Abstract
Background:
Ulcerative colitis (UC), a chronic immune-mediated colon inflammation, impacts patients’ quality of life. Immunosuppressive-treated UC patients are prone to opportunistic infections like cytomegalovirus (CMV) infection, which exacerbates UC, causes steroid resistance, and elevates surgery and mortality risks. Identifying CMV colitis from UC exacerbation is difficult due to overlapping symptoms and low biopsy detection rates.
Objectives:
To develop an artificial intelligence (AI)-based multimodal model for early identification of UC with concomitant CMV colitis.
Design:
This was a retrospective diagnostic study.
Methods:
A total of 174 moderate to severe UC patients (87 with CMV colitis) from 2015 to 2023 in Peking Union Medical College Hospital were enrolled retrospectively. A total of 3345 colonoscopy images were collected. The dataset was split into training (70%) and testing (30%) sets. A multimodal dynamic affine transformation (DAFT) model integrating clinical biomarkers and endoscopic images was constructed, along with ResNet and SeNet models. Model performance was evaluated using accuracy, sensitivity, specificity, positive and negative predictive values from the confusion matrix.
Results:
UC patients with CMV colitis had distinct clinical characteristics. The multimodal DAFT model outperformed ResNet and SeNet in distinguishing UC with CMV colitis, with higher accuracy (0.91), sensitivity (0.87), and specificity (0.93).
Conclusion:
AI application offers a promising way to enhance early identification of UC with CMV colitis. The multimodal model combining clinical and endoscopic data can assist clinicians in accurate and timely diagnosis.
Keywords: artificial intelligence, cytomegalovirus colitis, deep learning, multimodal model, ulcerative colitis
Plain language summary
Artificial intelligence based multimodal model for the identification of ulcerative colitis with concomitant cytomegalovirus colitis
Background: Ulcerative colitis (UC), a chronic immune-mediated colon inflammation, impacts patients’ quality of life. Immunosuppressive-treated UC patients are prone to opportunistic infections like cytomegalovirus (CMV) infection, which exacerbates UC, causes steroid resistance, and elevates surgery and mortality risks. Identifying CMV colitis from UC exacerbation is difficult due to overlapping symptoms and low biopsy detection rates.
Aim: To develop an AI-based multimodal model for early identification of UC with concomitant CMV colitis.
Methods: 174 moderate to severe UC patients (87 with CMV colitis) from 2015 - 2023 in Peking Union Medical College Hospital were enrolled retrospectively. 3345 colonoscopy images were collected. The dataset was split into training (70%) and testing (30%) sets. A multimodal DAFT model integrating clinical biomarkers and endoscopic images was constructed, along with Resnet and Senet models. Model performance was evaluated using accuracy, sensitivity, specificity, positive and negative predictive values from the confusion matrix.
Results: UC patients with CMV colitis had distinct clinical characteristics. The multimodal DAFT model outperformed Resnet and Senet in distinguishing UC with CMV colitis, with higher accuracy (0.91), sensitivity (0.87), and specificity (0.93).
Conclusion: AI application offers a promising way to enhance early identification of UC with CMV colitis. The multimodal model combining clinical and endoscopic data can assist clinicians in accurate and timely diagnosis.
Background
Ulcerative colitis (UC) is a chronic immune-mediated disease characterized by inflammation of the colon that can have a prolonged and unremitting course, severely impacting patients’ quality of life. Data from the National Urban Employee Basic Medical Insurance Program in China showed that the incidence of UC increased from 8.1/100,000 in 2013 to 16.6/100,000 in 2016. 1 The rapid increase in the number of UC cases highlights the importance of prevention and active treatment. Patients with UC who receive treatment with steroids, immunosuppressants, and biologics have a 3–5 times higher risk of opportunistic infections. 2 Of these infections, cytomegalovirus (CMV) infection in UC patients requires particular attention. Research has shown that CMV infection can induce disease activity in UC patients, cause steroid resistance, and increase surgery and mortality rates in patients with CMV colitis and UC. 3 Early recognition and treatment of CMV colitis are necessary to improve steroid resistance and reduce the risk of colectomy in UC patients.4,5 However, acute UC flare and CMV colitis share very similar clinical presentations, including hematochezia, diarrhea, and fever, which increases the difficulty of identification. Currently, the diagnosis of CMV colitis relies on pathological examination, including searching for viral inclusions in colonic mucosal tissue using HE staining, immunohistochemical (IHC) staining, and CMV-DNA in situ hybridization. 6 HE staining has a high specificity, ranging from 92% to 100%, but its sensitivity is in a wide fluctuation from 10% to 87%, 7 and the diagnosis is observer-dependent. IHC staining increases the diagnostic sensitivity (78%–93%) and specificity (92%–100%) 8 but to some extent, it is time-consuming and also depends on the location and number of the biopsy. 9 Some characteristic endoscopic features like punched-out ulcers, irregular ulceration, and cobblestone-like appearance may imply CMV colitis. 10 A study investigating physicians’ opinions on the endoscopic features of CMV colitis with UC patients revealed that punched-out ulcerations were the most agreed-upon feature that indicated CMV colitis. 11 Targeting these endoscopic features of CMV colitis when biopsy is benefit to increase the sensitivity of pathological examination. Identifying these features under endoscopy requires repeated training and enriched endoscopic experience, and inexperienced endoscopists may overlook these features. Therefore, a diagnostic tool is needed to provide clues for tissue biopsy that help better identify CMV colitis earlier.
