Abstract
Objective:
To develop scanning electron microscopy-based Trainable Weka (Waikato Environment for Knowledge Analysis) Intelligent Segmentation Technology (SEMTWIST), an open-source software tool, for structural detection and rigorous quantification of wound biofilm aggregates in complex human wound tissue matrix.
Approach:
SEMTWIST model was standardized to quantify biofilm infection (BFI) abundance in 240 distinct SEM images from 60 human chronic woundedge biospecimens (four technical replicates of each specimen). Results from SEMTWIST were compared against human expert assessments and the gold standard for molecular BFI detection, that is, peptide nucleic acid fluorescence in situ hybridization (PNA-FISH).
Results:
Correlation and Bland–Altman plot demonstrated a robust correlation (r = 0.82, p < 0.01), with a mean bias of 1.25, and 95% limit of agreement ranging from −43.40 to 47.11, between SEMTWIST result and the average scores assigned by trained human experts. While interexpert variability highlighted potential bias in manual assessments, SEMTWIST provided consistent results. Bacterial culture detected infection but not biofilm aggregates. Whereas the wheat germ agglutinin staining exhibited nonspecific staining of host tissue components and failed to provide a specific identification of BFI. The molecular identification of biofilm aggregates using PNA-FISH was comparable with SEMTWIST, highlighting the robustness of the developed approach.
Innovation:
This study introduces a novel approach “SEMTWIST” for in-depth analysis and precise differentiation of biofilm aggregates from host tissue elements, enabling accurate quantification of BFI in chronic wound SEM images.
Conclusion:
Open-source SEMTWIST offers a reliable and robust framework for standardized quantification of BFI burden in human chronic wound-edge tissues, supporting clinical diagnosis and guiding treatment.
Keywords: human chronic wound, biofilm infection, machine learning, scanning electron microscopy, wheat germ agglutinin, culture
INTRODUCTION
Biofilms in chronic wounds act as hostile microbial aggregates as they remain encased within their polymeric matrix cocoons, making them resistant to both antimicrobial therapy and host immune defenses.1 Biofilms create a persistent inflammatory state that disrupts normal wound healing processes by impairing reepithelialization and angiogenesis.2 Chronic wounds with biofilms are more likely to develop recalcitrant severe infections, leading to complications such as amputation and tissue necrosis.3 Without a proper diagnosis, clinicians may rely on systemic antibiotics, which are often ineffective against biofilm-associated bacteria due to their protective extracellular polymeric substance (EPS) matrix.4 This can lead to prolonged antibiotic use, contributing to antimicrobial resistance without effectively resolving the infection.5 Biofilms exhibit up to 1,000-fold increased resistance to antibiotics compared with planktonic bacteria, making treatment challenging.6
Traditional culture-based methods often fail to detect biofilm infection (BFI) in wound tissues, as the biofilm-associated bacteria enter a dormant state, rendering them undetected using conventional microbiological diagnostic techniques.7 Diagnosing BFI in chronic wounds remains challenging, with no standardized clinical protocol currently established for clinical research practice.8-11 Reliable and robust clinical assessment of BFI burden in wounds is critical to determining treatment strategy.12,13 The extent to which the BFI burden negatively impacts wound healing outcomes depends on the balance between BFI threat and countering defenses, including host factors and therapeutic intervention.1,3,14,15 The conventional methods, such as colony forming unit (CFU) assays and bacterial culture, can only identify planktonic bacterial growth and may not be reliable in detecting BFI in wounds because of the slow and fastidious characteristics of the growth of biofilm bacteria.12,16 In the fastidious state, bacteria within biofilm exhibit metabolic dormancy, which slows down their growth. As a result, biofilm bacteria may not reach detectable levels within the set incubation period of 24–48 h of the standard culture method, leading to falsenegative results.17-19 Of various methods available for qualitative structural BFI detection on abiotic or biotic surfaces, scanning electron microscopy (SEM)-based detection of BFI structure is considered the gold standard.20-22 The European Society of Clinical Microbiology and Infectious Diseases recommends using SEM as one of the most reliable diagnostic techniques for detecting biofilms in wounds.23,24 The high resolution of SEM allows detailed analysis of biofilm structures, allowing direct visualization by achieving a resolution of 50 to 100 nm.25 This resolution allows for in-depth examination of fine bacterial surface features, biofilm architecture, and nanoscale bacterial interactions.21 However, in the present study, our primary focus was on identifying biofilm aggregates within human chronic wound-edge tissue. Therefore, the images were captured with consideration of the size of host tissue features observed, which were crucial for training the algorithm for biofilm quantification.
One of the challenges of SEM-based BFI detection in wound-edge tissue is the lack of availability of robust and objective quantification methods. Although the visual system of humans, with expert knowledge, can interpret and provide relative BFI abundance in SEM images from wound-edge tissue,26 such assessments are primarily subjective, time-intensive, limited by the availability of expert raters, and prone to interobserver variability.27-30 This variability often stems from differences in expertise, image noise, and the complex anatomical architecture of tissues. Quantitative methods, such as machine learning (ML)- and artificial intelligence (AI)-based models31-33 for image analysis and processing, may help minimize bias and improve reproducibility in biofilm quantification of wound SEM images. Weka, an open-source software tool that enables users to perform ML-based image segmentation, object classification, and feature extraction with minimal programming experience,33-35 has been widely used in biological and clinical research to quantitatively analyze various types of microorganisms, cells, and tissues.36-40
This study aimed to develop and standardize an objective ML-based approach using the trainable Weka intelligent segmentation technology (TWIST) tool capable of grading BFI burden in the chronic wound-edge tissues into the following clinically relevant categories: none, low, intermediary, or high BFI abundances.
Various ML algorithms within the TWIST framework have been used for several pixel-based binary segmentation tasks across different image modalities, including bacterial counting, growth assessment, and cell segmentation.41
This gradation of BFI abundance is essential to diagnose BFI promptly, allowing for the implementation of effective treatment strategies and significantly reducing the risk of poor outcomes. A reliable quantification approach is also essential for clinical research studies evaluating antibiofilm treatments, where BFI reduction is a primary outcome. The article has been designed as per the Strengthening the Reporting of Observational Studies in Epidemiology guidelines (Fig. 1).
Figure 1.
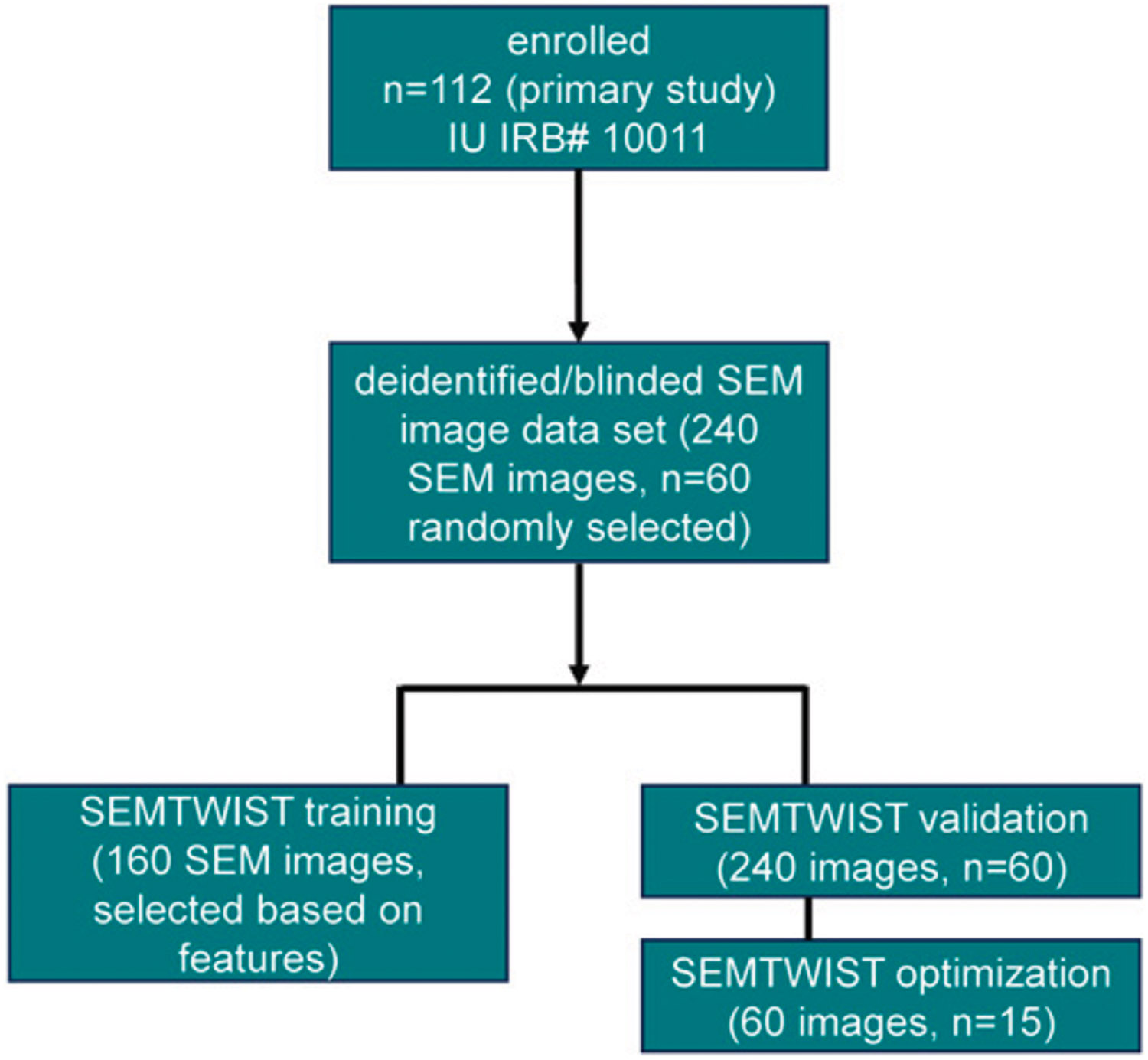
Schematic illustrating a total of 112 patients enrolled in an ongoing, unreported clinical trial (IRB# 10011, Indiana University). From this primary cohort, a blinded subset of 240 SEM images (four technical replicates from each participant) were obtained from n = 60 (randomly selected) participants. A set of 160 SEM images (130 unique) were used to train SEMTWIST for identifying biofilm and nonbiofilm features within SEM images. Following the training phase, the deidentified dataset of 240 images (n = 60) was used to validate SEMTWIST for quantifying biofilm infection abundance in the chronic wound tissue. A random subset (n = 15 of the n = 60) was selected to optimize SEMTWIST’s performance by comparing its results with those from human rater assessments, WGA staining, culture pathology, and PNA-FISH techniques (bottom right), as specified in subsequent figures. PNA-FISH, peptide nucleic acid fluorescence in situ hybridization; SEM, scanning electron microscopy; SEMTWIST, scanning electron microscopy-based Trainable Weka (Waikato Environment for Knowledge Analysis) Intelligent Segmentation Technology; WGA, wheat germ agglutinin.
