Abstract
Sortase (SrtA), an enzyme that anchors surface proteins to the cell wall of Gram-positive bacteria, cleaves sorting signals at the LPXTG motif. We have identified a second sortase (SrtB) in the Gram-positive pathogen Staphylococcus aureus that is required for anchoring of a surface protein with a NPQTN motif. Purified SrtB cleaves NPQTN-bearing peptides in vitro, and a srtB mutant is defective in the persistence of animal infections. srtB is part of an iron-regulated locus called iron-responsive surface determinants (isd), which also contains a ferrichrome transporter and surface proteins with NPQTN and LPXTG motifs. Cell wall-anchored surface proteins and the isd locus seem involved in a novel mechanism of iron acquisition that is important for bacterial pathogenesis.
Molecular iron is an essential cofactor for bacterial growth, because microbes use iron during aerobic respiration, DNA replication, and other biological processes (1, 2). The importance of iron as a cofactor is demonstrated further by the fact that bacteria have evolved processes for the acquisition of iron from their environment (3). Siderophores are secreted molecules that bind free iron and then are captured by bacterial surface receptors that deliver iron for transport across the cytoplasmic membrane (4). In addition to retrieving free iron, bacteria also can acquire iron that is bound by various proteins (4, 5). Iron acquisition is critical for the establishment of human infections, because several host proteins sequester iron and reduce the concentration of free iron within tissues in an effort to prevent bacterial growth. Thus, to establish an invasive infection, the Gram-positive human pathogen Staphylococcus aureus must have evolved mechanisms for acquiring iron from the host environment (6).
Surface proteins play important roles in the pathogenesis of Gram-positive bacterial infections (7) and are anchored to the cell wall by a mechanism requiring a C-terminal sorting signal (8). Sorting signals are composed of an LPXTG motif, a hydrophobic domain, and a tail of positively charged residues (9, 10). The hydrophobic domain and charged tail function to retain surface proteins temporarily within the secretory pathway and provide for the recognition of the LPXTG motif (10). Sortase, an extracellular transpeptidase, catalyzes cell wall anchoring by cleaving the LPXTG motif between the threonine and glycine residues (11–13). The carboxyl group of threonine (T) at the C-terminal end of surface proteins is subsequently amide-linked to cross-bridges in the bacterial cell wall (14–16). Deletion of the sortase gene (srtA) in S. aureus or other bacterial pathogens results in severe virulence defects (17, 18). Sortase and surface proteins with LPXTG motifs are found in all Gram-positive bacteria (7, 19). Thus, it seems that the sortase-mediated mechanism of protein anchoring is a universal strategy of Gram-positive bacteria to display virulence factors on the microbial surface. We report here the identification of anchored surface proteins that are regulated in response to iron during the pathogenesis of S. aureus infections. Further, we have characterized a cell wall-anchoring pathway mediated by sortase B (SrtB) that cleaves surface protein at an NPQTN signal and anchors the polypeptide to the cell wall envelope.
Experimental Procedures
Bacterial Strains and Plasmids.
S. aureus strains RN4220 (res−; ref. 20) and Newman (21) have been described. srtA and srtB knockout mutations were generated by allelic exchange with the tetM and ermC genes, respectively. Briefly, flanking sequences were cloned into the temperature-sensitive shuttle vector pTS1, and marker exchange was induced with a temperature shift to 43°C and antibiotic selection. S. aureus fur knockout strains were generated by phage transduction using the mutant allele of S. aureus MJH010 (22) and selection on tetracycline plates. Gene fusions were generated by ligating PCR-amplified DNA fragments at unique restriction sites. Cell wall sorting signals were amplified by using primers with abutted KpnI and BamHI restriction sites and ligated into pSeb-staphylococcal protein A (Spa; ref. 10) cut with the same enzymes. The recombinant pSeb shuttle vector was transformed into Escherichia coli, sequenced, and electroporated into S. aureus. PCR amplification was used to introduce the FLAG epitope into isdC and to clone the recombinant gene under the control of the srtA promoter. Bacterial strains and plasmids used in this report are listed in Table 1.
Table 1.
