Abstract
TOR (target of rapamycin) protein kinases were identified in yeasts, mammals, and Drosophila as central controllers of cell growth in response to nutrient and growth factors. Here we show that Arabidopsis thaliana possesses a single TOR gene encoding a protein able to complex with yeast 12-kDa FK506-binding protein and rapamycin despite the insensitivity of Arabidopsis vegetative growth to rapamycin. Analysis of two T-DNA insertion mutants shows that disruption of AtTOR leads to the premature arrest of endosperm and embryo development. A T-DNA-mediated translational fusion of AtTOR with the GUS reporter gene allows us to show that AtTOR is expressed in primary meristem, embryo, and endosperm, but not in differentiated cells. The implications of these features for the plant TOR pathway are discussed.
TOR (target of rapamycin) proteins are large eukaryotic proteins belonging to the family of the phosphatidylinositol 3-kinase related kinases (1). Despite having a kinase domain related to phosphatidylinositol 3-kinase, TOR proteins are thought to be protein kinases. TOR proteins are the targets of the antiproliferative drug rapamycin produced by the bacteria Streptomyces hygroscopicus (1). Their inactivation by rapamycin is mediated by formation of a ternary complex where rapamycin forms noncovalent links between the peptidyl-prolyl isomerase 12-kDa FK506-binding protein (FKBP12) and the FKBP-rapamycin-binding domain (FRB) of TOR proteins (2). In Saccharomyces cerevisiae, two TOR genes, TOR1 and TOR2, have been identified in screens for mutants resistant to rapamycin (3–5). They modulate cell growth in relation to nutrient availability and are functionally redundant for the regulation of protein synthesis necessary for the G1 to S progression in the cell cycle. This regulation occurs by activation of the eukaryotic translation initiation factor eIF4F and of RNA polymerases I and III (1). Other TOR functions were also described in the budding yeast where TOR1 and TOR2 control the regulation of nutrient metabolism by sequestering transcription factors in the cytoplasm, and TOR2 is involved in the organization of the actin cytoskeleton (1). Two TOR genes have been recently characterized in the fission yeast Schizosaccharomyces pombe, where the situation is different from S. cerevisiae, because SpTOR proteins are not redundant and are both resistant to rapamycin. SpTOR2 is essential for growth, whereas SpTOR1 is required for response to nitrogen starvation and stresses (6). A mammalian TOR, named mTOR (or FRAP/RAFT/RAPT), has also been identified (7–9) that activates protein synthesis in response to nutrient and growth factors in at least two ways: one is the activation of capped mRNA translation through deactivation of the eIF4E-binding protein 1 (4E-BP1), which sequestrates the initiation factor eIF4E; the other is the activation of translation by activation of the 40S ribosomal protein ribosomal protein S6 kinase (S6K) S6K1 (also named p70S6K) (1). In Drosophila, which possess a single TOR gene, overexpression of dS6K was found to rescue the viability of leaky TOR mutants arrested at the pupal stage (10).
Several observations suggest that nutrient perception is a major effector of TOR activation. First, Drosophila dTOR mutant cells have a phenotype similar to those that are starved (i.e., decreasing nucleolar size, fat body vesicle formation, and endoreplicative cell cycle arrest) (10). Second, in the budding yeast and in mammalian cells, rapamycin triggers autophagy, a typical starvation response (1). Finally, in mammalian cells, rapamycin blocks the amino acid-dependent phosphorylation of S6K1 and 4E-BP1 proteins, two downstream targets of TOR (1).
The phylogenetic conservation of the TOR pathway in both yeast and animals and its central role in regulating cell growth as a function of nutrient availability suggest that it may be conserved in all eukaryotes, including plants. Plant growth is a result of cell growth coupled with cell division and of cell expansion. In specialized zones, meristems, cell growth, and division occur to produce new tissues and organs. Postmitotic differentiated cells can reach large sizes, such as in the hypocotyl or in mature leaves and roots. Meristem activity and embryonic development depend on the availability of nutrients provided by other parts of the plant. There is very little knowledge of the perception and integration of nutrient supplies at the level of dividing embryonic and meristematic cells. As coordination of nutrient sensing with cell growth and division is a very basic requirement, it probably involves evolutionary conserved pathways on which new functions were added during the ecophysiological history of organisms. The specific nutritional capacities of plants, such as carbon autotrophy, make them attractive models for the comparative molecular genetics of these regulations. The TOR pathway can play an important role in the generation of the form of multicellular organisms during embryonic and adult development by relaying the perception of nutrients furnished by source tissues into growing (sink) zones. With this in mind, we undertook an analysis of the TOR pathway in Arabidopsis thaliana.
This work describes the identification, functional analysis by reverse genetics, and pattern of expression of AtTOR, the single Arabidopsis homolog of animal and yeast TORs.
Materials and Methods
Molecular Biology.
