Abstract
Reverse genetics is a powerful tool for understanding gene functions and their interactions in the mosquito innate immunity. We took the transgenic approach, in combination with the RNA interference (RNAi) technique, to elucidate the role of mosquito REL1, a homolog of Drosophila Dorsal, in regulation of Toll immune pathway in the mosquito Aedes aegypti. By transforming the mosquitoes with ΔREL1-A or a double-stranded RNA construct of REL1 driven by the female fat body-specific vitellogenin (Vg) promoter with the pBac[3xP3-EGFP, afm] vector, we generated two different transgenic mosquito strains, one with overexpressed AaREL1 and the second with AaREL1 knockdown. Both strains had a single copy of the respective transgene, and the expression in both transgenic mosquitoes was highly activated by blood feeding. Vg-ΔREL1-A transgenic mosquitoes activate Toll immune pathway in the fat body by blood feeding. The overexpression of both isoforms, AaREL1-A and AaREL1-B, in Vg-ΔREL1-A transgenic mosquitoes resulted in the concomitant activation of Aedes Spätzle1A and Serpin-27A, independent of septic injury. The same phenotype was observed in the mosquitoes with RNAi knockdown of an Aedes homolog to Drosophila cactus, an IκB inhibitor of Drosophila Toll pathway. The effect of the transgenic RNAi knockdown of AaREL1 on mosquito innate immunity was revealed by increased susceptibility to the entomopathogenic fungus Beauveria bassiana and the reduced induction of Spz1A and Serpin-27A gene expression after fungal challenge. These results have proven that AaREL1 is a key downstream regulator of Toll immune pathway in the mosquito A. aegypti.
Keywords: REL1, cactus, spätzle1A, serpin-27A
Mosquitoes transmit numerous human diseases, including malaria, which is responsible for >2 million deaths per year (1, 2). Lymphatic filariasis, a nematode-based disease transmitted by mosquitoes, affects millions of people in tropical regions of the world (3). Diseases caused by Dengue fever are reaching disastrous levels in Central and South America and Southeast Asia (4). The West Nile encephalitis virus has killed hundreds of people in even the most developed countries such as the United States (5). This catastrophic situation indicates that there is an urgent need to explore every possible avenue for developing novel control strategies against these mosquito-borne diseases.
Innate immunity plays a pivotal role in the interaction between a pathogen and its mosquito host. In Anopheles gambiae, 242 genes from 18 gene families have been implicated in innate immunity by comparative genomic analysis of the Drosophila immune systems (6). The most recent data have clearly implicated several mosquito immune genes in parasite lysis and melanization (7). The silencing of leucine-rich repeat immune protein 1, which was shown to be up-regulated by bacterial challenge (8), causes a substantial increase in the number of Plasmodium oocysts (9). The knockdown of the TEP1 gene, which encodes a protein that has been shown to be involved in the phagocytosis of bacteria (10), greatly increases the number of developing oocysts (11). A recent study demonstrated that the mosquito strain refractory to parasite transmission had an elevated level of reactive oxygen species as another factor contributing to parasite melanotic encapsulation (12).
In Drosophila, induction of the Toll pathway leads to the activation of two NF-κB proteins, Dorsal and Dorsal-related immune factor (Dif), which results in the production of antifungal peptides such as drosomycin. The IMD pathway requires another NF-κB protein, Relish, which induces the expression of genes encoding antibacterial peptides such as diptericin and attacin. Microarray studies have indicated that, in addition to AMP genes, many other genes are markedly up-regulated as a consequence of the challenge-dependent activation of Toll and/or IMD pathway, including those encoding recognition molecules such as peptidoglycan-recognition protein and Gram-negative bacteria-binding protein (13). These recognition molecules are involved in the IMD and Toll pathways; they are also implicated in the protease cascade of the phenoloxidase or blood clotting pathways.
The Drosophila Toll pathway is involved in many aspects of the immune response. In addition to activating antifungal defenses, this pathway is also required for survival after some Gram-positive bacteria infections (14). Drosophila Toll pathway is also involved in cellular immune response against invaders and in humoral antifungal response. Drosophila mutants that constitutively activate the Toll pathway, such as necrotic, Toll10b, and cactus, all show a hemocyte phenotype with overreactive blood cells that form melanotic masses (15), similar to the cellular response to parasites (16).
In Drosophila Toll pathway, signaling from the membrane receptor, Toll, activates two closely related NF-κB proteins: Dif in adults, and Dorsal and/or Dif in larvae in Drosophila. In the naïve state, both Dorsal and/or Dif are retained in inactive forms in the cytoplasm by the inhibitory protein, cactus, which is a member of the I-κB family of NF-κB inhibitors. Toll signaling activates the phosphorylation of cactus, resulting in ubiquitination and proteasome-dependent degradation of cactus. Dorsal and/or Dif then translocate to the nucleus, where they activate the transcription of target genes. The remarkable structural and functional similarities in the mechanisms of activation of Dorsal during Drosophila morphogenesis and of NF-κB during the mammalian acute phase response have implicated Dorsal and the related protein, Dif, in the host defense of Drosophila (17, 18). However, the analyses of Drosophila Dif/Dorsal double mutants have revealed that Dif is the essential regulator of the Drosophila immune pathway in adult flies (19). Dif mediates the Toll-dependent control of inducible Drosomycin expression, and Dif alone is required for the antifungal response in adult flies. Dif ortholog has not been found in mosquitoes. The published genome sequence of the mosquito A. gambiae harbors only two NF-κB genes, REL1 (Gambf1) and REL2 (Relish), which are homologues of Drosophila Dorsal and Relish (6), respectively. Our previous study has shown that Aedes aegypti REL1 regulates antifungal immune response and functions as a Dif analog of the Toll-mediated antifungal immune pathway in adult female mosquitoes (20).
