Abstract
Chloride intracellular channels (CLICs) are a family of six evolutionarily conserved proteins with diverse functions. Previously, we identified CLIC2, as the fifth-ranked master regulator associated with diffuse-type gastric cancer (dGC) showing increased expression in tumors. Here we used bulk, as well as single cell sequencing datasets of dGC, to demonstrate for the first time a direct association of CLIC2 with the microsatellite stable GC and, furthermore, the expression of CLIC2 in macrophages (MCs), and endothelial cells (ECs) populating gastric tissue. We generated CLIC2 knock-out THP-1 monocytic cells (THP-1CLIC2_KO) determining that while CLIC2 deletion had no observable effect on monocytes, THP-1CLIC2_KO macrophages exhibited significant morphological changes, including increased membrane protrusions, and upregulated expression of CD11b, CD11c, CD80, and CD86 markers. Furthermore, cytokine secretion profiling of THP-1CLIC2_KO differentiated cells revealed elevated secretion of CCL8, alongside reduced secretion of IL-1β, IL-6, and osteoprotegerin (OPG). Additionally, we observed increased phosphorylation of Shp1 phosphatase with the concomitant absence of Stat3 phosphorylation, which impaired downstream signaling, in line with the evidence that Clic2 interacts with both Shp1 and Stat3. Based on these findings, we suggest that CLIC2 plays a pivotal role in regulating monocyte-to-macrophage differentiation by modulating the Stat3 signaling pathway, thus enhancing gastric cancer progression by establishing a tumor-permissive microenvironment.
Supplementary Information
The online version contains supplementary material available at 10.1186/s13062-025-00666-3.
Keywords: Gastric microenvironment, Tumor microenvironment, Gastric cancer, Clic2, Macrophages, Differentiation, Signaling
Introduction
Gastric cancer (GC) is the fifth most common cancer and the fifth leading cause of cancer-related deaths globally, with 70% of incidence and mortality recorded in Asian countries (GLOBOCAN 2022). Even if many classification systems have been adopted for this extremely heterogeneous neoplasia, GC is still predominantly diagnosed by its histological pattern into intestinal (iGC), diffuse (dGC), and mixed (mGC) types according to the Lauren classification [1]. iGC show several clinical and phenotypical differences from dGC, it accounts for most of the GC cases, is characterized by localized lesions composed of well-differentiated tumor cells still retaining gastric gland architecture and is associated with a better prognosis. Conversely, dGC is less frequent, characterized by undifferentiated cells that infiltrate the submucosal layers of stomach and a worse prognosis. If examining the immune systems of the GC histotype, dGC tumors usually show an altered tumor microenvironment with deficient immune activation [2]. Furthermore, dGC is also associated with genomic stability [3] leading to low mutational burden and less neoantigen production, which contributes to the low immunogenicity of this histotype [4]. Previously, our group dissected the molecular features of GC, identifying, among the others, Chloride intracellular channel 2 (CLIC2) to be strongly associated with GC [5]. Chloride intracellular channels (CLICs) are a family of six evolutionary conserved proteins with a molecular mass of approximately 30 kDa, which have extremely heterogeneous functions [6, 7] and that have been found related to several diseases, including cancer [8]. CLICs are found in several tissues and intracellular locations, as they can exist as soluble globular and integral membrane proteins and have also been proposed to regulate ion channels rather than forming them themselves [7]. Previous studies have shown that CLIC2 has several cellular functions beyond ion exchange [9] with roles that vary depending on the organ and cellular context [10]. The impact of CLICs expression levels on survival rates varies significantly with the type of cancer, indicating that different cancers may employ distinct mechanisms for progression, metastasis, or invasion [7]. CLICs have long been elusive, with CLIC2 being the least investigated due to the absence of this gene in the murine genome, which prevented the use of knockout mice models for functional studies [11, 12]. CLIC2 is scarcely localized in membranous structures including plasma and organellar membranes, suggesting that the majority of CLIC2 does not form ion channels [13]. Our work contributes to clarify CLIC2 role in GC, exposing new evidence about CLIC2 expression and functions. While other studies so far explored CLIC2 influences on cancer cells [7, 12, 14], here we demonstrated for the first time that CLIC2 increased expression in GC do not account to gastric tumor cells, but instead it derives from immune cells directly or indirectly supporting the development of the surrounding cancer cells.
Results
CLIC2 regulon is intimately linked to diffuse-type gastric cancer and to genomically stable disease
In a previous study [5], we compared the gene expression profiles of the The Cancer Genome Atlas Project (TCGA) Stomach adenocarcinoma (STAD) dataset to dissect the molecular heterogeneity of diffuse-type gastric cancer (dGC) in comparison to intestinal-type gastric cancer (iGC). By this analysis we identified, for the first time, a gene network (defined as a regulon) associated with the gene CLIC2, which was more active in patients with dGC compared to those with iGC (Fig. 1a, Supplementary Table 1). To further elucidate the relationships linking the CLIC2 regulon to dGC, we searched for other covariates associated with the expression of this regulon and with CLIC2 gene expression. We found that both the active core of CLIC2 regulon (Fig. 1b) and the CLIC2 gene (Fig. 1c) were significantly upregulated in patients with microsatellite stable (MSS) disease compared to those with microsatellite instable disease (both low and high instability, MSI-L and MSI-H, respectively). Furthermore, by analyzing the network of genes comprising the CLIC2 regulon, we discovered that they were interconnected through categories that included the SHP2_PATHWAY (weight 22.91), HEMOSTASIS (weight 11.15), CELL SURFACE INTERACTIONS AT THE VASCULAR WALL (weight 9.31), and HALLMARK_IL2_STAT5_SIGNALING (weight 5.53) among others (Fig. 1d, Supplementary Table 2).
Fig. 1.