Strains and plasmids used in this study
| Property | Reference | |
|---|---|---|
| Strains | ||
| S. aureus Newman | Human clinical isolate, wild-type srtA and srtB | 21 |
| S. aureus SKM7 | Isogenic Newman variant, Δ(srtB) | This study |
| S. aureus RN4220 | Restriction deficient laboratory strain, wild-type srtA and srtB | 20 |
| S. aureus SKM1 | Isogenic variant of RN4220, Δ(srtA) | 17 |
| S. aureus SKM5 | Isogenic variant of RN4220, Δ(srtB) | This study |
| S. aureus SKM15 | Isogenic variant of RN4220, Δ(fur) | This study |
| S. aureus SKM16 | Isogenic variant of RN4220, Δ(fur, srtB) | This study |
| E. coli BL21 (DE3) | E. coli B strain, λ lysogen, T7 expression | 23 |
| Plasmid | ||
| pSM74 | seb promoter and orf fused to 5′ of isdC sorting signal | This study |
| pSM75 | pSM74 plus srtB expressed from the srtA promoter | This study |
| pSM79 | pSeb-Spa plus srtB expressed from the srtA promoter | This study |
| pSM80 | seb promoter fused isdC orf carrying a FLAG epitope tag | This study |
| pSM83 | pSM80 plus srtB expressed from the srtA promoter | This study |
| pSM84 | pET16a derivative, srtBΔN expressed from the T7 promoter | This study |
| pSeb-Spa | seb promoter and orf fused to 5′ end of spa sorting signal | 10 |
| pSeb-FnbA | seb promoter and orf fused to 5′ end of fnbA sorting signal | 10 |
| pSeb-FnbB | seb promoter and orf fused to 5′ end of fnbB sorting signal | This study |
| pSeb-ClfA | seb promoter and orf fused to 5′ end of clfA sorting signal | 17 |
| pSeb-ClfB | seb promoter and orf fused to 5′ end of clfB sorting signal | This study |
| pSeb-SdrD | seb promoter and orf fused to 5′ end of sdrD sorting signal | This study |
| pSeb-SdrE | seb promoter and orf fused to 5′ end of sdrE sorting signal | This study |
| pSeb-SasA | seb promoter and orf fused to 5′ end of sasA sorting signal | This study |
| pSeb-SasB | seb promoter and orf fused to 5′ end of sasB sorting signal | This study |
| pSeb-SasC | seb promoter and orf fused to 5′ end of sasC sorting signal | This study |
| pSeb-SasE | seb promoter and orf fused to 5′ end of sasE sorting signal | This study |
| pSeb-SasF | seb promoter and orf fused to 5′ end of sasF sorting signal | This study |
| pSeb-SasG | seb promoter and orf fused to 5′ end of sasG sorting signal | This study |
| pSeb-SasH | seb promoter and orf fused to 5′ end of sasH sorting signal | This study |
| pSeb-SasJ | seb promoter and orf fused to 5′ end of sasJ sorting signal | This study |
Pulse-Labeling Experiments.
S. aureus strains were grown in tryptic soy broth, suspended in minimal medium with or without iron, and subjected to pulse labeling with 100 μCi (1 Ci = 37 GBq) of [35S]methionine. Samples were precipitated with trichloroacetic acid and either boiled directly in 4% SDS or first digested with lysostaphin or mutanolysin at 37°C. Samples were immunoprecipitated with α-SEB (Sigma) in radioimmunoprecipitation assay buffer with protein A-Sepharose (Sigma). Immunoprecipitates were washed in radioimmunoprecipitation assay, boiled in sample buffer, and analyzed by 14% SDS/PAGE and PhosphorImager (10).
Peptide Cleavage by SrtA and SrtB.
Reactions were performed in 250 μl containing 50 mM Tris⋅HCl buffer and 150 mM NaCl, pH 7.5, and as indicated 6.0 μM SrtBΔN or SrtAΔN with 3 μM NPQTN or LPXTG fluorescent peptides (DABCYL-KVENPQTNAGT-EDANS and DABCYL-LPETG-EDANS) and 5 mM [2-(trimethylammonium)ethyl]methanethiosulfonate or 10 mM DTT. Reactions were carried out for 2 h at 37°C and analyzed fluorometrically by using 350 nm for excitation and 500 nm for recordings. Expression and purification of SrtAΔN has been reported previously (13). srtBΔN was PCR-amplified by using primers that deleted the N-terminal 30 residues encompassing the membrane anchor sequence and introducing NdeI and BamHI sites. srtBΔN was cloned into the pET16b expression vector (Novagen) and transformed into E. coli BL21(DE3) (23). After isopropyl β-D-thiogalactoside induction of cultures, cells were lysed in a French press, and recombinant protein was purified by affinity chromatography on nickel-nitrilotriacetic acid.
Results and Discussion
SrtA Anchors LPXTG Motif-Bearing Surface Proteins.
Examination of the S. aureus genome for srtA homologs revealed a second sortase gene designated srtB (19, 24). SrtB is a polypeptide of 246 amino acids with a putative N-terminal membrane anchor and an active site cysteine located within the TLXTC signature motif of sortases (24, 25). We wondered whether each sortase enzyme, SrtA and SrtB, cleaves specific subsets of surface proteins and searched the staphylococcal genome for surface protein genes by using BLAST searches with known sorting signals as queries. Table 2 lists the sorting signals of 20 different surface protein genes (24). To test whether sorting signals were substrates for either sortase, isogenic mutants of S. aureus RN4220 were generated that lacked either srtA or srtB. Staphylococcal strains were transformed with plasmids that encoded hybrid proteins in which each sorting signal had been fused to the C terminus of enterotoxin B (Seb), a protein that normally is secreted into the extracellular environment (26). When analyzed in wild-type S. aureus, all 16 Seb hybrids examined were cleaved to generate mature anchored species (Table 2). A staphylococcal mutant with a chromosomal deletion of srtA (27) failed to anchor all 16 fusions, indicating that sorting signals with an LPXTG motif require SrtA for their anchoring to the cell wall envelope (Table 2). In contrast, mutants with a deletion of srtB anchored LPXTG motif-containing proteins to the cell wall in a manner that was indistinguishable from that of wild-type staphylococci (Table 2).