Reverse-genetic analyses, particularly the transgenic and RNA interference (RNAi) techniques, will now be necessary to provide an understanding of gene functions and their interactions in the mosquito innate immunity. We previously reported on the transgenic mosquitoes with stable, dominant negative immune-deficient phenotype for AaREL2 (Relish), which showed a marked susceptibility to Gram-negative bacteria infection and severely compromised induction in the expression levels of both defensin A (Def A) and cecropin A (Cec A) (21). Here, we constructed two different transgenic mosquito strains with overexpression or double-stranded RNA (dsRNA)-based knockdown of REL1, respectively, by combining a transgenic approach and RNAi techniques to elucidate the regulatory role of REL1 in more detail. In addition, we compared the immune phenotypes between these transgenic mosquitoes and the knockdown of a Toll immune pathway component gene, Aedes Cactus, showing that mosquito Toll immune pathway participates in the REL1-mediated regulation of mosquito immune genes and antifungal immune response.
Materials and Methods
Transformation Vector Construct and Germ-Line Transformation. The pBac[3xP3-EGFP, afm] transformation vector containing an eye-specific promoter (3xP3), in front of a TATA box and enhanced GFP, was described by Horn and Wimmer (22). A 1.5-kb size of ΔREL1-A lacks the major part of C terminus, which constitutes the glutamine-rich domain. The deletion of AaREL1-A from P452 to F566 were introduced by PCR-based mutagenesis (Pf primer, 5′-AGTAGATCTGTGCCCCCCAGCAGG; Pr primer, 5′-CATAGATCTGGAGGCCGAAACCCTC). The Vg-ΔREL1-A construct was made by linking the vitellogenin (Vg) promoter, ΔREL1-A, and SV40 polyadenylation element. Two 626-bp mosquito AaREL1-A fragments encoding the N-terminal region and a part of the Rel homology domain were obtained by PCR through the use of two pairs of primers [P1f primer, 5′-CGATCTAGACGAAAAGAAAGCCAC (XbaI); P1r primer, 5′-CATGGATCCGGAACACCTGGAAGC (BamHI); P2f primer, 5′-CGACTCGAGCGAAAAGAAAGCCAC (XhoI); P2r primer: 5′-CATCTGCAGGGAACACCTGGAAGC (PstI)] with the different restriction site sequences at either end, and these sites were used for generation of Vg-iREL1 transgenic mosquitoes. Each of the two fragments was then sequentially inserted into the corresponding restriction sites. The Vg-iREL1 construct was made by linking the Vg promoter with two copies of the 626-bp REL1 fragment, arranged as a head-to-head inverted repeat with a 0.2-kb intron spacer. These Vg-ΔREL1-A and Vg-iREL1 were introduced into the pBac[3xP3-EGFP, afm] plasmid at the AscI unique cloning site, respectively (Fig. 1 A and B). The resulting plasmids, pBac[3xP3-EGFP, afm, Vg-ΔREL1-A] and pBac[3xP3-EGFP, afm, Vg-iREL1] were used for injection of A. aegypti embryos, respectively. A. aegypti germ-line transformation was performed as described by Kokoza et al. (23).
Fig. 1.
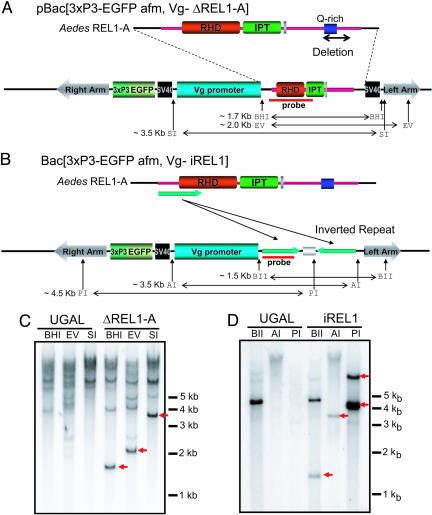
Construction of two different transgenic mosquitoes with altered REL1 activity. (A and B) Schematic diagrams of the pBac[3xp3-EGFP, afm, Vg-ΔREL1-A] and the pBac[3xp3-EGFP, afm, Vg-iREL1] transformation vector that were respectively transformed into the A. aegypti germ line. (C and D) Southern blot analyses of genomic DNA extracted from the Vg-ΔREL1-A transgenic mosquitoes, the Vg-iREL1 transgenic mosquitoes, and the parental UGAL strain, digested with BamHI (BHI), EcoRV (EV), SalI (SI), BglII (BII), ApaI (AI), and PstI (PI). The additional bands, indicated as arrows, were detected in the transgenic mosquitoes. The probe region and the restriction sites of the transformation vector are indicated in A and B.
