Abstract
Herbs have historically been central to medicinal practices, representing one of the earliest forms of therapeutic intervention. While synthetic drugs are often highly effective in treating acute conditions, their use is frequently accompanied by adverse side effects. In addition, the growing dependence on synthetic pharmaceuticals has raised concerns regarding affordability, thereby fostering a renewed interest in herbal medicine as a cost-effective and holistic alternative. In response to this need, the current study introduces a computer vision framework for accurate herb identification. A novel dataset, Herbify, was compiled from two different herb datasets and refined through rigorous cleaning, preprocessing, and quality control procedures. The resulting dataset underwent standardization via the Preprocessing Algorithm for Herb Detection (PAHD), producing a refined dataset of 6104 images, representing 91 distinct herb species, with an average of about 67 images per species. Utilizing transfer learning, the research harnessed pre-trained Convolutional Neural Networks (CNNs) and Vision Transformers (ViTs), then integrated these models into an ensemble framework that leverages the unique strengths of each architecture. Experimental results indicate that EfficientNet v2-Large achieved a noteworthy F₁-score of 99.13%, while the ensemble of EfficientNet v2-Large and ViT-Large/16, termed EfficientL-ViTL, attained an even higher F₁-score of 99.56%. Additionally, the research also introduces ‘Herbify’ application, an AI-driven framework designed to identify herbs using the developed model. By directly tackling the principal obstacles in herb identification, the proposed system achieves a highly accurate and operationally viable classification mechanism. The experimental outcomes showcase top-tier performance in herb identification and emphasize the transformative potential of AI-based solutions in supporting botanical applications.
Supplementary Information
The online version contains supplementary material available at 10.1186/s13007-025-01421-5.
Keywords: Medicinal plants, Herb classification, Convolutional neural networks, Vision transformers, Ensemble, Transfer learning, Deep Learning
Introduction
Herbs have historically been valued for their medicinal, nutritional, and cultural significance across various regions [1]. The global herbal medicinal market is substantial and continues to expand, driven by rising consumer interest in natural healthcare approaches [2]. Valued at USD 216.40 billion in 2023, this market is projected to reach USD 5 trillion by 2050 by the World Health Organization (WHO) [3]. According to a World Health Organization report, nearly 80% of the global population primarily relies on plant-based and traditional herbal remedies for their healthcare needs, underscoring their widespread application around the world [4].
Natural, plant-based medicines are gaining ground over the synthetic drugs. Global emphasis over sustainable practices is fostering efforts to explore herbal medicine as a promising alternative to synthetic pharmaceuticals [2, 5]. Although synthetic drugs are highly effective for acute conditions, they often carry adverse side effects, including kidney and liver damage. Furthermore, widespread reliance on synthetic drugs raises concerns regarding affordability, thereby positioning herbal medicines as a more cost-effective alternative [6, 7]. For instance, prolonged use of certain non-steroidal anti-inflammatory drugs (NSAIDs) can increase the risk of ulcers and kidney damage [8].
This growing emphasis on herbal medicine is largely attributable to the minimal side effects and non-toxic attributes of many naturally occurring medicinal plants [9–11]. For example, recent research has confirmed that curcumin, a key compound found in turmeric, exhibits anti-cancer properties, underscoring its significance in the healthcare sector [12].
By 2023, smartphones have become indispensable to everyday life, with over 6.92 billion users globally, roughly 86.29% of the world’s population [13], generating substantial opportunities for innovation, particularly in plant identification. Utilizing smartphones equipped with high-resolution cameras, powerful processors, and internet connectivity, to identify medicinal plants can offer significant benefits, especially in regions where access to expert knowledge or comprehensive reference materials is limited, opening the gateways for real-time applications such as herb identification [11]. Swift advancements in computer vision, especially through models such as convolutional neural networks (CNNs) and vision transformers (ViTs), have significantly enhanced the precision and efficiency of plant recognition systems. These models can be trained on large, diverse datasets to identify a wide array of species, including medicinal plants, even under varying environmental conditions.
Convolutional neural networks excel at extracting local features such as edges, textures, and shapes through their convolutional layers, while pooling layers provide translation invariance, thereby enhancing spatial understanding. Vision transformers, by contrast, adeptly capture global context using self-attention mechanisms to analyze long-range dependencies within an image [14]. Unlike CNNs, which rely on hierarchical feature maps, ViTs segment an image into patches and treat each patch as an input token, like words in natural language processing, thus offering a flexible method of representing visual data [15]. Hybrid models exploit the complementary strengths of these architectures, for instance, by using CNNs to extract local features and ViTs for broader contextual reasoning. In some designs, both architectures process the image concurrently, with their outputs fused for superior performance. This combination enables robust solutions in tasks such as image classification, object detection, and medical imaging. By integrating CNNs’ local feature extraction with ViTs’ global comprehension, hybrid models achieve state-of-the-art performance across diverse and challenging visual domains.
Many existing research efforts in medicinal plant identification are limited by the small scale of the datasets on which their models are trained. For example, the DeepHerb dataset—developed to address data scarcity—originally comprised only 2515 images, gathered primarily through scanners and mobile phones [10]. While some studies have employed CNNs to capture local features or ViTs to grasp global context, these models often fall short by using only one of the two techniques, thereby limiting classification robustness. Even when hybrid models have been attempted, ensemble learning strategies were frequently overlooked, leading to suboptimal performance. Moreover, real-time application of these models remains largely unexplored, with few, if any, studies offering a deployable interface for practical use.
To overcome these limitations and produce a more comprehensive, uniform, and high-quality resource, this study introduces a novel dataset named ‘Herbify.’ This dataset is formed by merging two distinct repositories—DIMPSAR [9] and DeepHerb [10], resulting in a total of 6104 images spanning 91 unique classes. This research also presents a hybrid ensemble model that combines the local feature extraction capabilities of CNNs with the global context awareness provided by ViTs, thereby enhancing overall classification accuracy.
This study aims to address practical constraints in real-world scenarios and existing research works by employing two core strategies. First, the newly consolidated ‘Herbify’ dataset offers extensive coverage of various medicinal plants, thereby mitigating issues related to limited data. The comprehensive preprocessing module PAHD has been developed to standardize the dataset and improve image quality, with a primary focus on minimizing background variance and noise, improving the overall results accuracy. Among the techniques used, data augmentation further enhances both the volume and diversity of the dataset. Secondly, the ensemble deep learning framework that capitalizes on CNNs for local feature extraction and ViTs for global context allows for precise herb identification. Finally, to ensure real-time usability, the study translates this framework into a fully functional web-based application that not only identifies medicinal plants but also provides the scientific name and a resemblance probability. This user-friendly interface paves the way for seamless, real-time herb identification, thereby extending the benefits of the research to a broader audience.
Literature review
In recent years, the detection and classification of herbs using advanced image-based techniques have gained significant attention for promoting sustainable agriculture and safeguarding global health. Integrating sophisticated image processing methods with deep learning architectures has shown considerable success in accurately identifying and categorizing herbs.
Murugan Prasathkumar et al. (2021) spotlight various Indian medicinal plants, discussing their phytocompounds and pharmacological properties, such as antimicrobial, antioxidant, antidiabetic, and anticancer effects, across 14 families. Among these, Senna auriculata is noted for its wide range of medicinal benefits, while B. asiatica, D. metal, and A. marmelos also exhibit notable therapeutic value. The authors investigate 20 Indian medicinal plants used in traditional polyherbal formulations within Ayurveda, Unani, and Siddha, thereby providing valuable insights for the advancement of ethno-medicine [16]. Subsequently, Pushpa et al. (2023) introduced the DIMPSAR dataset [17], comprising 5900 images of 40 plant species and 6900 single-leaf images spanning 80 species, all collected under real-time Indian conditions [9]. Leveraging this dataset, they developed Ayur-PlantNet, a lightweight CNN that achieves 92.27% accuracy rate in Ayurvedic plant identification [9, 18]. In a related effort, Roopashree et al. (2021) curated DeepHerb, a dataset with 2515 images covering 40 herb species, enabling a vision-based identification system that attained 97.5% accuracy. Their “HerbSnap” app demonstrates practical use, although DeepHerb faces challenges when dealing with compound leaves [10].
The study by B.R. Pushpa et al. (2024) presents a hierarchical classification model for 100 Indian medicinal plants, integrating convolutional and conventional features in conjunction with Random Forest classifiers. When evaluated on both a self-built leaf dataset (GSL100) and real-time datasets (RTL80 and RTP40), the model achieved a 94.54% accuracy on GSL100 and 75.46% on the real-time datasets, surpassing existing methods. This improvement underscores the model’s robustness and efficiency under real-world conditions [19]. Meanwhile, Chin Poo Lee et al. (2023) proposed a Plant-CNN-ViT ensemble that fuses Vision Transformer, ResNet-50, DenseNet-201, and Xception to address plant classification in data-scarce settings, achieving near-perfect accuracy on four plant leaf datasets: 100.00% on both the Flavia and Folio Leaf datasets, 100.00% on the Swedish Leaf dataset, and 99.83% on the MalayaKew Leaf dataset. Despite its higher complexity, the model effectively captures spatial details and mitigates overfitting, indicating its potential for broader applications, such as plant disease detection [20]. Mirzapour et al. (2023) proposed the Controllable Ensemble Transformer and CNN (CETC) model, which integrates CNNs and transformers to capture both local and global features for medical image classification. The architecture combines convolutional encoders, transposed-convolutional decoders, and a transformer-based classifier, achieving superior performance on two publicly available COVID-19 datasets [21]. Khan et al. (2025) proposed an optimized ensemble model for diabetic retinopathy detection, integrating advanced preprocessing (CLAHE, Gamma correction, and DWT) with three pre-trained CNNs (DenseNet169, MobileNetV1, Xception). The model uses the Salp Swarm Algorithm to dynamically assign ensemble weights, achieving 89.07% accuracy on the APTOS 2019 dataset and demonstrating strong performance across multiple evaluation metrics [22]. Hekmat et al. (2025) proposed a DE-optimized ensemble combining MobileNetV1, MobileNetV2, and ResNet50V2, achieving up to 98% accuracy in brain tumor detection, with improved interpretability using Grad-CAM [23]. Nanni et al. (2023) proposed ensembles of diverse CNN and transformer models optimized with novel Adam-based variants, demonstrating superior performance across multiple benchmarks and highlighting the benefits of combining varied architectures and optimizers for image classification [24]. Abulfaraj et al. (2025) proposed a multi-label image classification ensemble integrating a Vision Transformer (ViT) with enhanced MobileNetV2 and DenseNet201 models using parallel convolutional layers and a voting mechanism. The model achieved accuracies of 98.24% on VOC 2007, 98.89% on VOC 2012, 99.91% on MS-COCO, and 96.69% on NUS-WIDE 318. Their approach demonstrated superior performance over individual models, effectively combining global and local feature learning [25]. Amin et al. (2023) introduced a Deep Learning Based Active Learning (DLBAL) framework combining EfficientNet-B0 with an enhanced sample selection strategy to reduce manual annotation in image classification. Unlike conventional uncertainty-based methods, DLBAL incorporates high-confidence samples to improve learning efficiency. Experimental results on CACD and Caltech-256 show that the method outperforms existing approaches in both accuracy and annotation cost reduction [26]. In another research, Amin et al. (2022) proposed EADN, a lightweight deep learning model for anomaly detection in surveillance videos, combining shot segmentation, time-distributed CNN layers, and LSTM for spatiotemporal feature learning. The model achieved AUCs of 93% on UCSDped1, 97% on UCSDped2 and CUHK Avenue, and 98% on UCF-Crime, outperforming state-of-the-art methods while maintaining low computational cost and false alarm rates [27]. Similarly contributing to the world of research, Amin et al. (2023) proposed ADSV, an attention-based deep learning model combining LWCNN, LSTM, and attention mechanisms for anomaly detection in surveillance videos. Using shot segmentation and time-distributed layers, the model processes chronologically ordered frames to extract spatiotemporal features. Evaluated on CUHK-Avenue and UCF-Crime datasets, ADSV achieved accuracy improvements of 12.9 and 14.88% over existing methods, with reduced false alarms and a compact model size of 54.1 MB. The approach demonstrates strong generalization while maintaining computational efficiency [28]. In another study, N. Rohith et al. (2023) enhanced VGG-19-BN and ResNet101 by incorporating advanced attention modules like CBAM and ECANet, thereby allowing the networks to focus on critical regions of medicinal plant images. Employing the DIMPSAR dataset [9, 17], the authors curated a final dataset of 146 samples per class across 40 classes, demonstrating the benefits of attention mechanisms in improving classification performance [5].
Several hybrid models have also been introduced to bolster medicinal plant classification. Tiwari et al. (2024) developed MedLeafNet, a CNN-ViT architecture trained on a subset of the PlantVillage dataset containing 38 plant disease classes and 54,272 images. By concentrating on 20 targeted classes, they addressed dataset imbalance and achieved 95–96% accuracy, surpassing conventional techniques. MedLeafNet employs contrast boosting, sharpening, and color-based segmentation for preprocessing and leverages data resampling and augmentation to enhance generalization. This system is deployed as a web-based platform offering real-time disease identification and prevention recommendations [29]. Khan et al. (2025) proposed a hybrid ensemble model combining Vision Transformer (ViT-L16) with CNNs (ResNet50, EfficientNetB1, and a custom ProDense block) for breast cancer detection from mammograms. The model achieved 98.08% accuracy on the INbreast dataset, outperforming existing methods by leveraging both transformer-based global feature extraction and deep CNN features [30]. Similarly, Hajam et al. (2023) applied VGG16, VGG-19, and DenseNet201 with ensemble learning to classify leaves from 30 medicinal plant classes, comprising a total of 1835 images. Their best-performing ensemble—VGG-19 and DenseNet201, achieved a 99.12% test accuracy, underlining ensemble learning’s efficacy in reducing dependence on manual expertise for medicinal plant identification [31]. In another effort, Kunjachan et al. (2024) employed a CNN with four convolutional layers, pooling operations, a fully connected layer, and a soft-max classifier for herb recognition. Using the ReLU activation function and Adam optimizer with a time-based decay learning rate, the model achieved a 96.25% accuracy, affirming the utility of deep learning, preprocessing, and augmentation for medicinal plant applications [32]. Beyond pure image analysis, Liu et al. (2022) introduced TCMBERT, a two-stage transfer learning framework for traditional Chinese medicine (TCM) prescription generation from limited clinical records. This pioneering method effectively handles sequence generation tasks, outperforming existing models in both qualitative and quantitative evaluations [33]. Lastly, Mujahid et al. (2024) achieved a 92.66% accuracy in classifying Indonesian herbal leaves using CNNs on a dataset gathered via Bing Downloader Scraping, demonstrating low loss values and reliable feature extraction for herbal medicine research and biodiversity protection [34].