INNOVATION
SEM analysis reveals that wound biofilm, shaped by an iterative exchange between microbial and host-defense processes, is a polymicrobial aggregate including host cells. In the present work, we standardized the TWIST algorithm to robustly quantify BFI in SEM images from intricate human chronic wounds, characterizing biofilm distribution within the wound tissue in relation to the surrounding host tissue microenvironment and landscape features.
Clinical problem addressed
Wound biofilm aggregates in chronic human wounds exhibit unique characteristics, and their quantification in clinical samples has long been challenging due to limited understanding of the complex structural interactions between biofilm and the wound tissue. Fluorescence-based techniques such as wheat germ agglutinin (WGA) staining have shown promising results for in vitro biofilm experiments with limited efficacy for detecting biofilms within infected wound tissue samples.42,43 Advanced methods such as peptide nucleic acid fluorescence in situ hybridization (PNA-FISH)44 provide insights into the microbial composition and distribution but do not offer details about the structural and topographic interactions between biofilms and tissues that SEM provides. Similarly, reverse transcription-polymerase chain reaction evaluates biofilm-related gene expression but does not offer insights into the spatial organization of the EPS matrix and biofilm morphology.45-47 Understanding these complex structural interactions is crucial for accurate and reliable biofilm quantification, ultimately supporting improved diagnostics and treatment strategies.48
MATERIALS AND METHODS
Methods
Human subjects and tissue collection from chronic wounds.
Study design.
A blinded subset of 240 SEM images (n = 60 subjects) from an ongoing, unreported clinical trial (IRB# 10011, Indiana University) were used for SEMTWIST optimization and validation. These images were analyzed without the knowledge of subject identity or clinical outcome. This dataset was solely used for SEMTWIST development and is distinct from the primary analysis of this study, which will be reported separately upon completion of trial outcomes. All participants provided written informed consent before inclusion in the primary study (IRB# 10011).
Participants.
Subjects in the study were aged 18 years and older with infected chronic wounds from a prospective clinical study. Demographic characteristics of the 60 subjects whose SEM image data were used for SEMTWIST optimization and validation are summarized in Table 1.
Table 1.
Demographic characteristics and wound type of the human participants
| Population characteristics | Number |
|---|---|
| Age, years | |
| Mean | 56 |
| Range | 33–80 |
| Sex | |
| Female | 17 |
| Male | 43 |
| Type of wound | |
| Diabetic foot ulcer | 9 |
| Venous leg ulcer | 16 |
| Pressure ulcer | 7 |
| Surgical | 9 |
| Other | 19 |
SEM sample processing and imaging for biofilm detection.
Wound biopsies were collected in a 2.5% glutaraldehyde fixation buffer from patients at the Indiana University wound clinics. For processing, the samples were washed with phosphate-buffered saline and resuspended in ethanol, followed by dehydration using a graded ethanol series (50%, 70%, 90%, and 100%). Subsequently, the samples were treated with hexamethyldisilazane (HMDS) (Ted Pella, Inc., Redding, CA) in combination with ethanol at varying ratios: 100% ethanol: HMDS—3:1, 100% ethanol: HMDS—1:1, and 100% ethanol: HMDS—1:3. Finally, the samples were treated with pure HMDS and left to dry overnight under a hood.
After drying, the samples were desiccated in a vacuum chamber for at least 12 h before imaging. The dried samples were then gold-coated using a Cressington turbo-pumped sputter coater. For biofilm detection and tissue morphology examination, SEM images were obtained using a field emission scanning electron microscope (FESEM) (JEOL 7800F, JEOL Japan) at a beam energy of 5 kV and 5,000× magnification. Images were captured from four different regions/locations within the same tissue sample to ensure comprehensive coverage of the tissue regions. For selection of four regions of the tissue, a low-magnification (60×) image was obtained. The image was digitally segmented into four quadrants, and one representative high-magnification (5,000×) image from each quadrant was used.
Quantitative biofilm assessment in SEM images using TWIST.
SEM images obtained from patients’ wound biopsies were analyzed using TWIST software, a plug-in ImageJ-based Fiji software. Image analysis included three major steps as follows:
a. Image preprocessing.
All SEM images were saved in .tiff form at 5,000× magnification to have a broader field of view. In each image, the scale bar was removed using scripts with boundary detection function as part of the preprocessing. All the SEM images’ pixel dimensions (2,048 × 2,560) and bit depth (8 bits) were kept constant to enable batch processing. No background subtraction or filtering techniques were applied before segmentation, ensuring that the integrity of the tissue features was preserved for our study.
b. Image classification and segmentation by TWIST training.
The TWIST training was conducted using two primary types of SEM images: (i) SEM images without BFI (visible host tissue features only): Before biofilm segmentation, the TWIST software was utilized to highlight and elucidate the architectural features of host tissue in SEM images of wound-edge tissue obtained from patients.
These images served as a baseline for understanding the structural characteristics of the tissue without biofilm interference. (ii) SEM images with BFI (visible host tissue features and biofilm): The biofilm in the image was segmented using TWIST, which uses a suite of ML algorithms for effective segmentation. A suite consisted of a collection of supervised ML algorithms integrated with ImageJ, utilizing user-labeled training data to train a supervised ML classifier (specifically a fast random forest classifier in this article) for complex image segmentation. The algorithm integrated feature extraction functions that enhanced the detection of variations in important SEM image characteristics (such as edges, textures, and intensity). These extracted features served as inputs for classifiers, which learned to differentiate between biofilm and nonbiofilm surface patterns. The trained model was then applied to a new SEM image, enabling automated segmentation with improved accuracy. Relevant pixels within the image were selected and manually drawn using the drawing tool, dividing the image into biofilm and surface classes. The surface class encompassed complex tissue features with varying morphologies and characteristics. The size of the pixel regions was chosen to include all tissue and biofilm key features. The classifier was trained in a semisupervised manner using these samples, with the default classifier (fast random forest) used for all analyses in this study. The pixels could be adjusted or repositioned to retrain the classifier for improved segmentation accuracy. The following feature filters were selected to create a classifier for most images: Gaussian blur, Sobel filter, Hessian, difference of Gaussians, membrane projections, variance, mean, minimum, maximum, and median.38 This combination provided the most effective segmentation within the shortest time frame. Other Weka settings were kept at default (membrane thickness 1, membrane patch size 19, minimum sigma 1.0, and maximum sigma 16.0). These filters helped in identifying irregular surface features of bacterial biofilms, characterized by their clustered microbial cells and thick EPS matrix by emphasizing the subtle variations in topography, texture, intensity, and contrast within the image. This resulted in a reliable distinction between biofilm aggregates and surrounding host tissue features with enhanced efficiency.
Batch processing was feasible for some images, allowing a single trained classifier to be applied across multiple images for automatic segmentation. For example, one classifier could be created for images without biofilm, focusing solely on tissue features, and another for images with ≥70% biofilm coverage. Due to the complexity of tissue architecture and the variable locations of embedded biofilm, some images were processed using a customized training dataset that was tailored to include distinct features within the images. By selecting relevant pixels, the classifier learns to recognize and differentiate these patterns. This process was verified by carefully zooming in and out of the images to ensure proper feature recognition. A total of 160 SEM images were used to train the software for recognizing both with and without BFI features. Of the 240 images from which data have been provided in Fig. 1D, a set of 30 images, and an additional 130 unique images were used.
c. Image postprocessing.
The histogram of each image was analyzed to determine the number of white pixels representing the biofilm area. Using TWIST’s “get probability” feature, images were generated with black and white regions corresponding to non-BFI and BFI classes, respectively. Thresholding was then applied to assign distinct colors to each class in the histogram, yielding the percentage of biofilm coverage in each image. In addition, k-means clustering algorithm was used for postprocessing where the pixels were classified into a predefined number of color clusters. Each pixel was assigned to a cluster, and binary masks were generated for each color class. The percentage of biofilm (a predetermined color class in TWIST segmentation) was calculated by determining the proportion of pixels in that class relative to the total number of pixels in the image.
Quantitative biofilm assessment in SEM images by human raters.
SEM images were coded and provided to four biofilm experts (3–4 years of experience with wound BFI and SEM images) for blinded evaluation. A total of 60 SEM images from 15 subjects (four images per subject) were selected for rating. These images were taken from different regions of each sample (left, right, and center) to minimize bias. The experts assessed the biofilm abundance based on the percentage coverage in each image. The categories for biofilm abundance were as follows: ≤10% (no biofilm, with prominent host tissue features visible), 10% to <40% (low biofilm, characterized by few bacteria and associated EPS matrix observed), 40% to <70% (intermediate biofilm, with biofilm structures covering multiple tissue regions), and ≥70% (dense biofilm, where most tissue features are obscured by biofilm structures).
Tissue preparation for PNA-FISH analysis.
