Abstract
Background
Coronary heart disease (CHD) remains a global health threat. As antiplatelet therapy constitutes the cornerstone of CHD management, ticagrelor has been universally endorsed as a first-line agent in major clinical guidelines. However, the therapeutic efficacy of ticagrelor is compromised by interindividual variability in bleeding risk. Notably, while inherited genetic variations account for part of this heterogeneity, the dynamic regulatory role of modifiable epigenetic mechanisms—particularly DNA methylation in mediating platelet reactivity—remains inadequately characterized, presenting a critical knowledge gap in optimizing precision antiplatelet strategies.
Methods
We utilized the 850k methylation array to measure DNA methylation levels in blood samples from 47 healthy controls and 93 patients with CHD. Subsequently, epigenome-wide association study (EWAS), summary data-based Mendelian randomization (SMR), and heterogeneity in dependent instruments (HEIDI) analyses were applied to pinpoint critical methylation sites that influence gene expression, platelet function recovery, and bleeding risk. After developing a targeted cellular model using the CRISPR-dCas9-DNMT3A/Tet1CD-U6-sgRNA system and integrating with transcriptomic sequencing data, we conducted mechanistic cellular experiments to elucidate how these methylation sites affect platelet function recovery and bleeding risk. The findings were further validated through animal studies.
Results
Integrated analysis of EWAS and SMR-HEIDI revealed that hypermethylation at CpG site cg03230175 within the GPD2 gene promoter region was significantly associated with decreased GPD2 gene expression (P = 1.76E-18), delayed platelet functional recovery (P = 9.02 × 10–3), and elevated hemorrhagic risk (P = 2.71 × 10–2). Transcriptomic studies indicated that GPD2 gene (cg03230175) methylation affects mitochondrial function, nuclear factor kappa B (NF-κB) signaling pathway, reactive oxygen species metabolic process, and G protein-coupled receptor (GPCR) ligand binding. Cellular experiments demonstrated that the GPD2 gene (cg03230175) methylation inhibits coagulation function by suppressing reactive oxygen species (ROS) production, NF-κB activation, and P2Y12 gene expression (P2Y12 receptor plays a pivotal role in platelet activation, thrombus formation, and the pathogenesis of thrombotic disorders). The animal study results confirmed that GPD2 enzyme inhibition can indeed prolong the clotting time in mice.
Conclusions
GPD2 gene (cg03230175) methylation resulted in reduced gene expression levels, inhibited mitochondrial energy metabolism, decreased ROS levels, and affected P2Y12 gene expression through the NF-κB pathway, ultimately leading to inhibition of coagulation function.
Registry: The Impact of Genotype on Pharmacokinetics and Antiplatelet Effects of Ticagrelor in Healthy Chinese (IGPPT). Trial registration number: NCT03092076. Date of Registration: 09 March 2017, retrospectively registered. URL of trial registry record: https://clinicaltrials.gov/ct2/show/NCT03092076.
Supplementary Information
The online version contains supplementary material available at 10.1186/s11658-025-00759-x.
Keywords: GPD2, DNA methylation, Epigenetic mechanism, ROS, NF-κB, P2Y12
Background
Coronary heart disease (CHD) poses a severe threat to public health. Antiplatelet therapy plays a crucial role in the treatment of CHD, effectively reducing the incidence of major cardiovascular events [1]. Ticagrelor is the first-line antiplatelet medication recommended by authoritative treatment guidelines [2], forming the basis of dual antiplatelet therapy following acute coronary syndrome (ACS) and percutaneous coronary intervention (PCI), and it is the preferred alternative for patients with resistance to clopidogrel [3–5]. Nevertheless, patients with CHD may be at elevated bleeding risk following administration of ticagrelor, and there is a notable discrepancy in the platelet function recovery [6–8]. The underlying mechanism remains uncertain. Therefore, elucidating the mechanism responsible for the interindividual variability in bleeding risk following ticagrelor therapy is of critical clinical significance and practical utility.
DNA methylation (DNAm) represents a significant epigenetic regulatory mechanism. Although it does not alter the DNA sequence, it has the capacity to influence gene expression and plays a significant role in the onset and progression of cardiovascular diseases. The levels of DNAm may be subject to alteration in response to external stimuli. The factor II receptor-like 3 (F2RL3) gene exhibits significantly demethylation in smokers, which is associated with increased platelet reactivity and an elevated risk of myocardial infarction [9]. Hypomethylation of the P2Y12 promoter has been linked to elevated platelet reactivity and an elevated risk of ischemic events [10]. In patients with serum albumin levels < 35 g/L, smokers or drinkers, lower promoter DNAm of the P2Y12 gene was associated with an increased risk of clopidogrel resistance [11]. Hypomethylation of the ABCB1 gene promoter has been linked to a reduction in the antiplatelet effects of clopidogrel [12] and aspirin [13]. Demethylation at the PON1 promoter has been linked to an increased bleeding risk following PCI in patients undergoing dual antiplatelet therapy with clopidogrel [14]. It is evident that DNAm has significant implications for antiplatelet therapy. The process of DNAm is catalyzed by the DNA methyltransferases (DNMTs), with DNMT1 and DNMT3A/B being the most significant [15]. The process of endogenous DNA demethylation is primarily facilitated by the ten_eleven translocation family proteins (Tets) [16]. Researchers have engineered the Cas9 protein within the CRISPR-Cas9 framework into a catalytically inactive form known as dead Cas9 (dCas9), thereby creating an innovative system for transcriptional regulation that facilitates epigenetic modifications at specific genomic sites without inducing DNA cleavage. By conjugating dCas9 with the catalytic domains of DNMTs or Tets and using sgRNA to guide it to the designated position, it becomes possible to selectively target promoter sequences for methylation or demethylation [17].
Our analysis revealed that methylation at cg03230175 site of the GPD2 gene promoter region was associated with an elevated bleeding risk and a prolonged recovery of platelet function following ticagrelor administration. GPD2 gene (position chr2:157291462) encodes glycerol-3-phosphate dehydrogenase 2, which is situated in the inner mitochondrial membrane and serves as a crucial enzyme in the glycerophosphate shuttle (GPS). This enzyme plays an integral role in glycolysis, aerobic sugar oxidation, and the electron transport chain involving coenzyme Q within mitochondria. It catalyzes the conversion of dihydroxyacetone to glycerol 3-phosphate while facilitating the reduction of NAD+ to nicotinamide adenine dinucleotide (NADH), concurrently transferring protons from intracellular NADH to flavine adenine dinucleotide (FADH2) and shuttling electrons to mitochondrial coenzyme Q [18]. The activation of GPS helps to alleviate NADH accumulation in cytoplasm, thereby reducing the glycolytic flux and ensuring that pyruvate is channeled towards maintaining the tricarboxylic acid cycle in the mitochondrial matrix following glycolysis, rather than being converted into lactic acid.
This study investigated how methylation of GPD2 gene (cg03230175) inhibits platelet activation—a process characterized by high energy demands where glucose metabolism and fatty acid metabolism are primary energy sources. During platelet activation, ATP generated through glycolysis is preferentially utilized; this phase also witnesses a slight increase in mitochondrial oxygen consumption alongside enhanced uptake of exogenous glucose [19]. Furthermore, research indicates that elevated levels of exogenous glucose significantly upregulate P2Y12 expression on platelets, contributing to their hyperreactivity [20]. GPD2 enhances the production of ROS in mitochondrial complex I via reverse electron transport (RET), while also facilitating RET by preserving the reduced state of the coenzyme Q pool [21–23]. RET is a specialized process that reduces NAD+ to NADH as electrons from panthenol are transferred back to respiratory complex I, resulting in substantial reactive oxyigen species (ROS) generation. These ROS may modulate P2Y12 receptor expression, which is closely associated with platelet reactivity, through Nuclear factor kappa B (NF-κB) [20].
ROS are crucial signal transduction molecules whose production is stimulated by cell metabolism or external factors and that play a pivotal role in regulating cell proliferation, differentiation, and death by modulating the redox state within cells [24].
NF-κB is a pivotal redox-sensitive transcription factor, a protein complex ubiquitous in animal cell types. It plays a crucial role in regulating cellular transcriptional function, inflammatory response, and stress response, with its activation influenced by intracellular redox levels, and it is intimately associated with the mediation of ROS [25]. A slight elevation in ROS levels facilitate NF-κB activation and anti-apoptosis, while an excessive increase in ROS levels prevent NF-κB activation and promote apoptosis.
P2Y12 is a G-protein-coupled receptor that is activated following the binding of adenosine diphosphate (ADP) and is predominantly expressed in platelets and other cells of the immune system. The P2Y12 receptor binds to the Gi protein, which results in the inhibition of adenylate cyclase and a reduction in the level of cyclic adenosine monophosphate (cAMP) in platelets. This initiates a cascade of signal amplification reactions, ultimately leading to a change in platelet activity [26]. The most important finding was the presence of a binding site for NF-κB in the promoter region of the P2Y12 gene [27].
Methods
The overall idea and process of the study are shown in Fig. 1. Should other researchers require it, we will make available more detailed data, analytical procedures, and research materials to enable the reproduction of the findings or the replication of the study process.
Fig. 1.
Flowchart of the study
Study population
This study was a single-center, open trial including 47 healthy subjects and 93 patients with CHD. The inclusion and exclusion criteria for healthy subjects and patients with CHD are provided in Supplemental Methods.
Blood samples were collected from healthy subjects at specified time points to measure the ADP-induced platelet aggregation rate. Patients with CHD were followed up at 6 and 12 months post-PCI, primarily for bleeding incidents and medication usage.
Blood samples’ collection and processing
All participants fasted for at least 8 h. In the early morning, 4 mL of peripheral venous blood was collected in a vacuum blood collection tube containing ethylenediaminetetraacetic acid (EDTA)-K2 anticoagulant. The blood was then centrifuged at 2095 × g for 10 min at 4 °C. After centrifugation, the upper plasma and lower blood cells were separately transferred into cryopreservation tubes, uniformly numbered and labeled, and stored in an ultralow-temperature freezer at −80 °C for future use.
DNA extraction from blood sample
We thawed the blood cell samples and the TGuide Blood Genomic DNA Extraction Kit at room temperature after removal from the −80 °C freezer and centrifuged for 10 s. We transferred 140 μL of blood cell sample into a 1.5-mL Eppendorf (EP) tube, added 60 μL of GA buffer and 10 μL of proteinase K solution, gently mixed, and centrifuged. Using the TGuide M16 automatic nucleic acid extraction system, we ran program 102 with a sample volume of 200 μL and an elution volume of 80 μL to extract genomic DNA.