CLIC2 regulon and gene correlates with dGC and with genomically stable disease. a Heatmap visualization of the CLIC2 regulon gene core, showing expression levels of genes in relation to Lauren classification and master regulator activation. b Relative expression of the CLIC2 regulon in relation to the microsatellite stable gastric cancer (MSS) or microsatellite instable, low or high, gastric cancer (MSI-L and MSI-H). Orange boxes indicate patients expressing the active core of the CLIC2 regulon, green boxes indicate patients expressing genes not composing the CLIC2 regulon core, violet boxes indicate patients expressing the inactive core of the CLIC2 regulon. c Relative expression of the CLIC2 gene in relation to the microsatellite stable gastric cancer (MSS) or microsatellite instable, low or high, gastric cancer (MSI-L and MSI-H). Orange boxes indicate patients expressing the active core of the CLIC2 regulon, green boxes indicate patients expressing genes not composing the CLIC2 regulon core, violet boxes indicate patients expressing the inactive core of the CLIC2 regulon. d Gene network visualization of genes composing CLIC2 regulon. White circles indicate genes of the regulon, gray rhombus indicate categories from MSigDB, grey circles are secondary genes intercepted during analysis of connections, green lines are genetic interactions and grey lines are gene connections on reactome, MSigDB, InterPro and Wikipathways
CLIC2 expression is upregulated in diffuse-type gastric cancer and is peculiar to endothelial cells and macrophages
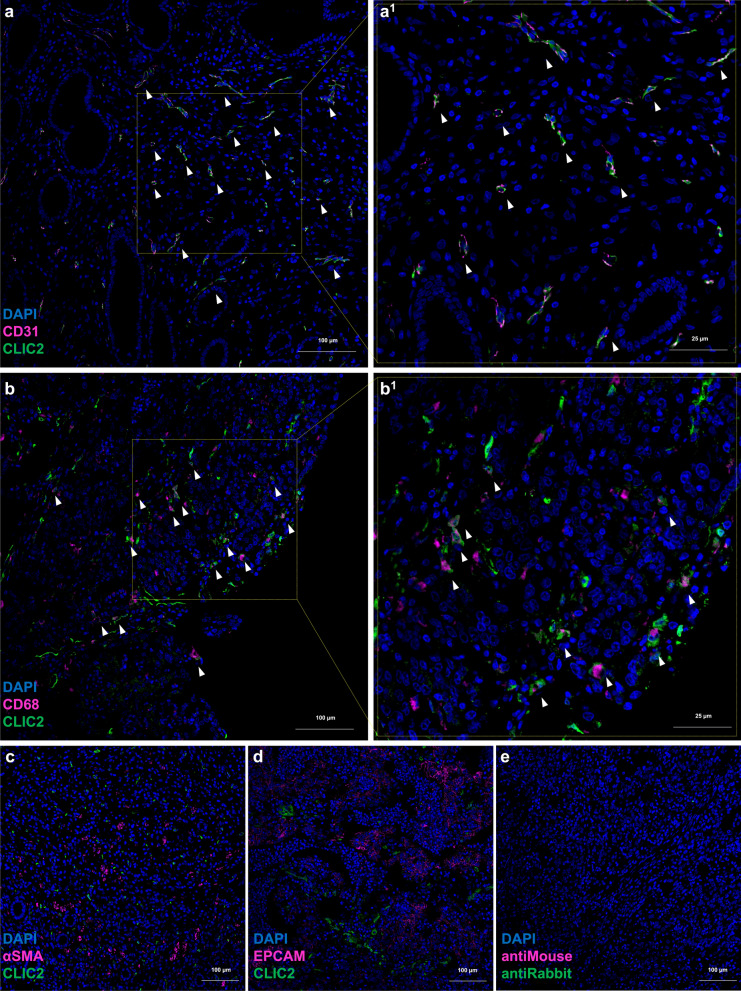
Due to the extreme heterogeneity of GC, we investigated whether CLIC2 gene expression could be specifically associated with the tumor itself or not. To infer tumor purity from bulk RNA sequencing data, we applied the ESTIMATE algorithm [15] as it was also implemented on TCGA Samples, retrieving IMMUNE (IS), STROMAL (SS), and ESTIMATE (ES) scores and subsequently, we correlated the expression of CLIC2 with these three scores. We found that CLIC2 gene expression was strongly and positively correlated with all three scores, with a Spearman’s rho of 0.743 with IS (Fig. 2a), 0.789 with SS (Fig. 2b), and 0.847 with ES (Fig. 2c), indicating that CLIC2 expression increases with the presence of immune and stromal cells in GC tissues. As a control of our approach, we repeated this analysis by comparing the expression of CD3 (a T-cell marker), ACTA2 (a fibroblast marker), and EPCAM (an epithelial cell marker) with the three scores and, as expected, we found that CD3 was significantly and positively related to IS (rho = 0.847), ACTA2 to SS (rho = 0.762), and EPCAM was significantly and negatively related to all the scores (rho = −0.250) (Supplementary Fig. 1). Next, to determine the cell types expressing CLIC2 in normal and dGC tumor tissues, we analyzed Kumar’s single cell sequencing gastric dataset [16], identifying 13 cell populations in normal (Fig. 2d) and tumor (Fig. 2e) samples. In tumor samples, 5 cell populations were statistically more represented respect to normal samples (Fig. 2h: Proliferative Cells—PC, B-cells, Enterocyte, Mast cell and Macrophage), while 3 cell populations were more represented in normal tissues compared to tumor samples (Fig. 2h: Gland Mucous Cells—GMC, Enteroendocrine and Pit Mucous Cells—PMC). Five populations did not show statistically significant changes. Interestingly, CLIC2 expression (Fig. 2f, g) was detected in more than 10% of the cells of a population in 3 cases: EC (endothelial cell), macrophage and fibroblast. Among these populations only macrophages were statistically over-represented in Tumor (Fig. 2h). To confirm this analysis, we performed multiple co-staining of Clic2 with main cytotype markers. We carried out immunofluorescence staining of tissue sections from formalin-fixed, paraffin-embedded human biopsies of seven dGC patients co-staining anti-Clic2 with anti-CD31 to reveal tissue endothelial cells or anti-CD68 for macrophages or anti-αSMA for fibroblasts or anti-EpCam for epithelial cells. As expected, in the disordered region of the tumor biopsy, Clic2 signal could be retrieved in association with CD31 (Fig. 3a) or CD68 (Fig. 3b) but not with αSMA (Fig. 3c) or EpCam (Fig. 3d), demonstrating that Clic2 is a protein characteristic of tumoral endothelial cells and tumor macrophages (Single channels images of Fig. 3 merges are shown in supplementary Fig. 2).
Fig. 2.
CLIC2 expression is gastric cancer tissues is restricted to endothelial cells and macrophages. Analysis of TCGA STAD dataset to infer tumor purity, applying ESTIMATE algorithm and correlating CLIC2 gene expression with a immune score, b stromal score, c ESTIMATE score. Gene expression based single-cell clustering of d normal gastric mucosa and e diffuse gastric cancer (see legend for more detail about color coding of cell populations; EC—Endothelial Cells, GMC—Gland Mucous Cells, PC—Proliferative Cells, PMC—Pit Mucous Cells, SM—Smooth Muscle Cells). CLIC2 presence (violet dots) or absence (grey dots) in f normal gastric mucosa and g diffuse gastric cancer single-cell clusters identified in d and e. h Odd ratio of cell populations presence in gastric cancer compared to normal samples
Fig. 3.
In dGC samples CLIC2 is expressed by tissue endothelial cells and macrophages. Confocal microscopy images (10X magnification) of immunofluorescence staining of representative tissue slides from 7 dGC patient samples. Images show co-staining of Clic2 with a CD31 as endothelial cells marker, b CD68 as pan-macrophages marker, c αSMA as fibroblast marker and d Epcam as epithelial cells marker. Panels “1” show a magnification of respective areas. White arrows indicate cells co-expressing both assayed proteins. e Negative controls
Clic2 intracellular expression follows the secretory pathway
Based on our previous findings we decided to investigate more about Clic2 expression and role in macrophages within the gastric cancer microenvironment. First, we aimed to investigate its intracellular localization to gain more insight into its function and, to this aim, we used the THP-1 cells as model, as they are monocytic cells that can be easily differentiated to macrophages. THP-1 monocytes were treated with low doses of PMA (5 ng/ml) to prevent pre-polarization towards either anti- or pro-inflammatory phenotypes [17], and then subjected to confocal immune fluorescence analysis (Fig. 4). Z-axis projection of images clearly revealed that Clic2 protein starts from the ER network, stained by the ER marker KDEL (Fig. 4, panel a), crosses over the Golgi complex (Fig. 4, panel b: compare anti-Clic2 to GM130 Golgi marker) and to reach the plasma membrane as shown by co-localization with the β1-integrin (Fig. 4, panel c), following the secretory pathway, in agreement with the literature [11] that documented Clic2 secretion in brain tumors.
Fig. 4.
