Abstract
Background
Ovarian cancer represents the most lethal gynecological cancer with poor response to checkpoint inhibitors. Human endogenous retroviruses (HERVs) are aberrantly expressed by tumor cells and may represent a source of shared T-cell epitopes for cancer immunotherapy regardless of the tumor mutational burden.
Methods
A transcriptomic analysis based on RNA sequencing was developed to quantify the expression of HERV-K sequences containing the selected epitopes. The presence of HERV-K/HML-2 Gag antigen was then assessed by immunohistochemistry (IHC) on tumor microarrays from ovarian cancer samples and normal ovarian tissues. A specific immunopeptidomics approach was developed to detect epitopes on human leukocyte antigens (HLA) molecules. Epitope-specific CD8+ T cells were quantified by multimer staining. HERV-specific T cells were obtained after in vitro stimulation of T cells from HLA-A2-positive healthy donors or patients with ovarian cancer, and in vitro target cell killing was evaluated using real-time analysis. In vivo antitumor efficacy of HERV-specific T cells was assessed in an avian embryo model.
Results
Epitope-containing HERV transcripts were significantly higher in ovarian cancers compared with normal tissues. The presence of the HERV-K/HML-2 Gag antigen was confirmed by IHC in 20/40 (50%) ovarian cancers while no Gag expression was found in normal ovarian tissue samples. Immunopeptidomics analysis revealed the presence of epitopes on HLA molecules on the surface of ovarian tumor cell lines but not on normal primary cells from critical tissues. Low percentages of HERV-specific T cells were detected among tumor-infiltrating lymphocytes from ovarian cancers. Furthermore, in vitro stimulation of patient T cells induced functional epitope-specific T cells, confirming the immunogenicity of these epitopes in patients with ovarian cancer. In vitro, HERV-specific T cells specifically killed ovarian cancer cells in an HLA class I-restricted manner while sparing normal HLA-A2-positive primary cells derived from critical tissues. Epitope-specific CD8+ T cells exhibited a strong antitumoral activity in vivo, inducing a highly significant decrease in tumor volume in comparison with control groups.
Conclusion
These results provide the preclinical rationale for developing T-cell-based approaches against HERV-K-derived epitopes in ovarian cancer.
Keywords: Immunotherapy, Ovarian Cancer, T cell
WHAT IS ALREADY KNOWN ON THIS TOPIC
Human endogenous retroviruses (HERV) expression has been described in different tumors including ovarian cancer. The characterization of HERV-derived tumor epitopes and of the corresponding T-cell responses remained to be performed to develop T-cell-based immunotherapies in ovarian cancer.
WHAT THIS STUDY ADDS
HERV-K antigens are expressed and give rise to epitopes presented on human leukocyte antigen molecules in ovarian cancer but not in normal cells. These HERV-K-derived epitopes induce high-avidity cytotoxic CD8+ T cells that recognize and kill ovarian cancer cells in vitro and in vivo while sparing normal cells. This study also shows that functional HERV-specific T cells can be obtained from patients with ovarian cancer after specific stimulation.
HOW THIS STUDY MIGHT AFFECT RESEARCH, PRACTICE OR POLICY
This new type of shared tumor epitopes can be used to develop vaccines, adoptive T-cell therapies or T-cell recruiting bispecific antibodies to treat a large population of patients with ovarian cancer.
Introduction
Ovarian cancer represents the most lethal gynecological cancer. The majority of patients are diagnosed at an advanced stage and relapse after first-line chemotherapy is common, leading to poor patient outcome.1 2 Reported results of programmed cell death 1 (PD-1) or programmed cell death ligand 1 (PD-L1) blocking antibodies have been disappointing with an objective response rate under 10%.3 Furthermore, combinations of anti-PD-L1 antibodies with chemotherapy have also shown negative results.4 New approaches are thus warranted to induce efficient antitumor immune responses in patients with ovarian cancer.
The efficacy of tumor-specific T cells relies on the recognition of tumor epitopes presented on human leukocyte antigen (HLA) molecules on the surface of cancer cells.5 In this context, neoepitopes derived from non-synonymous somatic mutations have been shown to induce high-avidity tumor-specific T cells.6 However, the development of personalized cancer vaccines or adoptive T-cell therapies targeting this type of epitopes remains challenging, especially in tumors characterized by a low mutational burden, such as ovarian cancer.7 Thus, there is still an unmet need to identify new tumor epitopes, ideally shared across patients, in ovarian cancer.
Human endogenous retroviruses (HERVs) represent 8% of the human genome. These DNA sequences are the legacy of multiple ancient germline infections by exogenous retroviruses.8,10 HERVs are mainly localized in heterochromatin and repressed by epigenetic silencing in normal cells, even if a transcriptional background has been described in normal tissues.11 Aberrant expression of HERVs has been reported in different tumors, including breast,12 ovarian,13 prostate14 and kidney cancers,15 melanoma,16 glioblastoma,17 as well as hematological malignancies.18 Some HERVs, especially the most recently integrated ones (such as HERV-K family), have retained partially conserved open reading frame (ORF) encoding Gag, Pol or Env proteins. We have previously demonstrated that HERVs represent a source of shared tumor epitopes capable of inducing T-cell clones with high functional avidity. We have further characterized CD8+ T cell clones specific to HLA-A2-restricted 9-mer epitopes derived from HERV-K and shown their antitumor activity in triple-negative breast cancer and acute myeloid leukemia.19 20 Combining a new transcriptomic quantification method with immunohistochemistry (IHC) and targeted immunopeptidomics, we report here that HERV-K-derived epitopes are specifically expressed in ovarian cancer cells compared with normal cells. We also demonstrate that HERV-specific T cells selectively kill ovarian cancer cells, both in vitro and in vivo, while sparing normal cells, providing the preclinical rationale for developing T cell-based approaches against these new targets in ovarian cancer.
Material and methods
In silico analyses
Datasets: RNA sequencing (RNA-seq) reads from The Cancer Genome Atlas (TCGA) were obtained from the Genomic Data Commons (GDC) portal. They were downloaded in the bam format (already aligned) and generated fastq (raw reads) using the PicardTools software’s SamToFastq (V.2.26.10) command. RNA-seq reads from Genotype-Tissue Expression (GTEx) V.8 (https://gtexportal.org/home/) were obtained using the Terra portal as indicated on their website, and generated fastq data similarly to TCGA sequences. Tumor mutational burdens (TMBs) from TCGA ovarian samples were retrieved from Wang et al.21
Tissue selection: From TCGA, we selected all available ovarian tumor samples (n=421) and selected peritumoral healthy samples from the seven critical tissues with at least 10 samples: Lung (n=110), kidney (n=129), liver (n=49), bladder (n=19), stomach (n=36), head and neck (n=44) and colon/rectum (n=51). From GTEx, we selected all the available samples of the same type of tissue obtained from TCGA allowing batch correction, and we added other critical normal tissues that were not available in TCGA.
Reference annotation: We used the University of California San Francisco (UCSF) hg38 as reference genome and Gencode V.33 for gene annotations. For HERV annotation, we extracted the retro.hg38.v1 version from the Telescope database.22 The two annotations were then merged. To avoid duplicates, we ran gffcompare23 to identify overlaps between regions. We then computed the Jaccard similarity between regions that overlapped; regions with a similarity above 0.5 were considered duplicates, and to maintain a consistent nomenclature of HERVs, only the ones originating from the Telescope annotation were kept. We then only considered protein-coding genes and HERV transcripts.
Transcript quantification: Similarly to SalmonTE24 and REdiscoverTE25 pipelines, Salmon26 (V.1.10.0) was used on the unaligned reads to quantify how many reads were associated with each locus.23 The reference index was built with the index option set at 15, and the reference sequences were generated using the gffread (V.0.12.8) (22) package. Mapping and quantification were then performed by calling salmon count with –validateMappings –l A options.
Expression normalization: The pseudo counts obtained for all samples were loaded into R (4.1.2) using the tximport function, from the tximport library.27 Before any normalization, we computed for each sample the sum of counts of all HERV loci with a potential ORF containing the epitope of interest (FLQ, RLI and YAM). These epitope-containing HERV transcript counts were added to the whole dataset, as with any transcript. The counts were then normalized using the median of ratio function from DESeq2 library and transformed into log scale with the vst function.28
Data exploration and analyses: All analyses were done using R (V.4.1.2) and RStudio server (2023.12.1-402). Differential expression analyses between ovarian tumor tissues and all selected peritumoral tissues were performed with DESeq2 (Wald test). P values were corrected for multiple testing using the Benjamini-Hochberg method. Batch corrections between TCGA and GTEx were run using the Combat function29 from the sva package, with TCGA as reference Batch and were done tissue per tissue as previously described in a study by Wang et al.30 Visualizations were done with ggplot231 and ggbiplot library.
