Abstract
Esophageal squamous cell carcinoma (ESCC) is characterized by a complex tumor microenvironment (TME). Cancer-associated fibroblasts (CAFs) play a crucial role in the TME that facilitate tumor progression via interactions with cancer cells. However, the mechanisms underlying the activation of CAFs in TME remain largely unknown. Here, we characterized the exosomes derived from normoxic and hypoxic ESCC cells using electron microscopy and western blot. The impact of exosomes on CAF activation and the motility of ESCC cells was examined in vitro. The molecular complex involving circNRIP1 was explored using RNA pull-down. We demonstrated that exosomes derived from ESCC cells, including KYSE-150 and TE-10 cells, exhibited a significantly increase in secretion under hypoxic conditions. These hypoxic exosomes were internalized by fibroblasts and further promoted the transformation of normal fibroblasts into CAFs, as evidenced by enhanced migration and secretion of pro-inflammatory cytokines. circNRIP1 was enriched in hypoxic exosomes, and its absence abolished the effect of hypoxic exosomes to activate CAFs. Furthermore, the CAFs activated by exosomal circNRIP1 further promoted the migration and invasion of ESCC cells. Mechanistically, circNRIP1 bound to the N1-methyladenosine (m1A) methyltransferase TRMT6 and activated CAFs in a TRMT6-dependent manner. This study revealed the role of hypoxia-induced exosomal circNRIP1 in the activation of CAFs, which contributes to ESCC development. These findings shed light on the mechanisms of the CAF activation in ESCC, positioning hypoxia-induced exosomal circNRIP1 as a potential molecular target for ESCC.
Supplementary Information
The online version contains supplementary material available at 10.1186/s12876-025-03978-w.
Keywords: Esophageal squamous cell carcinoma, Hypoxia, Exosome, Cancer-associated fibroblast, CircNRIP1
Esophageal squamous cell carcinoma (ESCC) is a highly prevalent and lethal malignancy of the digestive system, representing a major global health burden [1, 2]. Chemotherapy combined with immunotherapy can significantly prolong patients’ survival and progression-free survival by activating the immune response to eliminate cancer cells within the tumor microenvironment (TME). Despite the continuous advancements of diagnostic techniques and therapeutic methods, ESCC presents a dismal 5-year survival rate(< 20%) [3]. Therefore, an expanding understanding of the interaction of tumor cells and TME is urgently needed to facilitate the development of personalized treatment strategies and improve the prognosis of ESCC.
Tumor-stroma interactions are a major obstacle in cancer treatment, contributing to tumor development and impede effective cancer management in clinical practice [4, 5]. Within the TME, various stromal cells coexist with tumor cells, primarily cancer-associated fibroblasts (CAFs), endothelial cells, tumor-infiltrating lymphocytes, and myeloid cells [6]. CAFs represent heterogeneous components derived from tissue-resident fibroblasts, stellate cells, mesenchymal stem cells, adipocytes, and other cell types activated by malignant tumor cells [4, 7]. New insights have been shed on the interaction between tumor cells and CAFs, highlighting that tumor-cell-derived CXCL1 activates CAFs via the CXCR2-STAT3 pathway [8]. In addition, hypoxic exosomal miR-5100 promotes the activation of CAFs by orchestrating STAT3 signal transduction, which further facilitates head and neck squamous cell carcinoma metastasis [9]. The role of STAT3 as a key signaling molecule in exosomal circNRIP1-mediated CAF activation requires further validation. In turn, CAFs possess the capability to regulate tumor progression by remodeling the extracellular matrix (ECM) in ESCC [10]. CAFs can block angiogenesis and metastasis in ESCC via secreting protein [11]. CAFs-derived Exosomal miR-889-3p repress M1 macrophage polarization to boost ESCC development [12]. However, no prior studies have established a functional link between miR-889-3p and hypoxic exosomes or circNRIP1, though both have been independently implicated in CAF activation. Future investigations should determine whether hypoxic exosome-derived circNRIP1 modulates CAF activation through miRNA sponging, thereby amplifying oncogenic signaling in the tumor microenvironment. Due to the necessary supportive role in tumor development, targeting CAFs has an attractive prospect in the aspect of improving the efficacy of ESCC therapy [13]. However, the mechanisms underlying the activation of CAFs in the TME of ESCC remain unclear.
Exosomes, the essential components of the TME, exert pleiotropic effects to mediate information exchange between cancer cells and stromal cells by transferring pro-tumorigenic genetics, proteins, and lipids to the recipient cells [14]. The rapid growth of solid tumors creates insufficient blood supply, leading to hypoxia conditions [15], and hypoxic exosomes secreted by tumor cells and taken up by adjacent cells encourage tumorigenesis, angiogenesis, and metastasis [16]. In addition, hypoxia-induced exosome cargo changes also lead to tumor malignancy [17]. However, the mechanism by which hypoxic exosomes mediate stromal cell activation within the ESCC microenvironment remains poorly understood.