CLIC2 follows the canonical secretory pathway. Clic2 intracellular staining in THP-1 cells differentiated to unpolarized macrophages visualized by confocal microscospy (63X magnification). Dapi staining of nuclei is showed in blue (a1, b1, c1), Clic2 signal is showed in green (a2, b2, c2), cell compartments are showed in red (a3 for endoplasmic reticulum using anti-KDEL; b3 for Golgi apparatus using anti-GM130; c3 for plasma membrane using anti-Integrin β1). Merged signals are showed in a4, b4, c4. For the staining showed in panels a and b, differentiated cells were fixed and permeabilized, for the staining of panel c, differentiated cells were fixed without permeabilization
Clic2 influences monocyte to macrophages differentiation
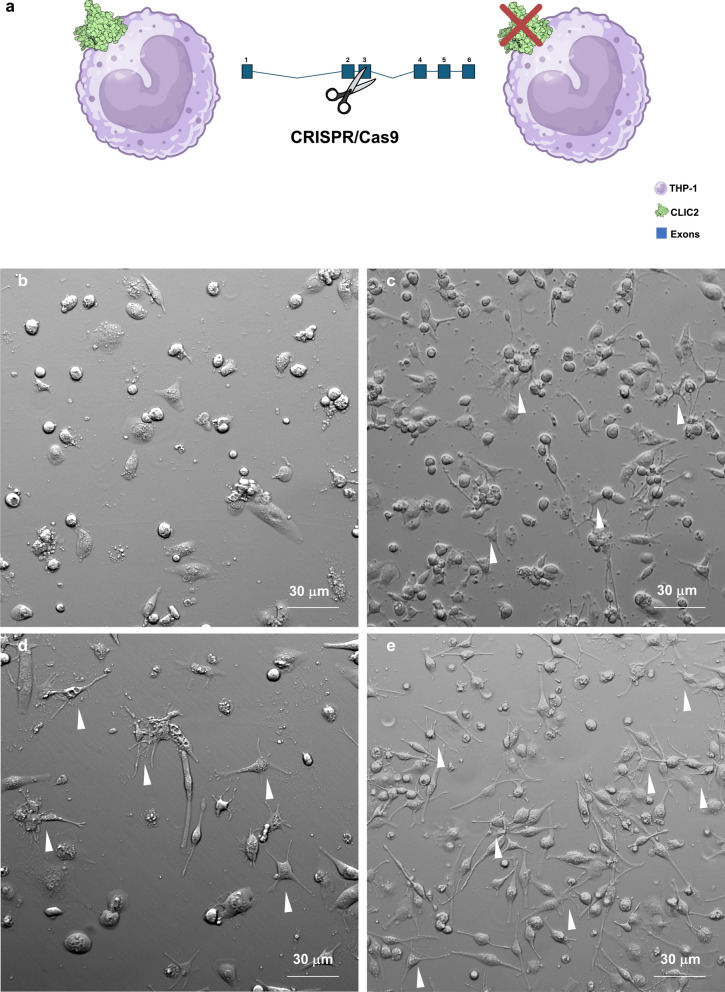
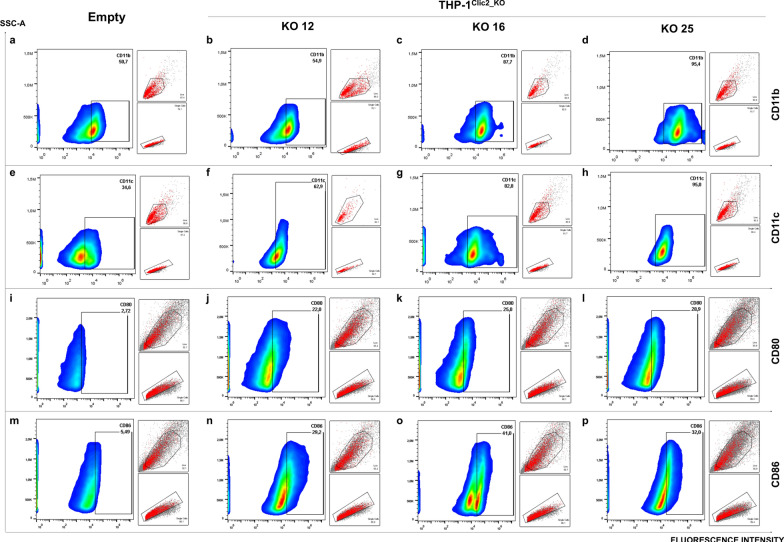
To investigate the functions of Clic2 in macrophages, we abrogated CLIC2 expression in THP-1 cells using the CRISPR/Cas9 approach (Fig. 5a). We checked cell viability with annexin V/PI staining, without measuring significant apoptosis in CLIC2 knockout (THP-1CLIC2_KO) cells compared to THP-1 control (LC) (Supplementary Fig. 3). Next, we randomly isolated three clones (12, 16, 25, Supplementary Fig. 4) for the next experiments. Surprisingly, upon differentiating both the THP-1 control (LC) and THP-1CLIC2_KO cells, we saw a notable change in morphology. Control cells showed the typical flat and round shape (Fig. 5b), while the THP-1CLIC2_KO cells displayed a spindly shape with multiple extensions of the plasma membrane (Fig. 5c–e). This suggested that CLIC2 could play a role in regulating the differentiation of monocytes and thus we measured membrane expression of some common markers. THP-1CLIC2_KO cells had increased expression of both CD11b (mean of the clones 81% vs 49% of LC, Fig. 6a–d) and CD11c (mean of the clones 81% vs 39% of LC, Fig. 6e–h) compared to LC, as well as for CD80 (mean of the clones 25% vs 3% of LC, Fig. 6i–l) and CD86 (mean of the clones 34% vs 5,5% of LC, Fig. 6m–p). These results showed that removing CLIC2 from monocytes driven cells differentiation to a dendritic cell-like phenotype, which likely reflects in an increased propensity to antigen presentation.
Fig. 5.
CLIC2 influences macrophages differentiation. a CLIC2 expression was abrogated in THP-1 cells using CRISPR/Cas9, then phenotype and molecular profiling was carried out. Control cells (b) and Clic2 knock out clones (c—KO 12; d—KO 16; e—KO 25), were differentiated to unpolarized macrophages and then observed to the microscope. Gradient contrast images (20X) shows altered morphology of THP-1CLIC2_KO cells after induction of differentiation to M0 macrophages. White arrows evidences dendritic-like macrophages
Fig. 6.