Peptide sequences matching Uniprot: The whole reviewed Human Uniprot database (Swiss-Prot) was downloaded from uniprot.org (release 2024_01). Exact sequence pattern matching was performed between each peptide sequence and Uniprot db, and no match was found with any known human protein except with HERV-K-derived proteins.
Immunohistochemistry
The OV1004a ovary tissue microarray (TMA) from TissueArray was selected for IHC analysis. This TMA contains a total of 40 ovarian tumor samples, each of them in duplicate (online supplemental table 2). Five normal ovarian tissues were also analyzed (from OV1004a FDA3331a, TissueArray; #T8234701-5, BioChain). When available, tumor samples from patients whose TILs were analyzed were also subjected to IHC staining for HERV-K/HML-2 Gag expression. IHC was performed on an automated immunostainer (Ventana Benchmark Ultra, Roche, France) using the ultraView Universal DAB Detection Kit. Slides were deparaffinized at 72°C using Ventana EZ Prep reagent (Roche, Cat. 950–102) and hydrated, followed by an antigen retrieval method using Ventana Tris-EDTA buffer pH 7.8 (Roche, Cat. 950–224) for 32 min at 95°C. Sections were incubated with the HERV-K/HML-2 Gag monoclonal antibody (diluted at 1:200, Austral Biologicals, Cat. HERM-1841–5) for 32 min and then with the ultraView Universal HRP Multimer. As a negative control, the microarray was processed similarly but incubated with the ultraView Universal HRP Multimer only. Staining was revealed with 3,3′-diaminobenzidine as a chromogenic substrate for 8 min. Then, the sections were counterstained with Gill’s Hematoxylin (Roche, Cat. 760–2021) for 8 min and counterstained with Bluing reagent (Roche, Cat. 760–2037) for 4 min.
For staining quantification, TMAs were scanned with panoramic scan II (3D Histech, HU) at 20×. Images were generated using Caseviewer software (3Dhistotech 2.4.0.119028) for coloration and IHC. Captured images were processed using HALO AI software (Indica Labs, V.3.6.4134) and adapted algorithms for quantification (Multiplex IHC V.3.4.9). Each spot was annotated by an anatomopathologist to outline the tumor and eliminate artifact zones. The Mininet neural network was used to train the software to discriminate stroma and tumor structures. The nuclear and cytoplasmic stains were quantified with an algorithm taking into account the percentage of positive cells per spot and cell staining intensity (three intensities used: low, moderate, high). Staining was also evaluated by two operators and scored manually (scores: 0, 1, 2 or 3) according to staining intensities. The final quantification considers both HALO AI software quantification and scores obtained by both operators. A score of 3 was classified as high, scores of 2 or 1 as moderate and a score of 0 as negative.
Immunofluorescence
For immunofluorescence (IF) staining, the OV1004a ovary TMA previously used for IHC staining was also selected. Slides were incubated 2 hours in 60°C and then placed in Bond RX automated immunostainer (Leica Biosystems). Multiplexed IF was performed using the OPAL 6-Plex detection kit (NEL821001KT, AKOYA biosciences) which uses tyramide signal amplification (TSA)-conjugated fluorophores. Slides were dewaxed at 72°C with Bond Dewax Solution (Leica Biosystems, AR9222) and antigen retrieval was performed using Bond Epitope Retrieval Solution 1 (ph6, Leica, AR9961) for 20 min at 95°C.
Sequential IF was performed according to the steps below. A 20-min stripping at 95°C corresponding to the antigen retrieval was performed before antibody staining. The following antibodies were used: Anti-FoxP3 (RTU, PA0263, Leica) followed by Anti-Mouse IgG1 HRP (1/250) (Jackson ImmunoResearch, 115-035-205); OPAL 520 Anti-CD20 (1/600, M0755, Dako); secondary antibody OPAL HRP Rabbit (Leica, ARR1001KT); OPAL 480 Anti-CD3 (1/100, A0452, Dako); secondary antibody OPAL HRP Rabbit (Leica, ARR1001KT); OPAL 690 Anti-CD8 (1/50, M7103, Dako) with secondary antibody Anti Mouse IgG1 HRP (1/250) (Jackson ImmunoResearch, 115-035-205) OPAL 620 (1/200) Anti-CD68 (1/100, M0876, Dako); secondary antibody Anti Mouse IgG total HRP (1/250) (Invitrogen, G21040); OPAL 570 Anti-CK (1/100, M3515, Dako) secondary antibody OPAL HRP Rabbit (Leica, ARR1001KT); TSA DIG (1/100) + OPAL 780 The slides were counterstained with DAPI and then mounted using Prolong Gold Antifade Reagent (Invitrogen, Ref# P36930).
Stained formalin-fixed paraffin-embedded (FFPE) slides were scanned on a Vectra Polaris Imaging System (Akoya Biosciences, Marlborough, Massachusetts, USA; RRID:SCR_023774) at ×20 magnification. Slides were then visualized with Phenochart (V.1.0.12, AKOYA Biosciences; RRID:SCR_019156) for multispectral imaging and were annotated and quantified using Halo AI software (Indica Labs, V.4.0), Phenochart and adapted algorithms for quantification (HighPlex FL).
Biological samples
Blood from healthy donors was obtained from the “Etablissement Français du Sang” (Lyon, France). For the TIL cohort, fresh high-grade serous ovarian carcinomas (n=13) were provided by the tissue bank of the Centre Léon Bérard (CLB) (BB-0033–00050, CRB - CLB, Lyon, France; French agreement number: AC-2013–1871). For the patient peripheral blood mononuclear cell (PBMC) cohort, frozen PBMCs from patients with high-grade serous ovarian carcinoma (n=7) were provided by Institut Universitaire du Cancer de Toulouse on approval by the Institutional Review Board and written informed consent under the DEcipher Cancer ImmunogEnicity (DECIdE) protocol (NCT03958240) and from the CLB (n=5, BB-0033–00050, CRB - CLB, Lyon, France; French agreement number: AC-2013–1871) after approval from the institutional review board and ethics committee (L-06–36 and L-11–26) and patient written informed consent. All samples were collected in accordance with the Declaration of Helsinki. All patients had histologically documented tumors, were≥18 years old at inclusion and were followed with a standard-of-care procedure. Exclusion criteria were: seropositivity for hepatitis B, hepatitis C, HIV or hantavirus; any condition incompatible with a blood sampling procedure; pregnancy or breastfeeding; and suspected or documented active autoimmune disease or use of immunosuppressive drugs.
Cell lines and primary cell cultures
T2 (Cat. ACC 598) cells were purchased from DSMZ (Germany), OVCAR3 (Cat. HTB-161), OV90 (Cat. CRL-3585) and SKOV3 (Cat. HTB-77) cells were purchased from The American Type Culture Collection (ATCC) and were cultured according to the manufacturer’s instructions with 1% penicillin/streptomycin (PS, Gibco Cat. 15140122) added to the culture medium. To generate HLA-A2+SKOV3, cells were transduced with a lentiviral vector encoding HLA-A2 (RD Biotech) containing a puromycin resistance gene (RD Biotech). Transduced cells were selected and maintained with 3 µg/mL of puromycin. HLA-A2+ normal primary cells HCM (Cat. C-12810, batch#463Z016.2, female donor DN000249), HBEpC (Cat. C-12640, batch#469Z016, female donor DN000331) and NHEK (Cat. C-12003, batch#467Z005.1, female donor DN000068) were purchased from Promocell GmbH (Germany), nHKPT cells (Cat. 3253021, batch#RPCT022823, male donor) from Tebubio (France) and HA cells (Cat. 1800-SC, batch#35382) from ScienCell (USA). All human normal primary cells were cultured following the supplier’s recommendations. All cell lines were cultured at 37°C in humidified atmosphere with 5% CO2. OVCAR5 tumor cell line was available at Complete Omics for mass spectrometry (MS)-immunopeptidomics analysis.
Epitope detection by mass spectrometry
Epitope validation by MS was performed by Complete Omics (Maryland, USA) according to the method previously described in a study by Douglass et al32 with additional modifications. Briefly, for sample preparation, at least 20 million cells were lysed and peptide-HLA complexes were isolated using in-house packed Valid-NEO neoantigen enrichment column preloaded with anti-human pan-HLA antibodies modified with conjugated chemical moieties that are designed to increase the binding efficacy between the column matrix and HLA-epitope complex molecules. The MaxRec technology was applied to increase the sensitivity of the detection by preventing sample loss. Several versions of MaxRec peptides were developed to compare their protection efficacy, together with chemically modified MaxRec peptides. After elution, dissociation, filtration and cleanup, epitope peptides were lyophilized before further analysis.
