Abstract
Aims
Vascular smooth muscle cells (VSMCs) accumulate in atherosclerotic plaques and exhibit remarkable phenotypic plasticity, contributing to both plaque growth and stability. The plaque-stabilizing fibrous cap is rich in VSMC-derived cells, yet the cellular transitions and regulatory mechanisms governing fibrous cap formation remain unclear. Here, we aimed to identify the VSMC phenotypic transitions associated with this critical process.
Methods and results
Mapping of lineage-traced VSMCs during plaque development revealed investment of VSMCs prior to fibrous cap formation. Using single-cell RNA-sequencing (scRNA-seq) profiles of lineage-traced VSMCs from atherosclerotic and acutely injured mouse arteries, we identified a disease-specific VSMC state co-expressing contractile genes with extracellular matrix (ECM) components (including fibrillar collagens and elastin) and NOTCH3, which are associated with fibrous cap formation. Computational trajectory analysis predicted that this proposed fibrous cap-related VSMC (fcVSMC) state arises from a previously described plastic, intermediate VSMC population expressing SCA1 and VCAM1. Clonal analysis further showed that NOTCH3+ fcVSMCs derive from intermediate VSMCs in both atherosclerosis and an acute vascular injury model, suggesting a conserved disease-relevant mechanism. The fcVSMCs were enriched in plaque fibrous caps compared to lesion cores, consistent with a role in fibrous cap formation. By combining scRNA-seq trajectory analysis and spatial transcriptomics of human atherosclerotic plaques, we identified protease-activated receptor-1 (PAR1) as a candidate regulator of fcVSMC generation. PAR1 was expressed by VSMCs in human plaque fibrous caps and PAR1 activation by thrombin induced expression of contractile genes and ECM components associated with the fcVSMC state in human VSMCs.
Conclusion
Our findings identify a VSMC transition linked to fibrous cap formation in atherosclerosis and show this is modelled by vascular injury. We identify VSMC-expressed PAR1 as a potential therapeutic target for promoting plaque stability by driving the transition to the matrix-producing, fibrous cap-associated VSMC state.
Keywords: Vascular smooth muscle cells, Atherosclerosis, Fibrous cap, Phenotypic switching, Protease-activated receptor-1
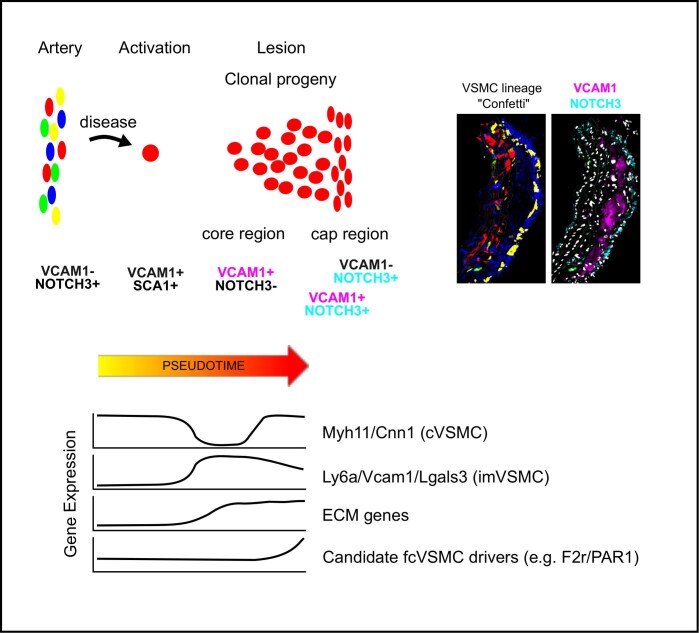
Graphical Abstract
Graphical Abstract.
Time of primary review: 72 days
1. Introduction
Vascular smooth muscle cell (VSMC) accumulation is a hallmark of atherosclerosis1,2 and neointima formation following vessel occlusion and injury.3,4 VSMCs are quiescent and express components of the contractile machinery (such as MYH11, CNN1, and ACTA2) to control blood pressure and flow in healthy arteries.1 However, these cells retain remarkable plasticity and dedifferentiate in response to injury or inflammation. This involves down-regulating the contractile apparatus and increasing migration, proliferation, and extracellular matrix (ECM) production. In atherosclerotic plaques, VSMCs acquire a range of different phenotypes1,5,6 and generate both the plaque-stabilizing fibrous cap and cells in the lesion core that have characteristics of mesenchymal cells, phagocytes, and osteochondrocytes, which may contribute to lesion destabilization.2,6 Delineating the governance of these distinct VSMC-derived cells in disease will therefore aid understanding of the functional role of these cells in disease.
VSMC-contribution to atherosclerotic plaques and injury-induced neointimal lesions in disease models is oligoclonal.7–10 Very few pre-existing VSMCs give rise to all lesional VSMCs, and the range of plaque VSMC phenotypes can be generate from the clonal progeny of individual cells.7,11 In mice, VSMCs with reduced levels of contractile markers that express stem cell antigen 1 (SCA1, encoded by Ly6a) and Vcam1 have been suggested to represent a plastic VSMC state capable of giving rise to diverse plaque VSMC phenotypes.11,12 Humans, which do not have a SCA1/Ly6a homologue, also have VSMC populations with an equivalent transcriptional profile to mouse SCA1+ VSMCs.13 This plastic VSMC state is rarely detected in healthy arteries and is induced upon VSMC de-differentiation in response to disease stimuli.11,13 In atherosclerosis, such cells showed substantial overlap in the transcriptional signature of ‘activated’ VSMC-derived cells expressing osteochondrocyte and phagocyte markers.11 Computational inference of VSMC transdifferentiation pathways, functional in vitro experiments with isolated SCA1-expressing VSMCs and use of a dual lineage tracing system have all provided substantial evidence that VSMCs co-expressing SCA1, VCAM1, and LGALS3 act as intermediates and give rise to the diverse VSMC states observed in atherosclerotic plaques.12,14,15 Accordingly, this ‘intermediate modulated’ VSMC (imVSMC) population could represent a novel therapeutic target, whereby its disease-induced transdifferentiation to diverse cell states could be biased towards those states which are beneficial to disease outcome.
The fibrous cap structure is imperative to mitigating plaque rupture and downstream cardiovascular events, and is rich in VSMCs expressing typical contractile VSMC markers, as well as ECM components such as fibrillar collagens and elastin.1,16,17 Previous work has highlighted several candidate factors involved in promoting the formation of a VSMC-rich fibrous cap, such as PDGFRB,18 TCF21,19 and retinoic acid.12 Compelling evidence suggests that although loss of NOTCH3 signalling characterises VSMC lesion investment, subsequent reactivation is critical in the formation of a fibrous cap with contractile marker-expressing VSMCs.20 However, the VSMC states underpinning fibrous cap formation and how cap formation is related to the imVSMC state remains to be elucidated.
Here, we investigated the regulation of VSMCs underlying fibrous cap formation in atherosclerosis. By combining scRNA-seq analysis, and immunostaining in disease models with clonal VSMC lineage tracing we find a transcriptional VSMC state with characteristics of fibrous cap cells and show evidence that this is generated from the intermediate VSMC state. Interestingly, this transition was detected in both atherosclerotic and acutely injured arteries, suggesting importance for general VSMC regulation. We identify candidate regulators of this fibrous cap-associated transition and highlight VSMC-expressed thrombin receptor as a candidate driver of fibrous cap formation.
2. Methods
Experimental details are provided as Supplementary Information.
2.1. Animals and procedures
Experiments, licenced under UK Home Office regulations (P452C9546 and PP7513347), were approved by the University of Cambridge Ethical Reviews Committee. Tamoxifen-induced VSMC lineage-labelling of Myh11-CreERT2; Rosa26-Confetti (Myh11Confetti) and Myh11-CreERT2; Rosa26-eYFP (Myh11eYFP) animals was done before disease induction. Animals received pre-operative analgesic (∼0.1 mg/kg body weight, buprenorphine), were anaesthetized with 2.5%–3% isoflurane by inhalation (1.5 L/min) and the left carotid artery ligated with a 6–0 silk suture. Myh11Confetti; Apoe−/− mice were fed a high fat diet (HFD, Special Diets Services; 21% fat, 0.2% cholesterol). Animals were euthanized by cervical dislocation or CO2 asphyxiation.
2.2. Human tissue
Human tissue was acquired under informed consent using protocols approved by the Cambridge or Huntingdon Research Ethical Committee, conforming to the principles of the Declaration of Helsinki. Anonymized samples were obtained from patients undergoing carotid endarterectomy, cardiac transplant, valve replacement or by donation after circulatory death. Human arterial sections were co-stained for αSMA (DAKO, M0851, 1:400) and PAR1 (Novus Biologicals, N2–11, 1:200). A plaque-containing aortic sample from a 54-year old male organ donor was analysed by spatial transcriptomics (see Supplementary Information).
2.3. Arterial VSMC analysis
Plaque VSMC infiltration was analysed using 100 μm sections from the ascending and descending aorta, aortic arch and carotid arteries of 7 animals (4 at week 6.5 and 3 at week 11), previously used to assess VSMC clonality.13 A total of 150 unique plaques were analysed (see Supplementary material online, Table S1, plaques with suboptimal clearing were excluded). Lesion classification was based on DAPI signals, and Confetti signals subsequently used to score VSMC location, number, and clonality. Thin (14 μm) sections were stained with antibodies to NOTCH3 (Abcam, ab23426, 10 μg/mL), VCAM1 (BioLegend, 105702, 10 μg/mL) or isotype control antibodies, DAPI-stained (1 μg/mL), and mounted in RapiClear 1.52. VCAM1 and/or NOTCH3 expression by individual Confetti+ VSMCs was assessed by Z-plane navigation, and scored for 16 plaques (26 clones) after 11 weeks HFD (n = 3, Supplementary material online, Table S1) and vessels from control or injured animals (three per time point).
2.4. Cell culture and molecular analysis
Human VSMCs (hVSMCs), cultured in SMC-GM2 medium (Promocell) supplemented with 100 U/mL penicillin, 100 µg/mL streptomycin, were analysed at passages 2–10 and treated with thrombin (0.5, 1, or 2 U/mL) and Factor Xa (2.5 μg/mL) in serum-free media. Cells were transfected with 50 nM human PAR1-specific siRNA (ON-TARGETplus® SMART Pool, Dharmacon, L-005094–00-0005) or non-targeting control siRNA (Dharmacon, D-001810-10-05) for 72 h, before thrombin treatment. Western blotting was done using primary antibodies for PAR1 (Santa Cruz, ATAP2, 1:2000), and GAPDH (Cell Signaling Technologies, 2118, 1:4000). Primer sequences for reverse transcription, quantitative PCR (RT-qPCR) are listed in Supplementary material online, Table S2.