DNA quality assessment
We used the Nanodrop 2000 spectrophotometer to measure the OD260/OD280 and OD260/OD230 optical density (OD) ratios to determine the DNA concentration. To ensure accuracy, we repeated the operation twice and calculated the average value. Additionally, we conducted a DNA purity test with the following steps:
(i) Preparation of 1 × Tris–acetate–EDTA (TAE) buffer: We diluted 50 × TAE buffer 50-fold to prepare 1 × TAE buffer for use. (ii) Preparation: We weighed out 1.0 g of agarose and added it to 100 mL of 1 × TAE buffer, then mixed well. We added 10 μl of GelRed dye and heated in a microwave until clear and transparent. (iii) Preparation of the agarose gel plate: We allowed the agarose gel solution to cool to 60–65 °C and poured it into the gel casting mold. We inserted the comb and waited for it to solidify. We placed the gel in the electrophoresis tank and added enough 1 × TAE buffer. (iv) Sample loading and electrophoresis: We mixed the DNA size marker with loading buffer and loaded it into each sample well. We set the voltage and time to complete the electrophoresis. After electrophoresis completed, we turned off the power. (v) Observation and photography: We removed the gel and placed it under a gel documentation system for observation and photography. We observed the DNA bands and judged their correctness on the basis of the marker. DNA with high purity and good integrity should display bright, single bands without smearing or trailing.
DNAm measurement
For the DNA samples from 47 qualified healthy subjects and 93 patients with CHD that passed quality control, the Infinium Methylation EPIC BeadChip (Illumina, USA) was used to assess the methylation levels at specific sites. This array encompasses over 850,000 methylation sites, covering a wide range of regions in the human genome including promoters, enhancers, transcription factor binding sites, and noncoding RNAs, providing important information about gene expression and regulation, cellular differentiation, and disease susceptibility. To minimize batch effects, samples were randomized before testing.
The data processing, quality control, and normalization details for the specific cohort are provided in the Supplementary Methods.
EWAS and SMR-HEIDI
RT50, defined as the time required for healthy subjects to recover to 50% of peak pharmacodynamic effect following ticagrelor administration, serves as a quantitative biomarker of platelet function restoration.
In the EWAS investigating RT50 in healthy subjects, we employed multivariable linear regression analysis to identify differentially methylated positions (DMPs), adjusting for sex and age as covariates. For nominally significant methylation sites, we further conducted multivariable logistic regression analyses to assess their associations with bleeding risk in patients with CHD. These analyses incorporated established bleeding risk factors from the PRECISE-DAPT score model post-PCI, including age, hemoglobin levels, white blood cell count, and creatinine clearance. Then, SMR-HEIDI analysis was conducted to screen out the key CpG sites that were significantly associated with gene expression, RT50 in healthy subjects, and the bleeding risk in patients with coronary heart disease.
The detailed analysis process can be found in the Supplementary Methods.
Cell culture
The GPD2 gene is highly expressed in platelets and megakaryoblast leukemia cells [28]. However, it is challenging to culture platelets in vitro, so we selected a human megakaryoblast leukemia cell line (MEG-01). MEG-01 cells, which can produce platelet-like particles (PLPs) under standard culture conditions, are a derivative of transformed bone marrow megakaryocytes derived from a 55-year-old male patient with chronic myeloid leukemia (CML) [29, 30].
MEG-01 cells (CL-0498) were kindly provided by Wuhan Pricella Biotechnology Co., Ltd., cultured in Roswell Park Memorial Institute (RPMI) 1640 medium supplemented with 10% fetal bovine serum (FBS) and 1% penicillin–streptomycin solution (P/S) and maintained in an incubator at 37 °C and 5% CO2 atmosphere.
Knockdown and overexpression of GPD2 gene
Small interfering RNAs (siRNAs) targeting the GPD2 gene for knockdown were synthesized by Guangzhou IGE Biotechnology Co., Ltd. When the MEG-01 cells reached approximately 70% confluence, the culture medium was replaced with OPTI-MEM I reduced serum medium. The siRNAs and Lipofectamine 3000 solutions were diluted separately according to the manufacturer’s instructions, mixed gently, and incubated at room temperature for 20 min. The mixture was then added to the cells and incubated for 6 h before the medium was replaced with fresh one. The cells were cultured for an additional 2 days.
Plasmids for overexpression of the GPD2 gene were synthesized by Guangzhou IGE Biotechnology Co., Ltd. When the cells reached approximately 70% confluence, the culture medium was replaced with antibiotic-free RPMI 1640 medium. The plasmid solution was diluted separately according to the manufacturer’s instructions, mixed gently, and incubated at room temperature for 20 min. The mixture was then added to the cells and incubated for 6 h before the medium was replaced with fresh one. The cells were cultured for an additional 2 days.
Methylation and demethylation of the GPD2 gene (cg03230175)
Lentiviruses containing the CRISPR-dCas9-DNMT3A/Tet1-U6-sgRNA system were constructed by Shanghai Genechem Co., Ltd. When the cells reached approximately 70% confluence, the culture medium was replaced with antibiotic-free RPMI 1640 medium. The lentivirus solution was diluted according to the manufacturer’s instructions and then gently mixed with the cells. After 4–6 h, the medium was replaced with fresh culture medium, and the cells were cultured for an additional 2 days.
The functional elements of the CRISPR-dCas9-DNMT3A/Tet1-U6-sgRNA constructs and their intended uses in this study are as follows:
CRISPR-dCas9: By targeting specific DNA sequences with the inactivated Cas9 protein (dCas9) and achieving precise regulation of gene expression with the aid of fused functional proteins, there is no need to cut DNA.
DNMT3A/Tet1: To achieve methylation/demethylation modification.
U6-sgRNA: To guide CRISPR-dCas9-DNMT3A/Tet1 to the designated position.
Regulatory elements: Puromycin resistance gene (for stable cell line selection).
Bisulfite sequencing polymerase chain reaction (BSP) assay
We engaged Guangzhou SaiCheng Co., Ltd. to perform a BSP assay to confirm the methylation status of the GPD2 gene (cg03230175). Comprehensive details of the methodology are described in the Supplementary Methods.
RNA sequencing
Total RNA from two groups of MEG-01 cells (control and methylation-treated groups; four replicates per group; eight samples in total) was extracted using TRIzol reagent (Invitrogen) following the manufacturer’s instructions. RNA quality and quantity were assessed using a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific). The RNA samples were then sequenced using NovaSeq6000 (Illumina, USA). RNA expression was quantified as transcripts per million (TPM) across all genes, followed by log2(TPM + 1) transformation for subsequent analysis.
Details for RNA sequencing are provided in the Supplementary Methods.
Real-time polymerase chain reaction
Primers for genes GPD2, P2Y12, CD62P, VAMP8, VASP, and β-actin were synthesized by Ruibiotech Co., Ltd. Total RNA was extracted from MEG-01 cells using TRIzol reagent (Invitrogen) and reverse-transcribed into complementary DNA (cDNA). Real-time polymerase chain reaction (PCR) was performed with the addition of the fluorescent dye SYBR Green I to the reaction mixture. Fluorescent signals during the PCR reaction were detected using a real-time PCR instrument. Quantification of target genes and the reference gene was carried out using the 2-ΔΔCT method.
Detailed primer sequences can be found in the Supplementary Methods.
Immunoblotting
Antibodies against GPD2, P2Y12, CD62P, VAMP8, VASP, β-actin, Histone H3, α-tubulin, NF-κB P65, p-P65, and p-IκBα were purchased from the MedChemExpress platform. Total protein was extracted from MEG-01 cells using radioimmunoprecipitation assay (RIPA) lysis buffer, and protein concentration was quantified by the bicinchoninic acid (BCA) method. A 6–15% resolving gel and a 5% stacking gel were prepared. After protein loading, the stacking gel was run at 70 V for 30 min, followed by continued electrophoresis at 120 V until the bromophenol blue dye reached an appropriate position in the resolving gel. Proteins were then transferred to a polyvinylidene fluoride (PVDF) membrane at a constant current of 230 mA for 100 min. The membrane was blocked with 5% nonfat milk at room temperature for 1 h and then incubated with the primary antibodies overnight at 4 °C with gentle agitation. The membrane was washed three times with Tris-buffered saline with Tween-20 (TBST) for 5 min each time before incubation with the secondary antibodies at room temperature for 1 h. After three additional washes with TBST, the membrane was incubated in the developing solution in the dark for 3 min. The protein bands were visualized using an imaging system, and intensities were analyzed using ImageJ software.
Flow cytometry
First, the stock solution of 2′,7′-dichlorodihydrofluorescein diacetate (DCFH-DA) was diluted 1:1000 in serum-free RPMI 1640 medium to prepare a working solution with a final concentration of 10 μM DCFH-DA. Subsequently, MEG-01 cells were collected into Eppendorf tubes and centrifuged at 1000 rpm for 5 min. After washing twice with PBS, 400 μL of the DCFH-DA working solution was added to the Eppendorf tubes, which were then placed in a 37 °C incubator for 30 min of dark cultivation. The tubes were inverted every 3–5 min to ensure thorough contact between the probe and the cells. Following incubation, the cells were washed three times with serum-free cell culture medium to remove any unincorporated DCFH-DA. Finally, the cell suspension was transferred to a flow cytometry tube and analyzed as soon as possible using a flow cytometer.
Platelet-like particles (PLPs) functional studies
Details of PLP experiments are described in the Supplementary Methods.
Chromatin immunoprecipitation (ChIP) assay
The ChIP assay was conducted to investigate NF-κB binding to the promoter region of the P2Y12 gene in MEG-01 cells. The primers necessary for the ChIP Assay were synthesized by Wuhan Boerfu Biotechnology Co., Ltd.
Detailed experimental procedures can be found in the Supplementary Methods.
Transmission electron microscopy experiments
After the intervention, MEG-01 cells were first harvested into 15-mL centrifuge tubes and centrifuged at 9000 g for a duration of 5 min. The supernatant was then carefully removed, and electron microscopy fixative was added delicately to the cell pellet, ensuring that the sample was protected from light during the 1-h fixation period at room temperature. Subsequently, the fixed cell pellet was gently resuspended in a minimal volume of fixative to achieve an even distribution of the sample. The resuspended cells were applied to a carbon-coated copper grid and left to adsorb for the appropriate duration. Excess fixative was subsequently blotted off with filter paper, and the prepared grid was then transferred to the transmission electron microscope for detailed observation.
