Abstract
Food production systems may act as transmission routes for antimicrobial-resistant (AMR) bacteria and AMR genes (AMRGs) to humans. However, the food resistome remains poorly characterized. Here 1,780 raw-material (milk, brine, fresh meat and so on), end-product (cheese, fish, meat products and vegetables) and surface (processing, cooling, smoking, ripening and packing rooms) samples from 113 food processing facilities were subjected to whole-metagenome sequencing. Assembly-free analyses demonstrated that >70% of all known AMRGs, including many predicted to confer resistance to critically important antibiotics, circulate throughout food production chains, with those conferring resistance to tetracyclines, β-lactams, aminoglycosides and macrolides being the most abundant overall. An assembly-based analysis highlighted that bacteria from the ESKAPEE group, together with Staphylococcus equorum and Acinetobacter johnsonii, were the main AMRG carriers. Further evaluation demonstrated that ~40% of the AMRGs were associated with mobile genetic elements, mainly plasmids. These findings will help guide the appropriate use of biocides and other antimicrobials in food production settings when designing efficient antimicrobial stewardship policies.
Subject terms: Microbiome, Antimicrobial resistance
Whole-metagenome sequencing of 1,780 raw-material, end-product and surface samples from 113 food processing facilities reveals the occurrence of antimicrobial resistance determinants in foods and their processing environments.
Main
The rapid emergence and spread of antimicrobial (AM) resistance are among the most urgent threats to health, as many infections are becoming refractory to AM treatment1. It has been estimated that 4.95 million deaths were caused in 2019 by AM-resistant (AMR) micro-organisms2, which places AMR as the third greatest cause of death globally3, and it is estimated that these figures will increase if a global and coordinated action plan is not implemented4,5.
The misuse of AMs, especially in clinical settings and by the society, but also in agriculture, farming and other agrifood-associated settings, has been identified as one of the major drivers of AMR increase6. In fact, an important share of the AM sold globally is used for farming7. However, despite the predicted high impact that the microbial content of foods may have on the human microbiome and health8, the diversity of resistant bacteria and AMR genes (AMRGs) in food production systems remains poorly characterized, with the available studies focusing exclusively on certain pathogen–drug combinations9,10.
Some micro-organisms are able to survive and persist in food processing environments, despite the cleaning and disinfection practices that take place routinely11, and can serve as natural inoculants for foods12. While many environmental micro-organisms thriving in food processing plants play beneficial roles in food production, others can act as food spoilers and/or can pose a threat to human health, including through the transmission of AMRGs to the human gut microbiome13,14. However, the diversity of AMRGs circulating through processing environments from different food production sectors, and the impact on the resistome of end food products, is yet to be unravelled.
The possibilities offered by metagenomics for the in-depth characterization of microbial communities and their genomic content has created new opportunities to decipher resistomes and ascertain their risks, including their potential for mobilization15. The location of AMR determinants within mobile genetic elements (MGEs) represents a major concern, as MGE-mediated horizontal transfer facilitates AMRG spread16. This is especially true for plasmids, which have facilitated a tenfold increase in the carriage of AMR determinants from 2000 to 202017.
The hypothesis of this study is that a very diverse and sector-specific resistome circulates daily through food production settings and that the resistome of end food products is markedly shaped by production environments and the ecologic pressures imposed by processing. To test this hypothesis, 1,780 raw-material, end-product and surface samples from 113 food processing facilities were subjected to whole-metagenome sequencing by the MASTER EU Consortium (https://www.master-h2020.eu/), as summarized in Fig. 1. The analyses presented here provide an in-depth characterization of the occurrence of AMR determinants in foods and their processing environments, highlighting potential hotspots for their transfer and disclosing information on their carrier taxa and association with MGEs.
Fig. 1. Sampling strategy and raw output of the metagenomic analysis.
On the left side, countries of the visited industries are reported. For each industry type, the general process and sampling strategy is represented. The values indicate the total number of samples collected from raw materials, FCSs and NFCSs, and end food products. Also, for each industry type, pie and bar charts represent the proportion of contigs and the raw number of MAGs reconstructed from each sample type (that is, raw materials, final products, intermediate products and industry surfaces). Created with BioRender.com.
Results
Production surfaces show the highest AMRG load and diversity
Overall, 2.7 million sequences (0.006% of the total number of sequences) aligned to 528 different AMRG clusters, which corresponds to 72.8% of all known AMRG clusters in the ResFinder database. The occurrence differed widely among AMRG classes, from 55.6% for glycopeptides to 87.9% for amphenicols (Supplementary Table 1). Remarkably, only 20 AMRG clusters were considered as prevalent (that is, >0.1 counts per million reads (CPM) in >10% samples), representing 79.4% of all AMRG reads (Extended Data Fig. 1). The occurrence, diversity and richness of AMRGs were significantly higher in food contact surfaces (FCSs) and non-food contact surfaces (NFCSs) than in raw materials and/or end food products, with end products showing a higher AMRG load, in CPM, than raw materials (Fig. 2a). Regardless of sample type, AMRG loads and richness were significantly higher in meat production than in other production sectors (Extended Data Fig. 2a).
Extended Data Fig. 1. Main prevalent AMRG.
Mean values of prevalent AMRG ( > 0.1 CPM in >10% samples) found a) by sample type and industry type and b) percentage of prevalent AMRG found globally.
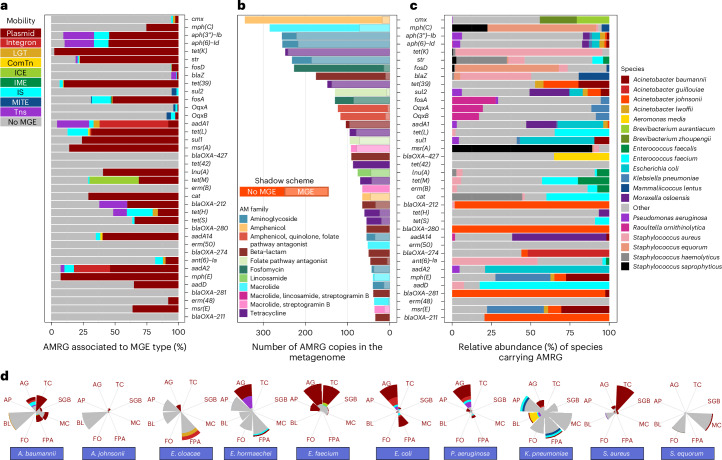
Fig. 2. Quantitative overview of AMRG occurrence in foods and food processing environments.
a, Total AMRG counts, richness index and Simpson’s evenness index for AMRGs. n values were 294, 662, 489 and 335 for raw materials, final products, FCSs and NFCSs, respectively. The lower, middle and upper hinges in the box plots correspond to the first, second and third quartiles (the 25th, 50th or median and 75th percentiles), respectively. The upper whisker extends from the hinge to the largest value no further than 1.5× IQR from the hinge (where IQR is the interquartile range, or distance between the first and third quartiles). The lower whisker extends from the hinge to the smallest value at most 1.5× IQR of the hinge. Only significative P values (<0.05) are indicated according to the Wilcoxon test. b, Principal coordinates analysis for AMRG counts normalized to CPM. PC, principal coordinate. c, The relative abundance of AMRGs associated with the ten most abundant antibiotic families detected. For those genes conferring resistance to more than one antibiotic family, each antibiotic family was counted separately. AMRGs not included within the former ten most abundant families are represented as ‘Other antibiotic families’. d, Average abundance of the main AMRG on each industry type and sample type, indicated by column and point colours, respectively, except for the first column, which shows a global comparative analysis among industry types. The coloured box next to each AMRG name corresponds to the antibiotic family the gene confers resistance to (OqxB is associated with resistance against three antibiotic families), according to the legend in the bottom left part of the figure. AMRG groups obtained at 90% identity clustering by CD-HIT were used for the analysis shown in this figure. n values for b–d are as follows: Meat – Raw material, 47; Final product, 38; Food contact, 67; Non food contact, 68. Cheese-dairy – Raw material, 229; Final product, 605; Food contact, 390; Non food contact, 240. Processsed fish – Raw material, 12; Final product, 13; Food contact, 20; Non food contact, 22. Vegetables – Raw material, 6; Final product, 6; Food contact, 12; Non food contact, 5.
Extended Data Fig. 2. Quantitative overview of AMRG occurrence in foods and food processing environments.
a) Total AMRG counts, richness index and Simpson’s evenness index for AMRG. b) Principal Coordinates Analysis for AMRG counts normalised to CPM.
In a principal coordinates analysis based on between-sample dissimilarity (beta-diversity) in relative abundance of AMRG clusters according to the sample type, all comparisons (for meat, dairy, fish and vegetable production sectors) resulted in statistically significant differences (adonis P value of 0.001) (Fig. 2b). Significant differences in beta-diversity were also observed among industry categories for all sample types (Extended Data Fig. 2b).
AMRGs associated with resistance to tetracyclines, β-lactams and aminoglycosides dominated overall, although the patterns differed widely among industry and sample types (Fig. 2c). Tetracyclines AMRGs were particularly dominant in the meat industry and mainly in FCS (Fig. 2c and Extended Data Fig. 3). β-Lactam and aminoglycoside AMRGs were also abundant on meat processing surfaces, but not on food products or raw materials. In dairies, surfaces also showed the highest load of AMRGs, with β-lactam and aminoglycoside AMRGs being the most predominant, followed by tetracycline AMRGs, which were instead most prevalent in raw materials and final products (Fig. 2c). The lowest AMRG loads were found in fisheries, with near absence of AMRGs in raw materials and final products and with β-lactam and aminoglycoside AMRGs being dominant on surfaces (Fig. 2c). Finally, raw materials and FCSs contained the highest AMRG loads for vegetable factories, with notable variations in the resistome composition among sample types (β-lactam AMRGs were the most abundant on FCSs, while amphenicol, streptogramin and lincosamide AMRGs dominated in raw materials) (Fig. 2c). Interestingly, AMRGs from some AM families showed a higher occurrence in surfaces and final products (for example, macrolide AMRGs for all sectors; β-lactam AMRGs in facilities processing vegetables; fosfomycin AMRGs in meat production) and were almost absent in raw materials, suggesting that processing environments and not the input materials may act as the source of these AMRGs that contaminate the final products.