Detection parameters for each epitope were examined and curated through a Valid-NEO method builder bioinformatics pipeline developed by Complete Omics. Precursor or fragmented ions with excessive noise due to environment or coelution with impurities were excluded from the detection. To boost detectability, a series of recursive optimizations for the significant ions were conducted.
HERV-K/HML-2 Gag protein detection by western blot
Ovarian cancer cells were lysed in cold radioimmunoprecipitation assay (RIPA) buffer (R0278, Merck) supplemented with a protease inhibitor cocktail (#11697498001, Merck) and phosphatase inhibitor cocktail (#04906837001, Roche) for 30 min on ice. Cell lysates were cleared by centrifugation (13,000 g, 15 min, 4°C), and protein concentration was determined by a bicinchoninic acid (BCA) protein assay (#23227, Thermo Fisher). Proteins (20 µg) were resolved on sodium dodecyl-sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and transferred onto polyvinylidene fluoride (PVDF) membranes by electroblotting. Non-specific binding sites were blocked using Tris-buffered saline containing 0.1% Tween-20 and 5% (w/v) dry milk. Mouse monoclonal IgG1 anti-HERV-K Gag (clone 4D4/F7, #HERM1841-5, AMSBIO Europe BV, Netherlands), and mouse monoclonal IgG1 anti-actin (clone C4, #MAB1501, Millipore) were incubated overnight at 4°C. After incubation with an anti-mouse secondary antibody (Peroxidase AffiniPure Donkey Anti-Mouse IgG (H+L) #Ref 715-035-151, Jackson Immunoresearch) conjugated to horseradish peroxidase, blots were revealed using the electrochemiluminescence (ECL) reagents (#170–5060, Bio-Rad) and acquired on a Chemidoc (Bio-Rad) imaging system.
Fresh tumor dilaceration and expansion of tumor-infiltrating lymphocytes (TILs)
Tumor tissues were dissected into fragments of approximately 1 mm3 and digested with collagenase IV (Sigma, Cat. C2674) and DNase I (Sigma, Cat. D4513) during mechanical dilaceration, for 45 min in Roswell Park Memorial Institute (RPMI) containing 20% Fetal Bovine Serum (FBS, Eurobio, France, Cat. CVFSVF00-01, batch S80412-4602). The tumor lysate was centrifuged at 1500 rpm for 5 min, counted and resuspended in 5% human serum enriched RPMI (Gibco Cat. 52400041). Cells were counted and plated in a flat bottom 96-well plate with anti-CD3 anti-CD28 Dynabeads (Dynabeads Human T-Expander CD3/CD28 Cat. 11 141D, Gibco, France) and interleukin (IL)-2 (Proleukin, Vidal Cat. at 100 IU/mL) in a beads to cells ratio of 1:4, to expand tumor-infiltrating lymphocytes (TILs). Dextramer (Immudex ApS, Denmark) staining was performed on TILs expanded for 14 days after tumor dilaceration at room temperature prior to viability and surface marker staining. A dextramer complexed to a non-natural irrelevant peptide (ALIAPVHAV, HLA-A2 neg. control Cat. WB02666 Immudex ApS, Denmark, European Union) was used as a negative control.
All samples were analyzed on a LSR-Fortessa (BD Biosciences, France) with calibrated settings to compare samples throughout the entire study. Data were analyzed using FlowJo Software (Tree Star V.10.4, New Jersey, USA).
In vitro priming assays and generation of epitope-specific CD8+ T cells
PBMCs were obtained by Ficoll density gradient centrifugation (Eurobio, France, Cat. CMSMSL01-01). For healthy donor PBMCs, monocytes were isolated and cultured for 6 days in RPMI medium in the presence of GM-CSF (PeproTech, Cat.300–03) 200 ng/mL and IL-4 (PeproTech, Cat. 200–04) 50 ng/mL for the differentiation of monocyte-derived dendritic cells (MoDCs). MoDCs were matured with tumor necrosis factor alpha (TNF-a, PeproTech Cat. 300–01A 20 ng/mL) and Polyinosinic:polycytidylic acid (PolyIC, Invivogen Cat. tlrl-PIC-5 40 µg/mL) 16 hours before pulsing with the peptide of interest and cultured with autologous T cells in a MoDC:T ratio of 1:10. Cells were co-cultured for 14 days in AimV medium enriched with IL-7 (Miltenyi Cat.130-095-363) and IL-15 (PeproTech Cat.200–1 s), changing media periodically and restimulating a second time with peptide-pulsed MoDCs after 7 days. For patients, PBMCs (from blood EDTA-tubes) were washed twice in phosphate buffered saline (PBS) and resuspended in complete Xvivo-15 medium (Lonza, Cat.02-060Q) supplemented with 1% Penicillin/Streptomycin (P/S) and a cocktail of cytokines comprizing granulocyte-macrophage colony-stimulating factor (GM-CSF, 100 ng/mL, PeproTech, Cat.300–03), IL-4 (100 ng/mL PeproTech, Cat. 200–04) and IL-7 (100 ng/mL, Miltenyi, Cat.130-095-363). One day after, PBMCs were stimulated according to the protocol described by Roudko et al.33 Briefly, cells were stimulated with peptides at a final concentration of 10 µM, in the presence of lipopolysaccharide from E.Coli (LPS, 100 ng/mL, Invivogen, Cat. tlrl-3pelps), R848 (3.5 µg/mL, Invivogen, Cat.tlrl-r848-5) and IL-1β (10 ng/mL, Biotechne, Cat. 201-LB). Cultures were maintained at 37°C in a humidified 5% CO₂ atmosphere. The next day and every 2–3 days, the medium was refreshed with complete RPMI 1640 medium supplemented with 10% human serum (sAb, Biowest, Cat.S4190-100), 1% L-glutamine, 1% PS, IL-2 (10 IU/mL, Proleukin, Vidal Cat. at 100 IU/mL) and IL-7 (10 ng/mL, Miltenyi Cat.130-095-363). On days 10–12, cells were harvested and analyzed for peptide-specific T-cell responses by flow cytometry or enzyme-linked immunospot assay (ELISpot) before sorting.
Tetramers were produced by conjugating biotinylated HLA-A*02:01 peptide-specific monomers (P2R Facility, Nantes) with phycoerithrin (PE)-streptavidin (BioLegend, USA, Cat. 405204) or allophycocyanin (APC)-streptavidin (BioLegend, Cat.405207) and used to evaluate the presence of specific cells by fluorescence-activated cell sorting (FACS) analysis together with a viability dye (Zombie NIR Fixable Viability Kit, Cat.423106, BioLegend) and cell surface markers, including anti-human CD3 (BV421, 1/40, Cat.306718, BioLegend) and anti-human CD8 (APC, 1/100, Cat.344722, BioLegend).
After expansion, antigen-specific T cells were isolated from the in vitro cell culture by magnetic monomer sorting. Positive and negative fractions were cultured separately and expanded on feeders composed of 35 Gy-irradiated allogeneic PBMCs and B-lymphoblastoid cell lines (kindly provided by Dr Henri Vie CRCI2NA-Equipe12 INSERM UMR 1307/CNRS UMR6075). Feeder cells were plated in a 96-well round bottom plate at a concentration of 0.10×106 cells per well in RPMI 5% human serum with leukocyte phytohemagglutinin (PHA-L, 1.5 µg/mL, Merck KgAa), Germany, Cat.30852801), IL-2 150 IU/mL and IL-7 (10 ng/mL). Cells were cultured for 14 days, and medium was replaced when needed with IL-2-enriched and IL-7-enriched fresh RPMI 5% human serum. This process was repeated if needed to obtain a specific population (>80%).
T cell functional avidity and IFNγ ELISpot
To analyze antigen-specific responses, 5×104 peptide-stimulated patient PBMCs were co-cultured with 3×10⁴ T2 cells pre-pulsed with 10⁻⁵ M of HERV cognate peptide or an irrelevant control peptide. The co-culture was performed in interferon-gamma (IFN-γ) ELISpot plates to assess T-cell responses at the individual well level (IFN-γ ELISpot, Diaclone, France, Cat.856-051-020). A well was considered positive when more than 50 spots were detected and when the spot count was at least twice that observed in the corresponding well stimulated with the irrelevant peptide.