Bulk RNA-seq was conducted using female hVSMCs from three different donors, following 24-h thrombin/vehicle treatment in serum-free medium and data analysed using the limma package21 with cell line donor as a covariate.
ScRNA-seq profiles of VSMCs from mouse carotid arteries were generated as described13 after pooling single cell suspensions from 5 Myh11eYFP animals 11 days post-ligation or 2 control Myh11eYFP animals that did not undergo surgery. Gene count matrices for lineage-labelled VSMCs from a mouse atherosclerosis model12 were downloaded (GSE155513). Parameters for data processing and filtering are described in Supplementary material online, Table S3. Normalized, clustered and differential gene expression analysis was done in Seurat.22 Trajectory analysis was done using the Slingshot,23 Monocle3, or Partition-based graph abstraction24 in Scanpy25 with R-Python interfacing. Differential expression across pseudotime was performed with generalized additive modelling (GAM). Genes showing statistically significant pseudotime-dependent changes (logFC > 0.25) were clustered by Pearson correlation distance to reveal similar expression patterns. NicheNet (v2.2.0) was used in the sender-agnostic mode to identify potential ligands inducing the I-cluster 6 VSMC state.
Gene ontology (GO) analysis of genes differentially expressed between clusters, conditions, and cell states, or along trajectory path pseudotimes was carried out using gProfiler (https://biit.cs.ut.ee/gprofiler/gost) and the combined GO analysis was performed using the compareCluster function in clusterProfiler.26 Statistical significance was evaluated using cumulative hypergeometric probability testing with g:SCS multiple testing correction and 0.05 P-adj threshold. Circos plots were generated using the GOplot package (CRAN).
2.5. Statistical analysis
Statistical significance was assessed using Chi-squared (X2), Kruskal–Wallis (and Mann–Whitney between groups, with Bonferroni-correction, if P < 0.05), and cumulative hypergeometric probability testing, with g:SCS correction. Group sizes represent biologically independent samples (different donors, different mice, plaques or clones as specified), rather than technical replicates. An adjusted P-value (P-adj) <0.05 was considered statistically significant. Statistical testing information for single-cell and bulk RNA-seq dataset analysis is above and detailed in the Supplemental Information.
3. Results
3.1. Plaque VSMC infiltration leading to generation of the fibrous cap
To understand the kinetics of fibrous cap formation, we analysed developing plaques in arteries of Apoe−/− mice that were fed a HFD for 6.5 (W6.5) or 11 weeks (W11). To trace the clonal progeny of VSMCs in lesions, we combined a Myh11-driven, tamoxifen-inducible Cre recombinase with the Rosa26-Confetti reporter (Myh11Confetti), and induced stochastic expression of one of four fluorescent proteins specifically in VSMCs prior to HFD feeding. Confocal microscopy of thick (100 μm) arterial sections revealed that the proportion of lesions with longitudinally arranged nuclei adjacent to the endothelium at the luminal edge, indicating formation of a fibrous cap, increased with both time of HFD and plaque size (Figure 1A–C). This is consistent with fibrous cap development correlating with the duration of lipid exposure and plaque volume.16,27 We focused on plaques that contained lineage-labelled VSMCs (122/150). A cap was observed in most, but a substantial percentage of lesions containing Confetti+ cells (36%) did not have an obvious cap structure. Plaques lacking a fibrous cap were observed more frequently at the earlier time point (42% at W6.5 vs. 28% at W11, Figure 1C), indicating that VSMCs can enter the plaque prior to fibrous cap formation.
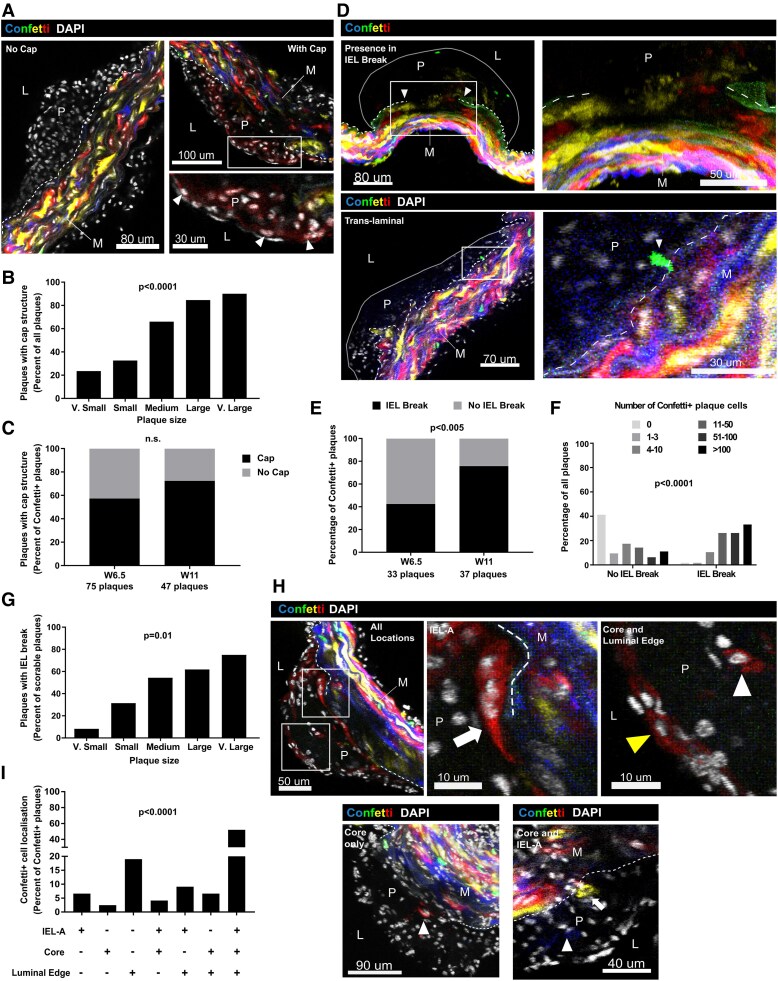
Figure 1.
VSMC infiltration in atherosclerosis occurs prior to generation of a fibrous cap. Analysis of Confetti + cells in plaques from VSMC-lineage-labelled Myh11Confetti; Apoe−/− animals fed a HFD for 6.5 or 11 weeks (W). (A) Representative confocal images (single Z-stack) of plaques without (left) and with fibrous cap structure (arrowheads, right). Magnified view of the boxed region (lower right). Scale bars = 80 μm (left), 100 μm (top right), and 30 μm (lower right). (B) Percent of all plaques with fibrous cap structure according to plaque size. n = 150 plaques; n = 17 very (v) small, 49 small, 47 medium, 27 large, 10 v. large. P < 0.001, X2, and df = 4. (C) Percent of plaques containing Confetti+ cells stratified by time of HFD feeding. n = 122 plaques; n = 75 plaques from four animals at W6.5 and n = 47 plaques from three animals at W11. P = 0.09, X2, and df = 1. (D) Representative plaques (max. projection) with Confetti+ cells at an IEL break (top) or ‘translaminal’ Confetti+ cells crossing an intact IEL (lower panels). Magnified views show boxed regions (right). Arrowheads: IEL breakpoints (top) and translaminal Confetti+ cell (lower). Scale bars = 80 μm (top-left), 50 μm (top-right), 70 μm (bottom-left), and 30 μm (bottom-right). (E) Proportion of Confetti+ cell-containing plaques with an IEL break, stratified by time of HFD. n = 70 plaques from four animals (W6.5) and 37 plaques from three animals (W11). P = 0.005, X2, and df = 1. (F) Distribution of plaques by number of Confetti+ cells per plaque, for lesions without (54 plaques) or with an IEL break (43 plaques). P < 0.0001, X2, and df = 5. (G) Percent of plaques with IEL break, stratified by plaque size (9 v. small, 29 small, 35 medium, 17 large, 7 v. large). P = 0.04, X2, and df = 4. (F, G) Plaques were excluded if IEL status could not be confidently scored. (H) Representative confocal images (top, single Z-plane; lower panels, max. projection) of plaques showing Confetti+ cells in all plaque regions (top), core only (lower left), or core and ‘IEL-adjacent’ (IEL-A, lower right). Magnified views of boxed regions show IEL-A (middle) and ‘core and luminal edge’ located Confetti+ cells (right). Confetti+ cells at the IEL-A (white arrows), Confetti+ cells at the luminal edge (yellow arrowhead), and Confetti+ cells in the plaque core (white arrowheads) are marked. (I) Distribution of plaque regions with Confetti+ cells for plaques with Confetti+ cell investment (n = 122 plaques). P < 0.0001, X2, and df = 6. (A, D, H) Confetti (CFP = blue, RFP = red, YFP = yellow, and GFP = green) and DAPI (white) signals are shown as indicated. Plaque outlines (grey line) and internal elastic lamina (IEL, dashed line) are marked. L, lumen; P, plaque, M, media.
Definitive scoring of internal elastic lamina (IEL) integrity was possible in 70 plaques with Confetti + cells, and an IEL break was observed in most of these (60%, Figure 1D and E). However, no IEL breaks were detected in 40% of lesions, and an intact IEL was most frequently observed in plaques at W6.5 (60%; Figure 1E). In two cases, we detected a translaminal lineage-labelled cell in lesions without IEL breaks (Figure 1D). This was a relatively rare event, but agree with migration of VSMCs through IEL fenestrations observed by high-resolution confocal microscopy.28 These findings suggest that VSMC plaque infiltration can occur both via cell migration through an intact IEL from the underlying media and through IEL breaks. Plaques without detectable IEL breaks tended to contain lower numbers of VSMC-lineage cells, whereas lesions with IEL breaks contained more VSMC lineage cells (Figure 1F). This suggests that VSMC investment through breaks in the IEL occurs in more advanced plaques, or that IEL breaks are secondary to VSMC investment. The frequency of IEL breaks for all (Confetti+ and Confetti−) plaques was higher at W11 (29/42, 69%) compared to W6.5 (14/52, 27%) and we observed a plaque size-dependent increase in IEL-break frequency (Figure 1G), further suggesting that IEL breaks are associated with plaque maturity.