In our earlier work [35], we introduced an integrated framework that combines convolutional neural networks and vision transformers for precise soil‐type classification, alongside a fuzzy-logic engine to generate crop‐recommendation profiles. These methods were encapsulated in a mobile application capable of both rapid soil analysis and tailored crop‐suitability guidance. In this study, we build upon that foundation by adapting and extending our computer-vision and decision-support pipeline to the automated identification of medicinal herbs.
Despite notable progress in automated herb recognition, current research is still hampered by several persistent limitations. First, there is an acute shortage of well-curated, herb-specific image corpora. The few public collections that do exist are often small, imbalanced, and contain mislabeled or poor-quality images, all of which undermine the training of data-hungry deep networks. Second, most studies adopt either convolutional neural networks to exploit local, fine-grained texture cues or vision transformers to capture long-range contextual relationships, but rarely integrate the complementary strengths of both families of models. Consequently, many published systems perform well under controlled conditions yet generalize poorly to the heterogeneous settings encountered in the field. Finally, even when technical accuracy is satisfactory, research prototypes seldom mature into user-friendly, real-time tools, limiting their practical value for agronomists, clinicians, and end-users.
In response, the present work makes three intertwined contributions. (i) We assemble and rigorously standardize a large-scale medicinal-herb image repository, cleaning labels and removing low-quality samples to ensure reliable supervision. (ii) We introduce PAHD, a preprocessing pipeline that enhances color constancy and mitigates background noise, and we propose a hybrid ensemble in which CNN backbones and ViT components are jointly optimized to fuse local and global features. (iii) We deploy this model in “Herbify”, a cross-platform application that delivers fast, on-device inference and an intuitive interface for real-time herb identification.
Methodology
The study aims to develop an advanced artificial intelligence framework designed for the accurate identification of common herbs. Figure 1 presents a comprehensive overview of the workflow underlying the methodology. The methodology structured into two primary phases, each focusing on distinct objectives essential for achieving a robust AI model. Phase 1 centers on the creation of a comprehensive and large-scale herb dataset, named the ‘Herbify.’ This phase involves the integration and meticulous cleaning of the sub-datasets, which are processed to form the complete Herbify dataset. The dataset creation process emphasizes rigorous data standardization to ensure high-quality inputs for the model’s subsequent training and application stages. Phase 2 focuses on constructing and deploying the AI framework. It begins with normalizing the Herbify dataset into a consistent format that facilitates effective model training. Following this standardization, the dataset is partitioned into subsets for training, validation, and testing. The main emphasis of this phase is on model training, which employs state-of-the-art techniques such as convolutional neural networks (CNNs) and vision transformers (ViTs). Thereafter, the models are trained and rigorously tested to assess their performance in herb classification. Following training, the framework transitions to the application stage, where CNNs and ViTs are combined through ensembling techniques to optimize classification accuracy. The best performing ensemble model is then incorporated into a fully functional, web-based application designed for end-user accessibility, providing a seamless interface for practical herb identification.
Fig. 1.
The overall two-phase workflow adopted in this study’s methodology. In Phase I (Processing), the raw DIMPSAR corpus is first cleaned and quality-verified, then standardized using the PAHD algorithm. The standardized DIMPSAR data is subsequently merged with the DeepHerb dataset to create the Herbify dataset, which undergoes further manual verification, cleaning and normalization to ensure consistency. In Phase II (Development), the datasets are preprocessed and partitioned into training, validation and test splits; multiple candidate models are then trained and evaluated. The models are then combined into various ensembles. Finally, the top-performing ensemble is selected and deployed within the herb-identification application
The study introduces several notable works, including an advanced preprocessing pipeline, referred to as PAHD (Preprocessing Algorithm for Herb Detection), and the novel integration of the Herbify dataset. It also pioneers the development of hybrid models, created through the ensembling of state-of-the-art CNNs and ViTs, which enhances model performance by leveraging the unique strengths of each architecture. The final product, a user-friendly web application, offers reliable herb recognition capabilities, making the AI framework accessible for practical, real-world applications.
Herb datasets
The study employs two primary sub-datasets—DIMPSAR [9] and DeepHerb [10], along with the newly processed and merged Herbify dataset. The DIMPSAR dataset was individually preprocessed, formatted, and then merged with DeepHerb dataset to create the comprehensive Herbify dataset used for the final analyses. Each dataset is organized in a standardized format, with folders representing herb classes, and all images for a given class stored within the respective folder. Overall, the research encompasses three distinct dataset variations: the processed DIMPSAR dataset, the DeepHerb dataset, and the final, integrated Herbify dataset. This progression from individual datasets to a unified, comprehensive dataset underpins the study’s robust framework for medicinal herb classification.
The herbs and their plants in the datasets possess medicinal properties, with their leaves offering significant benefits for treating various human ailments. Moreover, components of these plants—including the bark, fruits, and seeds are crucial in producing a diverse array of medicinal compounds [10]. Their simple cultivation process makes them ideal candidates for growth in home or community gardens with minimal effort. Disseminating knowledge about these medicinal herbs and their practical applications is essential for increasing public awareness and promoting their use in natural health practices [10, 36].
DIMPSAR dataset
The DIMPSAR dataset comprises images of Indian medicinal plants, curated through fieldwork conducted in various botanical gardens across Karnataka and Kerala, India [9]. To emulate a smartphone-based image acquisition approach, a group of smartphone users captured the images, ensuring the dataset reflects real-world conditions with diverse perspectives, variations in leaf color, multiple resolutions, and differing capture distances. Images were acquired using fixed-lens smartphone cameras, with spatial resolutions ranging from 2560 × 1920 to 5312 × 2988 pixels. This unique acquisition process contributes to the dataset’s authenticity, capturing the challenges of real-time conditions such as fluctuating lighting, occlusions, and variable backgrounds. As a result, the DIMPSAR dataset offers substantial value for applications in plant analysis and recognition in realistic environments. To address class imbalance issues inherent in the dataset, the dataset’s authors employed image augmentation techniques. The original DIMPSAR dataset contained two main image sets: 5900 images representing forty plant species, and a second set comprising single-leaf images of eighty species with a total of 6900 samples [9, 17]. Since this study specifically focuses on single-leaf image-based herb detection, only the latter set, which originally contained eighty plant species, was selected. Notably, this set included duplicate classes, which, after removing similar images, was reduced to seventy-nine distinct plant species. The original dataset presented several quality concerns, including low-resolution images, noise, background biases, motion artifacts, out-of-focus regions, suboptimal cropping, and occlusions. To address these issues, a thorough manual cleaning process was conducted. Images with particularly low resolution, high background bias, significant motion artifacts, or severe occlusions were removed, while those with minor issues, such as out-of-focus regions or poor cropping, were manually corrected. Given the dataset’s varied backgrounds and inherent inconsistencies, a novel preprocessing algorithm named PAHD was introduced to standardize the images, ensuring a consistent format across samples.
The resulting DIMPSAR dataset variation produced by this study consists of 4735 images spanned across seventy-nine classes, each herb class manually cleaned and standardized using the PAHD preprocessing algorithm. This enhanced dataset provides a reliable foundation for herb detection and medicinal plant classification in real-world applications. The scientific names of the herb species included in the final, refined version of the DIMPSAR dataset are as follows: Allium cepa, Aloe barbadensis miller, Andrographis paniculata, Annona squamosa, Artocarpus heterophyllus, Azadirachta indica, Bacopa monnieri, Bambusa vulgaris, Basella alba, Brassica oleracea, Calotropis gigantea, Capsicum annuum, Cardiospermum halicacabum, Carica papaya, Catharanthus roseus, Chamaecostus cuspidatus, Cinnamomum camphora, Citrus limon, Citrus medica, Coffea arabica, Coleus amboinicus, Colocasia esculenta, Coriandrum sativum, Cucurbita, Curcuma longa, Cymbopogon, Ducati Panigale, Eclipta prostrata, Eucalyptus globulus Labill, Euphorbia hirta, Gomphrena globosa, Graptophyllum pictum, Hibiscus rosa-sinensis, Hymenaea courbaril, Ixora coccinea, Jasminum, Justicia adhatoda, Lantana camara, Lawsonia inermis, Leucas aspera, Magnolia champaca, Mangifera indica, Manilkara zapota, Mentha, Momordica dioica, Morinda citrifolia, Moringa oleifera, Murraya koenigii, Neolamarckia cadamba, Nerium oleander, Nyctanthes arbor-tristis, Ocimum basilicum, Ocimum tenuiflorum, Papaver somniferum, Phaseolus vulgaris, Phyllanthus emblica, Piper betle, Piper nigrum, Pisum sativum, Pongamia pinnata, Psidium guajava, Punica granatum, Radermachera xylocarpa, Raphanus sativus, Ricinus communis, Rosa rubiginosa, Ruta graveolens, Saraca asoca, Saraca asoca, Solanum lycopersicum, Solanum nigrum, Spinacia oleracea, Syzygium cumini, Tagetes, Tamarindus indica, Tecoma stans, Tinospora cordifolia, Wrightia tinctoria, and Zingiber officinale.
DeepHerb dataset
The DeepHerb dataset was developed to address the lack of available datasets specifically for medicinal herbs. This dataset was collected from various locations across Karnataka, India, with images captured either by mobile phone or scanner. The original dataset contained 2515 images at a resolution of 1600 × 1200 pixels, encompassing forty distinct classes [10]. The openly available iteration of the DeepHerb dataset has been refined to 1835 images spanning thirty classes from the original forty [37]. Since the DeepHerb dataset was already in a cleaned and standardized format suitable for the study’s requirements, no additional cleaning or processing was necessary. It was incorporated directly in its existing form in the current study. The scientific names of the herb species included in the current version of the DeepHerb dataset are as follows: Alpinia Galanga, Amaranthus Viridis, Artocarpus Heterophyllus, Azadirachta Indica, Basella Alba, Brassica Juncea, Carissa Carandas, Citrus Limon, Ficus Auriculata, Ficus Religiosa, Hibiscus Rosa-sinensis, Jasminum, Mangifera Indica, Mentha, Moringa Oleifera, Muntingia Calabura, Murraya Koenigii, Nerium Oleander, Nyctanthes Arbor-tristis, Ocimum Tenuiflorum, Piper Betle, Plectranthus Amboinicus, Pongamia Pinnata, Psidium Guajava, Punica Granatum, Santalum Album, Syzygium Cumini, Syzygium Jambos, Tabernaemontana Divaricata, andTrigonella Foenum-graecum.
Herbify dataset
The Herbify dataset represents an advanced, comprehensive, and expanded compilation of medicinal herb images, derived from the cleaned and processed versions of the DIMPSAR and DeepHerb datasets. This merged dataset underwent additional manual verification, with any anomalies or problematic images carefully removed. Herb species common to both DIMPSAR and DeepHerb were consolidated to create a unified dataset consisting of 91 herb species. Images within the Herbify dataset vary in spatial resolution from 103 × 94 pixels to 4236 × 4447 pixels, with an average resolution of approximately 1267 × 1135 pixels. The dataset includes a total of 6104 images, representing ninety-one unique herb species. Each species class contains between 7 and 163 images, with an average of 67 images per class.
Appendix provides a detailed description of the Herbify dataset. For each herb, the sample image, scientific name, common name, number of available samples, the corresponding sub-dataset, threat status, medicinal properties, and its typical location are detailed in the section.
Dataset preparation
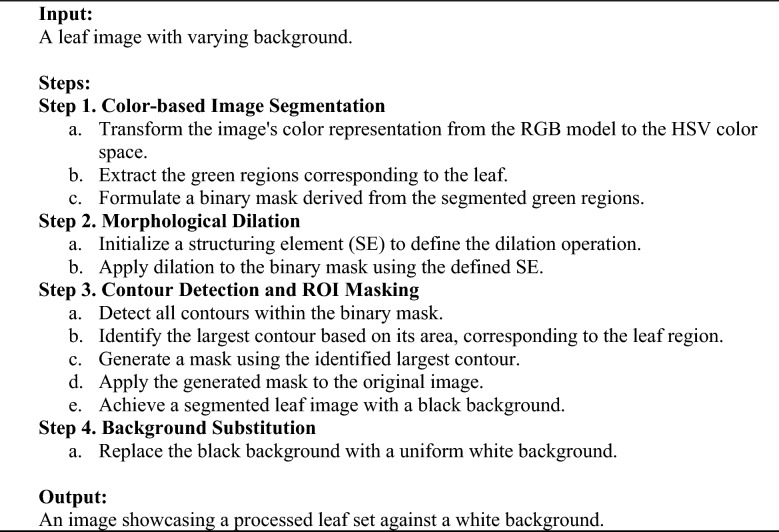
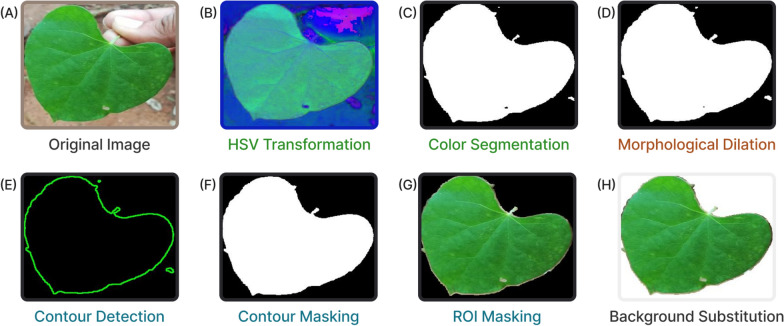
Data preprocessing represents a critical phase in deep learning pipelines, particularly when handling datasets like DIMPSAR, which present challenges due to substantial background variability, irregularities, and extraneous elements. The application of a robust preprocessing algorithm effectively addresses these issues, enhancing image quality and consistency prior to input into deep learning models. For example, images within the DeepHerb dataset are standardized, with only the herb visible against a uniform white background. In contrast, the DIMPSAR dataset presents highly varied backgrounds, often including additional elements such as human hands and other non-essential objects that introduce noise and visual clutter. To address these challenges, a novel preprocessing approach was implemented, referred to as the Preprocessing Algorithm for Herb (object) Detection (PAHD). The PAHD algorithm systematically reduces background interference and extraneous elements in DIMPSAR images, facilitating clearer input for model training. Figure 2 illustrates the step-by-step application and outcome of PAHD, while Algorithm 1 provides a detailed breakdown of each stage. An example of background clutter is shown in Fig. 2 (A), where a leaf is depicted along with a visible hand holding it. Such extraneous details can introduce inconsistencies in learning processes during model training, potentially affecting model performance.