Table 2.
SrtA-mediated anchoring of surface proteins with LPXTG sorting signals
| Protein | Sorting signal* | % anchored protein†
|
||
|---|---|---|---|---|
| wt | srtA− | srtB− | ||
| Spa | LPETG EENPFIGTTVFGGLSLALGAALLAGRRREL | 100 | 0 | 98 |
| FnbA | LPETG GEESTNKGMLFGGLFSILGLALLRRNKKNHKA | 98 | 0 | 99 |
| FnbB | LPETG GEESTNNGMLFGGLFSILGLALLRRNKKNHKA | 98 | 0 | 99 |
| ClfA | LPDTG SEDEANTSLIWGLLASIGSLLLRRKKENKDKK | 71 | 0 | 75 |
| ClfB | LPETG DKSENTNATLFGAMMALLGSLLLFRKRKQDHKEKA | 99 | 0 | 100 |
| SdrC | LPETG SENNNSNNGTLFGGLFAALGSLLSFGRRRKKQNK | N/D | N/D | N/D |
| SdrD | LPETG NENSGSNNAATLFGGLFAALGSLLLFGRRKKQNK | 99 | 0 | 100 |
| SdrEa | LPETG SENNGSNNATLFGGLFAALGSLLLFGRRKKQNK | 98 | 0 | 98 |
| Pls | LPDTG NDAQNNGTLFGSLFAALGGLFLVGRRRKNKNNEEK | N/D | N/D | N/D |
| SasA | LPDTG DSIKQNGLLGGVMTLLVGLGLMRKKKKDENDQDDSQA | 62 | 0 | 66 |
| SasB | LPDTG MSHNDDLPYAELALGAGMAFLIRRFTKKDQQTEE | 89 | 0 | 92 |
| SasD | LPAAG ESMTSSILTASIAALLLVSGLFLAFRRRSTNK | 41 | 0 | 40 |
| IsdA(SasE) | LPKTG LTSVDNFISTVAFATLALLGSLSLLLFKRKESK | 44 | 0 | 45 |
| SasF | LPKAG ETIKEHWLPISVIVGAMGVLMIWLSRRNKLKNKA | 32 | 0 | 24 |
| SasG | LPKTG LESTQKGLIFSSIIGIAGLMLLARRRKN | 81 | 0 | 77 |
| SasH | LPKTG TNQSSSPEAMFVLLAGIGLIATVRRRKAS | 84 | 0 | 80 |
| SasI | LPKTG ETTSSQSWWGLYALLGMLALFIPKFRKESK | N/D | N/D | N/D |
| IsdB(SasJ) | LPQTG EESNKDMTLPLMALLALSSIVAFVLPRKRKN | 27 | 0 | 23 |
| Cna | LPKTG MKIITSWITWVFIGILGLYILRKRFNS | N/D | N/D | N/D |
Sorting signals were fused to the C terminus of enterotoxin B (Seb), a protein that normally is secreted into the extracellular medium of staphylococci. Sorting signals are aligned at the LPXTG motif, and the hydrophobic domains are underligned.
Pulse-labeled staphylococcal cultures were incubated for 5 min, samples were digested with lysostaphin, and Seb fusion protein was immunoprecipitated and analyzed by SDS/PAGE and PhosphorImager. The percentage amount of mature (anchored) protein as compared to the total amount of P1 and P2 precursor and mature protein is indicated.
srtB and the isd Operon.
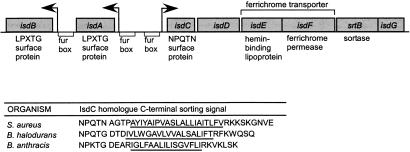
Unlike S. aureus srtA, which seems to be transcribed as a single gene (27), srtB is located within an operon that also contains three genes with unknown function and a putative iron acquisition system consisting of a lipoprotein homologous to hemin-binding proteins and a ferrichrome transporter (refs. 28 and 29; Fig. 1). Upstream of this transcriptional unit is a Fur box (30), a DNA sequence to which the ferric uptake repressor (Fur) binds and inhibits transcription when staphylococci are grown in the presence of high extracellular iron concentrations (22, 31). Adjacent to this operon and divergently transcribed from independent promoters are two surface proteins with LPXTG motif-containing sorting signals whose expression also appears to be regulated by Fur. This locus was named isd, for iron-responsive surface determinant. isdC, the first gene of this operon, encodes a polypeptide with an N-terminal signal peptide, a C-terminal hydrophobic domain, and charged tail, resembling sorting signals. The sequence NPQTN is located upstream of the hydrophobic domain replacing the LPXTG motif of srtA-type sorting signals. The isd operon is conserved in B. halodurans and B. anthracis the isdC homologues of which encode the NPQTG and NPKTG sequences, respectively (Fig. 1).