Synthesis and Microinjection of dsRNA. Synthesis of dsRNA was accomplished by simultaneous transcription of both strands of template DNA with a HiScribe RNAi Transcription Kit (NEB, Beverly, MA). The treatment and injection into mosquito of the RNA samples were performed as described by Shin et al. (20).
Southern and Northern Hybridization. Genomic DNA from transgenic and control mosquitoes was purified by using the method of Bender et al. (24). Total RNAs from the previtellogenic and vitellogenic mosquitoes [at 12 and 24 h after blood meal (PBM)] were prepared by using the TRIzol method (Invitrogen) and used for the expression study of Vg-ΔREL1-A, Vg-iREL1, and tested immune genes. For Northern hybridization in response to septic injury, the vitellogenic mosquitoes at 24 h PBM were injected with a stationary-phase culture of Enterobacter cloacae or Micrococcus luteus, or the spores of Beauveria bassiana (5 × 107 per ml of viable spores), and collected at 5, 24, and 48 h after septic injury.
Quantitative Real-Time PCR Analysis. Total RNA of the mosquito fat bodies was prepared by using the TRIzol technique (Invitrogen). Contaminating genomic DNA was removed by treatment with RNase-free DNase I (Invitrogen). Reverse transcription was carried out by using an Omniscript reverse transcriptase kit (Qiagen) in a 20-μl reaction mixture, containing Oligo-dT primer and 2 μg of total RNA at 37°C for 1 h. Two microliters of cDNA from this reaction was subjected to real-time PCR, which was performed by using the iCycler iQ system (Bio-Rad). Reactions were performed in 96-well plates with a QuantiTect SYBR PCR kit (Qiagen), and each sample was analyzed in triplicate and normalized to the internal control, actin mRNA. The following primers were used for amplification and quantification of the expression of AaREL1-A and AaREL1-B genes: AaREL1-A forward primer, 5′-ATAGGCGAGATCAACATCAGCAGC; AaREL1-A reverse primer, 5′-CGTTGCTGTTCCTGCTTCATATCG; AaREL1-B forward primer, 5′-CGCCAAACGACCAACTACCGAATCC; AaREL1-B reverse primer, 5′-AGAGACAGATGAGTGCCGCTTCCG.
Septic Injury and Survival Experiments. Septic injury and survival experiments were performed in the same way as described in ref. 21. Septic injury was produced by pricking mosquitoes in the rear part of the abdomen with an acupuncture needle (0.20 × 25 mm) dipped into a concentrated culture of E. cloacae and M. luteus or a spore suspension (5 × 107 per ml of viable spores) of B. bassiana stain GHA. Three groups of 30 vitellogenic mosquitoes at 24 h PBM after a second blood feeding were challenged.
Results
Construction of Transgenic Mosquitoes with Altered Toll Immune Response. Drosophila dorsal, in similarity with mammalian NF-κB/REL genes, controls its own transcription (25). The Rel homology domain has been found to be important for transcriptional activation, in addition to being responsible for DNA binding and nuclear import (26). We designed transgenic mosquitoes to test whether AaREL1 Rel homology domain possessed similar properties and controlled transcription of the AaREL1 gene(s). To accomplish this experiment, we generated a mutagenized construct of AaREL1-A, ΔREL1-A, in which we deleted the C-terminal domain (CTD) that included a glutamine-rich region (Fig. 1 A). The CTD reportedly has dual motifs for the activation and repression (27). The ΔREL1-A construct was subcloned, together with the Vg promoter, to the pBac[3xP3-EGFP, afm] vector that was used for germ-line transformation of A. aegypti (Fig. 1 A). From 500 embryos injected with pBac[3xP3-EGFP, afm, Vg-ΔREL1-A], 68 survived into adulthood, and 1 yielded EGFP-positive progeny. A stable transgenic line was established from this progeny and characterized as containing the transgene in the mosquito genome (Fig. 1C). Southern blotting with probes from the left and right arms of the vector confirmed that this transgenic line had a single-copy insertion (data not shown). Northern blot analysis indicated that Vg-ΔREL1-A transcript was greatly induced by blood feeding in transgenic mosquitoes from 12 h to 24 h PBM (Fig. 2A). The endogenous REL1 transcripts that corresponded to both of the isoforms, REL1-A and REL1-B, were significantly elevated in these transgenic mosquitoes, showing that we obtained the transgenic mosquitoes with the overexpression phenotype of REL1 transcripts. These results demonstrated that AaREL1-A Rel homology domain was involved in transcriptional activation of both the AaREL1-A and AaREL1-B transcripts. However, a precise mechanism of its action remains to be resolved.
Fig. 2.