Functional avidity of sorted specific CD8+ T-cells was assessed using IFN-γ ELISpot assay (IFN-γ ELISpot, Diaclone, France, Cat.856-051-020). Epitope-specific CD8+ T cells were restimulated with T2 cells pulsed with decreasing concentrations of the cognate peptide (from 10−6 to 10−13 M) or with an irrelevant peptide (10−6 M) used as a negative control in a T cell:T2 cell ratio of 1:10 in RPMI medium (Gibco, France, European Union) supplemented with 8% human serum. After 18 hours, supernatants were removed and ELISpot was performed. The minimal peptide concentration required to achieve a significant cytokine response was determined (>30 spots and twofold change in comparison to the negative control). The peptide concentration required to achieve a half-maximal cytokine response (EC50) was determined using GraphPad Prism (V.6.0 for Windows was used for the 50% EC (EC50) determinations, R>0.98).
FACS analysis of target cell death
Cytotoxic activity of patient-derived specific CD8+ T cells was evaluated after co-culture at a 5:1 Effector (E):Target (T) ratio with T2 target cells labeled with carboxyfluorescein succinimidyl ester CFSE (1/1000, C34554, Thermo Fisher Scientific) and pulsed with either the cognate HERV peptide or an irrelevant peptide at a concentration of 10−6 M. After 24 hours, the percentage of CFSE+target cell death was analyzed by multiparametric flow cytometry using a viability dye (Zombie NIR Fixable Viability Kit, Cat.423106, BioLegend).
Real-time killing assay
Killing assays were performed using an xCELLigence RTCA eSight Cell Analyzer (Agilent, France). OVCAR3 tumor cell line and normal primary cells were seeded in E-plates 16 (Agilent) and cultured overnight before adding T cells in an E:T ratio of 2:1 or 5:1. When required, synthetic peptides at a final concentration of 1 nM for 2 hours at 37°C, blocking anti-major histocompatibility complex-I (MHC-I) antibody (50 µg per well; clone W6/32, Bio X Cell, Cat. BE0079) or isotype control (final volume 100 µL) were added. Annexin V fluorescein isothiocyanate (FITC, BioLegend, France, Cat. 640945) was added to each well (f.d. 1/1000) for cell death analysis in live imaging.
Impedance variation (cell index) measurement and live imaging acquisition were performed in real time, every 15 min for 48 hours, after the addition of T cells and each condition was performed in triplicate. The Normalized Cell Index (NCI) was calculated at the time of T cell addition. Percent of lysis was calculated using RTCA Software Pro Immunotherapy Module and normalized against unspecific T cell. Annexin V staining was measured on images (green object count/image) at different time points (0, 6, 24 and 48 hours after the beginning of the co-culture) using the RTCA eSight software.
In vivo anti-tumor efficacy
Measurement of the in vivo antitumoral activity of HERV-specific CD8+ T cells was performed by Oncofactory (Lyon, France). Embryonated eggs were obtained from a local supplier (EARL Les Bruyeres, France). The sanitary status of laying hens was regularly checked by the supplier according to French laws. Development of embryos and in ovo xenografts was performed as previously described in a study by Jarrosson et al.34 Eggs were incubated at 38.5°C in a humidified incubator until the desired developmental stage (stage HH14, 2 days post-fertilization). Embryos were randomized in each experimental group and were harvested at embryonic day 4 (stage HH25, 4 days post-fertilization).
OVCAR3 cells were labeled with 7 µM CFSE solution (Life Technologies, Cat. C34554) and mixed with T cells at an E:T of 5:1 before co-engraftment. At stage HH14 chick embryos were co-engrafted with mixed cells in presumptive somitic areas with a glass capillary connected to a pneumatic PicoPump (PV820, World Precision Instruments) under a fluorescence stereomicroscope. Targeted tissue areas for the graft were visualized under the stereomicroscope. Co-transplanted embryos were harvested after 48 hours (at stage HH25) and the in vivo antitumoral effect of CD8+ T cells was evaluated by assessing the tumor volume using three-dimensional (3D) light-sheet imaging. For this analysis, tissues from n=45 chicken embryos were cleared and imaged using whole-mount selective plane illumination microscopy (SPIM) imaging with volumetric analyses performed as previously described in a study by Jarrosson et al.34
One-way analysis of variance post-hoc Dunnett’s test or pairwise Wilcoxon rank test was used for multigroup comparisons. P values were considered statistically significant if p<0.05, with ***p<0.001.
Statistical analyses
All the statistical analyses were performed using R statistical software V.4.4.3 and are presented in supplementary statistical data analysis document in the online supplemental data file.
Results
HERV-K antigens are selectively expressed in tumor cells compared with normal cells
Based on our previous study,19 we selected three HLA-A2-restricted epitopes derived from cancer-associated HERV-K sequences, which were shown to be immunogenic and able to induce high-avidity cytotoxic T cells. The two 9-mer epitopes FLQFKTWWI (FLQ) and RLIPYDWEI (RLI) stem from HERV-K Gag and YAMSNLFSI (YAM) from HERV-K Pol. To quantify the expression of these epitopes at the transcriptomic level in ovarian cancer, we first identified all the HERVs with a predicted ORF containing the corresponding epitope sequences. The sequences of the FLQ and RLI epitopes were found in 33 and 37 HERVs, respectively, while the sequence of the YAM epitope was detected in 43 HERVs (online supplemental table 1). Importantly, these peptides were not found in any known protein sequence in Uniprot except for HERV-K-derived proteins. Differential expression analysis performed between ovarian cancer and critical peritumoral tissues from TCGA normalized RNA-seq data (figure 1A) showed that the majority of the HERV loci containing one or several epitope sequences are more abundantly expressed in ovarian tumor samples than in all peritumoral tissues (figure 1B). Of these, HERV-K/HML-2_19q11, HERV-K/HML-3_10q22.1, and both HERV-K/HML-2_19q11 and HERV-K/HML-2_11q12.3b were the most differentially expressed transcripts containing FLQ, YAM and RLI peptides, respectively (figure 1C). We then computed the sum of the raw counts for all the transcripts containing a given epitope sequence in each sample to compare its expression (figure 1A). The sum of epitope-containing HERV transcripts was significantly higher (p<0.01) in ovarian cancer for all three peptides compared with the seven normal tissues considered to be critical tissues, with more than one log2-fold change in most tissues (figure 1D). To verify that the peritumoral tissues available in TCGA are representative of normal critical tissues, we also processed normal samples from the GTEx RNA-seq database in the same way as for TCGA. The expression of epitope-containing HERVs in GTEx healthy critical tissues was no higher in brain, blood vessels, blood, heart, skin or muscle than in the other critical tissues also available in TCGA (online supplemental figure 1A). To validate that the epitope-containing HERVs were more abundantly expressed in ovary tumors than in normal tissues, we incorporated GTEx-related samples to the TCGA dataset, although, as shown by the uniform manifold approximation and projection (UMAP) and HERV expression levels (online supplemental figure 1B,C), GTEx and TCGA datasets could not be compared directly. We therefore ran a batch correction on the whole dataset using ComBat29 for each tissue independently (except for ovarian peritumor samples that are not available in TCGA) (online supplemental figure 1D). This approach confirmed that the expression levels were significantly higher (p<0.01) in ovary tumors than in any normal peritumoral tissue for all three peptides (online supplemental figure 1E). Interestingly, the expression of epitope-containing HERVs in ovarian cancer was not associated with mutational burden (online supplemental figure 2).
Figure 1. Transcriptomic analysis of HERV epitope expression. (A) Schematic representation of data processing. Salmon was used to map and quantify reads to a merged reference transcript annotation containing both genes and HERV annotations. Transcriptomic expression of the epitope was estimated using the sum of all HERV loci with a potential ORF containing its sequence (FLQ, RLI, and YAM). Data were normalized and log transformed (vst) using DESeq2. (B) Volcano plot showing differentially expressed HML-2/HERV-K loci in ovarian cancer against peritumoral tissues from TCGA. Dark-gray dots represent the HML-2/HERV-K loci differentially expressed (log2FC>1 and FDR<0.01, Wald test from DESeq2). Red dots represent the HML-2/HERV-K loci that are differentially expressed and contain FLQ, RLI and/or YAM epitopes in a potential ORF. (C) Differential expression of epitope-containing HERV loci in ovarian cancer against peritumoral tissues from TCGA. Red and blue indicate, respectively, over-expressed or under-expressed HERV loci in ovarian cancer compared with peritumoral tissues; color intensity represents the degree of perturbation (log2FC), and the size of dots the statistical significance (FDR). (D) Epitope-containing HERVs in ovarian cancer (red) and peritumoral tissues from TCGA. Epitope expression levels represent the sum of epitope-containing HERVs normalized and log2 transformed (vst) using DESeq2. The number of HERV loci containing each epitope is represented on top (HERV loci). The number of samples for the corresponding tissues is indicated in the legend (n). Expression levels in ovarian cancer were compared with each tissue for all three peptides. (***log2FC>1 and p value<0.01, **log2FC>0.75 and p value<0.01, *log2FC>0.5 and p value<0.01, Wilcoxon test). FC, fold change; FDR, false discovery rate; HERVs, human endogenous retroviruses; ORF, open reading frame; RNAseq, RNA-sequencing; TCGA, The Cancer Genome Atlas.