THP-1CLIC2_KO macrophages display enhanced differentiation and increased antigen presentation markers. THP-1 control cells (a, e, i, m) and three THP-1.CLIC2_KO clones (b–d; f–h j–l; n–p), were differentiated to unpolarized macrophages and analyzed by flow cytometry for common membrane markers such as CD11b (a–d), CD11c (e–h), CD80 (i–l), CD86 (m–p). Graphs show fluoresce intensity in relation to SSC-A physical parameter, smaller boxes display gating strategies (first gating on live cells FSC-A/SSC-A, second gating on single cells FSC-A/FSC-H)
Clic2 influences cytokine secretion and Jak/Stat signaling in macrophages
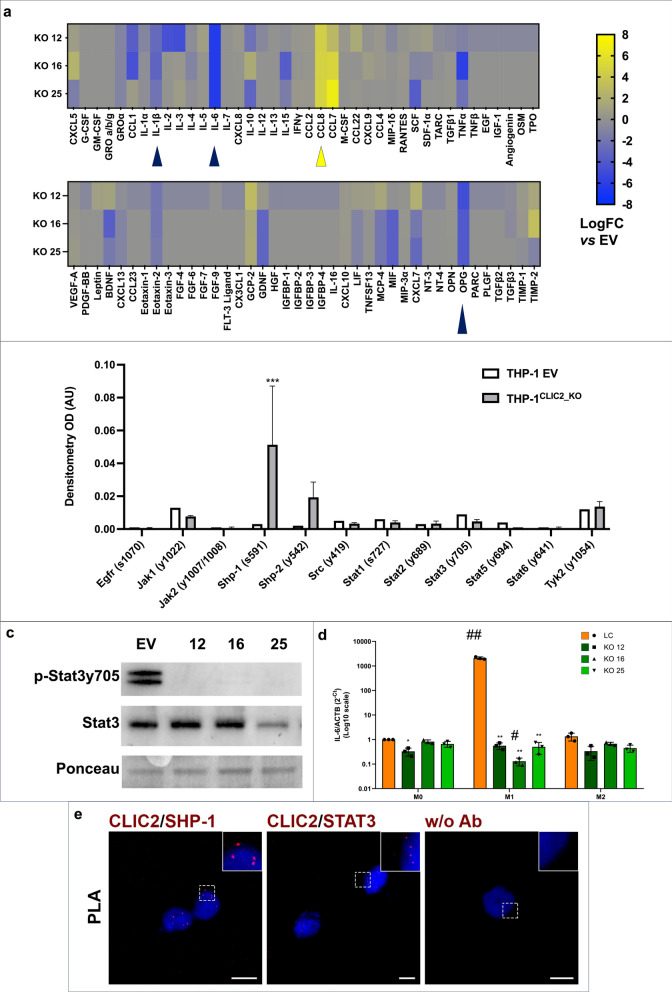
Macrophages are among the most influencing cells of the tumor microenvironment (TME) and cytokine secretion by macrophages can boost or totally impair the anti-tumor response by immune system [18]. Given the above described in-silico and in-vitro results, we wondered whether CLIC2 absence in differentiated THP-1 cells could somehow affect cytokine secretion. To this aim, we collected conditioned media of differentiated LC and THP-1CLIC2_KO cells to simultaneously analyze 80 secreted cytokines (Supplementary Table 4). We only focused on cytokines up- or down- regulated at least 2 LogFC compared to control, in all three clones. In THP-1CLIC2_KO cells compared to LC, we found increased secretion of CCL7 and CCL8, two powerful chemoattractants and immune activators (Fig. 7a, Supplementary Fig. 5) [19], with CCL8 being statistically significant (p = 0.0126). Conversely, reduced secretion of the tumor-supporting IL-6 (p < 0.0001), Osteoprotegerin (OPG, p = 0.0003) and IL-1β (p = 0.0066) were measured (Fig. 7a, Supplementary Fig. 5) [20]. Furthermore, as the Jak/Stat pathway handles the regulation of monocyte-to-macrophage differentiation and polarization, we monitored its status in differentiated LC and THP-1CLIC2_KO cells by simultaneously measuring the phosphorylation of 12 proteins of this signaling network (Supplementary Table 5). Interestingly, we observed in all clones a significant increase in phosphorylation (and thus activation) of the membrane-associated phosphatase Shp1 (p = 0.0007) and an increase of Shp2 phosphorylation even if not statistically significant (Fig. 7b). This was accompanied by the consequent absence of their main target Stat3, that resulted not phosphorylated (Fig. 7c) and, consequently, not transcriptionally active. Stat3 is, indeed, the main transcription factor regulating IL6 transcription. CLIC2 absence in differentiated macrophages impaired Stat3 signaling and IL6 transcription (Fig. 7d), as also demonstrated during macrophages polarization to pro- or anti-inflammatory phenotype (respectively M1 and M2).
Fig. 7.
THP-1CLIC2_KO macrophages can positively influence the tumor microenvironment. a THP-1 control cells and three THP-1CLIC2_KO clones were differentiated to unpolarized macrophages and 80 cytokines were simultaneously measured in their culture media using antibody arrays. Dot blot signals of all samples were normalized and compared to those of controls, then LogFC of THP-1CLIC2_KO clones vs controls was calculated. Blue arrows indicate cytokines secreted at least −2 LogFC from all three clones respect to controls. Yellow arrows indicate cytokines secreted at least + 2 LogFC from all three clones respect to controls. The details of data normalization are reported in materials and methods. Statistical significance was calculated using GraphPad Prism 10, unpaired t-test of densitometry of CLIC2-KO clones vs LC (CCL7 p = 0.5188; CCL8 p = 0.0126; IL-1β p = 0.0066; IL-6 p < 0.0001; OPG p = 0.0003). b THP-1 control cells and three THP-1CLIC2_KO clones were differentiated to unpolarized macrophages and Jak/Stat pathway phosphorylation was explored using antibody arrays. Densitometric analysis was carried out and mean of KO clones signals were visualized as histograms. Statistical analysis was performed with GraphPad Prism 10, 2-way Anova, Shp-1 p = 0.0007. c Western blot analysis of Stat3 phosphorylation in THP-1 controls and THP-1CLIC2_KO clones differentiated to M0 macrophages. d Real-time PCR analysis of IL6 transcription in THP-1 controls and THP-1CLIC2_KO clones differentiated to M0 macrophages and further polarized to M1 or M2 phenotype. e Proximity Ligation Assay (PLA) to assess Clic2/Shp-1 and Clic2/Stat3 proximity in fixed and permeabilized THP-1 cells differentiated to unpolarized macrophages. (e1, e2) Representative images of PLA signals (red puncta) in THP-1 cells differentiated to unpolarized macrophages. The primary antibodies used are indicated in red. Nuclei were counterstained with DAPI (blue). In (e3) “w/o Ab” indicates the negative control performed in the absence of primary antibodies. Scale bars, 10 µm
CLIC2 physically associates with Shp1 and Stat3 in differentiated macrophages
To investigate whether Clic2 directly interacts with components of the Shp1-Stat3 signaling axis, we performed in situ proximity ligation assays (PLA) in THP-1 cells differentiated into M0 macrophages. As shown in Fig. 7e, PLA signals were clearly detected in both Clic2/Shp1 and Clic2/Stat3 pairs, indicating that Clic2 is part of a protein complex involving these two key signaling molecules. The presence of punctate fluorescent signals confirms close physical proximity (<40 nm), strongly supporting a direct molecular association under physiological conditions. These results suggest that Clic2 may act as a scaffold or stabilizing partner for Shp1 and Stat3 in macrophages, thereby modulating downstream signaling responses.