To understand the dynamics of VSMC localization within plaques during lesion development, we scored the presence of Confetti+ cells in each of three plaque regions: ‘IEL-adjacent’ (IEL-A: plaque core, directly contacting the IEL), non-attached cells in the ‘core’ and ‘luminal edge’, which include the fibrous cap (Figure 1H). Plaques with VSMCs in all three locations were most common (63 out of 121 scorable Confetti+ lesions), although VSMC investment was limited to specific regions in 48% the plaques (Figure 1I). VSMCs were selectively located at the luminal edge in 23 plaques (19%, Figure 1I), which is in keeping with a previous study that suggested VSMCs initially migrate through the lesion via the luminal edge.10 However, lineage-labelled VSMC contribution at the luminal edge was absent in 16 lesions (13%), including 5 lesions with labelled cells selectively in the region adjacent to the IEL (IEL-A, Figure 1I). Overall, these findings suggest that, while migration via the luminal edge is prevalent, alternative routes exist and demonstrates that VSMCs are present in plaques before cap formation.
3.2. Inference of a VSMC transition that gives rise to a fibrous cap-associated cell state in vascular disease models.
While the VSMC origin of fibrous cap cells is well documented, the pathways underlying their generation are not fully understood. To capture active VSMC transitions underlying fibrous cap development, we therefore examined scRNA-seq profiles of lineage-traced VSMCs from experimental atherosclerosis at early-, mid- and late-stage (8, 16, and 22 weeks of HFD on Apoe−/− background; GSE15551312). Disease-specific VSMC states were identified by integration with profiles of VSMCs from healthy vessels of control Apoe+/+ mice. Contractile VSMC (cVSMC) markers (Myh11, Acta2, and Cnn1) were highly expressed in cell clusters found in both healthy and atherosclerotic arteries [Athero (A)-cluster 0, 1, 7, 9, and 11, Figure 2A–C; Supplementary material online, Figure S1 and Table S4]. Disease-associated states14,29 included Sox9+, Col2a1+, and Chad+ chondromyocytes (CMCs; A-cluster 2 and 10), fibroblast-like (FB-like, A-cluster 8) cells expressing high levels of Pi16, Igfbp4, and Dpep1 and a distinct cell population expressing Cd68 and other macrophage markers (MΦ-like; A-cluster 3 and 12; Figure 2A and B). We also detected activated, or intermediate modulated VSMCs (imVSMCs), characterized by expression of Ly6a, Vcam1, Tnfrsf11b, Lgals3, and ECM genes (A-cluster 4 and 5; Figure 2B, also been termed ‘SEM’,12,14 fibromyocytes,19 or pioneer cells15). A small population of proliferating (Prlf) VSMCs (Ccnd1+, Pcna+, and Mki67+) co-expressed imVSMC markers (A-cluster 13, Figure 2A and B), in keeping with previous work suggesting that the imVSMC state is associated with proliferation.12
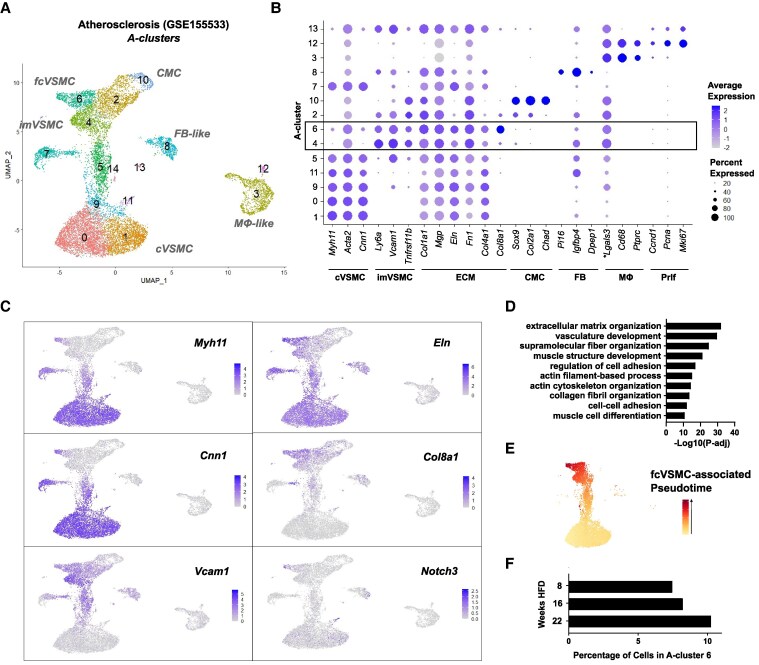
Figure 2.
Identification of a proposed fibrous cap-associated VSMC state derived from intermediate modulated VSMCs. (A) Uniform manifold approximation and projection (UMAP) showing scRNA-Seq profiles of VSMC-lineage label-positive cells from atherosclerotic arteries of Myh11-CreERt2; reporter-ZsGreen; Apoe−/− animals (GSE155513)20 and healthy control Apoe+/+ carotid arteries. Atherosclerosis cell clusters (A-clusters) and VSMC states are annotated. (B) Dot plot showing expression of VSMC-state marker genes in clusters using a grey (low) to blue (high) scale and the percentage of cells with detected marker expression for each cluster indicated by dot size. cVSMC, contractile VSMC; imVSMC, intermediate modulated VSMC; ECM, extracellular matrix; CMC, chondromyocyte; FB, fibroblast; MΦ, macrophage; Prlf, proliferation. *Lgals3 is also expressed by imVSMCs. (C) UMAP feature plots showing expression levels for cVSMC (Myh11, Cnn1), ECM (Eln, Col8a1, Col4a1), the imVSMC gene Vcam1, and Notch3 using grey (low) to blue (high) scales. (D) GO terms enriched for genes up-regulated in A-cluster 6 compared to A-cluster 4 cells. P-adj: cumulative hypergeometric probability testing with g:SCS correction. (E) UMAP showing the cells included in the inferred trajectory generating A-cluster 6 cells. Pseudotime is indicated using a yellow–red scale. (F) Proportion of VSMC-lineage cells mapping to A-cluster 6 at each HFD time point.
An unannotated disease-specific cell cluster (A-cluster 6) co-expressed contractile VSMC markers (Myh11, Acta2, and Cnn1) with several atherosclerosis-induced ECM genes (Col1a1, Mgp, Eln, Fn1, Col4a1, and Col8a1) and had reduced levels of Ly6a, Vcam1, and Tnfrsf11b compared to imVSMCs (Figure 2A–C; Supplementary material online, Figure S1). Genes that were up-regulated in A-cluster 6 cells compared to A-cluster 4/imVSMCs included Notch3 that has been shown to play an important role in fibrous cap formation,20 and were enriched for GO terms associated with contractile/muscle function (muscle structure development, actin cytoskeleton organization, and muscle cell differentiation), and genes associated with ECM production (ECM organization, collagen fibril organization; Figure 2C and D; Supplementary material online, Table S5 and S6). The co-expression of the broader contractile gene programme with a synthetic phenotype led us to hypothesize that A-cluster 6 cells were associated with the fibrous cap (referred to as fcVSMCs). Independent trajectory inference analyses suggested that imVSMCs are intermediates for fcVSMC formation (Figure 2E; Supplementary material online, Figure S1C). Furthermore, A-cluster 6 cells were present at a greater proportion at the later time point of HFD feeding (22 weeks; Figure 2F; Supplementary material online, Figure S1), aligning with fibrous cap formation occurring at later disease stages16,27 (Figure 1B and C). This is consistent with the idea that imVSMCs represent dedifferentiated VSMCs (or fibromyocytes) that migrate into lesions and give rise to the other VSMC-lineage plaque cells.
We next compared the imVSMCs-to-fcVSMC transition to ‘re-differentiation’ of VSMCs after injury, using carotid ligation surgery, which results in reproducible and acute VSMC de-differentiation, formation of a VSMC-rich intimal lesion and subsequent upregulation of contractile VSMC markers.30,31 We performed scRNA-seq on VSMC lineage+ (eYFP+) cells sorted from ligated arteries of Myh11-CreERt2/Rosa26-eYFP (Myh11eYFP) mice 11 days post-injury (DPI), when neointimal lesions are developing and VSMC re-differentiation is ongoing30,31 (Figure 3A; Supplementary material online, Figure S2).
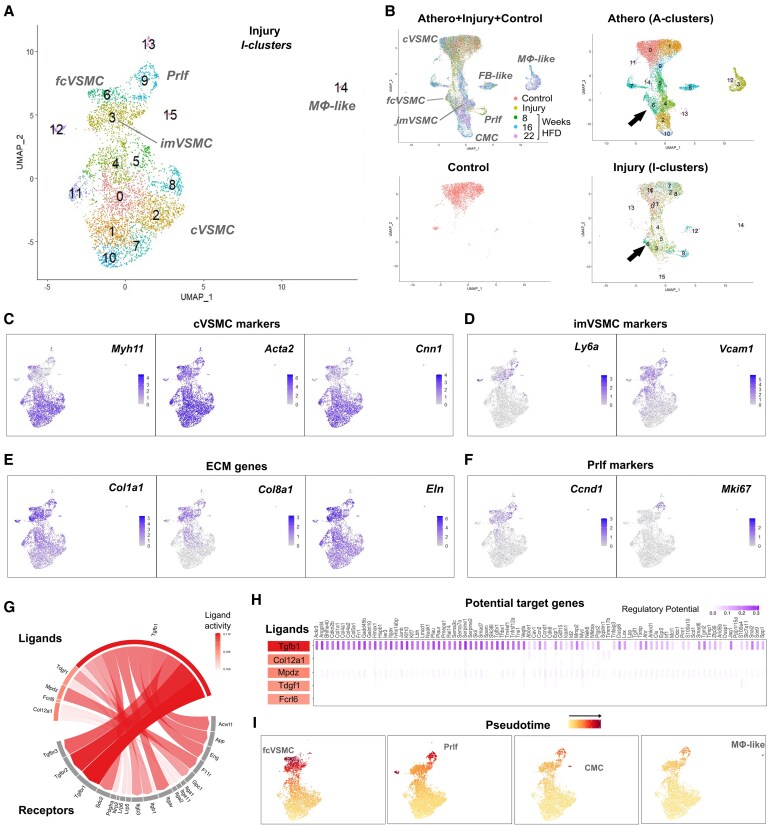
Figure 3.
