ABSTRACT
Charge heterogeneity in monoclonal antibodies (mAbs), caused by post-translational modifications, remains a substantial obstacle to ensuring consistent, stable, and effective therapeutics. Conventional optimization techniques, such as one-factor-at-a-time and design of experiments, often fail to capture the complex, nonlinear interactions between culture parameters (e.g. pH, temperature, duration) and medium components (e.g. glucose, metal ions, amino acids). This review highlights machine learning (ML) as a powerful approach for modeling these relationships and forecasting charge variant profiles in CHO cell-based mAb process development. We summarize supervised learning and regression methods used to link process conditions with charge heterogeneity and present case studies showing ML’s role in reducing acidic and basic variants. We also discuss challenges related to data quality, model interpretability, scalability, and regulatory compliance. Finally, we propose a roadmap for adaptive, ML-driven optimization strategies for bioprocess development, aligned with Quality-by-Design principles.
KEYWORDS: Monoclonal antibody, charge heterogeneity, machine learning, medium optimization, bioprocessing, quality by design
Introduction
Monoclonal antibodies (mAbs) are invaluable biopharmaceuticals, being essential for treating complex conditions such as cancer, autoimmune disorders, and infectious disease.1 MAbs stand out as exceptional candidates for precision therapies and large-scale production. Unlike small-molecule drugs, they offer increased specificity, fewer off-target effects, reduced risk of drug interactions, and superior therapeutic efficacy, making them a cornerstone of modern medicine.2,3 Over half of recent biopharmaceutical approvals are attributed to mAbs.4 Their adoption has considerably advanced innovation and strengthened the global market, driven by engineering breakthroughs, rising demand for biosimilars, and the increasing prevalence of chronic diseases.5,6 While mAbs have revolutionized modern medicine, maintaining their consistent quality remains a major challenge because of the complex manufacturing processes and product variations that can affect their safety and effectiveness.7,8
One of the biggest challenges in maintaining consistent quality of mAbs is controlling charge variants, which are a critical quality attribute (CQA) that can greatly affect their stability, bioavailability, pharmacokinetics, efficacy, and safety.9,10 Heterogeneity in charge variants mainly arises from post-translational modifications (PTMs), such as deamidation, glycosylation, and oxidation, forming both acidic and basic charge variants.11 These charge variants can disrupt the structural integrity and biological activity of mAbs, influencing their tissue distribution, pharmacokinetics, and overall effectiveness as therapeutic agents.10,12 Charge variation is among the most common sources of heterogeneity in recombinant mAbs, presenting substantial challenges for ensuring batch-to-batch consistency and streamlining downstream processing.9,13 Given its critical impact on drug safety and efficacy, regulatory agencies like the Food and Drug Administration (FDA) and European Medicines Agency (EMA) have established strict guidelines to closely monitor and strategically control charge variation during process development and optimization of mAbs.14,15
Several key factors influence the quality of recombinant proteins in biopharmaceutical manufacturing, including the choice of cell lines, the culture media, and the optimization of key process conditions.16 Among these factors, culture conditions and medium components are critical, as they directly impact cellular metabolism.17 Culture conditions, such as pH,18 temperature,19 ambient light,20 and cell culture duration,21 are critical in driving non-enzymatic reactions that influence cellular metabolism and PTMs.16 Modifying these parameters can either amplify or diminish unwanted changes, directly affecting the distribution of charge variants and the overall quality of the final product.9 The composition of the culture medium, including feed components and specific chemical supplements, also plays a pivotal role in modulating CQAs, such as glycosylation patterns, aggregation, and charge heterogeneity.22–24 Effective management of these factors throughout the bioreactor process is essential for minimizing the impact of accumulated byproducts and ensuring consistent product quality.25,26
In recent years, several approaches have been used to optimize culture parameters. This includes traditional methods like one factor at a time (OFAT) and design of experiments (DOE), more advanced techniques such as multivariate data analysis (MVDA) and Systems Biology (SB), and cutting-edge tools like machine learning (ML).1,27 Although traditional methods like OFAT and DOE have been essential for optimizing culture parameters, they often fail to address the complex interactions between culture parameters and cellular processes.28 Advanced techniques such as MVDA and SB have provided invaluable insights into these complexities, but, they still encounter challenges, including limited scalability and the inability to directly optimize specific properties, such as charge variants.29,30 Conversely, ML utilizes large datasets and advanced algorithms to uncover hidden patterns and predict the optimal conditions for protein production, even when the underlying mechanisms are not fully understood.1 By analyzing factors such as pH, temperature, cell culture duration, and nutrient composition, ML provides important insights into the intricate connections between culture parameters and CQAs, including charge variants.21,31,32 ML’s innovative capabilities can compensate for shortcomings of traditional methods and enable a new era of precision and efficiency in bioprocess optimization.
Here, we examine the crucial role of culture conditions and medium components in influencing charge heterogeneity, which is an important quality attribute in mAb bioprocess optimization. We investigate the causes and effects of charge variants and emphasize the transformative potential of ML in optimizing culture parameters and improving CQAs. Through case studies, we demonstrate how ML effectively links culture parameters to charge variants and outline a proposed workflow for ML-driven optimization in industrial settings. Finally, we discuss the challenges and opportunities of implementing ML in scalable, efficient, and high-quality biopharmaceutical processes, providing a roadmap for future innovations.
Charge heterogeneity in MAbs
Charge heterogeneity in mAbs refers to various molecular variants within the same product batch, exhibiting distinct net surface charges. These variants are typically categorized into three primary groups: main species, acidic species, and basic species.10 The main species is the dominant form of a mAb, typically appearing as the primary peak in chromatographic analysis.33 Although regarded as the desired product, it often contains common modifications, such as converting N-terminal glutamine to pyroglutamate, removing C-terminal lysine from the heavy chain, and adding neutral oligosaccharides to the standard asparagine site within the CH2 domain.34 Acidic variants, which possess a more negative charge than the main species, frequently arise from PTMs such as the deamidation of asparagine residues, widely regarded as the primary driver of these variants. Acidic species also stem from sialylation of glycans, glycation, or the presence of terminal acidic groups.35–37 In contrast, basic variants have a higher positive charge than the main species due to incomplete removal of C-terminal lysine, along with modifications such as C-terminal amidation, formation of succinimide, N-terminal glutamate conversion, or other structural changes.33 The proportions of these charge variants can change based on the production process, cell line, and storage conditions.9
The charge heterogeneity seen in mAbs results mainly from various PTMs, both enzymatic and non-enzymatic, that occur during biomanufacturing and storage, in addition to inherent contributions from the host cell line.38–40 Acidic variants often arise when asparagine residues undergo deamidation, a process that converts them into aspartic acid or isoaspartic acid, adding negative charges to the molecule. This process is accelerated under elevated pH and temperature.13,33 Other important factors that contribute to acidic species include the enzymatic addition of sialic acids to N-glycans and the non-enzymatic glycation of lysine residues, both of which alter the net charge by increasing negative charges or masking positive ones.15,22 Additionally, reactive carbonyl species like methylglyoxal can modify arginine residues, which further decreases their basicity.41 In contrast, basic variants usually arise when C-terminal lysine residues are not fully removed or when N-terminal glutamine fails to cyclize into pyroglutamate, both of which result in the retention of positive charges.40,42 Other modifications, such as succinimide formation, which temporarily neutralizes negative charges, and enzymatic C-terminal amidation, can also help increase base specificity.15,38 Furthermore, oxidative changes to methionine and tryptophan residues, along with variations in disulfide bond patterns, can lead to conformational changes that subtly affect chromatographic behavior, even if they do not directly modify the molecule’s overall charge.9,33,43 Cell line-specific factors, including glycosylation profiles, enzyme activity levels, and metabolic state, play a crucial role in influencing the distribution of charge variants.16,31 A comprehensive overview of these mechanisms and their implications is presented in Table 1.
Table 1.
Characteristics and impact of charge variants in mAbs.
| Feature | Acidic | Main | Basic | Ref. |
|---|---|---|---|---|
| Net Charge | More negative/Lower pI | Expected net charge/pI for the target molecule profile | More positive/Higher pI | Xie et al.44 |
| Chromatographic Profile | Elutes earlier in Cation Exchange Chromatography (CEX)/Migrates towards anode in cIEF | A principal peak in CEX/cIEF; Represents the target elution/migration time | Elutes later in CEX/Migrates towards cathode in cIEF | Beck et al.9 Xie et al.44 |
| Representative PTMs | Deamidation (Asn → Asp/isoAsp) - Sialylation (N-glycans) - Glycation - Trp Oxidation (sometimes) - Arg Modification (e.g., by methylglyoxal) | N-terminal Pyroglutamate Formation - Complete C-terminal Lys Removal - Core/Neutral N-glycosylation (e.g., G0F, G1F) | Incomplete C-term Lys Removal - Incomplete N-term PyroGlu Formation - Succinimide Formation - C-terminal Amidation - Asp Isomerization to isoAsp (sometimes) - Met Oxidation | Brorson and Jia38 Du et al.33 Gangwar et al.32 |
| Key Origin Drivers | High Temperature; High pH (esp. for deamidation); Long culture duration; High glucose/sugar; Oxidative stress (DO, light, certain metals); Nutrient depletion; Cell stress/lysis | Optimal biosynthesis, Efficient enzymatic processing, Controlled process conditions | Suboptimal enzymatic processing (esp. carboxypeptidase, glutaminyl cyclase), Low pH (esp. isoAsp/succinimide), Specific metal ion levels (e.g., Zn/Cu affecting Cp), Cell line-specific enzyme levels | Gangwar et al.29 Ha et al.32 Sissolak et al.36 Weng et al.21 |
| Analytical Detection Methods | CEX, cIEF/iCIEF, LC-MS, Peptide Mapping | CEX, cIEF/iCIEF, LC-MS, Peptide Mapping | CEX, cIEF/iCIEF, LC-MS, Peptide Mapping | Gupta et al.35 Horvath et al.45 Wagner-Rousset et al.15 |
| Stability Profile | Often linked to degradation pathways (deamidation, oxidation); Potential for increased aggregation/fragmentation propensity | Represents target stability profile; Generally stable within specifications | May contain degradation intermediates (succinimide, isoAsp); Can impact conformation/solubility | Hazeltine et al.46 Kaur et al.26 Luo et al.42 Zheng et al.47 |
| Functional Implications | Altered antigen binding (if in CDR), ↓ ADCC/CDC (sialylation), altered PK, immunogenicity (sometimes) | Represents the target activity profile; Benchmark for comparison. | Altered FcRn binding/PK (C-term Lys), potentially altered potency or clearance (conformational variants), immunogenicity (sometimes, e.g., leader seq.) | Chen et al.48 Hong et al.49 Zhang et al.50 Zheng et al.51 |
| Regulatory Perspective | CQA; Must be characterized, monitored, and controlled within justified limits. | Defined as the Reference Standard profile; CQA; Must be characterized, monitored, and controlled within justified limits | CQA; Must be characterized, monitored, and controlled within justified limits | Horvath et al.45 Rouiller et al.52 Xu et al.53 |
Even at low levels, charge variants can affect the therapeutic effectiveness, stability, and regulatory compliance of mAbs.10 These variants typically arise from PTMs that can affect pharmacokinetics, pharmacodynamics, and immunogenicity, collectively influencing the clinical effectiveness of mAbs.4,36 For example, acidic variants of antibodies may decrease their ability to bind to target molecules, reducing their therapeutic effectiveness.13,39 Conversely, basic variants might change how antibodies interact with Fc receptors, potentially shortening their half-life or causing undesirable immune reactions.54 Even at low levels, these charge variations can affect the treatment’s effectiveness and stability, making it more challenging to maintain consistency across batches and adding complexity to the development and regulatory approval processes.22,54,55 Given these potential impacts, regulatory bodies such as the FDA and EMA recognize charge heterogeneity as a CQA that requires careful control.15,56 As such, addressing these variations effectively is key to achieving reliable and scalable mAb production with consistent quality.9
Understanding the charge variability in mAbs relies on various advanced analytical techniques.40 Capillary isoelectric focusing (cIEF) provides a high-resolution view of charge variants based on isoelectric points, making it essential for analyzing acidic, basic, and main species in mAb formulations.40,54 Its accuracy makes it invaluable in both development and quality control. Ion exchange chromatography (IEX), especially cation exchange chromatography (CEX), refines the separation of different charge variants by capitalizing on subtle charge differences.33 Through carefully tuned pH and salt gradients, it can tease apart distinct molecular species with remarkable precision.54 In addition, liquid chromatography-mass spectrometry (LC – MS) delivers a high-resolution snapshot of PTMs that affect mAb charge profiles.39 Complementary techniques like electrospray ionization (ESI) and detailed peptide mapping take it a step further, uncovering deeper molecular details for a more thorough and precise analysis.9,39 Innovative techniques such as imaging capillary IEF (icIEF) and integrated chromatography-mass spectrometry methods are advancing charge variant analysis by providing enhanced sensitivity and resolution, ultimately leading to improved stability, safety, and efficacy of mAbs.54,55
Culture condition and media component optimization for mAbs
Optimizing culture conditions and culture medium components is a primary strategy for reducing charge heterogeneity in mAb production, as it directly affects the PTMs that determine the charge profile. Culture conditions can be categorized into physical (e.g., temperature, pH), operational (e.g., culture duration), and environmental (e.g., ambient light, dissolved gases), all of which affect the cellular metabolic state and the kinetics of PTMs.18,20,31,37,44 Temperature can modulate charge heterogeneity; reducing the culture temperature (for instance, to 32–34 °C) can decrease metabolic activity and slow non-enzymatic reactions like deamidation, thus lowering the formation of acidic species.37,47 However, this drop in temperature may also affect enzymes such as carboxypeptidases, which could lead to incomplete removal of C-terminal lysine and, as a result, an increase in basic variants.50
pH, though easier to control, plays an equally important role. Higher pH levels typically accelerate deamidation and reduce sialylation, both leading to an increase in acidic species.31 On the other hand, lowering the pH can encourage sialylation and enhance glutamine cyclization at the N-terminus, which in turn shifts the charge profile toward more basic species.18,44 Besides temperature and pH, factors like culture duration also influence charge heterogeneity. For instance, in GS-CHO cells, longer culture periods (e.g., from day 5 to day 14) result in the accumulation of PTMs, which in turn increase the proportion of acidic variants.18 Environmental factors such as ambient light and dissolved gases (pCO₂ and pO₂) also influence PTMs.20,31 For instance, light exposure can lead to the oxidation of residues such as tryptophan, which increases acidic variants.20 Higher levels of pCO₂ lead to the formation of acidic species, while variations in pO₂ levels affect the production of basic compounds.31 Table 2 provides a comprehensive summary of studies analyzing these parameters over various Chinese hamster ovary (CHO) cell lines.
