhibitor can affect other tissues and cells in addition to the cancer cells, making it hard to interpret the results. To minimize the risk and determine the exact roles of ALK7 in PDAC cells, we performed short hairpin RNA (shRNA) mediated ALK7 knockdown (KD) in PDAC cells prior to inoculating them into animals (Fig. 2A). Two different shRNAs against ALK7, including shALK7#1 and shALK7#2, were tested and showed approximately 40% and 60% KD efficiency, respectively, in western blotting (Supplementary Fig. S4A). We further examined PDAC cell phenotypes with or without ALK7 KD in vitro. ALK7 KD (shALK7#2) led to approximately a 30% reduction in cell proliferation, a 70% reduction in cell invasion, a significant delay in wound-healing processes, and a more than 50% reduction in colony formation efficiency (Supplementary Fig. S4B-E). Since shALK7#2 showed greater KD efficiency and phenotypic changes in PDAC cells, the shALK7#2 clone was selected for in vivo experiments. To further assess the ALK7 gene dosage effect, isogenic PDAC cell lines were generated with defined ALK7 expression levels: wild-type (WT; full expression), knockdown (KD; intermediate expression), and knockout (KO; minimal expression). Functional assays revealed a progressive suppression of aggressive tumor phenotypes with decreasing ALK7 expression. Specifically, proliferation rates, invasive capacity through Matrigel, wound healing ability, and colony formation efficiency were all significantly reduced in a gene dose–dependent manner (Supplementary Fig. S5). These findings support the conclusion that ALK7 expression is positively associated with proliferative and invasive behaviors in PDAC cells.
Fig. 2.
Genetic ALK7 inhibition suppresses spontaneous PDAC metastasis in an orthotopic PDAC model in vivo. (A) A schematic of the orthotopic PDAC spontaneous metastasis model with or without ALK7 knockdown (KD). PDAC cells were treated with shALK7#2, a shRNA targeting ALK7 (shALK7KD), or with a scrambled control shRNA (shControl). shALK7KD and shControl PDAC cells were injected into the tail of the pancreas of NSG mice (NOD-scid IL2Rgammanull) (n = 10), and tumor progression was observed for 3 weeks. (B) Primary tumor growth monitored with an IVIS imager (photons/s) at week 2. (C) IVIS images of PDAC tumor-bearing mice at week 3. (D) Quantification of bioluminescence signal from all primary tumors in shALK7KD and shControl mice at week 3 (n = 10). (E) Quantification of the weight of excised primary tumors obtained from the shALK7KD and shControl mice (n = 10). (F) Quantification of bioluminescence signal from the lungs (n = 10). IVIS images of metastatic PDAC cells in the lungs obtained from shALK7KD and shControl mice (organs obtained at week 3). (G) IVIS images of metastatic PDAC cells in the lungs obtained from shALK7KD and shControl mice (organs obtained at week 3). (H) Quantification of bioluminescence signal from the livers (n = 10). (I) IVIS images of metastatic PDAC cells in the livers obtained from shALK7KD and shControl mice (organs obtained at week 3). (J) IVIS images and total flux from CTCs ex vivo obtained from shControl- vs. shALK7KD PDAC tumor-bearing mice (n = 10). (K) The Cancer Genome Atlas (TCGA) survival analysis of human PDAC patients with upregulated ALK7 signaling compared to those with normal ALK7 signaling, Kaplan-Meier analysis with log-rank test in the regular ALK7 expression (n = 158, overall survival = 21.71 months) or ALK7 up-regulation (n = 19, overall survival = 15.11 months). O.S. indicates Overall Survival. * (p < 0.05) indicates statistical significance. ‘ns’ indicates not significant
To assess the role of ALK7 in tumor growth and metastasis in vivo, we established an orthotopic PDAC model by injecting wild type- or ALK7KD PDAC cells (shALK7KD) into the tail of the pancreas of 5-6-week-old immunodeficient NSG mice (NOD-scid IL2Rgammanull) (Fig. 2A). We monitored primary tumor growth weekly for 3 weeks. After 2 weeks of tumor inoculation, ALK7 KD did not affect PDAC tumor growth compared to controls (shControl: scramble shRNA), evidenced by quantification of the bioluminescence signal from primary tumors (p = 0.68) (Fig. 2B). We imaged tumors in the third week (Fig. 2C), similarly measured bioluminescence signal, and showed no significant difference between two groups; however, the ALK7 KD cohort had smaller tumors (p = 0.067) (Fig. 2D). We then sacrificed the animals and harvested their primary tumors, lungs and livers. We found no difference in tumor weights (p = 0.42) (Fig. 2E). Next, we assessed the effect of ALK7 knockdown (ALK7 KD) on lung metastasis. Lung metastasis was significantly reduced by ALK7 KD (p = 0.045) (Fig. 2F), suggesting that ALK7-specific inhibition limits distant lung dissemination, as shown in representative IVIS images (Fig. 2G). We also observed a trend toward reduced metastatic burden in the liver (p = 0.063) (Fig. 2H, I). Interestingly, bioluminescence signals from the liver were much lower (ranging from 105 to 106) compared to those from the lungs (106 to 107), suggesting that this disease model favors lung metastasis over liver metastasis. We next sought to determine whether PDAC intravasation was affected by ALK7 knockdown. To answer this question, we sought to detect circulating tumor cells (CTCs) in the blood circulation of the tumor-bearing mice inoculated with shALK7KD PDAC cells or shControl PDAC cells (n = 10). At week 3, we collected 350–550 µl of blood from two groups of mice and imaged CTCs after 24 h of plating. We quantified relative numbers of CTCs by measuring total bioluminescent flux in the samples, showing a trend of fewer CTCs in the ALK7 KD group compared to the control group, although there was no significant difference between groups (p = 0.119) (Fig. 2J, right).
Although these in vivo results suggested a promise that ALK7 inhibition can limit PDAC metastasis in animals, it remained unclear whether this is valid in human PDAC patients. To explore whether our findings may be relevant to human PDAC, we sought to determine a relationship between ALK7 and overall survival in human PDAC patients by using existing data sets generated by the Cancer Genome Atlas (TCGA) consortium and analyzed the relationship between patient survival and the expression of ALK7-related genes (ALK7, inhibin A, inhibin B) in patient tumors [21]. A total of 19 out of 177 (11%) patients were found to have upregulated expression of either ALK7 (ACVR1C), inhibin A (INHBA), or inhibin B (INHBB) in their tumor samples. Inhibin-βA and Inhibin-βB, respectively encoding activin A and -B, serve as ligands for ALK7. We included them all to better analyze functional aspects of the ALK7 with its ligand expression. On average, these patients survived for a significantly shorter period after their initial diagnosis than patients whose tumors expressed normal levels of ALK7-related genes (15.11 vs. 21.71, in months by Kaplan-Meier analysis; p = 0.0325 by log-rank test), suggesting that ALK7 signaling plays a critical pro-tumorigenic role worsening the outcomes in PDAC patients (Fig. 2K). Taken together, genetic ALK7 knockdown in PDAC blocks PDAC spread to the lungs in animal models, and its pro-tumorigenic role was further evidenced by human survival data.
A non-canonical ALK7/β-catenin axis promotes EMT and aggressiveness in PDAC
While our in vivo data suggested that PDAC can metastasize to distant organs in an ALK7-dependent manner, the mechanistic roles of ALK7 in PDAC metastasis are still unclear. To investigate the ALK7-mediated molecular processes that might be responsible for PDAC metastasis, we examined both the canonical and non-canonical downstream signaling pathways of TGFβ receptor in shControl (shC) and shALK7KD (shALK7) PDAC cells using western blot analysis (Fig. 3A). In either condition, endogenous phosphorylation of SMAD2/3, the critical components of the canonical pathway, was not observed. On the other hand, ALK7-mediated non-canonical signaling pathways such as RhoA/MLC2, AKT, and ERK (MAPK) were found to be constitutively activated but perturbed by ALK7 KD. ALK7 KD most dramatically downregulated the levels of RhoA, phospoh-MLC2, phospho-AKT, and phospho-JNK (Fig. 3A). The RhoA, AKT, and JNK pathways play a crucial role in mediating crosstalk between TGFβ signaling and epithelial-mesenchymal transition (EMT), which is the key driver of the aggressive phenotypes in metastatic cancer [22, 23]. As a common downstream target of the AKT, JNK, and RhoA pathways, GSK3β was analyzed by measuring the expression levels of its phosphorylated form along with total β-catenin, a key downstream effector. Western blot analysis showed that the phosphorylated GSK3β level was downregulated by ALK7 KD, accompanying a decrease in the β-catenin level (Fig. 3B). ALK7 KD downregulated nuclear expression of β-catenin, which was demonstrated by both immunofluorescent microscopy (Fig. 3C) and western blot analysis using fractionated cytoplasmic and nuclear proteins (Fig. 3D). Since β-catenin is a potent EMT regulator, the expressions of major EMT-related transcription factors were assessed. ALK7 KD downregulated the expression levels of ZEB1, SLUG, SNAIL, and TWIST1 (Fig. 3E).
Fig. 3.
A non-canonical ALK7/β-catenin pathway promotes EMT and aggressiveness in PDAC. (A) Western blotting to evaluate the activation status of canonical and non-canonical TGFβ receptor downstream pathways. Smad2/3 is associated with the canonical pathway, while RhoA, MLC2, AKT, ERK (MAPK), JNK, and p38 are associated with the non-canonical pathway. (B) Western blotting to assess the GSK3β/β-catenin axis. (C) Immunofluorescence to examine nuclear translocation of β-catenin. (D) Western blotting of β-catenin after cell fractionation: total, cytosolic, and nuclear fractions of cell lysates. (E) Western blotting to evaluate the expression of epithelial-mesenchymal transition (EMT) markers (ZEB1, SLUG, SNAIL, and TWIST1). (F) Western blotting to assess the expression of EMT markers in ALK7 KD PDAC cells, with or without transfection of β-catenin expressing plasmids. (G) Western blotting to examine known β-catenin target genes involved in cell proliferation (c-Myc and CyclinD1), stemness (CD44, SOX2, and OCT4), and cell migration (N-cadherin and Fibronectin) in ALK7 KD PDAC cells, with or without transfection of β-catenin expressing plasmids. (H) A summary of the findings. Scale bars (C) = 10 μm. * (p < 0.05), ** (p < 0.01), *** (p < 0.001) indicate statistical significance. ‘ns’ indicates not significant.Immunoblotting experiments were independently performed in triplicate (n = 3)
Based on these results, we hypothesized that β-catenin is a mediator for EMT characteristics of PDAC. To validate whether β-catenin is mainly responsible for ALK7 KD-mediated loss of EMT factors, a β-catenin expressing plasmid was additionally introduced into ALK7 KD PDAC cells. Western blot analysis showed that the induced β-catenin expression fully restored the expression of EMT-related transcription factors in ALK7 KD PDAC cells (Fig. 3F). β-catenin is not only the regulator of EMT but also directly regulates many essential genes, which collectively contribute to the aggressive phenotypes of cancers. Western blot analysis showed that ALK7 KD strongly downregulated the expression of cell proliferation-related genes (c-Myc and CyclinD1), drug-resistance/stemness-related genes (CD44, SOX2, and OCT4), and cell migration-related genes (N-cadherin and Fibronectin). When β-catenin overexpression was induced, the decreased expression of β-catenin target genes was fully restored (Fig. 3G). The restoration of aggressive phenotypes was further evaluated in ALK7 knockdown PDAC cells overexpressing β-catenin. Reintroduction of β-catenin substantially rescued proliferative capacity, invasive potential, migratory ability (wound closure), and clonogenic growth, all of which had been markedly impaired by ALK7 knockdown (Supplementary Fig. S6). Taken together, our findings support the idea that a non-canonical ALK7/β-catenin signaling pathway contributes to the EMT and aggressiveness in PDAC (Fig. 3H).