Circular RNAs (circRNAs) derived from exosomes are abnormally expressed in various cancer cells and transported to recipient cells, supporting diverse critical cancer-promoting functions [18, 19]. circNRIP1, otherwise known as hsa_circ_0004771, is mapped to chromosome 21 at the locus (16,386,664–16,415,895) and is transcribed from the exon2 and exon3 of NRIP1 gene. circNRIP1 is a newly discovered biomarker for colorectal cancer, closely related to various cancers, including ESCC [20]. circNRIP1 knockdown significantly inhibits the proliferation, migration, and invasion of ESCC cells and promotes cell apoptosis [21]. However, the role of exosomal circNRIP1 from ESCC cells has not been elucidated.
In this study, we investigated the role of hypoxia-induced exosomes in the activation of CAFs in ESCC. We characterized the features of hypoxia-ESCC cell-derived exosomes and explored their impact on the activation of CAFs and the migration and invasion of ESCC cells. Furthermore, we found that circNRIP1 is enriched in hypoxic exosomes and elucidated the potential mechanisms by which exosomes regulate the activation of CAFs through circNRIP1. Our results showed the function and underlying mechanism of hypoxic exosomes in activating CAFs, providing new ideas for the therapy of ESCC.
Method
Cell culture and treatment
The human ESCC cell lines KYSE-150 (iCell, iCell-h245, China) and TE-10 (iCell, iCell-h418, China) were used in this study. These cell lines were cultured in RPMI-1640 medium supplemented with 10% fetal bovine serum (FBS, 10,099–141, Gibco, USA) and 1% penicillin–streptomycin (PS). Human embryonic lung fibroblasts MRC5 cells (Sciencell Company, USA) were cultured in MEM medium supplemented with 10% FBS and 1% PS. A hypoxic environment was established using a three-gas incubator (96% nitrogen, 5% carbon dioxide, and 1% oxygen, SQ-80N, Shanghai mousse Experimental Equipment Co., Ltd). KYSE-150 and TE-10 cell lines were cultured in the tri-gas incubator for 48 h to generate hypoxic exosomes. All cells were routinely authenticated by short tandem repeat (STR) analysis, and regularly treated with a mycoplasma-removing agent.
Plasmid transfection and lentiviral infection
Short hairpin RNA (shRNA) was applied to knockdown circNRIP1 in ESCC cell lines. Three shRNAs targeting the back-splice junction of circNRIP1 (shcircNRIP1-1, shcircNRIP1-2, shcircNRIP1-3) and non-targeting control (shNC) were designed using the pLKO.1-puro system (Sigma-Aldrich). TE-10 cells were infected with shcircNRIP1 or shNC lentivirus (MOI = 50) in RPMI-1640 with 8 μg/mL polybrene (Sigma-Aldrich) following the manufacturer’s instructions. After 48 h, the cells were selected with 2 μg/mL puromycin (Thermo Fisher) for 72 h. The expression of circNRIP1 was confirmed by real-time PCR and normalized to GAPDH.
MRC5-induced fibroblasts were transduced with shTRMT6 or shNC lentivirus (MOI = 10) with 8 μg/mL polybrene followed by selection in 2 μg/mL puromycin for 72 h. The transfection efficiency was confirmed by western blot.
To generate a stable overexpression cell strain, the full-length of TRMT6 was cloned into a lentiviral vector containing a CMV promoter and puromycin resistance gene. MRC5-induced fibroblasts were infected with the lentivirus (MOI = 10) and selected with 2 μg/mL puromycin. The expression of TRMT6 was confirmed by real-time PCR and western blot.
Exosome isolation
Exosomes were separated by ultracentrifugation. RPMI-1640 medium without FBS was applied when the cells reached 80% confluence. After 48 h, the culture medium of hypoxic and normoxic cells was obtained and centrifuged separately at 300 g for 10 min at 4℃. The supernatant collected was centrifuged at 2,000 g for 10 min to remove cellular fragments. The supernatant collected was centrifuged at 10,000 g for 30 min to withdraw large pellets. Finally, the supernatant was centrifuged at 100,000 g for 60 min to collect exosomes. The sediments were washed in PBS buffer and resuspended in 100 μl of PBS buffer. The exosomes were preserved at − 20℃ or − 80℃ for later use.