Table 2.
Key culture parameters influencing charge Heterogeneitya.
| Parameter | Cell line | Rangeb | Effectc |
Ref. | |
|---|---|---|---|---|---|
| Acidic | Basic | ||||
| Temperature (ºC) | CHO-K1 | 31, 34, 37 | Affected | Increases | Sissolak et al.36 |
| CHO-DG44 | 35, 36.5 | Decreases | Affected | Weng et al.21 | |
| CHO | 29, 32, 35, 37 | Increases | N/A | Tang et al.37 | |
| CHO | 33, 35, 37 | Decreases | N/A | Kishishita et al.57 | |
| CHO-K1 HP & LP | 33 → 37 | Decreases | Increases | Zhu et al.19 | |
| CHO-K1 | 30, 32, 34 | Increases | Decreases | Zheng et al.47 | |
| CHO | 33, 35, 37 | N/A | Increases | Luo et al.42 | |
| CHO | 32, 35, 37 | Decreases | Increases | Zhang et al.50 | |
| CHO | 31 → 35 | Decreases | N/A | Yang et al.58 | |
| CHO | 34 → 36.5 | Increases | N/A | Chung et al.25 | |
| CHO-K1 GS | 30, 32, 33.5, 35, 36.5 | Decreases | Increases | McHugh et al.59 | |
| rCHO | 30, 33, 37 | Decreases | Increases | Yoon et al.60 | |
| GS CHO1, DG44 CHO2 | 32, 33, 34, 35, 36.5, 37.5, 38.5 | Decreases | Increases | Xu et al.61 | |
| CHO | 36, 37, 38 | Increases | N/A | Horvath et al.45 | |
| pH | CHO | 6.8, 7, 7.2 | Increases | Affected | Brunner et al.31 |
| CHO-S | 7.05, 7.15 | Increases | Decreases | Rouiller et al.52 | |
| GS‐CHO | 6.7, 6.9, 7.1 | Increases | Decreases | Lee et al.18 | |
| CHO | 5.5 → 7 | Affected | N/A | Trexler‐Schmidt et al.62 | |
| GS-CHO | 6.75±0.1 → 6.95±0.1 | Decreases | N/A | Xie et al.44 | |
| CHO | 6.85±0.05 → 7.15±0.05 | N/A | N/A | Zheng et al.51 | |
| CHO | 7 → 7.3 | Increases | N/A | Horvath et al.45 | |
| Culture duration (days)d | CHO-DG44 | 11, 12, 13, 14, 15 | Increases | N/A | Weng et al.21 |
| CHO | 3, 6, 9, 12, 15 | Increases | N/A | Tang et al.,37 | |
| CHO | 12 vs. 14 | N/A | Decreases | Luo et al.42 | |
| GS-CHO | 5, 8, 10, 12, 14 | Increases | N/A | Lee et al.18 | |
| CHO-K1 GS | 3, 8, 12 | Increases | N/A | Park et al.63 | |
| CHO-K1 GS | 3, 6, 9 | Increases | N/A | McHugh et al.59 | |
| Ambient Light (lux) | CHO | 1000, 3000, 10000 | Increases | Increases | Mallaney et al.20 |
| Feed media age (weeks)e | CHO | 3 → 10 | Increases | N/A | Chung et al.25 |
| Passage number | GS-CHO | PN-10 → PN-60 | Decreases | Decreases | Kaur et al.26 |
| CHO DG44 | P5 → P33 | N/A | Increases | Xu et al.53 | |
| pCO2%) | CHO | 5 → 20 | Affected | Increases | Brunner et al.31 |
| pO2%) | CHO | 10 → 40 | N/A | Increases | Brunner et al.31 |
| Process mode | CHO | Traditional Fed-batch vs. Concentrated Fed-batch | Decreases | N/A | Yang et al.58 |
| Seed density (PCV%) | CHO | 0.12 → 0.36 | Affected | N/A | Horvath et al.45 |
aAll products are mAbs, and all media used are chemically defined.
b“→” indicates a change from one value to another; “vs.” compares two values; commas separate multiple values for the same parameter.
cEach parameter’s impact on charge variants is classified as Increase, Decrease, Affected (without a consistent trend), or N/A (no data or negligible impact reported).
dAll the studies included in this table for culture duration employ a fed-batch cultivation method.
eThis parameter primarily refers to the duration of storage for prepared liquid media, which can affect stability and component integrity. Dry powder media are generally unaffected until reconstitution.
The effects of culture parameters on mAb charge heterogeneity, as summarized in Table 2, often differ across studies. These variations are due to the inherent complexity of bioprocesses and context-specific factors such as the CHO cell line or clone, the structural features of the expressed antibody, and interactions among variables like pH, temperature, and nutrient composition.10,21,36 Process configurations (e.g., fed-batch regimes) and media-related factors (e.g., age and formulation) also significantly influence the outcomes.25,26 Rather than indicating contradictions, this variability highlights the biological diversity of upstream systems and emphasizes the importance of case-specific, data-driven process development strategies.9,56
Along with the culture conditions, the composition of the culture medium and feed is key in shaping the charge variant profile of mAbs. Ingredients like glucose and trace metals in the media can either promote or hinder specific PTMs, influence metabolic pathways, and ultimately shape the overall charge variant distribution.22,23,31,36 Amino acids are essential building blocks for protein synthesis and key modulators of PTMs.64 For instance, adequate levels of asparagine act as a substrate for deamidation and promote acidic variants.52 In contrast, high levels of lysine and arginine can inhibit carboxypeptidases, restrict lysine cleavage at the C-terminal, and increase basic variants.24 Similarly, trace metal ions such as zinc, copper, iron, and manganese are critical cofactors for enzymes involved in a range of PTMs, such as carboxypeptidase activity, amidation, and glycosylation, while also catalyzing oxidation reactions.22,37,65 Most importantly, their equilibrium and speciation can affect enzyme activity and charge profile. For example, excess copper (e.g., 0.004–0.4 mM) can catalyze oxidative reactions, contributing to charge heterogeneity.21,66 Moreover, reducing sugars, mainly glucose and galactose, at high concentrations (> 15 g/L) promote the formation of acidic species via non-enzymatic glycation and influence the distribution of charge species via sialylation.36,52,67 Furthermore, certain vitamins, supplements, hydrolyzates, sodium butyrate, and accumulated metabolic byproducts can modulate cellular metabolism, redox state (as antioxidants or prooxidants), and PTM enzymatic activities, adding further layers of complexity to the optimization of the charge profile.25,29,44,49,68 Table 3 summarizes key findings from studies on the effects of various parameters on mAb charge variants in different CHO expression systems.
Table 3.
Key media components influencing charge Heterogeneitya.