To assess the pharmacological inhibition of ALK7 signaling, downstream signaling alterations induced by SB431542 treatment in PD7591 cells were compared with those observed in ALK7 knockdown cells. Similar to ALK7 knockdown, SB431542 markedly reduced phosphorylation of MLC2, AKT, and p38 with moderate decreases in phospho-JNK and β-catenin. Overall, ALK7 knockdown led to a broader and more pronounced suppression of both canonical and non-canonical ALK7 downstream signaling pathways. Notably, phospho-p38 suppression was more prominent with SB431542, while reduction of phospho-ERK and RhoA was specific to ALK7 knockdown (Supplementary Fig. S7A). These findings were further validated in a 3D spheroid culture of PD7591 cells. SB431542 treatment in 3D culture robustly reduced the phosphorylation of MLC2 and AKT, along with moderate decreases in phospho-p38, phospho-JNK, and β-catenin. Consistent with the 2D results, no reduction in phospho-ERK was observed under these conditions (Supplementary Fig. S7B). In contrast, analysis of downstream signaling pathways in spheroids of shALK7 knockdown PD7591 cells revealed that ALK7 depletion led to a robust decrease in phospho-MLC2, phospho-AKT, phospho-JNK, phospho-ERK, and β-catenin, with a moderate reduction in RhoA—findings that were similar to those from the 2D culture. Notably, ALK7 knockdown resulted in a more pronounced reduction of phospho-ERK in the 3D setting compared to 2D (Supplementary Fig. S8).
The overall expression patterns of canonical and non-canonical ALK7 downstream molecules were comparable between 2D and 3D conditions, with only minor differences in the levels of phospho-JNK and phospho-p38.
ALK7/β-catenin signaling upregulates MMP7/9 expression and enhances PDAC invasiveness
Although these data suggest that the non-canonical ALK7/β-catenin signaling pathway promotes tumor EMT and aggressiveness in PDAC cells, the functional roles of ALK7 in PDAC are unclear. To understand the molecular function of ALK7 in PDAC without bias, we performed bulk RNA-sequencing of PDAC cells after transfection with control (shControl) and ALK7 shRNA vectors (shALK7KD) (Fig. 4A). Using the Kyoto Encyclopedia of Genes and Genomes (KEGG), one of the gene ontology (GO) analyses, we found that genes involved in interactions between focal adhesions, receptors on the cell surface, pathways such as phosphatidylinositol 3’-kinase (PI3K)-AKT signaling, and transendothelial migration were significantly downregulated in ALK7 KD as compared to control cells (Supplementary Fig. S9A), which is consistent with our data described in Fig. 3. Interestingly, our further GO analysis of molecular function (MF), biological processes (BP), and UniProtMF showed downregulation of cell migration, proteolysis, metalloprotease activity, and angiogenesis (Supplementary Fig. S9A-C).
Fig. 4.
ALK7/β-catenin signals upregulate MMP7/9 expression, enhancing invasiveness in PDAC cells. (A) A bulk RNA-sequencing (RNA-seq) and gene set enrichment analyses in ALK7-knockdown or control PDAC cells. The count means gene numbers categorized in a specific pathway identified. (B) Western blotting to assess MMP7 and MMP9 expression in wild-type PDAC or ALK7-silenced PDAC cells with or without β-catenin induction. (C) RT-qPCR to evaluate MMP7 and MMP9 mRNA transcription in wild-type PDAC or ALK7-silenced PDAC cells with or without β-catenin induction (n = 3). One-way ANOVA with Tukey’s multiple comparisons (D) Representative immunofluorescent (IF) images of PDAC cells (anti-GFP, green; phalloidin, red; and merged images with DAPI, blue), embedded in 2.5 mg/mL collagen with or without Batimastat (20 μM) treatment (24 h). (E) Number of invadosome-like protrusions in PDAC cells with or without Batimastat treatment (n = 4). (F) Protrusion lengths in PDAC cells with or without Batimastat treatment (n = 4). (G) Percent of circular cells with or without Batimastat treatment (n = 4). Scale bars (D) = 50 μm; ** (p < 0.01), *** (p < 0.001) **** (p < 0.0001) indicate statistical significance. ‘ns’ indicates not significant. Immunoblotting experiments were independently performed in triplicate (n = 3).
Given the prominent emergence of proteolytic and metalloprotease activities in our functional screening, we next analyzed the mRNA expression of 14 matrix metalloproteinases (MMPs) previously implicated in cancer invasion and metastasis. These MMPs were selected based on their reported or predicted regulation by TGF-β superfamily signaling, including ALK7–SMAD and β-catenin pathways [24, 25], with particular focus on MMP7 and MMP9, which are strongly linked to PDAC progression [26, 27]. Among the 14 MMPs examined, seven—MMP15, MMP9, MMP25, MMP19, MMP23, MMP3, and MMP7—were markedly downregulated in ALK7 knockdown PDAC cells (Supplementary Fig. S10). These include membrane-type MMPs (MMP15, MMP25, MMP23), which mediate local ECM degradation, and secreted MMPs (MMP9, MMP19, MMP3, MMP7) involved in basement membrane breakdown and metastasis. Notably, MMP7 is a direct transcriptional target of β-catenin, while MMP9 is indirectly regulated by β-catenin or TGF-β signaling in several cancers. To further delineate the gene dosage relationship between ALK7 and its downstream effectors, we employed an isogenic PDAC cell model with graded ALK7 expression levels—wild-type (high), knockdown (~ 40% of baseline), and knockout (minimal). Expression of β-catenin, MMP7, and MMP9 decreased proportionally with ALK7 expression, supporting a gene dosage-dependent regulatory axis (Supplementary Fig. S5 and S11). Importantly, the suppressed expression of MMP7 and MMP9 in ALK7 knockdown cells was fully restored at both the mRNA and protein levels upon β-catenin overexpression (Fig. 4B and C), confirming β-catenin as a key mediator of ALK7-dependent MMP regulation and reinforcing the biological relevance of MMP7 and MMP9 as central effectors in ALK7-driven PDAC invasiveness.
Given that MMP7/9 are regulated by ALK7/β-catenin signaling pathways, we sought to determine the formation of invadosome-like protrusions in PDAC cells treated with the broad-spectrum protease inhibitor Batimastat in a 3D culture system. (Fig. 4D). We seeded wild-type PDAC cells within a 3D collagen gel (2.5 mg/ml) and treated them with Batimastat (20 µM) or vehicle (DMSO) for a day. After 24 h of cell seeding and drug treatments, PDAC cells were fixed and stained with phalloidin. Cells extended their invadosome-like protrusions in collagen; however, Batimastat inhibited cell protrusions and made cells more circular (Fig. 4D). Indeed, our image analyses showed that Batimastat treatment reduced the number of protrusions per cell (p = 0.007) (Fig. 4E) and protrusion lengths (p = 0.0014) (Fig. 4F). We also confirmed more circular cells per image in the group treated with Batimastat (p = 0.0078) (Fig. 4G). Taken together, MMP7/9 are upregulated by ALK7/β-catenin pathways, and PDAC cells treated with Batimastat exhibited less invasive phenotypes.
MMP inhibition minimally affects PDAC extravasation
While these data suggest that ALK7/β-catenin-mediated MMP7 and MMP9 expression promotes the formation of invadosome-like protrusions by PDAC cells in 3D hydrogels, it remains unclear how these cells interact with blood vessels. To examine PDAC interactions with a blood vessel in 3D, in particular, tumor extravasation, we employed microfluidic blood vessel (BV)-on-chip devices, which have been developed and published in our lab [28, 29]. To form a healthy engineered blood vessel in the device, we seeded human umbilical vein endothelial cells (HUVECs) as a monoculture (without PDAC) in a hollow channel fully embedded in Collagen I (Col I, 2.5 mg/mL) (Fig. 5A). During the three days of culture on a rocking platform introducing shear stress, HUVECs in the device form a perfusable, engineered BV and deposit their own basement membrane (BM) proteins, collagen IV (Col IV) [30], which is distinct from Col I initially used in our bulk ECM (Fig. 5A). After 3 days of culture, we fixed the cells and stained them with anti-CD31 and anti-Col IV antibodies to identify CD31 + engineered BV and Col IV + vascular BM. We observed an engineered BV and basement membrane (BM) deposited in the basal side of the BV at Day 3 (Fig. 5B). To examine PDAC extravasation, we introduced PDAC cells into the BV lumens through the media reservoirs on Day 3, when the engineered BV was fully formed and the BMs were deposited. After 1 day of PDAC introduction, we fixed the device and observed the tumor-vessel interactions (Fig. 5C). In the experiment, we tested Batimastat to understand whether MMP blockade prevents PDAC extravasation. Batimastat (20 µM) or DMSO (a vehicle) was treated at Day 3 when PDAC cells were introduced to the BV lumen. After 24 h of co-culture and drug treatment, we fixed the cells and visualized the BM, endothelial cells, and PDAC cells, using anti-Collagen IV, anti-CD31, and anti-GFP antibodies (Fig. 5D). We observed extravasation in both groups, showing loss of collagen IV + BM and CD31 + HUVECs where tumor cells invade the vessel lumen (Fig. 5D). We quantified the percent PDAC invasion (Fig. 5E), extravasated PDAC cells (Fig. 5F), and remaining CD31 + BV (Fig. 5G) and Col IV + BM areas (Fig. 5H); surprisingly, we showed minimal effects of Batimastat in the prevention of PDAC extravasation and breakdown of the BVs and their BMs. Taken together, MMP inhibition minimally affects PDAC extravasation.
Fig. 5.