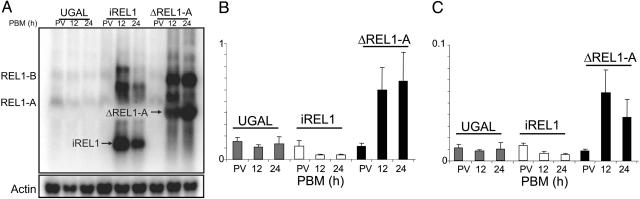
The expression profiles of Vg-ΔREL1-A and Vg-iREL1 transgene and transgenic effects on the endogenous REL1 gene expression. (A) Northern blot analysis showing the Vg promoter-mediated activation of the transgenes after blood feeding. Endogenous REL1 transcripts, REL1-A and REL1-B, were detected in the parental mosquitoes at any stage. The additional bands, which appeared after blood feeding, were detected in the Vg-ΔREL1-A and Vg-iREL1 transgenic mosquitoes, respectively. (B and C) Real-time PCR analysis showing the relative level of endogenous REL1 transcripts in transgenic and parental mosquitoes. After blood feeding, the Vg-ΔREL1-A transgenic mosquitoes showed the overexpression of REL1 transcripts, whereas the Vg-iREL1 transgenic mosquitoes showed the knockdown of both REL1 transcripts, REL1-A (B) and REL1-B (C). PV, previtellogenic.
We constructed the transgenic mosquitoes with impaired Toll-mediated systemic immunity by using RNAi techniques. A fold-back construct of the double-stranded AaREL1 RNA (iREL1) were inserted in front of the Vg promoter. This iREL1 construct consists of a 626-bp fragment from the N terminus region of AaREL1. This fragment was placed in the sense conformation followed by the same fragment inserted in the anti-sense conformation (Fig. 1B). An intron from A. aegypti clathrin heavy chain gene (28) was placed as a spacer. Introns have been shown to increase the effectiveness of these types of constructs, possibly because of the spliceosome facilitating the formation of the double-stranded RNAi structure (29, 30). When expressed, the homologous regions of the construct fold into a double-stranded structure. A Vg-iREL1 construct, made by linking the Vg promoter, was inserted into the pBac[3xP3-EGFP, afm] vector (Fig. 1B), and the transformation vector, pBac[3xP3-EGFP, afm, Vg-iREL1], was transformed into the germ line of mosquito A. aegypti embryos. From 400 embryos injected with pBac[3xP3-EGFP, afm, Vg-iREL1], 45 survived into adulthood and 1 yielded EGFP-positive progeny. The stable transgenic line from this progeny contained a single copy insertion into mosquito genome, which was confirmed by Southern blotting with probes from the left and right arms of the vector (Fig. 1D and data not shown).
Northern blot analysis showed that the transcript of iREL1 was significantly induced by blood feeding in the Vg-iREL1 transgenic mosquitoes. The appearance of two bands larger than REL1-A or REL1-B, respectively, interfered with the observation of whether endogenous REL1-A and/or REL1-B transcripts were knocked down by overexpression of effector dsRNA, iREL1. Indeed, real-time PCR showed that the expressions of both the REL1-A and REL1-B transcripts were knocked down by the overexpression of iREL1 (Fig. 2 B and C). In addition, Northern blot analysis with a probe from the left arm of the vector indicated that the two interfering bands were originated from the read-through of Vg-iREL1 transcripts (data not shown).
Aedes Spätzle1A and Serpin-27A Gene Expression in the Vg-ΔREL1-A Transgenic Mosquitoes. Drosophila immune genes dependent on a Toll pathway show a specific activation profile by fungal challenge. The microarray analysis shows that the Toll pathway controls most of the late genes induced by fungal infection (13). In our previous paper, we showed that Aedes Serpin-27A was specifically induced by fungal challenge and transcriptionally regulated by AaREL1 (20).
In Drosophila, Toll activation requires the proteolytic processing of the secreted polypeptide spätzle (Spz), a member of the cysteine-knot family of growth factors, and the cleaved form of this molecule acts as a ligand for Toll. A recent study showed that the direct interaction of Drosophila Spz with Toll receptor established Toll signaling (31). Based on the partial nucleotide sequence published in the The Institute for Genomic Research A. aegypti gene index (The Institute for Genomic Research Gene Index Databases, The Institute for Genomic Research, Rockville, MD, www.tigr.org/tdb/tgi), a 195-bp DNA fragment was obtained by RT-PCR from A. aegypti mRNA. The complete 1,613-bp-long transcript, which has a 5′ UTR and 3′ UTR, including a poly (A) sequence, was obtained by 5′-RACE and 3′-RACE PCR. The 260-residue protein encoded by this transcript, so-called Aedes Spz1A, contained a C-terminal domain highly homologous to the cysteine-knot domain of the Drosophila Spz (32). We tested the expression profiles of Aedes Spz1A in A. aegypti mosquitoes after infection with Gram-negative bacteria (E. cloacae), Gram-positive bacteria (M. luteus), and fungal spores (B. bassiana). The specific response of A. aegypti Spz1A was elicited only by the fungal challenge (data not shown), the same response as that of Aedes Serpin-27A, which was shown to be regulated by Aedes REL1 NF-κB transcription factor (20).