Discussion
Our study provides the first evidence that Clic2 in gastric cancer governs macrophage differentiation, thereby promoting tumor progression through modulation of the tumor microenvironment. Additionally, we reveal a strong correlation between CLIC2 gene expression and genomically stable GC, indicating that Clic2-positive macrophages may play a key role in establishing the immunosuppressive phenotype characteristic of this tumor type. We observed CLIC2 gene expression in macrophages, as well as in endothelial cells and fibroblasts, underscoring its significant influence in maintaining the balance within the tumor microenvironment. Finally, we demonstrate that Clic2 supports Stat3 signaling, a prominent pathway in tumor progression. In the heterogeneous context of gastric cancer, we identified a gene network intersecting with CLIC2 that closely clustered with the diffuse histotype of gastric cancer (Fig. 1a, Supplementary Table 1), but even more interestingly, this network was significantly linked to patients with microsatellite stable (MSS) disease (Fig. 2b). This finding was particularly interesting as microsatellite stability in tumors is closely linked to the immune system's ability to efficiently respond to cancer cell proliferation [21]. MSS tumors are less immunoreactive compared to microsatellite instability (MSI) tumors, which can more effectively trigger immune system activation, also thanks to a more efficient antigen presentation of the dendritic cells to the lymphocytes [22]. As our experiments demonstrate that removing CLIC2 from THP-1 cells increases the expression of antigen presentation markers in unstimulated macrophages, it could be possible that the increased expression of CLIC2, specifically measured in dGC and in patients with MSS disease, could be related to the reduced capability of activation of the immune system characteristic of this subset of GC patients. This observation aligns with the understanding that diffuse-type gastric cancer is considered a"cold"tumor, where the immune system is less effective in controlling tumor development, and immunotherapy is often ineffective [2]. In addition, many categories that connected the genes composing the CLIC2 regulon suggested some functional relation of this regulon with immune cells, as indicated for example by the top-ranked SHP2_PATHWAY, HEMOSTASYS, CELL SURFACE INTERACTIONS AT THE VASCULAR WALL, HALLMARK_IL2_STAT5_SIGNALING, and HALLMARK_INTERFERON_GAMMA_RESPONSE. Effectively, we could demonstrate Clic2 expression in endothelial cells and in macrophages composing the tumor microenvironment of dGC patients (Fig. 3). CLIC2 expression in endothelial cells and in myeloid cells was also evidenced in normal colon tissues surrounding colon cancers [23], although in that context it was demonstrated that CLIC2 was mainly related to the correct maintenance of tight junctions in endothelial cells. Collectively, our data indicated the following: a) The genes in the CLIC2 regulon were directly and indirectly linked to the immune system; b) Both the CLIC2 regulon and the CLIC2 gene were overexpressed and strongly associated with genomically stable, diffuse-type gastric cancer; c) There was a strong relationship between CLIC2 and the tumor microenvironment (TME). These findings strongly suggested that CLIC2 could contribute to immune regulation within the TME by promoting an immune suppressive environment. When we investigated the intracellular localization of Clic2 to deduce its molecular function, our results were partially consistent with previous findings by Ozaki et al. [11], who described Clic2 secretion in brain tumors, where it inactivates MMP14, thus inhibiting cell migration and invasion. Our data confirm that Clic2 follows the canonical secretory pathway (Fig. 4), but we propose a different role for Clic2 in macrophages. In our model of CLIC2 knockout monocytic cells, we observed that the absence of CLIC2 during the differentiation of monocytes to macrophages resulted in cells adopting a dendritic-like phenotype. These macrophages exhibited several membrane protrusions resembling dendrites (Fig. 5) and were more efficiently differentiated and prone to antigen presentation at the basal state (Fig. 6). Notably, THP-1CLIC2_KO macrophages increased the secretion of strongly chemoattractant cytokines CCL7 and CCL8 (Fig. 7), which can efficiently attract several immune cells [24], even if they can display wide effect with controversial contributions, as they also have been reported to sustain tumor growth [25]. Additionally, THP-1CLIC2_KO macrophages produce lower levels of OPG that was reported to cooperate with IL-1b to promote breast cancer progression [26]. Finally, THP-1CLIC2_KO macrophages also showed a negatively regulation of the IL6/STAT3 pathway, a major tumor-sustaining and immunoinhibitory signaling pathway in the TME [27], through the hyperactivation of the phosphates Shp1, that controls many of the intracellular pathways of macrophages [28]. To better define the molecular link between Clic2 and this pathway, we performed proximity ligation assays (PLA) in differentiated THP-1 macrophages and demonstrated that Clic2 physically interacts with both Shp1 and Stat3 (Fig. 7e), supporting the existence of a functional complex. This observation suggests that Clic2 may act as a molecular scaffold or stabilizer that modulates the spatial proximity between Shp1 and Stat3, possibly limiting Shp1 access to Stat3 under physiological conditions. In the absence of Clic2, Shp1 may gain greater accessibility to Stat3, leading to enhanced dephosphorylation and consequent downregulation of Stat3-dependent transcriptional programs. Collectively, our findings are coherent and robust, validated by our bioinformatic and biological approaches. We propose a new role for CLIC2 in diffuse gastric cancer, highlighting its central contribution to macrophage differentiation toward an immune suppressive phenotype. Future research should focus on investigating whether it is possible to interfere with CLIC2 in-vivo to revert the immunological state of diffuse gastric cancer and improve host antitumor immune response. However, the absence of a murine model due to the lack of the CLIC2 gene in mice is a significant obstacle. This challenge could be addressed using ex-vivo models, such as patient-derived organoids (PDOs) which retain the microenvironment composition in cell culture [29].
Materials and methods
Regulon analysis
A regulon is a group of genes that are regulated as a unit, generally controlled by the same regulatory gene that expresses a protein acting as a repressor or activator. Elective regulatory genes are transcription factors which bind to specific regions of the DNA and together with cofactors and other proteins influence the transcriptional rate of a specific set of target genes (TGs) collectively known as the TF’s regulon. The combined interactions of all TFs to their target genes are often referred to as a gene regulatory network (GRN), a simplified representation of the underlying regulatory circuits. Coupling GRNs with activity inference algorithms can facilitate the interpretation of transcriptomics data and provide a more effective means of understanding the underlying regulatory mechanisms in the system of interest [30]. About inference algorithms ARACNe [31] a well-known algorithm to reconstruct GRN from transcriptomic data. Because regulation can occur by both direct (TF-target) and indirect interactions (TF-TF-target), the ARACNe algorithm uses the data processing inequality (DPI) theorem to enrich the regulons with direct TF-target interactions. Carro et al. showed that a C/EBPβ and STAT3 regulon is linked to the ‘mesenchymal’ gene expression signature of High-grade gliomas (HGGs) [32]. Finally, Alvarez et al. [33] propose that the expression of the transcriptional targets of a protein, collectively referred to as its regulon, represent an optimal multiplexed reporter of its activity extending the concept of regulon to every protein targets. As previously described [5], we selected the regulon of CLIC2 and plotted the expression of its genes clustering the samples according to Ward agglomeration method considering a Euclidean distance. Positive normalized enriched score (NES) highlights a more active subnetwork associated with CLIC2. We evaluated, by linear modeling, the association of CLIC2’s single sample NES with clinical features and depicted its associations with microsatellite instability. Wilcoxon rank-sum test was performed to determine a statistical difference (p-value < 0.05) in the expression of CLIC2 between the normal tissue and GC and among Normal tissue, dGC and iGC two by two.
Network and expression level analysis
Network visualization was carried out by Cytoscape platform and its apps (https://cytoscape.org). A specifically table report was obtained with the relative open access online Enrichment Databases results and ordered by decreased weight in the network (Supplementary Table 2). We checked the difference distribution between Normal Tissue Samples and Tumor Samples by Wilcoxon Rank Sum Test and between Normal Tissue Samples, Intestinal and Diffuse Samples subgroups by the same test. Only p-value < 0.05 was considered statistically significant. All analyses were performed by R/Bioconductor environment (https://www.r-project.org/; https://bioconductor.org/).
ESTIMATE correlation
The CLIC2 expression level on TCGA Samples was correlated with the ESTIMATE (Estimation of STromal and Immune cells in MAlignant Tumor tissues using Expression data) tool’s three scores respectively, as described by Yoshihara et al. [15]. The correlation between data series was investigated by Spearman’s Rank Test, statistical significance was established on p-value < 0.05.