The PNA-FISH was performed to detect biofilm in formalin-fixed and paraffin-embedded (FFPE) tissue sections. Briefly, the FFPE tissue sections were deparaffinized through xylene and ethanol to water before probe hybridization. Then, the tissue sections were fixed with 4% PFA for 10 min at RT, followed by incubation of 10 min with 50% ethanol. Afterward, the Staphylococcus aureus- and Pseudomonas aeruginosa-specific hybridization solution was added to the section, and hybridization was performed at 57°C for 90 min. Afterward, the tissue sections were washed in a wash solution for 30 min at 57°C. Finally, the nuclei were stained with DAPI solution for 5 min at room temperature, and the slides were mounted with ProLong™ gold antifade mount medium (Invitrogen: P36930). The images were captured using a confocal laser scanning microscopy (LSM 880, Zeiss) and AxioScan Z1 (Zeiss).
Biofilm matrix staining.
Biofilm staining using the WGA-Alexa Fluor 488 conjugate was conducted to detect biofilm in FFPE human woundedge tissue sections, following the previously published protocol.43
Culture pathology.
Wound tissues were obtained from each patient after cleaning the wound surface with saline. The sample was kept in a sterile container and transported to a clinical pathology laboratory for standard bacterial culture-based analysis by inoculating the samples on tryptic soy agar to detect bacterial load.
Image adjustment analysis.
We performed sensitivity analysis for TWIST performance on five randomly selected SEM images. In each image, the annotation bar was removed using scripts with boundary detection function as part of the preprocessing using MATLAB script. Using the bicubic interpolation method,49 the images were downscaled or upscaled to 320 × 256, 480 × 384, 640 × 512, 960 × 768, 1,280 × 1,024, and 2,560 × 2,048 pixels while maintaining the original aspect ratio. The TWIST segmentation was trained on the same instance locations for two classes (BFI and non-BFI) to obtain classification results. The output classified figure from TWIST was processed using k-means clustering.
Statistical analysis
The raw data (n = 15, 60 SEM images) were compiled and categorized into two groups—human raters and TWIST—to evaluate correlation and agreement for BFI abundance data. Regression analysis and correlation methods in GraphPad Prism and Microsoft Excel were used to assess the correlation between groups. Bland–Altman plot50 was made in R version 4.3.2 to determine agreement levels between individual human rater’s BFI data and the rater average and TWIST. Limits of agreements (LoAs) were calculated as the mean difference ± (1.96 × standard deviation of the difference). In the Bland–Altman plot, the y-axis represents the differences between two measurements, while the x-axis shows their mean value.
RESULTS
SEM-ML tool for BFI assessment
To quantify biofilm infections in chronic wound-edge tissues using SEM images, we used an ML-based approach and validated this method using a known set of biofilm infected wound-edge tissues. SEM images were obtained and analyzed by experts alongside our SEM-ML system (Fig. 2A). To enhance the clinical relevance of the assessment for treatment strategy, the % BFI abundance data output from ML-based quantification was classified into four grades: (i) no BFI, (ii) low (BFIlo), (iii) intermediate (BFIint), or (iv) high (BFIhi) (Fig. 2B). Representative raw SEM images and the corresponding ML-processed images of the wound BFI gradations are shown in Fig. 2C. We observed that SEM-ML analysis was able to reliably detect all the abovementioned grades of BFI by accurately segmenting BFI and non-BFI regions. To establish the feasibility of the BFI gradation approach, we performed an SEM-ML assessment of BFI abundance in 240 unique SEM images obtained from human (n = 60) chronic wound-edge tissues that included four technical replicates for each biospecimen (Fig. 2D). Recognizing that human expert raters are the standard for assessing BFI in SEM images,26 we conducted a comparison between the results of SEM-ML analysis and those from human assessments using data from randomly selecting 60 images from a total set of 240 images.
Figure 2.
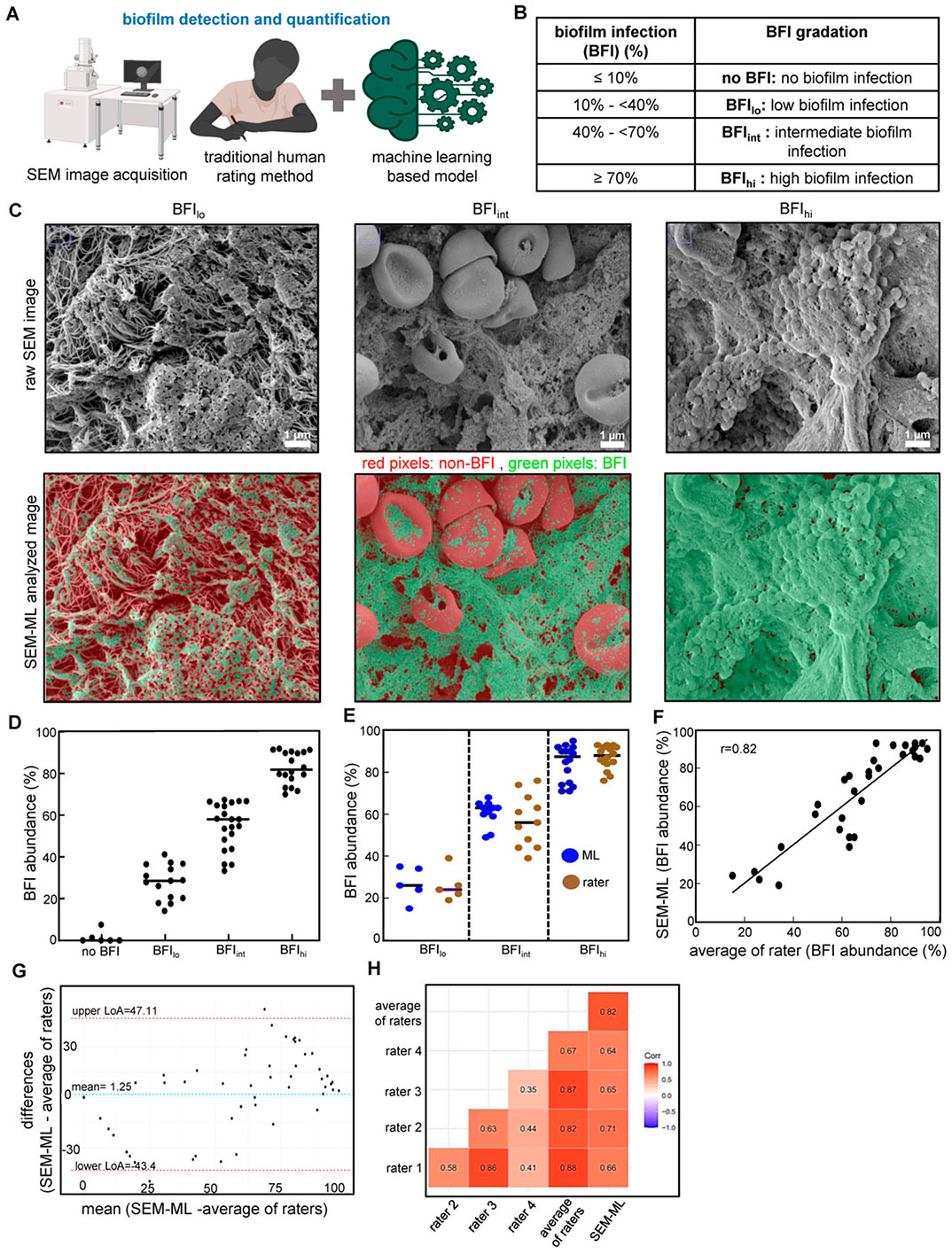
Quantification assessment of biofilm infection (BFI) abundance in human chronic wound edge using scanning electron microscopy (SEM)-machine learning (ML). (A) Schema showing BFI detection and quantification in human chronic wound-edge tissues using SEM image acquisition. BFI quantification was performed using (i) traditional human rater or (ii) novel ML-based approaches; (B) Classification of % BFI abundance obtained using ML and corresponding classification of BFI abundance used by expert human raters: no BFI, low (BFIlo), intermediate (BFIint), or high (BFIhi); (C) raw SEM images (top panel) of human wound-edge tissue samples, BFI abundance was processed and quantified using ML (processed image, bottom panel) approach and classified to BFI abundance. Representative images from BFIlo, BFIint, or BFIhi are shown. (D) Human chronic wound-edge tissue BFI classification using SEM-ML demonstrates the feasibility of the approach for clinical application. The data presented were obtained from human chronic wound edges, with a total of 240 images, unique SEM images quantified (n = 60), including four technical replicates from each wound-edge specimen; (E) a subset of 60 images (n = 15 subjects) was selected from the primary cohort (n = 60) for validation of the quantitative SEM-ML data against a traditional “Expert Rater based” subjective assessment of BFI abundance; this assessment utilized 4 rater expert in biofilm biology; (F) regression analysis showing a positive correlation (p < 0.05), with a correlation coefficient (r = 0.82) between the rater-generated results and the ML-generated data; (G) Bland–Altman plot indicating good agreement between BFI abundance data between human raters and SEM-ML analysis; and (H) correlation plot illustrating interrater variability while a strong correlation between the raters′ mean value and ML quantification.
As shown in Fig. 2E, F, a strong correlation (r = 0.82, p < 0.01) was observed between the results from human experts and those produced by SEM-ML analysis, which was further validated by the Bland–Altman plot (Fig. 2G). In the plot, the mean bias and 95% LoA was 1.25 with −43.40 to 47.11, between the average BFI abundance for expert human raters versus the data from SEM-ML analysis. Nonetheless, correlation analysis revealed interrater variability in the human analysis, highlighting the subjective nature of human assessment (Fig. 2H). Substantial variability between the trained human raters was also noted in the Bland–Altman plot as shown by mean bias and the wide LoAs 17.8 (−33.34 to 68.93) (Supplementary Fig. S1). These findings suggest low reliability and reproducibility of BFI assessment of SEM image datasets by human experts. Interestingly, a strong correlation was noted between the human expert rater averages and the SEM-ML approach, demonstrating the validity and robustness of the SEM-ML-based assessment of clinical BFI abundance in human wound specimens.