We used both Aedes Spz1A and Serpin-27A gene expression to characterize the Vg-ΔREL1-A transgenic mosquitoes. These transgenic mosquitoes, which overexpressed both REL1-A and REL1-B transcripts (Fig. 2), showed greatly increased transcripts of Aedes Spz1A and Serpin-27A after blood feeding, indicating that the increase in AaREL1 transcripts had an effect on that of the Aedes Spz1A and Serpin-27A transcripts (Fig. 3A). The expression levels of both immune genes by blood feeding were similar to those after septic injury with the mixture of E. cloacae and M. luteus, showing that the Vg-ΔREL1-A transgenic mosquitoes could fully activate these immune genes without immune challenge (Fig. 3B). These results that overexpressed AaREL1 in the Vg-ΔREL1-A transgenic mosquitoes activated Aedes Spz1A and Serpin-27A, which are specifically activated by fungal challenge, clearly confirm that AaREL1 is a key transcriptional regulator of mosquito antifungal immune response.
Fig. 3.
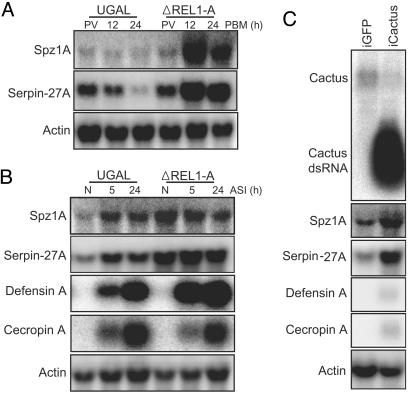
The effects of the transgenic Vg-ΔREL1-A and Cactus dsRNA treatments on the expression of Aedes Spz1A and Serpin-27A. (A) Northern blot analysis showing the activation of Aedes Spz1A and Serpin-27A after blood feeding in Vg-ΔREL1-A transgenic mosquitoes. (B) The expression profiles of the Aedes Spz1A and Serpin-27A 5 h and 24 h after septic injury in the Vg-ΔREL1-A transgenic mosquitoes and the parental UGAL strain 24 h PBM. (C) The increased expression of the Aedes Spz1A and Serpin-27A genes in Cactus dsRNA-treated mosquitoes (iCactus). GFP dsRNA-treated mosquitoes (iGFP) were used as a negative control. PBM (h), hours after blood meal; ASI (h), hours after septic injury.
Toll Immune Pathway in the Regulation of Mosquito Immune Genes. To verify whether AaREL1 is directly involved in mosquito Toll immune pathway, we tested the gene expression of Aedes Spz1A and Serpin-27A in Cactus dsRNA-treated mosquitoes. The dsRNA complementary to the A. aegypti homologue (The Institute for Genomic Research A. aegypti Gene Index, TC47332) of Drosophila cactus, an inhibitor κB of both Dif and Dorsal, was synthesized in vitro and injected into the thorax of newly emerged female mosquitoes. When Aedes Cactus dsRNA was introduced into the mosquitoes, the mRNA level of Aedes Cactus greatly decreased (Fig. 3D). The mRNA quantities of Aedes Spz1A and Serpin-27A in Cactus dsRNA-treated mosquitoes significantly increased, suggesting that AaREL1, which was freed in the cactus knockdown mosquitoes, could activate these immune genes without any challenge (Fig. 3C). This gene expression profile, which was similar to that in the Vg-ΔREL1-A transgenic mosquitoes, confirmed that the phenotype observed in the Vg-ΔREL1-A transgenic mosquitoes was the result of overexpressed AaREL1-A and AaREL1-B.
Interestingly, we found weak activation of the Def A and Cec A genes in the Cactus dsRNA-treated mosquitoes (Fig. 3C). The same levels of Def A and Cec A gene expression were observed in the Vg-ΔREL1-A transgenic mosquitoes at 24 h PBM without septic injury (data not shown), indicating that the mosquito Toll immune pathway could affect the activation of the Def A and Cec A genes and Spz1A and Serpin-27A. However, the expression levels of the Def A and Cec A genes in the Vg-ΔREL1-A transgenic mosquitoes by blood feeding were incomparably weaker than those by septic injury, whereas Spz1A and Serpin-27A genes were fully activated in the Vg-ΔREL1-A transgenic mosquitoes by blood feeding (Fig. 3B). These results suggest that Toll immune pathway may weakly affect the regulation of Def A and Cec A genes, whereas the Imd/Relish pathway mainly regulates the expression of these immune genes (21).
The Increased Susceptibility to B. bassiana and the Reduced Immune Activation of Spz1A and Serpin-27A Genes in the Vg-iREL1 Transgenic Mosquitoes. We applied a survival test to examine the effect of transgenic alteration of Aedes REL1 on mosquito immunity. The fungal susceptibility of the Vg-ΔREL1-A and Vg-iREL1 transgenic mosquitoes was compared with wild-type UGAL after the challenge of B. bassiana spores at 24 h PBM, which showed the REL1 knockdown in the Vg-iREL1 transgenic mosquitoes and the REL1 overexpression in the Vg-ΔREL1-A transgenic mosquitoes (Fig. 2). The Vg-iREL1 transgenic mosquitoes after a second blood feeding were significantly more susceptible to the fungal infection than wild-type and Vg-ΔREL1-A transgenic mosquitoes, when we preactivated the Vg-iREL1 expression by a first blood feeding (Fig. 4). In contrast, we did not find any increased susceptibility to three Gram-positive bacteria (M. luteus, Staphylococcus aureus, and Bacillus subtilis) tested in the Vg-iREL1 transgenic mosquitoes (data not shown). To address the question of whether Def A could play any role in the antifungal defense, we used the Vg-Def A transgenic mosquitoes (23). We injected REL1 and control dsRNAs (Mal, REL2) into Vg-Def A transgenic and wild-type mosquitoes and tested them for fungal susceptibility. REL1 dsRNA-treated, blood-fed Vg-Def A transgenic and wild-type mosquitoes exhibited similar increased susceptibility to fungal infection in contrast to mosquitoes treated with control dsRNAs (data not shown). This experiment demonstrated that Def A is not involved in antifungal defense in A. aegypti.