Fig. 2.
Workflow of the Preprocessing Algorithm for Herb Detection (PAHD), comprising RGB → HSV conversion, color-based leaf segmentation, morphological dilation, contour detection and masking, ROI extraction, and background substitution
Data augmentation is a fundamental preprocessing strategy that significantly boosts both the volume and diversity of data available for model training. Applying these transformations exclusively to the training set yields multiple variants of the original data, thereby strengthening the model’s resilience and improving its generalization capabilities. Moreover, deep learning architectures typically demand input images to conform to specific size, quality, and format requirements; preprocessing steps guarantee that these conditions are satisfied [38].
This preparatory stage is vital for boosting the model’s ability to detect patterns and extract salient features, even when the dataset possesses inherent limitations. Overall, preprocessing enhances model robustness and improves generalization to unseen data.
The subsequent sections offer a comprehensive overview of the preprocessing methods utilized in this study.
Algorithm 1.
Preprocessing Algorithm for Herb Detection (PAHD)
PAHD: color-based image segmentation
One major challenge in processing herb images is the frequent occurrence of various objects in both the foreground and background, beyond just the leaf itself. Addressing this issue effectively requires leveraging color information, which can be done through transformations in color space. The first step in the PAHD algorithm is thus color-based image segmentation, where green regions are extracted to target the segmentation of the leaf. The process starts by converting the input image from the RGB (Red, Green, and Blue) model into the HSV (Hue, Saturation, and Value) format.
The RGB color space, while prevalent in digital imaging, is suboptimal for color-based segmentation tasks due to its sensitivity to lighting variations, which often leads to inconsistent pixel intensity values for similarly colored objects. By contrast, the HSV color space separates color into three distinct channels—hue, saturation, and value offering more stable and reliable segmentation under variable lighting conditions. In particular, hue indicates the specific color (for example, red, green, or blue), saturation measures how vivid or pure that color is, and value reflects its brightness or lightness [39]. The equations for converting an image from RGB to HSV are provided in Eqs. 1–5, wherein the RGB color model, the values R, G, and B indicate the intensity of the red, green, and blue channels, respectively. Conversely, in the HSV model, H, S, and V correspond to hue, saturation, and value [40]. This separation enhances segmentation accuracy, making it especially useful in real-world images with uncontrolled lighting. Figure 2 (B) displays an example of an image converted to HSV.
| 1 |
| 2 |
| 3 |
| 4 |
| 5 |
Once the image is converted to HSV, segmentation focuses on isolating green regions that represent the leaf while excluding extraneous elements. Since green is the dominant color in plant life, a defined range of HSV values captures the varied hues, saturations, and brightness levels of green typically found in vegetation. In particular, the following HSV thresholds are defined to encompass a wide range of green hues, the lower bound hue is set at 35°, corresponding to a conventional green in the color spectrum. Saturation and value for the lower bound are set to 40 and 50, respectively, to eliminate darker regions and low-chroma areas, such as background noise or shadows. For the upper bound, the hue extends to 85°, encompassing a wider range of green tones. The saturation and value are set to their maximum (255), allowing the algorithm to capture all chromatic variations and ensure that both dark and light green shades are effectively included. Figure 2 (C) illustrates a sample binary mask of a green leaf, extracted through these bounds. Applying these defined bounds in the HSV color space generates a binary mask that isolates the green regions associated with the leaf, effectively filtering out non-green elements such as the background and other extraneous objects.
PAHD: morphological dilation
The next stage of the PAHD algorithm involves applying morphological dilation, a technique essential for refining the extracted binary mask to achieve accurate representation of the regions of interest (ROIs) and address any minor gaps or imperfections. In image processing, morphological techniques like dilation, erosion, opening, and closing are extensively employed to adjust the structure of objects in binary images [41]. Dilation is a morphological operation that fuses two sets by performing vector addition with a given structuring element (SE). When this process is applied to an image I using the SE, it results in a new binary image B, as defined by Eq. 6 [42]:
| 6 |
In simpler terms, each white (foreground) pixel in the binary mask is expanded outward, filling small gaps and linking fragmented sections of the leaf. The structuring element used for dilation—often a small matrix, such as a (3 × 3) or (5 × 5) square or circular shape, defines the extent of this expansion. This element acts as a neighborhood radius for each pixel and determines the degree of foreground region enlargement. The dilation operation is further described by Eq. 7, which highlights how bright regions are enhanced and expanded [43]:
| 7 |
Applying dilation to the binary mask of the leaf’s green regions provides several key benefits. First, it helps smooth jagged or irregular edges that may result from noise in the original image or imperfect segmentation thresholds. Dilation also effectively bridges small, disconnected green regions, which may occur due to image noise, shadow effects, or incomplete segmentation. Furthermore, it incorporates smaller or fragmented leaf sections that might have been missed due to thresholding errors or low contrast, thereby creating a more continuous and complete representation of the leaf structure. Finally, dilation enhances the mask’s contours, facilitating more accurate contour detection in subsequent processing stages. Figure 2 (D) displays an instance of the mask after the dilation process, illustrating the improved continuity and clarity of the leaf structure.
PAHD: contour detection and ROI masking
Following morphological dilation, the next critical stage in the PAHD pipeline is contour detection within the processed binary mask, alongside Region of Interest (ROI) masking. Contours are essential for accurately delineating the boundaries within an image, facilitating the isolation of the leaf as the primary region of interest. By extracting contours from the dilated binary mask, the algorithm precisely outlines the leaf, effectively excluding any remaining background artifacts or non-leaf regions. The contour detection process focuses on identifying only the outermost contours, which simplifies the task by capturing the primary boundary of the leaf without unnecessary complexity [44]. Figure 2 (E) illustrates an example of the contours that have been detected in the sample leaf image.
After identifying the contours, they are sorted according to their area, calculated using Green’s Algorithm [45]. The formula for area calculation via Green’s theorem is shown in Eq. 8 [46]:
| 8 |
The largest contour, typically corresponding to the main object of interest (in this case, the leaf), is selected. Selecting the largest contour enhances robustness by reducing the likelihood of including noise or smaller, irrelevant regions, such as residual green fragments caused by lighting variations or image noise.
With the largest contour identified, a binary mask is generated to enclose this contour, resulting in a refined representation of the leaf boundary. This contour mask is created by filling in the largest contour, producing a binary mask where all pixels within the contour are marked as foreground (white), and pixels outside the contour are designated as background (black). This high-precision mask precisely delineates the leaf and is prepared for the final ROI extraction stage [47]. Figure 2 (F) presents a sample of the binary mask.
In the final stage of this process, the original image is filtered with the contour mask to isolate the leaf using ROI masking. By overlaying the contour mask onto the original image, only the pixels corresponding to the leaf contour are preserved, while all other areas are effectively masked out, leaving a clean image of the leaf alone [48]. The equation for generating the ROI-extracted image, where I is the image and mask is the binary mask, is shown in Eq. 9:
| 9 |
The resulting image, depicted in Fig. 2 (G), displays the detected leaf against a black background. This final output presents a segmented image where the leaf stands out prominently, with all non-leaf areas removed.
PAHD: background substitution
In the final stage of the PAHD process, the isolated leaf previously set against a black background is transferred onto a white background to ensure consistency with the formatting of the DeepHerb dataset. This background replacement involves substituting all black pixels in the segmented image with white pixels, while preserving the color and detail of the leaf itself. The transformation is achieved by creating a uniform white background layer and applying it to areas of the image where the contour mask does not encompass the leaf. This white background provides a neutral, uniform backdrop, minimizing any background variation and enhancing the contrast between the leaf and the surroundings. This process is essential for maintaining consistency. Additionally, this step mitigates the potential influence of background color on learning algorithms, as it reduces the likelihood of overfitting to background patterns or inconsistencies, which can be especially problematic in complex deep learning models [49]. Figure 2 (H) illustrates the final result image, highlighting the major difference compared to the original image, with background inconsistencies removed and the target object, the leaf, properly isolated.
Once background replacement is complete, the processed DIMPSAR dataset variant once again undergoes a rigorous review and cleaning process to ensure quality and consistency across all images. This inspection process verifies that no background elements or artifacts are present and confirms that each leaf is fully isolated with sharp, accurate contours. Therefore, rigorous quality control is essential for maintaining the dataset’s integrity. Even slight discrepancies or subtle background variations can adversely influence the performance of machine learning models that depend on this data. Furthermore, the review process enables the identification and correction of any errors in segmentation that may have occurred during the PAHD algorithm application.
Augmentation
A primary obstacle in crafting high-performance deep neural networks lies in sourcing sufficiently large and heterogeneous datasets [50]. These models depend on abundant training data to deliver reliable, accurate, and robust outcomes. Comprehensive datasets, by offering a wide spectrum of examples, enhance the network’s learning process and its ability to generalize. To meet these requirements, data augmentation has emerged as an indispensable preprocessing technique in neural network workflows. This method artificially enlarges the training set by applying diverse transformations to existing samples, thus increasing both the size and variability of the dataset [51].
In this study, multiple data augmentation techniques were systematically applied to the training subsets of all datasets, with transformations carefully selected to enhance the model’s adaptability to varied inputs. The applied techniques included horizontal and vertical flips, image transposition, adjustments in brightness and contrast, modifications to tone curves, gamma adjustments, and the addition of blurring effects. Noise, such as ISO and Gaussian noise, was also introduced to simulate real-world conditions [52]. These augmentations were implemented using the Albumentations library in Python, which supports a range of robust, efficient image transformation methods [53].
Adjusting image dimensions
Before training commenced, every image in the dataset was rescaled to the spatial specifications demanded by the respective architectures. Convolutional neural networks processed inputs standardized at 256 × 256 pixels, whereas vision transformers operated on frames resized to 224 × 224 pixels—the resolution conventionally prescribed for these models. Enforcing these uniform dimensions ensured consistent data flow during both training and evaluation and bolstered each model’s capacity to generalize when faced with unseen, real-world imagery.
Image normalization
Normalization serves as another core preprocessing strategy, particularly beneficial for image datasets, and is anticipated to enhance the quality of the herb dataset [54]. This process involves standardizing a tensor image by adjusting it based on its mean and standard deviation, which ensures that the input features are uniformly scaled and centralized. Such standardization is critical for achieving improved convergence during model training. By choosing defined mean and standard deviation values, i.e., reflecting the first and second order statistics for each channel, the z-score for the data can be computed on a channel-specific basis. This process is commonly known as standardization [55], expressed as
| 10 |
Here, inputchannel denotes the input data, meanchannel represents the channel mean, and stdchannel refers to the channel standard deviation. By adjusting each channel of the image data to mimic a standard normal distribution, normalization will significantly enhance the model’s ability to generalize across diverse herb image sets.
Dataset splitting
To optimize herb identification with CNN and ViT architectures, the dataset was stratified into three subsets. Seventy percent was devoted to training, while the remaining thirty percent was split evenly—fifteen percent for validation and fifteen percent for testing. This structured allocation enabled evaluation at successive phases: the training portion furnished ample leaf images for pattern learning; the validation portion tracked performance during training and highlighted generalization behavior; and the independent test portion provided a rigorous assessment on unseen herb images, delivering an unbiased estimate of real-world applicability [35].
Models development
This study investigates sophisticated deep-learning schemes for identifying herbs, leveraging convolutional neural networks alongside vision transformers to achieve high-accuracy classification. The CNN suite comprises MobileNet v3-Large, VGG-19, ResNet-152, and EfficientNet v2-Large, whereas the transformer lineup includes ViT-Base/16 and ViT-Large/16. To further elevate predictive power, we constructed ensembles that merged outputs from selected CNN and ViT models, thereby yielding a sturdier and more precise classifier. Figure 3 summarizes the core architectural elements of the CNN and ViT approaches.
Fig. 3.
Schematic depiction of (A) Convolutional Neural Network architectures and (B) Vision Transformer architectures
Convolutional networks
Deep learning has driven transformative advances in many fields, with convolutional neural networks at the forefront of these achievements. CNNs have been extensively refined and widely adopted for image‐analysis tasks [56]. Their hallmark convolutional layers excel at capturing local spatial patterns in visual data. A typical CNN consists of four principal layer classes—convolution, max-pooling, fully connected, and output—arranged sequentially [35, 57]. This modular design affords substantial flexibility, allowing models to be tailored to domain-specific goals such as automated herb classification. Figure 3 (A) illustrates the core configurations of CNNs.
Consequently, CNNs operate simultaneously as sophisticated feature encoders and discriminative classifiers. The feature value at location (i, j) in the k-th map of l-th layer, [58] is given by
| 11 |
Applying the nonlinear activation a(.) produces
| 12 |
Let pool(.) denote the pooling operator and the neighborhood centered at (i, j). The pooled response for feature map is then expressed as [59]
| 13 |
Through the combined action of convolutional and pooling layers, CNNs extract salient representations from the input, while fully connected layers leverage these abstractions for classification. To model complex patterns, CNNs employ diverse activation functions and span architectures of varying depth, width, and parameter counts [35, 60].