In recent years, artificial intelligence (AI) technology has been widely utilized in the field of gastroenterology, greatly improving the clinical diagnosis efficiency of gastrointestinal diseases. In the field of endoscopy, AI has been used to identify early-stage tumors and depth of tumor invasion in the upper gastrointestinal tract, identify and classify colon polyps, and predict Helicobacter pylori infection. 12 AI also has enormous potential in the field of inflammatory bowel disease (IBD). Cai et al. applied traditional machine learning algorithms, including random forests (RFs), support vector machines (SVMs), and eXtreme gradient boosting (XGBoost), to train to predict disease activity in IBD patients. 13 Compared to traditional machine learning, deep learning exhibits a powerful ability to learn the essential features of a dataset from a small sample set, with a higher accuracy for classification or prediction. Guimaraes et al. applied convolutional neural networks (CNNs), one of the representative algorithms of deep learning, based on endoscopic images and clinical data to differentiate inflammatory bowel disease, infectious colitis, and ischemic colitis. 14 However, there is little research on deep learning models for differentiation of UC and CMV colitis.
The purpose of this study is to establish a multimodal model trained with clinical biomarkers and endoscopic images, based on a deep learning algorithm for early recognition and diagnosis of CMV colitis.
Method
Dataset description
This study consecutively included 87 patients with moderate to severe active UC and 87 patients with moderate to severe UC combined with CMV colitis who underwent colonoscopy at Peking Union Medical College Hospital from July 2015 to June 2023. UC diagnosis was made based on the “Chinese clinical practice guideline on the management of UC (2023, Xi’an) 15 ” and “European Crohn’s and Colitis Organization (ECCO) Consensus on Diagnosis and Management of Ulcerative Colitis.” 16 Moderate to severe activity was defined as a modified Mayo score of ⩾6. 17 CMV colitis diagnosis was based on pathological examination: finding CMV inclusion bodies in colonic mucosal HE staining, positive CMV Immunohistochemical staining, or positive CMV-DNA in situ hybridization. A total of 87 patients with matched age and modified Mayo score who had moderate to severe UC without CMV colitis were also collected as a control group. Colonoscopy images and the following laboratory tests were collected for each patient: white blood cell count, neutrophil count, hemoglobin, eosinophil count, lymphocyte count, platelet count, erythrocyte sedimentation rate (ESR), and immunoglobulin A (IgA).
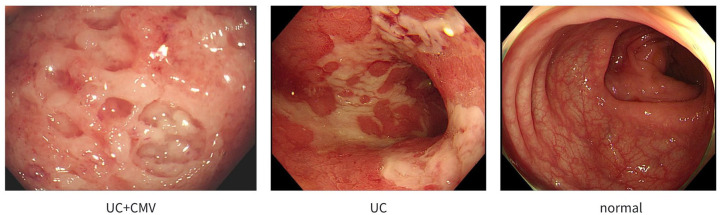
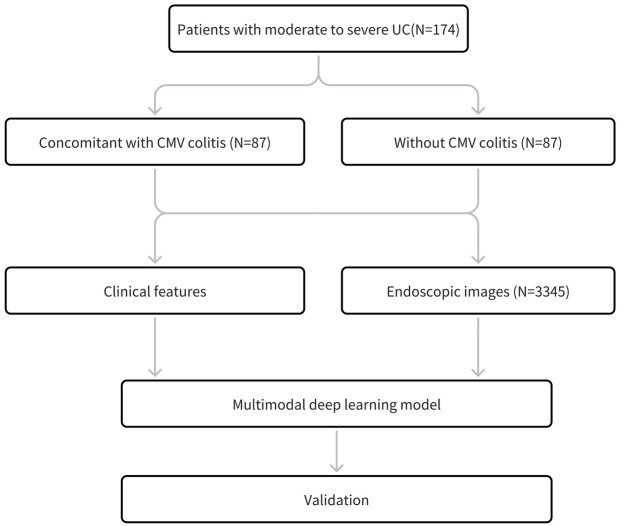
A total of 3345 colonoscopy images were included and classified into three categories by two experienced endoscopists: CMV colitis (CMV, n = 1115), UC without CMV colitis (UC, n = 1115), and normal (N, n = 1115). When developing and evaluating the model, all the images were randomly divided into two subsets: training set (70% of the sample size) and testing set (30% of the sample size). The definitions of the endoscopic classification are detailed in Figure 1. The flow chart of the study is shown in Figure 2.
Figure 1.
The endoscopic classification of each category.
UC + CMV, UC concomitant with CMV colitis.
CMV, cytomegalovirus; UC, ulcerative colitis.
Figure 2.
Study flow diagram.
Data extraction
Data extraction was performed by removing interfering information such as patient age, gender, and examination time from colonoscopy images. The specific steps involved findcontours function in OpenCV 18 to identify the contour of the colon, and obtain the coordinates of the rectangular box using the x, y coordinates of the point set. The region corresponding to the rectangular box with four vertices (x_min, y_min), (x_min, y_max), (x_max, y_min), and (x_max, y_max) was taken as the image used for diagnosis.
Model architecture
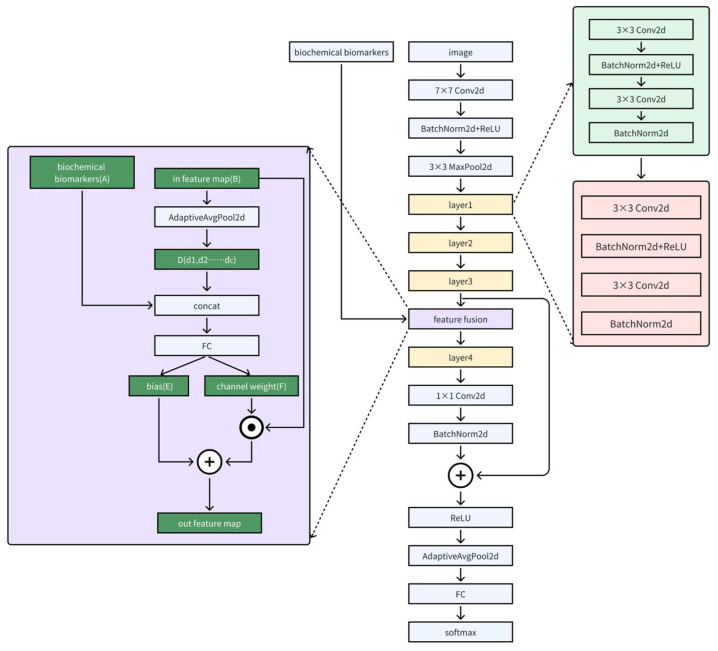
The model architecture consisted of a main network utilizing the structure of residual networks (ResNet): a 7 × 7 convolutional layer, batch normalization, ReLU activation function, and max pooling layer were used to extract shallow features of the image, followed by four layers to extract deep features. Each layer consisted of two Basic Blocks, of which the structure is shown on the right side of Figure 3. The feature maps from layer4 and layer3 were merged after being downsampled by residual structure, then global pooling, a fully connected layer, and a softmax function were applied to obtain the final classification results.