Animal studies
Eight-week-old C57BL/6 wild-type mice, each weighing about 20 g, were purchased from Zhuhai BesTest Bio-Tech Co., Ltd. and raised in the specific-pathogen-free (SPF) environment of the Institute of Biological and Medical Engineering, Guangdong Academy of Sciences. The mice were randomly assigned to one of five groups and received daily injections of either saline, vehicle (dimethyl sulfoxide), KM04416 (20 mg/kg), ticagrelor (10 mg/kg), or a combination of KM04416 and ticagrelor. The injections were administered for a period of five consecutive days.
The small-molecule inhibitor KM04416 was purchased from the MCE platform, and its in vivo formulation was prepared according to the manufacturer’s protocol. The formulation comprises the following components:
Dimethyl sulfoxide (DMSO) serves as the primary solvent and penetration enhancer, effectively dissolving hydrophobic compounds and facilitating transmembrane delivery (with its concentration strictly maintained below 10% to mitigate cytotoxicity);
PEG300 enhances drug solubility through molecular encapsulation, prolongs release kinetics, and stabilizes molecular integrity, with its molecular weight modulating solution viscosity and osmotic pressure;
Tween-80 acts as a surfactant to reduce interfacial tension, thereby improving dispersion homogeneity, enhancing absorption efficiency;
Saline ensures physiological compatibility by matching the osmotic pressure of bodily fluids (~ 290 mOsm/L) and providing weak alkaline pH buffering, thereby minimizing injection-induced irritation and diluting the toxicity of organic solvents.
This formulation aligns with established pharmacokinetic optimization principles for small-molecule inhibitors in preclinical studies.
All animal experiments have been approved by the Ethics Review Committee of Guangdong Provincial People’s Hospital and comply with the guidelines for the care and use of laboratory animals by the National Institutes of Health of China.
Tail-clip bleeding time assay
Tail-clip bleeding time was measured by clipping 2 mm from the tip of the tail of mice. The tail was placed in prewarmed saline at 37 °C, and the time to cessation of bleeding was measured for 1000 s.
Activated clotting time (ACT) assay
Initially, we prepared three 1.5-mL EP tubes and labeled them as 1, 2, and 3. To each tube, we pipetted 0.1 mL of ACT kaolin solution and 0.1 mL of ACT cephalin solution, then gently mixed the contents. Following this, we obtained 1 mL of blood from the medial canthus of the mouse and added 0.3 mL of this blood to each tube, ensuring a gentle mix. We placed the tubes in a 37 °C water bath for 3 min, tilting tube 1 every 30 s until the blood remained stationary when the tube was inverted. After observing clot formation in tube 2, we monitored tube 3. The time at which clot formation occurred in tube 3 was noted as the activated clotting time.
Statistical analysis
Results are summarized as count (percentage) for categorical variables and mean ± standard deviation (SD) for continuous variables. Data were analyzed using an unpaired t-test for comparisons between means, one-way analysis of variance with Bonferroni correction for comparisons between > 2 groups, or two-way analysis of variance (ANOVA) followed by Bonferroni post test if necessary. Statistical analysis was performed using GraphPad Prism software (GraphPad Software, Inc., La Jolla, CA).
A statistically significant difference was present when p < 0.05. The following symbols are used to denote the level of statistical significance: *p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001.
Results
EWAS results identify the critical CpG site
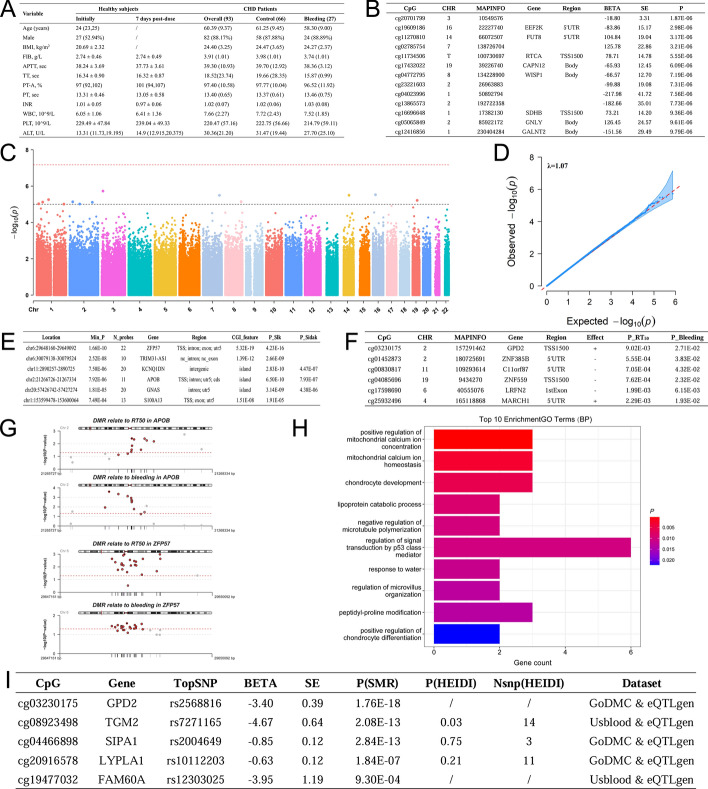
Figure 2A presents a baseline characteristics table for healthy individuals and patients with CHD.
Fig. 2.
The results of EWAS. A Baseline characteristics of the study population. B EWAS results for RT50 and annotation information. MAPINFO: chromosomal coordinates of the CpG (build 37); Region: gene region feature category describing the CpG position from UCSC; TSS200: 0–200 bases upstream of the transcriptional start site (TSS); TSS1500: 200–1500 bases upstream of the TSS, 5′UTR: within the 5′ untranslated region between the TSS and the ATG start site; Body: between the ATG and stop codon; irrespective of the presence of introns, exons, TSS, or promoters; 3′UTR: between the stop codon and poly A signal. C Manhattan plot of EWAS results for RT50. Each dot represents a CpG site. The red dashed line denotes the significance threshold after Bonferroni correction, which is P = 6.74 × 10−8, whereas the black dashed line signifies the threshold at P = 1 × 10−5. D QQ plots of EWAS results for RT50. Each dot represents a CpG site. E The DMR results of RT50 and annotation information. Slk_P: uncorrected Stouffer–Liptak–Kechris p values; Sidak_P: p values corrected for multiple testing are reported. F The CpG sites that are in the promoter region and have the same effect on RT50 and bleeding risk. P_RT50: p value for RT50; P_Bleeding: p value for bleeding risk. G DMRs associated with RT50 and bleeding risk. The red dashed line represents the threshold of nominal significance, p = 0.05. Each red dot indicates a CpG site within DMRs. The purple dots represent the CpG sites with the most significant associations with the phenotype within DMRs. H The top 10 GO enrichments of key CpG sites. A more intense red color indicates a smaller p-value. I The pleiotropic associations between CpG sites and annotated genes. P(SMR): p-value from SMR; P(HEIDI): p-value from HEIDI test; Nsnp(HEIDI): number of single-nucleotide polymorphisms (SNPs) used in the HEIDI test
To identify the key DNAm sites affecting RT50, an EWAS was performed. After cleaning the raw data, the remaining 741,318 CpG sites were analyzed using multiple linear regression with RT50 as the dependent variable. No statistically significant sites were found after Bonferroni correction (P < 6.74 × 10−8, 0.05/741,318). Functional annotations for sites meeting P < 1 × 10−5 are presented (Fig. 2B, C). The QQ plot indicates that the observed p-values for several sites in the tail are less than the theoretical p-values; however, this discrepancy is not significant (Fig. 2D). To delve into the DNAm characteristics affecting RT50, we performed a differentially methylated regions (DMR) analysis (Fig. 2E). After applying hypothesis testing corrections, 21 DMRs with P_Sidak (p value after Sidak correction) < 0.05 were detected. These DMRs encompassed between 2 and 22 methylation sites, with 10 DMRs located within CpG islands and 8 DMRs associated with promoter regions. The complete results are provided in Supplementary Table S1.
CpG sites with P_RT50 (p value for RT50) < 0.05 were then analyzed using multiple logistic regression, with bleeding risk as the dependent variable (Fig. 2F). A total of 194 sites in the promoter region, with P_Bleeding (p value for bleeding risk) < 0.05, aligning with the RT50 effect direction, are detailed in Supplementary Table S2. Then, we identified two DMRs associated with both bleeding and RT50, located on the APOB and ZFP57 genes, respectively (Fig. 2G). Subsequently, we conducted Gene Ontology (GO) analysis and identified the top 10 biological processes (Fig. 2H), which revealed that these CpG sites were mainly involved in positive regulation of mitochondrial calcium ion concentration (p = 2.36 × 10−4) and mitochondrial calcium ion homeostasis (P = 2.84 × 10−3). The complete results are provided in Supplementary Table S3.
Pleiotropy refers to the phenomenon in which a single gene or genetic variant influences multiple phenotypes. By using SMR-HEIDI tests, we performed a pleiotropic analysis to evaluate whether the 194 CpG sites share causal genetic variants with their cis-regulated genes (i.e., testing for shared causal variants influencing both DNAm and gene expressions). As a result, only 45 were matched with GWAS data in the meQTL (Godmc) and eQTL (eQTLgen) summary datasets. After Bonferroni correction (P < 0.001, 0.05/45), only five CpG sites showed pleiotropic associations with their corresponding gene expression levels (Fig. 2I). Interestingly, GPD2 gene (cg03230175) methylation was significantly negatively correlated with the gene expression and positively associated with both RT50 and bleeding risk.
The impact of epigenetic modifications of the GPD2 gene (cg03230175) on MEG-01 cells
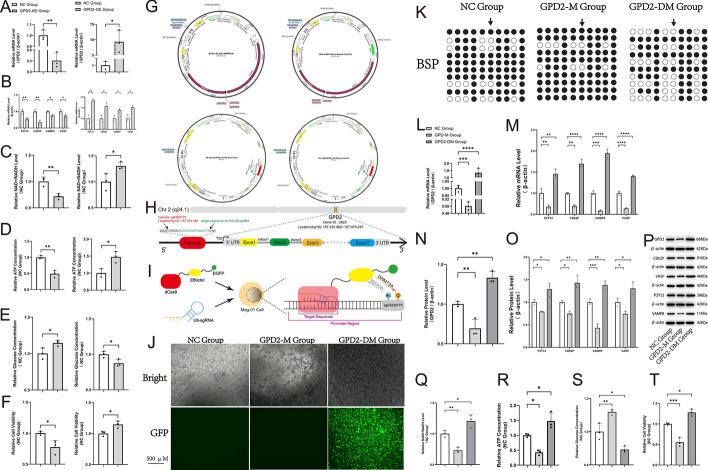
Initially, specific knockdown of the GPD2 gene (GPD2-KD group) was performed on MEG-01 cells (Fig. 3A). The results revealed a significant reduction in the expression levels of certain genes associated with coagulation function (Fig. 3B). Among these, P2Y12 is the target for ticagrelor’s antiplatelet effect, and the decreased expression of its gene directly indicates a reduced reactivity of platelets [31]. CD62P is a glycoprotein on the cell membrane surface on which platelet activation depends and plays a key role in the coagulation process [32]; VAMP8 is a membrane protein that facilitates platelet granule exocytosis, and the microRNA regulating VAMP8 expression is associated with high platelet reactivity [33]; VASP is a regulator of platelet aggregation, involved in platelet aggregation and adhesion to the vascular wall [34]. The levels of these three markers can also indirectly reflect the reactivity of platelets. Additionally, the NAD+/NADH ratio and cell viability also decreased (Fig. 3C, F). After 48 h of culture, the intracellular ATP concentration in the GPD2-KD group was significantly lower than that in the negative control (NC) group, while the residual glucose concentration in the culture medium was significantly higher (Fig. 3D, E). Conversely, the results observed in the GPD2 gene-specific overexpressing group (GPD2-OE group) exhibited a contrasting pattern.