Figure 1.
Genomic organization of S. aureus isd. srtB is located in an operon with a putative iron transporter (isdE, isdF) and a surface protein with a C-terminal NPQTN sorting signal (isdC). Adjacent to this operon are two LPXTG-containing surface proteins that are substrates for srtA-mediated anchoring as shown in Table 2. All transcriptional units contain a Fur box regulatory element upstream of their putative promoters (arrows). The peptide sequence of C-terminal sorting signals of IsdC from S. aureus is compared with that of Bacillus halodurans and Bacillus anthracis.
SrtB Cleaves IsdC.
We wondered whether IsdC represents a surface protein substrate for SrtB. The putative sorting signal was fused to the C terminus of Seb (10), staphylococci expressing the hybrid protein were pulse-labeled with [35S]methionine, and Seb-IsdC was immunoprecipitated with antibodies raised against Seb. The cytoplasmic (P1) and secreted (P2) precursors of Seb-IsdC could be distinguished from the mature sortase-cleaved species after separation on SDS/PAGE (Fig. 2 A and B). When grown in iron-rich media, S. aureus RN4220 cannot process Seb-IsdC unless SrtB is overexpressed from a constitutive promoter such as the srtB promoter (Fig. 2B). Under iron-restrictive conditions, small amounts of the fusion protein are cleaved to generate the mature species, and this activity is abolished when srtB is deleted (Fig. 2C). Seb-IsdC cleavage in cells that overexpress srtB from a constitutive promoter is not affected by changes in extracellular iron concentration (Fig. 2C).
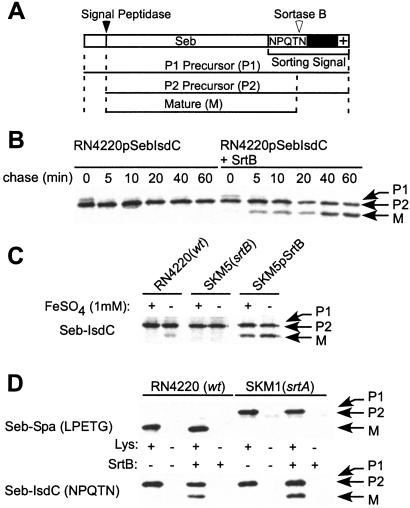
Figure 2.
IsdC contains a C-terminal NPQTN sorting signal and is cleaved by SrtB. (A) The drawing shows the fusion of the C-terminal sorting signal of IsdC to Seb. The cytoplasmic precursor (P1) is cleaved by signal peptidase, generating the extracellular P2 precursor. SrtB cleaves P2 and anchors the mature (M) species to the cell wall. (B) S. aureus RN4220 harboring pSM74 or pSM75 was subjected to pulse labeling with [35S]methionine, and Seb-IsdC was immunoprecipitated and analyzed on SDS/PAGE. pSM74 encodes seb-isdC, whereas pSM75 encodes seb-isdC as well as srtB under control of the srtA promoter. (C) S. aureus RN4220 (pSM74), SKM5 (srtB) (pSM74) and SKM5 (pSM75) were pulse-labeled in the presence or absence of 1 mM FeSO4 and incubated for 30 min. wt, wild type. (D) S. aureus RN4220 and SKM1 (srtA) harboring pSM74, pSM75, pSM79, or pSeb-Spa (Seb with the LPXTG sorting signal or protein A) were pulse-labeled in the presence or absence of 100 mM NH2OH. pSM79 encodes the seb-spa fusion of pSeb-Spa and srtB under control of the srtA promoter. After trichloroacetic acid precipitation, samples were treated with 10 μg/ml lysostaphin or left untreated and then boiled in 4% SDS buffer. Immunoprecipitated Seb-Spa and Seb-IsdC were analyzed by SDS/PAGE and PhosphorImager.
Treatment of acid-precipitated proteins in staphylococcal cultures with hot SDS solubilizes extracellular polypeptides, whereas cytoplasmic-, membrane-, or cell wall-anchored proteins are not extracted by this method, because these proteins reside within the murein sacculus (8). However, digestion of the cell wall with lysostaphin allows cellular proteins to be extracted with hot SDS (8). Pulse-labeled Seb-Spa, bearing a C-terminal sorting signal with an LPXTG motif, or Seb-IsdC is not solubilized by SDS extraction unless the cell wall was digested with lysostaphin (Fig. 2D). Hydroxylamine (NH2OH) is a strong nucleophile that releases surface protein hydroxamate into the extracellular medium by substituting for the physiological peptidoglycan substrate, i.e., the amino groups of cross-bridges (13). Treatment of wild-type staphylococci with hydroxylamine released Seb-Spa into the extracellular medium, whereas srtA mutant strains failed to release the surface protein hydroxamate. Hydroxylamine treatment also released mature Seb-IsdC into the extracellular medium, suggesting the formation of surface protein hydroxamate. Hydroxylaminolysis was stimulated by overexpression of SrtB, whereas deletion of srtA had no effect on the cleavage of the NPQTN sorting signal (Fig. 2D). Together these results suggest that NPQTN sorting signals are sufficient to anchor Seb reporter proteins to the envelope by a mechanism requiring srtB but not srtA. Further, srtB overexpression does not affect the cell wall anchoring of LPXTG motif-containing surface proteins.