This study evaluates several representative CNN architectures. MobileNet v3-Large, selected for its lightweight depth-wise separable convolutions, offers rapid inference with minimal computational overhead [61]. VGG-19, developed by the Visual Geometry Group and comprising 19 layers with over 180 million parameters, is computationally demanding but delivers high precision in capturing fine textures and intricate structures, making it well suited to herb recognition [62, 63]. ResNet-152, a deep Residual Network, utilizes skip connections to mitigate vanishing‐gradient issues and effectively learns complex feature hierarchies [64, 65]. EfficientNet v2-Large balances depth, width, and resolution to achieve strong accuracy while maintaining relatively modest parameter and FLOP counts [66].
Vision transformers
Transformers’ extraordinary achievements in natural-language processing have redirected computer-vision research, inspiring a detailed examination of their capacity to solve sophisticated visual tasks [67]. Their principal strength lies in modelling long-range dependencies while executing computations in parallel—something conventional recurrent neural networks (RNNs) cannot do efficiently [68, 69]. In vision, their popularity stems from the self-attention mechanism, which encodes extended contextual relationships far more effectively than RNNs or LSTMs, whose performance typically degrades when sequences grow long. By explicitly capturing pairwise interactions, self-attention yields pronounced performance gains across numerous applications [70].
Recent advances—most notably the Vision Transformer (ViT) family—demonstrate that self-attention-centric models can outperform classical convolutional neural networks on a breadth of vision benchmarks [71]. ViT splits an image into fixed-size patches, flattens each patch into a vector of raw pixel intensities, and linearly projects these vectors to the model’s input dimension. Treating the resulting sequence analogously to words in a sentence allows the network to exploit self-attention for learning spatial relationships [35, 72]. Figure 3 (B) illustrates the vision transformers architecture.
Let denote a sequence of n tokens , each embedded in d dimensions. Self-attention projects the sequence into query, key, and value spaces using trainable matrices , , and (with ). The projections are , , and [73]. The layer output is
| 14 |
which enables every token to be refined by global contextual information.
In this work we employ two ViT variants: ViT-Base/16 (ViT-B/16) and ViT-Large/16 (ViT-L/16). ViT-B/16 comprises 12 Transformer encoder blocks and offers a computationally economical choice, whereas ViT-L/16 doubles the depth to 24 encoders, trading greater resource demands for superior accuracy [74]. Consequently, ViT-B/16 is well-suited to settings with restricted hardware budgets, while ViT-L/16 targets scenarios that prioritize higher capacity and precision.
Training procedure
Developing a reliable herb-identification system necessitates training learning algorithms on extensive, high-quality image repositories. Running large-capacity networks on limited data, however, incurs significant computational costs and heightens the danger of overfitting [75]. To address these issues, the study adopts transfer learning procedure, which re-purposes the models pretrained on massive datasets such as ImageNet so that their broad visual representations can be specialized for herb imagery [76].
CNNs and ViTs were initialized with ImageNet weights, after which their output layers were replaced with fully connected layers sized for each dataset: 79 classes for DIMPSAR, 30 for DeepHerb, and 91 for Herbify. An overview of modulation is shown in Fig. 4. The approach further employed fine-tuning, allowing gradients to adjust a large subset or all of the pretrained parameters rather than limiting updates to the added layers alone. Fine-tuning is especially effective when the target domain diverges substantially from the pretraining domain, as is the case here [77].
Fig. 4.
Transfer learning pipeline for herb classification: the pre-trained CNN/ViT’s original classification head is removed, replaced with a newly initialized head, and the entire model is then fine-tuned on the herb dataset
Adaptive Moment Estimation (Adam) optimized the CNNs, leveraging its ability to combine AdaGrad and RMSProp advantages for sparse-gradient, non-stationary tasks [78]. For ViTs, the AdamW variant was used because decoupling weight decay from the gradient step enhances regularization, stabilizes transformer training, and curtails overfitting [79].
Learning rate and weight decay, two crucial hyperparameters governing convergence were exhaustively explored for Herbify via Grid Search, which systematically tests predefined combinations to locate the optimal setting for the architecture and dataset [80]. Meticulous tuning of these values markedly improved training efficiency and predictive accuracy.
Training relied on the multi-class cross-entropy loss [72], expressed as
| 15 |
where is the ground-truth indicator for sample i and class j, is the predicted probability, N is the number of samples, and M is the number of herb species [81]. Minimizing this loss via the chosen optimizer progressively refines network parameters and elevates classification accuracy.
Convolutional neural networks and vision transformers ensembles
Once separate convolutional neural networks and vision transformer models had been fine-tuned on the herbify dataset, we merged them using an ensemble strategy. Harnessing the complementary strengths of CNNs and ViTs markedly improves classification accuracy—critical when an error could misidentify an herb species and propagate inaccurate medical information. CNNs specialize in capturing local structure through hierarchical convolutions, proving invaluable for detection and segmentation tasks [82, 83]. Fusing CNN-derived local features with the wide-context representations of ViTs yields a generalizable model that performs robustly across varied data distributions [84].
In the ensemble, the original classification heads were removed and replaced with layers dedicated to feature fusion. Feature maps from the CNN and ViT branches were concatenated, then passed into a fully connected layer generating 1024-dimensional representations.
These activations were fed through a Rectified Linear Unit (ReLU), enabling non-linear modelling [85]:
| 16 |
A second fully connected layer projected the 1024 features to the exact class count for each dataset—79 for DIMPSAR, 30 for DeepHerb and 91 for Herbify. A soft-max layer then converted the logits into probabilities [86]:
| 17 |
where N is the number of classes and the i-th logit.
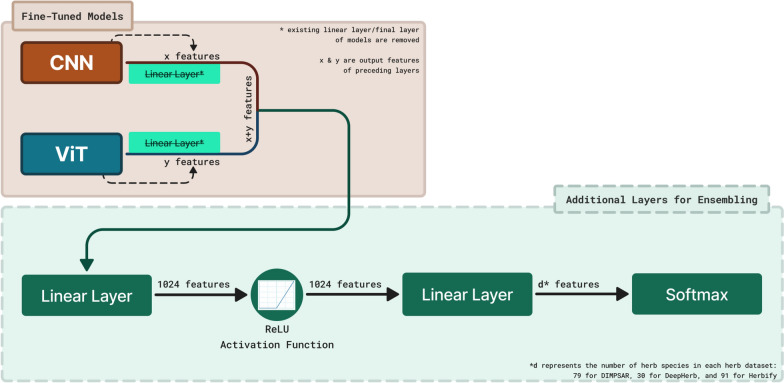
Figure 5 illustrates the ensemble model’s overall structure, demonstrating the integration of CNN and ViT models for enhanced herb classification. For the ensemble models, only the newly added layers were trained by applying the transfer learning procedure. The pre-trained layers were frozen to ensure that no updates were made to the previously learned features. This approach preserves the knowledge acquired during fine-tuning [35].
Fig. 5.
Ensemble architecture combining CNN and ViT backbones: trained classification heads are removed, backbone feature embeddings are combined and fed into a series of MLP layers (Linear → ReLU → Linear), with a soft-max output for final herb classification
Eight ensemble variants were evaluated, each engineered to capitalise on the unique capabilities of its component networks (Table 1). By aggregating multiple learners, every ensemble delivers heightened performance and resilience across diverse deployment scenarios.
MobileL-ViTB (MobL-VB) combines MobileNet v3-Large and ViT-Base/16, striking an optimal balance between efficiency and rich feature representation—ideal for light-scale cloud services demanding fast yet accurate inference.
VGG19-ViTB (V19-VB) couples VGG-19 with ViT-Base/16, excelling when meticulous, context-aware analysis is paramount and model size is a secondary concern.
Res152-ViTL (R152-VL) unites ResNet-152 with ViT-Large/16, offering deep capacity for complex imagery, while EfficientL-ViTL (EffL-VL) blends EfficientNet v2-Large with ViT-Large/16 to provide a scalable compromise between accuracy and efficiency. Both serve large-scale cloud deployments requiring state-of-the-art accuracy with moderate resource use.
MobileL-VGG19-ViTB (MobL-V19-VB) and Res152-EfficientL-ViTL (R152-EffL-VL) extend capacity further by integrating multiple high-capacity CNNs with ViTs, delivering superior precision where both speed and accuracy are critical.
The most comprehensive ensembles, MobileL-VGG19-EfficientL-ViTB (MobL-V19-EffL-VB) and VGG19-Res152-EfficientL-ViTL (V19-Res152-EffL-VL), combine virtually all leading CNN families with ViTs. These configurations target high-stakes scenarios—such as advanced herb recognition research—where maximum predictive power is prioritised and ample computational resources are available.
Table 1.
Structural overview of ensemble models used in the study
| Models | Ensemble Model | |||||
|---|---|---|---|---|---|---|
| Convolutional Neural Networks | Vision Transformers | |||||
| MobileNet v3-Large | VGG-19 | ResNet-152 | EfficientNet v2-Large | ViT-Base/16 | ViT-Large/16 | |
| ✓ | × | × | × | ✓ | × | MobileL-ViTB |
| × | ✓ | × | × | ✓ | × | VGG19-ViTB |
| × | × | ✓ | × | × | ✓ | Res152-ViTL |
| × | × | × | ✓ | × | ✓ | EfficientL-ViTL |
| ✓ | ✓ | × | × | ✓ | × | MobileL-VGG19-ViTB |
| × | × | ✓ | ✓ | × | ✓ | Res152-EfficientL-ViTL |
| ✓ | ✓ | × | ✓ | ✓ | × | MobileL-VGG19-EfficientL-ViTB |
| × | ✓ | ✓ | ✓ | × | ✓ | VGG19-Res152-EfficientL-ViTL |
Together, the ensemble suite addresses deployment scenarios ranging from resource-constrained edge devices to high-throughput, high-resolution analytical platforms, providing adaptable solutions for diverse operational requirements [20, 87].
Performance evaluation
The devised models underwent a rigorous examination using a broad spectrum of metrics to ensure a thorough appraisal of their herb-classification capability [88]. Core indicators comprised Accuracy, Sensitivity, Specificity, Precision (also known as Recall), and the F₁-Score [89]. To counter the skew often present in herb datasets, the Geometric Mean (G-Mean) was likewise calculated, as it reliably reflects performance across imbalanced class distributions [90]. Together, these measures offer a multidimensional view of predictive effectiveness, each highlighting a distinct facet of classifier behavior. Equations 18–23 formalize these metrics.
| 18 |
| 19 |
| 20 |
| 21 |
| 22 |
| 23 |
where TP, TN, FP, and FN denote true positives, true negatives, false positives, and false negatives, respectively [91].
Additionally, plots of training accuracy and loss were produced to enhance interpretability. These visualizations track learning progress over time, revealing trends that help detect overfitting or underfitting and thus deepen insights into learning stability, model behavior, and anticipated generalizability [92].
Proposed application
This research introduces “Herbify,” a cross-platform web application tailored to botanists, specialists, and plant enthusiasts. Illustrated in Fig. 6, the system emphasizes an intuitive, streamlined interface that enables straightforward navigation and rapid inference. Given the widespread availability of smartphones, including in remote areas, the application is optimized for mobile use to maximize reach and usability across diverse populations [93]. Core web technologies—HTML, CSS, and JavaScript—constitute the front end. Backend tasks are orchestrates by a Flask server. To compensate for the limited computational power of mobile devices, resource-intensive operations such as image processing and herb identification are delegated to cloud servers. Users simply capture a photograph of a specimen; the image is then transmitted to the cloud for high-accuracy analysis and inference.
Fig. 6.
Pipeline overview of Herbify, a web application for herb identification. A user-captured image is uploaded to the cloud, standardized by the PAHD algorithm, classified by an ensemble model, and the predicted herb is displayed to the user
The processing pipeline commences with the Preprocessing Algorithm for Herb Detection (PAHD), which isolates the leaf, suppresses extraneous noise and background elements, and substitutes the background with a uniform white plane to prepare the image for recognition. For classification, the server deploys an advanced ensemble of CNNs and ViT, accurately identifying the plant and detailing its characteristics. After inference, the application returns the refined image alongside the specimen’s scientific and common names, furnishing users with a comprehensive identification report [94]. By combining affordability, accessibility, and ease of use, “Herbify” aspires to become a pivotal resource for botanical research, enhancing field studies through an efficient digital platform.
Setup
The development and training was carried out on a desktop workstation featuring an NVIDIA RTX 3080 Ti GPU, an Intel® Core™ i9-10900 K CPU, 64 GB of RAM, and 1 TB of SSD storage. Data preprocessing was performed using the OpenCV and Scikit-image libraries [95, 96]. Deep learning architectures—including convolutional neural networks, vision transformers, and ensemble constructs—were developed and trained within the PyTorch framework [97]. Evaluation metrics were computed with the Scikit-learn library [98], while graphical visualizations were created using the Matplotlib and Seaborn libraries [99, 100]. This setup facilitated efficient model training and evaluation, leveraging the capabilities of both hardware and software to achieve high-performance results.
Results and discussion
In this section, we deliver a detailed synthesis of the study’s achievements, emphasizing the herb datasets, the stages of model development, the experimental outcomes and insights gained, and the evaluation of various deep learning architectures. We examine convolutional neural networks (CNNs), vision transformers (ViTs) and their ensemble strategies, as well as the design and implementation of a dependable, high-precision herb recognition application.
Herb datasets
The study utilized three herb datasets: the DIMPSAR dataset with 79 herb species, the DeepHerb dataset with 30 herb species, and the cleaned and merged Herbify dataset containing 91 herb species. On the DIMPSAR dataset, the Preprocessing Algorithm for Herb Detection (PAHD) was applied after initial cleaning, cropping, and error correction to bring the DIMPSAR dataset to a unified standard. For the Herbify dataset, an extensive manual inspection was conducted after the merger to ensure its resilience and reliability.
We employed a uniform partitioning scheme for all datasets, designating 70% of the samples for training, 15% for validation, and the remaining 15% for testing. To bolster model resilience, the training subsets were subjected to comprehensive augmentation. These procedures ranged from fundamental operations—such as scaling and rotation—to more advanced manipulations, including noise injection to emulate real‐world variability. Table 2 summarizes the original dataset volumes (pre‐augmentation), the exact sample counts in each partition, and the expanded sizes of the augmented training sets across the three datasets.