Extended Data Fig. 3. Quantitative overview of CIA AMRG occurrence in foods and food processing environments.
a) Global CIA AMRG abundance as CPM. b) Differences in CIA AMRG counts among sample types from the same industry type (top boxplot) and among industry types for the same sample type (bottom boxplot). c) Relative abundance of the 20 most abundant CIA AMRG.
The most abundant AMRGs (>0.5 mean CPM per sample) found include various AMRGs of resistance to tetracyclines (tet(S) and tet(M), mainly in dairy; tet(L), tet(K) and tet(39), mainly in meat) and aminoglycosides (str, aph(6)-Id and aph(3″)-Ib; mainly in dairy), together with some individual genes associated with resistance to either folate pathway antagonists, macrolides, fosfomycin, amphenicols or beta-lactams (for example, sul2, fosD, mph(C), oqxB, cmx), and were in most cases more abundant on food processing surfaces than on raw materials and end products (Fig. 2d and Extended Data Fig. 1a).
A total of 665 different AMRGs, grouped in 121 gene clusters, associated with resistance to critically important antibiotics (CIAs), were detected in our study, which correspond to 69.5% and 72%, respectively, of all those included in the Resfinder database (Supplementary Table 1). Only mph(C) and msr(A) genes were within the most prevalent ARGs (Extended Data Fig. 1). Fish industries presented significantly lower CPM values for CIA-AMRGs than other industries (Extended Data Fig. 4a). Moreover, an increasing trend of CIA-AMRG abundance from raw materials to processing surfaces was observed in meat and dairy factories, with final products showing intermediate CPM values (Extended Data Fig. 4b). Among the CIA-AMRGs, mph(C), msr(E), msr(A), erm(B), mecB, mecD and mph(E) were the most abundant (>0.1 mean CPM per sample) and showed a differential distribution among sample and industry types (Extended Data Fig. 4c).
Extended Data Fig. 4. Antibiotic families distribution along different rooms.
For both FC and NFC samples in each industry type studied.
Diverse non-ESKAPEE taxa contribute to the food resistome
An assembly-based analysis rendered 8,584 contigs harbouring AMRGs from 1,152 samples. Among these, 5,446 contigs (63.4%) assigned to the species level were considered for downstream analysis. They were associated with 618 different species from 215 genera and were classified into 481 unique AMRGs (representing 19% of all AMRGs in Resfinder), associated with resistance against 18 different AM classes (Supplementary Table 1). β-Lactam AMRGs were those most frequently recovered (24.2% of all AMRGs), followed by aminoglycoside (20.8%), tetracycline (15.6%), amphenicol (12.5%) and macrolide (11.6%) AMRGs.
Staphylococcus, particularly the human-associated opportunistic pathogen Staphylococcus aureus, showed the greatest number of AMRG-carrying contigs (22.7% and 8.3%, respectively, of all AMRG-carrying contigs). This species belongs to the clinically relevant ESKAPEE group, which overall accounted for 18.9% of all AMRG-carrying contigs (Fig. 3 and Supplementary Table 2). Other ESKAPEE members, including Acinetobacter baumannii (2.3%), Enterococcus faecium (2.3%), Escherichia coli (2.3%), Klebsiella pneumoniae (1.4%) and Pseudomonas aeruginosa (1.1%), were also among the 20 main AMRG carriers. However, this indicates that the greatest contribution to the food resistome comes from non-ESKAPEE food-associated or commensal bacteria such as Staphylococcus equorum (6.6%), Acinetobacter johnsonii (5.9%), Staphylococcus haemolyticus (2.1%), Staphylococcus saprophyticus (1.7%), Acinetobacter lwoffii (1.7%), Raoultella ornithinolytica (1.5%) and Brevibacterium zhoupengii (1.5%) (Fig. 3 and Supplementary Table 2).
Fig. 3. The most relevant AMRG-carrier bacterial species in the different food production sectors, with indication of their resistome profile.
Heat map values were calculated by dividing the number of AMRG-encoding contigs (from each sample and industry type) assigned to the species indicated in the row by the total number of AMRG-encoding contigs (for the sample and industry type indicated in each column), to reduce overestimation in cheese and dairy industries due to unbalanced sampling. The left bar plot indicates the percentage of contigs assigned to each species on total number of AMRG-encoding contigs, while the right bar plot indicates the AMRG family distribution within each species, by percentage of total contigs. The most abundant species plus Enterobacter species, belonging to the ESKAPEE group, were plotted. Members of the ESKAPEE group are reported in red.
Some AMRGs were almost exclusively associated with certain taxa, such as tetK, blaZ and ant(6)-Ia with S. aureus, or blaOXA-211, blaOXA-212, blaOXA-280 and blaOXA-281 with A. johnsonii. Moreover, some micro-organisms with recognized positive effects in food production contributed with particular AMRGs, such as S. equorum carrying mph(C) and fosD, or Brevibacterium aurantiacum and B. zhoupengii carrying cmx, the most abundant AMRG in the study.
The distribution of AMRG-carrying species varied widely among production sectors and sample types (Fig. 3). Thus, while a wide range of species were identified as AMRG carriers in samples from dairy and meat production, the resistome of fish factories was mainly associated with A. johnsonii and that of vegetable facilities with A. lwoffii and A. johnsonii. Remarkably, considering the profile of AMRG-carrying taxa, in meat and dairy industries, processing surfaces and end products clustered together and separate from raw materials. This is explained by the relevance as carrier taxa exclusively in end products and production surfaces of S. aureus, S. equorum, E. faecium, E. coli and K. pneumoniae in meat production and S. aureus, S. equorum, S. saprophyticus and B. zhoupengii in dairy production.
The food resistome is highly linked to MGEs
The most abundant AMRGs identified by the assembly-based approach (those representing >2% of the total AMRGs), and their association with MGEs, are shown in Fig. 4. Overall, 31.7% of the AMRGs were identified to be plasmid associated, with 23% of them also presenting a plasmidic replicon (Supplementary Fig. 1). Other MGEs identified as harbouring AMRGs included unit transposons (Tns, 4.9%), insertion sequences (ISs, 4.0%), integrons (2.9%), miniature inverted repeats (MITEs, 0.7%), integrative conjugative elements (ICEs, 0.6%), composite transposons (ComTns, 0.2%) and integrative mobilizable elements (IMEs, 0.03%). Furthermore, WAAFLE18 predicted the mobilization via potential lateral gene transfer (LGT) of 37 AMRGs (0.6%). Notably, when considering all these MGE types, 37.8% of all AMRGs were considered to be mobile owing to their association with a MGE (Supplementary Fig. 1a). The proportion of AMRGs contained in MGEs varied greatly among companies, rooms, surfaces and products, with meat and cheese final products and raw materials harbouring in general a higher ratio of AMRGs associated with MGEs (49.1% and 40.9% in meat and cheese products, respectively; Extended Data Fig. 5). It also differed widely among AM classes (Fig. 4a and Supplementary Fig. 1a) and ranged from 5.1% and 7.5% of AMRGs occurring in MGEs for fosfomycin and β-lactam resistance, respectively, to 65.8% and 71.0% for tetracycline and aminoglycoside resistance, respectively. Aminoglycoside resistance was mostly associated with plasmids (56.8%), Tns (17.3%), integrons (11.6%) and ISs (9.6%); tetracycline AMRGs were very frequently associated with plasmids (62.4%), ISs (4.0%) and ICEs (3.8%); and mobile β-lactam AMRGs were mainly associated with Tns (2.8%), MITEs (2.0%) and plasmids (1.6%) (Supplementary Fig. 1a).
Fig. 4. The most abundant AMRGs occurring in foods and food processing environments, with indication of their distribution across different taxa.
a, The percentage of the most abundant AMRGs occurring in the different MGE types. b, The number of contigs harbouring the most abundant AMRGs found in the metagenomes analysed. Bars are coloured according to the AM families each AMRG confers resistance to. Shadow bars for each AMRG represent the number of those AMRGs that were identified to occur in any MGE. c, Taxonomic assignment of contigs carrying the most abundant AMRGs, expressed as relative abundance (% of contigs carrying the AMRG indicated on the y axis). d, The association of AMRGs with MGEs by AMRG class in ESKAPEE key taxa and other main AMRGs carrying taxa: A. johnsonii and S. equorum. The size of the pie graph portions indicates the number of AMRGs found for each AMRG class, while the colour indicates the proportion of AMRGs found associated with MGEs for the corresponding species and AMRG class, according to the legend shown on the bar plot figure. AG, aminoglycosides; AP, amphenicols; BL, β-lactams; FPA, folate pathway antagonists; FO, fosfomycin; MC, macrolides; TC, tetracyclines; SGB, streptogramin B.
Extended Data Fig. 5. Ratio of AMRG-carrying contigs.
Association with MGE in the different surfaces and rooms in meat (a) and dairy (b) industry.