These findings also indicate that human expert rating is only valid when performed by >3 raters. In this study, a stepwise analysis procedure of SEMTWIST has been described (Fig. 3). All segmentation results were saved as MACRO test files with MATLAB scripts, which as needed can be reopened and used for future reference. The detailed outline of each step used for processing is described in Supplementary Fig. S2.
Figure 3.
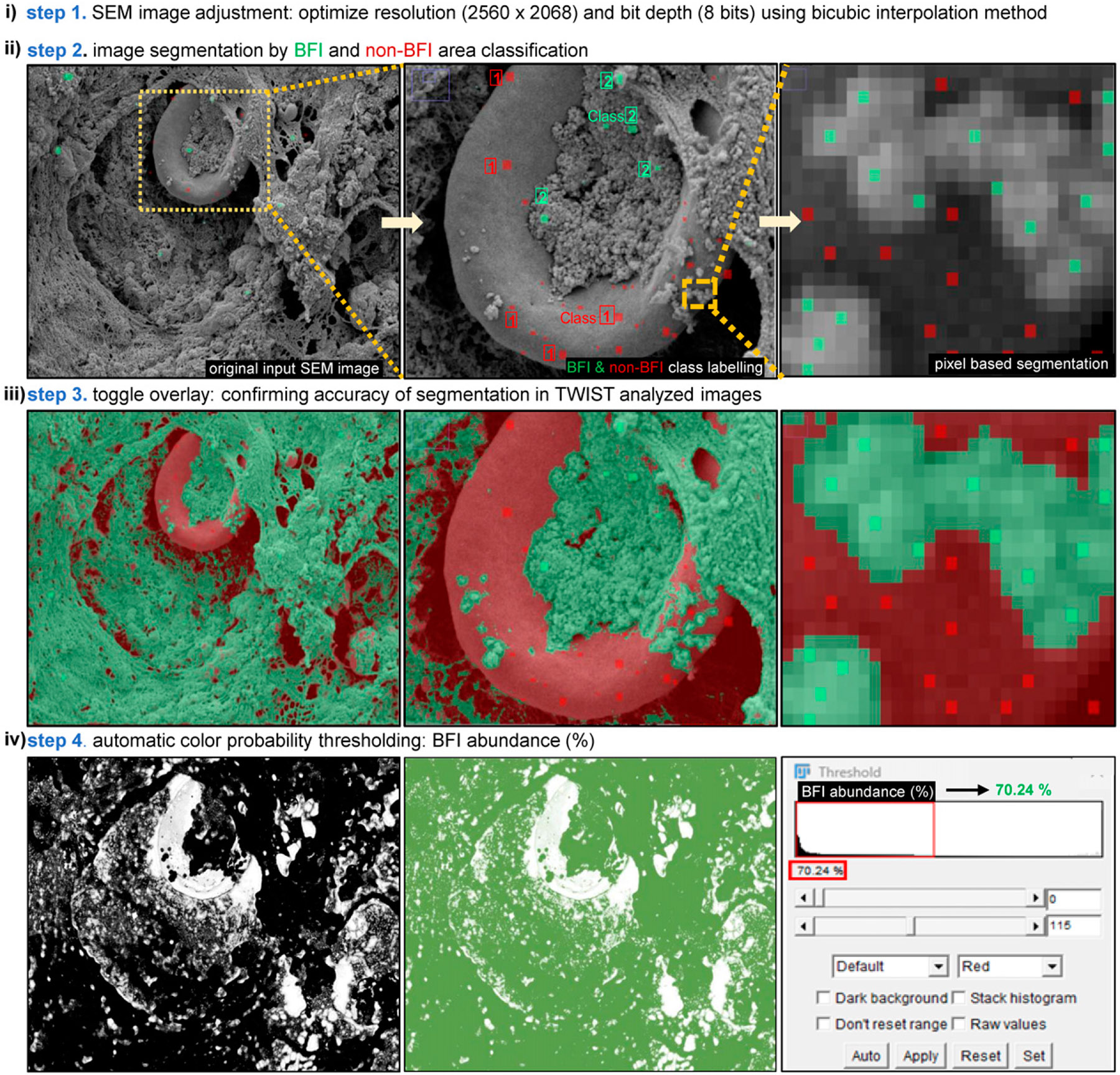
Stepwise implementation of biofilm infection (BFI) detection and quantification using trainable Weka intelligent segmentation technology (TWIST), an ML-based approach. A stepwise procedure of BFI quantitative assessment using TWIST (ML) approach, including (i) adjustment and optimizing of the resolution of the raw image obtained using SEM; (ii) input of the raw image to TWIST for segmentation of BFI (green) and non-BFI elements (red), this process is completed in high magnification of the parent raw image for clear visualization; (iii) toggle overlay allows to visualize and confirm the BFI-positive area is adequately marked; (iv) probability thresholding step provides a map of BFI-infected area (black color pixels) and a quantitative% of BFI abundance.
Analysis of image sensitivity in SEMTWIST segmentation
High-resolution SEM images are essential for improving image segmentation accuracy.51,52 To optimize the resolution of SEM images for input to TWIST analysis, we performed sensitivity analyses (Fig. 4 and Supplementary Fig. S3). Representative raw SEM images and their corresponding TWIST-segmented images at resolutions of 320 × 256, 960 × 768, and 2,560 × 2,048 pixels showed BFI abundances of 42.6%, 26.4%, and 25.8%, respectively. These results demonstrate how changes in pixel density can impact segmentation accuracy and BFI detection within each image, thus influencing quantitative outcomes in BFI analysis (Fig. 4A). In each figure, TWIST was trained on the same instance locations with only variation in resolution to obtain classification results. TWIST performance sensitivity was assessed under conditions of different SEM image resolution. In general, the performance of the ML algorithm decreased at lower pixel densities (Fig. 4B). For high-resolution SEM images (pixel density of 2,560 × 2,048), TWISTbased detection has high accuracy regardless of the extent of biofilm abundance in the image. In cases of low-resolution SEM images, TWIST-based detection was less susceptible to errors when biofilm abundance in the image is high, that is, when >75% of the SEM image is covered by biofilm structures (image 2, Supplementary Fig. S3). However, under conditions of low-resolution SEM images and the rare (<50%) presence of biofilm, TWIST-based detection may not be reliable (Fig. 4C) (image 1, Supplementary Fig. S3). Upscaling images to 2,560 × 2,048 will improve the diagnostic accuracy when comparing images of different pixel densities, particularly when the training data lack sufficient spatial resolution.51,52
Figure 4.
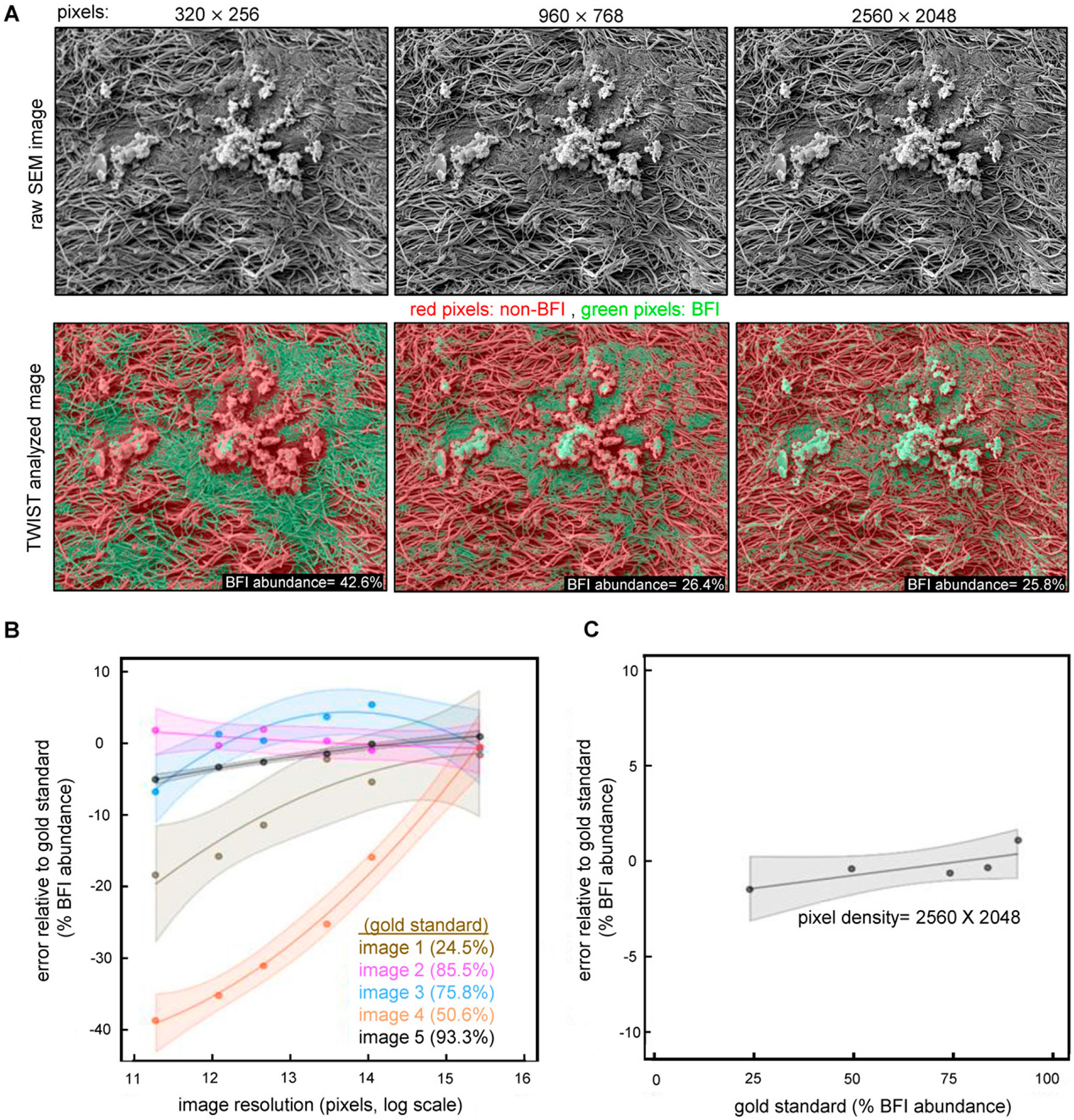
The image pixel density is critical for SEM image biofilm quantification using TWIST. (A) the SEM images were downscaled or upscaled to 320 × 256, 960 × 768, and 2,560 × 2,048 pixels while maintaining the original aspect ratio; (B, C) sensitivity analysis for TWIST performance on five randomly selected SEM images (Supplementary Fig. S3) was performed. The greatest errors occur at lower pixel densities at about 50% (BFIint) biofilm density, (C) images with higher biofilm densities (i.e., >75%, BFIhi) have little or no susceptibility to errors due to pixel density.