MMP inhibition minimally affects PDAC extravasation. (A) A schematic of the bioengineered blood vessel (BV)-on-chip. (B) HUVECs in monoculture stained with anti-CD31 (red), anti-Collagen IV (yellow) antibodies, and DAPI (blue). Collagen IV, a vascular basement membrane (BM) protein, is secreted by HUVECs and deposited in the basal side of the engineered BV (side view, left: apical side, right: basal side). (C) A schematic of the extravasation-on-chip workflow. After 3 days of HUVEC seeding, HUVECs form an engineered BV, fully depositing BM. Then, PDAC cells were introduced into the BV lumens through the media reservoirs. Batimastat (20 µM) or DMSO (a vehicle) was treated on Day 3 when PDAC cells were introduced. (D) Representative immunofluorescent (IF) images of PDAC extravasation, staining for PDAC (green, GFP+), Collagen IV (yellow), CD31 (red), and merged images. (E) Percent PDAC invasion in the BV (n = 6). (F) Extravasated PDAC cells (n = 6). (G) Percent CD31 + area out of the total area of the BVs (n = 6). (H) Percent Col IV + area out of the total area of the BVs (N = 6). Scale bars (B) = 50 μm; (D) = 250 μm; ‘ns’ indicates not significant
MMP inhibition effectively blocks PDAC intravasation by preserving basement membrane integrity
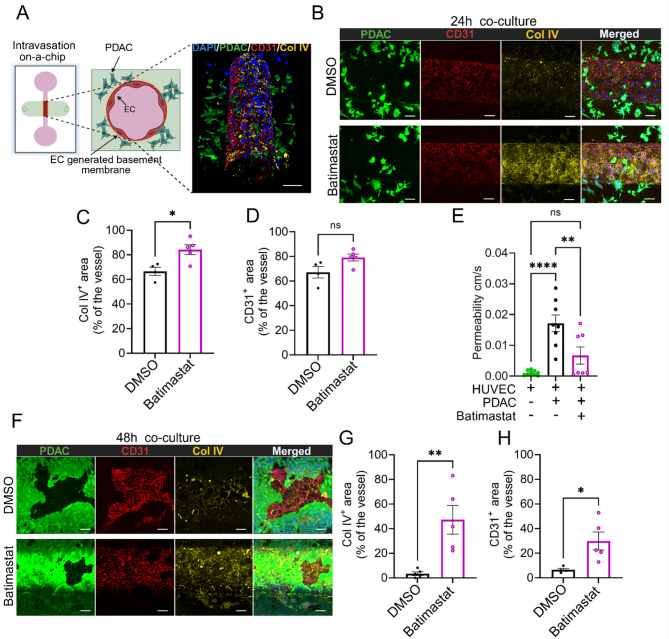
PDAC metastasis is highly dependent on intravasation, as tumor cells must first overcome the fibrotic stroma and breach the vascular barrier to enter circulation [5, 6]. During intravasation, cancer cells, to cross the endothelium, must breach the vascular basement membrane (BM) [31]. We hypothesized that PDAC cells utilize MMPs to breach BM and intravasate into BVs and MMP inhibition blocks the intravasation process. To test the hypothesis, we utilized the BV-on-chip by including PDAC cells in the Col I ECM bulk (1.5 × 106 cells/mL) to surround a HUVEC-generated bioengineered BV (Fig. 6A). After co-culturing them with or without Batimastat for 24 h, the devices were fixed and immunostained with anti-Col IV, anti-CD31, and anti-GFP antibodies to visualize the vascular BM, endothelial cells, and PDAC cells, respectively (Fig. 6B). At 24 h, the integrity of BVs was compromised by PDAC invasion in the DMSO-treated control group, as indicated by the loss of the CD31 and Col IV signals when there are PDAC cells in proximity. Here, Batimastat prevented the BM breakdown and loss of endothelium (Fig. 6B). We quantified the percent Col IV + BM areas, showing that BVs co-cultured with PDAC cells in the presence of Batimastat (20 µM) were resistant to BM breakdown, as indicated by the higher Col IV + area under Batimastat treatment (p = 0.014) (Fig. 6C). However, we did not observe a significant difference in CD31 + endothelial areas between the two groups; however, the Batimastat-treated group showed a trend in more preserved CD31 + endothelial areas (p = 0.056) (Fig. 6D). We next examined vascular permeability (Supplementary Fig. S12). Endothelial barrier permeability is governed by intercellular junctions and other components such as the basement membrane, glycocalyx, and supporting pericytes [32]. To investigate vascular barrier function, we assessed the vascular permeability of BVs in our microphysiological model as we had previously performed [29]. Briefly, we introduced fluorescently labeled 70 kDa dextran into the vessel lumen of intact and PDAC-invaded BVs treated with or without Batimastat (20 µM) and compared their barrier function by measuring the diffusion coefficient of the dextran out of the vessel into the interstitium. Intact BVs without PDAC served as robust vascular barriers and kept dextran molecules within the vessel lumen. Conversely, dextran readily leaked out of PDAC-invaded BVs; here, Batimastat rescued the barrier function of PDAC-invaded BVs (Fig. 6E, Supplementary Fig. S12B). We quantified PDAC cells surrounding the vessels to isolate the Batimastat impact from PDAC cell numbers; overall, Batimastat treatment did not impact PDAC cell numbers (Supplementary Fig. S12C). We next extended the experiment to 48 h. Noteworthily, PDAC invasion of the bioengineered vessel was dramatically delayed by Batimastat, compared to DMSO controls (Fig. 6F). Col IV + BM remained protected from PDAC invasion when treated with Batimastat (20 µM) (p = 0.0059) (Fig. 6G). CD31 + endothelial areas were also preserved by Batimastat treatment (p = 0.0263) (Fig. 6H). There was no significant difference found in PDAC cells surrounding the vessels in Batimastat- vs. DMSO-treated groups, indicating that observed differences in BM degradation result from MMP inhibition, not by different PDAC cell numbers around the vessels (Supplementary Fig. S12D).
Fig. 6.
MMP inhibition effectively limits PDAC intravasation by preserving BM integrity. (A) A schematic of the PDAC intravasation-on-chip. PDAC cells were seeded in the Col I bulk at 1.5 × 106 cells/mL; immediately after, HUVECs were seeded into microchannels. Devices were treated with Batimastat or DMSO for 1 or 2 days. (B) Representative images for PDAC (green), CD31 (red), and Col IV (yellow) at 24 h. (C) Percent Col IV + area out of the total area of the BVs with or without Batimastat at 24 h (n = 4–5). (D) Percent CD31 + area out of the total area of the BVs with or without Batimastat at 24 h (n = 4–5). (E) Vessel permeability in normal and PDAC conditions with or without Batimastat at 24 h (n = 7–9). (F) Representative images for PDAC (green), CD31 (red), and Collagen IV (yellow) at 48 h. (G) Percent Col IV + area out of the total area of the BVs with or without Batimastat at 48 h (n = 5). (H) Percent CD31 + area out of the total area of the BVs with or without Batimastat at 48 h (n = 5). Scale bars (A) = 75 μm, (B, F) = 100 μm; * (p < 0.05), ** (p < 0.01), **** (p < 0.0001) indicate statistical significance. ‘ns’ indicates not significant
Genetic inhibition of ALK7 preserves basement membrane integrity and endothelial barrier, limiting PDAC intravasation
While the results showed that vascular BM integrity was preserved by Batimastat during PDAC intravasation, Batimastat is a broad-spectrum MMP inhibitor, and it is still unclear whether ALK7 inhibition could show the similar effects. To test the roles of the ALK7 receptor in PDAC intravasation, we co-cultured control (shControl) or ALK7 knockdown (shALK7KD) PDAC cells with engineered blood vessels. Control and ALK7 KD PDAC cells were introduced in the Col I bulk at a 0.7 × 106 cells/mL concentration. Shortly after ECM polymerized, we seeded HUVECs into the hollow channel. We fixed the devices at 24 h and 48 h and immunostained the cells with anti-GFP (PDAC), anti-CD31 (HUVEC), anti-Col IV (BM) antibodies, and DAPI (Fig. 7A and F). At 24 h, HUVECs co-cultured with control PDAC cells (Fig. 7A) lost their endothelial barrier, and were devoid of BM; here, HUVECs co-cultured with ALK7 KD PDAC cells relatively better preserved blood vessels, as shown in the percentage of CD31 + vessel area (not significant) (Fig. 7B) and Col IV + BM area (p = 0.024) (Fig. 7C). ALK7 inhibition did not affect PDAC cell numbers at 24 h, quantified as PDAC cell area (Fig. 7D). Our comparison of the Col IV + and CD31 + area in each vessel co-cultured with control PDAC cells demonstrated a higher CD31 + over the Col IV + area (p = 0.007) (Fig. 7E, left). By contrast, HUVECs co-cultured with ALK7 KD PDAC cells showed preserved the ratio of Col IV + over CD31 + signal (p = 0.4) (Fig. 7E, right), indicating that because cancer cells first came in contact with the BM and later with endothelial cells, the ALK7 KD condition better preserve the BM + area. We quantified the protrusion length and number in control and ALK7 KD PDAC cells. In support BM integrity in vessels co-cultured with ALK7 KD cells, we found that these cells exhibited fewer protrusions (p = 0.045), although the length of protrusions was not significantly different (Fig. 7F). These findings suggest a link between reduced protrusion formation and decreased matrix metalloproteinase (MMP) activity. At 48 h, ALK7 KD PDAC invasion of the bioengineered vessel was even more dramatically delayed (Fig. 7G). Both the CD31 + endothelial area (p = 0.037) and BM layer remained protected from PDAC cell invasion (p = 0.0086) (Fig. 7H and I). 7 Control and ALK7 KD PDAC cell numbers were similar throughout the experiment, quantified as PDAC area (Fig. 7J). We noticed a similar trend to the 24 h timepoint, with excessive breakdown of BV and BM in control PDAC, compared to the ALK7 KD PDAC group (Fig. 7K). We sought to determine whether SB431542 treatment has a comparable effect on blood vessel integrity in a PDAC extravasation and intravasation. Supporting our data regarding the role of the ALK7 receptor in extravasation and intravasation, SB431542 did not maintain barrier integrity in the extravasation assay (Supplementary Fig. S13A). However, the SB431542 inhibitor maintained the vascular barrier function of HUVECs against tumor intravasation (p < 0.0001; Supplementary Fig. S13B). To determine the effect of ALK7 inhibition on tumor vascular barrier function in vivo, we inoculated PD7591 cells into mice. Seven days after the tumor inoculation, we administered either SB431542 (25 mg/kg/day, i.p. injection) or DMSO daily. After 2 weeks of drug/vehicle treatment, we intravenously injected 70 kDa dextran tracer and assessed the permeability of tumor blood vessels. Control mice treated with DMSO were not able to carry the 70 kDa dextran within tumor blood vessels. Instead, the tracers readily diffused out to the tumor interstitium (Supplementary Fig. S14A). Conversely, the interstitial leakage of 70 kDa dextran was well restricted within tumor vessels harvested from SB431542-treated mice (Supplementary Fig. S14B). Taken together, our data showed that ALK7 inhibition in PDAC tumor intravasation normalized the barrier function of tumor blood vessels and prevented vascular leakage in vitro and in vivo. Overall, our data demonstrate that ALK7 inhibition potently preserved the BM integrity and endothelial barrier, blocking the intravasation process, suggesting that ALK7 signaling is directly linked to MMP secretion, BM breakdown, and PDAC intravasation (Fig. 7L).