The entire AtTOR coding sequence was reverse transcribed from 1 μg of total RNAs from wild-type roots (ecotype Columbia) with reverse transcriptase from avian myeloblastosis virus (Roche Molecular Biochemicals) and primer TOR-R1 (5′-GCGGCCGCAAATGCAAATTAGTTGA-3′). The RT product was amplified by PCR (9 min of elongation) with the Expand Long Template System (Roche Molecular Biochemicals) and primers TOR R1 and TOR 7 (5′-CCTGCATCCATGGCTTCCCCTTC-3′). The PCR product was cloned into the pCR-XL-TOPO vector (Invitrogen). A cDNA clone was entirely sequenced, and discrepancies with the genomic sequence and other partially sequenced clones were corrected by replacing restriction fragments.
The 5′ end of the AtTOR transcript was defined by 5′ rapid amplification of cDNA ends (GIBCO/BRL) by using 1.9 μg of root total RNA and primer 5′-GCCGGTATTCTTAACACAGTAAGAA-3′ as the gene specific primer. A 500-bp product from a nested PCR was cloned into pGEM-T-easy vector (Promega) and sequenced.
Sequence alignments were performed with genestream software [Institut de Génétique Humaine, Toulouse, France (http://xylian.igh.cnrs.fr/bin/align-guess.cgi)]. The SwissProt accession nos. are P42345 (mTOR/FRAP), P35169 (TOR1), and P32600 (TOR2). The dTOR GenPept accession no. is AAF53237.
The TOR-FRB probe (722 bp) used for Southern blot was PCR amplified from wild-type ecotype Wassilewskija genomic DNA, with primers 5′-AGGGTTGCCATACTTTGGCATG-3′ and 5′-GGCTAGCTGTTTGTCAATCCG-3′. The β-glucuronidase (GUS) probe is a 2.4-kb XbaI-SacI fragment from the pGKB5 vector, corresponding to the right border and the GUS gene (11).
Two-Hybrid Experiments.
S. cerevisiae strain SMY87–4 (MATa trp1–901 leu2–3, 112 ura3–52 his3–200 ade2 gal4Δ gal80Δ LYS2∷GAL-HIS3 GAL2-ADE2 met2∷GAL7-lacZ fpr1∷hisG), a gift from J. Heitman, was used for two-hybrid experiments with rapamycin. This strain is a derivative of the two-hybrid host strain PJ69–4A in which the FKBP12-encoding gene was disrupted (12). SMY87–4 cells were cotransformed with three plasmids: pTR17 (URA3) (13) to express a dominant rapamycin resistant allele of the TOR2 gene (TOR2–1), the plasmid containing the GAL4 DNA-binding domain [GAL4(BD)]∷AtFRB (TRP1) fusion, and the plasmid carrying the appropriate FKBP12 fused to GAL4 activation domain [GAL4(AD)] (LEU2). After selection for the presence of the three plasmids, cotransformed yeast strains were grown overnight, resuspended in top agar (0.7% agar in water), and spread on solid medium lacking leucine, tryptophan, uracil, and adenine. Rapamycin (1 μg diluted in 10% Tween-20/90% ethanol) or dilution solution alone was deposed on Whatman paper discs, on the surface of the agar, and cells were incubated at 30°C for 5 days. To construct the plasmid expressing the GAL4(AD)∷AtFRB fusion, the AtFRB sequence (corresponding to amino acid residue 1851 to 2050 of AtTOR) was PCR amplified from expressed sequence tag W43444 with primers 5′-CCGAATTCATACATTCTAATAATCGTGCTG-3′ and 5′-CCGCGGCCGCAGGAACTGCTAGCTCCAAGTCA-3′ and inserted downstream of the GAL4(BD) in pBI880 (14). The plasmid expressing the GAL4(AD)∷ ScFKBP12 fusion was obtained by subcloning the ScFKBP12 coding sequence excised from pSBH1 (15) into pBI880 (14).
Plant Material and Growth Conditions.
The T-DNA insertion mutants are in the Wassilewskija ecotype (11, 16). Plants were grown in a phytotron at 25°C under an 8-h light/16-h dark regime (23/18°C). For GUS staining in roots, plants were grown in vitro with the same photoperiod on 2-fold diluted Hoagland solution with 0.7% agar and 50 μg/ml of kanamycin. To produce callus, lacerated leaves were transferred on solid Gamborg B5 medium supplemented with 2,4-dichlorophenoxyacetic acid (0.5 mg/liter), kinetin (0.05 mg/liter), and sucrose (20 g/liter).
Staining, Binocular Optics, and Microscopy.
For embryo observation, immature siliques were dissected under binocular optics, and seeds were cleared in Hoyer's solution (7.5 g of arabic gum/100 g of chloral hydrate/5 ml of glycerin in 30 ml of water). For high-resolution pictures, seeds were stained with the Feulgen/LR White method as described (17) and observed with a Zeiss confocal microscope (LSA510) at an exciting wavelength of 488 nm and emission with a long-pass filter of 530 nm.
GUS staining was performed as described (18) with a 4-h incubation at 37°C. For seed staining, opened young siliques were incubated 1 h in 90% acetone (−20°C) followed by two 1-h vacuum infiltration in ferri-ferrocyanide solution [4 mM K4Fe(CN)6/4 mM K3Fe(CN)6/100 mM sodium phosphate, pH 7.0] and addition of the coloration solution (4 mM 5-bromo-4-chloro-3-indolyl-β-d-glucuronic acid/10 mM EDTA/0.1% triton/100 mM sodium phosphate, pH 7.0) before a 14-h incubation at 37°C. Seeds were dissected from siliques and placed in Hoyer's solution. Observations were performed with a Leica MZ FL3 binocular for leaves, flowers, and callus, and with a Leica DMRXA microscope for roots and seeds.