The association of AMRGs with MGEs was particularly relevant for ESKAPEE bacteria, especially E. coli (84.8% of AMRGs associated with MGEs), S. aureus (68.1%) and P. aeruginosa (67.9%), mostly linked to plasmids (Fig. 4d). Interestingly, even though β-lactam AMRGs were generally not associated with MGE in the study, in K. pneumoniae 75% of them were associated with MGEs, mainly ComTns carrying blaSHV-27. Aminoglycoside AMRGs were highly associated with integrons, ISs and Tns in A. baumannii, E. coli and P. aeruginosa (Fig. 4d). Meanwhile, AMRGs carried by the most relevant non-ESKAPEE bacteria (for example, S. equorum, A. johnsonii, A. lwoffii, R. ornithinolytica and B. zhoupengii) were rarely associated with MGEs (Fig. 4d and Supplementary Fig. 1b), with some non-ESKAPEE Staphylococcus, including S. haemolyticus and S. saprophyticus, representing an exception, as they harboured their AMRGs mainly in plasmids (Supplementary Fig. 1b).
As a potential indication of AMRG transfer events, we analysed the AMRGs shared by different species in the same production facility by aligning the AMRGs and their surrounding regions with high confidence thresholds. We found a high occurrence of intergenera AMRG sharing in specific taxa pairs, such as Jeotgalicoccus–Staphylococcus (associated with β-lactam AMRGs), Escherichia–Pseudomonas and Acinetobacter–Psychrobacter (mainly associated with folate pathway antagonist AMRGs). (Extended Data Fig. 6). Eight of these AMRG sharing events involving large regions (harbouring >30 coding sequences (CDSs)) were found between Corynebacterium and Rothia. Shared regions carried mainly aminoglycoside, folate pathway antagonist, β-lactam, aminoglycoside, tetracycline and amphenicol AMRGs. Remarkably, they were harboured mainly in samples from processing surfaces and final products (Extended Data Fig. 6).
Extended Data Fig. 6. Inter-genera AMRG sharing events detected.
a) Number of AMRG sharing events found and number of CDS within each AMRG sharing region. The point shape indicates AM class and the color indicates the genera-pair involved in the AMRG sharing events. b) AMRG sharing events (genera-pair) detected between chromosomes, chromosomes-plasmids and plasmids. c) AMRG sharing events abundance according to plasmidic (plasm), chromosomic (chrom) or hybrid (both) gene interchange. Small chord diagrams indicated AMRG shared events between sample type.
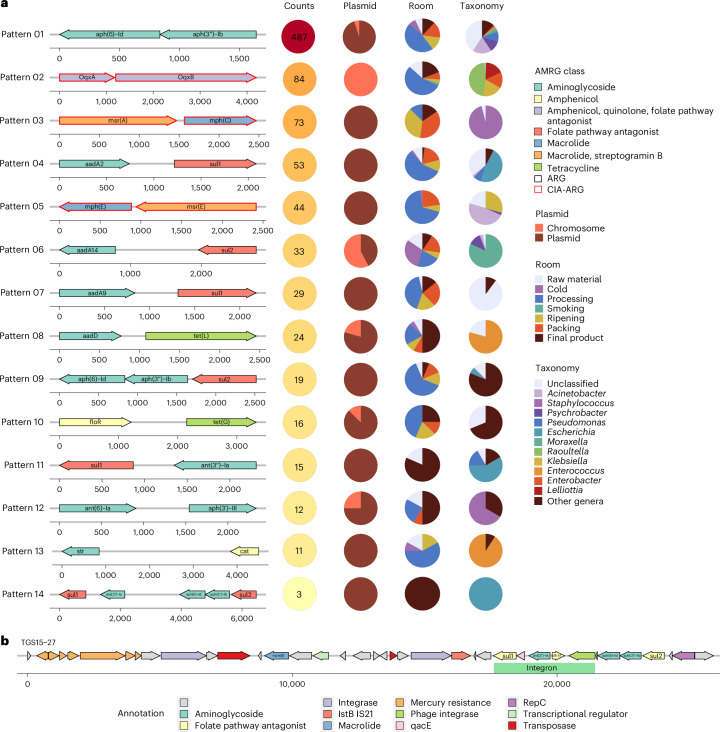
We also evaluated the potential of different AMRGs to be co-selected or transferred together by analysing their co-occurrence in the same contig in close proximity. In total, 18.1% of the AMRG-carrying contigs were identified to harbour two or more AMRGs. Among the most abundant patterns of co-occurrence, three involved CIA genes: OqxA–OqxB, found only on non-MGE contigs assigned to Raoultella, Enterobacter, Lelliottia and Klebsiella mainly from processing rooms; mph(C)–msr(A), found only on plasmidic contigs, mainly from ripening and packing rooms, assigned to Staphylococcus; and mph(E)–msr(E), found only on plasmidic contigs, mainly from processing rooms, assigned to Acinetobacter and Klebsiella (Fig. 5a). Some genes were largely found co-occurring with other AMRGs rather than as unique AMRGs in the contigs. These included aph(3″)-Ib, aph(6)-Id, OqxA, OqxB, msr(A), aadA2, mph(E), msr(E), aadA9, tet(G) and aph(3′)-III (Fig. 5a). Except for the OqxA–OqxB and aadA14–sul2 pairs, all the most prevalent co-occurrence gene patterns were mainly or entirely found in plasmid-related contigs, mainly in processing rooms and/or final products (Fig. 5a). The co-occurrence of AMRGs was primarily observed in ESKAPEE genera, Psychrobacter, Raoultella and Moraxella (Fig. 5a). Two AMRG patterns were found combined together in three final product samples from the same facility in a 26-kbp shared region that also contains seven genes related to mercury resistance and several regions associated with MGEs, as an integron containing sul1, ant(3″)-Ia and dfrA1 (Fig. 5b). All AMRG patterns obtained have been found previously, mainly in plasmidic regions (except pattern 2 and pattern 10) (Supplementary Table 3).
Fig. 5. Co-occurrence of AMRGs within contigs.
a, Circles indicate how often (%) the contigs carrying each co-occurrence pattern were assigned as plasmidic (‘Plasmid’ column); and the room and taxonomic assignment for the contigs (‘Room’ and ‘Taxonomy’ columns, respectively). Only those co-occurrence patterns with more than ten counts are indicated. AMRGs conferring resistance to CIAs are marked with red arrow lines. b, A gene location map representation of the aph(3″)-Ib-aph(6)-Id-sul2-ant(3″)-Ia-sul1 pattern, found three times in contigs with identity >99.9% on the plotted region.
Meat and dairy contact surfaces are main sources of AMRGs in food
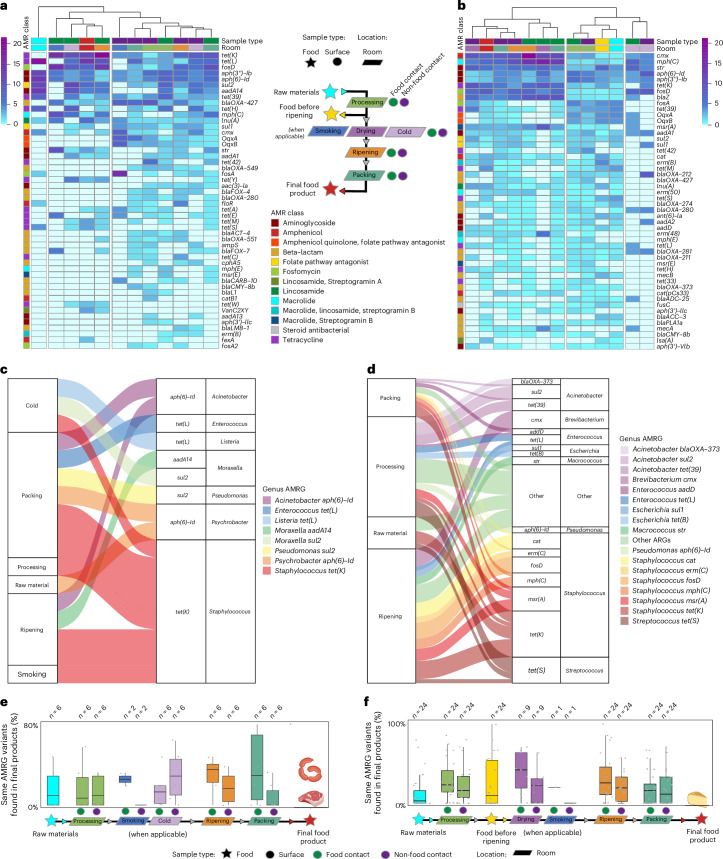
Meat production facilities were the most heterogeneous with respect to manufacturing procedures and nature of the final products (that is, fresh meat products, cured meats, aged meat and dry fermented sausages; Extended Data Fig. 7 and Supplementary Table 4). A comparison of the resistome profile of the different meat-production samples showed that raw materials clustered separately to all other sample categories, mainly due to the absence or very low occurrence of several AMRGs frequently found in surfaces and/or final products, such as tet(K), fosD, blaOXA-427, mph(C), str, sul1 and aadA1 (Fig. 6a). FCSs from ripening, smoking and cold rooms clustered with final products, mainly due to the high occurrence of tet(K) and fosD genes (Fig. 6a), assigned to S. aureus and S. equorum, respectively (Fig. 4). We further aimed to infer through contig alignment if some specific taxa were responsible for these AMRG-sharing events towards the final products and linked them mainly to Acinetobacter (originating from packing and ripening rooms, carrying aph(6)-Id), Moraxella (from ripening and cold rooms, carrying addA14 and/or sul2), Psychrobacter (from packing rooms and raw materials, carrying aph(6)-Id) and Staphylococcus (present ubiquitously and carrying tet(K)) (Fig. 6c). To infer the potential origin of the AMRGs in the final meat products, we analysed the occurrence of the same AMRG variants in both the final products and other sources, on each individual facility, and represented it as a ratio of coincidence (Fig. 6e). It was observed that FCSs from the latest steps of production (that is, ripening and packing) harboured the greatest proportion of the AMRGs found in the final meat products.