Fig. 7.
Genetic ALK7 inhibition preserves basement membrane (BM) integrity and endothelial barrier, limiting PDAC intravasation. (A) Representative images for PDAC (green), CD31 (red), and Col IV (yellow) in the intravasation-on-chip at 24 h. Control vs. ALK7 KD PDAC cells were seeded in Col I bulk at 0.7 × 106 cells/mL. (B) Percent CD31 + area out of the total area of the BVs with PDAC control and ALK7 KD cells at 24 h (n = 5). (C) Percent Col IV + area out of the total area with PDAC control and ALK7 KD cells at 24 h (n = 5). (D) Quantification of PDAC cells on the image at 24 h of co-culture. (E) Correlation between Col IV and CD31 expression at 24 h. (F) Quantification of protrusion numbers and length of control and ALK7 KD PDAC cells. (G) Representative images for PDAC (green), CD31 (red), and Col IV (yellow) at 48 h. (H) Percent CD31 + area out of the total area of the BVs with PDAC control and ALK7 KD cells at 48 h (n = 5). (I) Percent Col IV + area out of the total area with PDAC control and ALK7 KD cells at 48 h (n = 5). (J) Correlation between Col IV and CD31 expression at 48 h (K) Quantification of PDAC cells on the image at 48 h of co-culture (n = 5). (L) Summary of our findings on the mechanisms of ALK7 in PDAC metastasis. PDAC cells that express ALK7 receptor invade into ECM, break BM, and intravasate into blood vessels via GSK3β/β-catenin signaling that mediates MMP secretion and EMT transition. Scale bars (A, F) = 50 μm; * (p < 0.05), ** (p < 0.01), indicate statistical significance. ‘ns’ indicates not significant
Discussion
In this study, we report non-canonical ALK7 pathways that promote PDAC metastasis through β-catenin/MMP-mediated basement membrane breakdown and intravasation. Through 3D in vitro tumor-on-chip models and in vivo orthotopic PDAC mouse models of spontaneous metastasis, ALK7 was shown to enhance cancer cell invasiveness via non-canonical β-catenin signaling, promoting EMT. Simultaneously, ALK7 upregulates MMP expression, facilitating basement membrane degradation and weakening vascular integrity, thereby promoting PDAC intravasation. These findings highlight ALK7 as a dual modulator of metastasis, amplifying both intrinsic tumor cell mobility and extrinsic microenvironmental remodeling to drive invasive progression.
The roles of ALK7 in cancer progression have been unclear. Some studies have shown that ALK7 could be anti-tumorigenic and anti-metastatic based on ALK7-mediated cancer cell death [9, 10, 33]. However, some other studies suggested that ALK7 might be pro-tumorigenic and pro-metastatic by showing that ALK7 promoted tumor invasion, proliferation, and relapse [11–14]. Metastatic diseases are multi-factorial, involving many steps of the metastatic cascade, including primary tumor growth, tumor migration, vascular invasion, intravasation, CTCs, extravasation, tumor dormancy, and colonization. All these have different biological and biophysical processes with many different signaling pathways, and often, the same molecule results in different outcomes depending on what biological contexts the molecule is tested and observed. Widely used experimental metastasis models, such as tumor cell injection into the bloodstream, skip vascular invasion, and intravasation, could miss the essential role of ALK7 in tumor intravasation. To address this question, we elucidated PDAC interactions with BVs, especially PDAC intravasation and extravasation, using an in vivo orthotopic PDAC mouse model of spontaneous metastasis and 3D in vitro tumor-on-chip models.
Our in vivo orthotopic spontaneous metastasis experiments demonstrated that both pharmacological and genetic inhibition of ALK7 significantly reduced the dissemination of PDAC cells to distant organs, such as the lungs. Mechanistically, ALK7 appears to drive PDAC invasiveness through two interrelated processes: (1) enhancing tumor cell motility via β-catenin-mediated EMT activation [34] and (2) remodeling the tumor microenvironment through MMP-driven ECM degradation [35]. Given that mesenchymal-like migration relies on both cytoskeletal remodeling for increased motility and proteolytic ECM degradation for invasive capacity, we investigated whether ALK7 distinctly regulates these two processes. ALK7 knockdown significantly downregulated key EMT transcription factors, including ZEB1, SNAIL, SLUG, and TWIST1, which was reversed by β-catenin overexpression, suggesting that ALK7 promotes EMT via β-catenin signaling to enhance cytoskeletal remodeling and cell motility. In parallel, ALK7 was also found to regulate ECM degradation potentially by facilitating invadosome formation, specialized adhesion structures that enable cancer cells to degrade the surrounding matrix. RNA-seq analysis revealed that ALK7 knockdown led to significant downregulation of ECM receptor interactions and cell adhesion pathways, indicating that ALK7 plays a crucial role in tumor-ECM dynamics. ALK7 inhibition reduced MMP activity which could lead to disrupted protrusive structures, impairing ECM remodeling and weakening the invasive capacity of PDAC cells. These findings suggest that ALK7 simultaneously enhances tumor cell motility through β-catenin-driven EMT and facilitates ECM breakdown via MMP-dependent invadosome activity, collectively driving PDAC invasiveness and metastatic dissemination.
Our observations highlight the role of ALK7 in early metastasis dissemination by affecting intravasation but minimally influencing later step like extravasation. The reason is possibly because prior to intravasation, cancer cells invade into ECM, where they remodel surrounding tissue inducing local angiogenesis. We speculate that PDAC could activate MMPs to first degrade the basement membrane of tumor vasculature and penetrate blood vessels. During extravasation, CTCs encounter intact blood vessels in distant organs, which requires quite different mechanisms to exit the circulation. During the extravasation, CTCs adhere to the endothelium and migrate through the endothelial barrier to reach the underlying tissues [36, 37], which can be achieved even without breaching the basement membrane.
It is also important to note that Batimastat treatment induced morphological changes in PDAC cells, specifically reducing the length and number of protrusions, suggests that MMP activity influences cytoskeletal dynamics. Interestingly, a study by Harada et al., identified ARL4C–IQGAP1–MMP14, which is activated explicitly at invasive pseudopodia of pancreatic cancer cells, upstream of KRAS, where β-catenin inhibition downregulated ARL4C expression [38]. Most importantly, increased matrix stiffness is a common characteristic of solid tumors [39]. Previous studies indicated that increased matrix stiffness may promote malignant cell behaviours [40–42]. While two-dimensional (2D) studies indicated that increased ECM stiffness promotes increased degradation of ECM by invadopodia [43, 44], studies done with 3D cultures rich with basement membrane components restricted invadopodia extension and subsequent migration [45]. Our findings also highlight a potential link between ALK7 and Rho, suggesting that invadosome formation in PDAC could be facilitated by the ALK7/Rho/MMP axis. Additionally, ALK7-driven β-catenin signaling regulates MMP7/9 expression, increasing ECM degradation and tumor invasiveness. These findings suggest that ALK7 inhibition could suppress invadosome formation, thereby reducing PDAC cell intravasation and metastatic dissemination.
In the present study, we demonstrated that ALK7 inhibition slowed down the dissemination of PDAC cells to distant organs, such as lungs. Furthermore, the pro-metastatic role of ALK7 signaling in PDAC was confirmed by clinical data from the TCGA database, which indicated that patients with upregulated ALK7 signaling experience accelerated disease metastasis. Taken together, our report suggests that ALK7 signaling may induce tumor vessel dysfunction and metastasis by promoting PDAC cells to adopt a migratory, invasive phenotype. Our results highlight a role for ALK7 in early metastatic dissemination by affecting, particularly, intravasation. We speculate that the PDAC could activate MMPs to first degrade the ECM and BM of tumor vasculature and penetrate the BVs of vasculature. Although Batimastat is a broad-spectrum MMP inhibitor, its relatively low IC₅₀ values for MMP7 and MMP9 (6.3 nM and 4.4 nM, respectively) suggest that it may preferentially affect these targets under our experimental conditions, thereby reinforcing the biological relevance of focusing on MMP7 and MMP9 in this study. Nonetheless, future studies should investigate whether inhibiting MMPs with more selective and potent inhibitors—rather than earlier broad-spectrum agents—at early stages of pancreatic cancer would improve patient outcomes, and should also include functional validation experiments, such as individual rescue or knockdown of MMP7 and MMP9, to definitively confirm their specific contributions to ALK7-mediated PDAC invasion and metastasis [46]. MMPs are particularly important during cancer cell migration through the stroma, which explains the lack of success in initial trials involving patients with advanced metastatic cancers [47, 48].
PDAC is often diagnosed at an advanced stage due to the absence of specific symptoms and the lack of reliable early biomarkers, which significantly limits the opportunity for curative treatment. Although recent advances in diagnostic approaches such as endoscopic ultrasound, magnetic resonance imaging-based screening, and molecular assays involving circulating tumor DNA and exosome profiling offer promise for earlier detection in high-risk individuals, early intervention remains difficult in most clinical settings. In this context, inhibiting ALK7, a key modulator of tumor cell intravasation, may serve as an effective therapeutic strategy to suppress early metastatic dissemination before widespread disease occurs. Furthermore, even in patients diagnosed at advanced stages who are not eligible for complete surgical resection, therapeutic targeting of ALK7 could help prevent further metastatic spread by limiting tumor cell entry into the circulation. These considerations suggest that ALK7-targeted strategies may hold clinical utility across both early and advanced stages of pancreatic cancer progression.
Due to the lack of specific inhibitors, this study has certain limitations associated with the use of broad-spectrum agents such as Batimastat, which targets multiple MMPs, and SB431542, which inhibits ALK7 but also has known off-target activity against ALK4 and ALK5. While both compounds provided valuable mechanistic insights, their limited target selectivity may confound interpretation and reduce translational relevance. Therefore, future studies should employ more selective inhibitors—both for MMPs and ALK7—to better delineate receptor-specific mechanisms and enhance therapeutic potential. While this study primarily focused on the interaction between PDAC cells and endothelial cells during early metastatic dissemination, further studies are needed to compare how ALK7 modulates the characteristics of primary and metastatic tumors. Future work incorporating single-cell transcriptomic approaches or spatial profiling could provide a more comprehensive understanding of ALK7-dependent tumor–microenvironment interactions across different stages of disease progression.