Results
Molecular Characterization of AtTOR.
In a quest for plant homologs of the mammalian mTOR/FRAP and yeast TOR (TOR1 and TOR2) genes, an Arabidopsis expressed sequence tag (EST) (accession no. W43444) was identified bearing similarities with the C-terminal part of mammalian and yeast TOR proteins, including the kinase domain. Sequencing of the 2.5-kb cDNA fragment has further confirmed the close relationship of this encoded amino acid sequence with those of TOR proteins (49% similarity with amino acid residues 1,702–2,249 of mTOR and 1,659–2,474 of TOR2) and reveals the presence of a domain similar to the FRB domain, which is a hallmark of mammalian and yeast TORs. Considering this high degree of similarity with known TOR proteins, the protein corresponding to this EST was considered as an homolog of mammalian and yeast TORs and named AtTOR, for A. thaliana TOR.
The Arabidopsis TOR gene was found to map on the lower arm of chromosome 1. Most of the AtTOR genomic sequence was obtained from bacterial artificial chromosome (BAC) F20C18 (accession no. B18861). However, because F20C18 was truncated at the 3′ end of AtTOR, the remainder of the genomic sequence was obtained from DNA fragments amplified by PCR with primers derived from the expressed sequence tag cDNA sequence. The AtTOR genomic sequence was later found in BAC F2J10 (gene F2J10.9; accession no. AC015445). Southern blot hybridization (data not shown and Fig. 3A, first lane) and searches in the complete Arabidopsis genomic sequence show that AtTOR is a unique gene in this species.
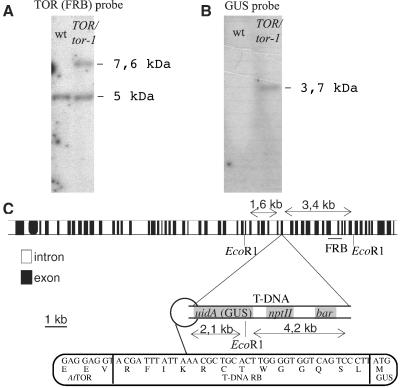
Figure 3.
The T-DNA insertion in the tor-1 mutant. (A and B) Southern blot analysis of genomic DNA from the heterozygousTOR/tor-1 line compared with wild type (wt). Five micrograms of DNA was digested with EcoRI, electrophoresed, transferred to a nylon membrane, and hybridized with a AtTOR-FRB (A) or a GUS (B) probe. (C) Representation of the T-DNA insertion site in tor-1. uidA is the sequence coding for GUS; npt II and bar are the coding sequences of the kanamycin and basta resistance genes, respectively.
Using primers derived from the genomic sequence, a partial 7.4-kb cDNA was cloned by RT-PCR. A 5′ RACE experiment allowed identification of a 230-base 5′ mRNA leader containing an upstream ORF (2 codons). The cDNA sequence was compared with the genomic sequence, and errors introduced during the RT-PCR were corrected by replacing restriction fragments by others obtained from independent PCR reactions and carrying no mutations. The final assembly was then entirely resequenced and deposed into GenBank (accession no. AF178967). The comparison of the genomic and the cDNA sequences revealed that AtTOR contain 56 exons and 55 introns, and that the AtTOR gene spans at last 17 kb of genomic DNA.
The AtTOR protein sequence deduced from the cDNA sequence contains 2,481 amino acid residues with a predicted molecular mass of 279 kDa. Alignment of AtTOR with TOR protein sequences from others organisms (Fig. 1) shows a high degree of conservation of the FRB and kinase domains as well as in a short C-terminal domain named FATC, which is conserved in the phosphatidylinositol 3-kinase related kinases FRAP, ATM, and TRRAP (1). Highly conserved stretches of amino acids are also present throughout the N-terminal two-thirds of the sequence, presumably reflecting functional or structural conservations. This part of AtTOR contains 12 motifs (20–40 residues), named HEAT repeats (found in Huntingtin, Elongation factor 3, A subunit of protein phosphatase 2A, and TOR1), which are found in all TOR proteins and have been proposed to be involved in proteins interactions (1). Protein sequence alignments also show that mTOR is the closest homolog of AtTOR (Fig. 1) and that AtTOR is closer to TOR2, the yeast TOR protein involved in cytoskeleton organization (1), than to TOR1 (Fig. 1).
Figure 1.
Comparison of the AtTOR protein sequence to the TOR protein sequences from other organisms. Each value indicates the percentage of identity with the corresponding domain sequence of AtTOR. In AtTOR, the FRB, kinase, and FATC domains correspond to residues 1930–2022, 2092–2340, and 2451–2481, respectively. At, A. thaliana; Hs, Homo sapiens; Sc, S. cerevisiae (Sc 1 for TOR1 and Sc2 for TOR2), Dm, Drosophila melanogaster. The number of amino acid residues of each protein is in brackets.
AtTOR Binds Yeast FKBP12 in the Presence of Rapamycin.