Fig. 4.
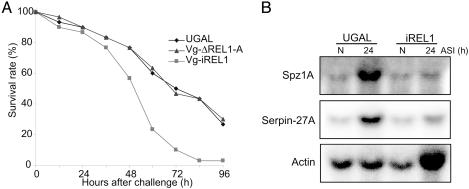
The increased susceptibility to B. bassiana (A) and the reduced immune activation of Spz1A and Serpin-27A genes in the Vg-iREL1 transgenic mosquitoes (B). (A) The Vg-ΔREL1-A and Vg-iREL1 transgenic and wild-type UGAL mosquitoes were blood fed at 2-3 days after emerging, forced to lay eggs at 3 days PBM, and blood fed again the next day. At 24 h PBM, B. bassiana spores were challenged. The result is representative of three independent experiments. (B) The Vg-iREL1 transgenic and wild-type UGAL mosquitoes were forced to lay eggs at 3 days after the second blood feeding and then challenged with B. bassiana spores the next day. The immune activation of Spz1A and Serpin-27A were impaired in fat bodies of the Vg-iREL1 transgenic mosquitoes. ASI (h), hours after septic injury.
To test the effects of transgenic REL1 knockdown on Spz1A and Serpin-27A gene expression, the mosquitoes were forced to lay eggs at 3 days after the second blood feeding and were then challenged with B. bassiana spores the next day. The knockdown of REL1 transcripts in the Vg-iREL1 transgenic mosquitoes (Fig. 2 B and C) negatively affected the induction of Spz1A and Serpin-27A gene expression in mosquito fat bodies (Fig. 4B), confirming that AaREL1 is involved in the induction of the Aedes Spz1A and Serpin-27A genes.
Discussion
We have investigated the contribution of REL1 in the mosquito Toll immune pathway by combining the transgenic and RNAi approaches. The overexpression of REL1 in the Vg-ΔREL1-A transgenic mosquitoes resulted in the activation of the downstream immune genes Aedes Spz1A and Serpin-27A. The expression levels of these genes elevated by blood feeding of the Vg-ΔREL1-A transgenic mosquitoes is similar to those activated by fungal challenge, indicating involvement of AaREL1. The fungal-specific activation of the immune genes, and the involvement with the antifungal immune response, are manifestations showing that AaREL1 is the key transcriptional regulator in the mosquito Toll immune response, much like Dif in the Drosophila Toll immune response (20). The fungal-specific induction of Drosomycin expression and antifungal activity against B. bassiana are characteristics of the Drosophila Toll immune pathway (14).
In the Drosophila loss-of-function mutant of cactus, which is a Drosophila I-κB inhibitor specific to Toll pathway, the Drosomycin genes are constitutively transcribed (33, 34), and the overexpression of Drosophila Dorsal could fully activate the Drosomycin expression without immune challenge (35). Here, we showed that the expressions of the Aedes Spz1A and Serpin-27A genes were constitutively activated in the RNAi-mediated knockdown of a mosquito cactus and that this phenotype was similar to that found in Vg-ΔREL1-A transgenic mosquitoes. These experiments clearly demonstrate the interaction of REL1 with cactus as a specific NF-κB transcription factor of the mosquito Toll immune pathway. In turn, this finding indicates conservation of the Toll immune pathway in the fat bodies of A. aegypti mosquito and Drosophila.
Examination of the Vg-ΔREL1-A transgenic Aedes mosquitoes has revealed that their PBM phenotype is the dramatic increase of REL1-A and REL1-B transcripts as well as the overexpression of the Serpin-27A and Spz1A genes without immune challenge. It is likely that the translated product of Vg-ΔREL1-A transgene does not directly activate these immune genes, because in vitro study by using a mosquito cell line showed that both REL1-A and REL1-B isoforms were necessary to fully activate the Aedes Serpin-27A genes (20). A number of studies pointed to different ways in which rel genes were autoregulated. Dorsal in Drosophila, like other rel proteins, was shown to regulate the level of its own transcription (25). We speculate that overexpression of ΔREL1-A in transgenic Aedes mosquitoes results in the blocking of the autoregulation of the REL1 gene that, in turn, leads to elevated expression of the REL1 isoforms REL1-A and REL1-B and consequently Serpin-27A and Spz1A.