| Components | Cell line | Rangeb | Effectsc |
Ref. | |
|---|---|---|---|---|---|
| Acidic | Basic | ||||
| Glucose (g/L) | CHO-K1 | 2 → 15 | Increases | N/A | Sissolak et al.36 |
| CHO | 11.5–23 | Increases | N/A | Quan et al.67 | |
| Copper (mM) | CHO-DG44 | 0.004, 0.04, 0.15, 0.4 | N/A | Increases | Weng et al.21 |
| CHO-GS | 0 → 0.0315 | Decreases | N/A | Gangwar et al.22 | |
| CHO-DXB11 | 0, 0.00015, 0.00025, 0.0005, 0.001, 0.002, 0.003 | Decreases | Increases | Yuk et al.43 | |
| CHO | 1X → 4X | N/A | Increases | Luo et al.42 | |
| CHO-K1 | 0.0004, 0.00055, 0.00075, 0.001 | Affected | Increases | Kaschak et al.40 | |
| CHO-S | 0x, 1x, 3x | Affected | N/A | Rouiller et al.52 | |
| CHO-K1 | 0, 0.005, 0.05, 0.1 | Decreases | N/A | Chaderjian et al.66 | |
| CHOK1SV | N/A (increased) | Decreases | Decreases | Hazeltine et al.46 | |
| Iron (mM) | CHO-GS | 0.0197, 0.3008, 0.3581, 1.6240 | Increases | N/A | Gangwar et al.22 |
| CHO/dhFr | 0.01, 0.018, 0.075 | Increases | N/A | Vijayasankaran et al.68 | |
| CHO-S | 0, 0.1343, 0.2686 | Increases | N/A | Rouiller et al.52 | |
| CHO DG44 | 0.01, 0.02, 0.03, 0.05, 0.11 | Increases | Affected | Xu et al.53 | |
| CHO | Control Medium vs. Control Medium + 0.08 mM Iron | Decreases | N/A | Chung et al.25 | |
| CHO-GS | 0.1, 0.2, 0.4 | Increases | Increases | Gangwar et al.32 | |
| Zinc (mM) | CHO-DG44 | 0, 0.004, 0.03, 0.04, 0.07, 0.08, 0.1, 0.13, 0.15 | Decreases | N/A | Weng et al.21 |
| CHO-GS | 0.0061, 0.0138, 0.1530, 0.0306 | Increases | Decreases | Gangwar et al.22 | |
| CHO | 1X → 12X | N/A | Decreases | Luo et al.42 | |
| CHO-S | 0X, 1X, 3X | Affected | N/A | Rouiller et al.52 | |
| CHO-GS | 0.060, 0.090, 0.180 | Decreases | Decreases | Gangwar et al.32 | |
| Manganese (mM) | CHO-DG44 | 0, 0.004, 0.040 | N/A | N/A | Weng et al.21 |
| CHO-GS | 0.00091, 0, 0.00182, 0.0273 | Increases | N/A | Gangwar et al.22 | |
| CHO-S | 0, 0.0002, 0.001 | Affected | Affected | Rouiller et al.52 | |
| CHOK1SV | N/A (increased) | Decreases | Decreases | Hazeltine et al.46 | |
| Magnesium (mM) |
CHO-GS | 0.8023, 0.8147, 0.8393, 0.8517 | N/A | N/A | Gangwar et al.22 |
| Cobalt (mM) | CHO-GS | 0, 0.0017, 0.00509 | Increases | N/A | Gangwar et al.22 |
| Nickel (mM) | CHO-GS | 0, 0.017, 0.0341 | Increases | Increases | Gangwar et al.22 |
| Sodium butyrate (mM) | rCHO | 0, 0.01, 0.05, 0.2, 1, 2, 4 | Decreases | Increases | Hong et al.49 |
| Selenium (mM) | CHO-S | 0, 0.0002, 0.001 | Affected | N/A | Rouiller et al.52 |
| Biotin (mM) | CHO-GS | 0, 0.0073, 0.0094 | Decreases | Increases | Gangwar et al.22 |
| Ascorbic acid | CHO | 0, 0.568, 5.678, 17.034 | Increases | N/A | (Chumsae et al.41 |
| Surfactants (%v/v, %w/v) | CHO-DUX B11, CHO-K1 | Type: PS80, PS20, P188 | N/A | N/A | Hossler et al.69 |
| Concentration: 0.0025% − 10% | Decreases | Increases | Gangwar et al.22 | ||
| Choline (mM) | CHO-GS | 0.2414, 0.6615, 0.873, 0.912 | N/A | Affected | |
| Carboxypeptidase B (w/w) | CHO-GS, CHO-DG44 | 0, 1:100, 1:10000 | Increases | Decreases | Xu et al.70 |
| CHO | 0, 1:125 | Affected | Decreases | Kim et al.71 | |
| Methylglyoxal (mM) | CHO | 0 → 2.8 | Increases | N/A | Chumsae et al.39 |
| Folic acid (mM) | CHO-GS | 0.0062, 0.008, 0.0332, 0.0419 | N/A | N/A | Gangwar et al.22 |
| CHO/dhFr | 0, 0.2232, 0.4463 | Increases | N/A | Vijayasankaran et al.68 | |
| Dextran sulfate (g/L) | CHO-K1, DG44 | MW: 4000, 15000, 40000 | Affected | Decreases | Hyoung Park et al.72 |
| Concentration: 0.1, 0.3, 0.5, 1 | |||||
| Hydrolyzate (g/L) | CHO-K1 | 0 → 0.1 | Decreases | Increases | Zheng et al.47 |
| Taurine (mM) | CHO | 0 → 13.2 | N/A | Decreases | Liu et al.73 |
| Pantothenic acid (mM) | CHO-GS | 0.0084, 0.0138, 0.0214, 0.0559 | N/A | Decreases | Gangwar et al.22 |
| Arginine (mM) | CHO | 2 → 10 | N/A | Increases | Zhang et al.24 |
| Rutine (g/L) | CHO | 0.005 → 1 | Decreases | Increases | Hossler et al.55 |
| Rosmarinic acid (g/L) | CHO | 0 → 0.36 | Decreases | N/A | Chung et al.25 |
| Lysine (mM) | CHO | 2 → 10 | N/A | Increases | Zhang et al.24 |
| Epigallocatechin gallate (g/L) | CHO | 0.1 → 1 | Decreases | Increases | Hossler et al.55 |
| Uridine (mM) | CHO | 0 → 6 | Decreases | Increases | Niu et al.74 |
| Thiamine (mM) | CHO-GS | 0.0129, 0.0134, 0.0159, 0.0273 | N/A | Affected | Gangwar et al.22 |
| Galactose (g/L) | CHO | 0, 8, 19.50 | Increases | N/A | Quan et al.67 |
| CHO-S | 0, 1.802, 3.603 | Affected | N/A | Rouiller et al.52 | |
| β-glycerol phosphate (g/L) | CHO DG44 | 0 → 0.9 | N/A | Increases | Xu et al.70 |
| Spermine (mM) | CHO-S | 0, 0.075, 0.2 | Increases | Affected | Rouiller et al.52 |
| Asparagine (mM) | CHO-S | 0, 10, 20 | Decreases | Decreases | Rouiller et al.52 |
| Pyridoxine (mM) | CHO/dhFr | 0, 0.2, 0.4 | Increases | N/A | Vijayasankaran et al.68 |
| CHO-GS | 0, 0.02, 0.0211, 0.0287 | N/A | N/A | Gangwar et al.22 | |
| Riboflavin (mM) | CHO/dhFr | 0, 0.0133, 0.0266 | Increases | N/A | Vijayasankaran et al.68 |
| Cysteine (mM) | CHOK1SV | 1×(historical media) to 1×/1.5×(CD media) | Decreases | Decreases | Hazeltine et al.46 |
| Medium (%) | CHO | 85, 100, 115 | Decreases | N/A | Horvath et al.45 |
| CHO-DUX B11 and CHO-K1 | 1X CDFM, 2X CDFM (X) | Decreases | Increases | Hossler et al.69 | |
| CHO | BM-1, BM-2، BM-3، BM-4 | Affected | N/A | Tang et al.37 | |
| Cobalamin (mM) | CHO/dhFr | 0, 0.0177, 0.0354 | Increases | N/A | Vijayasankaran et al.68 |
| Tryptophan (mM) | CHOK1SV | 1×(historical media) to 0.7×/0.4×(CD media) | Decreases | Decreases | Hazeltine et al.46 |
aAll products are mAbs, and all media used are chemically defined.
b“→” indicates a change from one value to another; “vs.” compares two values; commas separate multiple values for the same parameter.
cEach parameter’s impact on charge variants is classified as Increase, Decrease, Affected (without a consistent trend), or N/A (no data or negligible impact reported).
Historically, managing charge heterogeneity by optimizing culture conditions and media components has predominantly relied on experimental and statistical methods.21,57 These conventional techniques, including the DoE, statistical modeling, OFAT, and process controls, have formed the backbone of optimization efforts to manage charge heterogeneity.29,44 OFAT refers to an approach in which only one variable is changed while others are held constant, making it easy to implement but limited in detecting interactions. DoE, in contrast, enables the simultaneous investigation of multiple variables and their interactions using structured experimental designs.75 Early efforts often used a simple OFAT approach, although its inability to assess complex parameter interactions posed limitations.76 As revealed in studies such as Wang et al.21 and Horvath et al.,45 DOE enables the simultaneous evaluation of multiple factors, such as temperature, pH, and zinc levels, to identify optimal conditions and interactions, but it is still limited by the number of testable variables and the complexity of nonlinear biological systems.77,78
Furthermore, statistical modeling techniques such as MVDA and partial least squares are widely used to predict parameter effects and analyze complex relationships, but their reliance on large datasets and linear assumptions limits their accuracy in capturing nonlinear biological responses and PTM dynamics underlying charge heterogeneity.20,25,27,31,79 Nevertheless, studies such as Chung et al. have demonstrated the use of PLS-based MVDA to correlate oxidative stress markers with acidic charge variants, providing actionable insights for media optimization and feed strategy adjustment.25 MVDA encompasses statistical methods such as principal component analysis (PCA) and partial least squares that handle multiple input variables simultaneously to uncover patterns and correlations, often serving as exploratory or dimensionality-reduction tools.80 SB combines omics data with computational modeling to optimize culture media and better understand cellular behavior.1 Despite its potential, the complexity, data demands, and inability to directly optimize specific traits, like charge heterogeneity, of SB have limited its application to charge variant optimization in mAb bioprocess development.81,82 Consequently, while traditional process controls (e.g., online pH and DO monitoring) help stabilize culture conditions during development, they are insufficient for resolving complex nonlinear interactions driving charge heterogeneity, thus highlighting the need for advanced modeling approaches like ML.16,29,56 Figure 1 offers a comparative overview of the capabilities of OFAT, DoE, MVDA, and ML across key dimensions.
Figure 1.
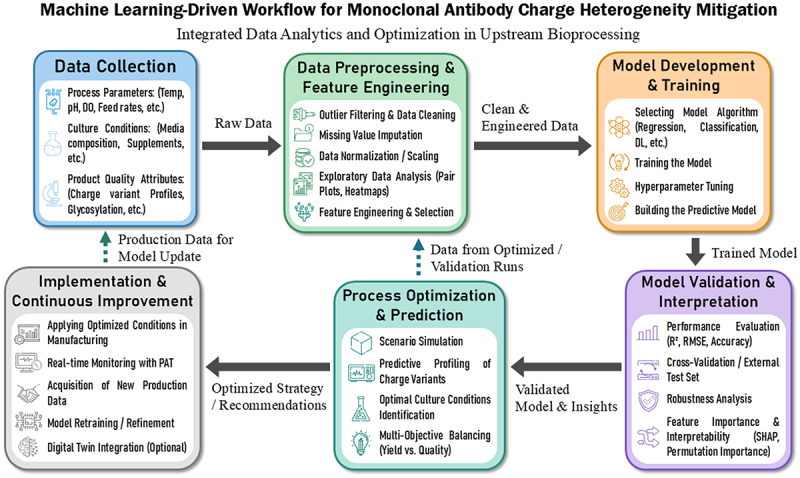
Comparative evaluation of OFAT, DoE, MVDA, and ML capabilities for bioprocess optimization. This radar chart compares four approaches, OFAT, DoE, MVDA, and ML, across six key dimensions relevant to CHO cell culture optimization for charge heterogeneity: process integration, predictive accuracy, result interpretability, industrial scalability (higher values preferred), and modeling complexity, data requirements (lower values preferred). Process integration refers to how well a method fits into the full development pipeline, including monitoring and control, while industrial scalability reflects its applicability from lab to production scale. ML (yellow) shows strong performance in accuracy and integration, though it demands more data and modeling complexity. SB offers deep biological insights but has limited direct use in charge variant prediction. The assessment is qualitative, based on published experimental and industrial studies.