Table 2.
Summary of herb datasets: total size and distribution across training (with and without augmentation), testing, and validation subsets
| Dataset | Total Number of Samples | Training Split (original) | Training Split (augmented) | Validation Split | Testing Split |
|---|---|---|---|---|---|
| DIMPSAR | 4735 | 3279 | 19,674 | 746 | 710 |
| DeepHerb | 1835 | 1273 | 7638 | 290 | 272 |
| Herbify (merged) | 6104 | 4233 | 25,398 | 958 | 913 |
Hyperparameters
Hyperparameters play a critical role in the training and optimization of deep learning models. To ensure optimal performance, hyperparameters were selected with meticulous care. Consistent hyperparameters for CNNs, ViTs, and ensemble models on the Herbify dataset are presented in Table 3. However, for CNN models on the DIMPSAR and DeepHerb datasets, variations in the learning rate and weight decay were applied, tailored to the specific dataset size and characteristics to mitigate overfitting and underfitting issues. These values are provided in Table 5.
Table 3.
Fixed hyperparameter settings for all models across herb datasets
| Hyperparameters | CNN | ViT | Ensemble model |
|---|---|---|---|
| Optimizer | Adam | AdamW | Adam |
| Batch Size | 16 | 16 | 16 |
| Max Epochs | 30 | 30 | 15 |
Table 5.
Learning rate and weight decay configurations for models over different herb datasets
| Hyperparameters | DIMPSAR | DeepHerb | Herbify (optimal) | ViT (All datasets) | Ensemble M. (Herbify DS) |
|---|---|---|---|---|---|
| Learning Rate (LR) | 1 × 10–4 | 1 × 10–4 | 1 × 10–5 | 1 × 10–5 | 1 × 10–4 |
| Weight Decay (WD) | 1 × 10–7 | 1 × 10–8 | 1 × 10–8 | 1 × 10–7 | 1 × 10–8 |
To optimize hyperparameters on the Herbify dataset, we conducted a grid search to pinpoint the optimal learning rate and weight decay. We selected the ResNet-50 architecture for convolutional models due to its favorable trade-off between computational cost and accuracy, and the ViT-Base/16 design for vision transformers. As detailed in Table 4, our grid spanned three candidate values for both learning rate and weight decay, yielding nine unique settings. We trained each configuration for ten epochs, with training times varying between approximately 45 and 120 min per run, depending on the model. Throughout the search, we consistently used the Adam optimizer for ResNet-50 and AdamW for ViT-Base/16, and maintained a batch size of 16.
Table 4.
Hyperparameter ranges for CNN and ViT architectures used in Grid Search
| Hyperparameters | Values |
|---|---|
| Learning Rate | 1 × 10–3, 1 × 10–4, 1 × 10–5 |
| Weight Decay | 1 × 10–7, 1 × 10–8, 1 × 10–9 |
The optimal hyperparameter configurations for CNNs are detailed in Table 5. For ViTs, the previously selected learning rate (1 × 10⁻5 or 1e-05) and weight decay (1 × 10⁻⁷ or 1e-07), utilized for training models over the DIMPSAR and DeepHerb dataset turned out to be the optimal hyperparameters, therefore these values remained unchanged across all datasets.
We applied similar hyperparameter tuning procedure to the ensemble architectures. By training only the newly appended layers, we preserved the pretrained representations and thus mitigated the risk of catastrophic forgetting. The hyperparameter values ultimately chosen for these ensembles, listed in Table 5, embody an optimal trade-off that enables each constituent model to contribute its strengths while upholding computational efficiency.
This structured approach to dataset preparation, augmentation, and hyperparameter tuning underpinned the robust performance observed across all models, laying the foundation for the successful deployment of an accurate herb recognition system.
Training and evaluation of models
Once preprocessing was complete, deep learning architectures were trained on three herb image collections via a fine-tuning strategy applied to both convolutional neural networks and vision transformers. The CNN architectures comprised MobileNet v3-Large, VGG-19, ResNet-152, and EfficientNet v2-Large, while the ViT variants included ViT-Base/16 and ViT-Large/16. Each model underwent fine-tuning under a standardized protocol with uniform hyperparameter settings. The DIMPSAR, DeepHerb, and Herbify datasets were split into training, validation, and test subsets, and model performance was evaluated using F₁-score, accuracy, precision, recall, specificity, and G-Mean. Due to pronounced class imbalance and distributional heterogeneity in the data, the F₁-score was selected as the primary metric for comparative analysis.
Training times varied based on the complexity (size and structure) of the models and the size of the datasets. For the DIMPSAR dataset, training durations ranged from 4 to 12 h. The DeepHerb dataset required 2–8 h, whereas Herbify, being the largest dataset, demanded 6–14 h. The most resource-intensive models were EfficientNet v2-Large and ViT-Large/16, while MobileNet v3-Large was the most computationally efficient due to its smaller architecture and fewer parameters. Model checkpoints were saved at each epoch upon achieving high validation accuracy, and the best-performing checkpoint over the validation split was used for testing. A visual representation of the F₁-scores of all models across the datasets in the form of bar graphs is provided in Fig. 7.
Fig. 7.
Comparative analysis of F₁-Scores for CNNs, ViTs, and ensemble models (on the Herbify dataset) across herb datasets
Performance analysis on the DIMPSAR dataset
The performance of the models on the DIMPSAR dataset is summarized in Table 6. Among all models, EfficientNet v2-Large achieved the highest F₁-score of 98.16%, alongside excellent accuracy (98.16%), precision (98.41%), sensitivity (98.16%), specificity (99.97%), and a G-Mean of 0.9906. These results underscore its capacity to effectively capture complex features within the dataset.
Table 6.
Evaluation metrics for convolutional neural networks and vision transformers architectures applied to the DIMPSAR dataset
| Model | F₁-Score (%) | Accuracy (%) | Precision (%) | Sensitivity (%) | Specificity (%) | G-Mean |
|---|---|---|---|---|---|---|
| MobileNet v3-Large | 97.45 | 97.46 | 97.79 | 97.46 | 99.96 | 0.9870 |
| VGG-19 | 94.60 | 94.78 | 95.01 | 94.78 | 99.93 | 0.9732 |
| ResNet-152 | 97.36 | 97.32 | 97.70 | 97.32 | 99.96 | 0.9863 |
| EfficientNet v2-Large | 98.16 | 98.16 | 98.41 | 98.16 | 99.97 | 0.9906 |
| ViT-Base/16 | 97.03 | 97.04 | 97.33 | 97.04 | 99.96 | 0.9849 |
| ViT-Large/16 | 96.76 | 96.76 | 97.16 | 96.76 | 99.95 | 0.9834 |
MobileNet v3-Large and ResNet-152 followed closely with F₁-scores of 97.45% and 97.36%, respectively. Comparable specificity (99.96%) and G-Means (0.9870 for MobileNet v3-Large and 0.9863 for ResNet-152) were exhibited by both model. Notably, MobileNet v3-Large outperformed ResNet-152 slightly in sensitivity (97.46% vs. 97.32%), indicating a better ability to identify true positive cases.
The comparison between CNN-based models and Vision Transformers revealed valuable insights. While the CNN models, particularly EfficientNet v2-Large, ResNet-152, and MobileNet v3-Large, demonstrated superior performance, the ViT architectures were competitive. ViT-B/16 achieved an F₁-score of 97.03%, marginally outperforming ViT-L/16, which scored 96.76%. Both ViT models exhibited high specificity (99.96% and 99.95%, respectively), reflecting strong capabilities in identifying negative samples. However, their slightly lower sensitivity and G-Mean values compared to the top-performing CNN models suggest that ViTs may be less effective at recognizing positive cases—a crucial factor given the dataset’s imbalance.
The observed differences between ViT-Base/16 and ViT-Large/16 are likely attributable to their architectural distinctions. ViT-B/16, with fewer parameters, demonstrated marginally better sensitivity and F₁-score, implying that a smaller ViT model generalize better to the dataset. Conversely, the larger capacity of ViT-L/16 did not yield improved performance, indicating inefficiencies in handling this particular dataset.
VGG-19 performed the least out of all models, achieving the least accuracy of 94.78%, with a F₁-score of 94.60%, highlighting the limitations of model’s ability to handle class imbalances. Furthermore, the trade-offs between precision and sensitivity are evident in some models. MobileNet v3-Large and ResNet-152 exhibited slightly higher precision than sensitivity, indicating a marginal bias toward correctly identifying negative samples. Specificity remained consistently high across all models, with values exceeding 99.93%, suggesting that false positives were rare for DIMPSAR dataset.
Figure 7 (A) provides a bar plot comparison of model performance on the DIMPSAR dataset. The training and validation performance trends for the DIMPSAR dataset are illustrated in Fig. 8 (A). Training accuracy curves indicate that most models converged by the third epoch, maintaining high performance thereafter. Simultaneously, all loss values plateaued by the fifth epoch, reflecting the stability and effectiveness of the training process.
Fig. 8.
Training accuracy and loss curves for all models over: (A) DIMPSAR dataset, (B) DeepHerb dataset, and (C) Herbify dataset
Performance analysis on the DeepHerb dataset
The evaluation of deep learning models on the DeepHerb dataset demonstrates exceptional accuracy, with several models achieving near-perfect results. Table 7 presents the performance metrics for the tested models. Among these, EfficientNet v2-Large delivered flawless outcomes, achieving perfect scores across all metrics: F₁-score, accuracy, precision, sensitivity, and specificity, each at 100 (with G-Mean being 1). This impeccable performance underscores the model’s unparalleled ability to accurately classify both positive and negative samples without error, and once again establishing itself as the benchmark. The model’s architecture, incorporating compound scaling and advanced feature extraction techniques, likely underpins its outstanding results, showcasing its capacity to handle even the dataset’s most nuanced variations.
Table 7.
Evaluation metrics for convolutional neural networks and vision transformers architectures applied to the DeepHerb dataset
| Model | F₁-Score (%) | Accuracy (%) | Precision (%) | Sensitivity (%) | Specificity (%) | G-Mean |
|---|---|---|---|---|---|---|
| MobileNet v3-Large | 99.27 | 99.26 | 99.36 | 99.26 | 99.97 | 0.9961 |
| VGG-19 | 99.27 | 99.26 | 99.37 | 99.26 | 99.97 | 0.9961 |
| ResNet-152 | 99.63 | 99.63 | 99.67 | 99.63 | 99.98 | 0.9980 |
| EfficientNet v2-Large | 100 | 100 | 100 | 100 | 100 | 1 |
| ViT-Base/16 | 98.50 | 98.52 | 98.71 | 98.52 | 99.94 | 0.9923 |
| ViT-Large/16 | 99.63 | 99.63 | 99.66 | 99.63 | 99.98 | 0.9980 |
Both ResNet-152 and ViT-Large/16 delivered exceptional results, each achieving an F₁-score of 99.63% and a geometric mean (G-Mean) of 0.9980. Both models maintained high precision (99.67% for ResNet-152 and 99.66% for ViT-L/16) and sensitivity (99.63%), indicating their effectiveness in balancing accurate positive and negative classifications.
MobileNet v3-Large and VGG-19 performed identically, attaining F₁-scores of 99.27% and G-Means of 0.9961. Although they did not surpass the top-performing models, their high specificity (99.97%) and competitive precision and sensitivity values affirm their reliability. MobileNet v3-Large, in particular, stands out for its lightweight architecture, making it a practical choice for deployment in resource-constrained environments such as mobile or edge devices.
ViT-Base/16 achieved the least F₁-score of 98.50% among all the models, which, while slightly lower than the top-tier models, still reflects excellent performance. Its marginally reduced precision (98.71%) and sensitivity (98.52%) suggest a limitation in capturing subtle feature representations compared to larger models such as ViT-L/16 or CNN-based architectures like EfficientNet v2-Large. Nevertheless, its high G-Mean of 0.9923 and specificity of 99.94% confirm its strong generalization ability, particularly in identifying negative samples accurately.
Across all models, specificity exceeded 99.94%, indicating a consistently low false-positive rate. The alignment of high precision and sensitivity values demonstrates a balanced ability to minimize false positives and correctly identify true positives.
These outcomes verify that the DeepHerb dataset constitutes a rigorously compiled and preprocessed resource for herb recognition, enabling multiple models to attain near-flawless performance. In Fig. 7 (B), a bar chart compares each model’s metrics on DeepHerb, while Fig. 8 (B) traces their training accuracy and loss curves. By epoch two, most networks have converged, achieving starting accuracies above 80% and loss values below one (with VGG-19 as the lone exception), thereby outperforming counterparts trained on the DIMPSAR dataset. Although all architectures exhibit high performance, VGG-19 requires additional epochs to stabilize, as its loss curve plateaus later.
Performance analysis on the herbify dataset
The Herbify dataset, created through the integration and further manual cleaning of the DIMPSAR and DeepHerb datasets, serves as a comprehensive benchmark for herb recognition tasks. The manual cleaning process significantly enhanced data quality, enabling models to extract critical features more effectively. Table 8 summarizes the performance of CNN and ViT models on this dataset, which consistently achieved high accuracy across all metrics.
Table 8.
Evaluation metrics for convolutional neural networks and vision transformers architectures applied to the Herbify dataset
| Model | F₁-Score (%) | Accuracy (%) | Precision (%) | Sensitivity (%) | Specificity (%) | G-Mean |
|---|---|---|---|---|---|---|
| MobileNet v3-Large | 98.90 | 98.90 | 99.01 | 98.90 | 99.98 | 0.9944 |
| VGG-19 | 97.17 | 97.26 | 97.33 | 97.26 | 99.96 | 0.9860 |
| ResNet-152 | 99.01 | 99.01 | 99.13 | 99.01 | 99.98 | 0.9949 |
| EfficientNet v2-Large | 99.13 | 99.12 | 99.26 | 99.12 | 99.99 | 0.9955 |
| ViT-Base/16 | 98.84 | 98.90 | 98.87 | 98.90 | 99.98 | 0.9944 |
| ViT-Large/16 | 98.90 | 98.90 | 99.02 | 98.90 | 99.98 | 0.9944 |
EfficientNet v2-Large and ResNet-152 once again emerged as the top performers, achieving F₁-scores of 99.13% and 99.01%, respectively. With EfficientNet v2-Large once again maintaining its lead in all key metrics, including precision (99.26%), sensitivity (99.12%), and specificity, which was nearly perfect at 99.99%. Its G-Mean of 0.9955, the highest among all models, underscores its exceptional balance between sensitivity and specificity, making it the most reliable choice for herb classification. ResNet-152 closely followed, with a specificity of 99.98%.