Extended Data Fig. 7. AMRG-carrier species distribution along meat production surfaces and relationship with their resistome profile.
Relative abundance of the 50 most abundant AMRG in meat facilities. The type of meat product elaborated in each facility is indicated as “Industry_type2”. The AM class of each gene is colour-indicated on the left part, while sample type and facility room are colour-indicated on top. The heatmap colour scale indicates the number of AMRG identified for each surface.
Fig. 6. Spread of AMRGs across meat and dairy production sites and to end products.
a,b, The relative abundance of the 50 most abundant AMRGs in meat (a) and dairy (b) facilities. The AM class of each gene is colour-indicated on the left part, while sample type and facility room are colour-indicated on top. The heat map colour scale indicates the number of AMRGs for each surface. The flow chart legend represents the flow that the food products (stars) follow from raw materials to final products along the food production system and the different rooms (trapezes). Swab samples were collected from surfaces (circles), coloured green if in contact with food products and purple if not. c,d, Resistome spread events between raw materials/processing environments and end products in meat (c) and dairy (d) producing facilities as revealed through the alignment analysis of AMRG-carrying contigs. The size of the boxes and lines are proportional to the number of spread events found. The right part of the figures corresponds to final product samples, where AMRG and genus are indicated. e,f, The percentage of the same AMRG variant found in the final products and other sample types from the same facility by using assembly-based approaches for meat (e) and dairy (f) producing facilities. Only those facilities where at least five AMRG variants were found in the final products were used for this analysis. The lower, middle and upper hinges in the box plots correspond to the first, second and third quartiles (the 25th, 50th or median and 75th percentiles), respectively. The upper whisker extends from the hinge to the largest value no further than 1.5× IQR from the hinge (where IQR is the interquartile range, or distance between the first and third quartiles). The lower whisker extends from the hinge to the smallest value at most 1.5× IQR of the hinge.
In cheese production, cheese samples collected before ripening and raw materials clustered together and close to FCSs and NFCSs from processing and cold rooms, while final products clustered closer to samples from FCSs and NFCSs from later stages of production, including ripening and packing rooms (Fig. 6b). Remarkably, some AMRGs that were absent or rarely occurring on raw materials, the processing area or the freshly processed cheese showed a high occurrence in final products and industry surfaces. For example, cmx was assigned mainly to Brevibacterium zhoupengii (25.3% of all cmx genes), Brevibacterium aurantiacum (21.6%) and Micrococcus luteus (10.6%); mph(C) to S. equorum (65.9%) and S. saprophyticus (24.8%); tetK to S. aureus (93.8%); fosD to Staphylococcus species (96.0%), particularly S. equorum (67.8%); blaZ to S. aureus (46.2%), Jeotgalicoccus sp. (22.2%) and Mammaliicoccus lentus (19.9%); and msrA to Staphylococcus species (100%), particularly S. saprophyticus (89.1%). By contrast, other AMRGs were almost exclusively found in raw materials and end products, highlighting their persistence throughout cheese processing and ripening, such as tet(M), assigned mainly to Lactococcus lactis (22.6%), Enterococcus faecalis (22.6%) and E. faecium (21.0%); tet(S) to Streptococcus parauberis (63.8%); erm(B) to Lactiplantibacillus plantarum (22.6%), Aerococcus urinaeequi (14.5%) and Staphylococcus pseudintermedius (12.9%); and ant(6)-Ia to S. aureus (54.5%) (Fig. 3). A company-per-company analysis, performed as described above and shown in Fig. 6b, highlighted the role of both FCS and NFCS from the latest stages of cheese production (ripening and packing rooms) as the main source of AMRG-carrying bacteria in final products, while the influence was lower for the raw materials or the initial processing areas. AMRG spread events, identified through contig alignment, mainly involved Staphylococcus originating from ripening rooms (carrying tet(K), msr(A), mph(C), fosD, erm(C) and/or cat genes), and Acinetobacter and Streptococcus originating mainly from processing rooms (carrying tet(39), sul2, blaOXA-373 and/or tet(S)) (Fig. 6d). Overall, FCSs from drying and ripening rooms showed a strong influence on the resistome of the final products (Fig. 6f).
More than 700 AMRG-carrying MAGs were reconstructed
Overall, 713 high-quality MAGs containing AMRGs were reconstructed. They were assigned to 85 known genera (mainly Staphylococcus, 13.5% of all AMRG-carrying MAGs; Acinetobacter, 10.4%; and Raoultella, 5.9%) and 264 different species (including 51 species not assigned to the genus level, coming from uncultured species genome bins), with S. equorum, Acinetobacter guillouiae and Lactococcus lactis being the most represented. MAGs from some species, such as Raoultella ornithinolytica and Shewanella putrefaciens, carried a very conserved resistome, while those from S. equorum, A. guillouiae and L. lactis had a more variable AMRG profile (Extended Data Fig. 8a). As could be expected, the number of different AMRGs and AMRG clusters found in MAGs was much lower than that found in contigs and in the assembly-free analysis (Extended Data Fig. 8b).
Extended Data Fig. 8. Analysis of AMRG-carrying MAGs.
a) Percentage of MAGs obtained per each species (y-axis) that carried the most abundant AMRG (x-axis). Only species with more than 10 AMRG-carrying MAGs were represented (K. pneumoniae, S. aureus and P. aeruginosa were also represented to cover the ESKAPEE group). The number written before each species name indicates the number of AMRG-carrying MAGs found for each species. b) Venn diagram comparing the number of AMRG-clusters found when assessing the resistome using the reads, the contigs or the MAGs datasets.
Discussion
This work provides a comprehensive survey of the resistome in foods and throughout the food production chain, based on a large number of samples collected (expanding the publicly available food-related metagenomes by ~3×), a wide diversity of food types and industrial surfaces sequenced, and a sequencing approach with increased depth (~2× depth, which yielded ~4× number of MAGs19). Detailed information was obtained on the main AMRGs occurring in food production systems, their possible association with specific taxa and MGEs and their spread across production sites. The application of whole-metagenome sequencing approaches in food industry settings faces important challenges, the most relevant one being the recovery of a sufficient amount of microbial DNA from the frequently low-biomass niches sampled. Another limitation is that sequences can originate from both dead and alive cells, although the detection of an AMRG still represents a potential risk for AMR spread20. In addition, the limited sensitivity of the technology for the detection of low-abundance micro-organisms might be overcome through the sequencing of enrichment cultures, although culturing steps introduce important biases in studies like ours focused on studying the biodiversity of systems as enrichments favour fast growers outcompeting other representative members of the community21,22. The MASTER EU Consortium has validated improved sampling and DNA extraction protocols for microbiome analysis tailored to food-processing environments23. Their application, combined with the bioinformatic tools and resources (cFMD and foodGenVir databases) described here and by Carlino et al.19, and available in public repositories24–26, greatly improves sequencing outcomes and will speed up the uptake of microbiome and resistome mapping approaches by the food research community and by industry stakeholders.
Our results indicate that the majority of known AMRGs (>70%), including many conferring resistance to CIAs, circulate daily throughout food production chains, although for most AMRGs the prevalence and abundance is overall low. We identified AMRGs associated with resistance to almost all known AM classes, with a higher representation of those linked to the AM classes most widely consumed in the European Union, that is, tetracyclines, β-lactams, aminoglycosides and macrolides27. While the results suggest that the resistome load and diversity is greater in the production of foods of terrestrial animal origin, in agreement with previous studies28, this would require further confirmation, as our study was unbalanced towards a higher proportion of cheese and meat producers and was limited to a few European countries. Additional studies including more facilities from a wider range of food production sectors and countries would be highly beneficial, as countries have different policies regarding AM usage in agriculture and farming, which will probably be reflected in differences in resistome composition29.
Some AMR ESKAPEE species are considered antibiotic-resistant priority pathogens posing the greatest threat to human health by the World Health Organization, prompting urgent calls for research into the development of new antibiotics effective against them30. We identified bacterial species belonging to the ESKAPEE group among the main carriers of AMRGs, mainly in cheese and meat factories and almost ubiquitously across all samples collected. Among them, S. aureus, a common member of the human skin microbiota that might appear in the processing environment through handling31, was the most frequent AMRG carrier. However, other non-ESKAPEE micro-organisms that are frequently found in food production environments32 and are not so well known or studied as carriers of AMRGs, such as S. equorum, S. saprophyticus, A. johnsonii, A. lwoffii, R. ornithinolytica or B. zhoupengii, were also identified as major contributors to the food production resistome. These findings should be the starting point for additional integrative initiatives specifically targeting these less known food-resistome-associated species aiming to better understand the risk they pose for AMR spread.
MGEs (mainly plasmids) play a crucial role in AMRG spread. As widely described before in other environments17,33, a notable proportion (37.8%) of the AMRGs identified here in food production systems are potentially mobilizable. This was particularly relevant for some ESKAPEE species (E. coli, P. aeruginosa and S. aureus) and AM classes (aminoglycosides and tetracyclines), which thus pose the highest risk for the spread of AMR in food processing environments and to consumers. By contrast, most non-ESKAPEE species were rarely found carring MGE-associated AMRGs. For instance, A. johnsonii, a non-ESKAPEE bacterium associated with food spoilage and just occasionally described as an opportunistic pathogen34, was widely distributed across foods and associated processing environments from all production sectors as a carrier of mainly chromosomal β-lactam blaOXA-type AMRGs; and various members of the genus Staphylococcus (S. equorum and S. saprophyticus) frequently found in food production environments32 were abundant carriers of particular chromosomal AMRGs (mph(C) and fosD in S. equorum and mph(C) and msr(A) in S. saprophyticus), with low occurrence in raw materials and an enhanced representation in processing surfaces and final products.