IsdC Is Anchored to the Cell Wall by a Mechanism Requiring srtB.
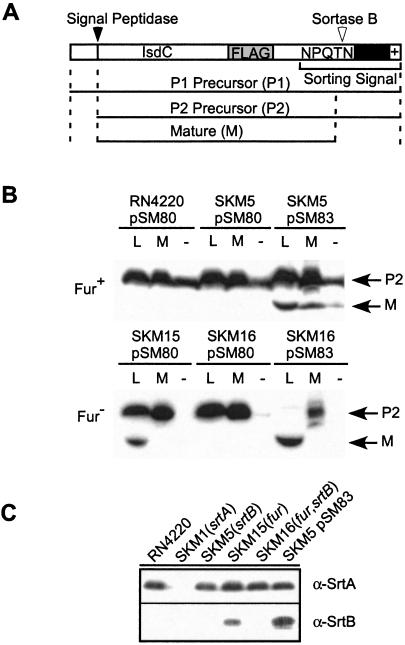
To test whether IsdC is a substrate for SrtB-mediated anchoring to the cell wall envelope, we constructed isdCFLAG, a mutant gene with a DNA insertion encoding a FLAG epitope tag upstream of the NPQTN sorting signal (Fig. 3A). The cell walls of staphylococci overexpressing IsdCFLAG from the seb promoter were digested with two different enzymes, and IsdCFLAG was detected by immunoblotting. Lysostaphin cuts the pentaglycine cross-bridge of the cell wall (32) close to the anchoring point of surface proteins bearing the LPXTG motif (33). Mutanolysin, an N-acetylmuramidase, cleaves the β1–4 glycosidic bond between N-acetylmuramic acid and N-acetylglucosamine (34). Mutanolysin releases cell wall-anchored surface proteins as a spectrum of fragments with linked peptidoglycan, all of which migrate more slowly on SDS/PAGE than the lysostaphin-released counterpart (Fig. 3B; ref. 35). When expressed in S. aureus RN4220 under iron-replete conditions, the mobility of IsdCFLAG on SDS/PAGE was not altered after digesting the cell wall with lysostaphin or muramidase. However, expression of IsdCFLAG in the fur mutant caused some but not all IsdCFLAG to migrate faster on SDS/PAGE than the P2 species observed in wild-type staphylococci (Fig. 3B). These species therefore represented mature, cell wall-anchored surface protein, because solubilization with mutanolysin resulted in their altered (slowed) mobility on SDS/PAGE. Cell wall anchoring of IsdCFLAG depends on srtB, because neither srtB nor srtB/fur double mutants generated the mature species (Fig. 3B). Further, overexpression of srtB in fur mutants caused all IsdCFLAG to be linked to the cell wall, whereas overexpression of SrtB in wild-type cells resulted in cell wall anchoring of some but not all IsdCFLAG (Fig. 3B). Thus, IsdC is a cell wall-anchored surface protein that requires SrtB for proper attachment to the envelope.
Figure 3.
Fur-regulated cell wall anchoring of IsdC requires SrtB. (A) The FLAG epitope tag was introduced into the coding sequence of isdC, and the gene was expressed from the seb promoter in pSM80. The drawing shows the primary structure of IsdCFLAG. The cytoplasmic precursor (P1) is cleaved by signal peptidase, generating extracellular P2. The P2 precursor is cleaved by SrtB, tethering the mature (M) species to the cell wall. (B) After transformation with pSM80 or pSM83, a pSM80 derivative encoding srtB under control of the srtA promoter, S. aureus cultures were precipitated with trichloroacetic acid and treated with the muralytic enzymes lysostaphin (L) and mutanolysin (M) or left untreated (−). Proteins were separated on SDS/PAGE, and IsdCFLAG was identified by immunoblotting with FLAG-specific monoclonal antibody. (C) Immunoblotting of proteins from lysostaphin-treated staphylococcal extracts with specific rabbit antibody reveals the expression of srtA and srtB in various S. aureus strains.
Expression of isd Is Controlled by Fur.
To test whether expression of the isd operon is controlled by Fur, staphylococcal srtB expression was measured by immunoblotting with specific antibody (Fig. 3C). S. aureus RN4220 did not express srtB in the presence of extracellular iron; however, deletion of the fur gene caused the mutant staphylococcal strain SKM15 to express srtB even in the presence of iron. Thus, Fur controls the expression of the isd operon, which is induced under iron-limiting conditions, as is typically encountered when the invasive pathogen S. aureus enters a human host.