VSMCs generate the transcriptional fibrous cap-associated VSMC state 11 days after vascular injury. (A) Uniform manifold approximation and projection (UMAP) showing injury (I) cell cluster annotation and VSMC states for scRNA-Seq analysis of VSMC-lineage label positive cells from ligated left carotid arteries of Myh11eYFP animals 11 days post-injury (DPI). (B) Integration of injury, atherosclerosis, and healthy control carotid arteries scRNA-Seq datasets. Top left: all cells coloured by dataset identity; top right: cells from the atherosclerosis data only, A-clusters annotated (see Figure 2A); lower left: cells from healthy arteries only, lower right: injury data, I-clusters annotated (panel A). (C-F) UMAP feature plots showing expression levels for markers of cVSMC (C), imVSMC (D), ECM (E), and proliferation (F) using grey (low) to blue scales (high). (G, H) Top 5 ligands and their associated receptors (G, arrow width indicates predicted strength) or regulatory potential for top 100 gene targets (H). Predicted ligand strength is indicated by shades of orange/red, regulatory potential in shades of purple. (I) UMAP showing the cells included the inferred trajectories leading to generation of a Prlf (I-cluster 9), CMC (I-cluster 15), MΦ-like or fibrous cap-associated (fcVSMC; I-cluster 6) cell state. Pseudotime is indicated using a yellow–red scale. cVSMC, contractile VSMC; imVSMC, intermediate modulated VSMC; CMC, chondromyocyte; FB, fibroblast; MΦ, macrophage; Prlf, proliferation.
Interestingly, integration of the 11 DPI transcriptional profiles with atherosclerosis data (Figure 3B; Supplementary material online, Figure S3), suggested that the range of VSMC states induced by carotid ligation injury at this time point overlaps those observed in atherosclerosis. As shown in Figure 2A and C-F, injured arteries contain imVSMCs expressing Ly6a+, Vcam1+, and Lgals3+ [injury cell cluster (I-cluster) 3 and 4] and Prlf cells expressing Ccnd1+, Pcna+, and Mki67+ (I-cluster 9), in addition to cVSMCs with no, or low, response to injury (I-cluster 0, 1, 2, 7, 10, and 11), as expected.13,32 Interestingly, we also detected cells with high levels of Sox9, Col2a1, and Chad, that were similar to CMCs (I-cluster 15, Supplementary material online, Figure S3). Rare cells expressing FB-like (I-cluster 12, Pi16+, Igfbp4+, and Dpep1+) or MΦ-like genes (I-cluster 14, Lgals3+, Cd68+, and Ptprc+) had low expression of the eYFP lineage label (see Supplementary material online, Figure S3), suggesting that these cells were not derived from VSMCs, similar to what has been proposed in atherosclerosis.29
Additionally, a cell population co-expressing contractile genes with key ECM genes (Col1a1, Mgp, Eln, Fn1, Col4a1, and Col8a1) and Notch3, mapped together with the fcVSMC population from the atherosclerosis dataset (I-cluster 6; Figure 3A and B; Supplementary material online, Figure S3). This population was also present in an independent experiment at 11 DPI (see Supplementary material online, Figure S4), but was not detected at earlier timepoints (5 and 7 DPI),13 suggesting that these cells are derived from modulated cells and represent VSMC re-differentiation. Intercellular communication analysis predicted that TGF-beta, an important determinant of fibrous cap formation, is the top ligand for I-cluster 6 cells and that it has high regulatory potential for I-cluster 6 marker genes (Figure 3G and H). Computational trajectory analysis indicated that, similar to in atherosclerosis,12,14 imVSMCs act as an intermediate state after injury that generate the fcVSMCs as well as proliferating cells and other VSMC-derived subsets (Figure 3I; Supplementary material online, Figure S2B and C). Together, these data support the idea that VSMCs transition to disease-relevant states via an imVSMC state and suggest that these transitions can be investigated using the reproducible vascular injury model. Importantly, our analysis identified a novel imVSMC-derived fcVSMC state in both models, that we hypothesise is implicated in fibrous cap formation.
3.3. VSMC transition to a NOTCH3+ fibrous cap-associated cell state occurs in the fibrous cap and injury-induced lesions
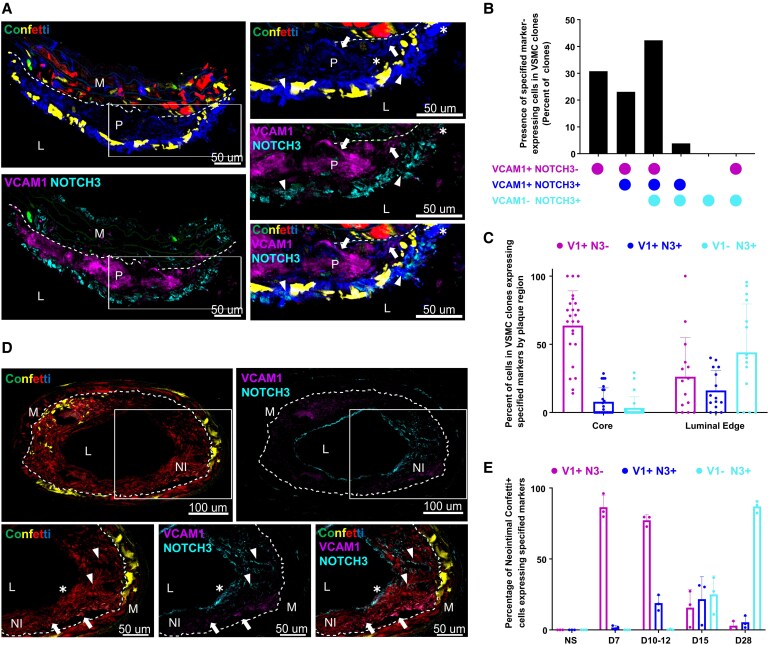
We then spatially mapped the imVSMC-to-fcVSMC transition, using Myh11Confetti mice where cell patches expressing the same fluorescent reporter protein represent clonally related VSMC-derived cells. Notch3 expression was largely confined to fcVSMCs in both atherosclerosis and injury (Figure 2C; Supplementary material online, Figure S2D), whereas Vcam1 was highly expressed by imVSMCs and down-regulated in fcVSMCs (Figures 2C and 3D). Hence, VCAM1+NOTCH3− cells highlighted the imVSMC state and VCAM1+NOTCH3+ represent cells at the cluster boundary (A-cluster 4-to-6 or I-cluster 3-to-6). VCAM1 and NOTCH3 expression was scored in Confetti+ cells in 26 individual clonal patches of VSMC-derived cells (16 plaques from three Myh11Confetti; Apoe−/− animals, Figure 4A–C; Supplementary material online, Figure S5A). VSMC clones containing cells with all marker combinations (VCAM1+NOTCH3−, VCAM1+NOTCH3+ and VCAM1−NOTCH3+ cells) were most frequently observed (42%; Figure 4B), showing that the progeny of a single VSMC can generate these three cell states. We did not observe VSMC clones containing only VCAM1−NOTCH3+ cells in any plaques, whilst VCAM1 was detected without NOTCH3 in 31% of clones (Figure 4B). This is consistent with the idea that VCAM1 induction is associated with VSMC entry into the plaque and that cells subsequently transition to the NOTCH3+ state.
Figure 4.
VSMCs in the intermediate modulated and fibrous cap-associated state co-occur in clonally related cells within lesions. (A-C) Immunofluorescence staining for VCAM1 and NOTCH3 in cryosections of a plaque from VSMC-lineage-labelled Myh11Confetti; Apoe−/− animals. (A) Representative image of plaque, showing the entire plaque (left) or magnified views of boxed region (right). Examples of VCAM1+ (arrows), NOTCH3+ (arrowheads), and VCAM1+NOTCH3+ cells (asterisk) are indicated. Scale bars = 50 µm. (B) Percentage of plaque-located Confetti+ clones that contain cells expressing specified combinations of markers. Dots represent clones (n = 26) from 16 plaques in three animals. (C) Percentage of cells with specified combinations of marker expression quantified separately for either the core or luminal edge portion of individual Confetti+ clones. n = 23 (core region) and 15 clones (luminal edge). Bars show mean, error-bars SD. (D, E) Immunofluorescence staining for VCAM1 and NOTCH3 in cryosections of the ligated artery of VSMC-lineage-labelled Myh11Confetti animals. (D) Representative image (15 days post-injury) showing entire section (top; scale bar = 100 µm) and magnified views of boxed regions (bottom; scale bar = 50 µm). Examples of VCAM1+ cells in the neointima (arrows), NOTCH3+ cells in the neointima (arrowheads) and cells expressing both VCAM1 and NOTCH3 (asterisk) are marked. (E) The proportion of neointimal Confetti+ cells with specified marker expression at different time points after injury (n = 3 animals per time point). Bars show mean, error-bars SD. (A, D) Signals show VCAM1 (magenta), NOTCH3 (cyan), and Confetti (CFP: blue, RFP: red, YFP: yellow, and GFP: green) as indicated. Dashed lines outline the internal elastic lamina. L, lumen; P, plaque, M, media; NI, neointima.
To assess whether the local plaque environment impact the likelihood of generating the imVSMC and fcVSMC states, we quantified marker-expressing cells separately for the luminal and core regions of each VSMC clone. In keeping with previous observations,20 we found abundant luminal NOTCH3 expression by VSMCs, whereas VCAM1 expression was more prominent in the core region of VSMC clones (Figure 4A and C). Similar proportions of the three cell states were detected in the luminal region, suggesting this is the site of imVSMC-to-fcVSMC state transition. Together, these data show that the computationally inferred transition from imVSMC to fcVSMC state occurs in vivo, and that this represents formation of the fibrous cap in atherosclerotic plaques.