Transmission electron microscopy
Pipette 5–10 μl of the resuspended exosome solution onto a copper electron grid. Allow the sample to settle for 1 min, then use filter paper to gently blot away the excess liquid from the edge of the grid. Wash with the appropriate amount of PBS. Apply 10 μl of phosphotungstic acid to the copper grid, settle samples for 1 min, and then use filter paper to remove the excess liquid. Samples were air-dried at room temperature for 2 min, then examined using a transmission electron microscope JEM-1200EX at an acceleration voltage of 100 kV.
Western Blot
The total protein of cells was extracted using RIPA reagent (Thermo, USA). Separated protein samples were loaded on 10% SDS-PAGE gel and transferred to PVDF membranes. Membranes were then treated with primary antibodies against ALIX (Proteintech, 12,422–1-AP, USA), TSG101 (Abcam, ab125011, USA), CD81 (Proteintech, 27,855–1-AP, USA), α-SMA (Proteintech, 14,395–1-AP, USA), COL1A1 (Proteintech,14,695–1-AP, USA), COL3A1 (Proteintech, 21,898–1-AP, USA), TRMT6 (Proteintech,16,727–1-AP,1:6000), GAPDH (Proteintech, 60,004–1-Ig, USA) at 4℃ overnight. PVDF membranes were washed with tris-buffered saline with 0.1% Tween 20 detergent (TBST) and then incubated with Goat Anti-Rabbit IgG H&L (Beyotime, A0208, USA) or Goat Anti-Mouse IgG H&L (Beyotime, A0216, USA) for 1 ~ 2 h at room temperature. The signal of protein bands was detected with enhanced chemiluminescence reagent (Thermo Fisher Scientific, USA) and photographed using a Chemi Doc MP system (Bio-Rad, Hercules, USA). The optical density was analyzed using Image J software.
Exosome tracing experiment
Fluorescence imaging was used to detect exosomes internalized by fibroblasts. Harvested exosomes were incubated with the fluorescent lipophilic dye Dil (Beyotime, C1036, China) for 2 h. Fluorescent-labeled exosomes were then added to fibroblast cultures. Subsequently, these fibroblasts in coverslips were fixed with 4% paraformaldehyde at 4℃ for 15 min. After washing with PBS three times, coverslips were permeabilized with 0.2% Triton X-100 in PBS for 5 min at room temperature and then blocked with 1% BSA for 1 h. Subsequently, cells were mounted with 90% glycerol containing DAPI (Invitrogen, USA) for 10 min. The cells were examined by fluorescence microscopy (Nikon, USA).
Transwell assay
The ability of cell migration and invasion was determined by using a transwell assay. In the migration assay, 1.5 × 10^5 fibroblasts or ESCC cells suspended in 500 μl serum-free RPMI-1640 were seeded in the upper chamber of the 0.8 μm Transwell insert (353,097, FALCON, USA) and 700 μl RPMI-1640 with 20% FBS were added to the lower compartment of the chamber. After being cultured for 96 h at 37 °C, the upper chamber was fixed by 4% paraformaldehyde at room temperature for 30 min and stained with crystal violet at room temperature for 30 min. Then, the slides were mounted on a slide and sealed with neutral balsam. Three random fields from both the central and peripheral regions of the membrane were selected for analysis. Subsequently, the cells within each field were enumerated employing Image J software.
For the invasion assay, 0.8 μm Transwell insert pre-coated with Matrigel (354,480, BioCoat, USA) was used. All other processes were the same as for the transwell migration assay.
ELISA assay
The level of pro-inflammatory factor IL-1β, IL-6, and IL-8 in the fibroblasts treated with hypoxic or normoxic exosomes was measured using IL-1β ELISA kit (ml058059, Mlbio, China), IL-6 ELISA kit (mlml038115, Mlbio, China), and IL-8 ELISA kit (ml103387, Mlbio, China), TGF-β ELISA kit (ml064258, Mlbio, China) according to the manufacturer’s protocol. The OD values of each well were measured at a wavelength of 450 nm using a microcoder (ThermoFisher, FC). The concentrations of the target substances in the samples were calculated based on a standard curve.
Exosomal circRNA screening
The ESCC exosomal circRNA data were extracted from the Homo sapiens_ exosomal circRNA dataset of exoRBaseV2 (http://www.exoRBase.org), specifically from 6 ESCC patient plasma samples. To identify key exosomal circRNAs in ESCC, we screened for circRNAs with median and mean expression levels greater than 1000 in exosomes isolated from the plasma of 6 ESCC patients, yielding a total of 52 circRNAs. We presented the top 35 of these circRNAs in a heatmap (as displaying all 52 circRNAs would be unclear), while the expression profiles of all 52 circRNAs are provided in the Supplementary Table S1. Subsequently, we integrated these 52 circRNAs with 19 circRNAs that have been previously reported to be upregulated in ESCC [22]. This integration resulted in the identification of a single overlapping circRNA, circNRIP1 (exo_circ_38881 in the exoRBaseV2 database).