Conclusions
We demonstrate that ALK7 receptor inhibition enhances the metastatic potential of pancreatic cancer cells by modulating GSK3β/β-catenin signaling, thereby regulating stemness and epithelial-to-mesenchymal transition (EMT). Additionally, ALK7 downregulation alters the expression of matrix metalloproteinases (MMPs), further contributing to metastatic progression. Using an in vivo spontaneous metastasis model, we show that pharmacological inhibition of ALK7 effectively disrupts the pancreatic ductal adenocarcinoma (PDAC) metastatic cascade. Metastasis is a complex, multistep process, making it challenging to study in live animal models. To overcome this limitation, we developed a 3D microfluidic platform that recapitulates pancreatic cancer cell interactions with the vasculature, enabling the real-time investigation of intravasation and extravasation. Our findings reveal that ALK7 blockade significantly impairs intravasation while having minimal impact on extravasation, likely due to the critical role of MMPs in the intravasation process. These results suggest that targeting ALK7 may be most effective in inhibiting early-stage metastatic dissemination by preventing intravasation rather than later-stage extravasation.
Materials and methods
Cell culture
The primary murine pancreatic cancer cell line, YFP PD7591 (a gift from Dr. Ben Stanger, University of Pennsylvania), was isolated from primary tumors of a genetically engineered mouse model (Kras-G12D p53-/- YFP Rosa) and cultured in DMEM supplemented with 10% fetal bovine serum (FBS), 50 µg/mL streptomycin and 50 I.U./mL penicillin. PD7591 was selected for this study because it originates from a model that closely recapitulates key features of human PDAC, including desmoplastic stroma and spontaneous metastasis [49–51]. HUVECs (Cell Systems) were cultured in EGM-2 (Lonza) supplemented with 2% FBS and the Microvascular Endothelial Cell Growth Medium-2 BulletKit (CC-4176, Lonza). EGM-2 was used in all conditions. All cells in this study were incubated in a standard tissue culture incubator at 37 °C, 95% humidity, and 5% CO2. HUVECs were used within 8 passages. PDAC cells were tested negative for Mycoplasma.
Spheroid preparation and drug treatment
To generate multicellular tumor spheroids, PD7591 cells (9 × 10³ cells/well) and LX-2 cells (1 × 10³ cells/well) were co-seeded in 96-well round-bottom ultra-low attachment microplates. The plates were incubated at 37 °C in a humidified incubator with 5% CO₂ for 3 days to allow spheroid formation. After spheroids were formed, cells were treated with 40 µM SB431542 and incubated for an additional 24 h under the same conditions. Following treatment, spheroids were collected and processed for downstream analyses.
Generation of a luciferase-expressing PDAC cell line
We generated our luciferase-expressing PDAC cell line by transducing PD7591 cells with a premade, firefly luciferase-expressing (CMV, puromycin) lentivirus (Kerafest, FCT005) according to the manufacturer’s transduction protocol. In brief, the lentivirus was added to a 6 µg/mL polybrene (Krackeler Scientific) containing PD7591 cells grown in cell growth medium at around 50-60% confluency overnight. The virus-containing medium was then removed and replaced with a puromycin (2 µg/mL) containing medium to select for successfully transduced PDAC cells. The multiplicity of infection (MOI) was 6.
Lentivirus transduction and generation of stable cell lines
PD7591 cells were seeded at a concentration of 3 × 105 cells in 6-well dishes and cultured in complete media. For transduction, shALK7KD (VB221019-1160hcn and VB221019-1161js) and scramble shRNA control (VB010000-0004axz) lentiviral particles were used according to the manufacturer’s instructions (Vectorbuilder, Chicago, USA). Briefly, PD7591 cells were incubated at 37 °C in a humidified 5% CO2 incubator until 30–50% confluency was reached. Cultured cells were infected with 10 MOI (multiplicity of infection) of lentiviral particles (> 108 TU/mL). After 48 h of infection, the cells were treated with G418 for two weeks to select stable KD of cells and examined by Western blotting.
Generation of ALK7 knockout (KO) PDAC cell lines
Large chromosomal deletions in the ALK7 gene were generated using the CRISPR-del strategy with recombinant Cas9 protein and two synthetic guide RNAs (gRNAs). Cas9/gRNA ribonucleoprotein (RNP) complexes were assembled by pre-incubating Cas9 protein with two gRNAs (1 µg each; gRNA#1: 5′-GCCCGTTGGTTAGCATGACGG-3′, gRNA#2: 5′-CCGTCATGCTAACCAACGGG-3′) in resuspension buffer R (Thermo Fisher Scientific). The complexes were delivered into PD7591 cells (2 × 10⁵ cells/µl) via electroporation using the Neon Transfection System (10 µl tip, 1150 V, 40 ms, 2 pulses) according to the manufacturer’s protocol. Following electroporation, cells were plated into 24-well plates for recovery, then serially diluted and seeded into 96-well plates at a density of 1 cell/well. Wells containing single-cell clones were identified by microscopic inspection and expanded in 12-well plates. Genomic DNA was extracted, and PCR-based genotyping was performed using primers flanking the targeted deletion region (forward: 5′-AGCACCACACTATAACTCCC-3′; reverse: 5′-ATGCTATCATCTTCCAAAGC-3′). PCR amplicons were subjected to Sanger sequencing to confirm deletion.
Animal study
To prepare an orthotopic mouse model of PDAC, 5 × 105 live PD7591 cells were suspended in 10 µL of sterile PBS and administered by intrapancreatic injection to 5-6-week-old NSG (NOD-scid IL2Rgammanull) mice. The cells were delivered around 2–3 mm from the tail of the pancreas using a 30-gauge insulin needle, as described previously [52, 53]. Briefly, small skin incisions were made over the abdominal wall near the spleen. The underlying muscle was opened, and the pancreas was gently exteriorized using a cotton bud. Injections were performed with the pancreatic tail supported on the cotton bud. The syringe was held in place for 30 s post-injection to minimize cell leakage. The abdominal cavity was closed with absorbable sutures, and the skin was closed with wound clips. Ketoprofen was administered for 2 days post-op at 5 mg/kg concentration. These animal models were treated with either SB431542 (25 mg/kg/day, i.p. injection) or DMSO (i.p. injection) 1 week after tumor inoculation for 2 weeks. In the study with ALK7 KD, we injected either shALK7KD cells or shControl PDAC cells into the tail of the pancreas. The animals were monitored for 3 weeks before organ collection. In the study with ALK7 KD, we injected either shALK7KD cells or shControl PDAC cells into the tail of the pancreas. The animals were monitored for 3 weeks before organ collection.
In vivo imaging of tumor metastasis
PD7591 cells were tagged with luciferase before being injected into our mouse models using lentiviral transduction. To image primary tumors in the PDAC model, 150 mg/kg of luciferin in sterile PBS was introduced to mice via i.p. injection. The injected luciferin reacts with tumor-expressing luciferase, giving off a bioluminescence signal that can be detected by an IVIS Xenogen imager, quantified as photons per second using the manufacturer’s imaging software.
For primary tumor imaging, mice were imaged for 10 min to the peak imaging time. To assess lung and metastasis, mice were sacrificed, and lungs and livers were collected. Tissues were incubated in a 150 µg/mL working solution of D-Luciferin in PBS on ice for 4 min prior to imaging. Each tissue was imaged separately with IVIS imager (Field of View: B; exposure set to auto, with three segments captured every minute). The bioluminescence signal was quantified using Free Living Imaging Software. Following the manufacturer’s recommendations, we set the detection threshold at 600 counts, with signals below this value considered background noise. Regions of interest (ROIs) were placed on signals exceeding 600 counts. Subsequently, the counts parameter was converted to radiance (photons/sec) in each pixel integrated over the ROI area (cm2) x 4π to obtain total flux.
Circulating tumor cell (CTC) enumeration
CTCs were quantified in mice with orthotopically injected PDAC ALK7 KD and PDAC shControl. Blood (350 to 550 µL) was collected via cardiac puncture using a 23-gauge needle attached to a 1-mL syringe. The blood was collected directly into tubes containing EDTA, and ACK lysis was performed to remove red blood cells. The cells were washed and seeded into a 24-well plate. 24 h later, PDAC cells were imaged with an IVIS imager. The blood was collected directly into tubes containing EDTA, and ACK lysis was performed to remove red blood cells. Since the PDAC cells expressed luciferase, circulating tumor cells (CTCs) were imaged using an IVIS imager (Field of View: C; exposure set to auto, with three segments captured every 1 min). Cells were incubated for 5 min in a 150 µg/ml working solution of D-luciferin prepared in pre-warmed tissue culture medium. The detection threshold was set at 600 counts, with signals below this value considered background noise. Regions of interest (ROIs) were placed on signals exceeding 600 counts. As a negative control, blood collected from naïve, non–tumor-bearing mice was analyzed (data not shown). Due to difficulties with collecting the same amount of blood from each animal, bioluminescence signals from each sample were normalized by the amount of blood collected.
In vivo tumor vascular permeability assay
Our PDAC mouse models were prepared and treated with SB431542 (25 mg kg− 1 day− 1, i.p. injection) or a DMSO vehicle (i.p. injection) as described below. Each group of mice was anesthetized using isoflurane and retro-orbitally injected with the 100 µl mixture of fixable Cascade blue-conjugated fixable biotin-conjugated 70 kDa dextran (5 mg mL− 1; Thermo Fisher) dissolved in sterile saline. 30 min after the dextran injection, the mice were quickly euthanized via cervical dislocation, and their tumors were resected and fixed in 4% PFA at room temperature for 18 h. Tumor samples were then incubated in 30% sucrose (in PBS) for 1 day and embedded in the OTC compound (Sakura, Japan) before cryosectioning. Sectioned tumor samples were kept in the dark at -20 °C. We then performed immunofluorescence microscopy and stained the tumor sections with FITC-conjugated anti-GFP antibody (Abcam, 1:500) to visualize GFP-positive tumor cells as well as Alexa 647 conjugated Streptavidin (Life Technologies) to visualize biotin-conjugated 70 kDa dextran. To quantify the vascular density in each tumor sample, rat anti-mouse CD31 (BD Biosciences) and Alexa 647 rabbit anti-rat antibodies (Jackson ImmunoResearch) were used to identify CD31-positive vasculature. Images were acquired with a Leica HC FLUOTAR L 40X/0.95.