To more directly test whether lesional NOTCH3+ fcVSMC cells are generated from VCAM1+ imVSMCs, we leveraged the acute and reproducible kinetics of lesion formation in the carotid ligation injury model. All three states (VCAM1+NOTCH3−, VCAM1+NOTCH3+, and VCAM1−NOTCH3+) were also detected within single VSMC clones in injury-induced lesions (Figure 4D), however, the frequencies varied over time. Initially, almost all lesional VSMCs were VCAM1+NOTCH3− (86%, 7 DPI, Figure 4E), but a substantial proportion of VCAM1+NOTCH3+ cells (19%) were observed in the growing intima at day 10–12. At 15 DPI, VCAM1+NOTCH3−, VCAM1+NOTCH3+, and VCAM1−NOTCH3+ states were detected at similar frequency (∼20% each; Figure 4E), whereas almost all lesional cells were VCAM1−NOTCH3+ at 28 DPI (87%; Figure 4E). For comparison, contractile gene re-expression was reported 2 weeks after injury.31 Together, these data demonstrate that VSMCs enter lesions in an imVSMC state and that clonal progeny of individual VSMCs transition to the fcVSMC state within injury-induced lesions. VSMCs in the medial layer of injured arteries expressed VCAM1 only at 7 DPI, NOTCH3 only at 15 and 28 DPI, and we did not detect VCAM1+NOTCH3+ cells at any time points (see Supplementary material online, Figure S5B), suggesting that the kinetics or underlying mechanisms may differ.
3.4. The thrombin receptor PAR1 is a candidate therapeutic target for promoting the fibrous cap-associated VSMC transition
To understand the regulatory mechanisms governing fcVSMC generation, we aimed to identify genes that are induced as cells undergo imVSMCs-to-fcVSMCs conversion. In total, 840 genes showed statistically significant pseudotime-dependent expression for the fcVSMC-generating trajectory in atherosclerosis [P-adj < 0.05, log(fold-change) > 0.25, Figure 5A]. These overlapped with pseudotime-associated genes for the equivalent trajectory in injury (59%; X2 test, P < 0.001; Figure 5B), further demonstrating the similarity of VSMC regulation in these models. Distinct pseudotime-dependent patterns were observed for Vcam1 and Notch3 (Figure 5C).
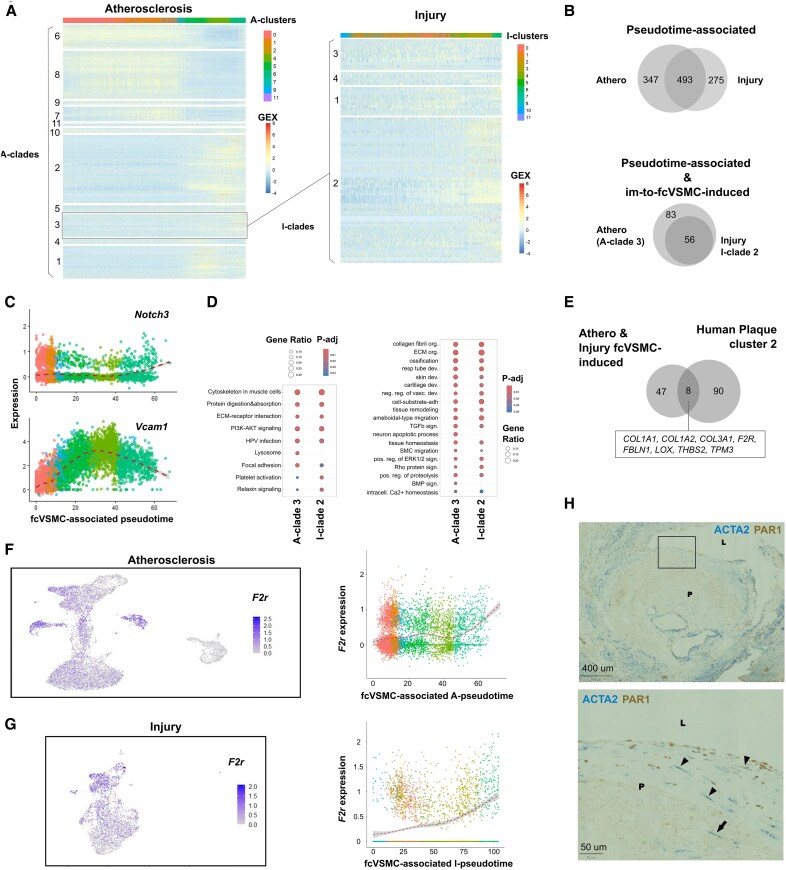
Figure 5.
Identification of PAR1 as a candidate regulator of VSMC differentiation to a fibrous cap-associated VSMC state. (A) Heatmap of pseudotime-dependent genes [P-adj < 0.05, log(fold-change) > 0.25] for the fcVSMC-associated trajectory in atherosclerosis clustered into ‘A-gene clades’ (left), and A-clade 3 genes clustered into ‘I-gene clades’ by expression along fcVSMC-associated pseudotime in injury (right). Cluster affiliation (top) is shown using Figures 2A (A-clusters, left) and 3A (I-clusters, right) colour scales. Blue (low)-to-red (high) colour-scale shows scaled gene expression. (B) Overlap of all fcVSMC pseudotime-associated genes in atherosclerosis and injury (top) and intersection of A-clade 3 and I-clade 2 genes (lower panel). (C) Notch3 and Vcam1 expression along pseudotime for fcVSMC generation in injury. Dashed line shows a GAM. (D) Selected gene ontologies enriched in A-clade 3 and I-clade 2 genes (see Supplementary material online, Tables S8-S11). P-adj: cumulative hypergeometric probability testing with g:SCS correction. (E) Venn diagram for common fcVSMC-induced genes (I-clade 2) and genes induced in human fibrous cap-located spatial transcriptomics capture spots. The eight overlapping genes are listed. (F, G) F2r feature plots (left) and pseudotime-dependent expression in atherosclerosis (F) and injury (G). (H) Representative immunohistochemistry staining for ACTA2 and PAR1 in sections of human carotid plaques (n = 5). Panels show a large region of the plaque (top) and a magnified view of the boxed region (bottom). Signals for ACTA2 (blue) and PAR1 (brown) are shown as indicated. Examples of ACTA2 + PAR1+ cells in the cap (arrowheads) and ACTA2 + PAR1+ cells (arrow) in the core are shown. Scale bars = 400 µm (top), 50 µm (bottom). L, lumen; P, plaque, M, media.
Hierarchical clustering of pseudotime-associated genes by expression pattern highlighted a gene subset (A-clade 3, boxed in Figure 5A) that is specifically up-regulated at the imVSMC-to-fcVSMC transition. A similar pseudotime-dependent induction was observed for 56 of the 84 A-clade 3 genes in the injury model (I-clade 2; Figure 5A and B). Pathway analysis indicated consistent enrichment of ECM organization, migration and adhesion for these ‘fcVMSC induced genes’ and highlighted signalling pathways, including TGF-beta, as potential regulators (Figure 5D). We focused on I-clade 2 genes, hypothesizing that the acute, accelerated kinetics of lesion formation in this model would enrich for genes that drive fcVSMC differentiation, rather than being expressed as a consequence of the transition.
The relevance to fibrous cap formation in human atherosclerosis was assessed using spatial transcriptomics. To capture the regulatory mechanisms driving ongoing fibrous cap development, we analysed an aortic sample with an early stage, fibrous cap-containing plaque (see Supplementary material online, Figure S6). A cluster of capture spots (human plaque [HP]-cluster 2, localized to the luminal edge of the lesion) had abundant levels of contractile markers and low expression of immune cell markers (CD68, CD163, and CD86) suggesting that these represent the fibrous cap cluster that contained mainly VSMCs. Eight genes of the 56 fcVSMC-induced genes were among the 90 marker genes for HP-cluster 2 (X2 test, P < 0.001), including fibrillar collagens33 (COL1A1, COL1A2, COL3A1), FBLN1, LOX, TPM3, THBS2, and F2R; Figure 5E; Supplementary material online, Figure S6C–E).
We selected F2R, which encodes the thrombin receptor or protease-activated receptor 1 (PAR1), for further analysis. F2r transcripts were detected at high levels in fcVSMC clusters in both atherosclerosis and injury models, with low expression in A-cluster 7 (termed minor VSMC12) FB-like A-cluster 8 cells, and imVSMCs (I-cluster 3) (Figure 5F and G). Immunostaining in human atherosclerosis revealed abundant PAR1-expressing ACTA2+ VSMCs in human plaque fibrous caps, and PAR1 protein was also detected at lower levels in the core (Figure 5H). Previous investigations have shown that thrombin promotes VSMC proliferation,34,35 but a role in VSMC re-differentiation and fibrous cap generation has not been investigated. We therefore assessed whether PAR1 functionally affects the imVSMC-to-fcVSMC transition, and thus potentially fibrous cap formation in primary human VSMC isolates. We observed a dose-dependent, statistically significant increase in CNN1 and ACTA2 expression in response to physiologically relevant thrombin concentrations (Figure 6A). Interestingly, the effect of thrombin was more profound in VSMCs isolated from female compared to male donors (Figure 6A; P = 0.008, Mann–Whitney U test).
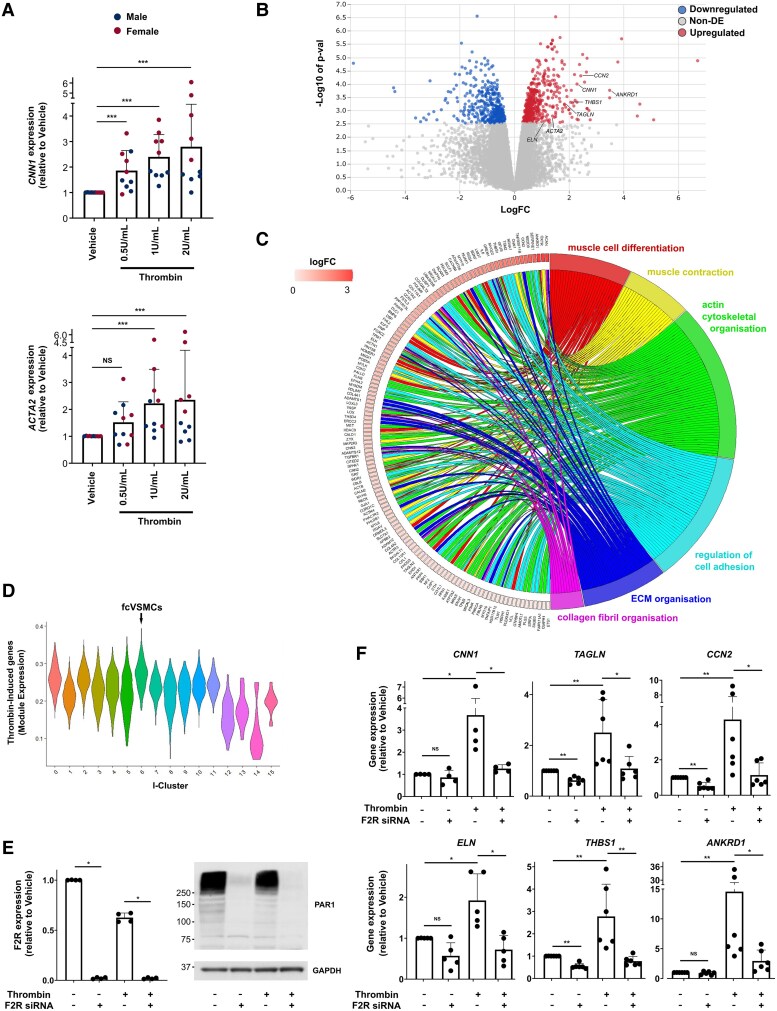
Figure 6.