Purified SrtB Cleaves NPQTN Peptide in Vitro.
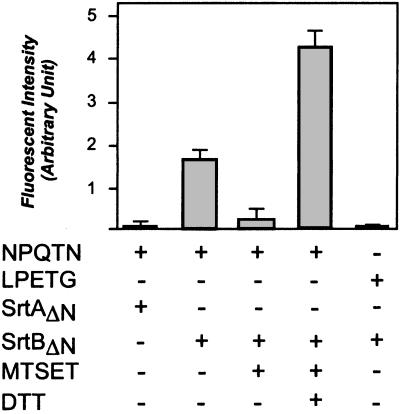
SrtAΔN and SrtBΔN, recombinant sortases with a six-histidine tag replacing the N-terminal membrane anchor, were purified from E. coli extracts by using affinity chromatography and incubated with peptide substrates containing either the LPETG or NPQTN motifs. Purified SrtAΔN but not purified SrtBΔN cut LPETG-containing peptides (Fig. 4). In contrast, purified SrtBΔN but not SrtAΔN cleaved the NPQTN peptides (Fig. 4). Similar to the previously reported SrtAΔN (13), SrtBΔN was inhibited by incubation with [2-(trimethylammonium)ethyl]methanethiosulfonate, suggesting that SrtB also employs an active site cysteine as a nucleophile to cleave its substrate (Fig. 4). The strong reducing agent DTT not only rescued [2-(trimethylammonium)ethyl]methanethiosulfonate-inhibited SrtBΔN but also stimulated cleavage of NPQTN peptides in the absence of the inhibitor (data not shown).
Figure 4.
Purified SrtB cleaves NPQTN peptides in vitro. SrtAΔN and SrtBΔN, recombinant sortases with a six-histidine tag replacing the N-terminal membrane anchor, were purified from E. coli extracts by using affinity chromatography. Sortase enzymes were incubated with the LPETG or NPQTN peptides, and cleavage was measured as an increase in fluorescence intensity. [2-(trimethylammonium)ethyl]methanethiosulfonate (MTSET) or DTT were added as indicated.
srtB Is Required for Staphylococcal Persistence During Host Infection.
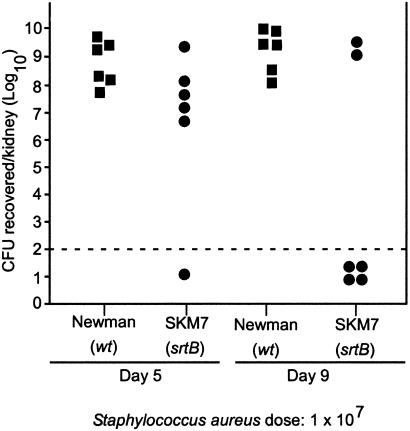
To analyze the role of srtB and SrtB-mediated anchoring of surface protein during infection, the virulence of staphylococcal strains was assessed by using a murine renal abscess model (36). Mice were injected into the tail vein with 1 × 107 colony forming units of S. aureus Newman, a clinical human isolate, or an isogenic derivative lacking srtB (SKM7). Mice were killed 5 days after infection, and homogenized kidneys were analyzed for abscess formation by incubation on agar medium and colony formation. Previous work showed that the srtA mutant S. aureus displays a 2-log reduction in virulence (>99% reduction of bacteria in kidneys) as compared with the wild-type strain Newman (17). No discernable defect in pathogenicity was detected for the srtB mutant strain (Fig. 5). However, incubation of mice for a period of 9 days revealed a significant reduction in the number of srtB mutant staphylococci within infected kidneys as compared with strain Newman (Fig. 5). Thus, although srtB may not be involved in the initial establishment of S. aureus disease, srtB may be required for staphylococcal persistence in infected tissues.
Figure 5.
Mutants in srtB are defective in persistence of renal infections of mice. Swiss–Webster mice were injected intravenously with 1 × 107 colony forming units (cfus) of the S. aureus clinical isolate Newman or the isogenic srtB mutant (SKM7). After 5 or 9 days, animals were killed, their kidneys were harvested, homogenized in 1% Triton-X100, and incubated on agar plates, and staphylococcal colonies were counted. Each symbol (■, wild type; ●, srtB mutant) represents the staphylococcal count in the kidneys of one animal. The dashed line represents the limit of detection for staphylococci in this assay system.
The Role of Sortases during S. aureus Infection.