Figure 3.
Deep learning architecture.
The feature fusion module is shown in the purple area on the left side of Figure 2. First, AdaptiveAvgPool2d was used to produce the channel-level feature vector D(d1,d2. . .dc) of the feature map B generated by layer3, where c was the number of channels. Then, the feature vector A corresponding to the medical index was concatenated with D and fused by a fully connected layer to obtain feature vectors E(e1,e2. . .ec) and F(f1,f2. . .fc), where E was the bias vector and F was the channel weight vector. Finally, the feature map G = B × F + E was used as the final fused feature.
The dynamic affine transformation (DAFT) module (G = B × F + E) provides the foundation for model decision interpretability: clinical biomarkers generate scaling factors F and offsets E through fully connected layers, directly modulating the channel-wise expression of image feature maps B. This mechanism enables key clinical features, such as low eosinophil count, to quantitatively influence the model’s interpretation of imaging manifestations, allowing clinicians to trace how biomarkers assist in endoscopic diagnosis.
To validate the independent contribution of clinical biomarkers to classification, this study employed a progressive ablation strategy to train three models. ResNet served as the baseline, processing only endoscopic images without attention mechanisms or clinical data to evaluate the performance of pure imaging features. The Squeeze-and-Excitation Networks (SeNet) model embedded an Squeeze-and-Excitation (SE) channel attention module into the ResNet architecture to enhance perception of critical imaging regions while excluding clinical biomarkers to test the effect of attention-based feature optimization. Building upon SeNet, DAFTNet integrated eight clinical biomarkers—white blood cell, neutrophil, eosinophil, lymphocyte, and platelet counts; hemoglobin, ESR, and IgA—with attention-optimized imaging features through the DAFT module, aiming to quantify the incremental value of clinical biomarkers in multimodal diagnosis.
Model training
Training was performed using a GeForce 3060 graphics card with 12 GB of memory. The cross-entropy cost function was chosen as the loss function to supervise the optimization direction of the model. Adam optimizer was used with a fixed learning rate of 1e-4, and gradient clipping was applied to prevent shaking caused by excessive gradient during training. A total of 2340 samples were collected as training data and trained for 200 epochs with a batch size of 10. To avoid overfitting, random horizontal and vertical flips and rotations were applied to increase the diversity of the dataset during training, and the best-performing model parameters were selected as the final result after convergence of the network.
Outcome definition
In this work, we investigated three major outcomes, namely (1) distinguishing healthy samples from pathological samples (healthy), (2) distinguishing UC from healthy and CMV colitis (UC), and (3) distinguishing CMV colitis from UC and healthy samples (CMV).
Performance evaluation metrics
The metrics used for assessing the model performance in test stages are accuracy, sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV), which are computed from the values of the confusion matrix, true positives (TP), true negatives (TN), false positives (FP), and false negatives (FN). The reporting of this study conforms to the Standards for Reporting Diagnostic Accuracy (STARD) statement (Supplemental Material). 19
Results
Table 1 presents the clinical characteristics of UC patients and UC patients with concomitant CMV colitis. Compared with UC patients without CMV colitis, those with CMV colitis were characterized by elderly age, more severe anemia, lower eosinophil, and platelet count.
Table 1.
Clinical characteristics.
| Characteristics | UC (n = 87) | UC + CMV (n = 87) | p Value |
|---|---|---|---|
| Age (years) | 39.2 ± 14.0 | 48.6 ± 14.0 | 0.000 |
| Gender (male/female, %) | 49/38 (56%/44%) | 56/31 (64%/36%) | 0.278 |
| White blood cell count (×109/L) | 7.8 ± 2.8 | 7.9 ± 3.9 | 0.958 |
| Neutrophil count (×109/L) | 5.3 ± 2.4 | 5.4 ± 3.2 | 0.727 |
| Hemoglobin (g/L) | 109.3 ± 24.4 | 101.2 ± 21.5 | 0.023 |
| Eosinophil count (×109/L) | 0.2 ± 0.2 | 0.1 ± 0.2 | 0.001 |
| Lymphocyte count (×109/L) | 1.8 ± 0.7 | 1.8 ± 1.0 | 0.851 |
| Platelet count (×109/L) | 364.6 ± 127.5 | 317.9 ± 137.2 | 0.022 |
| Erythrocyte sedimentation rate (mm/h) | 35.45 ± 28.2 | 36.35 ± 26.7 | 0.834 |
| Immunoglobulin A (g/L) | 2.3 ± 0.9 | 2.8 ± 5.5 | 0.523 |
UC + CMV: UC with concomitant CMV colitis.
CMV, cytomegalovirus; UC, ulcerative colitis.