Assigning taxonomy to contigs may have possible technical complications (especially if contigs are small or located within MGEs)35 and could lead to wrong interpretations due to the overrepresentation in public repositories of genomic regions from certain taxa (for example, clinically associated ESKAPEE genomes). To avoid these issues, we have been conservative in our analyses, eliminating short contigs (<1,000 bp; <3 CDS) not taxonomically assigned at the species level. This strategy minimizes assignment errors and yields reliable taxonomic assignment of AMRG-carrying contigs with known affiliation with a marginal proportion of false taxonomic assignments almost exclusively involving plasmidic contigs that were erroneously assigned mainly to closely related taxa, as described in detail in the Methods. Overall, despite the potential limitations, we believe the approach followed here offers important benefits. It provides insights into the genetic background of AMRGs that cannot be obtained through an assembly-free analysis (due to the inability to assign AMRGs to carrier taxa or MGEs), or from analyses focused on MAGs, as only a small proportion of contigs are ultimately clustered into MAGs36, making the reconstruction of MGEs particularly challenging37.
We obtained solid evidence highlighting the impact that some manufacturing processes and processing environments have on the resistome of end products, which overall had a higher amount and richness of AMRGs than raw materials, including genes not present at all in the latter and possibly introduced later in the production chain. Important differences were observed in the resistome composition of foods between pre- and postripening stages of production, with end products showing AMRG profiles closely resembling those from surfaces from the latest stages of manufacture (ripening and packing rooms). Our findings contradict those of some previous studies reporting a decrease in the amount and diversity of AMRGs along slaughterhouse processing lines38,39, suggesting that, although some processing procedures may be effective in reducing or containing the spread of resistant micro-organisms, new contamination events and other ecological forces along food processing lines may favour the introduction and/or expansion of other resistant micro-organisms and AMRGs. Overall, the facility surfaces and the microbial succession dynamics driven by processing conditions showed a greater impact on the resistome of end products than the raw inputs did. Factors such as temperature or humidity shifts, pH and water activity changes along fermentation and ripening, or frequency of cleaning and disinfection of surfaces, among others, can facilitate the expansion of specialized food-production-adapted AMR micro-organisms not present or present at very low abundance in raw materials, with good abilities to grow at low temperatures, pH and/or water activity, or to form biofilms in food production surfaces40. For instance, staphylococci, found here as main AMRG carriers in surfaces and end products, are well equipped to grow in products with intermediate or low water activity, such as fermented sausages or ripened cheeses41,42. Likewise, other main AMRG carriers, such as Acinetobacter, Pseudomonas or Moraxella, are frequently found in processing environments and are good biofilm formers, having representative species capable of growing at low temperatures43.
The evaluation of potential AMRG spread events also showed linkages between the resistome of final products and that of surfaces, especially FCSs from ripening and packing rooms. Although this influence might be bidirectional, the fact that processing surfaces were sampled here before processing started, thus reflecting the resident microbiome, suggests they may serve as an important source of AMRGs in foods. Previous investigations have described the importance of the house-specific microbiome for the proper maturation of some food products and the development of their organoleptic properties44. Our results support the concept of ‘natural contamination’ of food products from the production environment12, highlighting that processing plant microbiomes can greatly influence food safety by contributing to the spread of AMR bacteria that may cross-contaminate food products. Considering that industrial production of food relies on intense sanitation schemes to maintain a high level of hygiene as a means to ensure food quality and safety, and that some disinfectants and detergents used with this aim can impose selective pressures enhancing the chances of persistence of AMR micro-organisms and AMRG spread45,46, our findings emphasize the importance of also considering the appropriate use of these agents in food production settings when designing efficient AM stewardship policies.
Methods
Sample collection, DNA extraction and sequencing
A total of 1,780 samples, taken from 113 food industries, were analysed in this study, including 824 from processing environments, 254 from raw materials and 702 final product samples (Supplementary Table 4). Samples from processing environments included FCSs (n = 489) and NFCSs (n = 335), which were sampled using Whirl-Pak Hydrated PolyProbe swabs (Whirl-Pak) to cover an area of about 1 m2. Five swabs from each sample type (for example, floors and tables) were collected and pooled before DNA extraction. All the samples were stored at 4 °C and transported to the laboratory, where they were preprocessed within 24 h following the procedure described by Barcenilla et al.23.
The surface of vegetables, fish, fresh meat and cured meat (both raw materials and end products) was also swabbed in sterile conditions, and five swabs per sample type were also pooled together and processed following the same procedures as for the environmental swabs.
Ten millilitres of phosphate-buffered saline (PBS) 1× were added to each pool of five swabs. After homogenization in the Stomacher (300 rpm × 30 s), the cell suspension was collected and aliquoted in 15-ml sterile tubes (Eppendorf), which were centrifuged at 14,000g for 2 min. Samples were washed twice with 2 ml of sterile PBS, and cell pellets were then stored at −80 °C until further processing for DNA purification.
Other types of samples collected included raw milk (50 ml), brine (when available, 200 ml), whey (200 ml), cheeses before ripening (2 cheeses) and after ripening (2 cheeses) in cheese industries; and meat batter (50 g), sausages before ripening (2 sausages) and after ripening (2 sausages) in factories producing dry fermented sausages. Raw milk and whey samples were centrifuged at 5,000g for 15 min at room temperature. The fat layer was carefully removed, and the supernatant was decanted. Cell pellets were resuspended with a sterile PBS solution, followed by a new centrifugation under the same conditions. This washing step was repeated once, and cell pellets were stored at −80 °C until further processing for DNA purification. Brine samples were prefiltered using 10-µm filters to remove large particles, and then the permeate was filtered through a 0.22-μm filter membrane. Micro-organisms were recovered from the 0.22-μm filter by placing the filter in a sterile 50-ml Falcon tube and adding 0.5 ml of sterile PBS, pipetting up and down for 1 min. Then, the suspension was centrifuged at 5,000g for 5 min at room temperature, and cell pellets were stored at −80 °C until further processing for DNA purification. For meat batter, cheese and sausage samples, 10 g were homogenized with 90 ml of PBS using a Stomacher at maximum speed for 2 min. Afterwards, the homogenate was centrifuged at 5,000g for 15 min at room temperature. Three serial washes with PBS were performed as described for milk and whey samples. Finally, cell pellets were stored at −80 °C.
DNA extraction was performed from the cellular pellets using a DNA extraction procedure recently developed for microbiome analysis in food processing environments, based on the PowerSoil Pro Kit of Qiagen with modifications, to increase total microbial DNA yields in low-biomass samples23. The DNA concentration was quantified using the Qubit HS Assay (Thermo Fisher Scientific).
Metagenomic libraries were prepared using the Nextera XT Index Kit v2 (Illumina), and whole-metagenome sequencing was performed on an Illumina NovaSeq platform using 150-bp PE, generating an average of 7.5 Gb and ~50 million PE reads per sample.
Raw sequencing data preprocessing
Reads were quality filtered and adapters were trimmed using a preprocessing pipeline47, which includes TrimGalore48 using --nextera--stringency 5--length 75--quality 20--max_n 2--trim-n parameters.
Assembly-free analysis
Filtered reads were aligned against the ResFinder database49 (downloaded on 24 October 2022) using Bowtie250 with --very-sensitive--end-to-end parameters. Obtained sequence alignment map (SAM) files were filtered by an in-house ruby script51, which removes the gene overestimation occurring when forward and reverse reads are aligned within the same gene. The obtained counts matrix was processed to calculate the CPM adding a ‘bacterial marker’ modification according to
where ‘CPM’ is the total CPM value for each AMRG and ‘Bacterial marker alignments’ is the value obtained from viromeQC52, using --minlen 0--minqual 0 parameters, to normalize the dataset to consider only those reads derived from bacteria.
Reads are 150 bp in length and do not cover the entire length of the AMRG. As some AMRGs differ by just one or a few single-nucleotide polymorphisms, genes from the ResFinder database were clustered at 90% identity using CD-HIT53 with default values to reduce the chance of the same read aligning to multiple closely related genes. The gene name used for each cluster obtained (gene cluster name), which was chosen according to the most abundant gene within each cluster or by the first number of the cluster within bla and addA genes, is indicated in the ‘Genefam’ column in Supplementary Table 3.
The AMRG richness and the Simpson diversity index values were calculated using the R package vegan54 for alpha-diversity analysis, and statistical differences were evaluated with the Wilcoxon signed-rank test through the R package ggpubr55. Principal coordinates analyses with Bray–Curtis dissimilarities and Hellinger transformation using the vegdist function were used for beta-diversity analyses, and within-group dispersion was evaluated by the betadisper function. Both functions are located in the R package vegan54. Finally, the effects of industry type, surface and room on sample dissimilarities were determined by permutational multivariate analysis of variance using distance matrices (PERMANOVA) with the adonis function in the R package vegan54. The compare_means function in the R package ggpubr55 was used to include statistically significant differences in the box plot figures.
In addition, the same alpha- and beta-diversity analyses were repeated on a subset of AMRGs conferring resistance to CIAs, selected according to previous publications56–58.