Fig. 3.
Upon knockdown, overexpression, methylation, or demethylation of the GPD2 gene, significant alterations were observed in indices related to coagulation function and energy metabolism. A The relative expression level of the GPD2 gene. B The relative expression levels of genes associated with clotting. C The relative NAD+/NADH levels. D The relative intracellular ATP concentration. E The relative glucose concentration in medium. F The relative cell viability. G Schematic diagram of the CRISPR-dCas9-DNMT3A/TET1CD-U6-sgRNA system. H Schematic diagram showing the U6-sgRNA recognition position in the GPD2 gene promoter. I Schematic description of targeted methylation or demethylation via the CRISPR-dCas9-DNMT3A/TET1CD-U6-sgRNA system. J Fluorescence microscopy images. Cells emitting green fluorescence observed under green fluorescent protein (GFP) mode are deemed successfully infected. K Bisulfite sequencing. Hollow circles represent nonmethylated CpG, whereas solid circles represent methylated CpG motif. The black arrow points to the cg03230175 site. Each row represents an individual clone. L, M The relative messenger RNA (mRNA) levels of GPD2 gene and indicators associated with clotting. N–P The relative protein levels of GPD2 and indicators associated with clotting after methylation or demethylation of GPD2 gene. Q The relative NAD+/NADH level. R The relative intracellular ATP concentration. S The relative glucose concentration in medium. T The relative cell viability
Subsequently, two groups of MEG-01 cells were subjected to methylation (GPD2-M group) and demethylation (GPD2-DM group) treatments, respectively. The principles of these treatments are shown in Fig. 3G–I. Under bright-mode fluorescence microscopy (Fig. 3J), we observed that the cell count in the GPD2-M group was significantly lower compared with the NC group, while that in the GPD2-DM group was significantly higher; since only the vector without GFP was used for infection, no fluorescence was detected in the NC and GPD2-M groups under the GFP mode. Bisulfite sequencing results confirmed the successful specific methylation and demethylation of the GPD2 gene (cg03230175) in MEG-01 cells (Fig. 3K); the detection fragment designed by this method is approximately 234 bp in length.
Correlating these epigenetic changes with cellular outcomes, the results revealed that, in the GPD2-M group, the mRNA levels of both GPD2 gene and coagulation-related indicators were significantly reduced (Fig. 3L, M). Additionally, the corresponding protein expression levels (Fig. 3N–P), as well as the NAD+/NADH ratio (Fig. 3Q) and cell viability (Fig. 3T), were also markedly decreased. After 48 h of culture, the intracellular ATP concentration in the GPD2-M group was significantly lower than that in the NC group (Fig. 3R), whereas the residual glucose concentration in the medium was significantly higher (Fig. 3S).
RNA sequencing (RNA-seq) results reveal a potential pathway through which GPD2 gene (cg03230175) methylation affects coagulation function
To investigate the signaling pathway through which GPD2 gene (cg03230175) methylation affects the coagulation function, we subjected the NC and GPD2-M groups to transcriptomic sequencing.
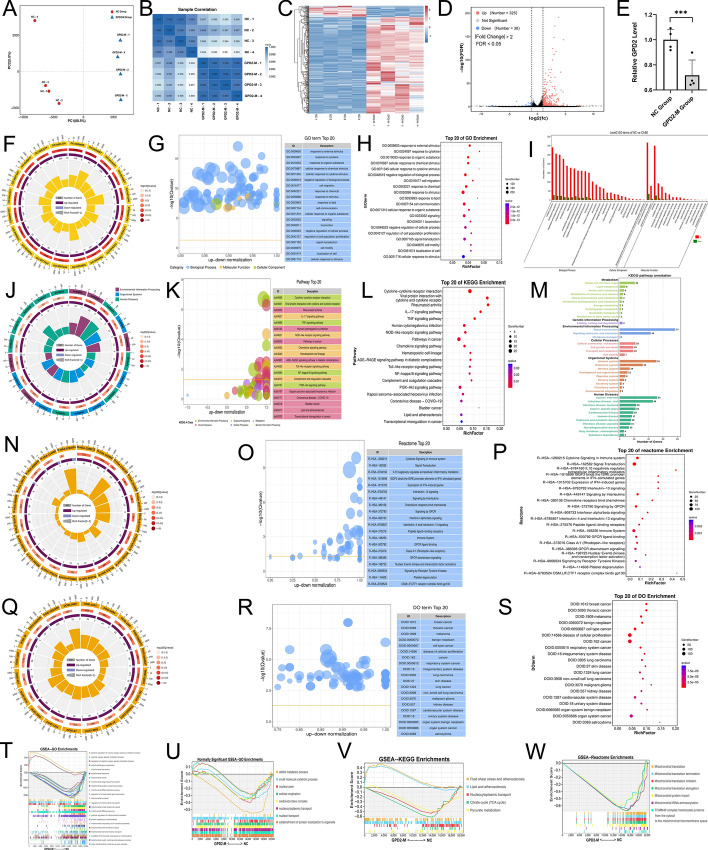
We assessed the consistency of gene expression across samples by principal component analysis (PCA) and Pearson correlation coefficients. The first principal component accounted for 88.5% of the variance, signifying that it represents the primary information in the dataset. The second component contributed an additional 5.5% to the variance, offering supplementary variability information (Fig. 4A). High intragroup correlations were observed within the NC and GPD2-M groups, while intergroup correlations, albeit lower, were still significant, with all R2 values above 0.99 (Fig. 4B). The analysis revealed 38 downregulated genes, including GPD2 (Fig. 4C–E).
Fig. 4.
The results of bulk RNA-seq. A PCA plot. B Correlation heatmap. C Complex heatmap. Red indicates upregulated genes, whereas blue represents downregulated genes. D Volcano plot. E Statistical chart of GPD2 gene expression; F GO enrichment circle plot. First circle: The top 20 enriched GO terms, with the scale for the number of DEGs on the outside. Different colors represent different ontologies. Second circle: The number of DEGs enriched in the GO term, along with the Q-value. The longer the bar, the more genes, and the redder the color, the smaller the Q-value. Third circle: bar graph of the proportion of upregulated and downregulated DEGs. Dark purple represents the proportion of upregulated genes, while light purple represents the proportion of downregulated genes; specific values are shown below. Fourth circle: the rich factor for each GO term, with background grid lines, each increment representing 0.1; G GO bubble plot. The y-axis represents −log10(Q-value), and the x-axis denotes the z-score value. The yellow line indicates the threshold at Q = 0.05. On the right is a list of the top 20 GO terms by Q-value. Different colors represent different ontologies; H GO enrichment bubble plot. The chart is constructed using the top 20 GO terms with the lowest Q-values. The y-axis represents the GO terms, and the x-axis indicates the rich factor. A larger bubble represents a greater number of genes, and the redder color means a smaller Q-value; I GO categorical bar chart. The x-axis represents secondary GO terms, and the y-axis indicates the number of DEGs within each term, with different colors denoting different GO terms. J KEGG enrichment circle plot. K KEGG bubble plot. L KEGG enrichment bubble plot. M KEGG categorical bar plot. The bold font on the y-axis indicates the KEGG_A level classification information, while the other colored fonts represent the B_level classification information. The x-axis shows the number of DEGs enriched in the pathway. N Reactome enrichment circle plot. O Reactome bubble plot. P Reactome enrichment bubble plot. Q DO enrichment circle plot. R DO bubble plot. S DO enrichment bubble plot. T GSEA-GO ES plot, this plot illustrates the enrichment score (ES) for the gene set associated with the GO term along the ranked list of genes. A positive ES value indicates that the gene expression associated with the GO term is upregulated in the GPD2-M group. Conversely, a negative ES value signifies downregulation. U GSEA-GO ES plot of GO terms centered around the GPD2 gene. V GSEA-KEGG ES plot. W GSEA-reactome ES plot
The GO analysis revealed that the differentially expressed genes (DEGs) are significantly enriched in several biological processes, including cell migration, cell communication, signaling, etc. Notably, the process of response to external stimulus was the most significantly enriched term (Fig. 4F–I). The Kyoto Encyclopedia of Genes and Genomes. (KEGG) pathway analysis identified several significantly enriched pathways among the DEGs, such as the interleukin (IL)-17 signaling pathway, the tumor necrosis factor (TNF) signaling pathway, the phosphoinositide 3-kinase/protein kinase (PI3K-Akt) signaling pathway, and notably, the NF-κB signaling pathway (Fig. 4J–M). In the reactome analysis, we identified several terms that are potentially associated with blood clotting function, including signaling by GPCR, GPCR ligand binding, GPCR downstream signaling, and platelet degranulation (Fig. 4N–P). In the Disease Ontology (DO) analysis, we identified a term pertinent to this study, specifically cardiovascular system disease (Fig. 4Q–S).
In gene set enrichment analysis (GSEA), we observed that all mitochondrial-related GO terms were downregulated (Fig. 4T). Additionally, GO terms centered around the GPD2 gene were significantly downregulated, including those related to alditol metabolism, nuclear pore, cellular respiration, oxidoreductase complex, nucleocytoplasmic transport, and nuclear transport (Fig. 4U), which may be associated with the nuclear translocation process of NF-κB and energy metabolism. Furthermore, we found that KEGG pathways associated with lipids and atherosclerosis were upregulated (Fig. 4V), whereas KEGG pathways related to energy metabolism and mitochondrial-related reactome pathways were downregulated (Fig. 4W).