Pallen et al. reported that Gram-positive bacterial genomes encode at least two, and in most cases several, different sortase genes (19, 37). However, previous work left the function of multiple sortase genes unresolved. It is shown here that S. aureus expresses two sortases that each function to anchor different classes of surface proteins. SrtA is a constitutively expressed sortase enzyme responsible for anchoring all LPXTG-containing surface proteins, whereas SrtB is specialized to carry out the specific iron-regulated cell wall sorting of a NPQTN signal. Both sortases are virulence factors; however, they seem to perform diverse roles during animal infections. S. aureus SrtA is known to anchor surface proteins involved in adherence to tissue and immune response evasion (38–40). SrtB seems to be dedicated to the essential process of iron acquisition during bacterial infection. The isd locus may function as a concerted unit to acquire iron from the environment, because it is conserved in the six available genome sequences of S. aureus strains as well as in several other Gram-positive bacteria. Many bacterial species express two or more sortases (19). Our results suggest that only some sortases are required for the anchoring of LPXTG motif-bearing polypeptides, leaving open the possibility that other sortases anchor surface proteins with hitherto unknown sorting signals (37). With cases of multiply antibiotic-resistant Gram-positive bacterial infections rising (41, 42), inhibition of the cell wall anchoring of surface proteins emerges as a possible target for the development of therapeutics against these pathogens.
Acknowledgments
We thank Simon Foster (University of Sheffield) for materials. S.K.M. acknowledges support from the Predoctoral Training Program in Genetics (T32GM07104) at the University of California, Los Angeles. Work in the O.S. laboratory is supported by National Institutes of Health-National Institute of Allergy and Infectious Diseases, Infectious Disease Branch Grant AI33987.
Abbreviations
- SrtA
staphylococcal surface protein sorting A
- SrtB
staphylococcal surface protein sorting B
- Spa
staphylococcal protein A
- Seb
staphylococcal enterotoxin B
- fur
ferric uptake repressor
- isd
iron-responsive surface determinant
Footnotes
This paper was submitted directly (Track II) to the PNAS office.
References
- 1.Braun V. Science. 1998;282:2202–2203. doi: 10.1126/science.282.5397.2202. [DOI] [PubMed] [Google Scholar]
- 2.Weinberg E D. Biometals. 2000;13:85–89. doi: 10.1023/a:1009293500209. [DOI] [PubMed] [Google Scholar]
- 3.Drazek E S, Hammack C A, Sr, Schmitt M P. Mol Microbiol. 2000;36:68–84. doi: 10.1046/j.1365-2958.2000.01818.x. [DOI] [PubMed] [Google Scholar]
- 4.Hantke K. Curr Opin Microbiol. 2001;4:172–177. doi: 10.1016/s1369-5274(00)00184-3. [DOI] [PubMed] [Google Scholar]
- 5.Schmitt M P. J Bacteriol. 1997;179:838–845. doi: 10.1128/jb.179.3.838-845.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Modun B, Evans R W, Joannou C L, Williams P. Infect Immun. 1998;66:3591–3596. doi: 10.1128/iai.66.8.3591-3596.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Navarre W W, Schneewind O. Microbiol Mol Biol Rev. 1999;63:174–229. doi: 10.1128/mmbr.63.1.174-229.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Schneewind O, Model P, Fischetti V A. Cell. 1992;70:267–281. doi: 10.1016/0092-8674(92)90101-h. [DOI] [PubMed] [Google Scholar]
- 9.Fischetti V A, Pancholi V, Schneewind O. Mol Microbiol. 1990;4:1603–1605. doi: 10.1111/j.1365-2958.1990.tb02072.x. [DOI] [PubMed] [Google Scholar]
- 10.Schneewind O, Mihaylova-Petkov D, Model P. EMBO J. 1993;12:4803–4811. doi: 10.1002/j.1460-2075.1993.tb06169.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Navarre W W, Schneewind O. Mol Microbiol. 1994;14:115–121. doi: 10.1111/j.1365-2958.1994.tb01271.x. [DOI] [PubMed] [Google Scholar]
- 12.Schneewind O, Fowler A, Faull K F. Science. 1995;268:103–106. doi: 10.1126/science.7701329. [DOI] [PubMed] [Google Scholar]
- 13.Ton-That H, Liu G, Mazmanian S K, Faull K F, Schneewind O. Proc Natl Acad Sci USA. 1999;96:12424–12429. doi: 10.1073/pnas.96.22.12424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ton-That H, Faull K F, Schneewind O. J Biol Chem. 1997;272:22285–22292. doi: 10.1074/jbc.272.35.22285. [DOI] [PubMed] [Google Scholar]