Structure determination of the human FKBP12–rapamycin–FRB complex shows that there are extensive rapamycin–protein interactions and relatively few interactions between FKBP12 and FRB (2). To confirm that the cloned cDNA was coding for a functional TOR protein, rapamycin-dependent FKBP12 binding was examined by using a yeast two-hybrid system (12, 13). SMY87–4 yeast cells containing a plasmid-borne rapamycin-resistant version of TOR1, deleted for the FPR1 gene (coding for endogenous yeast FKBP12) and coexpressing the GAL4(BD) fused to the AtTOR FRB domain (AtFRB) and the GAL4(AD) fused to yeast FKBP12 (ScFKBP12), were plated on selective media. Yeast growth on this medium depended on the expression of the GAL-ADE2 reporter gene. Yeast cells were then overlaid with small discs containing 1 μg of rapamycin or a control solution. After 5 days, colonies were readily observable around the disk containing rapamycin but not around the control disk (Fig. 2A). The two isogenic control strains coexpressing the unfused GAL4(BD) and the GAL4(AD)∷ScFKBP12 fusion or the GAL4(BD)∷AtFRB fusion and the unfused GAL4(AD) were not able to grow even around the rapamycin disk (Fig. 2 B and C), which shows that AtTOR is able to bind yeast FKBP12 but only in the presence of rapamycin.
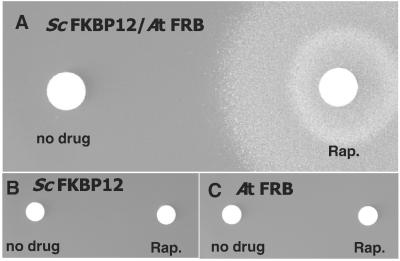
Figure 2.
Yeast two-hybrid assay showing that the AtTOR FRB domain is able to form a complex with rapamycin and ScFKBP12. (A) Two-hybrid strain SMY87–4 coexpressing the GAL4(BD)∷AtFRB and the GAL4(AD)∷ScFKBP12 fusion proteins was spread on medium lacking adenine. Formation of the FKBP12–rapamycin–FRB complex induces expression of the GAL-ADE2 reporter gene and is revealed by growth around the rapamycin (Rap.) disk (Right). (B and C) Same experiment as A, performed with control isogenic strains coexpressing the unfused GAL4(BD) and the GAL4(AD)∷ScFKBP12 fusion protein (B) or the GAL4(BD)∷AtFRB fusion protein and the unfused GAL4(AD) (C).
Identification of Two AtTOR Knockout Mutants.
To investigate AtTOR functions, a search for knockout mutants in the Institut National de la Recherche Agronomique T-DNA insertion library was performed (11, 16). One allele (tor-1) was identified in a screen based on the expression of the GUS reporter gene in the root (18). Southern analysis shows that tor-1 contains a single T-DNA insertion (Fig. 3 A and B). In this allele, the GUS reporter gene of the T-DNA is translationally fused to the TOR gene, leading to the creation of the translational fusion of the GUS protein and amino acid 1555 of AtTOR (Fig. 3C). A second TOR mutant (tor-2) was found in a PCR screen of the same collection. After three backcrosses to discard unlinked T-DNA insertions, a complex pattern of inserted T-DNA was found to be genetically linked to the AtTOR locus (data not shown). In both alleles, the T-DNA is inserted upstream of the FRB domain of AtTOR. In the progeny of TOR/tor-1 and TOR/tor-2 plants, a 2:1 ratio of segregation for kanamycin resistance (a marker carried by the T-DNA) was observed [respectively 420/226; χ2(2/1) = 0,72; P > 0.1 and 544/304; χ2(2/1) = 2,22; P > 0,05]. This segregation suggests a default of transmission of the tor alleles to the next generation (see below). Further analysis showed that, in both lines, one-quarter of the seeds from a heterozygous silique had aborted [aborted/normal seeds ratios of 186/594 into TOR/tor-1 siliques and 82/271 into TOR/tor-2 siliques lead to respective χ2(1/3) of 0.49 and 0.58 with P > 0.1]. The embryo-lethal seeds are randomly distributed along the silique (Fig. 4A). The absence of segregation distortion shows that tor alleles are equally well transmitted by male and female gametes. Heterozygous plants display no detectable phenotype.
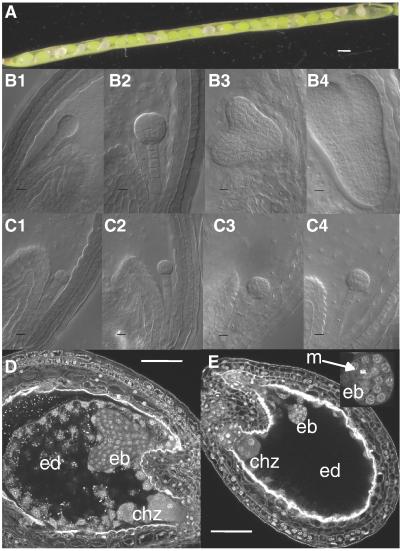
Figure 4.