Machine learning in bioprocessing
To address the challenges of optimizing mAb production in complex cell culture systems, where traditional methods often fail to capture nonlinear interactions and high-dimensional data, ML offers advanced tools for developing predictive models.83,84 As a subset of artificial intelligence, ML focuses on building algorithms that learn patterns from data, enabling them to make predictions or decisions without being explicitly programmed.1 These ML approaches, including supervised, unsupervised, and reinforcement learning, offer a robust toolkit for assaying complex bioprocessing data.85,86 Among these, supervised learning has the greatest application in biopharmaceuticals through the use of labeled datasets, as it directly links measured culture inputs, such as temperature, pH, and nutrient concentrations, to quantitative outputs such as titer or ratio of load types.83,84,87 Supervised learning algorithms are typically categorized into two main types based on the nature of the output. For regression tasks, algorithms like support vector regression, random forest, gradient boosting, and lasso/ridge regression are used to predict continuous outcomes, such as titer levels or percentages of charge variants.1 In contrast, classification tasks use methods like decision trees, support vector machines, and ensemble models, which combine the predictions of multiple individual models to improve overall performance and robustness (e.g., random forests and gradient boosting, which can also be applied to regression tasks, are specific types of ensemble models) to categorize samples into distinct qualitative classes or variant profiles.86,88 Both methods effectively model the nonlinear, multivariate relationships typical in mAb production systems.84
Beyond supervised learning, unsupervised learning is crucial for identifying hidden patterns in complex bioprocessing datasets, particularly when labeled data are limited or incomplete.85,89 Techniques such as PCA and clustering algorithms, including k-means and hierarchical clustering, identify metabolic bottlenecks, group similar culture conditions, and reduce data dimensionality for better interpretability.56,86,90 PCA, while a core technique in MVDA is also widely used in ML workflows as a preprocessing step to enhance model interpretability and manage high-dimensional inputs.56 For instance, PCA has helped reveal correlations between components of the culture medium and glycosylation profiles, facilitating the optimization of the culture medium without predefined outcomes.18 In parallel, reinforcement learning offers an adaptive, feedback-based approach that is ideal for dynamic optimization approaches in bioprocess development.84 It optimizes feeding strategies and environmental conditions to maintain CQAs, such as charge heterogeneity, during bioreactor runs.88 Although it is still relatively new in biomanufacturing, reinforcement learning holds great promise for making complex, time-sensitive decisions, especially when used alongside simulation tools to address the challenges of limited experimental data.1,86 These ML approaches provide essential tools for developing predictive models and optimizing inputs to achieve product quality.1,83 They are used to build models for critical quality attributes and to optimize culture conditions and media components.91
The development of predictive models for CQAs, which play a pivotal role in supporting quality by design (QbD) and process analytical technology (PAT) frameworks,84,88,92 has been strongly influenced by the application of ML. The use of proxy measurements that correlate with desired CQAs, permits effective assessment for QbD during process development,90,93 as charge heterogeneity in mAbs heavily depends on earlier process factors.56 Supervised learning models use labeled datasets to connect critical process parameters, such as temperature, pH, and dissolved oxygen, with critical material attributes, such as media composition28,85,94 This relationship helps predict the outcomes of CQAs, which can be continuous (e.g., percentage of acidic species) or categorical (e.g., compliant vs. non-compliant profiles).89 This approach enables early detection of deviations and proactive quality management within process development using the principles of QbD and PAT.90
Current applications of ML in optimizing cell culture conditions and medium components are proving invaluble in upstream bioprocessing for mAb process production.1,95,96 The complexity, high dimensionality, and dynamic nature of upstream cell‑culture bioprocesses, along with the increasing availability of rich process data, particularly benefit from more advanced ML approaches.97,98 A notable application is their use in predicting key performance indicators and critical process parameters.83,97 For instance, artificial neural networks and hybrid deep learning architectures, such as convolutional neural networks (CNN) and long short-term memory (LSTM), have been successfully applied to model and optimize CHO cell culture processes, demonstrating improved productivity predictions.96,97 Furthermore, ML can be used in media development and optimization. This involves applying multivariate analysis and active learning strategies, such as gradient boosting decision trees, to identify influential components and optimize their concentrations to enhance cell culture performance.83,99,100 These models can also predict the dynamic concentrations of essential media components, such as amino acids, and interpret the effects of complex raw materials, such as yeast extract, on protein expression.101,102
ML is not only limited to static optimization; it also encompasses dynamic optimization of cultivation parameters and establishes quantitative links between upstream conditions and product CQAs.95,98,103 Advanced techniques like deep reinforcement learning are being studied for dynamic adaptive optimization of multiple process parameters during development, allowing for dynamic adjustments during cultivation to maintain optimal conditions and balance objectives such as productivity and product quality.95 By combining process data with hybrid models that blend mechanistic insights and ML, these techniques improve prediction accuracy for CQAs like glycosylation and metabolic indicators, revealing how dynamic tweaks to culture and media conditions impact product quality.103,104 Importantly, ML models, often supported by PAT tools such as Raman spectroscopy, are increasingly capable of mapping upstream variables to CQAs, including glycosylation and, notably, charge heterogeneity.97,98,105 In this context, PAT tools serve as high-content sensors that facilitate data-driven modeling and upstream optimization during development, distinct from their typical use in real-time process control.106 ML enables proactive optimization and fine-tuning of upstream conditions in the upstream process by correlating culture conditions and growth medium components with charge variants.88,97 This approach empowers data-driven strategies that converge with QbD principles and improve management over critical quality characteristics such as glycosylation and charge heterogeneity.56,99
Integrating machine learning for charge heterogeneity mitigation
ML can help one understand, predict, and ultimately optimize charge heterogeneity in mAbs during upstream processing.107 Accordingly, ML, applied to charge heterogeneity, establishes quantitative relationships between upstream process parameters, medium composition, and resulting charge variant profiles by identifying complex patterns in large-scale bioprocess datasets.56 ML models are uniquely suited to this task, as they learn patterns from comprehensive datasets that pair controlled upstream variables (such as pH, temperature, feeding strategies, and media component concentrations) with high-resolution charge variant analysis data.37,45,95,107 By training supervised learning algorithms on this data, including regression models, classification models, and tree-based methods, the model learns the complex and often non-linear interplay between these factors and outcomes such as the percentage of acidic/basic species or the classification of acceptable profiles.29,108 This capability is vital because parameters such as pH and temperature are well-known to affect charge-shifting PTMs such as deamidation, and ML can model the combined and interacting effects of the entire upstream environment.109 Building on previous efforts in predicting PTMs like glycosylation, applying similar ML frameworks to charge variant data enables the identification of key upstream drivers of charge heterogeneity and supports targeted process optimization.1,110
Data-driven studies in CHO cells demonstrate the ability to quantitatively and clearly link specific upstream inputs such as medium composition and culture parameters to charge variant outcomes. For instance, Gangwar et al. introduced an ML framework for CHO-GS cells cultured in a chemically defined basal medium to predict CQAs and optimize the culture medium. Their approach used 15 different regression models, combined with feature selection techniques, to analyze the impact of metal ions (as inputs) on acidic and basic charge variants (as outputs) (Table 4). This work identified iron and zinc as input variables most strongly influencing charge heterogeneity, and by fine-tuning their concentrations, the researchers achieved a charge variant profile closely resembling the reference innovative product. Compared to the conventional DOE approach, which reported an R2 of 0.85, the ML-based optimization demonstrated superior predictive accuracy with an R2 of 0.95, highlighting the enhanced performance of ML models in predicting charge variant outcomes and optimizing medium composition.29
Table 4.
Machine learning/Data-driven optimization of culture parameters influencing charge variant.
| Cell line | Parameter | Media | Product | Algorithm | Charge Variant | Ref. |
|---|---|---|---|---|---|---|
| CHO-GS | Cobalt, Nickel, Magnesium, Manganese, Zinc, Iron, Copper | CD CHO® | IgG1 (Trastuzumab) | Random forest regression, linear regression, lasso regression, decision tree regression, extra tree regression, ridge regression, lasso least regression, Bayesian ridge, catboost regression, Huber regression, extreme gradient boosting, gradient boosting regression, elastic net, support vector regression, and k-nearest regression | Iron reduces acidic variants; Zinc increases basic variants. | (Gangwar et al.,29) |
| CHO-GS | Glucose, Lactate, pH, Dissolved Oxygen, Temperature, Ammonia | CD CHO® | IgG1 | CNN, LSTM, Bayesian Optimization, Random Forest, K-nearest Neighbor | 39.5% reduction in total acidic and basic variant levels | (Lu et al.,97) |
| CHO-S and CHO-GS | pH, temperature, initial seeding density | Serum-free commercial medium | IgG1, IgG4, (IgG+IgM) | Statistical Modeling, PCA, Transfer Learning, Active Learning | Higher pH increases Acidic Variants and decreases Basic Variants; Temperature shifts also inversely affect charge variants. | (Kumar et al.,111) |
Extending beyond medium composition, Lu et al. showcased how ML can effectively enhance the entire CHO cell culture process. They used a deep learning model – a combination of CNN-LSTM networks – to predict and fine-tune key process parameters such as nutrient levels, temperature, and pH (inputs). Applying this framework, they achieved significant improvements in product-related outputs, including a 28.1% increase in titer, a 19.1% increase in overall process yield, and a 39.5% reduction in charge variant percentage, demonstrating the comprehensive upstream process improvement enabled by ML. The model achieved an R2 of 0.956 and an RMSE of 0.082, demonstrating a substantial improvement in predictive accuracy compared to traditional models, which generally yield lower R2 values and higher error metrics.97 Furthermore, Kumar et al. presented a data-driven approach to optimize key physical process inputs – specifically pH, temperature shift, and initial seeding density – for achieving the desired product quality outputs, such as N-glycan and charge variant profiles, in mAb biosimilar production. Using multivariate statistical modeling across multiple CHO cell lines, they examined how variations in these input parameters affected the resulting CQAs. Their findings revealed consistent relationships among quality characteristics. By applying transfer learning and active learning from prior projects, they effectively reduced the experimental workload while ensuring that target output attributes were successfully achieved.111
Implementing an ML-based strategy to mitigate charge heterogeneity in mAb production typically follows a structured workflow that integrates data collection, model development, and process optimization.1,21 The first step involves data collection, where high-quality datasets from upstream bioprocesses, including CHO cell culture conditions and media compositions, are collected alongside analytical outputs such as charge variant profiles.112,113 Subsequently, this step involves the preparation of the gathered raw data through preprocessing techniques. This critical step ensures data quality and consistency by cleaning the dataset, handling missing or noisy values, normalizing variables, and applying feature engineering or selection to extract the most relevant inputs affecting charge heterogeneity.1,107,114,115 After preprocessing, appropriate ML models are selected and trained. Based on the specific goal, such as predicting the levels of acidic or basic species using regression models or classification models to classify heterogeneous profiles, appropriate algorithms are selected to achieve high prediction accuracy (e.g., R2 > 0.95) and interoperability.29,111,116 Figure 2 illustrates the typical stages involved, from initial data handling to operational implementation.
Figure 2.
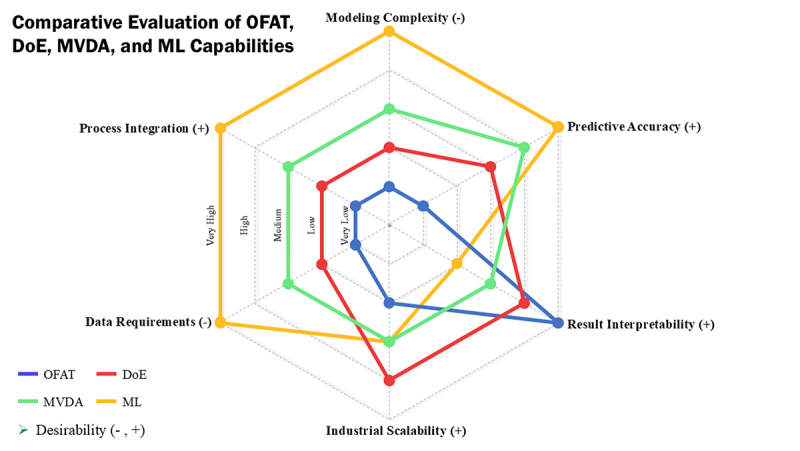
Machine learning-driven workflow for monoclonal antibody charge heterogeneity mitigation.