Methylation of the GPD2 gene (cg03230175) modulates ROS levels, which in turn influence the transcription of the P2Y12 gene through the NF-κB pathway
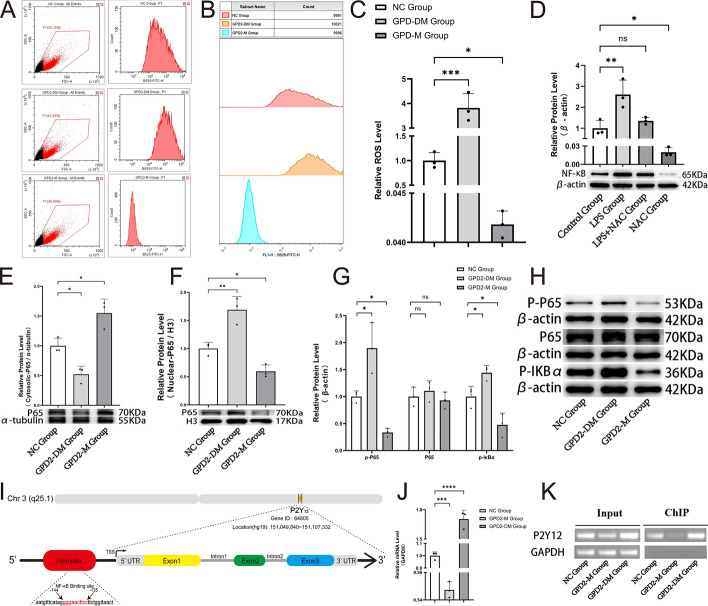
GPD2 is a “leaky” protein that produces large amounts of ROS [35, 36]. The glycerol 3-phosphate shuttle is a direct link between glycolysis and mitochondrial ROS release [37]. Therefore, to further explore the signaling pathway by which GPD2 gene (cg03230175) methylation affects blood coagulation function at the cellular level, we first detected the intracellular ROS levels using flow cytometry. The results showed that, compared with the NC group, the ROS level in the GPD2-M group significantly decreased, while the ROS level in the GPD2-DM group significantly increased (Fig. 5A–C). Subsequently, by adding lipopolysaccharides (LPS) and N-acetyl-l-cysteine (NAC) to the two groups of MEG-01 cells, we confirmed that changes in ROS levels indeed affected the activation of NF-κB (Fig. 5D). Then, we found that, compared to withthe NC group, the level of P65 in the cytoplasm of the GPD2-M group increased (Fig. 5E), while the level of P65 in the nucleus decreased (Fig. 5F), and the levels of p-P65 and p-IκBα in the entire cell also decreased (Fig. 5G, H), indicating that the activation process of NF-κB was inhibited, and the results in the GPD2-DM group were the opposite. Finally, ChIP analysis showed a significant increase in the binding of p65 to the P2Y12 promoter region (Fig. 5I–K). In summary, these results strongly suggest that methylation of the GPD2 gene leads to a decrease in ROS levels and subsequently inhibits the activation of NF-κB, leading to a decrease in P2Y12 gene expression.
Fig. 5.
GPD2 gene (cg03230175) methylation modulates intracellular ROS levels, which can influence P2Y12 gene expression by modulating the activation of NF-κB. NF-κB p65 binding to the P2Y12 promoter was determined by ChIP assay. A Flow cytometric analysis. The left panel shows a scatter plot of all events, highlighting P1 region. The right panel presents a histogram of fluorescence intensity for the B525-FITC-H channel within the P1 gate. B Summary flow cytometric analysis of cells from the three groups. Each panel represents the fluorescence intensity histograms for each group. C Statistical analysis bar plot of ROS levels. The ROS levels in each group are shown. Data expressed as mean ± standard error on the mean (SEM), n = 3. D Statistical chart of the relative NF-κB p65 protein levels after various interventions, accompanied by Western blot analysis. E Statistical chart of relative cytosolic NF-κB p65 protein level, accompanied by Western blot analysis. F Statistical chart of relative nuclear NF-κB p65 protein level, accompanied by Western blot analysis. G, H Statistical chart and Western blot analysis of relative levels of phosphorylated P65 (p-P65), total P65, and phosphorylated IκBα (p-IκBα) proteins. I Schematic diagram showing the NF-κB binding position in the P2Y12 promoter. J Statistical bar graph of the relative mRNA levels. K Gel electrophoresis banding pattern compares mRNA levels under input and ChIP conditions across the same three groups. The right panel shows the amplification of the P2Y12 promoter region containing the NF-κB binding motif after ChIP from MEG-01 cells, and the left panel shows the total DNA input. GAPDH was used as a control to demonstrate precipitation specificity
Methylation of the GPD2 gene (cg03230175) leads to mitochondrial damage, induces apoptosis, and inhibits the activity of PLPs
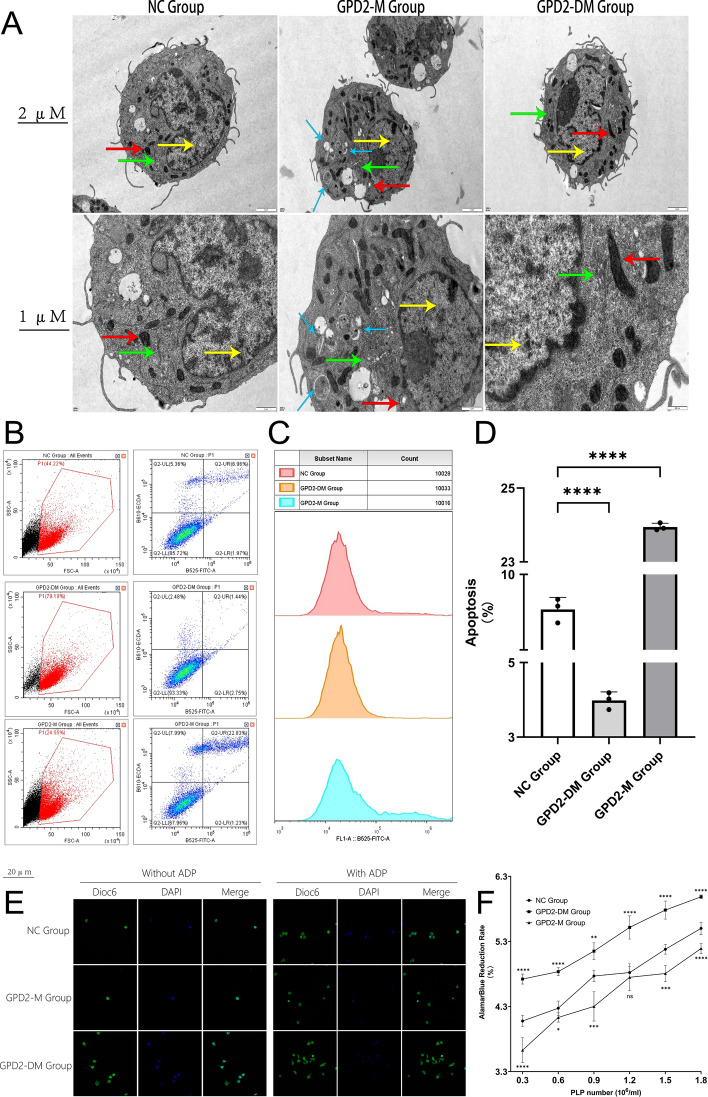
To observe the effects of GPD2 gene (cg03230175) methylation on the morphology of cells and mitochondria, we conducted electron microscopy analysis (Fig. 6A). The results revealed that the overall cell structure in the GPD2-M group was abnormal: the cell nuclei were elliptical with intact nuclear membranes, as indicated by the yellow arrows; the mitochondria were swollen, with some mitochondrial cristae fractured or disappeared, as indicated by the red arrows; a small amount of endoplasmic reticulum was dilated and vesiculated, as indicated by the green arrows; and a large number of autophagic bodies appeared, as indicated by the black arrows. In contrast, no abnormalities were observed in the NC and GPD2-DM groups. Subsequently, we used flow cytometry to detect apoptosis (Fig. 6B–D). The results showed that, compared with the NC group, the apoptosis rate in the GPD2-M group was significantly increased, whereas the opposite was true for the GPD2-DM group. Finally, we observed the PLPs produced by MEG-01 cells using laser confocal microscopy and detected PLPs activity with the AlamarBlue reagent. We found that GPD2 gene (cg03230175) methylation could reduce the number of PLPs and decrease their adhesion levels (Fig. 6E). Adding ADP or demethylating the GPD2 gene (cg03230175) could reverse these effects. Moreover, the PLP activity was also suppressed (Fig. 6F). For instance, when the PLP density reached 1.8 × 106/ml, the AlamarBlue reduction rate was 5.50% in the NC group, 5.99% in the GPD2-DM group (p < 0.0001), and just 5.19% in the GPD2-M group (p < 0.0001).
Fig. 6.