Fabrication of microfluidic devices
Our single-channel casting mold was fabricated using the microfabrication protocol described in our previous study [54], and microfluidic chip devices were then fabricated using soft lithography. First, polydimethylsiloxane (PDMS; Sylgard 184, Dow-Corning, DC2065622) was mixed with a curing reagent in a 10:1 weight ratio (base: curing agent), and the PDMS mixture was put in molds and cured overnight at 80 °C. The PDMS gaskets were then removed from the molds and bonded to a cover glass using plasma surface activation. The bonded chips were further cured at 80 °C for permanent bonding. The plasma etched chips were treated with 0.1 mg/mL poly-L-lysine (Sigma) for 1 h and 1% glutaraldehyde (Electron Microscopy Sciences) for 30 min, then rinsed with sterile water 3 times and immersed in a sterile water bath overnight. Steel acupuncture needles (0.25 × 50 mm, Hwato) were sterilized with 70% ethanol overnight. The next day, the needles were rinsed with sterile PBS, treated with BSA (1 mg/mL in PBS), and inserted into the microchannel in each device. After the devices were dried using an air gun, they were sterilized under UV light for 30 min. To cast the ECM bulk around each microchannel, 2.5 mg/mL of rat tail collagen I (Corning) dissolved in PBS at a pH of 8.0 was pipetted into each device for 1 h. The cell growth culture medium was added to the media reservoirs overnight, and the two acupuncture needles were carefully retrieved the next day, leaving one hollow cylindrical channel suspended in the interstitial collagen matrix. Devices were then placed on a platform rocker (Benchmark Scientific) overnight.
Invadosome-like protrusion formation assay
To measure PDAC plasticity in a 3D matrix, we encapsulated PDAC cells at cell density 0.5 × 106 cells/mL in 2.5 mg/mL collagen gel on a chip. 2.5 mg/mL collagen gel was prepared as described in the fabrication of the microfluidic device method section. PDAC cells were treated with 20 µM Batimastat or DMSO for 24 h. After the 24 h culture period, cells were fixed and stained for immunofluorescence microscopy. We manually picked single cells in ImageJ to analyze cell shape. PDAC cells within clusters were not quantified due to being unable to differentiate their shape, protrusion number, and length.
Histopathology hematoxylin and eosin staining
We performed Hematoxylin and eosin staining (H&E) on lungs, liver spleen and kidney of animals that are treated with vehicle (DMSO) or SB431542 for 14 days to assess inhibitor toxicity in the nonbearing tumor animals. Organs were collected and fixed for 24 h at 4 °C, then placed in 15% and subsequently 30% sucrose solutions until fully infiltrated. Tissues were embedded in Tissue-Tek OCT compound (Fisher), and 10 μm sections were cut at − 20 °C. Tissues were rehydrated in PBS and incubated for 10 with Harris’ Alum Hematoxylin (Harleco 638 A-71). The samples were washed with water and dipped in 70% ethanol for one minute. They were then stained with Eosin Y (1% alcoholic solution, Harleco 588X-75) for 2 min. After staining, the samples were dehydrated through two changes of 95% ethanol, followed by two changes of 100% ethanol (1 min each), and finally cleared with xylene for one minute.
Extravasation on-a-chip
HUVECs were suspended at 2 × 106 cells/mL in complete EBM-2 and seeded into the channel of the microfluidic device. HUVECs attached to the channel surface and formed a bioengineered vessel. 72 h later, single-cell PDAC suspension (0.2 × 106 cells/mL) was introduced into the media reservoir of the bioengineered vessel. PDAC cell suspension in EGM2 media contained either 20 µM Batimastat or DMSO. After seeding, PDAC cells started to attach to the luminal surface of the vessel. After 24 h of co-culture, cells were fixed and prepared for immunofluorescence (IF) microscopy. For SB431542 (5 µM) inhibitor studies, single-cell PDAC suspension (0.5 × 106 cells/mL) was introduced into the media reservoir of the vessel, co-cultured for 48 h and subsequently permeability assay was performed.
Intravasation on-a-chip
PDAC cells were seeded into 2.5 mg/mL collagen gels at 1.5 × 106 or 0.7 × 106 density. Immediately after, HUVECs were suspended at 2 × 106 cells/mL in EBM-2 and seeded into the channel of microfluidic devices. Batimastat at 20 µM concentration or DMSO vehicle control in EMB-2 was added to the media reservoir of the microfluidic devices. Cells were cultured for 24–48 h, subsequently fixed, and prepared for IF microscopy. For SB431542 (5 µM) inhibitor studies, PDAC cells were seeded at 0.5 × 106 density. Cells were co-cultured for 48 h, subsequently permeability assay was performed.
Immunofluorescent (IF) microscopy
Devices were fixed for IF staining and imaging with 4% paraformaldehyde (Electron Microscopy Sciences, 15710) in PBS for 1 h at room temperature. Fixed devices were permeated with PBST (0.3% Triton-X in PBS) for 45 min at room temperature, then blocked with 3% BSA in PBS overnight at 4 °C. They were then incubated with anti-VE-cadherin (Abcam, ab33168, 1:100), anti-CD31 (Dako, M0823, 1:200), and fluorescein isothiocyanate (FITC)-conjugated anti-GFP (Abcam, ab6662, 1:100) primary antibodies in blocking buffer overnight at 4 °C. The FITC-conjugated anti-GFP primary antibody was used to stain YFP cancer cells. After incubation with primary antibodies, the samples were washed overnight in PBS at 4 °C. Secondary antibodies (Life Technologies, 1:500) and DAPI (Sigma, 1:500) were added to the devices and incubated in a blocking buffer overnight at 4 °C in the dark. Before being imaged using Leica SP8 confocal microscopy (Leica, Germany), the secondary antibodies and DAPI were washed for 24 h in PBS at 4 °C in the dark to remove any unbound fluorescent background. Confocal images were acquired with the Leica SP8 confocal microscope with a 40× objective. Obtained fluorescent images were Z-stacked, 3D rendered, and adjusted for brightness and contrast using the ImageJ [55].
In vitro diffusive permeability quantification
To quantify the diffusive permeability of our engineered BVs, we assumed that the fluorescence intensity measured from a tissue sample is proportional to the number of tracer molecules in that sample. This assumption allowed us to quantify the diffusive permeability coefficient, Pd [56, 57].
 |
Where  is the mass flux of the solution,
is the mass flux of the solution,  is the concentration of the fluorescent tracer (dextran) in the vessel, and
is the concentration of the fluorescent tracer (dextran) in the vessel, and  is the concentration of the fluorescent tracer in the perivascular ECM. By tracking changes in the fluorescence intensity of the ECM over time, we are able to determine the mass flux and thus calculate the diffusive permeability coefficient for each vessel. We adopted the automated MATLAB code developed by Polacheck et al. to quantify diffusive permeability from time-lapse images [29]. For the permeability experiment, PDAC cells were seeded at 0.7 × 106 cells/mL in 2.5 mg/mL Collagen I. HUVECs were seeded at 1.5 × 106 cells/mL in microchannels. 24 h later, permeability was measured by adding 150 µL 70 kDa Texas Red Dextran (Thermo Fisher Scientific, D1864) into the media reservoirs of devices. Dextran concentration in media 25 µg/mL.
is the concentration of the fluorescent tracer in the perivascular ECM. By tracking changes in the fluorescence intensity of the ECM over time, we are able to determine the mass flux and thus calculate the diffusive permeability coefficient for each vessel. We adopted the automated MATLAB code developed by Polacheck et al. to quantify diffusive permeability from time-lapse images [29]. For the permeability experiment, PDAC cells were seeded at 0.7 × 106 cells/mL in 2.5 mg/mL Collagen I. HUVECs were seeded at 1.5 × 106 cells/mL in microchannels. 24 h later, permeability was measured by adding 150 µL 70 kDa Texas Red Dextran (Thermo Fisher Scientific, D1864) into the media reservoirs of devices. Dextran concentration in media 25 µg/mL.
Proliferation and colony formation assay
To perform the proliferation assay, 0.5 × 104 cells were seeded in each well of a 96-well dish (Corning). After 24 h, 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl-2 H-tetrazolium bromide (MTT) (5 mg/mL) was added to the plates and incubated at 37 °C for 4 h. After the formazan crystals were dissolved with DMSO, the OD at 570 nm was measured using a microplate reader. To perform the colony formation assay, shControl- and shALK7KD PD7591 cells (1 × 103 cells per well) were seeded into a 12-well dish. Five days later, cells were stained with 0.5% crystal violet (Sigma-Aldrich, St. Louis, MO, USA). Colonies were counted using the ImageJ software.
Wound-healing and invasion assays
For wound-healing assays, 0.5 × 106 cells were seeded in 60-mm dishes and cultured until confluent. Using a pipette tip, a straight scratch (wound) was generated, keeping the pipette tip at an angle of approximately 30°. Wound closure was then monitored over 48 h. For invasion assays, the QCM 24-Well Cell Invasion assay kit (Millipore, Burlington, MA, USA) was used according to the manufacturer’s guidelines. Briefly, the interiors were rehydrated with pre-warmed, serum-free media, and 2 × 105 cells were added to each insert. Complete EGM-2 (10% FBS) was added to the lower chambers, and cells were incubated for 24 h in a CO2 incubator. Cells were stained with a 0.5% crystal violet solution, and images were obtained using a 4× objective lens.
Western blotting
Whole-cell lysates were prepared using a passive lysis buffer (Promega, Madison, WI, USA) with a protease inhibitor cocktail (Roche, Basel, Switzerland). The proteins were electrophoresed and transferred to membranes. The membranes were incubated overnight at 4 °C with primary antibodies against phospho-Smad2/3, Smad2/3, RhoA, phospho-MLC2, MLC2, phospho-AKT, AKT, phospho-ERK, ERK, phospho-JNK, JNK, phospho-p38, p38, phospho-GSK3β, GSK3β, SNAIL, SLUG, TWIST1, c-Myc, CyclinD1, CD44, SOX2, OCT4, N-cadherin, Fibronectin, MMP7, MMP9 (all from Cell Signaling Technology, Beverly, MA, USA), β-catenin (BD Biosciences, San Jose, CA, USA), glyceraldehyde 3-phosphate dehydrogenase (GAPDH) (Trevigen, Gaithersburg, MD, USA), ALK7, ZEB1, α-tubulin, Lamin B1 (all from Santa Cruz Biotechnology, Santa Cruz, CA, USA). The membranes were then washed and incubated with the appropriate HRP-conjugated secondary antibodies (GeneTex, Irvine, CA, USA, Cat#GTX213111-01, GTX213110-01) for 1 h at room temperature before enhanced chemiluminescence substrate incubation (Santa Cruz Biotechnology, Cat#sc-2048). Western blot images were taken using a LAS 4000 mini camera (Fujifilm, Tokyo, Japan).