To determine whether this transcriptomic expression is associated with protein translation, we performed IHC experiments using a HERV-K/HML-2 Gag antibody on TMA of ovarian cancer samples (online supplemental table 2). Staining intensities were ranked as high, moderate or negative. A Gag staining was observed in 20 of the 40 (50%) tested samples. Six (15%) ovarian tumors were ranked as high, 14 (35%) as moderate and 20 (50%) were negative (figure 2A). No Gag staining was observed in normal ovarian tissue samples (n=5). Of note, no association was found between Gag protein expression and immune cell infiltration assessed by immunofluorescence (online supplemental figure 3A,B). These results suggest that the overexpression of HERV-K RNA observed in ovarian cancer is sufficient to induce the translation of a Gag protein detected by classical IHC in some cancer samples.
Figure 2. HML-2 antigens are selectively expressed in ovarian cancers. (A) Immunohistochemistry of HML-2 Gag antigen on tissue microarrays of ovarian cancer and healthy ovarian samples (20× at microscope and 22× for images, images processed with Halo AI software (Indica Labs, V.3.6.4134)). Representative examples of tumor and normal ovarian tissues and pie chart of HML-2 Gag expression level (low/negative, moderate and high) in the 40 tested cancer samples. (B and C) Targeted immunopeptidomics analysis (Max Rec_Ver2, Complete Omics Inc.) of FLQ and RLI presented on HLA molecules on the surface of the OVCAR3 (B), OVCAR5 (C) ovarian cancer cell lines. Heavy peptides are used as positive control (standard) in left panels. Endogenous targets are shown in right panels. Arrows indicate the peak of the specific epitopes and their molecular weight.
We then assessed whether the selected epitopes are efficiently presented on HLA molecules on the surface of tumor cells using a highly sensitive targeted MS-based immunopeptidomics analysis of HLA class I-eluted peptides from two HLA-A2-positive ovarian cancer cell lines from the NCI-60 panel, OVCAR3 and OVCAR5. The Valid-NEO method, which uses heavy isotope-labeled peptides to accurately characterize the presence of endogenous peptides by MS,35 was applied and optimized for FLQ and RLI, and allowed the detection of both peptides with a high confidence (figure 2B and C). A targeted Valid-NEO method for YAM analysis could not be optimized from a first series of experiments; therefore, the presence of YAM-HLA complexes on OVCAR3 and OVCAR5 could not be confirmed. To validate the tumor selectivity of these epitopes, peptide-HLA complexes were also investigated in five HLA-A2-positive primary normal cells from a selection of critical tissues: kidney proximal tubules (NhKPT), cardiomyocytes (HCM), astrocytes (HA), keratinocytes (NHEK) and bronchial epithelial (HBEpC) (online supplemental figure 4A). By using the same Valid-NEO method and MS sensitivity, no epitope-HLA complex was detected on any of the tested primary normal cells (online supplemental figure 4B).
T cells specific to HERV-K epitopes exist in patients with ovarian cancer and can be stimulated in vitro
To assess the immunogenicity of these HERV-derived epitopes in patients with ovarian cancer, we first investigated the presence of specific T cells among TILs from ovarian tumors (online supplemental table 3 and figure 5). Due to the limited number of cells obtained from tumor biopsies, we polyclonally expanded TILs in vitro without any addition of peptide before the detection of HERV-specific T cells by multimer staining in flow cytometry (online supplemental figure 5A). Multimer-positive T cells for at least one HERV epitope were detected at low frequencies in 10 of the 13 (77%) samples analyzed (online supplemental figure B,C and E), while no staining was observed on T cells from healthy HLA-A2-positive donors used as controls (online supplemental figure 5D,E).
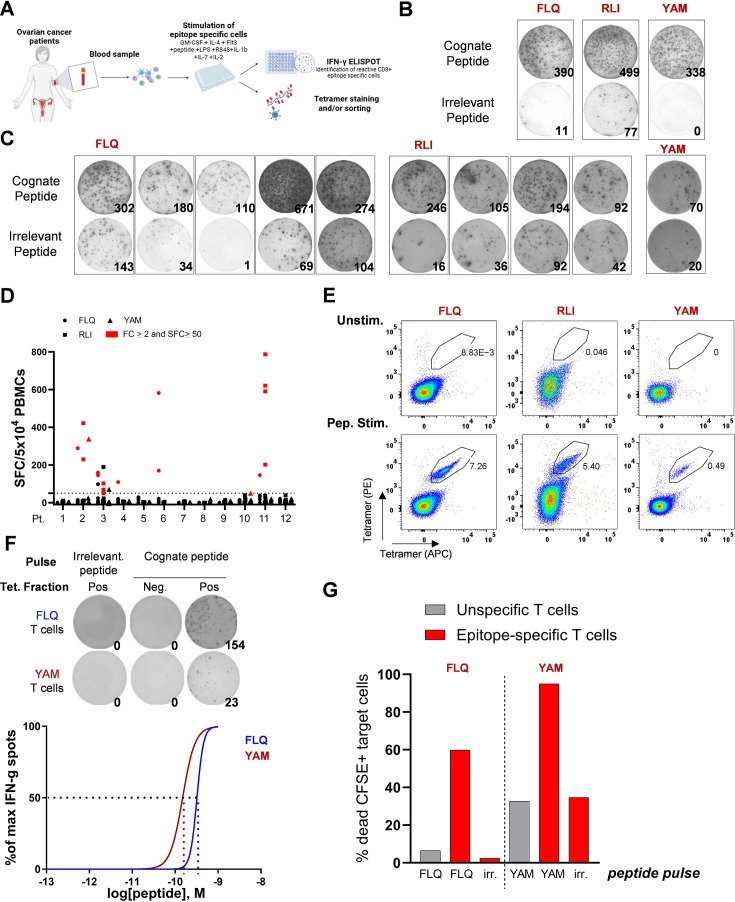
To further validate that these epitopes can induce specific and functional T cells in patients with ovarian cancer, we stimulated in vitro T cells from 12 HLA-A2-positive patients using a protocol of stimulation optimized for a low number of PBMCs (figure 3A, details in the Material and methods section). IFNγ ELISpot assay showed that epitope-specific T cells could be induced in 6 of the 12 patient samples (FLQ-specific, RLI-specific and YAM-specific T cells observed in five, three and two patient samples, respectively) (figure 3B–D). Multimer staining confirmed the presence of epitope-specific T cells (figure 3E), which could be amplified on feeder cells after immunomagnetic sorting for FLQ and YAM (online supplemental figure 6 A,B). ELISpot assays showed specific IFNγ secretion after restimulation of these selected T cells with T2 cells pulsed with the cognate peptide but not with an irrelevant peptide used as a control. These epitope-specific T cells were characterized by a high functional avidity, with an EC50 calculated at 3.2×10–10 and 1.5×10–10 M for FLQ and YAM-specific T cells, respectively (figure 3F). These HERV-K-specific T cells induced a specific killing of the T2 cells pulsed with the cognate epitope (figure 3G). Altogether, these results showed that stimulation with these HERV epitopes can induce specific and functional CD8+ T cells from patients with ovarian cancer.