We have been able to generate a transgenic mosquito with heritable dsRNA with a clear knockdown phenotype. The knockdown of REL1 transcripts in the Vg-iREL1 transgenic mosquitoes showed significantly increased susceptibility to fungal challenge and the reduced induction of Aedes Spz1A and Serpin-27A gene expression, the same phenotypes as the REL1 dsRNA-treated mosquitoes (20). Successful transgenic RNAi, in which the dsRNA trigger is expressed from a transgene, has been reported in Caenorhabditis elegans (36), Drosophila (37), and Arabidopsis (38). This report shows successful results of a dsRNA-mediated heritable knockdown experiment in mosquitoes. The recent approach to generating mosquitoes that constitutively express an anti-DENV effector RNAi construct resulted in the failure to develop the expected phenotype (39). We have observed the knockdown of target transcripts, which resulted in the phenotypes of increased fungal susceptibility and reduced induction of the immune gene expression.
Earlier, an A. gambiae REL1, Gambif1, was characterized, although no link to the Toll pathway has been established (40). In this work, we have established that mosquito REL1 is a downstream mediator of the Toll immune pathway. In particular, we have demonstrated that Vg-ΔREL1 transgenic mosquitoes, which overexpressed REL1, showed the same expression pattern of Aedes Serpin-27A and Spz1A as did Cactus dsRNA-treated mosquitoes. Furthermore, we have established a mosquito transgenic strain with heritable dsREL1 with the characteristic phenotypes of the impaired Toll immune pathway. Development of heritable transgenic mosquitoes with modified functions of REL1 will serve to further elucidate the role of the systemic immunity regulated by the Toll pathway in mosquito-pathogen interaction.
Whereas in Drosophila both fungal and Gram-positive bacterial infections are primarily controlled by the Toll pathway (41), no immune responses against Gram-positive bacteria have been found in REL1 dsRNA-treated (20) or Vg-iREL1-transgenic Aedes mosquitoes (this study). Recently, it has been found that A. gambiae REL2 is involved in defense against a Gram-positive bacteria S. aureus (42). Taken together, data indicate significant differences in specificities of major immune pathways between mosquitoes and Drosophila.
This paper was submitted directly (Track II) to the PNAS office.
Abbreviations: Dif, Dorsal-related immune factor; dsRNA, double-stranded RNA; RNAi, RNA interference; PBM, after blood meal; Vg, vitellogenin.
References
- 1.Beier, J. C. (1998) Annu. Rev. Entomol. 43, 519-543. [DOI] [PubMed] [Google Scholar]
- 2.Bremen, J. (2001) Am. J. Trop. Med. Hyg. 64, Suppl., 1-11. [Google Scholar]
- 3.Beaty, B. J. (2000) Proc. Natl. Acad. Sci. USA 97, 10295-10297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Wattam, A. R. & Christensen, B. M. (1992) Proc. Natl. Acad. Sci. USA 89, 6502-6505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Enserink, M. (2002) Science 297, 1988-1989. [DOI] [PubMed] [Google Scholar]
- 6.Christophides, G. K., Zdobnov, E., Barillas-Mury, C., Birney, E., Blandin, S., Blass, C. Brey, P. T., Collins, F. H., Danielli, A., et al. (2002) Science 298, 159-165. [DOI] [PubMed] [Google Scholar]
- 7.Meister, S., Koutsos, A. C. & Christophides, G. K. (2004) Int. J. Parasitol. 34, 1473-1482. [DOI] [PubMed] [Google Scholar]
- 8.Dimopoulos, G., Christophides, G. K., Meister, S., Schultz, J., White, K. P., Barillas-Mury, C. & Kafatos, F. C. (2002) Proc. Natl. Acad. Sci. USA 99, 8814-8819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Osta, M. A., Christophides, G. K. & Kafatos, F. C. (2004) Science 303, 2030-2032. [DOI] [PubMed] [Google Scholar]
- 10.Levashina, E. A., Moita, L. F., Blandin, S., Vriend, G., Lagueux, M. & Kafatos, F. C. (2001) Cell 104, 709-718. [DOI] [PubMed] [Google Scholar]
- 11.Blandin, S., Shiao, S. H., Moita, L. F., Janse, C. J., Waters, A. P., Kafatos, F. C. & Levashina, E. A. (2004) Cell 116, 661-670. [DOI] [PubMed] [Google Scholar]