PAR1 stimulation induces a transcriptional pattern associated with fibrous cap cells. (A) Transcript levels (qRT-PCR) of contractile genes CNN1 and ACTA2 in primary human VSMC (hVSMC) isolates from different donors (n = 5 male, blue, and 5 female donors, red) after 24 h vehicle or thrombin treatment (0.5, 1, or 2 U/mL). (B) Volcano plot showing differential gene expression between thrombin-treated (2 U/mL) vs. vehicle control samples (n = 3 female donors). Differentially expressed genes are coloured (P-adj < 0.05, red: induced by thrombin, blue: reduced by thrombin) and genes of interest indicated. See Supplementary material online, Table S12. (C) VSMC- and ECM-associated GO terms enriched in thrombin-induced genes and associated differentially expressed genes [white-red scale, log(fold-change)]. (D) Violin plot showing expression of thrombin-induced gene orthologues across the injury cell clusters. The arrow marks fcVSMC I-cluster 6. (E) F2R transcript levels (left, normalized to vehicle treated, n = 4) and western blot (right; molecular weight in kDa, n = 3) in primary hVSMCs ± thrombin with F2R-targeting (+) or control siRNA (−). (F) Transcript levels of candidate target genes in thrombin-treated hVSMCs relative to vehicle-treated samples with F2R-targeting (+) or control siRNA (−). n = 4–6. (A, E, F) Bars show mean ± SD relative to vehicle. *P-adj < 0.05, **P-adj < 0.01, ***P-adj < 0.001, NS: not significant (Mann–Whitney with Bonferroni-correction, following Kruskal–Wallis with P < 0.05). Replicates are hVSMC lines from different donors.
To comprehensively evaluate the transcriptional state induced by PAR1 stimulation, we profiled gene expression in thrombin-treated and untreated cells using RNA-seq. We confirmed up-regulation of contractile VSMC genes (including CNN1, ACTA2, and TAGLN) in thrombin-treated vs. control cells, and also detected increased expression of factors that have been associated with VSMC differentiation and/or fibrous cap development (Figure 6B). For example, connective tissue growth factor (CCN2) is involved in maintenance of the contractile VSMC state and is expressed in the fibrous cap36; elastin (ELN) is a key ECM component of the fibrous cap and also functions to maintain VSMC contractility37; Thrombospondin 1 (THBS1) has been implicated as a key factor involved in fibrous cap formation38; and Cardiac ankyrin repeat protein (ANKRD1) is expressed by VSMCs of the fibrous cap.39 Systematic analysis showed that the 492 thrombin up-regulated genes (P-adj < 0.05) were enriched for GO-terms associated with muscle cell differentiation, contraction, and focal adhesion as well as ECM organization, including collagen fibril organization, which is a key component of fibrous cap formation33 (Figure 6C; Supplementary material online, Table S12 and S13). A gene module comprising murine orthologues of thrombin-induced genes exhibited peak expression in the fcVSMC population identified in injured mouse arteries (I-cluster 6; Figure 6D). This suggests an overall induction of a fibrous cap-associated VSMC state by thrombin.
Thrombin-induced genes were also stimulated in response to another PAR1 protease, Factor Xa, although this did not reach statistical significance for all genes (see Supplementary material online, Figure S7A). To directly test whether protease-induced expression of contractile and fibrous cap-relevant genes was mediated by thrombin receptor stimulation, we targeted PAR1 activity using siRNA (Figure 6E; Supplementary material online, Figure S7B), which blunted or abolished thrombin-induced expression of CNN1, TAGLN, CCN2, ELN, THBS1, and ANKRD1 (Figure 6F). A PAR1-specific small molecular inhibitor, vorapaxar, also blocked thrombin-mediated gene induction (see Supplementary material online, Figure S7C). Interestingly, baseline expression of TAGLN, CCN2 and THBS1 was reduced by PAR1 knockdown, suggesting thrombin-independent regulation of these genes by PAR1 (Figure 6F). Together, these data indicate that thrombin receptor activation in VSMCs leads to induction of fcVSMC-associated genes.
4. Discussion
Here, we provide evidence that a novel VSMC sub-population (that we term fcVSMCs) is derived from VSMCs expressing a previously described intermediate modulated, or imVSMC, state, is present in the fibrous cap of plaques, and can be modelled in injury-induced lesions: (i) the expression of key fibrous cap-associated genes by fcVSMCs in both atherosclerosis and injury, (ii) trajectory analysis consistently predicts that fcVSMCs are derived from imVSMCs, (iii) co-detection of imVSMCs and fcVSMCs markers in clonally related cells in atherosclerotic lesions, (iv) initial absence of intimal NOTCH3-expression after injury, and (v) the fcVSMC marker, NOTCH3, shows abundant expression in the fibrous cap. We also identify candidate regulators of the proposed intermediate-to-fcVSMC transition, including F2R-encoded PAR1 and provide evidence suggesting that PAR1 stimulates the fibrous cap state. Finally, we show that PAR1 stimulation by thrombin in primary human VSMCs induces fibrous cap VSMC markers and a transcriptional profile matching that of fcVSMCs. We therefore suggest that PAR1 stimulation in VSMCs could promote fibrous cap formation.
The notion that an imVSMC state gives rise to fibrous cap VSMCs aligns with previous models. In particular, Ly6a/SCA1-expressing VSMCs are present prior to disease and have increased proliferative capacity,13 similar to the SCA1+ cells isolated from lesions.12 It is therefore conceivable that cells in an imVSMC state invade lesions and expand clonally to generate all VSMC-derived plaque cells, including those comprising the fibrous cap. Other groups have shown that manipulation of factors that promote the imVSMC state disrupt fibrous cap formation.18,19 NOTCH loss-of-function was required for VSMC plaque entry, whereas NOTCH3 reactivation is required for fibrous cap formation20 and NOTCH signalling is known to promote the contractile VSMC phenotype.40 This is consistent with our findings (i) that fcVSMCs up-regulate Notch3, (ii) our confirmation of NOTCH3 expression in fibrous cap VSMCs and delayed NOTCH3 expression when compared to VCAM1. Our findings are also consistent with dual lineage tracing experiments showing that Lgals3-expressing modulated cells generate all lesional VSMCs, including those in the fibrous cap at early atherogenesis.15 Alternative pathways may also contribute to fibrous cap formation, and could explain the presence of non-Lgals3-lineage cells at later plaque stages.15
PAR1 is expressed by VSMCs and thrombin is a well-known mitogen of these cells in vitro, putatively via PAR1.34,35 This proliferation-stimulating effect may exacerbate the response to vascular injury.41 VSMC proliferation has been detected in the fibrous cap10 and it is generally considered to indicate plaque stability, however, the role of PAR1 in fibrous cap formation has not been studied. Our finding that PAR1 stimulation induces expression of contractile genes in cultured VSMCs is consistent with studies in rat VSMCs42 and analysis of published microarray data from human aortic VSMCs after shorter thrombin treatment.43 Furthermore, previous work has shown that the PAR1 target CCN2, was also induced by thrombin in rat VSMCs.44
PAR1 is also expressed on platelets and by endothelial cells. Systemic inhibition of either thrombin or PAR1 attenuated atherosclerosis progression and even promoted features of plaque stability, due to effects on endothelial function, proinflammatory signalling and immune cell infiltration.41,45–48 Thus, cell type specific deletion of PAR-1 is needed to directly assess VSMC-specific effects in disease and, specifically, fibrous cap formation in atherosclerosis lesions. This could also hamper therapeutic translation of these findings. However, the identification of PAR1 as a candidate regulator of fibrous cap formation could therefore have several implications for the management of atherosclerosis. First, anticoagulants used to treat patients with atherosclerotic disease inhibits PAR1 (vorapaxar) or thrombin directly.49,50 Given the fibrous cap-inducing effect of VSMC-expressed PAR1 suggested here, vorapaxar treatment could have an unintended negative effect on plaque stability. Secondly, the induction of fcVSMC-associated genes by thrombin was more prominent in female than male VSMCs, and could contribute to the sex differences in plaque biology, particularly, the greater frequency of thin or ruptured fibrous caps in male patients.
Our observation that VSMC states induced by injury overlap those observed in atherosclerosis suggests that transitions to plaque VSMC states is an inherent characteristic of VSMCs in different physiological settings. Indeed, there are many commonalities between the two disease models. Both atherosclerotic and injured arteries show endothelial activation and a key role of inflammation, suggesting overlap in triggering mechanisms. The absence of a fibrous cap and high cholesterol in the injury model are notable differences. Recent evidence has suggested that plaque cholesterol induces VSMC responses similar to those of inflammation,51 which may explain the remarkable similarities of these vascular disease models. VSMC proliferation, migration and oligoclonal contribution to lesions is observed in atherosclerosis, aneurysm, and injury, and the substantial overlap in gene regulation highlighted here has also been found in other studies. For example, recent scRNA-seq analyses indicated that imVSMCs are present in both atherosclerotic and injured arteries.13,32,52 Our observation that an fcVSMC population is present in both models indicates that aspects of fibrous cap formation are analogous to VSMC re-differentiation in injury-induced lesions, although a fibrous cap structure is not formed. Importantly, this implies that the carotid ligation injury model, which is acute and has reproducible kinetics compared to experimental atherosclerosis, could be used to model the generation of fibrous cap cells. For instance, manipulation of candidate regulators, such as PAR1, could be used to screen factors for effects on the proportion of fcVSMCs in neointimal lesions at well-characterized time points.
The identification of a VSMC transition associated with fibrous cap formation provides exciting new insight into the mechanisms underpinning plaque stability. We propose that deeper understanding of this transition and its regulation could provide new drug targets that could be used for promoting fibrous cap formation and thus mitigation of plaque rupture.