Phenotype of the TOR/tor-1 mutant. (A) Heterozygous siliques of the TOR/tor-1 mutant before desiccation. Normal seeds are green, whereas aborted seeds are white. (B and C) Embryos from tor-1 heterozygous siliques observed with Nomarski optics: (B) normal embryo at the dermatogen (1 and 2), heart (3), and torpedo (4) stage; (C1–4) aborted mutant embryo from the same siliques as B1–4), respectively. (D and E) Normal (D) and aborted (E, with an enlargement of the embryo, Top Right) seeds from the same tor-1 silique observed by confocal microscopy after Feulgen staining. [Bar = 200 μm (A), 10 μdm (B and C), and 50 μm (D and E).] ed, endosperm; eb, embryo; chz, chalazal endosperm; m, metaphase.
To test whether these two mutant alleles belong to the same complementation group, emasculated TOR/tor2 flowers were pollinated with pollen from TOR/tor1 plants and GUS expression from the tor1 allele (which is expressed in embryos; see below) was used as a marker of crosspollinization. GUS-stained aborted embryos were found in the siliques resulting from these crosses, demonstrating that the two mutant alleles are not able to complement each other. It was therefore concluded that the embryo-lethal phenotype in these two lines was caused by the knockout of AtTOR.
Disruption of AtTOR Leads to the Premature Arrest of Endosperm and Embryo Development.
Closer observation of the immature aborted seeds revealed that both tor-1/tor-1 and tor-2/tor-2 embryos are arrested at the dermatogen stage (Fig. 4 B–E), still containing cells in metaphase (Fig. 4E). This observation shows that division itself is not inhibited as a consequence of AtTOR disruption.
In higher plants, the products of the double fertilization process are the embryo and the endosperm, a triploid tissue providing nutrients for embryo development and seed germination (19). During embryogenesis, the endosperm initially grows as a syncytium until it reaches around 200 nuclei, and then cellularization occurs. In the tor mutant endosperm, the number of nuclei was only 48 ± 13, and cellularization did not occur (compare Fig. 4 E with D). For comparison, in a wild-type seed, an endosperm with 44–48 nuclei corresponds to an embryo at the two-to-four-cells stage (20). This observation shows that endosperm is arrested at an earlier developmental stage than the embryo in tor seeds.
AtTOR Is Expressed in Embryos, Endosperm, and Primary Meristems.
TOR/tor-1 plants were used to monitor the pattern of AtTOR expression through different developmental stages. In these plants, GUS expression is driven by the AtTOR regulatory elements in their native genomic environment. One day after fertilization, GUS staining was detected in the endosperm, the embryo, and the chalazal proliferating tissue (a maternal tissue) (Fig. 5A). After the early globular stage, GUS staining is no longer detected in the endosperm but persists in the embryo (excluding the suspensor) up to the heart and torpedo stages (Fig. 5 B and C). This pattern correlates with a proliferation of nuclei in the endosperm that precedes active proliferation of embryonic cells. In the mature embryo, the apical and primary root meristems were stained as well as the dividing vascular tissues (Fig. 5D). In both seedling and adult plant, AtTOR is highly expressed in root meristems (Fig. 5 E–G) in the shoot apical meristem (Fig. 5I) and in floral buds (Fig. 5J). During lateral root formation, the expression is detected as soon as the first periclinal divisions of the lateral root primordia (Fig. 4E) and remains active in all cells during the formation of the emerging secondary root meristem (Fig. 4F). GUS staining was not detected in the pollen, stem, and non-meristematic cells of hypocotyl, roots, and leaves, including dividing stomatal precursors (Fig. 4H and data not shown). To further analyze the link between AtTOR expression and cell proliferation, callus formation was induced on explants of the heterozygous TOR/tor-1 line by hormonal treatment. Only the proliferating cells, at the surface of the callus, were found to express GUS activity (Fig. 4K). However, GUS staining was not found in dividing stomatal precursor cells of mature leaves and in secondary meristems, showing that AtTOR expression is not strictly correlated with cell division. In summary, AtTOR expression is detected in developing endosperm and embryo and in all of the primary meristems of the plant. Studies of the Arabidopsis leaf by using a Cdc2∷GUS reporter gene show that patterns of cell division are randomly distributed in a defined region at the base of the developing leaf (21). This region, which also contains many nondividing cells is almost superimposable on that of AtTOR expression (Fig. 4I).
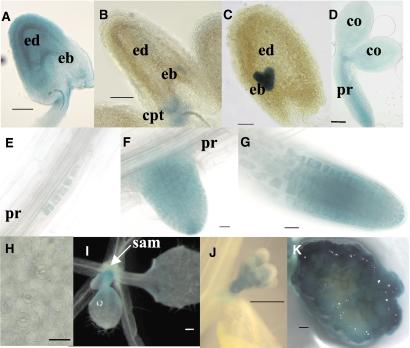
Figure 5.
AtTOR is expressed in endosperm, embryo, and primary meristems. Localization of the AtTOR∷GUS fusion protein in the TOR/tor-1 mutant. (A–C) Seeds containing a quadrant (A) to heart (C) stage embryo. (D) Mature embryo. (E–G) Lateral root development at the first layers (E), emergence (F), and elongation (G) stages. (H) Zoom of a leaf showing different stages of stomatal development. (I) Leaves primordium. (J) Floral buds. (K) One-month-old callus transversally cut before coloration. [Bar = 50 μm (A–D), 20 μm (E–H), and 500 μm (I–K)]. ed, endosperm; eb, embryo; cpt, chalazal proliferating tissue; pr, primary root; sam, shoot apical meristem.