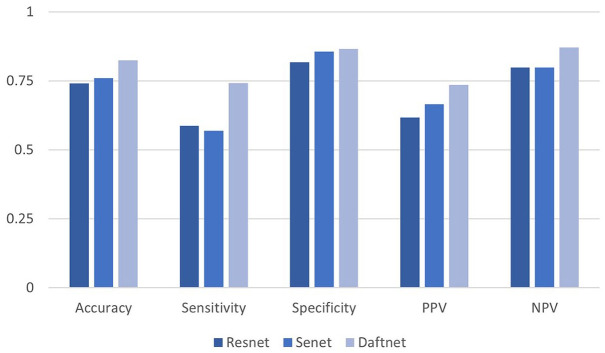
Table 2 reports the overall results in terms of accuracy, sensitivity, specificity, PPV, and NPV of the three models. Figures 4–6 show the performance of the model for all classification tasks. The SeNet model added an attention module SE Block on the basis of the classic ResNet model, and as Table 2 shows, the performance of the model for three tasks was generally improved. Compared to the ResNet and SeNet models, the DAFT Net 20 led to superior classification performance.
Table 2.
Metrics values for the trained models.
| Task | Data type | Model | Accuracy | Sensitivity | Specificity | PPV | NPV |
|---|---|---|---|---|---|---|---|
| healthy | Image-only | ResNet | 0.837 | 0.803 | 0.854 | 0.733 | 0.897 |
| SeNet | 0.851 | 0.818 | 0.867 | 0.755 | 0.905 | ||
| Image + table | DAFTNet | 0.865 | 0.788 | 0.903 | 0.802 | 0.895 | |
| UC | Image-only | ResNet | 0.741 | 0.588 | 0.818 | 0.618 | 0.799 |
| SeNet | 0.761 | 0.57 | 0.857 | 0.666 | 0.799 | ||
| Image + table | DAFTNet | 0.825 | 0.743 | 0.866 | 0.735 | 0.871 | |
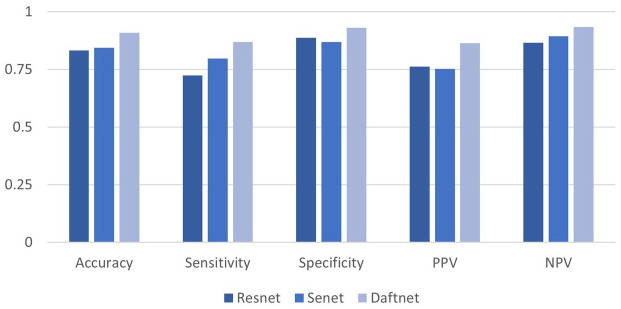
| UC + CMV | Image-only | ResNet | 0.833 | 0.725 | 0.887 | 0.762 | 0.866 |
| SeNet | 0.845 | 0.797 | 0.869 | 0.752 | 0.895 | ||
| Image + table | DAFTNet | 0.91 | 0.869 | 0.931 | 0.864 | 0.934 |
CMV, cytomegalovirus; NPV, negative predictive value; PPV, positive predictive value; UC, ulcerative colitis.
Figure 4.
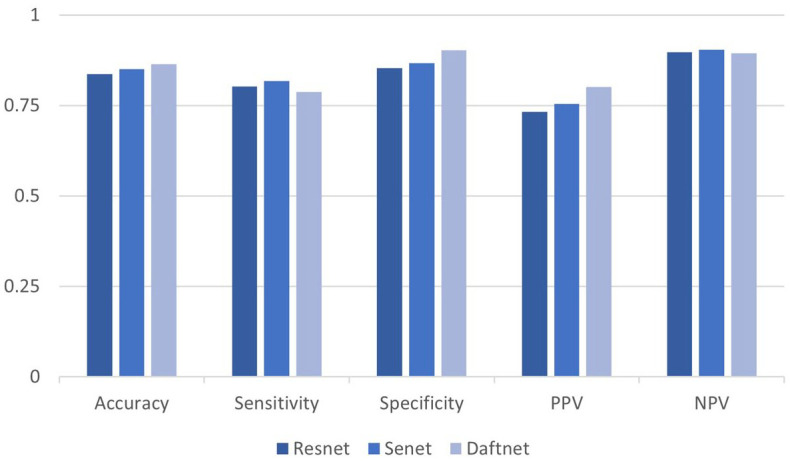
Healthy pathological classification results: distinguish healthy samples from pathological samples.
Figure 5.
UC classification results: distinguish UC from healthy and CMV colitis.
CMV, cytomegalovirus; UC, ulcerative colitis.
Figure 6.
UC + CMV classification results: distinguish CMV colitis from healthy and UC.
CMV, cytomegalovirus; UC, ulcerative colitis.
Ablation experiments demonstrated that clinical biomarkers were core determinants of model performance improvement. In the task of distinguishing UC from CMV colitis, DAFTNet achieved a 6.5% increase in accuracy and 7.1% increase in specificity compared to SeNet. This improvement confirms that biomarkers such as low platelet count provide independent diagnostic signals unrevealed by endoscopic images alone, playing a critical role in reducing false-positive rates.
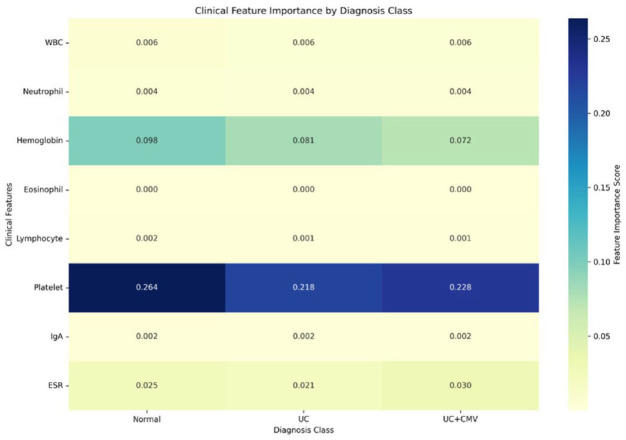
To further interpret how DAFTNet utilizes clinical biomarkers, we quantified the importance of each feature in the dynamic affine transformation. As shown in Figure 7, platelet count consistently demonstrated the highest importance across all diagnostic classes (Normal: 0.264, UC: 0.218, UC + CMV: 0.228), with statistically significant differences between Normal and inflammatory groups (p < 1e-10). Hemoglobin showed higher importance in Normal diagnosis (0.098) compared to UC (0.081) and UC + CMV (0.072) (p < 1e-8), suggesting its role in ruling out anemia-related mucosal changes. ESR importance was significantly elevated in UC + CMV (0.030) versus other classes (p < 0.01), reflecting systemic inflammation in CMV infection. The reduced importance of eosinophil count in UC + CMV (p < 0.05) aligns with the known phenomenon of eosinopenia during viral infections.