Single-cell analysis
10X data retrieved from Kumar 19 were processed according to best practice. Briefly, low-quality cells with a high percentage (> 15%) of reads from mitochondrial genes and low or high gene counts were removed from further analyses. Seurat v5 standard analysis workflow [34] run followed by integration of all the samples with rpca reduction. Optimization of cluster parameters and annotation according to ScType algorithm [35] given a list of 73 different cell population markers [35–37], was made resulting in 13 cell populations. The uniform manifold approximation and projection (UMAP) algorithm was used to depict the cell populations and for each of them Fisher test was performed to pinpoint the significant odd ratios (OR) (FDR-adj.p-value < 0.05 & abs (OR) > 1) between GC vs Normal.
Immunofluorescence on gastric sliced biopsies
Six biopsies were collected from patients affected by diffuse-type gastric cancer, in accordance with the approved guidelines of the World Medical Association’s Declaration of Helsinki and informed consent and ethical committee authorization with protocol number 20180042426 of the COMITATO ETICO UNICO REGIONALE PER LA BASILICATA. Immunofluorescences were performed as previously described [38], with some modification. Briefly, biopsies were fixed with formalin, embedded in paraffin and then sliced (thickness of the slices = 3 μm) using a Bio-Optica microtome HistoCore BIOCUT (RM2235). The samples were incubated overnight with primary antibodies in hystoblock (1:50 rabbit anti-Clic2 primary antibody—AB175230 Abcam, Cambridge, UK; 1:200 mouse anti-EpCAM primary antibody—2929 Cell Signaling, Massachusetts, USA; 1:1000 mouse anti-CD31 primary antibody—3528 Cell Signaling, Massachusetts, USA; 1:50 mouse anti-Smooth Muscle Actin primary antibody—sc53142 Santa Cruz Biotechnology, Dallas, Texas, USA; 1:50 mouse anti-CD68 primary antibody—168M9 Merck, Rahway, New Jersey, USA). Subsequently, the slices were incubated with secondary antibody Alexa Fluor anti-rabbit 488 (Cell Signaling 4412S) 1:1000 and with Alexa Fluor anti-mouse 647 (Invitrogen A21235) 1:1000 and finally with 4′,6-diamidino-2-phenylindole (DAPI—ThermoFisher D1306) 1:5000. The samples were analyzed through confocal microscopy using Olympus IX83 equipped with Olympus FLUOVIEW (FV3000) using two optical lenses, 10x (UPlanXApo 10X/0.40 ∞/0.17/OFN26.5) and 20X (UPlanXApo 20X/0.80 ∞/0.17/OFN26.5). fourfold magnifications of selected areas were performed during post production of images.
Cell cultures and macrophages differentiation
THP-1 monocytes were acquired from ATCC (ATCC® TIB-202; Manasas, VA, USA) and were cultured in RPMI-1640 with L-glutamine, supplemented with 10% inactivated fetal bovine serum (FBS; Gibco, Carlsbad, CA, USA), 100 U/mL penicillin, and 100 g/mL streptomycin (Gibco, Carlsbad, CA, SA), 1X Normocin (Invivogen, San Diego, USA) and 0.05 mM 2-mercaptoethanol (Gibco, Carlsbad, USA) in a 5%-CO2 humidified incubator at 37 °C. Cell concentration was maintained in a range from 2 × 105 cells/ml to 8 × 105 cells/ml. To induce cell attachment and M0 macrophage differentiation, cells were treated with 8 nM (5 ng/ml) PMA for 24 h, followed by a 72 h recovery phase [17]. When polarization to pro- (M1) or anti-inflammatory (M2) phenotype was requested, M0 macrophages were treated, respectively, with 250 ng/ml LPS and 20 ng/ml INFg for M1, or 20 ng/ml IL-4 and 20 ng/ml IL-13 for M2, for further 48 h, before analysis. THP-1CLIC2_KO cells were generated by lentiviral infection as previously described [39, 40]. Briefly, HEK293T cells (CRL-3216, ATCC Manasas, VA, USA), were used to produce lentiviral particles containing empty vector (lentiCRISPR v2 was a gift from Feng Zhang (Addgene plasmid # 52,961; http://n2t.net/addgene:52961; RRID:Addgene_52961) or the same vector, containing gRNA targeting exon 3 of CLIC2 mRNA (Clic2 gRNA sequence GAACGGAGGATTGGTACCTG, PAM sequence GGG) and lentiviral particles were then used to infect THP-1 cells by spinoculation. Transduced cells were selected by treating cells with 1 μg/ml puromycin for 2 weeks, then puromycin selection was removed and cells were plated in 96-well plate at concentration of 5 cells/ml in 100 μl of complete media, until visible clones could be detected and amplified for western blotting screening analysis (Supplementary Fig. 4).
Confocal microscopy and gradient-contrast imaging
To perform immunofluorescence analysis, THP-1 cells were plated in 12-well plates with coverslips on the bottom and differentiated to macrophages was induced as described above. Immunofluorescence was performed as previously described [41], briefly, cells were washed in phosphate-buffered saline (PBS), fixed in 4% paraformaldehyde and permeabilized with 0.1% Triton X-100 in PBS for 5 min. Thereafter, cells were stained with antibody anti-b1 integrin 1:50 (Santa Cruz Biotechnology, Dallas, USA) to visualize plasma membrane; antibody anti-GM130 1:100 (BD Transduction laboratories) was used to visualize the Golgi (BD Transduction Laboratory, Franklin Lakes, NJ, USA); antibody anti-KDEL 1:100 was used to visualize ER (Enzo Life Sciences, Farmingdale, NY, USA) and antibody anti-Clic2 1:100 (Abcam, Cambridge, UK) for 1 h. After that samples were stained with anti-rabbit Alexa-488 1:800 (Jackson Immunoresearch, West Grove, Pennsylvania, USA) or anti-mouse CY3 1:1500 (Jackson Immunoresearch, West Grove, Pennsylvania, USA) for 1 h. Finally, DAPI staining was used to visualize the nuclei. After washing, coverslips were mounted with a Vecta-mount medium (Vector laboratories, Newark, California, USA). Images were acquired with a laser scanning confocal microscope TCS SP5 (Leica MicroSystems, Mannheim, Germany) equipped with a plan Apo 63X, NA 1.4 oil immersion objective lens. Pictures were processed using LAS-AF Software (Leica MicroSystems, Germany) to reconstruct the x-axis projection using stack images. For gradient contrast images of THP-1 macrophages, cells were differentiated as described above and then were observed using APEXVIEW APX100 benchtop fluorescence microscope and images were taken using the Gradient Contrast Method using the optical lens 10x (UPlanXApo 10X/0.40 ∞/0.17/OFN26.5).
Proximity ligation assay (PLA)
PLA was carried out on fixed and permeabilized cells using the Duolink® In Situ Red Starter Kit (Mouse/Rabbit; Sigma-Aldrich), according to the manufacturer’s instructions. The following primary antibodies were used: 1:100 rabbit anti-Clic2 primary antibody - AB175230 Abcam, Cambridge, UK; 1:100 mouse anti-Stat3 primary antibody - 9139 Cell signaling, Massachusetts, USA; 1:25 mouse anti Shp-1 primary antibody - sc-7289 Santa Cruz Biotechnology, Dallas, USA; 1:100 rabbit anti-Jak1 primary antibody - 3344 Cell signaling, Massachusetts, USA. Images were acquired on a laser scanning confocal microscope (TCS SP5; Leica MicroSystems) equipped with a plan Apo 63X, NA 1.4 oil immersion objective lens.