Figure 3. Generation of functional epitope-specific CD8+ T cells from PBMCs of patients with ovarian cancer. (A) Schematic representation of immunogenicity assay used to evaluate HML-2 antigen-specific T cell responses. Patient (Pt.) PBMCs were expanded for 14 days in the presence of HERV epitopes before tetramer staining and IFN-γ ELISpot. (B and C) Representative IFN-γ ELISpot images for Pt. 2 (B) or for selected responsive patients (C). Expanded patient PBMCs were co-cultured with peptide-pulsed T2 cells before IFN-γ ELISpot. (D) Summary of ELISpot data (n=12). The selected wells with FLQ (circle) RLI (square) or YAM (triangle) reactive T cells are depicted in red. FC, fold change; SFC, spot forming cells. (E) Representative FACS plots of tetramer staining of PBMCs non-stimulated (Unstim, upper quadrants) or stimulated with the HERV peptides (Pep. Stim, bottom quadrants) in the same culture conditions. (F) Functional avidity assay. Top: Representative IFN-γ ELISpot of tetramer-positive and tetramer-negative sorted T cells cocultured with T2 cells pulsed with irrelevant or cognate peptide (10−9 M). Bottom: Functional avidity of FLQ-specific and YAM-specific CD8+ T cells, calculated as non-linear fit of normalized IFN-γ production. EC50 are represented by the interpolation of the dashed lines with the X-axis. 10−9 to 10−13 M range was used for the cognate peptide T2 pulsing. Neg, negative; Pos, positive; Tet, tetramer. (G) Quantification of unspecific or FLQ-specific and YAM-specific T cell-induced cell death of CFSE+T2 cells pulsed with cognate or irrelevant peptide (10−6 M). APC, allophycocyanin; CFSE, carboxyfluorescein succinimidyl ester; EC50, half-maximal cytokine response; Flt3, FMS-like tyrosine kinase 3; GM-CSF, granulocyte-macrophage colony-stimulating factor; HERVs, human endogenous retroviruses; IL, interleukin; LPS, lipopolysaccharide; IFN, interferon; PBMC, peripheral blood mononuclear cell; PE, phycoerythrin.
HERV-specific CD8+ T cells kill ovarian cancer cells while sparing normal cells
Next, we assessed the antitumor activity of HERV-K-specific T cells. CD8+ T cells from HLA-A2-positive healthy donors were stimulated in vitro using autologous dendritic cells pulsed with the cognate peptide and further amplified after immunomagnetic sorting. This approach generated a population of specific T cells containing a predominant clone, as described in our previous study19 (online supplemental figure 6A-C). As for the T cells derived from patients, these HERV-specific T cells were characterized by a high functional avidity, with IFNγ secretion still detected at peptide concentrations below 10–10 M, in line with an EC50 estimated at 1.6×10–10, 1.5×10–11 and 1.9×10–10 M for FLQ, RLI and YAM-specific T cells, respectively (online supplemental figure 6D). We then assessed the capacity of these HERV-specific CD8+ T cells to recognize and kill the ovarian cancer cell line OVCAR3. Tumor cell death was monitored in real time using the xCELLigence RTCA eSight technology. T cell clones induced a specific killing of OVCAR3 cells, as shown by the rapid decrease in the normalized cell index (NCI) and the corresponding increase in the percentage of cell lysis, whereas non-specific T cells (multimer-negative fractions) did not reduce the NCI. The effect was inhibited by an anti-HLA class I antibody, confirming that the observed cytotoxicity is HLA class I-dependent (figure 4A and online supplemental figure 6E,F). Annexin V staining was added to the medium and image analysis was performed alongside the impedance analysis to further validate the results. Imaging results showed a time-dependent and HLA-restricted increase in annexin V staining when ovarian cancer cells were co-cultured with HERV epitope-specific CD8+ T cells but not control T cells (figure 4B and C). We then confirmed that these HERV-specific T cells can kill other HLA-A2-positive ovarian cancer cell lines expressing HERV-K/HML-2 Gag, such as OV90 or HLA-A2-transduced SKOV3 (online supplemental figure 7, figure 4D). Finally, FLQ-specific T cells derived from a patient with ovarian cancer (figure 3E–G) were co-cultured with the ovarian cancer cell lines OVCAR3 and HLA-A2-transduced SKOV3. As shown in figure 4E, a specific killing of tumor cells was observed, validating the antitumor properties of HERV-specific T cells from patients.
Figure 4. HML-2 epitope-specific CD8+ T cells kill ovarian tumor cells in vitro. (A) Representative plots of tumor cell lysis kinetics of OVCAR3 cells in co-culture with FLQ- (left quadrant, E:T ratio 2:1), RLI- (central quadrant, E:T ratio 5:1) and YAM- (right left quadrant, E:T ratio 5:1) specific CD8+ T cells, or the corresponding tetramer-negative fractions (unspecific) assessed by measuring the cell index with the Xcelligence system. Results are presented as mean±SD of technical triplicates of a representative experiment of n=3 for FLQ and YAM, n=2 for RLI and YAM. (B) Representative 10× images of co-cultures of FLQ-T cells with OVCAR3 presented in A and C at 0 hour and 24 hours. Annexin V+ dead cells are depicted in green. (C) Cell death quantification of annexin V+ OVCAR3 cells (green object/image) at 0, 6, 24 and 48 hours after the addition of FLQ-specific, RLI-specific and YAM-specific CD8+ T cells or unspecific T cells. Results are expressed as mean±SD of two different images. (D and E) Maximal cell lysis of ovarian cancer cell lines (OV90, SKOV3) cocultured with epitope-specific T cells or the corresponding tetramer-negative fraction (unspecific) from healthy donors (D) or ovarian cancer patients (E). E:T ratio 5:1. In C, D and E, co-cultures with epitope-specific T cells are depicted in red and co-cultures with unspecific T cells are depicted in blue. In C, target cells alone are depicted in black, and the addition of the anti-HLA antibody to co-cultures with epitope-specific T cells is depicted with a red dotted line. E:T, effector:target; HLA, human leukocyte antigen.
We next analyzed the potential cytotoxicity of the specific T cell clones toward the panel of HLA-A2 primary normal cells derived from normal critical tissues. In agreement with the absence of epitope presentation on HLA molecules, we observed no significant effect of HERV-specific T cells on normal cells, confirming the tumor-selectivity of these epitopes (figure 5 and online supplemental figure 8).
Figure 5. HERV epitope-specific T cells are not cytotoxic toward healthy primary cells. (A) Viability of normal primary cells (HCM, NHEK, NHEK, HA, NHKpT, HBEpC) alone or co-cultured with unspecific T cells (negative tetramer fraction) or FLQ-specific, RLI-specific and YAM-specific T cells (5:1 effector to target cell ratio) monitored by normalized cell index through xCELLigence. OVCAR3 was used as a positive control. Data not interpretable for NHEK co-cultured with RLI-specific T cells (non-specific decrease of impedance). Results are mean±SD of technical triplicates. HA, human astrocytes; HBEpC, human bronchial epithelial cells; HERVs, human endogenous retroviruses; HCM, human cardiac myocytes; HLA, human leukocyte antigen; NCI, Normalized Cell Index; NHEK, normal human epithelial keratinocytes; NHKpT, normal human kidney proximal tubule cells.
In vivo antitumor efficacy of HERV-specific T cells
We then evaluated the in vivo antitumor efficacy of HERV-specific CD8+ T cells. We used the avian embryo tumor model that consists of implanting human cancer cells within selected embryonic tissues, leading to the establishment of 3D tumors within a few days34 36 (figure 6A). FLQ-specific CD8+ T cells co-transplanted with the ovarian cancer cell line OVCAR3 exhibited a strong antitumoral activity, inducing a highly significant decrease in tumor volume in comparison with both control groups in which tumor cells were transplanted alone or with polyclonal T cells (figure 6B and C). Indeed, a mean decrease of 63% (p<0.001) in normalized tumor volumes was observed in embryos transplanted with the epitope-specific T cells, compared with the control group transplanted with polyclonal unspecific T cells (figure 6B and online supplemental figure 9A,B). These latter results confirmed that HERV-specific T cells recognize and kill ovarian cancer cells in vivo in a 3D tumor environment.
Figure 6. HML-2 epitope-specific T cells elicit in vivo antitumor effects. (A) Schematic representation of the in vivo experiments. Effector to target (E:T) cell ratio used 5:1; E, embryonic day; HH, Hamburger-Hamilton stage. (B) Quantification of OVCAR-3 normalized tumor volumes using 3D light sheet microscopy of fluorescent target cells, after 48 hours co-engraftment with FLQ-specific CD8+ T cells (E:T=5:1). Tumor volumes are normalized against body surface area. Results are presented as mean±SD of 45 embryos (target cell alone n=13; unspecific T cells n=18; FLQ-specific CD8+ T cells n=14) ***p<0.001, one-way ANOVA Dunnett’s test. spec., specific; unspec., unspecific. (C) Representative images and volumes for three representative embryos at 48 hours of 3D views (light-sheet imaging) HH25 chick embryos co-engrafted with CFSE-labeled OVCAR3 alone (upper panels) and unspecific T cells (central panels) or FLQ-specific CD8+ T cells (lower panels). ANOVA, analysis of variance; CFSE, carboxyfluorescein succinimidyl ester; 3D, three-dimensional; V, volume.