MobileNet v3-Large achieved an impressive F₁-score of 98.90%, on par with ViT-Large/16 and slightly outperforming ViT-Base/16 (98.84%). Its precision (99.01%) and sensitivity (98.90%) indicate a balanced ability to classify true positives and negatives. Its G-Mean of 0.9944 further supports its robust performance, making it a strong contender for computationally efficient deployments. Both ViT-Base/16 and ViT-Large/16 achieved accuracy of 98.90%, with ViT-L/16 slightly surpassing ViT-B/16 in precision (99.02% vs. 98.87%) while matching in sensitivity and specificity (98.90% and 99.98%, respectively). Their G-Mean scores of 0.9944 highlight their reliability, even though they fall slightly behind CNN-based models.
Although VGG-19 achieved a solid F₁-score of 97.17%, it nonetheless trailed the other architectures on the majority of evaluation metrics. Its precision (97.33%) and sensitivity (97.26%) were comparatively lower, and its G-Mean of 0.9860 reflected less balanced performance. Despite this, VGG-19 remains a viable choice in scenarios prioritizing simpler architectures or minimal resource requirements.
Specificity across all models remained exceptionally high, exceeding 99.96%. The G-Mean metric further highlights balanced performance across metrics. These results confirm the Herbify dataset’s high quality, as the manual cleaning process further effectively eliminated noise and inconsistencies, allowing models to focus on essential features.
The Herbify dataset also demonstrated the capacity to train both CNN and ViT architectures effectively, with CNN models generally outperforming their transformer counterparts across most metrics. Figure 7 (C) shows bar plot comparisons for model performance on Herbify, while Fig. 8 (C) presents the training and loss curves. Training accuracy graphs indicate that most models converged by the ninth epoch, with loss plateauing by the tenth epoch. Initial training accuracy was lower (with around 60 for most models), and initial loss was higher (near 2.5), compared to other datasets. However, once convergence occurred, the training process was smoother and more stable than the counterparts and also yielding excellent performance despite the dataset’s larger size and greater complexity.
Performance analysis of ensemble models on the herbify dataset
Standalone convolutional neural networks and vision transformers attained near-perfect accuracy and F₁ scores on the Herbify benchmark, yet combining them in ensemble configurations yielded even greater robustness and reliability. By uniting CNNs—which excel at extracting fine-grained local patterns—with ViTs, capable of modeling long-range contextual relationships, the system leverages the complementary advantages of both architectures. This combination is especially suitable for navigating the dataset’s large scale and supporting real-time deployment. Training each ensemble required 40–120 min, contingent on the number and nature of the constituent networks, but the overhead remained modest because only a limited set of additional layers needed optimization.
Table 9 details the results for eight ensemble variants, all of which surpassed individual models on most evaluation metrics.
Table 9.
Evaluation metrics for ensemble architectures applied to the Herbify dataset
| Model | F₁-Score (%) | Accuracy (%) | Precision (%) | Sensitivity (%) | Specificity (%) | G-Mean |
|---|---|---|---|---|---|---|
| MobileL-ViTB | 99.17 | 99.23 | 99.20 | 99.23 | 99.9914 | 0.9961 |
| VGG19-ViTB | 98.85 | 98.90 | 98.92 | 98.90 | 99.9878 | 0.9944 |
| Res152-ViTL | 99.35 | 99.34 | 99.42 | 99.34 | 99.9926 | 0.9967 |
| EfficientL-ViTL | 99.56 | 99.56 | 99.61 | 99.56 | 99.9951 | 0.9978 |
| MobileL-VGG19-ViTB | 98.96 | 99.01 | 98.99 | 99.01 | 99.9890 | 0.9950 |
| Res152-EfficientL-ViTL | 99.46 | 99.45 | 99.51 | 99.45 | 99.9939 | 0.9972 |
| MobileL-VGG19-EfficientL-ViTB | 99.33 | 99.34 | 99.38 | 99.34 | 99.9926 | 0.9967 |
| VGG19-Res152-EfficientL-ViTL | 99.33 | 99.34 | 99.39 | 99.34 | 99.9926 | 0.9967 |
The ensembled models achieved F₁-scores ranging from 98.85 to 99.56%, compared to the individual models’ range of 98.90–99.13%. Notably, all ensembles, except for VGG19-ViTB and MobileL-VGG19-ViTB, exceeded the F₁-score of 99.17% on the Herbify dataset. Among them, the EfficientL-ViTL ensemble achieved the highest F₁-score of 99.56%, alongside perfect accuracy, precision, sensitivity, and G-Mean metrics, establishing it as the top-performing ensemble. Its near-perfect specificity of 99.9951% underscores its ability to minimize false positives, while its G-Mean of 0.9978 highlights its balanced performance. The synergy between EfficientNet v2-Large’s strong feature extraction capabilities and ViT-L/16’s global attention mechanism likely contributes to this exceptional performance.
The Res152-EfficientL-ViTL ensemble closely followed, with an F₁-score of 99.46% and a G-Mean of 0.9972. Although slightly behind EfficientL-ViTL in precision and sensitivity, its high specificity (99.9939%) and balanced metrics confirm its reliability. Similarly, the Res152-ViTL ensemble achieved an F₁-score of 99.35% and a G-Mean of 0.9967. Its high precision (99.42%) and sensitivity (99.34%), and its specificity of 99.9926% reflects its ability to minimize false positives. The ensemble effectively combines ResNet-152’s spatial feature extraction with ViT-L/16’s attention-based feature refinement.
Largest ensembles, such as MobileL-VGG19-EfficientL-ViTB and VGG19-Res152-EfficientL-ViTL, demonstrated comparable performance, achieving identical metrics, with F₁-score of 99.33%. Although they did not outperform EfficientL-ViTL or Res152-EfficientL-ViTL, they remain competitive alternatives with strong classification capabilities. These ensembles are particularly suitable for scenarios requiring advanced real-time integration, as they exhibit higher confidence in predictions.
Simpler ensembles like MobileL-ViTB and VGG19-ViTB achieved F₁-scores of 99.17% and 98.85%, respectively. While these results are commendable, they fall short of the top-performing ensembles. Their relatively lower precision and sensitivity suggest that these simpler combinations may lack the complementary depth and diversity offered by more complex ensembles like EfficientL-ViTL or Res152-EfficientL-ViTL. The MobileL-VGG19-ViTB ensemble, with an F₁-score of 98.96%, slightly outperformed VGG19-ViTB but remained behind the highest-performing ensembles. These findings suggest that increased complexity does not always guarantee proportional performance gains, particularly when the individual models in the ensemble have weaker or overlapping strengths.
Specificity across all ensembles remained exceptionally high, exceeding 99.9878%, ensuring minimal false positives—a critical requirement for herb classification tasks where errors could have significant downstream implications. The alignment between precision and sensitivity metrics for all ensembles indicates balanced performance in classifying both positive and negative samples. The consistently high G-Means highlight the effectiveness of ensemble learning in leveraging individual model strengths to achieve reliable and balanced classifications. Figure 7 (D) presents a bar plot comparing ensembled‐model performance on the Herbify dataset, with each model identified by its abbreviated name.
In terms of inference time, lighter ensembles like MobileL-ViTB and VGG19-ViTB demonstrated an average inference time of 1 to 3 s, making them ideal for deployment in edge devices. Larger ensembles, such as MobileL-VGG19-EfficientL-ViTB and VGG19-Res152-EfficientL-ViTL, required an average of 10–15 s per inference, with remaining ensembles having average inference time of 4–10 s. The best-performing ensemble, EfficientL-ViTL, struck an optimal balance between computational resource requirements and accuracy, with an average inference time of 2–4 s. All inference times were measured under the experimental setup described in the methodology.
Within both the ensemble and the individual classifiers, the predominant source of error stems from the high degree of morphological similarity among certain herb species—especially those represented by only a handful of training samples. When two species share nearly identical leaf shapes, venation patterns, or surface textures, the deep neural networks may struggle to extract sufficiently discriminative features. This challenge is compounded by severe class imbalance: small‐sample classes provide inadequate variance for the model to learn robust intra‐class representations, leading to overfitting on the limited examples available and, consequently, to misclassification when presented with novel instances.
In our experiments, the ensemble model most frequently confused the following pairs and groups of species: Annona squamosa, Artocarpus heterophyllus, Citrus limon, Citrus medica, Ducati panigale, Pongamia pinnata, Tagetes, and Wrightia tinctoria. All of these species pairs share overlapping visual characteristics which the current feature extractors handle inadequately. Moreover, many of these classes had fewer than fifty training images each, creating a high risk of poor generalization, and forcing the classifier to rely on coarse shape features that are often shared among closely related taxa.
Taken together, these failure modes highlight the need for further data diversity and feature‐learning strategies that prioritize inter-species distinctions. Addressing these issues—through larger sample size, targeted augmentation, weight specific loss (e.g. focal loss), and active sampling of challenging examples—will be crucial for reducing misclassification rates and improving overall model reliability.
Overall, ensemble techniques displayed greater confidence, resilience, and steadiness than individual predictors. By exploiting the distinct strengths of multiple model designs, ensemble learning enables each component to offset the shortcomings of others. As a result, this methodology produces more trustworthy outputs, yielding higher accuracy and improved generalization in the intricate challenge of herb identification.
Table 10 presents a detailed comparative analysis between this study and other related works. The table details key elements such as the methodology employed, dataset size, the number of species included, and the achieved accuracy. It demonstrates that the study attains high accuracy even when applied to a large number of herb species, highlighting the method’s enhanced precision and generalization capabilities in addressing the complex task of herb recognition.
Table 10.
Comparative analysis of the ‘Herbify’ study and related works
| References | Methodology | Dataset Size | No. of Species | Accuracy (%) |
|---|---|---|---|---|
| Pushpa et al. [18] | Ayur-PlantNet (lightweight deep CNN) | 4800 | 40 | 97.27% |
| Roopashree et al. [10] | DeepHerb using Xception features | 2515 | 40 | 97.5% |
| Pushpa et al. [19] | Hierarchical ML with convolution features | 13,536 | 100 | 94.54% on GSL100 leaf dataset & 75.46% on RTL80 and RTP40 dataset |
| Rohith et al. [5] | VGG-19 and ResNet101 with attention mechanism | 5840 | 40 | 97.8% |
| Tiwari et al.[29] | Hybrid CNN-ViT model | 54,272 | 38 (in total) but used 20 | 95–96% |
| Hajam et al. [31] | Ensemble Convolutional Learning with Fine-Tuning | 1835 | 30 | 99.12% |
| Herbify model (our) | Fast ensemble model combining CNNs and ViT | 6104 | 91 | 99.56% |
Herbify application
Herbify is a web-based mobile application that offers an easily accessible interface for herb identification. Its client side is built with HyperText Markup Language (HTML), Cascading Style Sheets (CSS), and JavaScript, while inference routines operate on a Flask server. Drawing on the methodological framework, the application delegates image preprocessing and model inference to cloud services, enabling seamless operation on both mobile and desktop devices. Figure 9 outlines the system’s primary elements.
Fig. 9.

Herbify: A web-based application for automated herb recognition
The home screen as shown in Fig. 9 (A) presents a clear, visually engaging layout. Core recognition functionality is shown in Fig. 9 (B), where users can submit an herb photograph as shown in Fig. 9 (B-i). After submission, the Preprocessing Algorithm for Herb Detection (PAHD) refines the image, and an ensemble deep-learning model determines the species. The selected ensemble was optimized to balance predictive precision with computational cost; specifically, the study employs the EfficientL-ViTL architecture, which integrates EfficientNet v2-Large (CNN) with ViT-Large/16 (ViT). This configuration achieves notable performance by simultaneously minimizing computational load, maximizing inference speed, and maintaining high classification accuracy. End-to-end, the cloud workflow—from image upload to prediction—takes approximately three-four seconds, with classification accounting for about 2–3.5 s.
The processed image and the top-5 classification results are presented in the interface, as shown in Fig. 9 (B-ii) and Fig. 9 (B-iii), respectively. The results include the probabilities, scientific names, and common names of the identified herbs. Additionally, a bar plot visualizing the top-5 identified species and their confidence levels is displayed, providing users with a clear representation of the model’s predictions.
To evaluate real-world performance, the application was tested with images that were deliberately selected to deviate from the Herbify dataset. These test images included varying conditions such as diverse lighting, complex backgrounds, and occluded or partially damaged leaves. Despite these challenges, the application achieved a top-1 accuracy of 90.2% across all test cases. When predictions were filtered to include only those with a confidence score above 85%, the accuracy stood at 84%. Notably, the top-5 accuracy increased to 95.5%, indicating the robustness of the ensembled model and the PAHD algorithm in handling real-world variability.
Overall,"Herbify"successfully leverages advanced ensembled deep learning models for herb recognition. The platform delivers fast inference times, high accuracy, and a user-friendly experience, making it a comprehensive and accessible tool for a wide range of users. Whether for agricultural experts, botanists, hobbyists, or researchers, the application offers a seamless way to obtain detailed information about herbs simply by uploading an image.
Conclusion
This study presents a resilient AI-driven framework that merges cutting-edge computer vision with advanced preprocessing and transfer learning methodologies to classify herb species from images with high precision and consistency. By delivering an accessible and reliable solution for herbal therapeutics, the approach meets the increasing demand for natural remedies while alleviating the side effects and economic burdens commonly associated with synthetic pharmaceutical treatments.
In pursuit of these objectives, the research was conducted in two distinct yet complementary phases. In the first phase, a novel and rigorously standardized dataset, Herbify, was developed through the systematic cleaning, merging, preprocessing, and refinement of existing herb datasets—DIMPSAR and DeepHerb. Central to this process was the Preprocessing Algorithm for Herb Detection (PAHD), which ensured data consistency and quality, resulting in a comprehensive dataset consisting of 6104 meticulously curated images representing 91 unique herb species.