Assembly-based analysis
For each sample, high-quality reads were assembled by using Megahit v1.1.159. Contigs’ files were imported into the TORMES v1.3.0 pipeline60 (parameters: --min_contig_len 1000--only_gene_prediction--prodigal_options ‘-p meta’ --gene_min_id 80--gene_min_cov 80), which discards contigs below 1,000 bp in length. The taxonomy of the remaining contigs was assigned using Kraken261 and the k2_pluspfp_20221209 database, and predicted and translated CDSs and proteins were extracted using Prodigal62. The screening of AMRGs in contigs was performed using BLASTN63, integrated in ABRicate (Seemann64), against the ResFinder database49 (downloaded on 24 October 2022). The output file was manually curated to solve BLASTN and ABRicate issues associated with their default option ‘culling_limit = 1’, which wrongly annotated the variants of the aminoglycosides resistance genes aadA as ant(3″)-Ia, even when the identity and the coverage of the latter was lower. Contigs with a length below 1,000 bp and fewer than 3 CDSs per contig (26.1% of the total number of contigs) were discarded. The thresholds were decided after thorough evaluation of the methodology being applied over (1) mock communities used as positive controls (ZymoBIOMICS Microbial Community Standard D6300; Zymo Research) in three different concentrations (that is, 10−2, 10−4 and 10−6) and in four different laboratories; (2) 4,229 genomes (belonging to the most abundant AMRG-carrying species found in this study, that is, A. baumannii, A. johnsonii, E. faecium, E. coli, S. aureus and S. equorum) downloaded from RefSeq; (3) assessing the confidence and root-to-leaf (RTL) scores provided by Kraken2 (after polishing confidence values with Conifer v1.0.265) accounting to taxonomic assignment per ranks and contig lengths; and (4) benchmarking compared with SqueezeMeta pipeline66. The methodology and results are thoroughly described and discussed in the Supplementary Information. Only the contigs assigned to the species level were considered for the downstream resistome and mobilome analysis. When possible, pipelines were parallelized to optimize computing resources by using GNU-Parallel67.
The potential presence of AMRGs in MGEs was evaluated by using different software and in-house scripts. The evaluation of plasmids was performed directly over the AMRG-carrying contigs by using Platon v1.668 and by enabling the option --meta and geNomad v1.6.169. In addition, the AMRG-carrying contigs were screened for the presence of plasmid replicons using the PlasmidFinder database70 and the BLASTN and ABRicate commands implemented in TORMES. Contigs were considered to be plasmidic when a plasmid replicon was present and/or when both Platon (enabling --meta option) and geNomad (enabling end-to-end option) detected a contig to be plasmidic. The occurrence of integrons was evaluated using IntegronFinder v2.0.171. Potential LGT events were evaluated using WAAFLE v1.0.018. The location of AMRGs within integron regions was confirmed using the ‘ARG-contig_mobilome_analysis’ pipeline72. MobileElementFinder v1.0.573 was used to screen AMRG-carrying contigs for the detection of MITEs, ISs, ComTns, Tns, ICEs, IMEs and cis-mobilizable elements (setting flag --min-coverage 0.8).
Finally, an approach developed by Groussin et al.74 was used for AMRG sharing events. AMRG-carrying contigs were aligned with themselves by BLASTN using 99.9% identity and 500-bp alignment length cut-offs, and aligned regions were extracted to confirm the presence of AMRGs by BLASTN against the Resfinder database using 80% identity and subject coverage cut-offs. Only those hits between two different samples from the same food processing facility were kept for further analysis of AMRG spread events (when query and subject sequences were assigned to the same genus) and genera AMRG sharing events (when query and subject sequences were assigned to different genera). The functional annotation of shared regions was performed by eggnog-mapper v2.1.1175 using MMseqs276 with eggNOG DB v5.0.2. Output files from eggnog-mapper were transformed using an in-house script77 into a gggenes formatted table, which was manually curated to then perform gene distribution plots by using the gggenes R package78. In addition, those contigs of particular interest (that is, carrying multiple AMRGs and/or MGEs) were aligned with BLASTN against the nt database79 to assess whether such contig structures had been previously found.
The reconstruction of metagenome-assembled genomes (MAGs) was performed as described by Carlino et al.19. In brief, high-quality reads from each sample were mapped to their corresponding assembled contigs by using bowtie2 v2.2.980, and by enabling parameters ‘--very-sensitive-local’ and ‘--no-unal’. Then, the jgi_summarize_bam_contig_depths script from MetaBAT v2.12.181 was used to calculate contig depth values from the SAM files obtained after bowtie2 alignment, as a preliminary step before contig binning with MetaBAT2. Only contigs longer than 1,500 bp were subjected to binning. The CheckM v1.0.13 ‘lineage_wf’ workflow82 was used to assess the quality of putative reconstructed genomes, and only those with completeness ≥50% and contamination < 5% (that is, medium- and high-quality MAGs, with high-quality MAGs being those with completeness >90% (ref. 83) were retained for further analyses. Average nucleotide identity among quality-controlled genomes was computed through Mash84. MAGs were then imported into the TORMES v1.3.0 pipeline60 for CDS prediction, annotation and resistome evaluation by using the same parameters described above for the contigs dataset.
The MAGs containing AMRGs were used to create the foodGenVir database, a database containing AMRG-carrying MAGs from food and food-related environments that is publicly available via Zenodo at ref. 85, together with information on their origin and AMR profile, among other information. It is important to acknowledge that many AMR determinants associated with MGEs may not be represented in the database, as non-chromosomally encoded contigs are normally not clustered into MAGs during binning.
Data visualization
Statistic analyses and data handling towards visualization were performed under R environment v4.3.186, by using circlize v0.4.1687, ComplexHeatmap v2.22.088, dplyr v1.1.489, ggalluvial v0,12.590, gggenes v0.5.178, ggplot2 v3.5.191, ggpubr v0.6.055, ggtree v3.14.092, khroma v1.16.093, pheatmap v1.0.1294, reshape2 v1.4.495, tidyr v1.3.196 and vegan v2.6.1054 packages. Antibiotic families from the ResFinder database were reorganized to facilitate and simplify data visualization. Only one-type antibiotic families were kept, and CPMs for each of these one-type antibiotic families were calculated by adding up the CPMs found on every ResFinder-Antibiotic family containing the one-type antibiotic family. As an example, the ‘Oxazolidinone’ family also contains those genes attached to the ‘Amphenicol, Oxazolidinone, Tetracycline’, ‘Oxazolidinone, Amphenicol’ and ‘Oxazolidinone, Amphenicol, Lincosamide, Streptogramin A, Pleuromutilin’ families from ResFinder.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Supplementary information
Supplementary Fig. 1 and discussion.
Supplementary Table 1. The number of AMRG and gene clusters per antibiotic class found on ResFinder and detected at the read and contig level. Values between brackets correspond to AMRG for CIAs. Supplementary Table 2. AMR gene clusters from ResFinder database used for short-read sequence alignment. Supplementary Table 3. The number of contigs carrying AMRG associated with the different species identified in the study. Supplementary Table 4. The relationship of samples collected through the MASTER project. Supplementary Table 5. Blastn versus nt (NCBI) results for 14 AMRG patterns found. Supplementary Table 6. The ratio of contigs correctly assigned to the different taxonomic ranks of the sequenced mock community (Zymo) as positive controls. Supplementary Table 7. The ratio of AMR-carrying contigs broken in silico to harbour three CDSs that were correctly assigned to the different taxonomic ranks of key organisms downloaded from RefSeq. Supplementary Table 8. Benchmarking of the pipeline followed for taxonomic assignment with the SqueezeMeta pipeline for AMRG-carrying contigs.
Acknowledgements
This work was funded by the European Commission under the European Union’s Horizon 2020 research and innovation programme under grant agreement no. 818368 (MASTER). N.M.Q. is currently funded under the Generation D initiative, promoted by Red.es, an organization attached to the Ministry for Digital Transformation and the Civil Service, for the attraction and retention of talent through grants and training contracts, financed by the Recovery, Transformation and Resilience Plan through the European Union’s Next Generation funds (ref. MMT24-IBFG-01). C.B. is grateful to Junta de Castilla y León and the European Social Fund for awarding her a predoctoral grant (grant no. BOCYL-D-07072020-6). The COMET-K1 competence centre FFoQSI (number 881882) is funded by the Austrian federal ministries BMK, BMDW and the Austrian provinces Lower Austria, Upper Austria and Vienna within the scope of COMET - Competence Centers for Excellent Technologies. The program COMET is handled by the Austrian Research Promotion Agency, FFG.
Extended data
Author contributions
M.L., M.P., V.T.M., M.W., A.M., N.S., P.D.C., D.E. and A.A.-O. conceived the study and obtained the funding. N.M.Q., J.F.C.-D., V.V., C.B., F.D.F., R.C.-R., N.C., M.D., I.C.-T., C.S., L.R., S.K. and S.S. performed the samplings at food-processing facilities. N.M.Q., V.V., C.B., F.D.F., R.C.-R., N.C., M.D., I.C-T., C.S., S.K. and S.S. performed samples preprocessing and DNA extraction. F.P. and N.S. sequenced the extracted DNA. N.C. and F.P. performed the initial bioinformatic analyses, from raw reads to filtered reads, assembled contigs and binned MAGs. N.M.Q., J.F.C.-D., V.V., R.C.R., N.C. and C.S. collated all the metadata information. N.M.Q., J.F.C.-D. and V.V. performed the downstream bioinformagic and resistome analyses and prepared the figures. N.M.Q, J.F.C.-D., V.V. and A.A.-O. wrote the manuscript with input from all authors. All authors read and approved the final manuscript.