β-catenin expression construct
For induced expression of β-catenin, a β-catenin pcDNA6 expression vector was purchased from Addgene (#Plasmid 123586, Watertown, MA, USA). ALK7 KD PD7591 cells were seeded in 6-well dishes, and then cells were transfected with the β-catenin expression plasmid using Lipofectamine 3000 (Thermo Fisher, San Jose, CA, USA). After 72 h, the cells were harvested, and western blotting assays were performed.
RNA sequencing and analyses
Total RNA was extracted from cultured cells using TRIzol Reagent (Invitrogen Life Technologies, Carlsbad, CA, USA) according to the manufacturer’s protocol. The quality of total RNA was assessed using an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA), and samples with an RNA integrity number (RIN) > 8 were selected for RNA-seq analysis. RNA-seq libraries were prepared using a TruSeq Standard mRNA Sample Prep Kit (Illumina) according to the manufacturer’s protocol. Paired-end sequencing with 100 cycles was performed using a NovaSeq 6000 platform (Illumina) instrument according to the manufacturer’s protocol. The quality of raw reads was assessed with FastQC (version 0.11.9), and the quality scores were > Q30, indicating high quality. Clean reads with high-quality scores were processed using the Tuxedo protocol [58] with TopHat2 (version 2.1.1) [59] and Cufflinks [60]. Reads for each sample were aligned to the mouse reference genome using TopHat2. Gene expression quantification was performed using Cufflinks, and transcript per million reads (TPM) was calculated as the expression value. Differentially expressed genes (DEGs) were selected as those with P < 0.001 and fold change > 1.5. The DAVID software (Database for Annotation, Visualization, and Integrated Discovery, v6.7; http://david.abcc.ncifcrf.gov) and and the BINGO 2.3 plugin for Cytoscape software (http://www.psb.ugent.be/cbd/papers/BiNG.
O/Home.html) were used for pathway and gene ontology analysis. RNA-seq data has been deposited in the NCBI Sequence Read Archive (SRA) with the accession number GSE290145.
Quantitative RT-PCR
Total RNA was extracted using the Illustra RNAspin Mini kit (GE Healthcare, Chicago, IL, USA) according to the manufacturer’s instructions. Reverse transcription was performed with 2 µg total RNA using M-MLV Reverse Transcriptase (Life Technologies). qRT-PCR was performed using SYBR Premix Ex Taq II (Takara Biotechnology, Dalian, China) and ABI PRISM 7500 Sequence Detector (Applied Biosystems, Carlsbad, CA, USA). The amount of target mRNA was normalized to that of GAPDH mRNA. The primer sequences used are listed in Table S1 in Supporting information.
TCGA survival analysis
Clinical and gene expression datasets from RNA-sequencing experiments were acquired from the Broad Institute’s Firehose Broad GDAC database and cbioportal.org. We defined the overexpression of three genes (ACVR1C, INHBA, and INHBB), each of which had a transcript abundance cutoff of RSEM greater than two standard deviations from the mean in each cancer type cohort. Inhibin-βA and Inhibin-βB, respectively encoding activin A and -B, serve as ligands for ALK7. We included them all to better analyze functional aspects of the ALK7 with its ligand expression. The overall survival outcomes were analyzed using the Kaplan-Meier method, and comparisons between overexpression and normal expression groups were made using a log-rank test.
Statistical analysis
Comparisons between two independent populations were conducted using an unpaired or paired two-tailed Student’s t-test, assuming a normal distribution. For group comparisons, one-way ANOVA was used with Tukey’s multiple comparisons. *P < 0.05 was the threshold for statistical significance. Details regarding the number of experimental replicates and sample sizes are provided in the figure legends. Statistical analyses were performed using GraphPad Prism. Data points on the graphs represent mean values, with error bars indicating the standard deviation (SD).
Electronic supplementary material
Below is the link to the electronic supplementary material.
Acknowledgements
Anna M. Kolarzyk and Yujin Kwon contributed equally to this work as co-first authors. Keng-Jung Lee and Elizabeth Oh contributed equally to this work as co-second authors. We thank Ben Z. Stanger for providing the pancreatic cancer cell line. We thank Claudia Fischbach-Teschl, Anushka Dongre, Hong-Hee Won, Chunyi Hu, and Ailong Ke for the helpful discussion. We thank Jake June-Koo Lee for analyzing the overall survival data of human pancreatic cancer patients in The Cancer Genome Atlas. We thank Cornell Institute of Biotechnology’s Imaging Facility, with NIH 1S10OD010605 funding, for the shared IVIS Spectrum. This work was performed at the Cornell NanoScale Facility (CNF), a member of the National Nanotechnology Coordinated Infrastructure (NNCI), which was supported by the National Science Foundation (Grant NNCI-1542081). This work was performed at the Cornell Biotechnology Resource Center (BRC) imaging facility supported by the NIH (S10OD025049) for the IVIS-Spectrum optical imager.
Abbreviations
- PDAC
Pancreatic Ductal Adenocarcinoma
- TGFβ
Transforming Growth Factor Beta
- ALK7
Activin Receptor-Like Kinase 7
- EMT
Epithelial-Mesenchymal Transition
- BM
Basement Membrane
- CTCs
Circulating Tumor Cells
- ECM
Extracellular Matrix
- 3D
Three-Dimensional
- shRNA
Short Hairpin RNA
- KD
Knockdown
- BV
Blood Vessel
- HUVECs
Human Umbilical Vein Endothelial Cells
- GO
Gene Ontology
- IF
Immunofluorescence
- FBS
Fetal Bovine Serum
- PDMS
Polydimethylsiloxane
Author contributions
A.M.K., W.K.K. and E.L. developed the concept and design of the study. A.M.K., Y.K., K.J.L., E.O., and S.Y.C. provided data and supported analysis. A.M.K., Y.K., I.C., R.L., T.J.K., and J.L. performed data analysis. A.M.K., Y.K., G.W., A.H.K., O.G., and M.H. provided resources. A.M.K., Y.K., W.K.K., and E.L. wrote the original manuscript draft. All authors reviewed and approved the final manuscript.
Funding
This study was supported by grants from the NIH (CA252162, CA279560, AI168886, and HL165135), NSF (Career Award, CBET 2338610), and the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (RS-2025-00515211). This study was also supported by the V Foundation Translational Research Award and the Multi-Investigator Seed Grant from the Office of the Vice Provost for Academic Integration at Cornell University.
Data availability
Datasets describing the genetics, epidemiology, and survival of patients with human pancreatic cancer at the time of the current study are available from the Cancer Genome Atlas Program (TCGA), which is maintained by the NIH NCI. The data supporting the findings of this study are available in the main text or in the supplementary information section. Additional data that support the finding of this study are available from the corresponding author upon reasonable request.
Declarations
Ethics approval and consent to participate
All animal experiments were approved by the Institutional Animal Care and Use Committee of Cornell University (protocol no. 2019-0072).
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Anna M. Kolarzyk and Yujin Kwon contributed equally to this work.
Contributor Information
Won Kyu Kim, Email: wkkim@kist.re.kr.
Esak Lee, Email: el767@cornell.edu.
References
- 1.Kamisawa T, Wood LD, Itoi T, Takaori K. Pancreatic cancer. Lancet. 2016;388:73–85. [DOI] [PubMed] [Google Scholar]
- 2.Sarantis P, Koustas E, Papadimitropoulou A, Papavassiliou AG, Karamouzis MV. Pancreatic ductal adenocarcinoma: treatment hurdles, tumor microenvironment and immunotherapy. World J Gastrointest Oncol. 2020;12:173–81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Masugi Y. The desmoplastic stroma of pancreatic cancer: multilayered levels of heterogeneity, clinical significance, and therapeutic opportunities. Cancers (Basel) 2022; 14. [DOI] [PMC free article] [PubMed]
- 4.Sahai E. Illuminating the metastatic process. Nat Rev Cancer. 2007;7:737–49. [DOI] [PubMed] [Google Scholar]
- 5.Orth M, Metzger P, Gerum S, Mayerle J, Schneider G, Belka C, Schnurr M, Lauber K. Pancreatic ductal adenocarcinoma: biological hallmarks, current status, and future perspectives of combined modality treatment approaches. Radiat Oncol. 2019;14:141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Thomas D, Radhakrishnan P. Tumor-stromal crosstalk in pancreatic cancer and tissue fibrosis. Mol Cancer. 2019;18:14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Pickup M, Novitskiy S, Moses HL. The roles of TGFβ in the tumour microenvironment. Nat Rev Cancer. 2013;13:788–99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hu T, Su F, Jiang W, Dart DA. Overexpression of activin Receptor-like kinase 7 in breast Cancer cells is associated with decreased cell growth and adhesion. Anticancer Res. 2017;37:3441–51. [DOI] [PubMed] [Google Scholar]
- 9.Michael IP, Saghafinia S, Tichet M, Zangger N, Marinoni I, Perren A, Hanahan D. ALK7 signaling manifests a homeostatic tissue barrier that is abrogated during tumorigenesis and metastasis. Dev Cell. 2019;49:409–e424406. [DOI] [PubMed] [Google Scholar]
- 10.Zeng F, Xu G, Zhou T, Yang C, Wang X, Peng C, Zhou H. Reduced expression of activin receptor-like kinase 7 in breast cancer is associated with tumor progression. Med Oncol. 2012;29:2519–26. [DOI] [PubMed] [Google Scholar]
- 11.Nguyen DT, Lee E, Alimperti S, Norgard RJ, Wong A, Lee JJ, Eyckmans J, Stanger BZ, Chen CS. A biomimetic pancreatic cancer on-chip reveals endothelial ablation via ALK7 signaling. Sci Adv. 2019;5:eaav6789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Principe M, Chanal M, Karam V, Wierinckx A, Mikaélian I, Gadet R, Auger C, Raverot V, Jouanneau E, Vasiljevic A, et al. ALK7 expression in prolactinoma is associated with reduced prolactin and increased proliferation. Endocr Relat Cancer. 2018;25:795–806. [DOI] [PubMed] [Google Scholar]
- 13.Asnaghi L, White DT, Key N, Choi J, Mahale A, Alkatan H, Edward DP, Elkhamary SM, Al-Mesfer S, Maktabi A, et al. ACVR1C/SMAD2 signaling promotes invasion and growth in retinoblastoma. Oncogene. 2019;38:2056–75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kashani E, Schnidrig D, Gheinani AH, Ninck MS, Zens P, Maragkou T, Baumgartner U, Schucht P, Rätsch G, Rubin MA, et al. Integrated longitudinal analysis of adult grade 4 diffuse gliomas with long-term relapse interval revealed upregulation of TGF-β signaling in recurrent tumors. Neuro Oncol. 2023;25:662–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Song W, Hu H, Yuan Z, Yao H. A prognostic model for anoikis-related genes in pancreatic cancer. Sci Rep. 2024;14:15200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Luo Q, Hu Z, Zhao H, Fan Y, Tu X, Wang Y, Liu X. The role of TGF-beta in the tumor microenvironment of pancreatic cancer. Genes Dis. 2023;10:1513–24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Homma Y, Endo I, Matsuyama R, Sho M, Mizuno S, Seyama Y, Hirano S, Aono T, Kitami C, Morita Y, et al. Outcomes of lung metastasis from pancreatic cancer: A nationwide multicenter analysis. J Hepatobiliary Pancreat Sci. 2022;29:552–61. [DOI] [PubMed] [Google Scholar]