Discussion
Features of AtTOR.
Our results show that the Arabidopsis TOR gene has features in common with other TOR genes. Domains known to be important for TOR function are conserved in the AtTOR protein, including the C-terminal region containing the kinase domain, the FRB domain, and the short C-terminal sequence. AtTOR appears to be a single gene in the Arabidopsis genome, a situation similar to that of animal genomes and in contrast with yeast in which TOR duplication has occurred (5, 6, 10). It remains to be determined whether the single animal and plant TOR genes can fulfill all of the functions of yeasts TOR1 and TOR2. Sequence comparisons show that the TOR2 protein of S. cerevisiae is most similar to plant and animal TOR proteins, suggesting that TOR1 appeared more recently in a common ancestor of yeasts.
The two-hybrid experiments have shown that the AtTOR FRB domain is able to form a complex with rapamycin and S. cerevisiae FKBP12. Formation of the FKBP12–rapamycin–FRB complex is responsible of the antiproliferative effect of rapamycin through inhibition of TOR proteins in a wide range of organisms and cells, including Drosophila, mammalian cells, S. cerevisiae, and Cryptococcus neoformans (1, 10, 13). In contrast, the vegetative growth of Arabidopsis was found to be insensitive to this drug even at concentrations up to 10 μM, which is 100 times the concentration inhibiting yeast growth (data not shown). Interestingly, rapamycin inhibits the growth of Chlamydomonas reinhardtii, a unicellular alga, but not of the bryophyte Physcomitrella patens, the monocot Oryza sativa, and the dicots Nicotiana tabacum and Brassica napus (Fabien Nogue, personal communication, and data not shown), suggesting that resistance to this drug may be a common feature of land plants. This resistance to rapamycin could be caused by the impossibility of the drug entering plant cells, but that is unlikely, because rapamycin is able to enter C. reinhardtii cells. A second explanation could be a default to form the FKBP12–rapamycin–FRB complex. This would likely involve the plant FKBP12 rather than the AtTOR-FRB. Indeed, the AtTOR-FRB is able to complex with rapamycin and ScFKBP12, whereas the FKBP12 from Vicia faba (broad beans) does not restore the sensitivity of the S. cerevisiae FKBP12 mutant to rapamycin (22). In addition, we were unable to detect complex formation between AtFKBP12, AtFRB, and rapamycin in the two-hybrid system (not shown). A third hypothesis may be the incapacity of the FKBP12–rapamycin–AtTOR complex to inhibit At TOR functions, as proposed for the resistance to rapamycin of S. pombe TOR proteins (6).
AtTOR Is Essential for Early Development.
Disruption of the TOR gene leads to embryonic arrest, indicating a crucial role of TOR during early development in both Drosophila and Arabidopsis (refs. 10 and 23; this study). Embryos homozygous for the AtTOR knockout mutations cannot develop past the dermatogen stage with cells still undergoing divisions. This result shows that AtTOR disruption is not inhibiting cell division itself. This phenotype is similar to that of growth-arrested Drosophila larvae, as a consequence of protein synthesis defects, which can resume the cell cycle on artificial stimulation (24). Indeed, the first cell divisions in the Arabidopsis embryo require little or no increase in overall cell mass and lead to a reduction of the amount of cytoplasm relative to the nucleus (25), supporting the idea that growth arrest occurs in tor−/− cells as the consequence of an incapacity to synthesize cell components during the transition between the two modes of cell division. Further support for this model is given by the genetic evidence that the haploid cell divisions occur correctly during the maturation of tor gametes. Cytological observations show that plant postmeiotic asymmetric cell divisions of the male gamete occur without net increase in the mass of the cytosol (26). In addition, growth of the pollen tube is polarized-tip growth involving essentially vacuole expansion and cell-wall synthesis (27). Similarly, in the female gamete, cytosolic growth occurs premeiotically, and the vacuole contributes most of the size of the embryo sac (28). A confirmation of this model would, however, require methods allowing a precise quantification of cytosolic mass and composition and metabolic activity in single cells.
Differentiated Plant Cells Do Not Express AtTOR.
AtTOR expression patterns are limited to the zones where cell proliferation is coupled to cytosolic growth, such as primary apical and root meristems. This pattern is in contrast with TOR expression in mammalian cells (8, 9) and in Drosophila larvae (E. Hafen, personal communication), which occurs in all tissues. The growth process of mature plant cells that do not express AtTOR (expanding root, leaf, and hypocotyl cells) is because of vacuole and cell-wall expansion and is not related to the growth process attributed to TOR in other organisms, which involves the synthesis of cytosolic components (1). However, given the central role of TOR proteins in relaying nutrient perception, the absence of AtTOR expression in differentiated cells also raises the question of how these cells perceive nutrient starvation. One hypothesis is that differentiated plant cells directly sense nitrogen and carbon at the level of the nitrate assimilation and carbon fixation pathways upstream of the amino acid perception, which was demonstrated to regulate TOR (1). Finally, the lack of AtTOR expression in stomatal precursors in which predivision growth occurs through vacuolar expansion (29) further supports the absence of an obligatory link between cell division and AtTOR activity, as discussed in the preceding section.