Peer review
Peer review information
Nature Microbiology thanks Franck Carbonero and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Data availability
Raw reads are available on the Sequence Read Archive of the National Center of Biotechnology Information (NCBI) under the BioProject numbers PRJNA897099 for vegetable facilities, PRJNA997800 for meat facilities and PRJNA997821 for cheese facilities, except those located in Ireland. Raw reads for fish processing factories and Irish cheese factories are available on the European Nucleotide Archive database under accession numbers PRJEB62794 and PRJEB63604, respectively. The specific BioSample, BioProject and public accession IDs for each sample, together with additional metadata are provided in Supplementary Table 4. The FoodGenVir database is publicly available via Zenodo at 10.5281/zenodo.8344969 (ref. 85).
Code availability
The pipelines and scripts used for reads filtering, assembly and binning, and resistome analysis are available via GitHub at https://github.com/SegataLab/MASTER-WP5-pipelines/tree/master/02-Preprocessing, https://github.com/SegataLab/MASTER-WP5-pipelines/tree/master/05-Assembly_pipeline, and https://github.com/nmquijada/MASTER-food-production-resistome, respectively.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
These authors contributed equally: Narciso M. Quijada, José F. Cobo-Díaz, Vincenzo Valentino.
Extended data
is available for this paper at 10.1038/s41564-025-02059-8.
Supplementary information
The online version contains supplementary material available at 10.1038/s41564-025-02059-8.
References
- 1.Fisher, M. C., Hawkins, N. J., Sanglard, D. & Gurr, S. J. Worldwide emergence of resistance to antifungal drugs challenges human health and food security. Science360, 739–742 (2018). [DOI] [PubMed] [Google Scholar]
- 2.Antimicrobial Resistance Collaborators. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet399, 629–655 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.GBD 2019 Diseases and Injuries Collaborators. Global burden of 369 diseases and injuries in 204 countries and territories, 1990-2019: a systematic analysis for the Global Burden of Disease Study 2019. Lancet396, 1204–1222 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.O’Neill, J. I. M. Antimicrobial resistance: tackling a crisis for the health and wealth of nations. (The Review on Antimicrobial Resistance, 2014); https://amr-review.org/sites/default/files/AMR%20Review%20Paper%20-%20Tackling%20a%20crisis%20for%20the%20health%20and%20wealth%20of%20nations_1.pdf
- 5.WHO: Antimicrobial Resistance, 2023 (WHO, 2024).
- 6.Kumar, A. & Pal, D. Antibiotic resistance and wastewater: correlation, impact and critical human health challenges. J. Environ. Chem. Eng.6, 52–58 (2018). [Google Scholar]
- 7.Van Boeckel, T. P. et al. Reducing antimicrobial use in food animals. Science357, 1350–1352 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Singh, R. K. et al. Influence of diet on the gut microbiome and implications for human health. J. Transl. Med.15, 73 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Temkin, E. et al. Estimating the number of infections caused by antibiotic-resistant Escherichia coli and Klebsiella pneumoniae in 2014: a modelling study. Lancet Glob. Health6, e969–e979 (2018). [DOI] [PubMed] [Google Scholar]
- 10.O’Neill, J. Tackling drug-resistant infections globally: final report and recommendations. (The Review on Antimicrobial Resistance, 2016); https://amr-review.org/sites/default/files/160518_Final%20paper_with%20cover.pdf
- 11.Buchanan, R. L., Gorris, L. G. M., Hayman, M. M., Jackson, T. C. & Whiting, R. C. A review of Listeria monocytogenes: an update on outbreaks, virulence, dose-response, ecology, and risk assessments. Food Control75, 1–13 (2017). [Google Scholar]
- 12.Quijada, N. M. et al. Autochthonous facility-specific microbiota dominates washed-rind Austrian hard cheese surfaces and its production environment. Int. J. Food Microbiol.267, 54–61 (2018). [DOI] [PubMed] [Google Scholar]
- 13.De Filippis, F., La Storia, A., Villani, F. & Ercolini, D. Exploring the sources of bacterial spoilers in beefsteaks by culture-independent high-throughput sequencing. PLoS ONE8, e70222 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Mulani, M. S., Kamble, E. E., Kumkar, S. N., Tawre, M. S. & Pardesi, K. R. Emerging strategies to combat ESKAPE pathogens in the era of antimicrobial resistance: a review. Front. Microbiol.10, 539 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Quijada, N. M., Hernández, M. & Rodríguez-Lázaro, D. High-throughput sequencing and food microbiology. Adv. Food Nutr. Res.91, 275–300 (2020). [DOI] [PubMed] [Google Scholar]
- 16.Peterson, E. & Kaur, P. Antibiotic resistance mechanisms in bacteria: relationships between resistance determinants of antibiotic producers, environmental bacteria, and clinical pathogens. Front. Microbiol.9, 2928 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Wang, X. et al. Global increase of antibiotic resistance genes in conjugative plasmids. Microbiol. Spectr.11, e0447822 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hsu, T. Y. et al. Profiling lateral gene transfer events in the human microbiome using WAAFLE. Nat. Microbiol.10, 94–111 (2025). [DOI] [PubMed] [Google Scholar]
- 19.Carlino, N. et al. Analysis of 2,500 food metagenomes reveals unexplored microbial diversity and links with the human microbiome. Cell187, 5775–5795 (2024). [DOI] [PubMed] [Google Scholar]
- 20.Pires, S. M., Duarte, A. S. & Hald, T. Source attribution and risk assessment of antimicrobial resistance. Microbiol. Spectr.6, 3 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.McHugh, A. J. et al. Microbiome-based environmental monitoring of a dairy processing facility highlights the challenges associated with low microbial-load samples. npj Sci. Food5, 4 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Prakash, O., Parmar, M., Vaijanapurkar, M., Rale, V. & Shouche, Y. S. Recent trend, biases and limitations of cultivation-based diversity studies of microbes. FEMS Microbiol. Lett.368, fnab118 (2021). [DOI] [PubMed] [Google Scholar]
- 23.Barcenilla, C. et al. Improved sampling and DNA extraction procedures for microbiome analysis in food-processing environments. Nat. Protoc.19, 1291–1310 (2024). [DOI] [PubMed] [Google Scholar]
- 24.Segata, N. cFMD. GitHubhttps://github.com/SegataLab/cFMD (2025).
- 25.Segata, N. MASTER-WP5-pipelines. GitHubhttps://github.com/SegataLab/MASTER-WP5-pipelines (2024).
- 26.Quijada, N.M. MASTER-food-production-resistome. GitHubhttps://github.com/nmquijada/MASTER-food-production-resistome (2025).
- 27.Antimicrobial Resistance in the EU/EEA—A One Health Response (ECDC, 2022).
- 28.Munk, P. et al. The European livestock resistome. mSystems9, e0132823 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Zhao, C., Wang, Y., Mulchandani, R. & Van Boeckel, T. P. Global surveillance of antimicrobial resistance in food animals using priority drugs maps. Nat. Commun.15, 763 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Summary for policymakers. In Intergovernmental Panel on Climate Change – Climate Change 2013 – The Physical Science Basis (ed. IMO) 1–30 (Cambridge Univ. Press, 2014); 10.1017/CBO9781107415324.004
- 31.Krismer, B., Weidenmaier, C., Zipperer, A. & Peschel, A. The commensal lifestyle of Staphylococcus aureus and its interactions with the nasal microbiota. Nat. Rev. Microbiol.15, 675–687 (2017). [DOI] [PubMed] [Google Scholar]
- 32.Marino, M., Frigo, F., Bartolomeoli, I. & Maifreni, M. Safety-related properties of staphylococci isolated from food and food environments. J. Appl. Microbiol.110, 550–561 (2011). [DOI] [PubMed] [Google Scholar]
- 33.Goren, M. G. et al. Transfer of carbapenem-resistant plasmid from Klebsiella pneumoniae ST258 to Escherichia coli in patient. Emerg. Infect. Dis.16, 1014–1017 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kämpfer, P. in Encyclopedia of Food Microbiology (eds Batt, C. A. & Tortorello, M. L.) 11–17 (Elsevier, 2014); 10.1016/B978-0-12-384730-0.00002-1
- 35.Fishbein, S. R. S., Mahmud, B. & Dantas, G. Antibiotic perturbations to the gut microbiome. Nat. Rev. Microbiol.21, 772–788 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Maguire, F. et al. Metagenome-assembled genome binning methods with short reads disproportionately fail for plasmids and genomic Islands. Microb. Genom.6, mgen000436 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Brown, C. L. et al. Critical evaluation of short, long, and hybrid assembly for contextual analysis of antibiotic resistance genes in complex environmental metagenomes. Sci. Rep.11, 3753 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Li, L. et al. Exploring the resistome, virulome, mobilome and microbiome along pork production chain using metagenomics. Int. J. Food Microbiol.371, 109674 (2022). [DOI] [PubMed] [Google Scholar]
- 39.Campos Calero, G. et al. Deciphering resistome and virulome diversity in a porcine slaughterhouse and pork products through its production chain. Front. Microbiol.9, 2099 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Li, S. et al. Microbiome-informed food safety and quality: longitudinal consistency and cross-sectional distinctiveness of retail chicken breast microbiomes. mSystems5, e00589–20 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Li, H., Zhu, Q., Chen, X., Zhou, J. & Wu, J. Isolation and characterization of coagulase negative staphylococci with high proteolytic activity from dry fermented sausages as a potential starter culture. Food Res. Int.162, 111957 (2022). [DOI] [PubMed] [Google Scholar]
- 42.Klempt, M., Franz, C. M. A. P. & Hammer, P. Characterization of coagulase-negative staphylococci and macrococci isolated from cheese in Germany. J. Dairy Sci.105, 7951–7958 (2022). [DOI] [PubMed] [Google Scholar]
- 43.Møretrø, T. & Langsrud, S. Residential bacteria on surfaces in the food industry and their implications for food safety and quality. Compr. Rev. Food Sci. Food Saf.16, 1022–1041 (2017). [DOI] [PubMed] [Google Scholar]
- 44.Bokulich, N. A. & Mills, D. A. Facility-specific “house” microbiome drives microbial landscapes of artisan cheesemaking plants. Appl. Environ. Microbiol.79, 5214–5223 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Botelho, J., Cazares, A. & Schulenburg, H. The ESKAPE mobilome contributes to the spread of antimicrobial resistance and CRISPR-mediated conflict between mobile genetic elements. Nucleic Acids Res.51, 236–252 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Palleja, A. et al. Recovery of gut microbiota of healthy adults following antibiotic exposure. Nat. Microbiol.3, 1255–1265 (2018). [DOI] [PubMed] [Google Scholar]
- 47.Segata, N. MASTER-WP5-pipelines. GitHubhttps://github.com/SegataLab/MASTER-WP5-pipelines/tree/master/02-Preprocessing (2024).