After training, the models undergo rigorous validation and evaluation using independent datasets distinct from the training dataset.107 This process assesses their predictive accuracy, robustness, and generalizability across different process conditions and scales, often using cross-validation techniques and real-time bioprocess data, as demonstrated by Kumar et al.110,111,115 Following this stage, model interpretation can be a valuable step to gain insight into how specific upstream parameters, such as pH or media components such as iron and zinc, affect charge heterogeneity, enhancing process understanding and guiding further optimization.18,52,107 Once model performance is validated, it is integrated into the bioproduction workflow for prediction and optimization purposes.1 The model can simulate different parameter combinations to predict charge profiles and identify optimal culture conditions and media compositions that minimize undesirable variants while meeting other production objectives.117,118 Optimal culture conditions are determined by generating candidate conditions, which are input into the trained ML model to predict charge profiles.28 The conditions yielding the best outcomes, such as minimal charge heterogeneity, are selected as optimal, enabling precise adjustments that enhance mAb production efficiency.29 Finally, continuous monitoring and improvement, incorporating active learning principles and feedback loops, ensure the long-term relevance of the model, where new process data is used to retrain and periodically update the model, aligning with Bioprocess 4.0 principles as a forward-looking framework for bioprocess optimization and product consistency.28,119,120 This iterative workflow enables proactive optimization of charge heterogeneity profiles, increasing yield and quality in mAb bioprocesses.121
Although promising, scaling and validating ML models to industrial bioreactors poses challenges. Scalability refers to the ability of models developed at laboratory scales to accurately predict charge variant profiles in large-scale production bioreactors.122 This transition is challenging due to scale-dependent differences in mixing, mass transfer, and changing process dynamics.93,123 These factors can all impact cellular metabolism and the kinetics of PTMs.1 In addition, models must be sufficiently robust to handle the increasing data volume and variability that arise from industrial operations.124 Validation is equally critical and requires rigorous evaluation using independent manufacturing datasets to demonstrate accuracy, robustness, and reliability across batches, raw materials lots, and operating sites.84,125 Validation strategies include k-fold cross-validation, use of dedicated test sets, and prospective studies to ensure that models meet stringent regulatory standards for biopharmaceutical applications.93,126 Ultimately, successfully scaling and validating ML models to reliably predict complex CQAs such as charge variants in industrial biomanufacturing is not straightforward.56,111 It requires a comprehensive strategy that effectively brings together robust data infrastructures, advanced computational and modeling approaches, and comprehensive data management strategies.124,127 Most importantly, the development of industry standards and clear regulatory frameworks will be important.88
Conclusion: challenges, opportunities, and future directions
MAbs are the dominant modern biopharmaceuticals, yet maintaining consistent product quality, especially in the case of charge heterogeneity, remains a major challenge. We showed that mAb charge heterogeneity primarily arises from a diverse set of PTMs that occur during the complex biomanufacturing process. These modifications are greatly impacted by culture conditions and media components. Traditional bioprocess optimization methods, such as OFAT, DoE, statistical modeling, and routine process controls, have been essential for optimizing cell growth and titer, and for gaining a basic process understanding. These methods, however, have inherent limitations in accurately optimizing the complex nature of the charge variants. This arises from the high dimensionality, non-linear relationships, and complex interactions between factors governing PTMs, along with the need for dynamic process management. ML has emerged as a powerful, data-driven approach that can overcome these traditional limitations by modeling complex biological dynamics and making accurate predictions. Notably, ML techniques, such as deep learning and hybrid modeling approaches integrated with PAT, are showing increasing effectiveness in quantitatively linking upstream process parameters and media components to product CQAs, including glycosylation and, in particular, charge variant distribution. This predictive capability, combined with ML-based optimization and advanced optimization frameworks, provides a promising path to proactively and strategically mitigate charge heterogeneity during upstream bioprocesses. This strategy also supports the integration of QbD principles into bioprocess development.
While ML holds promise for resolving charge heterogeneity in industrial mAb production, several barriers remain to its widespread adoption. A major hurdle lies in ensuring both the quality and accessibility of data. Training effective ML models requires large, diverse, and detailed datasets that capture the complex relationships between upstream conditions, medium composition, and charge variant profiles across batches and scales. Historical manufacturing data, however, are often incomplete, inconsistent, or lacking sufficient detail, and generating comprehensive experimental data from scratch can be time-consuming and expensive. Moreover, preparing this data, by correcting inconsistencies, handling missing entries, and managing variability, requires substantial manual effort.128 Scalability is another concern. Models that perform well in the lab or pilot are not always effective in industrial settings due to scale-dependent factors such as mixing, mass transfer, and operational variability. Furthermore, many ML models, especially deep learning approaches, are often seen as “black boxes,” making it difficult to interpret their predictions or link them to underlying biological mechanisms.129 Perhaps most importantly, the regulatory landscape for ML in Good Manufacturing Practice environments is still evolving. Standard guidelines on model validation, transparency, data integrity, and lifecycle management are needed to ensure trust and compliance. Without them, widespread adoption in the industry remains a challenge.
Despite challenges around data quality, model scalability, and regulatory acceptance, ML offers powerful new avenues for harnessing charge heterogeneity in mAb production. By integrating ML with real-time PAT tools, such as Raman spectroscopy, Fourier transform infrared spectroscopy, or fluorescence sensors, we can continuously feed high-resolution time-series data into predictive models. These models can actively adjust variables such as pH, temperature, or nutrient feeds to lock in target charge profiles. At the same time, automated micro‑bioreactor platforms and high-throughput analytics can generate the rich and diverse datasets that ML requires. Active learning strategies can focus experiments on the most informative conditions, reducing development time and cost compared to traditional factorial studies.130 On the modeling side, specialized algorithms tailored to individual PTMs, such as deamidation or oxidation pathways, can be paired with explainable Artificial Intelligence tools such as SHAP, and complemented by inherently interpretable tree-based algorithms (e.g., decision trees, random forests).29 The combination reveals exactly why and how specific process tweaks affect each modification. This transparency increases process understanding, supports regulatory communications, and enables more targeted cell line and media engineering. Moreover, ML can optimize multiple CQAs simultaneously, ensuring balanced control of charge variants, glycosylation, aggregation, and overall product quality.
Going forward, the computational approaches described here are being increasingly adopted to facilitate the establishment of a “smart factory” vision for bioprocessing. In the context of Bioprocessing 4.0, ML-based digital twins continuously analyze PAT streams and operational data to simulate and optimize every aspect of production, both upstream and downstream, in silico.119 Real-time predictive control will replace end‑point testing, automatically adjusting bioreactor settings to maintain CQAs within precise ranges, thereby reducing batch variability and the failure risk. The insights gained from this process will also lead to more rational designs of cell lines and culture media, embedding robust charge profiles from the very beginning of development. Together, these innovations promise better product stability, lower costs, faster time to market, and broader global access to life‑saving mAbs. Ultimately, this will make high-quality biologics more affordable, scalable, and responsive to emerging health needs.
Funding Statement
This work was supported by the NIHMS under Grant [R35 GM119850 (NEL)].
Disclosure statement
No potential conflict of interest was reported by the author(s).
CRediT authorship contribution statement
Hossein Kavoni: Conceptualization; investigation; writing-original draft; writing-review and editing. Iman Shahidi Pour Savizi: Investigation; writing-review and editing. Saratram Gopalakrishnan: writing-review and editing. Nathan E. Lewis: writing-review and editing. Seyed Abbas Shojaosadati: Conceptualization; project administration; supervision.
References
- 1.Kavoni H, Savizi ISP, Lewis NE, Shojaosadati SA.. Recent advances in culture medium design for enhanced production of monoclonal antibodies in CHO cells: a comparative study of machine learning and systems biology approaches. Biotechnol Adv. 2025;78:108480. doi: 10.1016/j.biotechadv.2024.108480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Castelli MS, McGonigle P, Hornby PJ. The pharmacology and therapeutic applications of monoclonal antibodies. Pharmacol Res Perspect. 2019;7(6). doi: 10.1002/prp2.535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Malhotra N, Chahal A, Jain A, Sharma P, Saini P, Hasan MR, Narang J. Monoclonal antibodies: purification, application in conventional methods and cutting edge technology. Biomed Mater Devices. 2024;3(1):367–20. doi: 10.1007/s44174-024-00203-2. [DOI] [Google Scholar]
- 4.Harris CT, Cohen S. Reducing immunogenicity by design: approaches to minimize immunogenicity of monoclonal antibodies. BioDrugs. 2024;38(2):205–226. doi: 10.1007/s40259-023-00641-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Aboul-Ella H, Gohar A, Ali AA, Ismail LM, Mahmoud AEE-R, Elkhatib WF, Aboul-Ella H. Monoclonal antibodies: from magic bullet to precision weapon. Mol Biomed. 2024;5(1):47. doi: 10.1186/s43556-024-00210-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Santos-Neto JF, Oliveira FO, Hodel KVS, Fonseca LMS, Badaró R, Machado BAS. Technological advancements in monoclonal antibodies. Scientific World. 2021;2021:1–19. doi: 10.1155/2021/6663708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Carrara SC, Ulitzka M, Grzeschik J, Kornmann H, Hock B, Kolmar H. From cell line development to the formulated drug product: the art of manufacturing therapeutic monoclonal antibodies. Int J Pharm. 2021;594:120164. doi: 10.1016/j.ijpharm.2020.120164. [DOI] [PubMed] [Google Scholar]