Discussion
HERVs are considered to be major contributors of innate immune priming. Like exogenous viruses, they activate antiviral responses triggered by endogenous nucleic acids, a process called “viral mimicry”. This influences the immune landscape of the tumor microenvironment by increasing tumor antigen presentation and T-cell activation. Furthermore, it has been shown that HERVs can trigger an adaptive immune response through B and T cell epitopes and thus represent a source of tumor antigens both in solid tumors and hematological malignancies.9 37
HERV expression has previously been reported in ovarian cancer. Pioneering works in the field were published by Wang-Johanning et al, showing the presence of HERV-K Env protein in ovarian cancer and the possibility of generating T cells with cytotoxicity against tumor cells using dendritic cells transfected with Env.13 38 Another paper recently published confirmed the expression of HERVs in ovarian cancer and proposed a HERV prognostic score associated with survival and correlated with T-cell infiltration.39 Consistent with these results, we demonstrated here that HERV-K-derived epitopes are selectively expressed and presented on HLA molecules on the surface of ovarian tumor cells, but not of normal cells from critical tissues. We also validated the cytotoxicity of high-avidity HERV-specific T cells against ovarian cancer cells, both in vitro and in vivo. New studies are expected to clarify the immune impact of other HERV families in ovarian cancer and potentially define additional epitopes of interest.
The transcriptomic quantification of this type of epitopes is challenging because the sequence of each epitope can be found in independent transcripts from different HERV loci. In this work, we used an approach based on the sum of all the transcripts containing a potential ORF encoding the epitope, as previously described in a study by Cuevas et al.40 This method can be considered one of the most accurate to quantify HERV expression at the family level.41 Consistently, we confirmed that epitope-containing HERVs are significantly overexpressed in ovarian cancer compared with normal tissues. Importantly, our study highlighted the bias existing when directly comparing datasets from different sources and databases (batch effect). Indeed, HERV counts are systematically higher in GTEx than in TCGA for comparable tissues. As a consequence, we performed a batch correction using ComBat29 for each tissue available in both TCGA and GTEx databases, as previously reported in a study by Wang et al.30 We used TCGA as the reference batch, so that overall expression values for each corresponding tissue from GTEx aligned with TCGA samples. After pooling the corrected batch samples, we added expression values from ovary tumor samples to compare with normal tissues. These results confirmed that the HERVs containing the RNA sequences coding for the three peptides are significantly overexpressed in ovarian cancer compared with normal tissues.
Importantly, we also showed that the overexpression of epitope-containing HERVs at the transcriptomic level is associated with the presence of HERV-K/HML-2 Gag protein in some ovarian cancer samples and with a selective epitope presentation on ovarian cancer cell lines. This supports our hypothesis that messenger RNA expression of HERVs is too low in normal tissues to achieve translation, whereas their overexpression in tumor cells is sufficient to generate epitopes efficiently presented to the immune system. Because the presence of the epitope is ultimately the main factor required for the activity of specific T cells, it is important to extend the immunopeptidomics analysis to tumor biopsies in the clinic.35
Due to their viral origin, HERV-derived epitopes are expected to be immunogenic. As observed in our previous work in triple-negative breast cancer,19 multimer-positive T cells were also detected in TILs from ovarian cancer samples. Similar results have been reported for tumor epitopes derived from mutations or cancer germline antigens.42 Unfortunately, the low number of these T cells did not allow us to select and expand them to confirm their specificity and evaluate their functionality. Furthermore, the initial step of polyclonal expansion precluded the analysis of their phenotype to assess exhaustion markers. Dedicated translational studies should be performed to better characterize the T cells spontaneously induced by HERV antigens in patients with cancer. Nevertheless, the low frequencies of specific T cells in tumors justify the therapeutic strategy of boosting this response with a vaccine or of inducing an epitope-dependent antitumor response using T-cell-based therapies. In this context, we showed that HERV-specific T cells can be induced using T cells from patients with ovarian cancer. These T cells are characterized by a high functional avidity (of the same order as that of T cells generated from healthy donors, even if no statistical comparison can be performed with the available data) and can kill ovarian cancer cells. Confirming the tumor selectivity of these targets, we provide evidence that HERV-specific T cells kill cancer cells in an HLA-restricted manner while sparing normal cells. This is in line with the immunopeptidomics data showing that HERV epitopes are presented on tumor cells but not on normal cells.
Because these epitopes are specific to humans, we developed an original in vivo model using avian embryos transplanted with ovarian tumor cells to confirm the antitumor activity of HERV-specific T cells. This model offers the possibility to use a low number of T cells and allows for the rapid establishment of the tumor with tumor-associated phenotypic features after engraftment in the embryonic tissues. The physiological relevance of the avian platform was demonstrated by the observation of similar responses to treatment when comparing clinical data and the parallel avian tumor replicates.34 36 In addition, it addresses the need to reduce animal use in the context of preclinical testing, as recently supported by regulatory agencies.43 However, the lack of long-term tumor establishment and the incomplete reconstitution of the tumor stroma are limitations that need to be taken into account. Some of them are shared with the classical subcutaneous xenografts in immunocompromised mouse models used for the preclinical testing of adoptive transfer therapies (low tumor stroma complexity, poor vascularization).
In summary, we showed that HERV-K-derived epitopes are selectively presented on tumor cells in ovarian cancer. They are immunogenic in patients and induce high-avidity CD8+ cytotoxic T cells that selectively kill ovarian tumor cells in vitro and in vivo. These data provide the preclinical rationale for developing T-cell-based therapies against these new targets, such as vaccines, adoptive T-cell therapies or T-cell-recruiting bispecific antibodies.
Supplementary material
Acknowledgements
We wish to thank the staff of the core facilities at the Cancer Research Center of Lyon (CRCL) for technical assistance, the BRC of the CLB, the Lyon Immunotherapy for Cancer Lab (LICL) of the CRCL for his assistance for mIF staining of immune cell infiltrates, as well as N Gadot and the research anatomopathology platform of the CLB. We thank T Andrieu, P Battiston-Montagne, and A Jambon for assistance in flow cytometry in the CRCL Cytometry platform. We thank B Manship for the editing review. The illustrations were created with https://BioRender.com. As citations were limited, we were unable to cite all available studies, and so apologize to any authors who feel their studies were not adequately represented in our accounting.
Footnotes
Funding: This project has been carried out thanks to the support of the Cancéropôle CLARA and La Région Auvergne-Rhône-Alpes as part of the Proof-of-Concept program (project)
Provenance and peer review: Not commissioned; externally peer reviewed.
Patient consent for publication: Not applicable.
Ethics approval: Not applicable.
Data availability statement
All data relevant to the study are included in the article or uploaded as supplementary information.