- 18.Yachida S, White CM, Naito Y, Zhong Y, Brosnan JA, Macgregor-Das AM, Morgan RA, Saunders T, Laheru DA, Herman JM, et al. Clinical significance of the genetic landscape of pancreatic cancer and implications for identification of potential long-term survivors. Clin Cancer Res. 2012;18:6339–47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Link JM, Eng JR, Pelz C, MacPherson-Hawthorne K, Worth PJ, Sivagnanam S, Keith DJ, Owen S, Langer EM, Grossblatt-Wait A, et al. Ongoing replication stress tolerance and clonal T cell responses distinguish liver and lung recurrence and outcomes in pancreatic cancer. Nat Cancer. 2025;6:123–44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Levine JM, Rompen IF, Franco JC, Swett B, Kryschi MC, Habib JR, Diskin B, Hewitt DB, Sacks GD, Kaplan B, et al. The impact of metastatic sites on survival rates and predictors of extended survival in patients with metastatic pancreatic cancer. Pancreatology. 2024;24:887–93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Cancer Genome Atlas Research Network. Electronic address aadhe, Cancer genome atlas research N. Integrated genomic characterization of pancreatic ductal adenocarcinoma. Cancer Cell. 2017;32:185–e203113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Xu J, Lamouille S, Derynck R. TGF-β-induced epithelial to mesenchymal transition. Cell Res. 2009;19:156–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Zhang YE. Non-Smad signaling pathways of the TGF-β family. Cold Spring Harb Perspect Biol 2017; 9. [DOI] [PMC free article] [PubMed]
- 24.Gonzalez-Avila G, Sommer B, Mendoza-Posada DA, Ramos C, Garcia-Hernandez AA, Falfan-Valencia R. vol 137, pg 57,. Matrix metalloproteinases participation in the metastatic process and their diagnostic and therapeutic applications in cancer (2019). Critical Reviews in Oncology Hematology 2019; 138:172–172. [DOI] [PubMed]
- 25.Gialeli C, Theocharis AD, Karamanos NK. Roles of matrix metalloproteinases in cancer progression and their Pharmacological targeting. FEBS J. 2011;278:16–27. [DOI] [PubMed] [Google Scholar]
- 26.He XJ, Jiang XT, Ma YY, Xia YJ, Wang HJ, Guan TP, Shao QS, Tao HQ. REG4 contributes to the invasiveness of pancreatic cancer by upregulating MMP-7 and MMP-9. Cancer Sci. 2012;103:2082–91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zhou DH, Trauzold A, Roder C, Pan G, Zheng C, Kalthoff H. The potential molecular mechanism of overexpression of uPA, IL-8, MMP-7 and MMP-9 induced by TRAIL in pancreatic cancer cell. Hepatobiliary Pancreat Dis Int. 2008;7:201–9. [PubMed] [Google Scholar]
- 28.Kwak TJ, Lee E. In vitro modeling of solid tumor interactions with perfused blood vessels. Sci Rep. 2020;10:20142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kwak TJ, Lee E. Rapid multilayer microfabrication for modeling organotropic metastasis in breast cancer. Biofabrication 2020; 13. [DOI] [PubMed]
- 30.Baluk P, Morikawa S, Haskell A, Mancuso M, McDonald DM. Abnormalities of basement membrane on blood vessels and endothelial sprouts in tumors. Am J Pathol. 2003;163:1801–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Jacob A, Prekeris R. The regulation of MMP targeting to invadopodia during cancer metastasis. Front Cell Dev Biol. 2015;3:4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Claesson-Welsh L, Dejana E, McDonald DM. Permeability of the endothelial barrier: identifying and reconciling controversies. Trends Mol Med. 2021;27:314–31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.HU T, SU F, JIANG W. Overexpression of activin Receptor-like kinase 7 in breast Cancer cells is associated with decreased cell growth and adhesion. Anticancer Res. 2017;37:3441–51. [DOI] [PubMed] [Google Scholar]
- 34.Carvalho TMA, Audero MM, Greco MR, Ardone M, Maggi T, Mallamaci R, Rolando B, Arpicco S, Ruffinatti FA, Pla AF, et al. Tumor microenvironment modulates invadopodia activity of Non-Selected and Acid-Selected pancreatic Cancer cells and its sensitivity to gemcitabine and C18-Gemcitabine. Cells. 2024;13:730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Karamanou K, Franchi M, Vynios D, Brézillon S. Epithelial-to-mesenchymal transition and invadopodia markers in breast cancer: lumican a key regulator. Semin Cancer Biol. 2020;62:125–33. [DOI] [PubMed] [Google Scholar]
- 36.Strilic B, Offermanns S. Intravascular survival and extravasation of tumor cells. Cancer Cell. 2017;32:282–93. [DOI] [PubMed] [Google Scholar]
- 37.Shenoy AK, Lu J. Cancer cells remodel themselves and vasculature to overcome the endothelial barrier. Cancer Lett. 2016;380:534–44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Harada A, Matsumoto S, Yasumizu Y, Shojima K, Akama T, Eguchi H, Kikuchi A. Localization of KRAS downstream target ARL4C to invasive pseudopods accelerates pancreatic cancer cell invasion. Elife; 2021. p. 10. [DOI] [PMC free article] [PubMed]
- 39.Liang R, Song G. Matrix stiffness-driven cancer progression and the targeted therapeutic strategy. Mechanobiol Med. 2023;1:100013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Levental KR, Yu H, Kass L, Lakins JN, Egeblad M, Erler JT, Fong SF, Csiszar K, Giaccia A, Weninger W, et al. Matrix crosslinking forces tumor progression by enhancing integrin signaling. Cell. 2009;139:891–906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Paszek MJ, Zahir N, Johnson KR, Lakins JN, Rozenberg GI, Gefen A, Reinhart-King CA, Margulies SS, Dembo M, Boettiger D, et al. Tensional homeostasis and the malignant phenotype. Cancer Cell. 2005;8:241–54. [DOI] [PubMed] [Google Scholar]
- 42.Provenzano PP, Inman DR, Eliceiri KW, Keely PJ. Matrix density-induced mechanoregulation of breast cell phenotype, signaling and gene expression through a FAK–ERK linkage. Oncogene. 2009;28:4326–43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Parekh A, Ruppender NS, Branch KM, Sewell-Loftin MK, Lin J, Boyer PD, Candiello JE, Merryman WD, Guelcher SA, Weaver AM. Sensing and modulation of invadopodia across a wide range of rigidities. Biophys J. 2011;100:573–82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Alexander NR, Branch KM, Parekh A, Clark ES, Iwueke IC, Guelcher SA, Weaver AM. Extracellular matrix rigidity promotes invadopodia activity. Curr Biol. 2008;18:1295–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Chang J, Pang EM, Adebowale K, Wisdom KM, Chaudhuri O. Increased stiffness inhibits invadopodia formation and cell migration in 3D. Biophys J. 2020;119:726–36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Yadav PK, Ghosh M, Kataria M. Matrix Metalloproteinases (MMPs) in Cancer Immunotherapy. In Handbook of Oxidative Stress in Cancer: Therapeutic Aspects. Edited by Chakraborti S. Singapore: Springer Singapore; 2021: 1–26.
- 47.Winer A, Adams S, Mignatti P. Matrix metalloproteinase inhibitors in Cancer therapy: turning past failures into future successes. Mol Cancer Ther. 2018;17:1147–55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Fields GB. The rebirth of matrix metalloproteinase inhibitors: moving beyond the dogma. Cells; 2019. p. 8. [DOI] [PMC free article] [PubMed]
- 49.Aiello NM, Maddipati R, Norgard RJ, Balli D, Li J, Yuan S, Yamazoe T, Black T, Sahmoud A, Furth EE, et al. EMT subtype influences epithelial plasticity and mode of cell migration. Dev Cell. 2018;45:681–e695684. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Brown BA, Myers PJ, Adair SJ, Pitarresi JR, Sah-Teli SK, Campbell LA, Hart WS, Barbeau MC, Leong K, Seyler N, et al. A histone Methylation-MAPK signaling Axis drives durable Epithelial-Mesenchymal transition in hypoxic pancreatic Cancer. Cancer Res. 2024;84:1764–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Rhim AD, Mirek ET, Aiello NM, Maitra A, Bailey JM, McAllister F, Reichert M, Beatty GL, Rustgi AK, Vonderheide RH, et al. EMT and dissemination precede pancreatic tumor formation. Cell. 2012;148:349–61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Kim MP, Evans DB, Wang H, Abbruzzese JL, Fleming JB, Gallick GE. Generation of orthotopic and heterotopic human pancreatic cancer xenografts in immunodeficient mice. Nat Protoc. 2009;4:1670–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Chai MG, Kim-Fuchs C, Angst E, Sloan EK. Bioluminescent orthotopic model of pancreatic cancer progression. J Vis Exp 2013. [DOI] [PMC free article] [PubMed]
- 54.Nguyen DH, Stapleton SC, Yang MT, Cha SS, Choi CK, Galie PA, Chen CS. Biomimetic model to reconstitute angiogenic sprouting morphogenesis in vitro. Proc Natl Acad Sci U S A. 2013;110:6712–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, Preibisch S, Rueden C, Saalfeld S, Schmid B, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Adamson RH, Lenz JF, Curry FE. Quantitative laser scanning confocal microscopy on single capillaries: permeability measurement. Microcirculation. 1994; 1:251–265. [DOI] [PubMed]
- 57.Polacheck WJ, Kutys ML, Tefft JB, Chen CS. Microfabricated blood vessels for modeling the vascular transport barrier. Nat Protoc. 2019;14:1425–54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL, Pachter L. Differential gene and transcript expression analysis of RNA-seq experiments with tophat and cufflinks. Nat Protoc. 2012;7:562–78. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013;14:R36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, Salzberg SL, Wold BJ, Pachter L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol. 2010;28:511–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Datasets describing the genetics, epidemiology, and survival of patients with human pancreatic cancer at the time of the current study are available from the Cancer Genome Atlas Program (TCGA), which is maintained by the NIH NCI. The data supporting the findings of this study are available in the main text or in the supplementary information section. Additional data that support the finding of this study are available from the corresponding author upon reasonable request.