- 8.Kozlowski S, Swann P. Current and future issues in the manufacturing and development of monoclonal antibodies. Adv Drug Deliv Rev. 2006;58(5–6):707–722. doi: 10.1016/j.addr.2006.05.002. [DOI] [PubMed] [Google Scholar]
- 9.Beck A, Nowak C, Meshulam D, Reynolds K, Chen D, Pacardo DB, Nicholls SB, Carven GJ, Gu Z, Fang J, et al. Risk-based control strategies of recombinant monoclonal antibody charge variants. Antibodies. 2022;11(4):73. doi: 10.3390/antib11040073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ha TK, Kim D, Kim CL, Grav LM, Lee GM. Factors affecting the quality of therapeutic proteins in recombinant Chinese hamster ovary cell culture. Biotechnol Adv. 2022;54:107831. doi: 10.1016/j.biotechadv.2021.107831. [DOI] [PubMed] [Google Scholar]
- 11.Shah A, Cui W, Harrahy J, Ivanov AR. Characterization of charge variants, including post-translational modifications and proteoforms, of bispecific antigen-binding protein by cation-exchange chromatography coupled to native mass spectrometry. Talanta. 2024;266:125062. doi: 10.1016/j.talanta.2023.125062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Boswell CA, Tesar DB, Mukhyala K, Theil F-P, Fielder PJ, Khawli LA. Effects of charge on antibody tissue distribution and pharmacokinetics. Bioconjug Chem. 2010;21(12):2153–2163. doi: 10.1021/bc100261d. [DOI] [PubMed] [Google Scholar]
- 13.Liu H, Nowak C, Shao M, Ponniah G, Neill A. Impact of cell culture on recombinant monoclonal antibody product heterogeneity. Biotechnol Prog. 2016;32(5):1103–1112. doi: 10.1002/btpr.2327. [DOI] [PubMed] [Google Scholar]
- 14.Goyon A, Excoffier M, Janin-Bussat M-C, Bobaly B, Fekete S, Guillarme D, Beck A. Determination of isoelectric points and relative charge variants of 23 therapeutic monoclonal antibodies. J Chromatogr B. 2017;1065–1066:119–128. doi: 10.1016/j.jchromb.2017.09.033. [DOI] [PubMed] [Google Scholar]
- 15.Wagner-Rousset E, Fekete S, Morel-Chevillet L, Colas O, Corvaïa N, Cianférani S, Guillarme D, Beck A. Development of a fast workflow to screen the charge variants of therapeutic antibodies. J Chromatogr A. 2017;1498:147–154. doi: 10.1016/j.chroma.2017.02.065. [DOI] [PubMed] [Google Scholar]
- 16.Torkashvand F, Vaziri B. Main quality attributes of monoclonal antibodies and effect of cell culture components. Iran Biomed J. 2017;21(3):131. doi: 10.18869/acadpub.ibj.21.3.131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Pereira S, Kildegaard HF, Andersen MR. Impact of CHO metabolism on cell growth and protein production: an overview of toxic and inhibiting metabolites and nutrients. Biotechnol J. 2018;13(3):13. doi: 10.1002/biot.201700499. [DOI] [PubMed] [Google Scholar]
- 18.Lee AP, Kok YJ, Lakshmanan M, Leong D, Zheng L, Lim HL, Chen S, Mak SY, Ang KS, Templeton N. Multi‐omics profiling of a CHO cell culture system unravels the effect of culture pH on cell growth, antibody titer, and product quality. Biotechnol Bioeng. 2021;118(11):4305–4316. doi: 10.1002/bit.27899. [DOI] [PubMed] [Google Scholar]
- 19.Zhu Z, Chen X, Li W, Zhuang Y-P, Zhao Y, Wang G. Intracellular CHO cell metabolite profiling and in vivo monitoring of redox state unravel the effect of temperature downshift on cell growth, antibody titer and product quality. 2022; doi: 10.22541/au.165401290.04491126/v1. [DOI] [PubMed]
- 20.Mallaney M, Wang S, Sreedhara A. Effect of ambient light on monoclonal antibody product quality during small‐scale mammalian cell culture process in clear glass bioreactors. Biotechnol Prog. 2014;30(3):562–570. doi: 10.1002/btpr.1920. [DOI] [PubMed] [Google Scholar]
- 21.Weng Z, Jin J, Shao C, Li H. Reduction of charge variants by CHO cell culture process optimization. Cytotechnology. 2020;72(2):259–269. doi: 10.1007/s10616-020-00375-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Gangwar N, Mishra R, Budholiya N, Rathore AS. Effect of vitamins and metal ions on productivity and charge heterogeneity of IgG1 expressed in CHO cells. Biotechnol J. 2021;16(8). doi: 10.1002/biot.202000464. [DOI] [PubMed] [Google Scholar]
- 23.Yuk IH, Zhang B, Yang Y, Dutina G, Leach KD, Vijayasankaran N, Shen AY, Andersen DC, Snedecor BR, Joly JC. Controlling glycation of recombinant antibody in fed‐batch cell cultures. Biotechnol Bioeng. 2011;108(11):2600–2610. doi: 10.1002/bit.23218. [DOI] [PubMed] [Google Scholar]
- 24.Zhang X, Tang H, Sun Y-T, Liu X, Tan W-S, Fan L. Elucidating the effects of arginine and lysine on a monoclonal antibody C-terminal lysine variation in CHO cell cultures. Appl Microbiol Biotechnol. 2015;99(16):6643–6652. doi: 10.1007/s00253-015-6617-y. [DOI] [PubMed] [Google Scholar]
- 25.Chung S, Tian J, Tan Z, Chen J, Zhang N, Huang Y, Vandermark E, Lee J, Borys M, Li ZJ. Modulating cell culture oxidative stress reduces protein glycation and acidic charge variant formation. Abingdon, OX, UK: Taylor & Francis; 2019. p. 205–216. doi: 10.1080/19420862.2018.1537533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kaur R, Jain R, Budholiya N, Rathore AS. Long term culturing of CHO cells: phenotypic drift and quality attributes of the expressed monoclonal antibody. Biotechnol Lett. 2023;45(3):357–370. doi: 10.1007/s10529-023-03346-2. [DOI] [PubMed] [Google Scholar]
- 27.Salim T, Chauhan G, Templeton N, Ling WLW. Using MVDA with stoichiometric balances to optimize amino acid concentrations in chemically defined CHO cell culture medium for improved culture performance. Biotechnol Bioeng. 2022;119(2):452–469. doi: 10.1002/bit.27998. [DOI] [PubMed] [Google Scholar]
- 28.Hashizume T, Ozawa Y, Ying B-W. Employing active learning in the optimization of culture medium for mammalian cells. NPJ Syst Biol Appl. 2023;9(1):20. doi: 10.1038/s41540-023-00284-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Gangwar N, Balraj K, Rathore AS. Explainable AI for CHO cell culture media optimization and prediction of critical quality attribute. Appl Microbiol Biotechnol. 2024;108(1):1–16. doi: 10.1007/s00253-024-13147-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Voit EO, Shah AM, Olivença D, Vodovotz Y. What’s next for computational systems biology? Front Syst Biol. 2023;3:1250228. doi: 10.3389/fsysb.2023.1250228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Brunner M, Fricke J, Kroll P, Herwig C. Investigation of the interactions of critical scale-up parameters (pH, p O 2 and p CO 2) on CHO batch performance and critical quality attributes. Bioprocess Biosyst Eng. 2017;40(2):251–263. doi: 10.1007/s00449-016-1693-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Gangwar N, Priyanka P, Rathore AS. Achieving charge variant profile of innovator molecule during development of monoclonal antibody based biosimilars–use of media components. Biochem Eng J. 2022;182:108438. doi: 10.1016/j.bej.2022.108438. [DOI] [Google Scholar]
- 33.Du Y, Walsh A, Ehrick R, Xu W, May K, Liu H. Chromatographic analysis of the acidic and basic species of recombinant monoclonal antibodies. Abingdon, Oxfordshire, UK: MAbs. Taylor & Francis; 2012. p. 578–585. doi: 10.4161/mabs.21328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Liu H, Ponniah G, Zhang H-M, Nowak C, Neill A, Gonzalez-Lopez N, Patel R, Cheng G, Kita AZ, Andrien B. In vitro and in vivo modifications of recombinant and human IgG antibodies. Mabs-austin. 2014;6(5):1145–1154. doi: 10.4161/mabs.29883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Gupta T, Kumar A, Seshadri S. Bioprocess challenges in purification of therapeutic protein charge variants. Biotechnol Bioprocess Eng. 2023;28(4):493–506. doi: 10.1007/s12257-023-0078-4. [DOI] [Google Scholar]
- 36.Sissolak B, Lingg N, Sommeregger W, Striedner G, Vorauer-Uhl K. Impact of mammalian cell culture conditions on monoclonal antibody charge heterogeneity: an accessory monitoring tool for process development. J Ind Microbiol Biotechnol. 2019;46(8):1167–1178. doi: 10.1007/s10295-019-02202-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Tang H, Miao S, Zhang X, Fan L, Liu X, Tan W-S, Zhao L. Insights into the generation of monoclonal antibody acidic charge variants during Chinese hamster ovary cell cultures. Appl Microbiol Biotechnol. 2018;102(3):1203–1214. doi: 10.1007/s00253-017-8650-5. [DOI] [PubMed] [Google Scholar]
- 38.Brorson K, Jia AY. Therapeutic monoclonal antibodies and consistent ends: terminal heterogeneity, detection, and impact on quality. Curr Opin Biotechnol. 2014;30:140–146. doi: 10.1016/j.copbio.2014.06.012. [DOI] [PubMed] [Google Scholar]
- 39.Chumsae C, Gifford K, Lian W, Liu H, Radziejewski CH, Zhou ZS. Arginine modifications by Methylglyoxal: discovery in a recombinant monoclonal antibody and contribution to acidic species. Anal Chem. 2013;85(23):11401–11409. doi: 10.1021/ac402384y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kaschak T, Boyd D, Lu F, Derfus G, Kluck B, Nogal B, Emery C, Summers C, Zheng K, Bayer R. Characterization of the basic charge variants of a human IgG1: effect of copper concentration in cell culture media. Abingdon, Oxfordshire, UK: MAbs. Taylor & Francis; 2011. p. 577–583. doi: 10.4161/mabs.3.6.17959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Chumsae C, Hossler P, Raharimampionona H, Zhou Y, McDermott S, Racicot C, Radziejewski C, Zhou ZS. When good intentions go awry: modification of a recombinant monoclonal antibody in chemically defined cell culture by Xylosone, an oxidative product of ascorbic acid. Anal Chem. 2015;87(15):7529–7534. doi: 10.1021/acs.analchem.5b00801. [DOI] [PubMed] [Google Scholar]
- 42.Luo J, Zhang J, Ren D, Tsai W, Li F, Amanullah A, Hudson T. Probing of C‐terminal lysine variation in a recombinant monoclonal antibody production using Chinese hamster ovary cells with chemically defined media. Biotechnol Bioeng. 2012;109(9):2306–2315. doi: 10.1002/bit.24510. [DOI] [PubMed] [Google Scholar]
- 43.Yuk IH, Russell S, Tang Y, Hsu W, Mauger JB, Aulakh RPS, Luo J, Gawlitzek M, Joly JC. Effects of copper on CHO cells: cellular requirements and product quality considerations. Biotechnol Prog. 2015;31(1):226–238. doi: 10.1002/btpr.2004. [DOI] [PubMed] [Google Scholar]
- 44.Xie P, Niu H, Chen X, Zhang X, Miao S, Deng X, Liu X, Tan W-S, Zhou Y, Fan L. Elucidating the effects of pH shift on IgG1 monoclonal antibody acidic charge variant levels in Chinese hamster ovary cell cultures. Appl Microbiol Biotechnol. 2016;100(24):10343–10353. doi: 10.1007/s00253-016-7749-4. [DOI] [PubMed] [Google Scholar]
- 45.Horvath B, Mun M, Laird MW. Characterization of a monoclonal antibody cell culture production process using a quality by design approach. Mol Biotechnol. 2010;45(3):203–206. doi: 10.1007/s12033-010-9267-4. [DOI] [PubMed] [Google Scholar]
- 46.Hazeltine LB, Knueven KM, Zhang Y, Lian Z, Olson DJ, Ouyang A. Chemically defined media modifications to lower tryptophan oxidation of biopharmaceuticals. Biotechnol Prog. 2016;32(1):178–188. doi: 10.1002/btpr.2195. [DOI] [PubMed] [Google Scholar]
- 47.Zheng C, Zhuang C, Qin J, Chen Y, Fu Q, Qian H, Wu T, Wang Y, Wu X, Qi N. Combination of temperature shift and hydrolysate addition regulates anti-IgE monoclonal antibody charge heterogeneity in Chinese hamster ovary cell fed-batch culture. Cytotechnology. 2018;70(4):1121–1129. doi: 10.1007/s10616-018-0192-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Chen F, Kou T, Fan L, Zhou Y, Ye Z, Zhao L, Tan W-S. The combined effect of sodium butyrate and low culture temperature on the production, sialylation, and biological activity of an antibody produced in CHO cells. Biotechnol Bioprocess Eng. 2011;16(6):1157–1165. doi: 10.1007/s12257-011-0069-8. [DOI] [Google Scholar]