Training SEMTWIST for specificity
For efficient performance of the TWIST, it is critical to train the software to distinguish between BFI and non-BFI structures in wound-edge tissue by specifically distinguishing host tissue structures. To conduct this, the images were classified into two categories: BFI and non-BFI (tissue features), resulting in binary pixel classification. The constituents of wound-edge tissue, including red blood cells (RBC), white blood cells (WBC), fibrin, activated platelets, and extracellular matrix (ECM), were distinguishable alongside the biofilm (Fig. 5A, B). Given that the biofilm may be embedded within, or interspersed between these structures, accurate segmenting for these structures is necessary. To achieve this, the training of TWIST is crucial to reliably identify and distinguish these tissue elements from the BFI40. Pixel-level training was conducted to ensure precise differentiation between BFI and non-BFI regions, thereby achieving high accuracy in region selection and classification. An example of the accuracy of the segmentation of tissue RBC structure and its discrimination from BFI is illustrated in Fig. 3. Given that the images are derived from chronic human wound-edge tissue samples, notable morphological alterations in RBC shape and ECM topography were observed, as depicted in Fig. 5A (Supplementary Fig. S4). RBCs typically appear as stomatocytes (concave-shaped) but may transform into echinocytes (shell-like with pointed projections), as observed in multiple patient samples.53,54 The ECM architecture is highly intricate and can sometimes be misinterpreted as biofilm due to protuberances or abnormal topographical modifications on its membrane that occur during ECM remodeling.55,56 Furthermore, other features observed included activated platelets that appeared as elongated spheres with spicules, neutrophils with a ruffled morphology, and monocytes with pseudopods such as protrusions. In addition, fibrins were identified as thread-like structures. Recognizing these morphological variations is critical for training the software to accurately distinguish biofilm dispersed within these tissue components in an SEM image.
Figure 5.
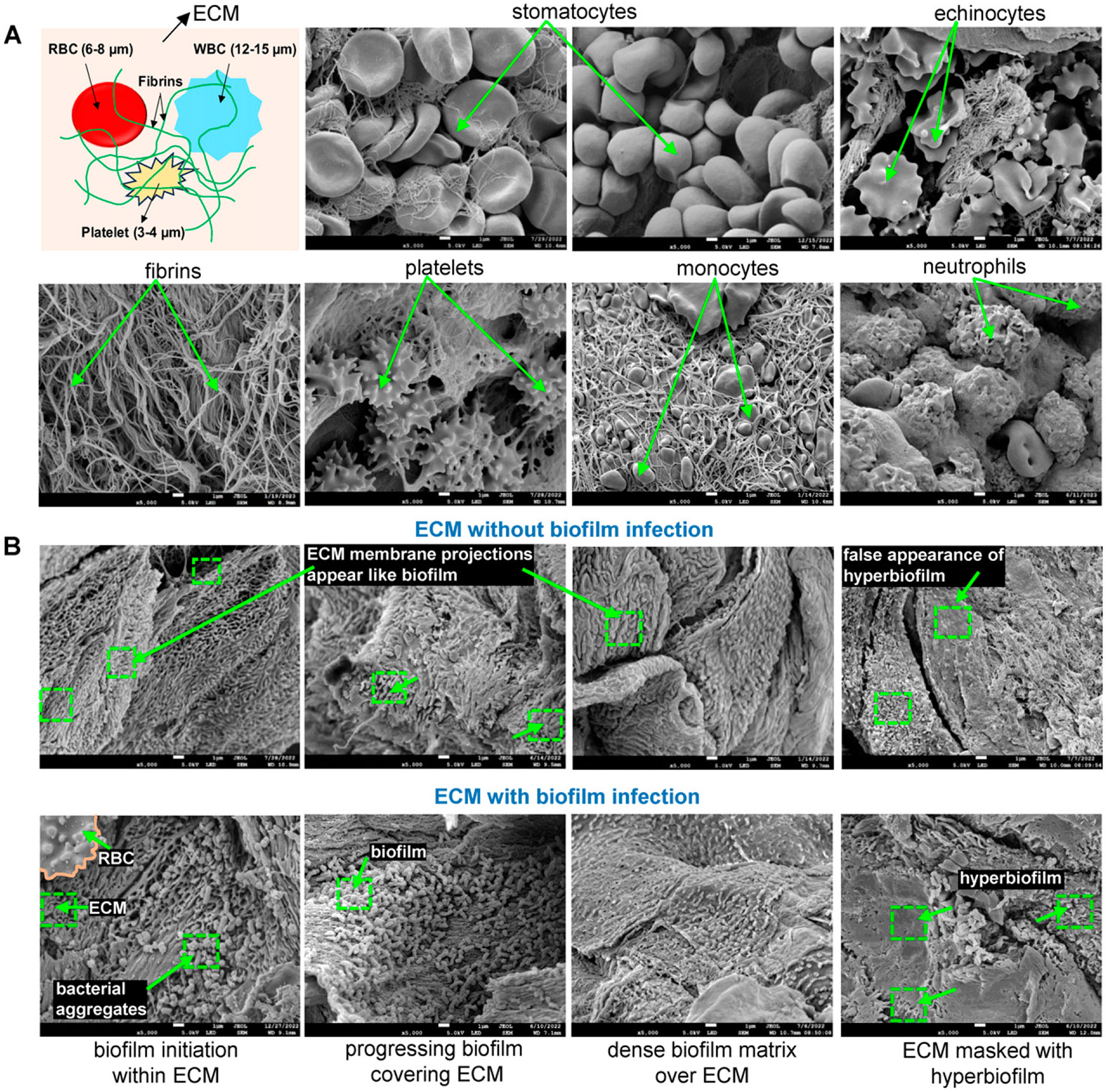

Training the TWIST to recognize the complex and diverse structure of host tissue in a chronic wound environment. (A) Representative SEM images serving as training set for TWIST to discern some common features in a complex chronic wound-edge tissue depicting noninfected red blood cell (RBC); stomatocytes and echinocytes were observed, fibrin network is dense and classified based on structure (thread-like), blood platelet features, and when activated in a wound environment changes its morphology, which helps in their classification and segmentation, white blood cell (WBC) such as monocytes and neutrophils are enormous in size, mostly embedded within fibrins and extracellular matrix (ECM). Monocytes are observed with pseudo-pod-like projections and blebs, whereas neutrophils have a ruffled morphology. (B) Representative additional images related to ECM used for training the TWIST: ECM images without biofilm infection (top panel) or with biofilm infection (bottom panel). Scale bar: 5,000×.
Validation using traditional biofilm methodologies
Microbial biofilm research has relied on CFU counts, culture-based assays, and biofilm staining as standard approaches for quantifying bacterial biofilm in both in vitro and clinical contexts.16,57 In a cohort of 15 patient samples analyzed in this study, eight showed negative results for the presence of any bacterial species in culture reports. Notably, five of these eight culture-negative biospecimens displayed dense biofilm structures upon SEM examination (Fig. 6A). PNA-FISH specifically identifies bacterial species and displays aggregation characteristics thus helping identify and characterize BFI.44 The biospecimens with negative culture reports showed the presence of polymicrobial BFI-like structures when examined using PNA-FISH (Fig. 6A), thus validating SEMTWIST findings and indicating that CFU-based culture pathology may not be reliable for the detection of BFI in woundedge tissues. We tested a widely used staining WGA-Alexa Fluor 488 conjugate for the detection of N-acetylglucosamine in biofilms42,43,58-60 and compared it with SEMTWIST BFI assessment (Fig. 6B). WGA staining was not necessarily specific for biofilm in human chronic wound-edge samples (Supplementary Fig. S5). BFI quantification using WGA or TWIST showed a weak correlation (r = 0.24).
Figure 6.