FACS staining and analysis
THP-1 macrophages were induced as described above, then cells were washed with PBS and gently collected by scraping. Cells were centrifuged 5’ at 300 g and then resuspended in PBS with 0.5% BSA, 2 mM EDTA and 10% FcR blocking reagent for 10’ at 4 °C in the dark. Subsequently, samples were left unstained for background fluorescence determination or stained with primary antibodies (final concentration 1:100, more details on antibody codes and producers in Supplementary Table 3) for further 45’ on ice in the dark. At the end of the incubation, samples were centrifuged and washed 3 times with PBS with 0.5% BSA, 2 mM EDTA and then acquired with a Beckman DxFlex (Beckman Coulter, Brea, USA) or with a BD Melody (BD Biosciences, Franklin Lakes, USA) flow cytometer. Analysis of the experiments and graph were produced with FLOWJO software (BD Biosciences, Franklin Lakes, USA), using unstained sample to setup background fluorescence. Gating workflow was the following: first gating on live cells FSC-A/SSC-A, then second gating on single cells FSC-A/FSC-H. This population was used for data analysis.
Cytokines and antibodies arrays
THP-1 macrophages were produced as described above, then cell culture conditioned media were collected, and centrifuged for 10’ at 300 g to remove cell debris, while macrophages were washed with PBS and then collected by scraping before lysis. Cytokine production was examined using Raybiotech Human Cytokine Array C5 (AAH-CYT-5-8) while the Jak/Stat pathway was examined using Human Phosphorylation Multi-Pathway Profiling Array C55 (AAH-PPP-1-4) following manufacturer instructions. Briefly, antibody array membranes were incubated with a blocking solution, then washed and incubated overnight at 4 °C with samples with gentle rocking. The day after, membranes were washed and subsequently incubated with primary antibodies cocktail for 2 h RT, then washed again before incubation with secondary antibodies for 2 h RT. Finally, membranes were washed again before incubation with detection buffer and imaged with Chemidoc XRS + system (BIORAD, Hercules, USA). Cell densities were measured by processing acquired images with Fiji Image J (https://imagej.net/software/fiji/) measuring the integral density of each spot. Data analysis for arrays comparisons, was performed following manufacturer instructions.
Western blotting
THP-1 macrophages were produced as described above, then cells were washed with PBS and collected by scraping in RIPA lysis buffer (Cell Signaling, Danvers, USA). Western blotting analysis was performed as previously described [42]. Briefly, whole cell lysates (30 µg) were resolved on Novex NuPage 12–4% (ThermoFisher Scientific, Carlsbad, USA) and then transferred to polyvinylidene difluoride membrane (Millipore, Bedford, USA). After transfer, membranes were incubated with Ponceau red solution (Sigma Aldrich, Sant Louis, USA) and images were acquired using Chemidoc XRS + system (Biorad, Hercules, USA). Membranes were then washed, blocked in TBS containing 5% non-fat dry milk (Biorad, Hercules, USA) before incubation with primary antibodies (1:1000) followed by incubation with horseradish-peroxidase-linked mouse or rabbit IgG (1:3000) (GE Healthcare Amersham, Little Chalfont, UK) in TBS containing 2% non-fat dry milk (Bio-Rad Laboratories, Hercules, USA). Protein signals were detected by chemiluminescence using the ECL System (Biorad, Hercules, USA) and a Chemidoc XRS + system (Biorad, Hercules, USA).
Real time
THP-1 macrophages were produced as described above, then cells were washed with PBS and collected by scraping in RNA extraction buffer, and RNA was isolated using PureLink RNA kit (Thermo Fisher Scientific), following manufacturer’s instruction. Reverse transcription and RT-PCR were performed as previously described [43, 44], as well as for Real-Time quantification of gene expression, using DDCt method [39]. Nucleotide sequences of primers used for Real-Time experiments are the following: human ACTB forward—5′ CCGACAGGATGCAGAAGGAG 3′; human ACTB reverse—5′ GCCTAGAAGCATTTGCGGTG 3′; human IL6 forward—5′ TCAATGAGGAGACTTGCCTG 3′; human IL6 reverse—5′ TGGGTCAGGGGTGGTTATTG 3′. Statistical analysis and graphical visualization were performed using GraphPad Prism 8.0.1. Statistical comparisons of CLIC2KO clones versus LC were made with 2-way ANOVA.
Supplementary Information
Author contributions
Vi.Lo., P.M. and Gio.Ca. contributed to the acquisition, analysis and interpretation of data; P.Z. contributed to the analysis of data and have drafted the work; G.D.P., Giu.Ce., M.L., C.S., G.A., V.P., contributed to the acquisition of data; S.R., S.L., T.N., contributed to drafting the work; G.P., C.B., contributed to the the analysis of data; Va.Lu., E.A., M.R.M., R.G., contributed to interpretation of data and revised the work; C.D.S., F.D.V. and P.R. revised the work; G.F. and F.A., contributed to the conception of the work and to the design of the experiments, to the interpretation of the data, to the draft and to the revision of the work. Each author have approved the submitted version (and any substantially modified version that involves the author's and have agreed both to be personally accountable for the author's own contributions and to ensure that questions related to the accuracy or integrity of any part of the work, even ones in which the author was not personally involved, are appropriately investigated, resolved, and the resolution documented in the literature.
Funding
The study was supported by Italian Ministry of Health, Finanziamento Ricerca Corrente 2023; Italian Ministry of Health, 5 × 1000 funds; Unione europea-Next Generation EU, CN00000041 “National Center for Gene Therapy and Drugs based on RNA Technology”; Unione europea-Next Generation EU, Missione 4 Componente 1 CUP B53C22001820006. The funders had no role in the study design, data collection and analysis, decision to publish or preparation of the manuscript.
Data availability
All data generated or analyzed during this study are included in this published article, and its supplementary information files, and are available upon reasonable request to the corresponding authors.
Declarations
Ethics approval and consent to participate
Human biopsies were collected from patients affected by diffuse-type gastric cancer, in accordance with the approved guidelines of the World Medical Association’s Declaration of Helsinki and informed consent and ethical committee authorization with protocol number 20180042426 of the COMITATO ETICO UNICO REGIONALE PER LA BASILICATA.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Viviana Longo, Pellegrino Mazzone, Giovanni Calice, Geppino Falco and Francesco Albano have contributed equally to the work.
Contributor Information
Geppino Falco, Email: geppino.falco@unina.it.
Francesco Albano, Email: francesco.albano@unina.i.