- 49.Hong JK, Lee SM, Kim K-Y, Lee GM. Effect of sodium butyrate on the assembly, charge variants, and galactosylation of antibody produced in recombinant Chinese hamster ovary cells. Appl Microbiol Biotechnol. 2014;98(12):5417–5425. doi: 10.1007/s00253-014-5596-8. [DOI] [PubMed] [Google Scholar]
- 50.Zhang X, Sun Y-T, Tang H, Fan L, Hu D, Liu J, Liu X, Tan W-S. Culture temperature modulates monoclonal antibody charge variation distribution in Chinese hamster ovary cell cultures. Biotechnol Lett. 2015;37(11):2151–2157. doi: 10.1007/s10529-015-1904-3. [DOI] [PubMed] [Google Scholar]
- 51.Zheng C, Zhuang C, Chen Y, Fu Q, Qian H, Wang Y, Qin J, Wu X, Qi N. Improved process robustness, product quality and biological efficacy of an anti-CD52 monoclonal antibody upon pH shift in Chinese hamster ovary cell perfusion culture. Process Biochem. 2018;65:123–129. doi: 10.1016/j.procbio.2017.11.013. [DOI] [Google Scholar]
- 52.Rouiller Y, Périlleux A, Vesin M, Stettler M, Jordan M, Broly H. Modulation of mAb quality attributes using microliter scale fed‐batch cultures. Biotechnol Prog. 2014;30(3):571–583. doi: 10.1002/btpr.1921. [DOI] [PubMed] [Google Scholar]
- 53.Xu J, Rehmann MS, Xu X, Huang C, Tian J, Qian N-X, Li ZJ. Improving titer while maintaining quality of final formulated drug substance via optimization of CHO cell culture conditions in low-iron chemically defined media. Abingdon, Oxfordshire, UK: MAbs. Taylor & Francis; 2018. p. 488–499. doi: 10.1080/19420862.2018.1433978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Yüce M, Sert F, Torabfam M, Parlar A, Gürel B, Çakır N, Dağlıkoca DE, Khan MA, Çapan Y. Fractionated charge variants of biosimilars: a review of separation methods, structural and functional analysis. Anal Chim Acta. 2021;1152:238189. doi: 10.1016/j.aca.2020.12.064. [DOI] [PubMed] [Google Scholar]
- 55.Hossler P, Wang M, McDermott S, Racicot C, Chemfe K, Zhang Y, Chumsae C, Manuilov A. Cell culture media supplementation of bioflavonoids for the targeted reduction of acidic species charge variants on recombinant therapeutic proteins. Biotechnol Prog. 2015;31(4):1039–1052. doi: 10.1002/btpr.2095. [DOI] [PubMed] [Google Scholar]
- 56.Walsh I, Myint M, Nguyen-Khuong T, Ho YS, Ng SK, Lakshmanan M. Harnessing the potential of machine learning for advancing “quality by design” in biomanufacturin. Abingdon, Oxfordshire, UK: MAbs. Taylor & Francis; 2022. p. 2013593. doi: 10.1080/19420862.2021.2013593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Kishishita S, Nishikawa T, Shinoda Y, Nagashima H, Okamoto H, Takuma S, Aoyagi H. Effect of temperature shift on levels of acidic charge variants in IgG monoclonal antibodies in Chinese hamster ovary cell culture. J Biosci Bioeng. 2015;119(6):700–705. doi: 10.1016/j.jbiosc.2014.10.028. [DOI] [PubMed] [Google Scholar]
- 58.Yang WC, Minkler DF, Kshirsagar R, Ryll T, Huang Y-M. Concentrated fed-batch cell culture increases manufacturing capacity without additional volumetric capacity. J Biotechnol. 2016;217:1–11. doi: 10.1016/j.jbiotec.2015.10.009. [DOI] [PubMed] [Google Scholar]
- 59.McHugh KP, Xu J, Aron KL, Borys MC, Li ZJ. Effective temperature shift strategy development and scale confirmation for simultaneous optimization of protein productivity and quality in Chinese hamster ovary cells. Biotechnol Prog. 2020;36(3):e2959. doi: 10.1002/btpr.2959. [DOI] [PubMed] [Google Scholar]
- 60.Yoon SK, Song JY, Lee GM. Effect of low culture temperature on specific productivity, transcription level, and heterogeneity of erythropoietin in Chinese hamster ovary cells. Biotechnol Bioeng. 2003;82(3):289–298. doi: 10.1002/bit.10566. [DOI] [PubMed] [Google Scholar]
- 61.Xu J, Tang P, Yongky A, Drew B, Borys MC, Liu S, Li ZJ. Systematic development of temperature shift strategies for Chinese hamster ovary cells based on short duration cultures and kinetic modeling. Mabs-austin. 2019;11(1):191–204. doi: 10.1080/19420862.2018.1525262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Trexler‐Schmidt M, Sargis S, Chiu J, Sze‐Khoo S, Mun M, Kao Y, Laird MW. Identification and prevention of antibody disulfide bond reduction during cell culture manufacturing. Biotech Bioeng. 2010;106(3):452–461. doi: 10.1002/bit.22699. [DOI] [PubMed] [Google Scholar]
- 63.Park JH, Jin JH, Lim MS, An HJ, Kim JW, Lee GM. Proteomic analysis of host cell protein dynamics in the culture supernatants of antibody-producing CHO cells. Sci Rep. 2017;7(1):44246. doi: 10.1038/srep44246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Ritacco FV, Wu Y, Khetan A. Cell culture media for recombinant protein expression in Chinese hamster ovary (CHO) cells: history, key components, and optimization strategies. Biotechnol Prog. 2018;34(6):1407–1426. doi: 10.1002/btpr.2706. [DOI] [PubMed] [Google Scholar]
- 65.Stone AT, Dhara VG, Naik HM, Aliyu L, Lai J, Jenkins J, Betenbaugh MJ. Chemical speciation of trace metals in mammalian cell culture media: looking under the hood to boost cellular performance and product quality. Curr Opin Biotechnol. 2021;71:216–224. doi: 10.1016/j.copbio.2021.08.004. [DOI] [PubMed] [Google Scholar]
- 66.Chaderjian WB, Chin ET, Harris RJ, Etcheverry TM. Effect of copper sulfate on performance of a serum‐free CHO cell culture process and the level of free thiol in the recombinant antibody expressed. Biotechnol Prog. 2005;21(2):550–553. doi: 10.1021/bp0497029. [DOI] [PubMed] [Google Scholar]
- 67.Quan C, Alcala E, Petkovska I, Matthews D, Canova-Davis E, Taticek R, Ma S. A study in glycation of a therapeutic recombinant humanized monoclonal antibody: where it is, how it got there, and how it affects charge-based behavior. Anal Biochem. 2008;373(2):179–191. doi: 10.1016/j.ab.2007.09.027. [DOI] [PubMed] [Google Scholar]
- 68.Vijayasankaran N, Varma S, Yang Y, Mun M, Arevalo S, Gawlitzek M, Swartz T, Lim A, Li F, Zhang B. Effect of cell culture medium components on color of formulated monoclonal antibody drug substance. Biotechnol Prog. 2013;29(5):1270–1277. doi: 10.1002/btpr.1772. [DOI] [PubMed] [Google Scholar]
- 69.Hossler P, McDermott S, Racicot C, Fann JCH. Improvement of mammalian cell culture performance through surfactant enabled concentrated feed media. Biotechnol Prog. 2013;29(4):1023–1033. doi: 10.1002/btpr.1739. [DOI] [PubMed] [Google Scholar]
- 70.Xu J, Santos J, Anderson NS, Borys MC, Pendse G, Li ZJ. Antibody charge variant modulation by in vitro enzymatic treatment in different chinese hamster ovary cell cultures. Biotechnol Prog. 2022;38(5). doi: 10.1002/btpr.3268. [DOI] [PubMed] [Google Scholar]
- 71.Kim DG, Kim HJ, Kim H-J. Effects of carboxypeptidase B treatment and elevated temperature on recombinant monoclonal antibody charge variants in cation-exchange chromatography analysis. Arch Pharm Res. 2016;39(10):1472–1481. doi: 10.1007/s12272-016-0818-5. [DOI] [PubMed] [Google Scholar]
- 72.Hyoung Park J, Sin Lim M, Rang Woo J, Won Kim J, Min Lee G. The molecular weight and concentration of dextran sulfate affect cell growth and antibody production in CHO cell cultures. Biotechnol Prog. 2016;32(5):1113–1122. doi: 10.1002/btpr.2287. [DOI] [PubMed] [Google Scholar]
- 73.Liu M, Wang J, Tang H, Fan L, Zhao L, Wang H-B, Zhou Y, Tan W-S. Cell culture medium supplemented with taurine decreases basic charge variant levels of a monoclonal antibody. Biotechnol Lett. 2018;40(11–12):1487–1493. doi: 10.1007/s10529-018-2606-4. [DOI] [PubMed] [Google Scholar]
- 74.Niu H, Wang J, Liu M, Chai M, Zhao L, Liu X, Fan L, Tan W-S. Uridine modulates monoclonal antibody charge heterogeneity in Chinese hamster ovary cell fed-batch cultures. Bioresour Bioprocess. 2018;5(1):1–8. doi: 10.1186/s40643-018-0228-2. [DOI] [Google Scholar]
- 75.Singh V, Haque S, Niwas R, Srivastava A, Pasupuleti M, Tripathi CKM. Strategies for fermentation medium optimization: an in-depth review. Front Microbiol. 2016;7:2087. doi: 10.3389/fmicb.2016.02087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Bhaturiwala R, Bagban M, Mansuri A, Modi H. Successive approach of medium optimization using one-factor-at-a-time and response surface methodology for improved β-mannanase production from Streptomyces sp. Bioresour Technol Rep. 2022;18:101087. doi: 10.1016/j.biteb.2022.101087. [DOI] [Google Scholar]
- 77.Huang Z, Xu J, Yongky A, Morris CS, Polanco AL, Reily M, Borys MC, Li ZJ, Yoon S. CHO cell productivity improvement by genome-scale modeling and pathway analysis: application to feed supplements. Biochem Eng J. 2020;160:107638. doi: 10.1016/j.bej.2020.107638. [DOI] [Google Scholar]
- 78.Zhou T, Reji R, Kairon RS, Chiam KH. A review of algorithmic approaches for cell culture media optimization. Front Bioeng Biotechnol. 2023;11:1195294. doi: 10.3389/fbioe.2023.1195294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Schaub J, Clemens C, Kaufmann H, Schulz TW. Advancing biopharmaceutical process development by system-level data analysis and integration of omics dataSystems biology of mammalian cell culture. In: Hu W, and Zeng A, editors. Advances in Biochemical Engineering/Biotechnology. Vol. 127. Berlin, Heidelberg: Springer; 2012. p. 133–163. doi: 10.1007/10_2010_98. [DOI] [PubMed] [Google Scholar]
- 80.Goldrick S, Sandner V, Cheeks M, Turner R, Farid SS, McCreath G, Glassey J. Multivariate data analysis methodology to solve data challenges related to scale‐up model validation and missing data on a micro‐bioreactor System. Biotechnol J. 2020;15(3):15. doi: 10.1002/biot.201800684. [DOI] [PubMed] [Google Scholar]
- 81.Germain RN, Meier-Schellersheim M, Nita-Lazar A, Fraser IDC. Systems biology in immunology: a computational modeling perspective. Annu Rev Immunol. 2011;29(1):527–585. doi: 10.1146/annurev-immunol-030409-101317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Kraikivski P. Mathematical modeling in systems biology. In: Entropy. Vol. 25. Basel, Switzerland: MDPI; 2023. p. 1380. doi: 10.3390/e25101380 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Hashizume T, Ying B-W. Challenges in developing cell culture media using machine learning. Biotechnol Adv. 2023;70:108293. doi: 10.1016/j.biotechadv.2023.108293. [DOI] [PubMed] [Google Scholar]
- 84.Rathore AS, Nikita S, Thakur G, Mishra S. Artificial intelligence and machine learning applications in biopharmaceutical manufacturing. Trends Biotechnol. 2023;41(4):497–510. doi: 10.1016/j.tibtech.2022.08.007. [DOI] [PubMed] [Google Scholar]
- 85.Khaleghi MK, Savizi ISP, Lewis NE, Shojaosadati SA. Synergisms of machine learning and constraint‐based modeling of metabolism for analysis and optimization of fermentation parameters. Biotechnol J. 2021;16(11):2100212. doi: 10.1002/biot.202100212. [DOI] [PubMed] [Google Scholar]
- 86.Lawson CE, Martí JM, Radivojevic T, Jonnalagadda SVR, Gentz R, Hillson NJ, Peisert S, Kim J, Simmons BA, Petzold CJ. Machine learning for metabolic engineering: a review. Metab Eng. 2021;63:34–60. doi: 10.1016/j.ymben.2020.10.005. [DOI] [PubMed] [Google Scholar]
- 87.Hernández E, Calzadilla L, Toledo A, Gozá O, Pietzke M, Vazquez A, Rodríguez G. Determination of best nutritional conditions for a monoclonal antibody-producing cell line based on a multivariate data analysis approach. Global J Res Eng. 2023;23:21–31. [Google Scholar]