Figure 7.
Clinical feature importance by diagnosis class.
Specifically, we can see from Table 2 and Figure 6, the DAFTNet model exhibited better performance in distinguishing UC combined with CMV colitis from UC and healthy samples, with a high accuracy of 0.91. The overall sensitivity of the model was relatively low; however, the sensitivity to identify CMV colitis reaches as high as 0.869.
The ROC analysis showed that the area under the curve of the three classification tasks was 0.90, 0.86, and 0.95, respectively (Figure 8), which implied a considerably excellent performance of our model in terms of predicting UC and CMV colitis.
Figure 8.
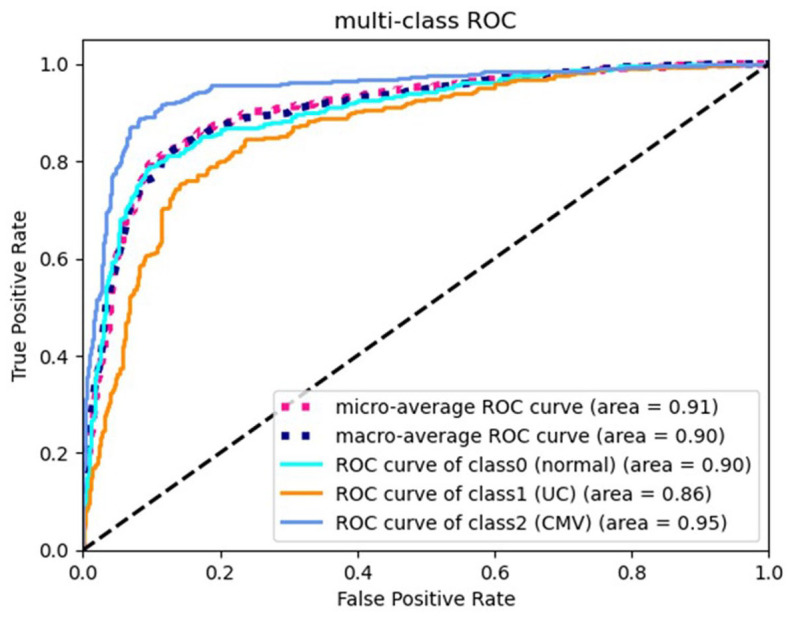
Performance curves.
Receiving operating characteristics (ROC) for the testing set.
Discussion
In clinical practice, the occurrence of concomitant CMV colitis in patients with UC is relatively common, which has an impact on the treatment and prognosis of UC. However, due to the similar clinical manifestations between the two, it is often challenging for clinicians to differentiate CMV colitis from UC flares. In this study, we have successfully developed three deep learning-based diagnostic models (ResNet, SeNet, and DAFTNet) for the identification of UC and CMV colitis. Among them, the DAFTNet model demonstrates better performance in distinguishing UC combined with CMV colitis from UC and healthy samples, with an accuracy of up to 0.91, and the sensitivity for identifying CMV colitis reaches 0.869. In addition, through comparison, our multimodal model (DAFT) outperforms the image-based ResNet and SeNet unimodal models with attention modules in all tasks and metrics, indicating that the multimodal model that combines clinical biomarkers and endoscopic image information can more effectively assist clinicians in making accurate diagnoses.
The application of AI in various tasks is becoming increasingly widespread in IBD. 21 Kraszewski et al. established a machine learning prediction model based on blood, urine, and fecal markers with 97% and 91% mean average precision for CD and UC, respectively. 22 Gui et al. developed a CNN-based model to detect neutrophils in biopsy slide images of UC patients and classify images into either a histologic remission or non-remission. 23 Chen et al. utilized multi-omics data to develop a diagnostic model for CMV reactivation in UC, identifying eight immune-related biomarkers via machine learning. 24 In this study, we have successfully developed three innovative deep learning diagnostic models focusing on real-time endoscopic guidance to identify UC and CMV colitis.
In our deep learning model, the recognition of images is based on the endoscopic features of CMV colitis. Yang et al. discovered that characteristic endoscopic features such as punch-out ulcerations, irregular ulcerations, and a cobblestone-like appearance may help predict the presence of CMV colitis. 10 Furthermore, we found that UC patients with CMV infection had a tendency to exhibit elderly age, more severe anemia, eosinopenia, and thrombocytosis. Previous studies showed that decreased peripheral blood absolute lymphocyte count, eosinophil count, IgA level, and higher platelet to lymphocyte ratio may serve as a predictive marker for CMV infection,25–27 which is consistent with our study. Therefore, clinical biomarkers may help predict CMV infection.