- 48.Krueger, F. TrimGalore. GitHubhttps://github.com/FelixKrueger/TrimGalore (2023).
- 49.Zankari, E. et al. Identification of acquired antimicrobial resistance genes. J. Antimicrob. Chemother.67, 2640–2644 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Langmead, B., Wilks, C., Antonescu, V. & Charles, R. Scaling read aligners to hundreds of threads on general-purpose processors. Bioinformatics35, 421–432 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Segata, N. MASTER-WP5-pipelines. GitHubhttps://github.com/SegataLab/MASTER-WP5-pipelines/blob/master/07-AMR_virulence_genes/count_reads.rb (2024).
- 52.Zolfo, M. et al. Detecting contamination in viromes using ViromeQC. Nat. Biotechnol.37, 1408–1412 (2019). [DOI] [PubMed] [Google Scholar]
- 53.Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics28, 3150–3152 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Oksanen, J. et al. vegan: Community Ecology Package. R version 4.3.1 (2022).
- 55.Kassambara, A. ggpubr: “ggplot2” Based Publication Ready Plots. R version 4.3.1 (2023).
- 56.Cobo-Díaz, J. F. et al. Microbial colonization and resistome dynamics in food processing environments of a newly opened pork cutting industry during 1.5 years of activity. Microbiome9, 204 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Young, C. C. W. et al. Antibiotic resistance genes of public health importance in livestock and humans in an informal urban community in Nepal. Sci. Rep.12, 13808 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.EFSA Panel on Biological Hazards (BIOHAZ). et al. Role played by the environment in the emergence and spread of antimicrobial resistance (AMR) through the food chain. EFSA J. 19, e06651 (2021). [DOI] [PMC free article] [PubMed]
- 59.Li, D., Liu, C.-M., Luo, R., Sadakane, K. & Lam, T.-W. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics31, 1674–1676 (2015). [DOI] [PubMed] [Google Scholar]
- 60.Quijada, N. M., Rodríguez-Lázaro, D., Eiros, J. M. & Hernández, M. TORMES: an automated pipeline for whole bacterial genome analysis. Bioinformatics35, 4207–4212 (2019). [DOI] [PubMed] [Google Scholar]
- 61.Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol.20, 257 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform.11, 119 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinform.10, 421 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Seemann, T. abricate. GitHubhttps://github.com/tseemann/abricate (2023).
- 65.Silamiķelis, I. Conifer. GitHubhttps://github.com/Ivarz/Conifer (2024).
- 66.Tamames, J. & Puente-Sánchez, F. Squeezemeta, a highly portable, fully automatic metagenomic analysis pipeline. Front. Microbiol.9, 3349 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Tange, O. Gnu Parallel 2018. Zenodo10.5281/zenodo.1146014 (2018).
- 68.Schwengers, O. et al. Platon: identification and characterization of bacterial plasmid contigs in short-read draft assemblies exploiting protein sequence-based replicon distribution scores. Microb. Genom.6, mgen000398 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Camargo, A. P. et al. Identification of mobile genetic elements with geNomad. Nat. Biotechnol.42, 1303–1312 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Carattoli, A. et al. In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob. Agents Chemother.58, 3895–3903 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Néron, B. et al. IntegronFinder 2.0: identification and analysis of integrons across bacteria, with a focus on antibiotic resistance in Klebsiella. Microorganisms10, 700 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Cobo-Díaz, J. F. ARG-contig_mobilome_analysis. GitHubhttps://github.com/JoseCoboDiaz/ARG-contig_mobilome_analysis (2023).
- 73.Johansson, M. H. K. et al. Detection of mobile genetic elements associated with antibiotic resistance in Salmonella enterica using a newly developed web tool: MobileElementFinder. J. Antimicrob. Chemother.76, 101–109 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Groussin, M. et al. Elevated rates of horizontal gene transfer in the industrialized human microbiome. Cell184, 2053–2067 (2021). [DOI] [PubMed] [Google Scholar]
- 75.Cantalapiedra, C. P., Hernández-Plaza, A., Letunic, I., Bork, P. & Huerta-Cepas, J. eggNOG-mapper v2: functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol.38, 5825–5829 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol.35, 1026–1028 (2017). [DOI] [PubMed] [Google Scholar]
- 77.Segata, N. MASTER-WP5-pipelines. GitHubhttps://github.com/SegataLab/MASTER-WP5-pipelines/blob/master/07-AMR_virulence_genes/emapper2gggenes.rb (2024).
- 78.Wilkins, D. gggenes: Draw Gene Arrow Maps in “ggplot2”. R version 4.3.1 (2023).
- 79.NCBI Resource Coordinators. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res.42, D7–D17 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods9, 357–359 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Kang, D. D. et al. MetaBAT 2: an adaptive binning algorithm for robust and efficient genome reconstruction from metagenome assemblies. PeerJ7, e7359 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res.25, 1043–1055 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Pasolli, E. et al. Extensive unexplored human microbiome diversity revealed by over 150,000 genomes from metagenomes spanning age, geography, and lifestyle. Cell176, 649–662 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Ondov, B. D. et al. Mash: fast genome and metagenome distance estimation using MinHash. Genome Biol.17, 132 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Quijada, N. M. et al. FoodGenVir database. Zenodo10.5281/zenodo.8344969 (2023).
- 86.R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2023).
- 87.Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. circlize implements and enhances circular visualization in R. Bioinformatics30, 2811–2812 (2014). [DOI] [PubMed] [Google Scholar]
- 88.Gu, Z. Complex heatmap visualization. iMeta1, e43 (2022). [DOI] [PMC free article] [PubMed]
- 89.Wickham, H., François, R., Henry, L., Müller, K. & Vaughan, D. dplyr: A Grammar of Data Manipulation. R version 4.3.1 (2023).
- 90.Brunson, J. C. ggalluvial: layered grammar for alluvial plots. J. Open Source Softw.5, 2017 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Wickham, H. ggplot2 – Elegant Graphics for Data Analysis (Springer, 2016); 10.1007/978-0-387-98141-3
- 92.Yu, G., Smith, D. K., Zhu, H., Guan, Y. & Lam, T. T. Y. ggtree: an R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol.8, 28–36 (2016). [Google Scholar]
- 93.Frerebeau, N. nfrerebeau/khroma v1.3.0. Zenodo10.5281/zenodo.1472077 (2019).
- 94.Kolde, R. pheatmap: Pretty Heatmaps. R version 4.3.1 (2019).
- 95.Wickham, H. Reshaping data with the reshape package. J. Stat. Softw.21, 1–20 (2007). [Google Scholar]
- 96.Wickham, H., Vaughan, D. & Girlich, M. tidyr: Tidy Messy Data. R version 4.3.1 (2023).
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Fig. 1 and discussion.
Supplementary Table 1. The number of AMRG and gene clusters per antibiotic class found on ResFinder and detected at the read and contig level. Values between brackets correspond to AMRG for CIAs. Supplementary Table 2. AMR gene clusters from ResFinder database used for short-read sequence alignment. Supplementary Table 3. The number of contigs carrying AMRG associated with the different species identified in the study. Supplementary Table 4. The relationship of samples collected through the MASTER project. Supplementary Table 5. Blastn versus nt (NCBI) results for 14 AMRG patterns found. Supplementary Table 6. The ratio of contigs correctly assigned to the different taxonomic ranks of the sequenced mock community (Zymo) as positive controls. Supplementary Table 7. The ratio of AMR-carrying contigs broken in silico to harbour three CDSs that were correctly assigned to the different taxonomic ranks of key organisms downloaded from RefSeq. Supplementary Table 8. Benchmarking of the pipeline followed for taxonomic assignment with the SqueezeMeta pipeline for AMRG-carrying contigs.
Data Availability Statement
Raw reads are available on the Sequence Read Archive of the National Center of Biotechnology Information (NCBI) under the BioProject numbers PRJNA897099 for vegetable facilities, PRJNA997800 for meat facilities and PRJNA997821 for cheese facilities, except those located in Ireland. Raw reads for fish processing factories and Irish cheese factories are available on the European Nucleotide Archive database under accession numbers PRJEB62794 and PRJEB63604, respectively. The specific BioSample, BioProject and public accession IDs for each sample, together with additional metadata are provided in Supplementary Table 4. The FoodGenVir database is publicly available via Zenodo at 10.5281/zenodo.8344969 (ref. 85).
The pipelines and scripts used for reads filtering, assembly and binning, and resistome analysis are available via GitHub at https://github.com/SegataLab/MASTER-WP5-pipelines/tree/master/02-Preprocessing, https://github.com/SegataLab/MASTER-WP5-pipelines/tree/master/05-Assembly_pipeline, and https://github.com/nmquijada/MASTER-food-production-resistome, respectively.