- 88.Mondal PP, Galodha A, Verma VK, Singh V, Show PL, Awasthi MK, Lall B, Anees S, Pollmann K, Jain R. Review on machine learning-based bioprocess optimization, monitoring, and control systems. Bioresour Technol. 2023;370:128523. doi: 10.1016/j.biortech.2022.128523. [DOI] [PubMed] [Google Scholar]
- 89.Daud KM, Ananda R, Zainudin S, Howe CW. Optimizing the production of valuable metabolites using a hybrid of constraint-based model and machine learning algorithms: a review. Int J Advan Comput Sci Appl. 2023;14(10). doi: 10.14569/IJACSA.2023.01410115. [DOI] [Google Scholar]
- 90.Rana P, Berry C, Ghosh P, Fong SS. Recent advances on constraint-based models by integrating machine learning. Curr Opin Biotechnol. 2020;64:85–91. doi: 10.1016/j.copbio.2019.11.007. [DOI] [PubMed] [Google Scholar]
- 91.Ramzi AB, Baharum SN, Bunawan H, Scrutton NS. Streamlining natural products biomanufacturing with omics and machine learning driven microbial engineering. Front Bioeng Biotechnol. 2020;8. doi: 10.3389/fbioe.2020.608918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Antonakoudis A, Barbosa R, Kotidis P, Kontoravdi C. The era of big data: genome-scale modelling meets machine learning. Comput Struct Biotechnol J. 2020;18:3287–3300. doi: 10.1016/j.csbj.2020.10.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Tsopanoglou A, Del Val IJ. Moving towards an era of hybrid modelling: advantages and challenges of coupling mechanistic and data-driven models for upstream pharmaceutical bioprocesses. Curr Opin Chem Eng. 2021;32:100691. doi: 10.1016/j.coche.2021.100691. [DOI] [Google Scholar]
- 94.Dan A, Liu B, Patil U, Manuraj BNM, Gandhi R, Buchel J, Chundawat SPS, Guo W, Ramachandran R. Machine learning model-based design and model predictive control of a bioreactor for the improved production of mammalian cell-based bio-therapeutics. Control Eng Pract. 2025;156:106198. doi: 10.1016/j.conengprac.2024.106198. [DOI] [Google Scholar]
- 95.Baako T-MD, Kulkarni SK, McClendon JL, Harcum SW, Gilmore J. Machine learning and deep learning strategies for Chinese hamster ovary cell bioprocess optimization. Fermentation. 2024;10(5):234. doi: 10.3390/fermentation10050234. [DOI] [Google Scholar]
- 96.Richter J, Wang Q, Lange F, Thiel P, Yilmaz N, Solle D, Zhuang X, Beutel S. Machine learning‐powered optimization of a CHO cell cultivation process. Biotechnol Bioeng. 2025;122(5):1153–1164. doi: 10.1002/bit.28943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Lu T, Meizhizi J, Yang M, Huang D. Deep learning-based prediction of critical parameters in CHO cell culture process and its application in monoclonal antibody production. Int J Multiling Advance In Applied Science Research. 2024; https://h-tsp.com/index.php/ijaasr/article/view/69. [Google Scholar]
- 98.Tanemura H, Kitamura R, Yamada Y, Hoshino M, Kakihara H, Nonaka K. Comprehensive modeling of cell culture profile using Raman spectroscopy and machine learning. Sci Rep. 2023;13(1):21805. doi: 10.1038/s41598-023-49257-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Grzesik P, Warth SC. One-time optimization of advanced T cell culture media using a machine learning pipeline. Front Bioeng Biotechnol. 2021;9:614324. doi: 10.3389/fbioe.2021.614324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Hong JK, Choi D-H, Park S-Y, Silberberg YR, Shozui F, Nakamura E, Kayahara T, Lee D-Y. Data-driven and model-guided systematic framework for media development in CHO cell culture. Metab Eng. 2022;73:114–123. doi: 10.1016/j.ymben.2022.07.003. [DOI] [PubMed] [Google Scholar]
- 101.Schinn S, Morrison C, Wei W, Zhang L, Lewis NE. A genome‐scale metabolic network model and machine learning predict amino acid concentrations in Chinese hamster ovary cell cultures. Biotechnol Bioeng. 2021;118(5):2118–2123. doi: 10.1002/bit.27714. [DOI] [PubMed] [Google Scholar]
- 102.Tachibana S, Chiou T, Konishi M. Machine learning modeling of the effects of media formulated with various yeast extracts on heterologous protein production in Escherichia coli. Microbiologyopen. 2021;10(3). doi: 10.1002/mbo3.1214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Okamura K, Badr S, Murakami S, Sugiyama H. Hybrid modeling of CHO cell cultivation in monoclonal antibody production with an impurity generation module. Ind Eng Chem Res. 2022;61(40):14898–14909. doi: 10.1021/acs.iecr.2c00736. [DOI] [Google Scholar]
- 104.Alam MN, Anurag A, Gangwar N, Ramteke M, Kodamana H, Rathore AS. Physics‐informed neural networks guided modelling and multiobjective optimization of a scpmAbscpproduction process. Can J Chem Eng. 2025;103(3):1319–1334. doi: 10.1002/cjce.25446. [DOI] [Google Scholar]
- 105.Lakshmanan M, Chia S, Pang KT, Sim LC, Teo G, Mak SY, Chen S, Lim HL, Lee AP, Bin Mahfut F, et al. Antibody glycan quality predicted from CHO cell culture media markers and machine learning. Comput Struct Biotechnol J. 2024;23:2497–2506. doi: 10.1016/j.csbj.2024.05.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Goldrick S, Umprecht A, Tang A, Zakrzewski R, Cheeks M, Turner R, Charles A, Les K, Hulley M, Spencer C, et al. High-throughput Raman spectroscopy combined with innovate data analysis workflow to enhance biopharmaceutical process development. Processes. 2020;8(9):1179. doi: 10.3390/pr8091179. [DOI] [Google Scholar]
- 107.Khuat TT, Bassett R, Otte E, Grevis-James A, Gabrys B. Applications of machine learning in biopharmaceutical process development and manufacturing: current trends, challenges, and opportunities. Comput Chem Eng. 2023;182:108585. [Google Scholar]
- 108.Sarker IH. Machine learning: algorithms, real-world applications and Research directions. SN Comput Sci. 2021;2(3):160. doi: 10.1007/s42979-021-00592-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Narayanan H, Dingfelder F, Butté A, Lorenzen N, Sokolov M, Arosio P. Machine learning for biologics: opportunities for protein engineering, developability, and formulation. Trends Pharmacol Sci. 2021;42(3):151–165. doi: 10.1016/j.tips.2020.12.004. [DOI] [PubMed] [Google Scholar]
- 110.Park Y. Machine learning-based approaches to integrate heterogeneous data for biological knowledge transfer [Doctoral dissertation]. Göttingen, Germany: Georg-August-Universität Göttingen; 2024. doi: 10.53846/goediss-10534. [DOI] [Google Scholar]
- 111.Kumar J, Sultana R, Saripalla D, Chopda V, Adhikary L. Optimizing mAb biosimilar production via transfer and active learning for fine-tuning physical process parameters for achieving targeted N-glycan and charge variant profiles: advancing bioprocess 4.0. 2025; doi: 10.22541/au.174083048.87047979/v1. [DOI]
- 112.Gyorgypal A, Chaturvedi A, Chopda V, Zhang H, Chundawat SPS. Evaluating the impact of media and feed combinations on CHO cell culture performance and monoclonal antibody (trastuzumab) production. Cytotechnology. 2025;77(1):40. doi: 10.1007/s10616-024-00690-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Mu’azzam K, Santos da Silva FV, Murtagh J, Sousa Gallagher MJ. A roadmap for model-based bioprocess development. Biotechnol Adv. 2024;73:108378. doi: 10.1016/j.biotechadv.2024.108378. [DOI] [PubMed] [Google Scholar]
- 114.Mou T, Pillai HS, Wang S, Wan M, Han X, Schweitzer NM, Che F, Xin H. Bridging the complexity gap in computational heterogeneous catalysis with machine learning. Nat Catal. 2023;6(2):122–136. doi: 10.1038/s41929-023-00911-w. [DOI] [Google Scholar]
- 115.Salehpour MJ, Hossain MJ. Leveraging machine learning for efficient EV integration as mobile battery energy storage systems: exploring strategic frameworks and incentives. J Energy Storage. 2024;92:112151. doi: 10.1016/j.est.2024.112151. [DOI] [Google Scholar]
- 116.Yaghoubi E, Yaghoubi E, Khamees A, Razmi D, Lu T. A systematic review and meta-analysis of machine learning, deep learning, and ensemble learning approaches in predicting EV charging behavior. Eng Appl Artif Intell. 2024;135:108789. doi: 10.1016/j.engappai.2024.108789. [DOI] [Google Scholar]
- 117.Sokolov M, Ritscher J, MacKinnon N, Souquet J, Broly H, Morbidelli M, Butté A. Enhanced process understanding and multivariate prediction of the relationship between cell culture process and monoclonal antibody quality. Biotechnol Prog. 2017;33(5):1368–1380. doi: 10.1002/btpr.2502. [DOI] [PubMed] [Google Scholar]
- 118.Sommeregger W, Sissolak B, Kandra K, von Stosch M, Mayer M, Striedner G. Quality by control: towards model predictive control of mammalian cell culture bioprocesses. Biotechnol J. 2017;12(7):12. doi: 10.1002/biot.201600546. [DOI] [PubMed] [Google Scholar]
- 119.Isoko K, Cordiner JL, Kis Z, Moghadam PZ. Bioprocessing 4.0: a pragmatic review and future perspectives. Digit Discov. 2024;3(9):1662–1681. doi: 10.1039/D4DD00127C. [DOI] [Google Scholar]
- 120.Ündey C, Ertunç S, Mistretta T, Looze B. Applied advanced process analytics in biopharmaceutical manufacturing: challenges and prospects in real-time monitoring and control. J Process Control. 2010;20(9):1009–1018. doi: 10.1016/j.jprocont.2010.05.008. [DOI] [Google Scholar]
- 121.Jakes C, Millán-Martín S, Carillo S, Scheffler K, Zaborowska I, Bones J. Tracking the behavior of monoclonal antibody product quality attributes using a multi-attribute method workflow. J Am Soc Mass Spectrom. 2021;32(8):1998–2012. doi: 10.1021/jasms.0c00432. [DOI] [PubMed] [Google Scholar]
- 122.Ott V, Ott J, Eibl D, Eibl R. Scaling fed-batch and perfusion antibody production processes in geometrically dissimilar stirred bioreactors. Processes. 2024;12(4):806. doi: 10.3390/pr12040806. [DOI] [Google Scholar]
- 123.Bielser J-M, Wolf M, Souquet J, Broly H, Morbidelli M. Perfusion mammalian cell culture for recombinant protein manufacturing – a critical review. Biotechnol Adv. 2018;36(4):1328–1340. doi: 10.1016/j.biotechadv.2018.04.011. [DOI] [PubMed] [Google Scholar]
- 124.Badr S, Sugiyama H. A PSE perspective for the efficient production of monoclonal antibodies: integration of process, cell, and product design aspects. Curr Opin Chem Eng. 2020;27:121–128. doi: 10.1016/j.coche.2020.01.003. [DOI] [Google Scholar]
- 125.Ahmed SF, Alam MSB, Hassan M, Rozbu MR, Ishtiak T, Rafa N, Mofijur M, Shawkat Ali ABM, Gandomi AH. Deep learning modelling techniques: current progress, applications, advantages, and challenges. Artif Intell Rev. 2023;56(11):13521–13617. doi: 10.1007/s10462-023-10466-8. [DOI] [Google Scholar]
- 126.Qiu J. An analysis of model evaluation with cross-validation: techniques, applications, and recent advances. Adv Econ Manag Political Sci. 2024;99(1):69–72. doi: 10.54254/2754-1169/99/2024OX0213. [DOI] [Google Scholar]
- 127.Aldoseri A, Al-Khalifa KN, Hamouda AM. Re-thinking data strategy and integration for artificial intelligence: concepts, opportunities, and challenges. Appl Sci. 2023;13(12):7082. doi: 10.3390/app13127082. [DOI] [Google Scholar]
- 128.Khuat TT, Bassett R, Otte E, Grevis-James A, Gabrys B. Applications of machine learning in antibody discovery, process development, manufacturing and formulation: current trends, challenges, and opportunities. Comput Chem Eng. 2024;182:108585. doi: 10.1016/j.compchemeng.2024.108585. [DOI] [Google Scholar]
- 129.Hassija V, Chamola V, Mahapatra A, Singal A, Goel D, Huang K, Scardapane S, Spinelli I, Mahmud M, Hussain A. Interpreting black-box models: a review on explainable artificial intelligence. Cognit Comput. 2024;16(1):45–74. doi: 10.1007/s12559-023-10179-8. [DOI] [Google Scholar]
- 130.Zhang L, Wen J. Active learning strategy for high fidelity short-term data-driven building energy forecasting. Energy Build. 2021;244:111026. doi: 10.1016/j.enbuild.2021.111026. [DOI] [Google Scholar]