- 15.Ton-That H, Schneewind O. J Biol Chem. 1999;274:24316–24320. doi: 10.1074/jbc.274.34.24316. [DOI] [PubMed] [Google Scholar]
- 16.Ton-That H, Mazmanian H, Faull K F, Schneewind O. J Biol Chem. 2000;275:9876–9881. doi: 10.1074/jbc.275.13.9876. [DOI] [PubMed] [Google Scholar]
- 17.Mazmanian S K, Liu G, Jensen E R, Lenoy E, Schneewind O. Proc Natl Acad Sci USA. 2000;97:5510–5515. doi: 10.1073/pnas.080520697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Bolken T C, Franke C A, Jones K F, Zeller G O, Jones C H, Dutton E K, Hruby D E. Infect Immun. 2001;69:75–80. doi: 10.1128/IAI.69.1.75-80.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Pallen M J, Lam A C, Antonio M, Dunbar K. Trends Microbiol. 2001;9:97–101. doi: 10.1016/s0966-842x(01)01956-4. [DOI] [PubMed] [Google Scholar]
- 20.Kreiswirth B N, Lofdahl S, Betley M J, O'Reilly M, Schlievert P M, Bergdoll M S, Novick R P. Nature (London) 1983;305:709–712. doi: 10.1038/305709a0. [DOI] [PubMed] [Google Scholar]
- 21.Duthie E S, Lorenz L L. J Gen Microbiol. 1952;6:95–107. doi: 10.1099/00221287-6-1-2-95. [DOI] [PubMed] [Google Scholar]
- 22.Horsburgh M, Ingham E, Foster S J. J Bacteriol. 2001;183:468–475. doi: 10.1128/JB.183.2.468-475.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Studier F W, Rosenberg A H, Dunn J J, Dubendorff J W. Methods Enzymol. 1990;185:60–89. doi: 10.1016/0076-6879(90)85008-c. [DOI] [PubMed] [Google Scholar]
- 24.Mazmanian S K, Ton-That H, Schneewind O. Mol Microbiol. 2001;40:1049–1057. doi: 10.1046/j.1365-2958.2001.02411.x. [DOI] [PubMed] [Google Scholar]
- 25.Ilangovan U, Ton-That H, Iwahara J, Schneewind O, Clubb R T. Proc Natl Acad Sci USA. 2001;98:6056–6061. doi: 10.1073/pnas.101064198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Tweten R K, Iandolo J J. J Bacteriol. 1983;153:297–303. doi: 10.1128/jb.153.1.297-303.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Mazmanian S K, Liu G, Ton-That H, Schneewind O. Science. 1999;285:760–763. doi: 10.1126/science.285.5428.760. [DOI] [PubMed] [Google Scholar]
- 28.Sebulsky M T, Hohnstein D, Hunter M D, Heinrichs D E. J Bacteriol. 2000;182:4394–4400. doi: 10.1128/jb.182.16.4394-4400.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Morrissey J A, Cockayne A, Hill P J, Williams P. Infect Immun. 2000;68:6281–6288. doi: 10.1128/iai.68.11.6281-6288.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Escolar L, Perez-Martin J, De Lorenzo V. J Bacteriol. 1999;181:6223–6229. doi: 10.1128/jb.181.20.6223-6229.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Xiong A, Singh V K, Cabrera G, Jayaswal R K. Microbiology. 2000;146:659–668. doi: 10.1099/00221287-146-3-659. [DOI] [PubMed] [Google Scholar]
- 32.Schindler C A, Schuhardt V T. Proc Natl Acad Sci USA. 1964;51:414–421. doi: 10.1073/pnas.51.3.414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Navarre W W, Ton-That H, Faull K F, Schneewind O. J Biol Chem. 1998;273:29135–29142. doi: 10.1074/jbc.273.44.29135. [DOI] [PubMed] [Google Scholar]
- 34.Yokogawa K, Kawata S, Nishimura S, Ikeda Y, Yoshimura Y. Antimicrob Agents Chemother. 1974;6:156–165. doi: 10.1128/aac.6.2.156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Sjöquist J, Meloun B, Hjelm H. Eur J Biochem. 1972;29:572–578. doi: 10.1111/j.1432-1033.1972.tb02023.x. [DOI] [PubMed] [Google Scholar]
- 36.Albus A, Arbeit R D, Lee J C. Infect Immun. 1991;59:1008–1014. doi: 10.1128/iai.59.3.1008-1014.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Ton-That H, Mazmanian S K, Schneewind O. Trends Microbiol. 2001;9:101–102. [Google Scholar]
- 38.McDevitt D, Francois P, Vaudaux P, Foster T J. Mol Microbiol. 1994;11:237–248. doi: 10.1111/j.1365-2958.1994.tb00304.x. [DOI] [PubMed] [Google Scholar]
- 39.Uhlén M, Guss B, Nilsson B, Gatenbeck S, Philipson L, Lindberg M. J Biol Chem. 1984;259:1695–1702. , and correction (1984) 259, 13628. [PubMed] [Google Scholar]
- 40.Signas C, Raucci G, Jonsson K, Lindgren P-E, Anantharamaiah G M, Hook M, Lindberg M. Proc Natl Acad Sci USA. 1989;86:699–703. doi: 10.1073/pnas.86.2.699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Neu H C. Science. 1992;257:1064–1073. doi: 10.1126/science.257.5073.1064. [DOI] [PubMed] [Google Scholar]
- 42.Sieradzki K, Roberts R B, Haber S W, Tomasz A. N Engl J Med. 1999;340:517–523. doi: 10.1056/NEJM199902183400704. [DOI] [PubMed] [Google Scholar]