Translational perspective.
Identifying regulators of the VSMC transition that generates fibrous cap cells could enable development of new interventions to stabilize atherosclerotic plaques and mitigate acute cardiovascular events resulting from plaque rupture.
Supplementary Material
Acknowledgements
We acknowledge the NIHR BRC Cell Phenotyping Hub for cell sorting, Darran Clements (the MRC/Wellcome Stem Cell Institute Advanced Imaging Facility) for assistance with confocal imaging, the Babraham Institute Sequencing Facility (UKRI-BBSRC Core Capability Grant), Simon Andrews and Felix Krueger (Babraham Bioinformatics) for data processing, Lauren Kitt for assistance with animal models. Data analysis were done using the High Performance Computer clusters at the University of Cambridge and Imperial College Research Computing Service (DOI: 10.14469/hpc/2232). We thank donors, their families, specialist nurses on organ donation (SNOD), and the Papworth Tissue Bank for human tissue samples. The Visium experiment was done by the Spatially Resolved Transcriptomics (SpaRTAN) group at Newcastle University.
Contributor Information
James C K Taylor, Section of Cardiorespiratory Medicine, Department of Medicine, University of Cambridge, Victor Phillip Dahdaleh Heart and Lung Research Institute, Papworth Road, Cambridge CB2 0BB, UK.
Matthew D Worssam, Section of Cardiorespiratory Medicine, Department of Medicine, University of Cambridge, Victor Phillip Dahdaleh Heart and Lung Research Institute, Papworth Road, Cambridge CB2 0BB, UK.
Sebnem Oc, Section of Cardiorespiratory Medicine, Department of Medicine, University of Cambridge, Victor Phillip Dahdaleh Heart and Lung Research Institute, Papworth Road, Cambridge CB2 0BB, UK; Functional Gene Control Group, MRC Laboratory of Medical Sciences and Institute of Clinical Sciences, Faculty of Medicine, Imperial College Du Cane Road, London W12 0NN, UK.
Jordi Lambert, Section of Cardiorespiratory Medicine, Department of Medicine, University of Cambridge, Victor Phillip Dahdaleh Heart and Lung Research Institute, Papworth Road, Cambridge CB2 0BB, UK.
Krishnaa T Mahbubani, Collaborative Biorepository for Translational Medicine, Department of Surgery, University of Cambridge and NIHR Cambridge Biomedical Research Centre, Cambridge CB2 0QQ, UK.
Kirsty Foote, Section of Cardiorespiratory Medicine, Department of Medicine, University of Cambridge, Victor Phillip Dahdaleh Heart and Lung Research Institute, Papworth Road, Cambridge CB2 0BB, UK.
Allie Finigan, Section of Cardiorespiratory Medicine, Department of Medicine, University of Cambridge, Victor Phillip Dahdaleh Heart and Lung Research Institute, Papworth Road, Cambridge CB2 0BB, UK.
Yee-Hung Chan, Section of Cardiorespiratory Medicine, Department of Medicine, University of Cambridge, Victor Phillip Dahdaleh Heart and Lung Research Institute, Papworth Road, Cambridge CB2 0BB, UK.
Nichola Figg, Section of Cardiorespiratory Medicine, Department of Medicine, University of Cambridge, Victor Phillip Dahdaleh Heart and Lung Research Institute, Papworth Road, Cambridge CB2 0BB, UK.
Murray C H Clarke, Section of Cardiorespiratory Medicine, Department of Medicine, University of Cambridge, Victor Phillip Dahdaleh Heart and Lung Research Institute, Papworth Road, Cambridge CB2 0BB, UK.
Martin R Bennett, Section of Cardiorespiratory Medicine, Department of Medicine, University of Cambridge, Victor Phillip Dahdaleh Heart and Lung Research Institute, Papworth Road, Cambridge CB2 0BB, UK.
Helle F Jørgensen, Section of Cardiorespiratory Medicine, Department of Medicine, University of Cambridge, Victor Phillip Dahdaleh Heart and Lung Research Institute, Papworth Road, Cambridge CB2 0BB, UK.
Supplementary material
Supplementary material is available at Cardiovascular Research online.
Authors’ Contributions
J.C.K.T. and H.F.J. conceived the study. J.C.K.T., M.D.W., J.L., and S.O. performed experiments and data analysis. N.F., K.F., A.F., and Y.-H.C. performed surgery and histology supervised by M.R.B. and M.C.H.C. K.T.M. provided human tissue samples. J.C.K.T. and H.F.J. wrote the manuscript with input from all authors.
Funding
This work was supported by grants from the British Heart Foundation (PG/19/6/34153, FS/15/62/32032, FS/15/38/31516, RM/13/3/30159, RE/13/6/30180, RE/18/1/34212, CH/2000003/12800 to J.C.K.T., M.D.W., H.F.J., and M.R.B.), the National Institute of Health Research (NIHR) Cambridge Biomedical Research Centre and the Chan Zuckerberg Initiative (2018-190766/RG98793, K.T.M., the Collaborative Bioresource for Translational Medicine).
Data availability
Bulk (GSE274393), spatial (GSE274572), and single-cell RNA-seq datasets generated in this study (GSE274572) have been deposited, and VSMC-lineage-labelled plaque cell profiles from HFD-fed animals (GSE155513) are available from the gene expression omnibus (GEO). Source data are available upon request.
References
- 1. Basatemur GL, Jørgensen HF, Clarke MCH, Bennett MR, Mallat Z. Vascular smooth muscle cells in atherosclerosis. Nat Rev Cardiol 2019;16:727–744. [DOI] [PubMed] [Google Scholar]
- 2. Gomez D, Shankman LS, Nguyen AT, Owens GK. Detection of histone modifications at specific gene loci in single cells in histological sections. Nat Methods 2013;10:171–177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Yang P, Hong MS, Fu C, Schmit BM, Su Y, Berceli SA, Jiang Z. Preexisting smooth muscle cells contribute to neointimal cell repopulation at an incidence varying widely among individual lesions. Surgery 2016;159:602–612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Nemenoff RA, Horita H, Ostriker AC, Furgeson SB, Simpson PA, VanPutten V, Crossno J, Offermanns S, Weiser-Evans MCM. SDF-1a; induction in mature smooth muscle cells by inactivation of PTEN is a critical mediator of exacerbated injury-induced neointima formation. Arterioscler Thromb Vasc Biol 2011;31:1300–1308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Gomez D, Owens GK. Smooth muscle cell phenotypic switching in atherosclerosis. Cardiovasc Res 2012;95:156–164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Shankman LS, Gomez D, Cherepanova OA, Salmon M, Alencar GF, Haskins RM, Swiatlowska P, Newman AAC, Greene ES, Straub AC, Isakson B, Randolph GJ, Owens GK. KLF4-dependent phenotypic modulation of smooth muscle cells has a key role in atherosclerotic plaque pathogenesis. Nat Med 2015;21:628–637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Chappell J, Harman JL, Narasimhan VM, Yu H, Foote K, Simons BD, Bennett MR, Jørgensen HF. Extensive proliferation of a subset of differentiated, yet plastic, medial vascular smooth muscle cells contributes to neointimal formation in mouse injury and atherosclerosis models. Circ Res 2016;119:1313–1323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Feil S, Fehrenbacher B, Lukowski R, Essmann F, Schulze-Osthoff K, Schaller M, Feil R. Transdifferentiation of vascular smooth muscle cells to macrophage-like cells during atherogenesis. Circ Res 2014;115:662–667. [DOI] [PubMed] [Google Scholar]
- 9. Jacobsen K, Lund MB, Shim J, Gunnersen S, Füchtbauer E-M, Kjolby M, Carramolino L, Bentzon JF. Diverse cellular architecture of atherosclerotic plaque derives from clonal expansion of a few medial SMCs. Jci Insight 2017;2:e95890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Misra A, Feng Z, Chandran RR, Kabir I, Rotllan N, Aryal B, Sheikh AQ, Ding L, Qin L, Fernández-Hernando C, Tellides G, Greif DM. Integrin beta3 regulates clonality and fate of smooth muscle-derived atherosclerotic plaque cells. Nat Commun 2018;9:2073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Dobnikar L, Taylor AL, Chappell J, Oldach P, Harman JL, Oerton E, Dzierzak E, Bennett MR, Spivakov M, Jørgensen HF. Disease-relevant transcriptional signatures identified in individual smooth muscle cells from healthy mouse vessels. Nat Commun 2018;9:1–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Pan H, Xue C, Auerbach BJ, Fan J, Bashore AC, Cui J, Yang DY, Trignano SB, Liu W, Shi J, Ihuegbu CO, Bush EC, Worley J, Vlahos L, Laise P, Solomon RA, Connolly ES, Califano A, Sims PA, Zhang H, Li M, Reilly MP. Single-cell genomics reveals a novel cell state during smooth muscle cell phenotypic switching and potential therapeutic targets for atherosclerosis in mouse and human. Circulation 2020;142:2060–2075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Worssam MD, Lambert J, Oc S, Taylor JCK, Taylor AL, Dobnikar L, Chappell J, Harman JL, Figg NL, Finigan A, Foote K, Uryga AK, Bennett MR, Spivakov M, Jørgensen HF. Cellular mechanisms of oligoclonal vascular smooth muscle cell expansion in cardiovascular disease. Cardiovasc Res 2022;119:1279–1294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Conklin AC, Nishi H, Schlamp F, Örd T, Õunap K, Kaikkonen MU, Fisher EA, Romanoski CE. Meta-analysis of smooth muscle lineage transcriptomes in atherosclerosis and their relationships to in vitro models. Immunometabolism 2021;3:e210022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Alencar GF, Owsiany KM, Karnewar S, Sukhavasi K, Mocci G, Nguyen A, Williams CM, Shamsuzzaman S, Mokry M, Henderson CA, Haskins R, Baylis RA, Finn AV, McNamara CA, Zunder ER, Venkata V, Pasterkamp G, Björkegren J, Bekiranov S, Owens GK. The stem cell pluripotency genes klf4 and oct4 regulate Complex SMC phenotypic changes critical in late-stage atherosclerotic lesion pathogenesis. Circulation 2020;142:2045–2059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Newby AC, Zaltsman AB. Fibrous cap formation or destruction—the critical importance of vascular smooth muscle cell proliferation, migration and matrix formation. Cardiovasc Res 1999;41:345–360. [PubMed] [Google Scholar]