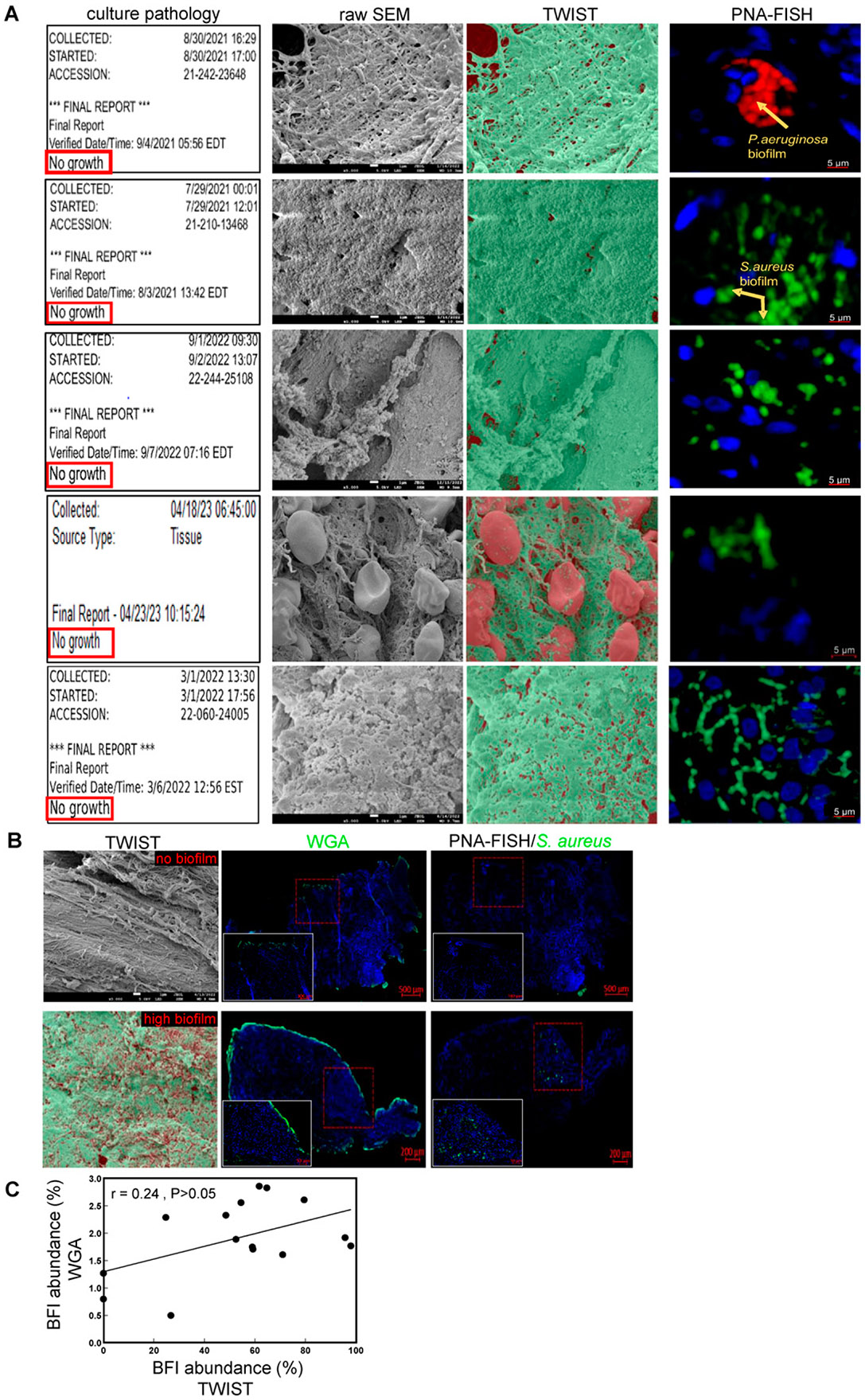
Standard wound culture-based analysis is not reliable to detect fastidious and slow-growing biofilm bacteria in human chronic wounds. (A) Clinical laboratory-based culture pathology reports from four human wound-edge tissue biospecimens out of the cohort of n = 15 showing no microorganism growth indicating negative culture reports. The SEM images with TWIST analysis of the same biospecimen revealed the presence of BFI in these samples. PNA-FISH confirmed the presence of Pseudomonas aeruginosa biofilm (stained red) and Staphylococcus aureus biofilm (stained green) in these samples, scale bar = 5 μm; (B) comparative analysis of wheat germ agglutinin (WGA) BFI staining with SEMTWIST or PNA-FISH assays indicating potential nonspecific detection of BFI with WGA in chronic wound tissues. Top panel: a representative wound biospecimen with no BFI as identified using SEMTWIST; bottom panel: a representative wound biospecimen with BFIhi as identified using SEMTWIST; (C) regression analysis indicating a weak correlation (r = 0.24, p > 0.05) between the WGA-generated and SEMTWIST-generated BFI abundance data.
DISCUSSION
Conventional SEM and FESEM are the two rigorous methods for high magnification and high-resolution visualization of biofilm morphology.21 Reading such images requires trained human experts limiting the significance of these analytical approaches as it relates to clinical research use. In recognition of the complex nature of tissue structures in the chronic wound SEM images, we used TWIST, an advanced segmentation, semisupervised ML technique.40 Deep-learning procedures based on convolutional neural networks have already been utilized to automate the SEM image analysis. However, such solutions have been practiced mostly in the context of material sciences.61-64 A few studies have utilized TWIST-based biofilm segmentation using SEM images but primarily for the abiotic surfaces.38,65 Chronic wound-edge SEM images are complex and consist of diverse features.53-56 This work represents a maiden effort seeking in-depth characterization using the SEM-ML approach to identify typical structures of BFI-infected or noninfected features in chronic wound SEM images. Tissue samples were sourced from chronic wound environments (Table 1), where various morphological modifications can be observed in micrographs captured using SEM. The training data construction for ML models accounted for how variability affects model performance.66 This study included a diverse set of tissue samples with different biofilm types and varying levels of complexity. Such an approach helped develop a robust model with generalizable application.
Human assessments are subject to bias and may overlook certain data types that otherwise can be effectively identified by ML algorithms, especially when performing pixel-level analysis. SEM images often contain noise, artifacts, or irregularities in tissue architecture that can obscure biofilm structures. Evaluators might interpret these visual distortions differently, leading to variability in biofilm quantification. For example, membrane projections or ECM features can be mistaken for biofilm, especially in samples with low biofilm abundance.55,56 Therefore, while human scores are valuable as a gold standard, it is essential to account for potential biases and discrepancies that may arise from human evaluations.67-69 By integrating software capabilities with human assessments and establishing rigorous gradation criteria, the BFI-assessing TWIST-model sets a new benchmark for biofilm quantification. To further strengthen our findings on the concordance between human evaluations and SEM data, this work relied on the highly specific PNA-FISH molecular technique widely recognized for its application to BFI detection.43 PNA-FISH specifically detected S. aureus and P. aeruginosa involvement in wound BFI.
WGA, a widely used lectin-based stain, is often used for the detection of EPS-encasing biofilm aggregates.42,43,70 It is important to recognize that WGA has a high affinity for N-acetylglucosamine residues and glycoproteins, which are present in abundance in both bacterial biofilm EPS and in human wound tissue.70 It is reported that WGA can bind ECM and epidermal cell membranes in tissue sections.71,72
Fluorescently tagged WGA lectin binds to connective tissues, including muscle, bone, ligament, and tendon.73,74 The specificity of WGA to specifically detect biofilm has therefore been questioned.75 While WGA may be useful to detect biofilm on abiotic structures, in a tissue environment it can crossreact with tissue components. In contrast, PNA probes using FISH have been designed and optimized for the detection of several bacteria. When PNA probes are attached to a fluorochrome dye, they can be detected by epifluorescence microscopy or flow cytometry using the FISH method. PNA-FISH has been established as a productive approach to detect and characterize wound biofilm in human tissues.76-78
The culture-based technique was used to assess biofilm presence; however, the primary issue in accurately determining biofilm presence using the culture technique can be attributed to the inherent complexity and heterogeneity of biofilm structures.79 Bacteria within biofilm aggregates are encased in a dense EPS, which poses challenges for effective bacterial separation and quantification. Even with mechanical disruption techniques such as vortexing or ultrasonication, biofilm aggregates often remain intact, leading to skewed results where clumps of bacteria may get erroneously counted as single colonies. Moreover, biofilm bacteria are adapted to anoxic conditions within the EPS matrix.12 When exposed to aerobic environments on nutrient-rich agar plates, these bacteria may undergo a rapid transition from anaerobic to aerobic conditions.12,80 This abrupt environmental shift can lead to bacterial stress, reduced viability, or even cell death.80 In some cases, bacteria may be viable but being fastidious strains, they are nonculturable under conventional laboratory conditions. Consequently, culture results, which typically require 1–3 days to obtain, can lead to an underestimation of biofilm abundance and inaccurate data interpretation, making this method unsuitable for early and precise diagnosis of biofilm presence in the clinical samples.
This work assigns importance to SEM image resolution as it relates to reliable TWIST-based detection of BFI. In this context, it is important to recognize that best results may be obtained from high-resolution images only when an appropriate classification model and interpolation methods are used.49 In certain medical applications, such as pathological imaging and digital mammography, high-resolution images often need to be downsampled due to the limitations of storage capacity. Therefore, selecting the appropriate image resolution for TWIST involves balancing the desired analysis with the available computational resources, as higher resolutions require significant processing time and power.81 Our investigation revealed that standardizing image resolution to 2,560 × 2,048 provided optimal results in the given context.
Generalizability
As demonstrated in the current study, where TWIST was trained to differentiate between biofilm and nonbiofilm features in wound SEM images, the same approach can be applied to specialized datasets for studies focused on other tissues such as bone and muscle. As an example for infected bone tissue sample, the TWIST ML algorithm needs to be trained for distinguishing specific features in the SEM images from bone tissue. The model must be trained to accurately identify key structural components, including the layered bone matrix (lamellae), honeycomb-like network (cancellous bone), osteoblasts, osteoclasts, osteocytes, mineralized bone matrix, and canaliculi.82 By applying the training strategy similar to the approach used that was used for classifying RBCs, WBCs, fibrin, and platelets in wound tissue SEM images, the infected bone model will be able to differentiate distinct features of that tissue, ensuring precise classification and segmentation of bone microstructures from BFI.
CONCLUSION
Complementing these findings, this work provided an ML-based quantitative tool for the determination of the extent of BFI burden in chronic wound-edge tissues leveraging the optimized SEM image resolution and segmentation technique that can deliver reliable quantitative insights into biofilm prevalence within intricate tissue environments.
Supplementary Material
Supplementary Figure S1
Supplementary Figure S2
Supplementary Figure S3
Supplementary Figure S4
Supplementary Figure S5
Supplementary Table S1
KEY FINDINGS.
Scanning electron microscopy-based Trainable Weka (Waikato Environment for Knowledge Analysis) Intelligent Segmentation Technology (SEMTWIST) provides precise biofilm infection (BFI) quantification wound tissue matrices, minimizing interrater variability inherent in manual assessments.