- 12.Kumar, S., Christophides, G. K., Cantera, R., Charles, B., Han, Y. S., Meister, S., Dimopoulos, G., Kafatos, F. C. & Barillas-Mury, C. (2003) Proc. Natl. Acad. Sci. USA 100, 14139-14144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.De Gregorio, E., Spellman, P. T., Tzou, P., Rubin, G. M. & Lemaitre, B. (2002) EMBO J. 21, 2568-2579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Brennan, C. A. & Anderson, K. V. (2004) Annu. Rev. Immunol. 22, 457-483. [DOI] [PubMed] [Google Scholar]
- 15.Qiu, P., Pan, P. C. & Govind, S. (1998) Development (Cambridge, U.K.) 125, 1909-1920. [DOI] [PubMed] [Google Scholar]
- 16.Carton, Y. & Nappi, A. J. (2001) Immunogenetics 52, 157-164. [DOI] [PubMed] [Google Scholar]
- 17.Reichhart, J. M., Georgel, P., Meister, M., Lemaitre, B., Kappler, C. & Hoffmann, J. A. (1993) C. R. Acad. Sci. Ser. III 316, 1218-1224. [PubMed] [Google Scholar]
- 18.Ip, Y. T., Reach, M., Engstrom, Y., Kadalayil, L., Cai, H., Gonzales-Crespo, S., Tatei, K. & Levine, M. (1993) Cell 75, 753-763. [DOI] [PubMed] [Google Scholar]
- 19.Meng, X., Khanuja, B. S. & Ip, Y. T. (1999) Genes Dev. 13, 792-797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Shin, S. W., Kokoza, V., Bian, G., Cheon, H. M., Kim, Y. J. & Raikhel, A. S. (2005) J. Biol. Chem. 280, 16499-16507. [DOI] [PubMed] [Google Scholar]
- 21.Shin, S. W., Kokoza, V., Lobkov, I. & Raikhel, A. S. (2003) Proc. Natl. Acad. Sci. USA 100, 2616-2621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Horn, C. & Wimmer, E. A. (2000) Dev. Genes Evol. 210, 630-637. [DOI] [PubMed] [Google Scholar]
- 23.Kokoza, V., Ahmed, A., Wimmer, E. A. & Raikhel, A. S. (2001) Insect Biochem. Mol. Biol. 31, 1137-1143. [DOI] [PubMed] [Google Scholar]
- 24.Bender, W., Spierer, P. & Hogness, D. (1983) J. Mol. Biol. 168, 17-33. [DOI] [PubMed] [Google Scholar]
- 25.Lemaitre, B., Meister, M., Govind, S., Georgel, P., Steward, R., Reichhart, J. M. & Hoffmann, J. A. (1995) EMBO J. 14, 536-545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Jia, S., Flores-Saaib, R. D. & Courey, A. J. (2002) Mol. Cell. Biol. 22, 5089-5099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Flores-Saaib, R. D., Jia, S. & Courney, A. J. (2001) Development (Cambridge, U.K.) 128, 1869-1879. [DOI] [PubMed] [Google Scholar]
- 28.Kokoza, V. A. & Raikhel, A. S. (1997) J. Biol. Chem. 272, 1164-1170. [DOI] [PubMed] [Google Scholar]
- 29.Smith, N. A., Singh, S. P., Wang, M. B., Stoutjesdijk, P. A., Green, A. G. & Waterhouse, P. M. (2000) Nature 407, 319-320. [DOI] [PubMed] [Google Scholar]
- 30.Lee, Y. S. & Carthew, R. W. (2003) Methods 30, 322-329. [DOI] [PubMed] [Google Scholar]
- 31.Weber, A. N., Tauszig-Delamasure, S., Hoffmann, J. A., Lelievre, E., Gascan, H., Ray, K. P., Morse, M. A., Imler, J. L. & Gay, N. J. (2003) Nat. Immunol. 4, 794-800. [DOI] [PubMed] [Google Scholar]
- 32.Parker, J. S., Mizuguchi, K. & Gay, N. J. (2001) Proteins 45, 71-80. [DOI] [PubMed] [Google Scholar]
- 33.Levashina, E. A., Ohresser, D., Lemaitre, B. & Imler, J. L. (1998) J. Mol. Biol. 278, 515-527. [DOI] [PubMed] [Google Scholar]
- 34.Nicolas, E., Reichhart, J. M., Hoffmann, J. A. & Lemaitre, B. (1998) J. Biol. Chem. 273, 10463-10469. [DOI] [PubMed] [Google Scholar]
- 35.Manfruelli, P., Reichhart, J. M., Steward, R., Hoffmann, J. A. & Lemaitre, B. (1999) EMBO J. 18, 3380-3391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Tavernarakis, N., Wang, S.L., Dorovkov, M., Ryazanov, A. & Driscoll, M. (2000) Nat. Genet. 24, 180-183. [DOI] [PubMed] [Google Scholar]
- 37.Kennerdell, J. R. & Carthew, R. W. (2000) Nat. Biotechnol. 18, 896-898. [DOI] [PubMed] [Google Scholar]
- 38.Chuang, C.-F. & Meyerowitz, E. M. (2000) Proc. Natl. Acad. Sci. USA 97, 4985-4990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Travanty, E. A., Adelman, Z. N., Franz, A. W., Keene, K. M., Beaty, B. J., Blair, C. D., James, A. A. & Olson, K. E. (2004) Insect Biochem. Mol. Biol. 34, 607-613. [DOI] [PubMed] [Google Scholar]
- 40.Brillas-Mury, C., Charlesworth, A., Gross, I., Richamn, A., Hoffmann, J. A. & Kafatos, F. C. (1996) EMBO J. 15, 4691-4701. [PMC free article] [PubMed] [Google Scholar]
- 41.Hoffmann, J. A. (2003) Nature 426, 33-38. [DOI] [PubMed] [Google Scholar]
- 42.Meister, S., Kanzok, S. M., Zheng, X. L., Luna, C., Li, T. R., Hoa, N. T., Clayton, J. R., White, K. P., Kafatos, F. C., Christophides, G. K. & Zheng, L. (2005) Proc. Natl. Acad. Sci. USA 102, 11420-11425. [DOI] [PMC free article] [PubMed] [Google Scholar]