The methylation status of the GPD2 gene (cg03230175) affects mitochondrial morphology, apoptosis, and number and aggregation of platelet-like particles (PLPs). A Transmission electron microscopy image. Yellow arrows indicate the nucleus; red arrows indicate mitochondria; green arrows indicate the endoplasmic reticulum; black arrows indicate autophagosomes. B, C Flow cytometry analysis of cell apoptosis via annexin V/propidium iodide (PI) staining. D Statistical analysis bar plot of apoptotic cells. The proportion of apoptotic cells in each group is shown. Data expressed as mean ± SEM, n = 3. E Laser scanning confocal microscopy image. Representation of PLPs observed under a laser scanning confocal microscope in each group. F Statistical analysis line graph. The relative fluorescence intensity of PLPs across each group at varying PLPs concentrations. Data presented as mean ± SEM, n = 6
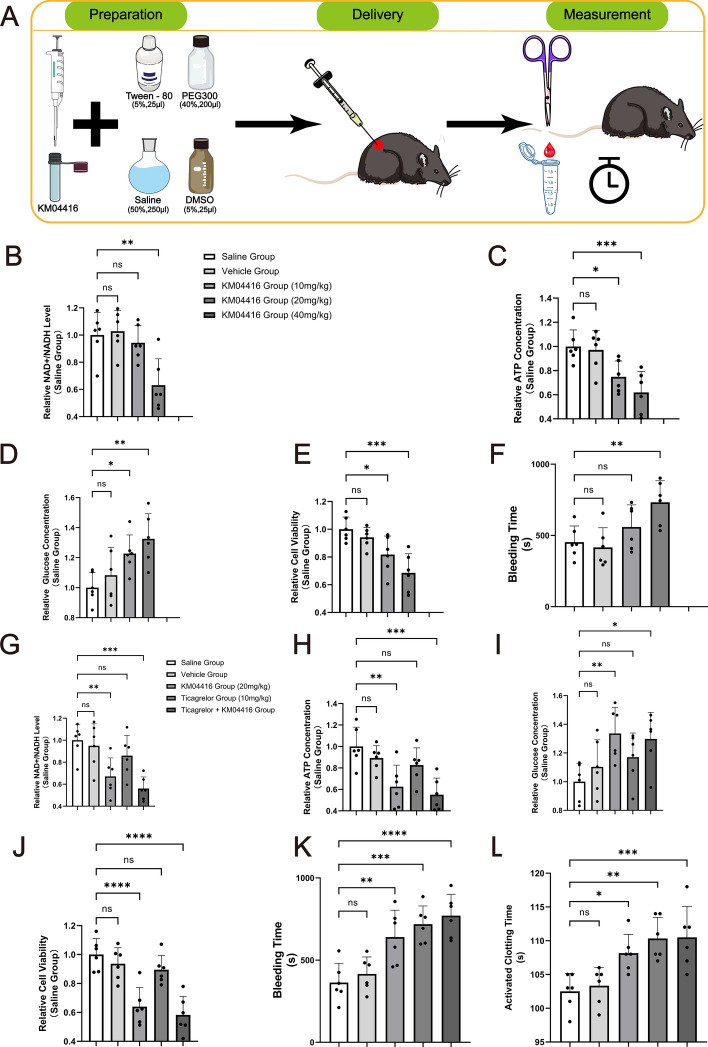
Effect of GPD2 inhibition on mice
To validate the role of the GPD2 enzyme in coagulation function, we administered the GPD2 inhibitor KM04416 via intraperitoneal injection to wild-type C57BL/6 mice (Fig. 7A). We first referred to the studies by Sehyun Oh et al. [38] and Gurmit Singh et al. [39], setting up three different concentrations for a preliminary experiment to determine the effective concentration of KM04416. The results showed that, at a dose of 20 mg KM04416 per kilogram of body weight (labeled as 20 mg/kg), the vitality of the mice was significantly reduced, and several indicators related to the GPD2 enzyme showed significant changes (Fig. 7B–E); in the 40 mg/kg group, all mice died. Furthermore, we followed the methodology of Jacob et al. [40] and Østergaard et al. [41] by clipping the terminal 2 mm of the mice tail and subsequently measuring the bleeding time. The results revealed that the bleeding time of mice in the 20 mg/kg group was significantly prolonged (Fig. 7F).
Fig. 7.
The GPD2 enzyme regulates coagulation function by affecting the mitochondrial energy metabolism process. A Illustrative diagram of the process of animal experiments. DMSO (5%) serves as the primary solvent and penetration enhancer; 40% PEG300 enhances drug solubility; 5% Tween-80 acts as a surfactant; 50% saline ensures physiological compatibility. B Relative NAD+/NADH levels in mice following injection of KM04416. C Relative ATP concentration in mice following injection of KM04416. D Relative glucose concentration in mice following injection of KM04416. E Relative cell viability in mice following injection of KM04416. F Tail-clip bleeding time in mice after the injection of KM04416. G Relative NAD+/NADH levels in mice following injection of KM04416 or ticagrelor. H Relative ATP concentration in mice following injection of KM04416 or ticagrelor. I Relative glucose concentration in mice following injection of KM04416 or ticagrelor. J Relative cell viability in mice following injection of KM04416 or ticagrelor. K Tail-cut bleeding time in mice after the injection of KM04416 or ticagrelor. L Activation clotting time in mice after the injection of KM04416 or ticagrelor
To further demonstrate that GPD2 enzyme regulates coagulation function by affecting the mitochondrial energy metabolism process, we introduced the P2Y12 inhibitor ticagrelor for corresponding animal experiments. The results showed that the vitality of the mice was significantly reduced, and several indicators related to GPD2 enzyme function in the 20 mg/kg group showed significant changes (Fig. 7G–J), while no such changes were observed in the ticagrelor group; however, the tail-clip bleeding time was significantly prolonged in both groups of mice (Fig. 7K). In addition, we also measured the activated clotting time (ACT) of mice (Fig. 7L). Measurement of ACT serves as another effective means to evaluate the anticoagulant effect of medications [42–44]. The results revealed that the ACT was also significantly prolonged in both groups of mice. This evidence suggests that GPD2 may regulate coagulation function by influencing the mitochondrial energy metabolism process.
Discussion
This study has several strengths: (i) Rich clinical sample cohort: The research leveraged a sizeable cohort of patients with CHD, yielding an extensive collection of clinical samples for analysis. (ii) Epigenetic revelation: We provided an epigenetic viewpoint by suggesting that GPD2 gene (cg03230175) methylation impacts coagulation function. (iii) Interdisciplinary strategy: Through the integration of epigenetics, transcriptomics, and molecular biology, we delineated the molecular mechanism by which GPD2 gene (cg03230175) methylation affects coagulation function via the ROS/NF-κB/P2Y12 signaling pathway.
While our study provides novel insights into the role of GPD2 gene (cg03230175) methylation in ticagrelor-associated bleeding, several limitations warrant cautious interpretation. First, the murine models employed pharmacological inhibition of GPD2 enzyme activity rather than direct manipulation of DNA methylation. Although this approach successfully recapitulated coagulation dysfunction phenotypes observed in human cohorts, it does not establish a causal link between GPD2 gene (cg03230175) methylation and bleeding risk at the animal level. Future studies using in vivo epigenetic editing tools such as CRISPR-dCas9-DNMT3A/Tet1CD-U6-sgRNA systems are required to dissect the direct contribution of GPD2 gene (cg03230175) methylation to coagulation function. Second, our animal experiments focused on acute pharmacological inhibition, whereas clinical antiplatelet therapy involves chronic ticagrelor exposure. Longitudinal studies assessing how prolonged GPD2 gene (cg03230175) methylation changes interact with drug metabolism and platelet functions may better model real-world scenarios. Nevertheless, in this study our integrative analysis suggests that GPD2 gene (cg03230175) methylation may serve as a potential biomarker for bleeding risk. Unlike static genetic variants, DNA methylation is reversible and may reflect dynamic interactions between antiplatelet therapy and metabolic stress. The robust association between GPD2 gene (cg03230175) methylation and bleeding events in ticagrelor-treated patients with CHD may guide personalized antiplatelet strategies.
Moving forward, future research directions could be enriched by considering a multi-omics approach that integrates various biological levels: (i) Integration of proteomics: To complement the epigenetic and transcriptomic data, proteomic analysis could be employed to investigate the downstream effects of GPD2 gene (cg03230175) methylation on protein expression and function. This would provide a more complete picture of how epigenetic changes translate into cellular and physiological outcomes. (ii) Metabolomics profiling: By analyzing the metabolic changes associated with GPD2 gene (cg03230175) methylation, metabolomics could uncover novel pathways and biomarkers related to coagulation dysfunction and CHD progression. This could lead to a better understanding of the systemic effects of epigenetic modifications. (iii) Genome-wide association studies (GWAS): Future studies could incorporate GWAS to identify additional genetic variants associated with GPD2 gene (cg03230175) methylation and coagulation function. This could reveal a broader genetic architecture that influences the risk of CHD. (iv) Functional validation in animal models: Overcoming the current limitations, the development of animal models with altered GPD2 gene (cg03230175) methylation status would allow for functional validation of the epigenetic findings. This would provide critical insights into the in vivo effects of GPD2 gene (cg03230175) methylation on coagulation function and its relevance to CHD pathogenesis. (v) Clinical translation studies: Large-scale, prospective clinical studies are needed to validate the predictive biomarker potential of GPD2 gene (cg03230175) methylation. Additionally, these studies could explore the efficacy of targeted therapies that modulate GPD2 gene (cg03230175) methylation in personalized medicine approaches. (vi) Single-cell omics: Advances in single-cell technologies could be leveraged to investigate epigenetic heterogeneity within patient samples. This would allow for more precise understanding of how different cell types contribute to the overall pathophysiology of CHD.
In summary, our investigation into the epigenetic regulation of GPD2 gene (cg03230175) methylation on coagulation function lays a foundational understanding that could guide the development of personalized medication regimens and enhance bleeding risk assessment for patients with CHD. The integration of multi-omics approaches in future research will be crucial for validating and expanding upon these findings. Such an integrative strategy holds the promise of not only enhancing our understanding of CHD but also paving the way for more precise and personalized medical interventions. Furthermore, future research should overcome the limitations faced in this study, particularly by establishing animal models to validate our findings and further explore the clinical implications of GPD2 gene (cg03230175) methylation in personalized medicine.
Conclusions
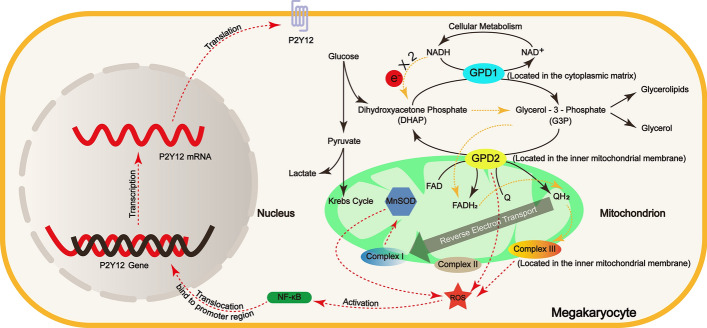
As shown in Fig. 8, this study delineated the impact of GPD2 gene (cg03230175) methylation on coagulation function, highlighting the pivotal role of the ROS/NF-κB/P2Y12 signaling pathway.
Fig. 8.
Graphical summary of the role of GPD2 in coagulation function. The solid black lines in the figure indicate normal metabolic processes. The dashed yellow line depicts the electron transfer pathway. The red dotted line illustrates the route through which the GPD2 enzyme influences blood clotting function
Although financial limitations precluded the creation of animal models, the integrative methodologies used offer profound insights into the molecular mechanisms underlying coagulation function impairments. Moreover, the results suggest that GPD2 gene (cg03230175) methylation could be a promising biomarker for predicting bleeding risk and a potential target for therapeutic strategies in the management of coronary artery disease.
Supplementary Information
Acknowledgements
We firstly extend our heartfelt gratitude to the participants for their contribution and cooperation. We are also deeply thankful to the implementers for their meticulous execution and unwavering commitment to the research process. Additionally, special thanks to Shanghai LabEx Biotech Co., Ltd. for supplying the professional reagents and consumables related to flow cytometry. Lastly, we acknowledge with profound respect the collective efforts of all the authors, whose intellectual contributions and collaborative spirit have been instrumental in bringing this work to fruition.