What Are the Components of the Plant TOR Pathway?
Known targets of mTOR in animal cells are the growth-regulating kinase S6K1 and the small 4E-BP1 translational regulators (1). Arabidopsis possesses two homologs of S6K1 but they do not have the N-terminal domain of mammalian S6K1, which is required for rapamycin sensitivity, and when expressed in mammalians cells, AtS6k2 is insensitive to rapamycin (30). Concerning 4E-BPs, no homolog can be found in the Arabidopsis genome. However, although no structural homolog of 4E-BPs exists in S. cerevisiae, a functional eIF4E-associated protein (Eap1p), containing the consensus eIF4E-binding domain and able to compete with eIF4G for binding to eIF4E, has been found to be related to the TOR pathway (31). The relation with TOR of potential eIF4E-associated proteins in plants, such as the Arabidopsis lipoxygenase 2 (32), remains, therefore, to be determined. The cyclin-dependent kinase inhibitor p27Kip1 and the α4 phosphatase, two others proteins that are probably downstream of mTOR (1), also have homologs in Arabidopsis (33, 34). Integration of cellular TOR activity at the level of the whole organism occurs through insulin and insulin-like receptors in animals (1). However, because no equivalents of insulin and insulin-like receptors have been found in Arabidopsis, the question of the signals regulating AtTOR is raised. Obvious candidates are phytohormones, particularly auxins and cytokinins, which are known to affect growth and cell proliferation (35). Answers to this question would support the theory that intracellular pathways remain quite well conserved during evolution and have been recruited by different extracellular systemic signaling systems, causing major evolutionary changes (36).
Acknowledgments
We thank Joseph Heitman (Duke University, Durham, NC) and Jean-Denis Faure [Institut National de la Recherche Agronomique (INRA), Versailles] for the gift of yeast strain and plasmids, Fabien Nogué (INRA) for P. patens experiments with rapamycin, Laurent Cournac [DEVM-Commissariat à l'Energie Atomique (CEA), Cadarache, France] for the gift of C. reinhardtii cultures, Fabienne Granier (INRA) for help in screening T-DNA lines, Marcel Salanoubat (Genoscope, Evry, France) for help in sequencing, and Ernst Hafen (Zoologisches Institut, Zürich) for providing unpublished results. B.M. is supported by an INRA-CEA doctoral fellowship.
Abbreviations
- TOR
target of rapamycin
- FRB
FKBP-rapamycin-binding domain of TOR
- FKBP12
12-kDa FK506-binding protein
- S6K
ribosomal protein S6 kinase
- eIF
eukaryotic initiation factor
- 4E-BP1
eIF4E-binding protein 1
- GUS
β-glucuronidase
- RT
reverse transcription
- GAL4(BD)
GAL4 DNA-binding domain
- GAL4(AD)
GAL4 activation domain
Footnotes
Data deposition: The sequence reported in this paper has been deposited in the GenBank database (accession no. AF178967).
References
- 1.Schmelzle T, Hall M N. Cell. 2000;103:253–262. doi: 10.1016/s0092-8674(00)00117-3. [DOI] [PubMed] [Google Scholar]
- 2.Choi J, Chen J, Schreiber S L, Clardy J. Science. 1996;273:239–242. doi: 10.1126/science.273.5272.239. [DOI] [PubMed] [Google Scholar]
- 3.Heitman J, Movva N R, Hall M N. Science. 1991;253:905–909. doi: 10.1126/science.1715094. [DOI] [PubMed] [Google Scholar]
- 4.Kunz J, Henriquez R, Schneider U, Deuter-Reinhard M, Movva N R, Hall M N. Cell. 1993;73:585–596. doi: 10.1016/0092-8674(93)90144-f. [DOI] [PubMed] [Google Scholar]
- 5.Helliwell S, Wagner P, Kunz J, Deuter-Reinhard M, Henriquez R, Hall M N. Mol Biol Cell. 1994;5:105–118. doi: 10.1091/mbc.5.1.105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Weisman R, Choder M. J Biol Chem. 2001;276:7027–7032. doi: 10.1074/jbc.M010446200. [DOI] [PubMed] [Google Scholar]
- 7.Sabatini D M, Erdjument-Bromage H, Lui M, Tempst P, Snyder S H. Cell. 1994;78:35–43. doi: 10.1016/0092-8674(94)90570-3. [DOI] [PubMed] [Google Scholar]
- 8.Brown E L, Albers M W, Shin T B, Ichikawa K, Keith C T, Lane W S, Schreiber S L. Nature (London) 1994;369:756–758. doi: 10.1038/369756a0. [DOI] [PubMed] [Google Scholar]