SEMTWIST outperformed traditional methods such as bacterial culture and wheat germ agglutinin staining by reliably detecting biofilm.
SEMTWIST enables BFI abundance grading, aiding clinicians in making informed decisions for effective BFI treatment strategies.
ACKNOWLEDGMENTS AND FUNDING SOURCES
This work was funded by the Naval Medical Research Command (NMRC)—Naval Advanced Medical Development (NAMD) under MTEC OTA project W81XWH-21-9-0020 to S.R., NIDDK 7R01DK125835-05 and R01DK125835 to C.K.S., and the NIH K25 grant GM143572 to Y.X.
Abbreviations And Acronyms
- AI
artificial intelligence
- CFU
colony forming unit
- ECM
extracellular matrix
- EPS
extracellular polymeric substance
- FESEM
field emission scanning electron microscopy
- FFPE
formalin-fixed paraffin embedded
- HMDS
hexamethyldisilazane
- LoA
limit of agreement
- ML
machine learning
- P. aeruginosa
Pseudomonas aeruginosa
- PNA-FISH
peptide nucleic acid fluorescence in situ hybridization
- RBC
red blood cell
- S. aureus
Staphylococcus aureus
- SEM
scanning electron microscopy
- WBC
white blood cell
- WGA
wheat germ agglutinin
Biographies
Surabhi Singh, PhD, is a Postdoctoral Associate at the McGowan Institute for Regenerative Medicine (MIRM), Department of Surgery at University of Pittsburgh Medical Center. Her research focuses on BFI in chronic wounds and novel approaches to quantify BIFs in chronic wounds.
Fabio Muniz De Oliveira, PhD, is a Postdoctoral Associate at MIRM, his research interests include understanding molecular mechanisms of host and biofilm interactions in chronic wounds.
Cong Wang, PhD, is a Postdoctoral Associate at MIRM with a keen interest in nanomaterials, nanofabrication, and developing AI algorithms.
Manoj Kumar, PhD, is a Postdoctoral Associate at MIRM with expertise in statistical analysis to predict the timing of disease progression.
Yi Xuan, PhD, is a Professor of Surgery with research interests in next-generation tissue nanotransfection devices, nanochips for exosome detection, and novel biochips for wound healing applications.
Deeptanker DeMazumder, MD, PhD, is an Associate Professor of Surgery and Internal Medicine (cardiology) at MIRM. His research employs novel artificial intelligence and machine learning (AI/ML) strategies in parallel studies of animal models and humans, including fundamental new AI/ML algorithms for personalized diagnosis and treatment of critical illness.
Chandan K. Sen, PhD, is the Director of MIRM. He is a world-renowned regenerative medicine expert and pioneer of novel wound care technologies.

Sashwati Roy, PhD, is a Professor of Surgery at MIRM. Her research interest focus is biofilm infection, inflammation, and macrophage biology in chronic wounds, more specifically diabetic ulcers.
Footnotes
AUTHOR DISCLOSURE AND GHOST WRITING
The authors declare no financial interests or conflicts of interest to disclose.
DISCLAIMER
Views expressed are those of the authors and do not reflect the official policy or position of the Department of Defense/U.S. Government. Identification of specific products or scientific instrumentation does not constitute an endorsement or implied endorsement on the part of the authors, Department of Defense, or any component agency.
REFERENCES
- 1.Sen CK, Roy S, Mathew-Steiner SS, et al. Biofilm management in wound care. Plast Reconstr Surg 2021;148(2):275e–288e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Versey Z, da Cruz Nizer WS, Russell E, et al. Biofilm-innate immune interface: Contribution to chronic wound formation. Front Immunol 2021;12:648554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Cavallo I, Sivori F, Mastrofrancesco A, et al. Bacterial biofilm in chronic wounds and possible therapeutic approaches. Biology (Basel) 2024;13(2):109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bjarnsholt T. The role of bacterial biofilms in chronic infections. APMIS 2013;121(136):1–51. [DOI] [PubMed] [Google Scholar]
- 5.Grooters KE, Ku JC, Richter DM, et al. Strategies for combating antibiotic resistance in bacterial biofilms. Front Cell Infect Microbiol 2024;14:1352273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Mah T-F. Biofilm-specific antibiotic resistance. Future Microbiol 2012;7(9):1061–1072. [DOI] [PubMed] [Google Scholar]
- 7.Khan J, Tarar SM, Gul I, et al. Challenges of antibiotic resistance biofilms and potential combating strategies: A review. 3 Biotech 2021;11(4):169–115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Liu Y, Long S, Wang H, et al. Biofilm therapy for chronic wounds. Int Wound J 2024;21(2):e14667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Falcone M, De Angelis B, Pea F, et al. Challenges in the management of chronic wound infections. J Glob Antimicrob Resist 2021;26:140–147. [DOI] [PubMed] [Google Scholar]
- 10.Sen CK. Human wound and its burden: Updated 2022 compendium of estimates. Adv Wound Care (New Rochelle) 2023;12(12):657–670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Xu Y, Dhaouadi Y, Stoodley P, et al. Sensing the unreachable: Challenges and opportunities in biofilm detection. Curr Opin Biotechnol 2020;64:79–84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Magana M, Sereti C, Ioannidis A, et al. Options and limitations in clinical investigation of bacterial biofilms. Clin Microbiol Rev 2018;31(3); doi: 10.1128/cmr00084-00016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Goswami AG, Basu S, Banerjee T, et al. Biofilm and wound healing: From bench to bedside. Eur J Med Res 2023;28(1):157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Cont A, Rossy T, Al-Mayyah Z, et al. Biofilms deform soft surfaces and disrupt epithelia. Elife 2020;9:e56533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Evelhoch SR. Biofilm and chronic nonhealing wound infections. Surg Clin North Am 2020;100(4):727–732. [DOI] [PubMed] [Google Scholar]
- 16.Wilson C, Lukowicz R, Merchant S, et al. Quantitative and qualitative assessment methods for biofilm growth: A mini-review. Res Rev J Eng Technol 2017;6(4). [PMC free article] [PubMed] [Google Scholar]
- 17.Donlan RM. Biofilms: Microbial life on surfaces. Emerg Infect Dis 2002;8(9):881–890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kulayta K, Zerdo Z, Seid M, et al. Biofilm formation and antibiogram profile of bacteria from infected wounds in a general hospital in southern Ethiopia. Sci Rep 2024;14(1):26359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Stewart PS, Franklin MJ. Physiological heterogeneity in biofilms. Nat Rev Microbiol 2008;6(3):199–210. [DOI] [PubMed] [Google Scholar]
- 20.Bossù M, Selan L, Artini M, et al. Characterization of Scardovia wiggsiae biofilm by original scanning electron microscopy protocol. Microorganisms 2020;8(6):807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Relucenti M, Familiari G, Donfrancesco O, et al. Microscopy methods for biofilm imaging: Focus on SEM and VP-SEM pros and cons. Biology (Basel) 2021;10(1):51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Golding CG, Lamboo LL, Beniac DR, et al. The scanning electron microscope in microbiology and diagnosis of infectious disease. Sci Rep 2016; 6(1):26516–26518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Schultz G, Bjarnsholt T, James GA, et al. ; Global Wound Biofilm Expert Panel. Consensus guidelines for the identification and treatment of biofilms in chronic nonhealing wounds. Wound Repair Regen 2017;25(5):744–757. [DOI] [PubMed] [Google Scholar]
- 24.Høiby N, Bjarnsholt T, Moser C, et al. ; ESCMID Study Group for Biofilms and Consulting External Expert Werner Zimmerli. ESCMID guideline for the diagnosis and treatment of biofilm infections 2014. Clinical Microbiology and Infection 2015;21(Suppl 1):S1–S25. [DOI] [PubMed] [Google Scholar]
- 25.Haddad G, Bellali S, Takakura T, et al. Scanning electron microscope: A new potential tool to replace gram staining for microbe identification in blood cultures. Microorganisms 2021;9(6):1170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Chan RK, Nuutila K, Mathew-Steiner SS, et al. A prospective, randomized, controlled study to evaluate the effectiveness of a fabric-based wireless electroceutical dressing compared to standard-of-care treatment against acute trauma and burn wound biofilm infection. Adv Wound Care (New Rochelle) 2024;13(1):1–13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Caccese JB, Kaminski TW. Comparing computer-derived and human-observed scores for the balance error scoring system. J Sport Rehabil 2016;25(2):133–136. [DOI] [PubMed] [Google Scholar]
- 28.Oka R, Kusumi T, Utsumi A. Performance evaluation of automated scoring for the descriptive similarity response task. Sci Rep 2024;14(1):6228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Makino T, Jastrzębski S, Oleszkiewicz W, et al. Differences between human and machine perception in medical diagnosis. Sci Rep 2022;12(1):6877. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Bryan RN, Davatzikos C, Herskovits EH, et al. Medical image analysis: Human and machine. Acad Radiol 2020;27(1):76–81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Zhang J, Wang Y, Donarski ED, et al. BCM3D 2.0: Accurate segmentation of single bacterial cells in dense biofilms using computationally generated intermediate image representations. NPJ Biofilms Microbiomes 2022;8(1):99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Jelli E, Ohmura T, Netter N, et al. Single-cell segmentation in bacterial biofilms with an optimized deep learning method enables tracking of cell lineages and measurements of growth rates. Mol Microbiol 2023;119(6):659–676. [DOI] [PubMed] [Google Scholar]
- 33.Mountcastle SE, Vyas N, Villapun VM, et al. Biofilm viability checker: An open-source tool for automated biofilm viability analysis from confocal microscopy images. NPJ Biofilms Microbiomes May 14 2021;7(1):44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Arzt M, Deschamps J, Schmied C, et al. LABKIT: Labeling and segmentation toolkit for big image data. Front Comput Sci 2022;4:777728. [Google Scholar]
- 35.Deter HS, Dies M, Cameron CC, et al. A cell segmentation/tracking tool based on machine learning. Computer Optimized Microscopy: Methods and Protocols 2019:399–422. [DOI] [PubMed] [Google Scholar]