Abbreviations
- ACS
Acute coronary syndrome
- ACT
Activated clotting time
- ATP
Adenosine triphosphate
- CHD
Coronary heart disease
- CML
Chronic myeloid leukemia
- CpG
Cytosine–phosphate–guanine
- CRISPR
Clustered regularly interspaced short palindromic repeats
- dCas9
dead Cas9 protein
- DMR
Differential methylation region
- DNAm
DNA methylation
- DNMTs
DNA methyltransferases
- DO
Disease Ontology
- eQTL
Expression quantitative trait loci
- EWAS
Epigenome-wide association studies
- FADH
Flavine adenine dinucleotide
- FBS
Fetal bovine serum
- GO
Gene ontology
- GPD2
Glycerol-3-phosphate dehydrogenase 2
- GPS
Glycerophosphate shuttle
- GWAS
Genome-wide association studies
- HEIDI
HEterogeneity in Dependent Instruments
- KEGG
Kyoto Encyclopedia of Genes and Genomes
- MEG-01
Megakaryoblast leukemia cell line
- meQTL
Methylation quantitative trait loci
- NAD
Nicotinamide adenine dinucleotide
- NF-κB
Nuclear factor kappa B
- PCI
Percutaneous coronary intervention
- PLPs
Platelet-like particles
- P/S
Penicillin–streptomycin
- RET
Reverse electron transport
- ROS
Reactive oxygen species
- SMR
Summary data-based Mendelian randomization
- Tets
Ten–eleven translocation family proteins
Author contributions
The study was designed and conducted by Jiajie Chen. Guifeng Xu was responsible for EWAS. Zhipeng Xie and Shaoxia Xie were involved in the recruitment of patients. Wenwei Luo revised and finalized the manuscript and acquired funding. Weihua Lai and Shilong Zhong undertook the role of manuscript reviser and acquired funding. All authors actively engaged in the research process, made significant contributions to manuscript revisions, and meticulously reviewed and approved the final version.
Funding
This study was supported by grants from the National Natural Science Foundation of China (82274016, 82473910, 82003746, and 81872934), Leading Talent of Guangdong Special Support Program (0720240120), Guangdong Basic and Applied Basic Research Foundation (2021A1515220031, 2023A1515011133, and 2025A1515012395), Science and Technology Program of Guangzhou (2023B03J1251), Key-Area Research and Development Program of Guangdong Province, China (2019B020229003), and National key research and development program (2017YFC0909301).
Data availability
Data were taken from published randomized controlled trials (NCT03092076), which were conducted according to the principles of the Declaration of Helsinki and with informed consent from participants. The datasets generated or analyzed during the study are available from the corresponding author on reasonable request.
Declarations
Ethics approval and consent to participate
The Ethics Review Committee of Guangdong Provincial People’s Hospital approved our human and animal studies (no. GDREC:KY2023-090-02) on 10 March 2023, in compliance with the “Ethical Review Measures for Biomedical Research Involving Human Subjects” and “Guidance on the Humane Treatment of Laboratory Animals” issued by the Ministry of Science and Technology of China, as well as relevant international animal welfare guidelines and regulations (including the ICLAS statement). Our research strictly adhered to the Declaration of Helsinki and other applicable guidelines. The study was registered with ClinicalTrials.gov (NCT03092076) and was strictly conducted in accordance with the relevant provisions of the Declaration of Helsinki and received approval from the Medical Research Ethics Committee of Guangdong Provincial People’s Hospital (no. GDREC2015143H(R1) and no. GDREC2017071H). Each patient was fully informed of the details of the research and the entire process. Furthermore, written informed consent was obtained from all participants (no. 20150115 and no.20170211).
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Wenwei Luo, Email: luowenwei@gdph.org.cn.
Shilong Zhong, Email: shz2020@qq.com.
Weihua Lai, Email: laiweihuax@163.com.
References
- 1.Levine GN, Bates ER, Bittl JA, Brindis RG, Fihn SD, Fleisher LA, Granger CB, Lange RA, Mack MJ, Mauri L, Mehran R, Mukherjee D, Newby LK, O’Gara PT, Sabatine MS, Smith PK, Smith SC Jr. 2016 ACC/AHA guideline focused update on duration of dual antiplatelet therapy in patients with coronary artery disease: a report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines. J Am Coll Cardiol. 2016;68(10):1082–115. 10.1016/j.jacc.2016.03.513. [DOI] [PubMed] [Google Scholar]
- 2.Corbett SJ, Ftouh S, Lewis S, Lovibond K, Guideline Committee. Acute coronary syndromes: summary of updated NICE guidance. BMJ. 2021;372:m4760. 10.1136/bmj.m4760. [DOI] [PubMed] [Google Scholar]
- 3.Kamran H, Jneid H, Kayani WT, Virani SS, Levine GN, Nambi V, Khalid U. Oral antiplatelet therapy after acute coronary syndrome: a review. JAMA. 2021;325(15):1545-1555. 10.1001/jama.2021.0716. Erratum in: JAMA. 2021 Jul 13;326(2):190. 10.1001/jama.2021.9484. [DOI] [PubMed]
- 4.Wallentin L, Becker RC, Budaj A, Cannon CP, Emanuelsson H, Held C, Horrow J, Husted S, James S, Katus H, Mahaffey KW, Scirica BM, Skene A, Steg PG, Storey RF, Harrington RA, Freij A, Thorsén M, PLATO Investigators. Ticagrelor versus clopidogrel in patients with acute coronary syndromes. N Engl J Med. 2009;361(11):1045–57. 10.1056/NEJMoa0904327. [DOI] [PubMed] [Google Scholar]
- 5.Valgimigli M, Bueno H, Byrne RA, Collet JP, Costa F, Jeppsson A, Jüni P, Kastrati A, Kolh P, Mauri L, Montalescot G, Neumann FJ, Petricevic M, Roffi M, Steg PG, Windecker S, Zamorano JL, ESC Scientific Document Group, ESC Committee for Practice Guidelines (CPG), ESC National Cardiac Societies. 2017 ESC focused update on dual antiplatelet therapy in coronary artery disease developed in collaboration with EACTS: the task force for dual antiplatelet therapy in coronary artery disease of the European Society of Cardiology (ESC) and of the European Association for Cardio-Thoracic Surgery (EACTS). Eur Heart J. 2018;39(3):213–60. 10.1093/eurheartj/ehx419. [DOI] [PubMed] [Google Scholar]
- 6.Zhu Q, Zhong W, Wang X, Mai L, He G, Chen J, Tang L, Liu S, Lai W, Zhong S. Pharmacokinetic and pharmacogenetic factors contributing to platelet function recovery after single dose of ticagrelor in healthy subjects. Front Pharmacol. 2019;10:209. 10.3389/fphar.2019.00209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Gurbel PA, Bliden KP, Butler K, Tantry US, Gesheff T, Wei C, Teng R, Antonino MJ, Patil SB, Karunakaran A, Kereiakes DJ, Parris C, Purdy D, Wilson V, Ledley GS, Storey RF. Randomized double-blind assessment of the ONSET and OFFSET of the antiplatelet effects of ticagrelor versus clopidogrel in patients with stable coronary artery disease: the ONSET/OFFSET study. Circulation. 2009;120(25):2577–85. 10.1161/CIRCULATIONAHA.109.912550. [DOI] [PubMed] [Google Scholar]
- 8.Storey RF, Bliden KP, Ecob R, Karunakaran A, Butler K, Wei C, Tantry U, Gurbel PA. Earlier recovery of platelet function after discontinuation of treatment with ticagrelor compared with clopidogrel in patients with high antiplatelet responses. J Thromb Haemost. 2011;9(9):1730–7. 10.1111/j.1538-7836.2011.04419.x. [DOI] [PubMed] [Google Scholar]
- 9.Corbin LJ, White SJ, Taylor AE, Williams CM, Taylor K, van den Bosch MT, Teasdale JE, Jones M, Bond M, Harper MT, Falk L, Groom A, Hazell GGJ, Paternoster L, Munafò MR, Nordestgaard BG, Tybjærg-Hansen A, Bojesen SE, Relton C, Min JL, Davey Smith G, Mumford AD, Poole AW, Timpson NJ. Epigenetic regulation of F2RL3 associates with myocardial infarction and platelet function. Circ Res. 2022;130(3):384–400. 10.1161/CIRCRESAHA.121.318836. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Li XG, Ma N, Wang B, Li XQ, Mei SH, Zhao K, Wang YJ, Li W, Zhao ZG, Sun SS, Miao ZR. The impact of P2Y12 promoter DNA methylation on the recurrence of ischemic events in Chinese patients with ischemic cerebrovascular disease. Sci Rep. 2016;6:34570. 10.1038/srep34570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Su J, Li X, Yu Q, Liu Y, Wang Y, Song H, Cui H, Du W, Fei X, Liu J, Lin S, Wang J, Zheng W, Zhong J, Zhang L, Tong M, Xu J, Chen X. Association of P2Y12 gene promoter DNA methylation with the risk of clopidogrel resistance in coronary artery disease patients. Biomed Res Int. 2014;2014:450814. 10.1155/2014/450814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Yang J, Zhou JS, Zhao YX, Yang ZH, Zhao HD, Zhang YD, Zou JJ. ABCB1 hypomethylation is associated with decreased antiplatelet effects of clopidogrel in Chinese ischemic stroke patients. Pharmazie. 2015;70(2):97–102. 10.1691/ph.2015.4756. [PubMed] [Google Scholar]
- 13.Li X, Zhao K, Ma N, Sun S, Miao Z, Zhao Z. Association of ABCB1 promoter methylation with aspirin exposure, platelet function, and clinical outcomes in Chinese intracranial artery stenosis patients. Eur J Clin Pharmacol. 2017;73(10):1261–9. 10.1007/s00228-017-2298-z. [DOI] [PubMed] [Google Scholar]
- 14.Lei HP, Yu XY, Wu H, Kang YH, Zhong WP, Cai LY, Zhang MZ, Chen JY, Mai LP, Ding QS, Yang M, Zhong SL. Effects of PON1 gene promoter DNA methylation and genetic variations on the clinical outcomes of dual antiplatelet therapy for patients undergoing percutaneous coronary intervention. Clin Pharmacokinet. 2018;57(7):817–29. 10.1007/s40262-017-0595-4. [DOI] [PubMed] [Google Scholar]