Our image-based ResNet and SeNet models performed similarly with previously established image-based CNNs in identifying UC 14 but they exhibited better performance in identifying CMV colitis. We acknowledge Kim et al.’s 28 valuable contribution to CMV detection using deep learning, where their ensemble approach achieved a sensitivity of 0.91 and an accuracy of 0.83. However, it is important to note that their best-performing single model achieved an accuracy of 0.792, comparable to our unimodal imaging models (ResNet: 0.833, SeNet: 0.845). This comparison underscores that ensemble techniques, while effective for performance gains, were essential for Kim et al.’s approach to outperform standalone models. By contrast, our multimodal model, DAFTNet, achieves competitive accuracy (0.91), specificity (0.931), and balanced sensitivity (0.869) by integrating multimodal data: endoscopic features (3345 images from 174 patients) and clinical biomarkers. This innovation addresses critical limitations of Kim et al.’s study, which relied solely on endoscopic features with a small sample size (n = 34, 86 images). Our DAFT module dynamically fuses these complementary data sources, enabling the model to leverage clinical biomarkers to refine image-based predictions. The 6.5% accuracy improvement of DAFTNet over SeNet confirms that our multimodal integration strategy—rather than model ensembling—drives performance gains, providing a more scalable and interpretable solution for clinical translation. We acknowledge that low sensitivity and PPV for the UC and CMV colitis groups are the main limitations of these two models. The multimodal model (DAFT) addressed this deficiency. Multimodal involves the collaborative analysis of various types of data, such as images, text, etc., to help AI more accurately complete classification and recognition tasks. The DAFT model presents several innovative aspects. 29 First, it enables a two-way exchange of information between image and tabular data by dynamically scaling and shifting the feature maps in a convolutional layer based on both the patient’s endoscopic images and clinical tabular information. This allows the CNN to leverage the complementary information from tabular data for improved prediction. Second, DAFT creates a bottleneck structure to compress the spatial dimensions of the image feature map through global average pooling, and then concatenates it with the tabular data. Through a series of fully connected layers and nonlinear transformations, it obtains the scaling and offset vectors for feature map transformation, which can dynamically adjust the feature maps according to the specific image and tabular information to better fuse the two types of data. In addition, DAFT is a versatile module that can be easily applied to any type of CNN for tabular information fusion, showing wide applicability in different application scenarios. Extensive experiments, including comparisons with multiple baseline methods, ablation studies, analysis of the contribution of tabular data features, and generalization and repeatability experiments on different datasets, demonstrate the effectiveness and superiority of DAFT in fusing image and tabular data, as well as its robustness to design choices. 29 In this study, our multimodal model (DAFT) integrates information from two modalities: clinical biomarkers and endoscopic images. The clinical biomarkers include those mentioned above that help predict cytomegalovirus infection: white blood cell count, lymphocyte count, eosinophil count, platelet count, and IgA level. The two modalities reflect complementary aspects of UC and CMV colitis. Obviously, our results demonstrate that the multimodal model, taking into account both image data and clinical biomarker data, outperforms the unimodal, image-based ResNet and SeNet with attention module across all tasks and metrics, which is consistent with a prior work utilizing a similar multimodal model. 20 Although a previous study has considered combining image and clinical parameters as a hybrid algorithm to improve the CNN’s poor sensitivity for IBD and low specificity for infectious colitis, 14 our multimodal model (DAFT) is superior in distinguishing CMV colitis and UC, with higher accuracy and ROC curve area.
The model’s primary goal is to guide endoscopists in identifying suspicious CMV colitis lesions during live examinations, ensuring no high-risk areas are missed. In this context, sensitivity—defined as the model’s ability to detect true-positive cases—directly addresses the clinical imperative to minimize underdiagnosis. A higher sensitivity reduces the likelihood of overlooking CMV-infected tissues, which could lead to delayed antiviral therapy. While specificity (0.931) remains important to limit unnecessary biopsies, the trade-off favors sensitivity in this pre-biopsy screening role, where the consequences of missed detection far outweigh the risks of additional sampling.
In the future, it is expected to develop a software system for identifying CMV colitis in UC patients based on existing models. This software can be used during colonoscopy to identify the manifestations of CMV colitis in real time, guide targeted biopsies, and improve the detection rate of CMV colitis. However, it is important to acknowledge the several limitations of the model.
Currently, despite the outstanding performance of the model, it cannot replace pathological examination as a gold standard in the diagnosis of CMV colitis, but it is an initial step in the process of development. The lack of external validation using independent datasets may affect the generalizability of our model. To establish more robust algorithms and validate the generalizability of our model, multicenter datasets with diverse endoscopic appearances and patient demographics are essential. In this study, we merely utilized images taken by endoscopists in real-life scenarios, without consistent information on localization. Our model is developed based on high-quality images, and its application under real-world suboptimal conditions, such as a blurry field of view due to poor intestinal preparation, may be affected. Furthermore, still images lack important information regarding the distribution of findings throughout the colon. Alternatively, implementation of video-based algorithms could be considered, but large datasets have yet to be established and linked to clinical data. It is important to bear in mind that in real-life scenarios, other infectious pathologies 30 (such as Epstein-Barr virus-induced colitis) may have a similar clinical and endoscopic manifestation to the CMV colitis studied here, which makes the classification task even more complex.
Conclusion
The increasing application of AI provides a promising opportunity to improve the diagnostic capabilities and clinical management of UC and coexisting CMV colitis. By combining clinical data and endoscopic images, the proposed multimodal model has the potential to assist clinicians in accurate and timely recognition and improving the prognosis of UC patients with concomitant CMV colitis.
Supplemental Material
Supplemental material, sj-docx-1-tag-10.1177_17562848251364194 for Artificial intelligence-based multimodal model for the identification of ulcerative colitis with concomitant cytomegalovirus colitis by Haozheng Liang, Yuxuan Tian, Gechong Ruan, Xiaoyin Bai, Wei Han, Xiangling Fu, Yuhang Wang, Jialin Shi, Yinghao Sun, Ji Wu, Chenyi Guo and Hong Yang in Therapeutic Advances in Gastroenterology
Acknowledgments
We thank the endoscopists at Peking Union Medical College Hospital for their assistance in data collection. Special thanks to Dr. Xiaowei Xue from the Department of Pathology for her expertise in confirming CMV viral inclusions in pathological sections, which was critical for the study’s diagnostic validation.