The second phase centered on the deployment of state-of-the-art deep learning architectures, including convolutional neural networks and vision transformers. Extensive experimental analysis demonstrated that the EfficientNet v2-Large model significantly outperformed conventional approaches, achieving remarkable accuracy across different datasets (98.16% for DIMPSAR, 100% for DeepHerb, and 99.12% for the unified Herbify dataset). Furthermore, the proposed ensemble model, EfficientL-ViTL, unifying EfficientNet v2-Large and ViT-Large/16, pushed performance boundaries further, achieving an exceptional accuracy of 99.56%. To enhance practical applicability, this powerful ensemble model was integrated into the Herbify web application, providing an accessible, reliable, and user-friendly tool for herb identification.
The contributions of this study are multifaceted. First, the establishment of the Herbify dataset and PAHD preprocessing pipeline provides a standardized, high-quality resource, facilitating future research in botanical image analysis and related fields. Second, the demonstrated efficacy of combining CNNs and ViTs within an ensemble approach highlights a promising direction for boosting performance in similar fine-grained visual classification tasks. Lastly, the developed AI-powered application underscores the significant potential of integrating sophisticated AI techniques into practical, real-world tools, thereby promoting public awareness and safe usage of herbal medicines.
Limitations and future work
While the results presented in this study demonstrate state-of-the-art performance for herb identification, several limitations warrant attention. Firstly, the Herbify dataset, despite being manually cleaned, rigorously curated and standardized, remains limited in its representation of geographic diversity. Most species included originate from specific regions, potentially limiting the model’s effectiveness in identifying herbs indigenous to other geographic areas not covered by the current dataset. Furthermore, although the dataset encompasses 91 species—when compared with contemporary large‐scale vision benchmarks that routinely exceed hundreds of thousands of samples, extending this number further would significantly enhance the robustness and applicability of the system in real-world contexts, where biodiversity is substantially greater.
Another limitation pertains to the deployment feasibility of the proposed models. The EfficientL-ViTL ensemble, while achieving superior accuracy, involves computationally intensive architectures (EfficientNet v2-Large and ViT-Large/16). Though we perform cloud-based computation for our application, such resource-intensive models might not perform optimally when requiring on-device prediction for devices with limited computational capabilities, such as low-power edge devices, thus restricting accessibility and usability in resource-constrained environments or mobile settings.
In terms of interpretability, the current AI framework primarily emphasizes prediction accuracy over explainability. Consequently, the black-box nature of deep learning models could limit user trust, particularly in critical area such as herbal safety, where confidence, interpretability, and transparency in decision-making processes are crucial.
Looking ahead, several promising avenues for future research and development can help overcome these limitations and further enhance the proposed framework. Expanding the Herbify dataset by incorporating additional herb classes from diverse geographic regions will significantly increase global relevance and performance. Such augmentation would facilitate more inclusive and robust identification capabilities, addressing geographical bias and better serving a global user base.
Future studies could also prioritize integrating Explainable AI (XAI) methods into the model architecture, such as attention visualization and saliency maps, to enhance transparency in model predictions. Increased model interpretability would build user trust, facilitate regulatory approval processes, and promote safer deployment, particularly in sensitive healthcare and agricultural applications.
Additionally, research into semi-supervised, self-supervised, and domain adaptation techniques could minimize reliance on extensive manual annotation, thereby reducing the dataset curation costs and effort. Investigating better but lighter and computationally efficient CNN and ViT architectures would further improve deployment potential across resource-limited platforms, enabling widespread adoption on mobile and embedded systems.
Finally, exploring the framework’s applicability to real-time scenarios and integration into broader applications, such as precision agriculture, botanical conservation initiatives, and herbal healthcare systems, represents an important trajectory for future research. Establishing real-time identification systems with rapid inference capabilities could significantly enhance practical usability and impact, ultimately advancing herbal medicine practices and botanical studies at a global scale.
By addressing these limitations and pursuing the outlined research directions, the presented AI-based herb identification framework has the potential to evolve into a robust, comprehensive, and globally applicable ecosystem, significantly contributing to both academic research and practical applications across numerous domains.
Data and code availability
The Herbify dataset—produced by cleaning, harmonizing and integrating two complementary sub-datasets—is hosted at Herbify-Dataset repository (https://github.com/Phantom-fs/Herbify-Dataset). This repository provides the processed data, full metadata descriptions, and step-by-step instructions for data loading and preprocessing to ensure full reproducibility.
The Herbify-Modules repository (https://github.com/Phantom-fs/Herbify-Modules) houses the essential modules for data preprocessing, augmentation, model training, model ensembling, and performance evaluation. Inside ready-to-run Jupyter notebooks, environment specifications, and pipelines are available.
A fully featured web interface for interactive herb identification—complete with front-end code, back-end API endpoints and deployment scripts—is available at Herbify repository (https://github.com/Phantom-fs/Herbify).
Every repository includes comprehensive documentation covering installation, setup, usage examples, and customization guidelines.
Supplementary Information
Acknowledgements
Not applicable.
Author contributions
Farhan Sheth: Conceptualization, Data Curation, Data Analysis, Implementation, Model Development, Formal Analysis, Methodology, Writing—Original Draft, Visualization; Ishika Chatter: Data Curation, Data Preprocessing, Writing—Review and Editing, Resources.; Manvendra Jasra: Investigation, Data Curation.; Gireesh Kumar: Methodology, Project Supervisor, Writing—Review and Editing.; Richa Sharma: Project Supervisor, Review, Writing—Review and Editing. All authors read and approved the manuscript.
Funding
Open access funding provided by Manipal University Jaipur. The work didn’t receive any funding.
Data availability
The Herbify dataset is hosted at Herbify-Dataset repository (https://github.com/Phantom-fs/Herbify-Dataset).
Declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Wikipedia contributors. Herb. In Wikipedia, the Free Encyclopedia. 2024. Retrieved 20:30, January 15, 2025, from https://en.wikipedia.org/w/index.php?title=Herb&oldid=1259464423
- 2.Bhattacharjee B, Sandhanam K, Ghose S, Barman D, Sahu RK. Market overview of herbal medicines for lifestyle diseases. In Role of Herbal Medicines: management of Lifestyle Diseases. Singapore: Springer Nature Singapore. 2024. pp. 597–614.
- 3.Parvin S, Reza A, Das S, Miah MMU, Karim S. Potential role and international trade of medicinal and aromatic plants in the world. Eur J Agricult Food Sci. 2023;5(5):89–99. [Google Scholar]
- 4.Ekor M. The growing use of herbal medicines: issues relating to adverse reactions and challenges in monitoring safety. Front Pharmacol. 2014;4:177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Rohith ND, Bhuvaneswari R. Medicinal plant classification using attention mechanism driven VGG-19 and Resnet101. In 2023 Global Conference on Information Technologies and Communications (GCITC). IEEE. 2023 pp. 1–7.
- 6.Ahmed SN, Ahmad M, Zafar M, Yaseen G, Iqbal N, Rashid N, Haroon A. Herbal drugs: safety, cost-effectiveness, regulation, current trends, and future directions. In Bioprospecting of Tropical Medicinal Plants. Cham: Springer Nature Switzerland. 2023. pp. 1479–1493.
- 7.Gantait S, Majumder J, Sharangi AB (Eds.). Biotechnology of Medicinal Plants with Antiallergy Properties: research Trends and Prospects. 2024.
- 8.Sohail R, Mathew M, Patel KK, Reddy SA, Haider Z, Naria M, Razzaq W. Effects of non-steroidal anti-inflammatory drugs (NSAIDs) and gastroprotective NSAIDs on the gastrointestinal tract: a narrative review. Cureus. 2023;15(4):1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Pushpa BR, Rani NS. DIMPSAR: dataset for Indian medicinal plant species analysis and recognition. Data Brief. 2023;49: 109388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Roopashree S, Anitha J. DeepHerb: a vision-based system for medicinal plants using xception features. Ieee Access. 2021;9:135927–41. [Google Scholar]
- 11.Deol S, Shukla V, Rastogi S, Kumar N. Integration of traditional knowledge and modern science: a holistic approach to identify medicinal leaves for curing diseases. In 2024 2nd International Conference on Disruptive Technologies (ICDT). IEEE. 2024. pp. 636–641.
- 12.Sharifi-Rad J, Rayess YE, Rizk AA, Sadaka C, Zgheib R, Zam W, Martins N. Turmeric and its major compound curcumin on health: bioactive effects and safety profiles for food, pharmaceutical, biotechnological and medicinal applications. Front Pharmacol. 2020;11: 550909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Flynn J. 20 vital smartphone usage statistics [2022]: facts, data, and trends on mobile use in the us. 2022.
- 14.Ali MAM, Aly T, Raslan AT, Gheith M, Amin EA. Advancing crowd object detection: a review of YOLO, CNN and ViTs hybrid approach. J Intell Learn Syst Appl. 2024;16(3):175–221. [Google Scholar]
- 15.Li J, Tan YA, Yang J, Li Z, Ye H, Xia C, Li Y. Unsupervised Adversarial Example Detection of Vision Transformers for Trustworthy Edge Computing. ACM Transactions on Multimedia Computing, Communications and Applications. 2024.
- 16.Prasathkumar M, Anisha S, Dhrisya C, Becky R, Sadhasivam S. Therapeutic and pharmacological efficacy of selective Indian medicinal plants—a review. Phytomed plus. 2021;1(2): 100029. [Google Scholar]
- 17.Pushpa BR, Rani S. Indian Medicinal Leaves Image Datasets. Mendeley Data, V3. 2023. 10.17632/748f8jkphb.3
- 18.Pushpa BR, Rani NS. Ayur-PlantNet: an unbiased light weight deep convolutional neural network for Indian Ayurvedic plant species classification. J Appl Res Med Aromat Plants. 2023;34: 100459. [Google Scholar]
- 19.Pushpa BR, Rani NS, Chandrajith M, Manohar N, Nair SSK. On the importance of integrating convolution features for Indian medicinal plant species classification using hierarchical machine learning approach. Eco Inform. 2024;81: 102611. [Google Scholar]
- 20.Lee CP, Lim KM, Song YX, Alqahtani A. Plant-CNN-ViT: plant classification with ensemble of convolutional neural networks and vision transformer. Plants. 2023;12(14):2642. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kaleybar JM, Saadat H, Khaloo H. Capturing local and global features in medical images by using ensemble CNN-Transformer. In 2023 13th International Conference on Computer and Knowledge Engineering (ICCKE). 2023. pp. 030–035.
- 22.Khan SUR, Asim MN, Vollmer S, Dengel A. AI-driven diabetic retinopathy diagnosis enhancement through image processing and Salp Swarm Algorithm-optimized ensemble network. 2025. arXiv https://arxiv.org/abs/2503.14209
- 23.Hekmat A, Zuping Z, Bilal O, Ullah I, Jalil A, Rehman A, Saba T. Differential evolution-driven optimized ensemble network for brain tumor detection. Int J Mach Learn Cybern. 2025. 10.1007/s13042-025-02629-6. [Google Scholar]
- 24.Nanni L, Loreggia A, Barcellona L, Ghidoni S. Building ensemble of deep networks: convolutional networks and transformers. IEEE Access. 2023;11:124962–74. [Google Scholar]
- 25.Abulfaraj AW, Binzagr F. A deep ensemble learning approach based on a vision transformer and neural network for multi-label image classification. Big Data Cognit Comput. 2025;9(2):39. [Google Scholar]
- 26.Amin SU, Hussain A, Kim B, Seo S. Deep learning based active learning technique for data annotation and improve the overall performance of classification models. Expert Syst Appl. 2023;228: 120391. [Google Scholar]
- 27.Amin SU, Ullah M, Sajjad M, Cheikh FA, Hijji M, Hijji A, Muhammad K. EADN: an efficient deep learning model for anomaly detection in videos. Mathematics. 2022;10(9):1555. [Google Scholar]
- 28.Amin SU, Kim Y, Sami I, Park S, Seo S. An efficient attention-based strategy for anomaly detection in surveillance video. Comput Syst Sci Eng. 2023;46(3):3939–58. [Google Scholar]
- 29.Tiwari RG, Maheshwari H, Gautam V, Agarwal AK, Trivedi NK, Jain V. MedLeafNet: a feature fusion network integrating transformer and CNN Modules for Indian Medical Leaf Classification. In 2024 2nd International Conference on Intelligent Data Communication Technologies and Internet of Things (IDCIoT). IEEE. 2024. pp. 999–1005.
- 30.Khan SUR, Asif S, Bilal O. Ensemble architecture of Vision Transformer and CNNs for breast cancer tumor detection from mammograms. Int J Imaging Syst Technol. 2025. 10.1002/ima.70090. [Google Scholar]
- 31.Hajam MA, Arif T, Khanday AMUD, Neshat M. An effective ensemble convolutional learning model with fine-tuning for medicinal plant leaf identification. Information. 2023;14(11):618. [Google Scholar]
- 32.Kunjachan S, Santhosh AM, Kala S. A Novel CNN Architecture for Herb Classification. In TENCON 2024–2024 IEEE Region 10 Conference (TENCON). IEEE. 2024. pp. 1385–1388.