- 9.Chiu M I, Katz H, Berlin V. Proc Natl Acad Sci USA. 1994;91:12574–12578. doi: 10.1073/pnas.91.26.12574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Zhang H, Stallock J P, Ng J C, Reinhard C, Neufeld T P. Genes Dev. 2000;14:2712–2724. doi: 10.1101/gad.835000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bouchez D, Camilleri C, Caboche M. C R Acad Sci Paris. 1993;316:1188–1193. [Google Scholar]
- 12.Arndt C, Cruz M C, Cardenas M E, Heitman J. Microbiology. 1999;145:1989–2000. doi: 10.1099/13500872-145-8-1989. [DOI] [PubMed] [Google Scholar]
- 13.Cruz M C, Cavallo L M, Gorlach J M, Cox G, Perfect J R, Cardenas M E, Heitman J. Mol Cell Biol. 1999;19:4101–4112. doi: 10.1128/mcb.19.6.4101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kohalmi S E, Nowak J, Crosby W L. In: Differentially Expressed Genes in Plants: A Bench Manual. Hansen E, Harper G, editors. London: Taylor & Francis; 1997. pp. 63–82. [Google Scholar]
- 15.Cardenas M E, Hemenway C, Muir R S, Ye R, Fiorentino D, Heitman J. EMBO J. 1994;13:5944–5957. doi: 10.1002/j.1460-2075.1994.tb06940.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Bechtold N, Ellis J, Pelletier G. C R Acad Sci Paris. 1993;316:1194–1199. [Google Scholar]
- 17.Braselton J P, Wilkinson M J, Clulow S A. Biotech Histochem. 1996;71:84–87. doi: 10.3109/10520299609117139. [DOI] [PubMed] [Google Scholar]
- 18.Sarrobert C, Thibaud M-C, Contard-David P, Gineste S, Bechtold N, Robaglia C, Nussaume L. Plant J. 2000;24:357–368. doi: 10.1046/j.1365-313x.2000.00884.x. [DOI] [PubMed] [Google Scholar]
- 19.Berger F. Curr Opin Plant Biol. 1999;2:28–32. doi: 10.1016/s1369-5266(99)80006-5. [DOI] [PubMed] [Google Scholar]
- 20.Boisnard-Lorig C, Colon-Carmona A, Bauch M, Hodge S, Doerner P, Bancharel E, Dumas C, Haseloff J, Berger F. Plant Cell. 2001;13:495–509. doi: 10.1105/tpc.13.3.495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Donnelly P M, Bonetta D, Tsukaya H, Dengler R E, Dengler N G. Dev Biol. 1999;215:407–419. doi: 10.1006/dbio.1999.9443. [DOI] [PubMed] [Google Scholar]
- 22.Xu Q, Liang S, Kudla J, Luan S. Plant J. 1998;15:511–519. doi: 10.1046/j.1365-313x.1998.00232.x. [DOI] [PubMed] [Google Scholar]
- 23.Oldham S, Montagne J, Radimerski T, Thomas G, Hafen E. Genes Dev. 2000;14:2689–2694. doi: 10.1101/gad.845700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Galloni M, Edgar B A. Development (Cambridge, UK) 1999;126:2365–2375. doi: 10.1242/dev.126.11.2365. [DOI] [PubMed] [Google Scholar]
- 25.Mansfield S G, Briarty L G. Can J Bot. 1991;69:461–476. [Google Scholar]
- 26.Owen H A, Makaroff C A. Protoplasma. 1995;185:7–21. [Google Scholar]
- 27.Hepler P K, Vidali L, Cheung A Y. Annu Rev Cell Dev Biol. 2001;17:159–187. doi: 10.1146/annurev.cellbio.17.1.159. [DOI] [PubMed] [Google Scholar]
- 28.Mansfield S G, Briarty L G, Erni S. Can J Bot. 1991;69:447–460. [Google Scholar]
- 29.Zhao L, Sack F D. Am J Bot. 1999;86:929–939. [PubMed] [Google Scholar]
- 30.Turck F, Kozma S C, Thomas G, Nagy F. Mol Cell Biol. 1998;18:2038–2044. doi: 10.1128/mcb.18.4.2038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Cosentino G P, Schmelzle T, Haghighat A, Helliwell S B, Hall M N, Sonenberg N. Mol Cell Biol. 2000;20:4604–4613. doi: 10.1128/mcb.20.13.4604-4613.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Freire M A, Tourneur C, Granier F, Camonis J, El Amrani A, Browning K S, Robaglia C. Plant Mol Biol. 2000;44:129–140. doi: 10.1023/a:1006494628892. [DOI] [PubMed] [Google Scholar]
- 33.De Veylder L, Beeckman T, Beemster G T, Krols L, Terras, Landrieu I, Van Der Schueren E, Maes S, Naudts M, Inze D. Plant Cell. 2001;13:1653–1668. doi: 10.1105/TPC.010087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Harris D M, Myrick T L, Rundle S J. Plant Physiol. 1999;121:609–617. doi: 10.1104/pp.121.2.609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Stals H, Inzé D. Trends Plant Sci. 2001;6:359–364. doi: 10.1016/s1360-1385(01)02016-7. [DOI] [PubMed] [Google Scholar]
- 36.Duboule D, Wilkins A S. Trends Genet. 1998;14:54–59. doi: 10.1016/s0168-9525(97)01358-9. [DOI] [PubMed] [Google Scholar]