Footnotes
ORCID iDs: Haozheng Liang  https://orcid.org/0009-0002-7894-2965
https://orcid.org/0009-0002-7894-2965
Xiaoyin Bai  https://orcid.org/0000-0002-9092-7592
https://orcid.org/0000-0002-9092-7592
Hong Yang  https://orcid.org/0000-0002-2986-7324
https://orcid.org/0000-0002-2986-7324
Supplemental material: Supplemental material for this article is available online.
Contributor Information
Haozheng Liang, Department of Gastroenterology, Peking Union Medical College Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing, China.
Yuxuan Tian, Department of Electronic Engineering and the Institute for Precision Medicine, Tsinghua University, Beijing, China.
Gechong Ruan, Department of Gastroenterology, Peking Union Medical College Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing, China.
Xiaoyin Bai, Department of Gastroenterology, Peking Union Medical College Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing, China.
Wei Han, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences, School of Basic Medicine, Peking Union Medical College, Beijing, China.
Xiangling Fu, School of Computer Science (National Pilot Software Engineering School), Beijing University of Posts and Telecommunications, Beijing, China.
Yuhang Wang, School of Computer Science (National Pilot Software Engineering School), Beijing University of Posts and Telecommunications, Beijing, China.
Jialin Shi, School of Computer and Communication Engineering, University of Science and Technology Beijing, Beijing, China.
Yinghao Sun, Department of Gastroenterology, Peking Union Medical College Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing, China.
Ji Wu, Department of Electronic Engineering and the Institute for Precision Medicine, Tsinghua University, Haidian District, Beijing, 100084, China.
Chenyi Guo, Department of Electronic Engineering and the Institute for Precision Medicine, Tsinghua University, Haidian District, Beijing, 100084, China.
Hong Yang, Department of Gastroenterology, Peking Union Medical College Hospital, Chinese Academy Medical Sciences & Peking Union Medical College, No.1 Shuaifuyuan Wangfujing Dongcheng District, Beijing 100730, China.
Declarations
Ethics approval and consent to participate: This study was approved by the Institutional Review Board (IRB) of Peking Union Medical College Hospital (approval number: I-23PJ1345). Due to the retrospective nature of the study and the use of anonymized clinical data, the IRB granted a waiver of informed consent in accordance with local regulations and the Helsinki Declaration (2013 Update).
Consent for publication: Not applicable (no identifiable patient data are included in this study).
Author contributions: Haozheng Liang: Data curation; Writing – original draft.
Yuxuan Tian: Formal analysis; Writing – original draft.
Gechong Ruan: Investigation; Project administration.
Xiaoyin Bai: Data curation; Project administration.
Wei Han: Investigation.
Xiangling Fu: Investigation.
Yuhang Wang: Formal analysis; Investigation.
Jialin Shi: Formal analysis.
Yinghao Sun: Writing – review & editing.
Ji Wu: Supervision.
Chenyi Guo: Supervision; Writing – review & editing.
Hong Yang: Funding acquisition; Supervision; Writing – review & editing.
Funding: The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Key R&D Program of China (2023YFC2507302), Capital Health Research and Development of Special Foundation (2022-2-4014), National High-Level Hospital Clinical Research Funding (2022-PUMCH-B-022), CAMS Innovation Fund for Medical Sciences (2022-I2M-C&T-B-011), State Key Laboratory Special Fund (2060204), National Key Clinical Specialty Construction Project (ZK108000) and Beijing Natural Science Foundation (7252113).
Competing interests: The authors declare that there is no conflict of interest.
Availability of data and materials: The datasets generated and analyzed during the current study are available from the corresponding author upon reasonable request, with restrictions to protect patient anonymity as approved by the IRB.
References
- 1. Yang H, Zhou R, Bai X, et al. Trend and geographic variation in incidence and prevalence of inflammatory bowel disease in regions across China: a nationwide employee study between 2013 and 2016. Front Med (Lausanne) 2022; 9: 900251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Beaugerie L, Rahier JF, Kirchgesner J. Predicting, preventing, and managing treatment-related complications in patients with inflammatory bowel diseases. Clin Gastroenterol Hepatol 2020; 18(6): 1324. [DOI] [PubMed] [Google Scholar]
- 3. Oh SJ, Lee CK, Kim YW, et al. True cytomegalovirus colitis is a poor prognostic indicator in patients with ulcerative colitis flares: the 10-year experience of an academic referral inflammatory bowel disease center. Scand J Gastroenterol 2019; 54(8): 976–983. [DOI] [PubMed] [Google Scholar]
- 4. Zhang M, Bai X, Zhang H, et al. The role of cytomegalovirus colitis on short- and long-term outcomes for patients with ulcerative colitis. Scand J Gastroenterol 2022; 57(3): 282–289. [DOI] [PubMed] [Google Scholar]
- 5. Kambham N, Vij R, Cartwright CA, et al. Cytomegalovirus infection in steroid-refractory ulcerative colitis: a case-control study. Am J Surg Pathol 2004; 28(3): 365–373. [DOI] [PubMed] [Google Scholar]
- 6. Kucharzik T, Ellul P, Greuter T, et al. ECCO guidelines on the prevention, diagnosis, and management of infections in inflammatory bowel disease. J Crohns Colitis 2021; 15(6): 879–913. [DOI] [PubMed] [Google Scholar]
- 7. Lawlor G, Moss AC. Cytomegalovirus in inflammatory bowel disease: pathogen or innocent bystander?. Inflamm Bowel Dis 2010; 16(9): 1620–1627. [DOI] [PubMed] [Google Scholar]