- 36.Mbiki S, McClendon J, Alexander-Bryant A, et al. Classifying changes in LN-18 glial cell morphology: A supervised machine learning approach to analyzing cell microscopy data via FIJI and WEKA. Med Biol Eng Comput 2020;58(7):1419–1430. [DOI] [PubMed] [Google Scholar]
- 37.Šimunić I, Jagčcić D, Isaković J, et al. Lusca: FIJI (ImageJ) based tool for automated morphological analysis of cellular and subcellular structures. Sci Rep 2024;14(1):7383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Vyas N, Sammons R, Addison O, et al. A quantitative method to measure biofilm removal efficiency from complex biomaterial surfaces using SEM and image analysis. Sci Rep 2016;6(1):32694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kataras TJ, Jang TJ, Koury J, et al. ACCT is a fast and accessible automatic cell counting tool using machine learning for 2D image segmentation. Sci Rep 2023;13(1):8213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Arganda-Carreras I, Kaynig V, Rueden C, et al. Trainable Weka Segmentation: A machine learning tool for microscopy pixel classification. Bioinformatics 2017;33(15):2424–2426. [DOI] [PubMed] [Google Scholar]
- 41.Kanuri N, Abdelkarim AZ, Rathore SA. Trainable WEKA (Waikato Environment for Knowledge Analysis) segmentation tool: Machine-learning-enabled segmentation on features of panoramic radiographs. Cureus 2022;14(1):e21777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Khan I, Minto RE, Kelley-Patteson C, et al. Biofilm-derived oxylipin 10-HOME–mediated immune response in women with breast implants. J Clin Invest 2023;134(3). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Das Ghatak P, Mathew-Steiner SS, Pandey P, et al. A surfactant polymer dressing potentiates antimicrobial efficacy in biofilm disruption. Sci Rep 2018;8(1):873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Barbosa A, Miranda S, Azevedo NF, et al. Imaging biofilms using fluorescence in situ hybridization: Seeing is believing. Front Cell Infect Microbiol 2023;13:1195803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.de Oliveira A, Pinheiro-Hubinger L, Pereira VC, et al. Staphylococcal biofilm on the surface of catheters: Electron microscopy evaluation of the inhibition of biofilm growth by RNAIII inhibiting peptide. Antibiotics 2021;10(7):879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Silva N, Marques L, Röder D. Diagnosis of biofilm infections: Current methods used, challenges and perspectives for the future. J Appl Microbiol 2021;131(5):2148–2160. [DOI] [PubMed] [Google Scholar]
- 47.Achinas S, Yska SK, Charalampogiannis N, et al. A technological understanding of biofilm detection techniques: A review. Materials 2020;13(14):3147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Flemming H-C, Wingender J. The biofilm matrix. Nat Rev Microbiol 2010;8(9):623–633. [DOI] [PubMed] [Google Scholar]
- 49.Keys R. Cubic convolution interpolation for digital image processing. IEEE Trans Acoust, Speech, Signal Process 1981;29(6):1153–1160. [Google Scholar]
- 50.Giavarina D. Understanding bland altman analysis. Biochem Med (Zagreb) 2015;25(2):141–151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Hirahara D, Takaya E, Kadowaki M, et al. Effect of the pixel interpolation method for downsampling medical images on deep learning accuracy. JCC 2021;09(11):150–156. [Google Scholar]
- 52.Hirahara D, Takaya E, Takahara T, et al. Effects of data count and image scaling on deep learning training. PeerJ Comput Sci 2020;6:e312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Silva AF, Sousa JS, Cunha PL, et al. Erythrocytes morphology and hemorheology in severe bacterial infection. Mem Inst Oswaldo Cruz 2019;114:e190326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Nicolescu CM, Moldovan SD. The influence of sepsis on erythrocytes morphology: Case report and literature review. Rom J Morphol Embryol 2024;65(1):119–123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Pfisterer K, Shaw LE, Symmank D, et al. The extracellular matrix in skin inflammation and infection. Front Cell Dev Biol 2021;9:682414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Castro-Manrreza M, Bonifaz L, Castro-Escamilla O, et al. Mesenchymal stromal cells from the epidermis and dermis of psoriasis patients: Morphology, immunophenotype, differentiation patterns, and regulation of T cell proliferation. Stem Cells Int 2019;2019(1):4541797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Thieme L, Hartung A, Tramm K, et al. Adaptation of the start-growth-time method for high-throughput biofilm quantification. Front Microbiol 2021;12:631248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Ahmad I, Nadeem A, Mushtaq F, et al. Csu pili dependent biofilm formation and virulence of Aci-netobacter baumannii. NPJ Biofilms Microbiomes 2023;9(1):101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Sinha M, Ghosh N, Wijesinghe DS, et al. Pseudomonas aeruginosa theft biofilm require host lipids of cutaneous wound. Ann Surg 2023;277(3):e634–e647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Skogman ME, Vuorela PM, Fallarero A. Combining biofilm matrix measurements with biomass and viability assays in susceptibility assessments of antimicrobials against Staphylococcus aureus biofilms. J Antibiot (Tokyo) 2012;65(9):453–459. [DOI] [PubMed] [Google Scholar]
- 61.Abeyrathna D, Ashaduzzaman M, Malshe M, et al. An ai-based approach for detecting cells and microbial byproducts in low volume scanning electron microscope images of biofilms. Front Microbiol 2022;13:996400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Bals J, Epple M. Deep learning for automated size and shape analysis of nanoparticles in scanning electron microscopy. RSC Adv 2023;13(5):2795–2802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Shah A, Schiller JA, Ramos I, et al. Automated image segmentation of scanning electron microscopy images of graphene using U-Net Neural Network. Materials Today Communications 2023;35:106127. [Google Scholar]
- 64.Iwata H, Hayashi Y, Hasegawa A, et al. Classification of scanning electron microscope images of pharmaceutical excipients using deep convolutional neural networks with transfer learning. Int J Pharm X 2022;4:100135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Ding H, Yang Y, Li X, et al. A simple AI-enabled method for quantifying bacterial adhesion on dental materials. Biomater Investig Dent 2022;9(1):75–83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Weidner L, Walton G. The influence of training data variability on a supervised machine learning classifier for Structure from Motion (SfM) point clouds of rock slopes. Engineering Geology 2021;294:106344. [Google Scholar]
- 67.Hagiwara A, Fujita S, Ohno Y, et al. Variability and standardization of quantitative imaging: Monoparametric to multiparametric quantification, radiomics, and artificial intelligence. Invest Radiol 2020;55(9):601–616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Kang C, Lee C, Song H, et al. Variability matters: Evaluating inter-rater variability in histopathology for robust cell detection. Paper presented at: European Conference on Computer Vision. 2022. [Google Scholar]
- 69.Obuchowicz R, Oszust M, Piorkowski A. Interobserver variability in quality assessment of magnetic resonance images. BMC Med Imaging 2020;20(1):109–110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Ryva B, Zhang K, Asthana A, et al. Wheat germ agglutinin as a potential therapeutic agent for leukemia. Front Oncol 2019;9:100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Clark RK, Galantino-Homer HL. Wheat germ agglutinin as a counterstain for immunofluorescence studies of equine hoof lamellae. Exp Dermatol 2014;23(9):677–678. [DOI] [PubMed] [Google Scholar]
- 72.Desantis S, Corriero A, Acone F, et al. Lectin histochemistry on the dorsal epidermis of the Breton dog. Acta Histochem 2003;105(1):73–79. [DOI] [PubMed] [Google Scholar]
- 73.Kostrominova TY. Application of WGA lectin staining for visualization of the connective tissue in skeletal muscle, bone, and ligament/tendon studies. Microsc Res Tech 2011;74(1):18–22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Emde B, Heinen A, Gödecke A, et al. Wheat germ agglutinin staining as a suitable method for detection and quantification of fibrosis in cardiac tissue after myocardial infarction. Eur J Histochem 2014;58(4). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Johnsen AR, Hausner M, Schnell A, et al. Evaluation of fluorescently labeled lectins for noninvasive localization of extracellular polymeric substances in Sphingomonas biofilms. Appl Environ Microbiol 2000;66(8):3487–3491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Lichtenberg M, Kirketerp-Møller K, Kvich LA, et al. Single cells and bacterial biofilm populations in chronic wound infections. APMIS 2024;132(12):1071–1077. [DOI] [PubMed] [Google Scholar]
- 77.Weaver AJ Jr, Brandenburg KS, Sanjar F, et al. Clinical utility of PNA-FISH for burn wound diagnostics: A noninvasive, culture-independent technique for rapid identification of pathogenic organisms in burn wounds. J Burn Care Res 2019;40(4):464–470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Malic S, Hill KE, Hayes A, et al. Detection and identification of specific bacteria in wound biofilms using peptide nucleic acid fluorescent in situ hybridization (PNA FISH). Microbiology (Reading) 2009;155(Pt 8):2603–2611. [DOI] [PubMed] [Google Scholar]
- 79.Welch K, Cai Y, Strømme M. A method for quantitative determination of biofilm viability. J Funct Biomater 2012;3(2):418–431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Monds RD, O’Toole GA. The developmental model of microbial biofilms: Ten years of a paradigm up for review. Trends Microbiol 2009;17(2):73–87. [DOI] [PubMed] [Google Scholar]
- 81.Rathee N, Pahal S, Sheoran D. Evaluating the uncertainty of classification due to image resizing techniques for satellite image classification. Mapan 2021;36(2):243–251. [Google Scholar]
- 82.Shah FA, Ruscsák K, Palmquist A. 50 years of scanning electron microscopy of bone—a comprehensive overview of the important discoveries made and insights gained into bone material properties in health, disease, and taphonomy. Bone Res 2019;7(1):15. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