- 15.Lyko F. The DNA methyltransferase family: a versatile toolkit for epigenetic regulation. Nat Rev Genet. 2018;19(2):81–92. 10.1038/nrg.2017.80. [DOI] [PubMed] [Google Scholar]
- 16.Kohli RM, Zhang Y. TET enzymes, TDG and the dynamics of DNA demethylation. Nature. 2013;502(7472):472–9. 10.1038/nature12750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Liu XS, Wu H, Ji X, Stelzer Y, Wu X, Czauderna S, Shu J, Dadon D, Young RA, Jaenisch R. Editing DNA methylation in the mammalian genome. Cell. 2016;167(1):233-247.e17. 10.1016/j.cell.2016.08.056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Mráček T, Drahota Z, Houštěk J. The function and the role of the mitochondrial glycerol-3-phosphate dehydrogenase in mammalian tissues. Biochim Biophys Acta. 2013;1827(3):401–10. 10.1016/j.bbabio.2012.11.014. [DOI] [PubMed] [Google Scholar]
- 19.Aibibula M, Naseem KM, Sturmey RG. Glucose metabolism and metabolic flexibility in blood platelets. J Thromb Haemost. 2018;16(11):2300–14. 10.1111/jth.14274. [DOI] [PubMed] [Google Scholar]
- 20.Hu L, Chang L, Zhang Y, Zhai L, Zhang S, Qi Z, Yan H, Yan Y, Luo X, Zhang S, Wang Y, Kunapuli SP, Ye H, Ding Z. Platelets express activated P2Y12 receptor in patients with diabetes mellitus. Circulation. 2017;136(9):817–33. 10.1161/CIRCULATIONAHA.116.026995. [DOI] [PubMed] [Google Scholar]
- 21.Langston PK, Nambu A, Jung J, Shibata M, Aksoylar HI, Lei J, Xu P, Doan MT, Jiang H, MacArthur MR, Gao X, Kong Y, Chouchani ET, Locasale JW, Snyder NW, Horng T. Glycerol phosphate shuttle enzyme GPD2 regulates macrophage inflammatory responses. Nat Immunol. 2019;20(9):1186–95. 10.1038/s41590-019-0517-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Robb EL, Hall AR, Prime TA, Eaton S, Szibor M, Viscomi C, James AM, Murphy MP. Control of mitochondrial superoxide production by reverse electron transport at complex I. J Biol Chem. 2018;293(25):9869–79. 10.1074/jbc.AAC119.008929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Chouchani ET, Pell VR, Gaude E, Aksentijević D, Sundier SY, Robb EL, Logan A, Nadtochiy SM, Ord ENJ, Smith AC, Eyassu F, Shirley R, Hu CH, Dare AJ, James AM, Rogatti S, Hartley RC, Eaton S, Costa ASH, Brookes PS, Davidson SM, Duchen MR, Saeb-Parsy K, Shattock MJ, Robinson AJ, Work LM, Frezza C, Krieg T, Murphy MP. Ischaemic accumulation of succinate controls reperfusion injury through mitochondrial ROS. Nature. 2014;515(7527):431–5. 10.1038/nature13909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Zorov DB, Juhaszova M, Sollott SJ. Mitochondrial reactive oxygen species (ROS) and ROS-induced ROS release. Physiol Rev. 2014;94(3):909–50. 10.1152/physrev.00026.2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Hayden MS, Ghosh S. Shared principles in NF-kappaB signaling. Cell. 2008;132(3):344–62. 10.1016/j.cell.2008.01.020. [DOI] [PubMed] [Google Scholar]
- 26.Cattaneo M. The platelet P2Y₁₂ receptor for adenosine diphosphate: congenital and drug-induced defects. Blood. 2011;117(7):2102–12. 10.1182/blood-2010-08-263111. [DOI] [PubMed] [Google Scholar]
- 27.Rauch BH, Rosenkranz AC, Ermler S, Böhm A, Driessen J, Fischer JW, Sugidachi A, Jakubowski JA, Schrör K. Regulation of functionally active P2Y12 ADP receptors by thrombin in human smooth muscle cells and the presence of P2Y12 in carotid artery lesions. Arterioscler Thromb Vasc Biol. 2010;30(12):2434–42. 10.1161/ATVBAHA.110.213702. [DOI] [PubMed] [Google Scholar]
- 28.Kammers K, Taub MA, Rodriguez B, Yanek LR, Ruczinski I, Martin J, Kanchan K, Battle A, Cheng L, Wang ZZ, Johnson AD, Leek JT, Faraday N, Becker LC, Mathias RA. Transcriptional profile of platelets and iPSC-derived megakaryocytes from whole-genome and RNA sequencing. Blood. 2021;137(7):959–68. 10.1182/blood.2020006115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Heo Y, Jeon H, Namkung W. PAR4-mediated PI3K/Akt and RhoA/ROCK signaling pathways are essential for thrombin-induced morphological changes in MEG-01 cells. Int J Mol Sci. 2022;23(2):776. 10.3390/ijms23020776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Takeuchi K, Satoh M, Kuno H, Yoshida T, Kondo H, Takeuchi M. Platelet-like particle formation in the human megakaryoblastic leukaemia cell lines, MEG-01 and MEG-01s. Br J Haematol. 1998;100(2):436–44. 10.1046/j.1365-2141.1998.00576.x. [DOI] [PubMed] [Google Scholar]
- 31.Chowdhury SK, Gemin A, Singh G. High activity of mitochondrial glycerophosphate dehydrogenase and glycerophosphate-dependent ROS production in prostate cancer cell lines. Biochem Biophys Res Commun. 2005;333(4):1139–45. 10.1016/j.bbrc.2005.06.017. [DOI] [PubMed] [Google Scholar]
- 32.Valgimigli M, Gragnano F, Branca M, Franzone A, da Costa BR, Baber U, Kimura T, Jang Y, Hahn JY, Zhao Q, Windecker S, Gibson CM, Watanabe H, Kim BK, Song YB, Zhu Y, Vranckx P, Mehta S, Ando K, Hong SJ, Gwon HC, Serruys PW, Dangas GD, McFadden EP, Angiolillo DJ, Heg D, Calabrò P, Jüni P, Mehran R, Single Versus Dual Antiplatelet Therapy (Sidney-3) Collaboration. Ticagrelor or clopidogrel monotherapy vs dual antiplatelet therapy after percutaneous coronary intervention: a systematic review and patient-level meta-analysis. JAMA Cardiol. 2024;9(5):437–48. 10.1001/jamacardio.2024.0133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.André P. P-selectin in haemostasis. Br J Haematol. 2004;126(3):298–306. 10.1111/j.1365-2141.2004.05032.x. [DOI] [PubMed] [Google Scholar]
- 34.Joshi S, Banerjee M, Zhang J, Kesaraju A, Pokrovskaya ID, Storrie B, Whiteheart SW. Alterations in platelet secretion differentially affect thrombosis and hemostasis. Blood Adv. 2018;2(17):2187–98. 10.1182/bloodadvances.2018019166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Benz PM, Laban H, Zink J, Günther L, Walter U, Gambaryan S, Dib K. Vasodilator-Stimulated Phosphoprotein (VASP)-dependent and -independent pathways regulate thrombin-induced activation of Rap1b in platelets. Cell Commun Signal. 2016;14(1):21. 10.1186/s12964-016-0144-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Willson JA, Arienti S, Sadiku P, Reyes L, Coelho P, Morrison T, Rinaldi G, Dockrell DH, Whyte MKB, Walmsley SR. Neutrophil HIF-1α stabilization is augmented by mitochondrial ROS produced via the glycerol 3-phosphate shuttle. Blood. 2022;139(2):281–6. 10.1182/blood.2021011010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kamiński MM, Sauer SW, Kamiński M, Opp S, Ruppert T, Grigaravičius P, Grudnik P, Gröne HJ, Krammer PH, Gülow K. T cell activation is driven by an ADP-dependent glucokinase linking enhanced glycolysis with mitochondrial reactive oxygen species generation. Cell Rep. 2012;2(5):1300–15. 10.1016/j.celrep.2012.10.009. [DOI] [PubMed] [Google Scholar]
- 38.Oh S, Jo S, Bajzikova M, Kim HS, Dao TTP, Rohlena J, Kim JM, Neuzil J, Park S. Non-bioenergetic roles of mitochondrial GPD2 promote tumor progression. Theranostics. 2023;13(2):438–57. 10.7150/thno.75973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Singh G. Mitochondrial FAD-linked Glycerol-3-phosphate dehydrogenase: a target for cancer therapeutics. Pharmaceuticals (Basel). 2014;7(2):192–206. 10.3390/ph7020192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Jacob S, Kosaka Y, Bhatlekar S, Denorme F, Benzon H, Moody A, Moody V, Tugolukova EA, Hull G, Kishimoto N, Manne BK, Guo L, Souvenir R, Seliger BJ, Eustes AS, Hoerger K, Tolley ND, Fatahian AN, Boudina S, Christiani DC, Wei Y, Ju C, Campbell RA, Rondina MT, Abel ED, Bray PF, Weyrich AS, Rowley JW. Mitofusin-2 regulates platelet mitochondria and function. Circ Res. 2024;134(2):143–61. 10.1161/CIRCRESAHA.123.322914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Østergaard H, Lund J, Greisen PJ, Kjellev S, Henriksen A, Lorenzen N, Johansson E, Røder G, Rasch MG, Johnsen LB, Egebjerg T, Lund S, Rahbek-Nielsen H, Gandhi PS, Lamberth K, Loftager M, Andersen LM, Bonde AC, Stavenuiter F, Madsen DE, Li X, Holm TL, Ley CD, Thygesen P, Zhu H, Zhou R, Thorn K, Yang Z, Hermit MB, Bjelke JR, Hansen BG, Hilden I. A factor VIIIa-mimetic bispecific antibody, Mim8, ameliorates bleeding upon severe vascular challenge in hemophilia A mice. Blood. 2021;138(14):1258–68. 10.1182/blood.2020010331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Handin RI. The history of antithrombotic therapy: the discovery of heparin, the vitamin K antagonists, and the utility of aspirin. Hematol Oncol Clin North Am. 2016;30(5):987–93. 10.1016/j.hoc.2016.06.002. [DOI] [PubMed] [Google Scholar]
- 43.Baluwala I, Favaloro EJ, Pasalic L. Therapeutic monitoring of unfractionated heparin - trials and tribulations. Expert Rev Hematol. 2017;10(7):595–605. 10.1080/17474086.2017.1345306. [DOI] [PubMed] [Google Scholar]
- 44.Penk JS, Reddy S, Polito A, Cisco MJ, Allan CK, Bembea M, Giglia TM, Cheng HH, Thiagarajan RR, Dalton HJ. Bleeding and thrombosis with pediatric extracorporeal life support: a roadmap for management, research, and the future from the pediatric cardiac intensive care society: part 2. Pediatr Crit Care Med. 2019;20(11):1034–9. 10.1097/PCC.0000000000002104. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Data were taken from published randomized controlled trials (NCT03092076), which were conducted according to the principles of the Declaration of Helsinki and with informed consent from participants. The datasets generated or analyzed during the study are available from the corresponding author on reasonable request.