- 17. Rekhter MD. Collagen synthesis in atherosclerosis: too much and not enough. Cardiovasc Res 1999;41:376–384. [DOI] [PubMed] [Google Scholar]
- 18. Newman AAC, Serbulea V, Baylis RA, Shankman LS, Bradley X, Alencar GF, Owsiany K, Deaton RA, Karnewar S, Shamsuzzaman S, Salamon A, Reddy MS, Guo L, Finn A, Virmani R, Cherepanova OA, Owens GK. Multiple cell types contribute to the atherosclerotic lesion fibrous cap by PDGFRβ and bioenergetic mechanisms. Nat Metab 2021;3:166–181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Wirka RC, Wagh D, Paik DT, Pjanic M, Nguyen T, Miller CL, Kundu R, Nagao M, Coller J, Koyano TK, Fong R, Woo YJ, Liu B, Montgomery SB, Wu JC, Zhu K, Chang R, Alamprese M, Tallquist MD, Kim JB, Quertermous T. Atheroprotective roles of smooth muscle cell phenotypic modulation and the TCF21 disease gene as revealed by single-cell analysis. Nat Med 2019;25:1280–1289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Martos-Rodríguez CJ, Albarrán-Juárez J, Morales-Cano D, Caballero A, MacGrogan D, de la Pompa JL, Carramolino L, Bentzon JF. Fibrous caps in atherosclerosis form by notch-dependent mechanisms common to arterial Media development. Arterioscler Thromb Vasc Biol 2021;41:e427–e439. [DOI] [PubMed] [Google Scholar]
- 21. Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. Limma powers differential expression analyses for RNA-Sequencing and microarray studies. Nucleic Acids Res 2015;43:e47–e47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Butler A, Hoffman P, Smibert P, Papalexi E, Satija R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol 2018;36:411–420. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Street K, Risso D, Fletcher RB, Das D, Ngai J, Yosef N, Purdom E, Dudoit S. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 2018;19:477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Wolf FA, Hamey FK, Plass M, Solana J, Dahlin JS, Göttgens B, Rajewsky N, Simon L, Theis FJ. PAGA: graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biol 2019;20:59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Wolf FA, Angerer P, Theis FJ. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol 2018;19:15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Yu G, Wang L-G, Han Y, He Q-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics J Integr Biol 2012;16:284–287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Alonso-Herranz L, Albarrán-Juárez J, Bentzon JF. Mechanisms of fibrous cap formation in atherosclerosis. Front Cardiovasc Med 2023;10:1254114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Lu YW, Lowery AM, Sun L-Y, Singer HA, Dai G, Adam AP, Vincent PA, Schwarz JJ. Endothelial myocyte enhancer factor 2c inhibits migration of smooth muscle cells through fenestrations in the internal elastic Lamina. Arterioscler Thromb Vasc Biol 2017;37:1380–1390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Sharma D, Worssam MD, Pedroza AJ, Dalal AR, Alemany H, Kim H-J, Kundu R, Fischbein MP, Cheng P, Wirka R. Comprehensive integration of multiple single-cell transcriptomic data sets defines distinct cell populations and their phenotypic changes in murine atherosclerosis. Arterioscler Thromb Vasc Biol 2024;44:391–408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Kumar A, Lindner V. Remodeling with neointima formation in the mouse carotid artery after cessation of blood flow. Arterioscler Thromb Vasc Biol 1997;17:2238–2244. [DOI] [PubMed] [Google Scholar]
- 31. Regan CP, Adam PJ, Madsen CS, Owens GK. Molecular mechanisms of decreased smooth muscle differentiation marker expression after vascular injury. J Clin Invest 2000;106:1139–1147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Warwick T, Buchmann GK, Pflüger-Müller B, Spaeth M, Schürmann C, Abplanalp W, Tombor L, John D, Weigert A, Leo-Hansmann M, Dimmeler S, Brandes RP. Acute injury to the mouse carotid artery provokes a distinct healing response. Front Physiol 2023;14:1125864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Jansen I, Cahalane R, Hengst R, Akyildiz A, Farrell E, Gijsen F, Aikawa E, van der Heiden K, Wissing T. The interplay of collagen, macrophages, and microcalcification in atherosclerotic plaque cap rupture mechanics. Basic Res Cardiol 2024;119:193–213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Fager G. Thrombin and proliferation of vascular smooth muscle cells. Circ Res 1995;77:645–650. [DOI] [PubMed] [Google Scholar]
- 35. Schrör K, Bretschneider E, Fischer K, Fischer J, Pape R, Rauch B, Rosenkranz A, Weber A-A. Thrombin receptors in vascular smooth muscle cells – function and regulation by vasodilatory prostaglandins. Thromb Haemost 2010;103:884–890. [DOI] [PubMed] [Google Scholar]
- 36. Cicha I, Yilmaz A, Klein M, Raithel D, Brigstock DR, Daniel WG, Goppelt-Struebe M, Garlichs CD. Connective tissue growth factor is overexpressed in complicated atherosclerotic plaques and induces mononuclear cell chemotaxis in vitro. Arterioscler Thromb Vasc Biol 2005;25:1008–1013. [DOI] [PubMed] [Google Scholar]
- 37. Karnik SK, Brooke BS, Bayes-Genis A, Sorensen L, Wythe JD, Schwartz RS, Keating MT, Li DY. A critical role for elastin signaling in vascular morphogenesis and disease. Development 2003;130:411–423. [DOI] [PubMed] [Google Scholar]
- 38. Moura R, Tjwa M, Vandervoort P, kerckhoven SV, Holvoet P, Hoylaerts MF. Thrombospondin-1 deficiency accelerates atherosclerotic plaque maturation in ApoE−/− mice. Circ Res 2008;103:1181–1189. [DOI] [PubMed] [Google Scholar]
- 39. de Waard V, van Achterberg TAE, Beauchamp NJ, Pannekoek H, de Vries CJM. Cardiac ankyrin repeat protein (CARP) expression in human and murine atherosclerotic lesions. Arterioscler Thromb Vasc Biol 2003;23:64–68. [DOI] [PubMed] [Google Scholar]
- 40. Tang Y, Urs S, Boucher J, Bernaiche T, Venkatesh D, Spicer DB, Vary CPH, Liaw L. Notch and transforming growth factor-β (TGFβ) signaling pathways cooperatively regulate vascular smooth muscle cell differentiation*. J Biol Chem 2010;285:17556–17563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Jaberi N, Soleimani A, Pashirzad M, Abdeahad H, Mohammadi F, Khoshakhlagh M, Khazaei M, Ferns GA, Avan A, Hassanian SM. Role of thrombin in the pathogenesis of atherosclerosis. J Cell Biochem 2019;120:4757–4765. [DOI] [PubMed] [Google Scholar]
- 42. Schauwienold D, Plum C, Helbing T, Voigt P, Bobbert T, Hoffmann D, Paul M, Reusch HP. ERK1/2-Dependent Contractile protein expression in vascular smooth muscle cells. Hypertens J Am Hear Assoc 2003;41:546–552. [DOI] [PubMed] [Google Scholar]
- 43. Janjanam J, Zhang B, Mani AM, Singh NK, Traylor JG, Orr AW, Rao GN. LIM and cysteine-rich domains 1 is required for thrombin-induced smooth muscle cell proliferation and promotes atherogenesis. J Biol Chem 2018;293:3088–3103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Ko W, Chen B, Hsu M, Tsai C, Hong C, Lin C. Thrombin induced connective tissue growth factor expression in rat vascular smooth muscle cells via the PAR-1/JNK/AP-1 pathway. Acta Pharmacol Sin 2012;33:49–56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Friebel J, Moritz E, Witkowski M, Jakobs K, Strässler E, Dörner A, Steffens D, Puccini M, Lammel S, Glauben R, Nowak F, Kränkel N, Haghikia A, Moos V, Schutheiss H-P, Felix SB, Landmesser U, Rauch BH, Rauch U. Pleiotropic effects of the protease-activated receptor 1 (PAR1) inhibitor, vorapaxar, on atherosclerosis and vascular inflammation. Cells 2021;10:3517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Pingel S, Tiyerili V, Mueller J, Werner N, Nickenig G, Mueller C. Thrombin inhibition by dabigatran attenuates atherosclerosis in ApoE deficient mice. Arch Méd Sci AMS 2014;10:154–160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Kalz J, ten Cate H, Spronk HMH. Thrombin generation and atherosclerosis. J Thromb Thrombolysis 2014;37:45–55. [DOI] [PubMed] [Google Scholar]
- 48. Spronk HMH, Borissoff JI, Soehnlein O, Koenen R, van Oerle R, Heeneman S, Daemen MJAP, Esmon CT, Degen JL, Weiler H, van Ryn J, Cate HT. Thrombin inhibition prevents against severe atherosclerosis progression in prothrombotic mice. Blood 2012;120:103. [Google Scholar]
- 49. Tantry US, Bliden KP, Chaudhary R, Novakovic M, Rout A, Gurbel PA. Vorapaxar in the treatment of cardiovascular diseases. Futur Cardiol 2020;16:373–384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Lee CJ, Ansell JE. Direct thrombin inhibitors. Br J Clin Pharmacol 2011;72:581–592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Carramolino L, Albarrán-Juárez J, Markov A, Hernández-SanMiguel E, Sharysh D, Cumbicus V, Morales-Cano D, Labrador-Cantarero V, Møller PL, Nogales P, Benguria A, Dopazo A, Sanchez-Cabo F, Torroja C, Bentzon JF. Cholesterol lowering depletes atherosclerotic lesions of smooth muscle cell-derived fibromyocytes and chondromyocytes. Nat Cardiovasc Res 2024;3:203–220. [DOI] [PubMed] [Google Scholar]
- 52. Lambert J, Oc S, Worssam MD, Häußler D, Solomon CU, Figg NL, Baxter R, Imaz M, Taylor JCK, Foote K, Finigan A, Mahbubani KT, Webb TR, Ye S, Bennett MR, Krüger A, Spivakov M, Jørgensen HF. Network-based prioritization and validation of regulators of vascular smooth muscle cell proliferation in disease. Nat Cardiovasc Res 2024;3:714–733. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Bulk (GSE274393), spatial (GSE274572), and single-cell RNA-seq datasets generated in this study (GSE274572) have been deposited, and VSMC-lineage-labelled plaque cell profiles from HFD-fed animals (GSE155513) are available from the gene expression omnibus (GEO). Source data are available upon request.