- 33.Liu Z, Luo C, Fu D, Gui J, Zheng Z, Qi L, Guo H. A novel transfer learning model for traditional herbal medicine prescription generation from unstructured resources and knowledge. Artif Intell Med. 2022;124: 102232. [DOI] [PubMed] [Google Scholar]
- 34.Mujahid PE, Manik R, Simbolon JS, Sinaga MRRS, Aisyah S, Nababan M, Harmaja OJ. Herbal Leaf Image Classification Using Convolutional Neural Network (CNN). J Sistem Inform Dan Ilmu Komputer. 2024;8(1):52–68. [Google Scholar]
- 35.Sheth F, Mathur P, Gupta AK, Chaurasia S. An advanced artificial intelligence framework integrating ensembled convolutional neural networks and Vision Transformers for precise soil classification with adaptive fuzzy logic-based crop recommendations. Eng Appl Artif Intell. 2025;158: 111425. [Google Scholar]
- 36.Tapsell LC, Hemphill I, Cobiac L, Sullivan DR, Fenech M, Patch CS, Inge KE. Health benefits of herbs and spices: the past, the present, the future. 2006. [DOI] [PubMed]
- 37.RoopashreeS, Anitha J. Medicinal Leaf Dataset. Mendeley Data, V1. 2020. 10.17632/nnytj2v3n5.1
- 38.Bhattacharyya S. A brief survey of color image preprocessing and segmentation techniques. J Patt Recogn Res. 2011;1(1):120–9. [Google Scholar]
- 39.Cheng HD, Jiang XH, Sun Y, Wang J. Color image segmentation: advances and prospects. Pattern Recogn. 2001;34(12):2259–81. [Google Scholar]
- 40.Ajmal A, Hollitt C, Frean M, Al-Sahaf H. A comparison of RGB and HSV colour spaces for visual attention models. In 2018 International conference on image and vision computing New Zealand (IVCNZ). IEEE. 2018. pp. 1–6.
- 41.Anishiya P, Joans SM. Number plate recognition for Indian cars using morphological dilation and erosion with the aid of ocrs. In International Conference on Information and Network Technology vol. 4. 2011.
- 42.Haralick RM, Sternberg SR, Zhuang X. Image analysis using mathematical morphology. IEEE Trans Pattern Anal Mach Intell. 1987;1987(4):532–50. [DOI] [PubMed] [Google Scholar]
- 43.Zhong XP, Chen HJ, Li MQ, Zeng WW. Understanding engineering drawing images from mobile devices. J Inform Sci Eng. 2021;37(1):1. [Google Scholar]
- 44.Catanzaro B, Su BY, Sundaram N, Lee Y, Murphy M, Keutzer K. Efficient, high-quality image contour detection. In 2009 IEEE 12th International Conference on Computer Vision. IEEE. 2009. pp. 2381–2388.
- 45.Federer H. The gauss-green theorem. Trans Am Math Soc. 1945;58(1):44–76. [Google Scholar]
- 46.Nykamp DQ. Using Green’s theorem to find area. From Math Insight. http://mathinsight.org/greens_theorem_find_area
- 47.Derraz F, Beladgham M, Khelif MH. Application of active contour models in medical image segmentation. In International Conference on Information Technology: Coding and Computing, 2004. Proceedings. ITCC 2004. (Vol. 2). IEEE. 2004. pp. 675–681.
- 48.Rudin S, Bednarek DR, Yang CYJ. Real-time equalization of region-of-interest fluoroscopic images using binary masks. Med Phys. 1999;26(7):1359–64. [DOI] [PubMed] [Google Scholar]
- 49.Sunil GC, Koparan C, Ahmed MR, Zhang Y, Howatt K, Sun X. A study on deep learning algorithm performance on weed and crop species identification under different image background. Artif Intell Agricult. 2022;6:242–56. [Google Scholar]
- 50.Van Der Ploeg T, Austin PC, Steyerberg EW. Modern modelling techniques are data hungry: a simulation study for predicting dichotomous endpoints. BMC Med Res Methodol. 2014;14:1–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Mikołajczyk A, Grochowski M. Data augmentation for improving deep learning in image classification problem. In 2018 international interdisciplinary PhD workshop (IIPhDW). IEEE. 2018. pp. 117–122.
- 52.Shorten C, Khoshgoftaar TM. A survey on image data augmentation for deep learning. J Big Data. 2019;6(1):1–48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Buslaev A, Iglovikov VI, Khvedchenya E, Parinov A, Druzhinin M, Kalinin AA. Albumentations: fast and flexible image augmentations. Information. 2020;11(2):125. [Google Scholar]
- 54.Kiflie MA, Sharma DP, Haile MA. Deep learning for Ethiopian indigenous medicinal plant species identification and classification. J Ayurv Integr Med. 2024;15(6): 100987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Saber A, Sakr M, Abo-Seida OM, Keshk A, Chen H. A novel deep-learning model for automatic detection and classification of breast cancer using the transfer-learning technique. IEEE Access. 2021;9:71194–209. [Google Scholar]
- 56.Yamashita R, Nishio M, Do RKG, Togashi K. Convolutional neural networks: an overview and application in radiology. Insights Imaging. 2018;9:611–29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Aghdam HH, Heravi EJ. Guide to convolutional neural networks. New York, NY: Springer. 2017;10(978–973):51. [Google Scholar]
- 58.O’Shea K. An introduction to convolutional neural networks. 2015. arXiv preprint arXiv:1511.08458.
- 59.Cong S, Zhou Y. A review of convolutional neural network architectures and their optimizations. Artif Intell Rev. 2023;56(3):1905–69. [Google Scholar]
- 60.Li Z, Liu F, Yang W, Peng S, Zhou J. A survey of convolutional neural networks: analysis, applications, and prospects. IEEE Trans Neural Netw Learn Syst. 2021;33(12):6999–7019. [DOI] [PubMed] [Google Scholar]
- 61.Howard A, Sandler M, Chu G, Chen LC, Chen B, Tan M, Adam H. Searching for mobilenetv3. In Proceedings of the IEEE/CVF international conference on computer vision. 2019. pp. 1314–1324.
- 62.Simonyan K, Zisserman A. Very deep convolutional networks for large-scale image recognition. 2014. arXiv preprint arXiv:1409.1556.
- 63.Calazans MAA, Ferreira FAB, Alcoforado MDLMG, Santos AD, Pontual ADA, Madeiro F. Automatic classification system for periapical lesions in cone-beam computed tomography. Sensors. 2022;22(17):6481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. In Proceedings of the IEEE conference on computer vision and pattern recognition. 2016. pp. 770–778.
- 65.Baaloul A, Benblidia N, Reguieg FZ, Bouakkaz M, Felouat H. An arabic visual speech recognition framework with CNN and vision transformers for lipreading. Multimed Tools Appl. 2024;83(27):69989–70023. [Google Scholar]
- 66.Tan M, Le Q. Efficientnetv2: Smaller models and faster training. In International conference on machine learning. PMLR. 2021. pp. 10096–10106.
- 67.Yu S, Ma K, Bi Q, Bian C, Ning M, He N, Zheng Y. Mil-vt: Multiple instance learning enhanced vision transformer for fundus image classification. In Medical Image Computing and Computer Assisted Intervention–MICCAI 2021: 24th International Conference, Strasbourg, France, September 27–October 1, 2021, Proceedings, Part VIII 24. Springer International Publishing. 2021. pp. 45–54.
- 68.Wu B, Xu C, Dai X, Wan A, Zhang P, Yan Z, Vajda P. Visual transformers: token-based image representation and processing for computer vision. 2020. arXiv preprint arXiv:2006.03677.
- 69.Han K, Wang Y, Chen H, Chen X, Guo J, Liu Z, Tao D. A survey on visual transformer. 2020. arXiv preprint arXiv:2012.12556.
- 70.Khan S, Naseer M, Hayat M, Zamir SW, Khan FS, Shah M. Transformers in vision: a survey. ACM Comput Surveys (CSUR). 2022;54(10s):1–41. [Google Scholar]
- 71.Lu K, Xu Y, Yang Y. Comparison of the potential between transformer and CNN in image classification. In ICMLCA 2021; 2nd International Conference on Machine Learning and Computer Application. VDE. 2021. pp. 1–6.
- 72.Descoteaux M, Maier-Hein L, Franz A, Jannin P, Collins DL, Duchesne S. (Eds.). Medical Image Computing and Computer Assisted Intervention− MICCAI 2017: 20th International Conference, Quebec City, QC, Canada, September 11–13, 2017, Proceedings, Part III (Vol. 10435). Springer. 2017.
- 73.Wolf T, Debut L, Sanh V, Chaumond J, Delangue C, Moi A, Rush AM. Transformers: State-of-the-art natural language processing. In Proceedings of the 2020 conference on empirical methods in natural language processing: system demonstrations. 2020. pp. 38–45.
- 74.Mao X, Qi G, Chen Y, Li X, Duan R, Ye S, Xue H. Towards robust vision transformer. In Proceedings of the IEEE/CVF conference on Computer Vision and Pattern Recognition. 2022. pp. 12042–12051.
- 75.Tajbakhsh N, Shin JY, Gurudu SR, Hurst RT, Kendall CB, Gotway MB, Liang J. Convolutional neural networks for medical image analysis: Full training or fine tuning? IEEE Trans Med Imaging. 2016;35(5):1299–312. [DOI] [PubMed] [Google Scholar]
- 76.Tan C, Sun F, Kong T, Zhang W, Yang C, Liu C. A survey on deep transfer learning. In Artificial Neural Networks and Machine Learning–ICANN 2018: 27th International Conference on Artificial Neural Networks, Rhodes, Greece, October 4–7, 2018, Proceedings, Part III 27. Springer International Publishing. 2018. pp. 270–279.
- 77.Sharif Razavian A, Azizpour H, Sullivan J, Carlsson S. CNN features off-the-shelf: an astounding baseline for recognition. In Proceedings of the IEEE conference on computer vision and pattern recognition workshops. 2014. pp. 806–813.
- 78.Kingma DP. Adam: a method for stochastic optimization. 2014. arXiv preprint arXiv:1412.6980.
- 79.Loshchilov I. Decoupled weight decay regularization. 2017. arXiv preprint arXiv:1711.05101.
- 80.Ahamad GN, Shafiullah, Fatima H, Imdadullah, Zakariya SM, Abbas M, Alqahtani MS, Usman M. Influence of optimal hyperparameters on the performance of machine learning algorithms for predicting heart disease. Processes. 2023;11(3):734. [Google Scholar]
- 81.De Boer PT, Kroese DP, Mannor S, Rubinstein RY. A tutorial on the cross-entropy method. Ann Oper Res. 2005;134:19–67. [Google Scholar]
- 82.Li LH, Tanone R. Ensemble Learning based on CNN and Transformer Models for Leaf Diseases Classification. In 2024 18th International Conference on Ubiquitous Information Management and Communication (IMCOM). IEEE. 2024. pp. 1–6.
- 83.Savarimuthu X, Subramani S, Raj ANJ. (Eds.). Artificial intelligence for multimedia information processing: tools and applications. CRC Press. 2024.
- 84.Jiang Z, Dong Z, Wang L, Jiang W. Method for diagnosis of acute lymphoblastic leukemia based on ViT-CNN ensemble model. Comput Intell Neurosci. 2021;2021(1):7529893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Schmidt-Hieber J. Nonparametric regression using deep neural networks with ReLU activation function. 2020. [DOI] [PubMed]
- 86.Alzubaidi L, Zhang J, Humaidi AJ, Al-Dujaili A, Duan Y, Al-Shamma O, Farhan L. Review of deep learning: concepts, CNN architectures, challenges, applications, future directions. J Big Data. 2021;8:1–74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Vasan D, Alazab M, Wassan S, Safaei B, Zheng Q. Image-Based malware classification using ensemble of CNN architectures (IMCEC). Comput Secur. 2020;92: 101748. [Google Scholar]
- 88.Liu J, Wang X. Plant diseases and pests detection based on deep learning: a review. Plant Methods. 2021;17:1–18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Kamilaris A, Prenafeta-Boldú FX. Deep learning in agriculture: a survey. Comput Electron Agric. 2018;147:70–90. [Google Scholar]
- 90.Espíndola RP, Ebecken NF. On extending f-measure and g-mean metrics to multi-class problems. WIT Trans Inform Commun Technol. 2005;35:25–34. [Google Scholar]
- 91.Dai W, Berleant D. Benchmarking contemporary deep learning hardware and frameworks: a survey of qualitative metrics. In 2019 IEEE First International Conference on Cognitive Machine Intelligence (CogMI). IEEE. 2019. pp. 148–155.
- 92.Zhou J, Gandomi AH, Chen F, Holzinger A. Evaluating the quality of machine learning explanations: a survey on methods and metrics. Electronics. 2021;10(5):593. [Google Scholar]
- 93.Montag C, Błaszkiewicz K, Sariyska R, Lachmann B, Andone I, Trendafilov B, Markowetz A. Smartphone usage in the 21st century: Who is active on WhatsApp? BMC Res Notes. 2015;8:1–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Grinberg M. Flask web development. “O’Reilly Media, Inc”. 2018.
- 95.Bradski G. Learning OpenCV: computer vision with the OpenCV library. O’REILLY Google Schola. 2008;2:334–52. [Google Scholar]
- 96.Van der Walt S, Schönberger JL, Nunez-Iglesias J, Boulogne F, Warner JD, Yager N, Yu T. scikit-image: image processing in Python. PeerJ. 2014;2: e453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Paszke A, Gross S, Massa F, Lerer A, Bradbury J, Chanan G, Chintala S. Pytorch: an imperative style, high-performance deep learning library. Adv Neural Inf Process Syst. 2019;32:1. [Google Scholar]
- 98.Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, Duchesnay É. Scikit-learn: machine learning in Python. J Mach Learn Res. 2011;12:2825–30. [Google Scholar]
- 99.Hunter JD. Matplotlib: a 2D graphics environment. Comput Sci Eng. 2007;9(03):90–5. [Google Scholar]
- 100.Waskom ML. Seaborn: statistical data visualization. J Open Source Softw. 2021;6(60):3021. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The Herbify dataset—produced by cleaning, harmonizing and integrating two complementary sub-datasets—is hosted at Herbify-Dataset repository (https://github.com/Phantom-fs/Herbify-Dataset). This repository provides the processed data, full metadata descriptions, and step-by-step instructions for data loading and preprocessing to ensure full reproducibility.
The Herbify-Modules repository (https://github.com/Phantom-fs/Herbify-Modules) houses the essential modules for data preprocessing, augmentation, model training, model ensembling, and performance evaluation. Inside ready-to-run Jupyter notebooks, environment specifications, and pipelines are available.
A fully featured web interface for interactive herb identification—complete with front-end code, back-end API endpoints and deployment scripts—is available at Herbify repository (https://github.com/Phantom-fs/Herbify).
Every repository includes comprehensive documentation covering installation, setup, usage examples, and customization guidelines.
The Herbify dataset is hosted at Herbify-Dataset repository (https://github.com/Phantom-fs/Herbify-Dataset).