- 8. Maresca R, Varca S, Di Vincenzo F, et al. Cytomegalovirus infection: an underrated target in inflammatory bowel disease treatment. J Clin Med 2023; 13(1): 130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Mccurdy JD, Enders FT, Jones A, et al. Detection of cytomegalovirus in patients with inflammatory bowel disease: where to biopsy and how many biopsies?. Inflamm Bowel Dis 2015; 21(12): 2833–2838. [DOI] [PubMed] [Google Scholar]
- 10. Yang H, Zhou W, Lv H, et al. The association between CMV viremia or endoscopic features and histopathological characteristics of CMV colitis in patients with underlying ulcerative colitis. Inflamm Bowel Dis 2017; 23(5): 814–821. [DOI] [PubMed] [Google Scholar]
- 11. Yang H, Ran Z, Jin M, et al. Current status of opportunistic infection in inflammatory bowel disease patients in asia: a questionnaire-based multicenter study. Gut Liver 2022; 16(5): 726–735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Okagawa Y, Abe S, Yamada M, et al. Artificial intelligence in endoscopy. Dig Dis Sci 2022; 67(5): 1553–1572. [DOI] [PubMed] [Google Scholar]
- 13. Cai W, Xu J, Chen Y, et al. Performance of machine learning algorithms for predicting disease activity in inflammatory bowel disease. Inflammation 2023; 46(4): 1561–1574. [DOI] [PubMed] [Google Scholar]
- 14. Guimaraes P, Finkler H, Reichert MC, et al. Artificial-intelligence-based decision support tools for the differential diagnosis of colitis. Eur J Clin Invest 2023; 53(6): e13960. [DOI] [PubMed] [Google Scholar]
- 15. Inflammatory Bowel Disease Group, Chinese Society of Gastroenterology, Chinese Medical Association; Inflammatory Bowel Disease Quality Control Center of China. Chinese clinical practice guideline on the management of ulcerative colitis (2023, Xi′an). Chin J Dig 2024; 44(2): 73–99. [Google Scholar]
- 16. Maaser C, Sturm A, Vavricka SR, et al. ECCO-ESGAR guideline for diagnostic assessment in IBD Part 1: initial diagnosis, monitoring of known IBD, detection of complications. J Crohns Colitis 2019; 13(2): 144–164. [DOI] [PubMed] [Google Scholar]
- 17. Wu K, Liang J, Ran Z, et al. Chinese consensus on diagnosis and treatment in inflammatory bowel disease (2018, Beijing). Chin J Digest 2018; 38(05): 292–311. [DOI] [PubMed] [Google Scholar]
- 18. Suzuki S, be Keiichi A. Topological structural analysis of digitized binary images by border following. Comp Vis Grap Image Process 1985; 29(3): 396. [Google Scholar]
- 19. Bossuyt PM, Reitsma JB, Bruns DE, et al. STARD 2015: an updated list of essential items for reporting diagnostic accuracy studies. BMJ 2015; 351: h5527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Wolf TN, Polsterl S, Wachinger C, et al. DAFT: a universal module to interweave tabular data and 3D images in CNNs. Neuroimage 2022; 260: 119505. [DOI] [PubMed] [Google Scholar]
- 21. Stafford IS, Gosink MM, Mossotto E, et al. A systematic review of artificial intelligence and machine learning applications to inflammatory bowel disease, with practical guidelines for interpretation. Inflamm Bowel Dis 2022; 28(10): 1573–1583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Kraszewski S, Szczurek W, Szymczak J, et al. Machine learning prediction model for inflammatory bowel disease based on laboratory markers. Working model in a discovery cohort study. J Clin Med 2021; 10(20): 4745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Gui X, Bazarova A, Del Amor R, et al. PICaSSO Histologic Remission Index (PHRI) in ulcerative colitis: development of a novel simplified histological score for monitoring mucosal healing and predicting clinical outcomes and its applicability in an artificial intelligence system. Gut 2022; 71(5): 889–898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Chen Y, Zheng Q, Wang H, et al. Integrating transcriptomics and proteomics to analyze the immune microenvironment of cytomegalovirus associated ulcerative colitis and identify relevant biomarkers. BioData Min 2024; 17(1): 26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. El Helou G, Lahr B, Razonable R. Absolute lymphocyte count as marker of cytomegalovirus and allograft rejection: is there a “Safe Corridor” after kidney transplantation?. Transpl Infect Dis 2021; 23(2): e13489. [DOI] [PubMed] [Google Scholar]
- 26. Yang H, Wu K, Zhang H, et al. IgA, albumin, and eosinopenia as early indicators of cytomegalovirus infection in patients with acute ulcerative colitis. BMC Gastroenterol 2020; 20(1): 294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Altunal LN, Ozel AS. and A K C. Cytomegalovirus reactivation in ulcerative colitis patients: early indicators. Niger J Clin Pract 2023; 26(6): 765–770. [DOI] [PubMed] [Google Scholar]
- 28. Kim JH, Choe AR, Byeon JR, et al. Enhancing the predictions of cytomegalovirus infection in severe ulcerative colitis using a deep learning ensemble model: development and validation study. JMIR Med Inform 2025; 13: e64987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Plsterl S, Wolf TN, Wachinger C. Combining 3D image and tabular data via the dynamic affine feature map transform. arXiv:2107.05990, 2021. [Google Scholar]
- 30. Liu Y, Li Y, Li Y, et al. Clinical features of intestinal ulcers complicated by Epstein-Barr virus infection: importance of active infection. Dis Markers 2021; 2021: 6627620. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental material, sj-docx-1-tag-10.1177_17562848251364194 for Artificial intelligence-based multimodal model for the identification of ulcerative colitis with concomitant cytomegalovirus colitis by Haozheng Liang, Yuxuan Tian, Gechong Ruan, Xiaoyin Bai, Wei Han, Xiangling Fu, Yuhang Wang, Jialin Shi, Yinghao Sun, Ji Wu, Chenyi Guo and Hong Yang in Therapeutic Advances in Gastroenterology