References
- 1.Torre LA, Trabert B, DeSantis CE, et al. Ovarian cancer statistics, 2018. CA Cancer J Clin. 2018;68:284–96. doi: 10.3322/caac.21456. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Narod S. Can advanced-stage ovarian cancer be cured? Nat Rev Clin Oncol. 2016;13:255–61. doi: 10.1038/nrclinonc.2015.224. [DOI] [PubMed] [Google Scholar]
- 3.Matulonis UA, Shapira-Frommer R, Santin AD, et al. Antitumor activity and safety of pembrolizumab in patients with advanced recurrent ovarian cancer: results from the phase II KEYNOTE-100 study. Ann Oncol. 2019;30:1080–7. doi: 10.1093/annonc/mdz135. [DOI] [PubMed] [Google Scholar]
- 4.Moore KN, Bookman M, Sehouli J, et al. Atezolizumab, Bevacizumab, and Chemotherapy for Newly Diagnosed Stage III or IV Ovarian Cancer: Placebo-Controlled Randomized Phase III Trial (IMagyn050/GOG 3015/ENGOT-OV39) JCO. 2021;39:1842–55. doi: 10.1200/JCO.21.00306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bonaventura P, Shekarian T, Alcazer V, et al. Cold Tumors: A Therapeutic Challenge for Immunotherapy. Front Immunol. 2019;10:168. doi: 10.3389/fimmu.2019.00168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Alcazer V, Bonaventura P, Tonon L, et al. Neoepitopes-based vaccines: challenges and perspectives. Eur J Cancer. 2019;108:55–60. doi: 10.1016/j.ejca.2018.12.011. [DOI] [PubMed] [Google Scholar]
- 7.Yarchoan M, Johnson BA, Lutz ER, et al. Targeting neoantigens to augment antitumour immunity. Nat Rev Cancer. 2017;17:209–22. doi: 10.1038/nrc.2016.154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Attermann AS, Bjerregaard A-M, Saini SK, et al. Human endogenous retroviruses and their implication for immunotherapeutics of cancer. Ann Oncol. 2018;29:2183–91. doi: 10.1093/annonc/mdy413. [DOI] [PubMed] [Google Scholar]
- 9.Kassiotis G. The Immunological Conundrum of Endogenous Retroelements. Annu Rev Immunol. 2023;41:99–125. doi: 10.1146/annurev-immunol-101721-033341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Liang Y, Qu X, Shah NM, et al. Towards targeting transposable elements for cancer therapy. Nat Rev Cancer. 2024;24:123–40. doi: 10.1038/s41568-023-00653-8. [DOI] [PubMed] [Google Scholar]
- 11.Burn A, Roy F, Freeman M, et al. Widespread expression of the ancient HERV-K (HML-2) provirus group in normal human tissues. PLoS Biol. 2022;20:e3001826. doi: 10.1371/journal.pbio.3001826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wang-Johanning F, Rycaj K, Plummer JB, et al. Immunotherapeutic potential of anti-human endogenous retrovirus-K envelope protein antibodies in targeting breast tumors. J Natl Cancer Inst. 2012;104:189–210. doi: 10.1093/jnci/djr540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Rycaj K, Plummer JB, Yin B, et al. Cytotoxicity of Human Endogenous Retrovirus K–Specific T Cells toward Autologous Ovarian Cancer Cells. Clin Cancer Res. 2015;21:471–83. doi: 10.1158/1078-0432.CCR-14-0388. [DOI] [PubMed] [Google Scholar]
- 14.Reis BS, Jungbluth AA, Frosina D, et al. Prostate Cancer Progression Correlates with Increased Humoral Immune Response to a Human Endogenous Retrovirus GAG Protein. Clin Cancer Res. 2013;19:6112–25. doi: 10.1158/1078-0432.CCR-12-3580. [DOI] [PubMed] [Google Scholar]
- 15.Smith CC, Beckermann KE, Bortone DS, et al. Endogenous retroviral signatures predict immunotherapy response in clear cell renal cell carcinoma. J Clin Invest. 2018;128:4804–20. doi: 10.1172/JCI121476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Krishnamurthy J, Rabinovich BA, Mi T, et al. Genetic Engineering of T Cells to Target HERV-K, an Ancient Retrovirus on Melanoma. Clin Cancer Res. 2015;21:3241–51. doi: 10.1158/1078-0432.CCR-14-3197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Shah AH, Rivas SR, Doucet-O’Hare TT, et al. Human endogenous retrovirus K contributes to a stem cell niche in glioblastoma. J Clin Invest. 2023;133:e167929. doi: 10.1172/JCI167929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Depil S, Roche C, Dussart P, et al. Expression of a human endogenous retrovirus, HERV-K, in the blood cells of leukemia patients. Leukemia. 2002;16:254–9. doi: 10.1038/sj.leu.2402355. [DOI] [PubMed] [Google Scholar]
- 19.Bonaventura P, Alcazer V, Mutez V, et al. Identification of shared tumor epitopes from endogenous retroviruses inducing high-avidity cytotoxic T cells for cancer immunotherapy. Sci Adv. 2022;8:eabj3671. doi: 10.1126/sciadv.abj3671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Alcazer V, Bonaventura P, Tonon L, et al. HERVs characterize normal and leukemia stem cells and represent a source of shared epitopes for cancer immunotherapy. Am J Hematol. 2022;97:1200–14. doi: 10.1002/ajh.26647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Wang H, Liu J, Yang J, et al. A novel tumor mutational burden-based risk model predicts prognosis and correlates with immune infiltration in ovarian cancer. Front Immunol. 2022;13:943389. doi: 10.3389/fimmu.2022.943389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Bendall ML, de Mulder M, Iñiguez LP, et al. Telescope: Characterization of the retrotranscriptome by accurate estimation of transposable element expression. PLoS Comput Biol. 2019;15:e1006453. doi: 10.1371/journal.pcbi.1006453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Pertea G, Pertea M. GFF Utilities: GffRead and GffCompare. F1000Res. 2020;9:ISCB Comm J-304. doi: 10.12688/f1000research.23297.2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Jeong H-H, Yalamanchili HK, Guo C, et al. An ultra-fast and scalable quantification pipeline for transposable elements from next generation sequencing data. Pac Symp Biocomput . 2018;23:168–79. [PubMed] [Google Scholar]
- 25.Kong Y, Rose CM, Cass AA, et al. Transposable element expression in tumors is associated with immune infiltration and increased antigenicity. Nat Commun. 2019;10:5228. doi: 10.1038/s41467-019-13035-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Patro R, Duggal G, Love MI, et al. Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods. 2017;14:417–9. doi: 10.1038/nmeth.4197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Soneson C, Love MI, Robinson MD. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Res. 2015;4:1521. doi: 10.12688/f1000research.7563.2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550. doi: 10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Johnson WE, Li C, Rabinovic A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics. 2007;8:118–27. doi: 10.1093/biostatistics/kxj037. [DOI] [PubMed] [Google Scholar]
- 30.Wang Q, Armenia J, Zhang C, et al. Unifying cancer and normal RNA sequencing data from different sources. Sci Data. 2018;5:180061. doi: 10.1038/sdata.2018.61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Wickham H. Ggplot2. Cham: Springer International Publishing; 2016. http://link.springer.com/10.1007/978-3-319-24277-4 Available. [Google Scholar]
- 32.Douglass J, Hsiue E-C, Mog BJ, et al. Bispecific antibodies targeting mutant RAS neoantigens. Sci Immunol. 2021;6:eabd5515. doi: 10.1126/sciimmunol.abd5515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Roudko V, Bozkus CC, Orfanelli T, et al. Shared Immunogenic Poly-Epitope Frameshift Mutations in Microsatellite Unstable Tumors. Cell. 2020;183:1634–49. doi: 10.1016/j.cell.2020.11.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Jarrosson L, Dalle S, Costechareyre C, et al. An in vivo avian model of human melanoma to perform rapid and robust preclinical studies. EMBO Mol Med. 2023;15:e16629. doi: 10.15252/emmm.202216629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Terai YL, Huang C, Wang B, et al. Valid-NEO: A Multi-Omics Platform for Neoantigen Detection and Quantification from Limited Clinical Samples. Cancers (Basel) 2022;14:1243. doi: 10.3390/cancers14051243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Jarrosson L, Costechareyre C, Gallix F, et al. An avian embryo patient-derived xenograft model for preclinical studies of human breast cancers. iScience. 2021;24:103423. doi: 10.1016/j.isci.2021.103423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Chour M, Porteu F, Depil S, et al. Endogenous retroelements in hematological malignancies: From epigenetic dysregulation to therapeutic targeting. Am J Hematol. 2025;100:116–30. doi: 10.1002/ajh.27501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Wang-Johanning F, Liu J, Rycaj K, et al. Expression of multiple human endogenous retrovirus surface envelope proteins in ovarian cancer. Int J Cancer. 2007;120:81–90. doi: 10.1002/ijc.22256. [DOI] [PubMed] [Google Scholar]
- 39.Natoli M, Gallon J, Lu H, et al. Transcriptional analysis of multiple ovarian cancer cohorts reveals prognostic and immunomodulatory consequences of ERV expression. J Immunother Cancer. 2021;9:e001519. doi: 10.1136/jitc-2020-001519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Cuevas MVR, Hardy M-P, Larouche J-D, et al. BamQuery: a proteogenomic tool to explore the immunopeptidome and prioritize actionable tumor antigens. Genome Biol. 2023;24:188. doi: 10.1186/s13059-023-03029-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kitsou K, Katzourakis A, Magiorkinis G. Limitations of current high-throughput sequencing technologies lead to biased expression estimates of endogenous retroviral elements. NAR Genomics and Bioinformatics. 2024;6:lqae081. doi: 10.1093/nargab/lqae081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Bobisse S, Genolet R, Roberti A, et al. Sensitive and frequent identification of high avidity neo-epitope specific CD8 + T cells in immunotherapy-naive ovarian cancer. Nat Commun. 2018;9:1092. doi: 10.1038/s41467-018-03301-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Wadman M. FDA no longer has to require animal testing for new drugs. Science. 2023;379:127–8. doi: 10.1126/science.adg6276. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data relevant to the study are included in the article or uploaded as supplementary information.