References
- 1.Laurén P. The two histological main types of gastric carcinoma: diffuse and so-called intestinal-type carcinoma. Acta Pathol Microbiol Scand. 1965;64:31–49. [DOI] [PubMed] [Google Scholar]
- 2.Iyer P, Moslim M, Farma JM, Denlinger CS. Diffuse gastric cancer: histologic, molecular, and genetic basis of disease. Transl Gastroenterol Hepatol. 2020;5:52–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bass AJ, et al. Comprehensive molecular characterization of gastric adenocarcinoma. Nature. 2014;513:202–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Yamaguchi H, Hsu J-M, Sun L, Wang S-C, Hung M-C. Advances and prospects of biomarkers for immune checkpoint inhibitors. Cell Rep Med. 2024;5: 101621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Russi S, et al. Gene regulatory network characterization of gastric cancer’s histological subtypes: distinctive biological and clinically relevant master regulators. Cancers. 2022;14:4961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Argenzio E, Moolenaar WH. Emerging biological roles of Cl− intracellular channel proteins. J Cell Sci. 2016;129:4165–74. [DOI] [PubMed] [Google Scholar]
- 7.Ozaki S, Mikami K, Kunieda T, Tanaka J. Chloride intracellular channel proteins (CLICs) and malignant tumor progression: a focus on the preventive role of CLIC2 in invasion and metastasis. Cancers. 2022;14:4890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Singh H. Two decades with dimorphic chloride intracellular channels (CLICs). FEBS Lett. 2010;584:2112–21. 10.1016/j.febslet.2010.03.013. [DOI] [PubMed] [Google Scholar]
- 9.Manori B, et al. Chloride intracellular channel (CLIC) proteins function as fusogens. Nat Commun. 2024;15:2085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Peretti M, et al. Chloride channels in cancer: Focus on chloride intracellular channel 1 and 4 (CLIC1 AND CLIC4) proteins in tumor development and as novel therapeutic targets. Biochim Biophys Acta (BBA): Biomembr. 2015;1848:2523–31. [DOI] [PubMed] [Google Scholar]
- 11.Ozaki S, et al. Chloride intracellular channel protein 2 is secreted and inhibits MMP14 activity, while preventing tumor cell invasion and metastasis. Neoplasia. 2021;23:754–65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ueno Y, et al. Chloride intracellular channel protein 2 in cancer and non-cancer human tissues: relationship with tight junctions. Tissue Barriers. 2019;7:1593775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Littler DR, et al. The enigma of the CLIC proteins: Ion channels, redox proteins, enzymes, scaffolding proteins? FEBS Lett. 2010;584:2093–101. [DOI] [PubMed] [Google Scholar]
- 14.Xu T, et al. Chloride intracellular channel protein 2: prognostic marker and correlation with PD-1/PD-L1 in breast cancer. Aging. 2020;12:17305–27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Yoshihara K, et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013;4:2612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kumar V, et al. Single-cell atlas of lineage states, tumor microenvironment, and subtype-specific expression programs in gastric cancer. Cancer Discov. 2022;12:670–91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Baxter EW, et al. Standardized protocols for differentiation of THP-1 cells to macrophages with distinct M(IFNγ+LPS), M(IL-4) and M(IL-10) phenotypes. J Immunol Methods. 2020;478: 112721. [DOI] [PubMed] [Google Scholar]
- 18.Chen S, et al. Macrophages in immunoregulation and therapeutics. Signal Transduct Target Ther. 2023;8:207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Metzemaekers M, Gouwy M, Proost P. Neutrophil chemoattractant receptors in health and disease: double-edged swords. Cell Mol Immunol. 2020;17:433–50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Qin X, et al. Cancer-associated fibroblast-derived IL-6 promotes head and neck cancer progression via the osteopontin-NF-kappa B signaling pathway. Theranostics. 2018;8:921–40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Greco L, Rubbino F, Dal Buono A, Laghi L. Microsatellite instability and immune response: from microenvironment features to therapeutic actionability—lessons from colorectal cancer. Genes. 2023;14:1169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Randrian V, Evrard C, Tougeron D. Microsatellite instability in colorectal cancers: carcinogenesis, neo-antigens. Immuno-Res Emerg Therapies Cancers. 2021;13:3063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ueno Y, et al. Chloride intracellular channel protein 2 in cancer and non-cancer human tissues: relationship with tight junctions. Tissue Barriers. 2019;7(1):1593775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Palomino DCT, Marti LC. Chemokines and immunity. Einstein. 2015;13:469–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Israr M, DeVoti JA, Papayannakos CJ, Bonagura VR. Role of chemokines in HPV-induced cancers. Semin Cancer Biol. 2022;87:170–83. [DOI] [PubMed] [Google Scholar]
- 26.Chung STM, Geerts D, Roseman K, Renaud A, Connelly L. Osteoprotegerin mediates tumor-promoting effects of Interleukin-1beta in breast cancer cells. Mol Cancer. 2017;16:1–14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Briukhovetska D, et al. Interleukins in cancer: from biology to therapy. Nat Rev Cancer. 2021;21:481–99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hong SY, et al. Targeting pathogenic macrophages by the application of SHP-1 agonists reduces inflammation and alleviates pulmonary fibrosis. Cell Death Dis. 2023;14(6):352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Seidlitz T, Koo BK, Stange DE. Gastric organoids: an in vitro model system for the study of gastric development and road to personalized medicine. Cell Death Differ. 2021;28:68–83. 10.1038/s41418-020-00662-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Müller-Dott S, et al. Expanding the coverage of regulons from high-confidence prior knowledge for accurate estimation of transcription factor activities. Nucleic Acids Res. 2023;51:10934–49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Margolin AA, et al. ARACNE: an algorithm for the reconstruction of gene regulatory networks in a mammalian cellular context. BMC Bioinf. 2006;7:S7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Carro MS, et al. The transcriptional network for mesenchymal transformation of brain tumours. Nature. 2010;463:318–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Alvarez MJ, et al. Functional characterization of somatic mutations in cancer using network-based inference of protein activity. Nat Genet. 2016;48:838–47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Hao Y, et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat Biotechnol. 2023;42:293–304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Ianevski A, Giri AK, Aittokallio T. Fully-automated and ultra-fast cell-type identification using specific marker combinations from single-cell transcriptomic data. Nat Commun. 2022;13:1-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zhang P, et al. Dissecting the single-cell transcriptome network underlying gastric premalignant lesions and early gastric cancer. Cell Rep. 2019;27:1934-1947.e5. [DOI] [PubMed] [Google Scholar]
- 37.Tsubosaka A, et al. Stomach encyclopedia: combined single-cell and spatial transcriptomics reveal cell diversity and homeostatic regulation of human stomach. Cell Rep. 2023;42: 113236. [DOI] [PubMed] [Google Scholar]
- 38.Guerriero I, et al. Exploring the molecular crosstalk between pancreatic bud and mesenchyme in embryogenesis: novel signals involved. Int J Mol Sci. 2019;4900(20):4900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Albano F, et al. Zfp36l2 role in thyroid functionality. Int J Mol Sci. 2021;22(17):9379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Albano F, et al. The expression of inhibitor of bruton’s tyrosine kinase gene is progressively up regulated in the clinical course of chronic lymphocytic leukaemia conferring resistance to apoptosis. Cell Death Dis. 2018;9(1):13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Filippone MG, Di Palma T, Lucci V, Zannini M. Pax8 modulates the expression of Wnt4 that is necessary for the maintenance of the epithelial phenotype of thyroid cells. BMC Mol Biol. 2014;15:1–11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Fiume G, et al. IBTK differently modulates gene expression and RNA splicing in hela and K562 cells. Int J Mol Sci. 2016;17(11):1848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Arcucci A, et al. Analysis of extracellular superoxide dismutase in fibroblasts from patients with systemic sclerosis. J Biol Regul Homeost Agents. 2011;25(4):647–54. [PubMed] [Google Scholar]
- 44.Colella M, et al. SMAD7 sustains XIAP expression and migration of colorectal carcinoma cells. Cancers. 2024;16(13):2370. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data generated or analyzed during this study are included in this published article, and its supplementary information files, and are available upon reasonable request to the corresponding authors.