RNA isolation and real-time PCR
Total RNA was extracted from each sample with TRIzol reagent. Extracted RNA was quantified using a TGem spectrophotometer (TIANGEN, China) at 260/280 nm. High-quality RNA was reversely transcribed to synthesize cDNA using a Reverse Transcription Kit (Thermo, #K1622, USA). PCR amplification was done with 2 × Master Mix (Roche, USA). The PCR protocol was 95 °C for 10 min, followed by 45 cycles of 95 °C for 15 s, 60 °C for 60 s and a final dissociation stage, using the ABI Q6 real-time PCR system (Applied Biosystems, USA). We assumed the amplification efficiency of the target and reference were approximately equal. The relative changes in gene expression were calculated by the 2 − ΔΔCT method. The PCR primer sequences are as follows.
COL1A1, forward, GTACCATGACCGAGACGTGT, reverse, GCAGTTCTTGGTCTCGTCAC. COL3A1, forward, GCCAAATATGTGTCTGTGACTCA, reverse, GGGCGAGTAGGAGCAGTTG; circNRIP1, forward, CCGGATGACATCAGAGCTACT, reverse, GTGCATCTTCTGGCTGTGTT; Actin, forward, AGCACAGAGCCTCGCCTTTG, reverse, CTTCTGACCCATGCCCACCA.
Divergent and convergent primers were designed for circNRIP1 validation.
Divergent-circNRIP1, forward, CCGGATGACATCAGAGCTACT, reverse, GTGCATCTTCTGGCTGTGTT; convergent-circNRIP1, forward, CTCCGGATGACATCAGAGCT, reverse, CGTCTGTCTCCAAGCTCTGA.
RNA pull-down
The streptavidin-coated magnetic beads were incubated with MRC-5 cell lysates at 4 °C for 1 h for pre-clearance. The 3′ biotin-labeled circNRIP1 probe was incubated with the beads at room temperature for 10 min for immobilization. The biotinylated beads were incubated with MRC-5 cell lysates at 4 °C overnight. The beads were then magnetically separated and washed five times. The beads were boiled in an SDS buffer for protein isolation. The band protein was analyzed using western blot.
Statistical analysis
All experiments were independently performed at least three times. Student’s t-test was performed for comparisons between two groups. One-way ANOVA with Tukey’s post-hoc test was conducted to access significant differences among multiple groups. GraphPad Prism V9.5.0 was used for statistical analyses, and the data were presented as the mean ± standard deviation (S.D.). In our study, p < 0.05 was considered statistically significant. Significant data are denoted with asterisks: *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Result
Hypoxia enhances exosome release
To investigate the mechanism of hypoxic exosomes mediating the motility of ESCC, the exosomes derived from the cellular media of ESCC cell lines KYSE-150 and TE-10 with normoxia or hypoxic treatment were isolated and characterized utilizing a Transmission Electron Microscope (TEM). TEM images showed that the exosomes were cup-shaped particles ranging from 30 to 200 nm in diameter (Fig. 1A). An increased number of exosomes were observed in the same field of view after hypoxic treatment. We also detected the expression of exosomal marker ALIX, TSG101, and CD81 in the equal concentrations of exosomes, confirming the extracellular enrichment of exosomes from normoxia or hypoxic cells (Fig. 1B, C). In contrast, the cytoplasmic GAPDH showed low or absent expression in exosome lysates. Moreover, we quantified normoxic exosomes (Nor-exo) and hypoxic exosomes (Hypo-exo) from equal cells using a BCA protein assay. Hypoxia treatment significantly enhanced exosome release from ESCC cells compared with normoxic ESCC cells (Fig. 1D, E). These studies demonstrate that hypoxic conditions enhanced the secretion of exosomes from ESCC cells.
Fig. 1.
Hypoxia enhances exosome release. A Observation of exosomes using TEM. B Western blot detected the protein expression of exosome marker ALIX, TSG101, and CD81 in KYSE-150 cells and the exosomes using equal total protein. C Western blot detected the protein expression of exosome marker ALIX, TSG101, and CD81 in TE-10 cells and the exosomes using equal total protein. D BCA assay detected the total protein of normoxic exosomes (Nor-exo) and hypoxic exosomes (Hypo-exo) from equal KYSE-150 cells. E BCA assay detected the total protein of normoxic exosomes and hypoxic exosomes from equal TE-10 cells. Data represent mean ± S.D. of three independent experiments. **P < 0.01
Hypoxic exosomes induce the activation of CAFs
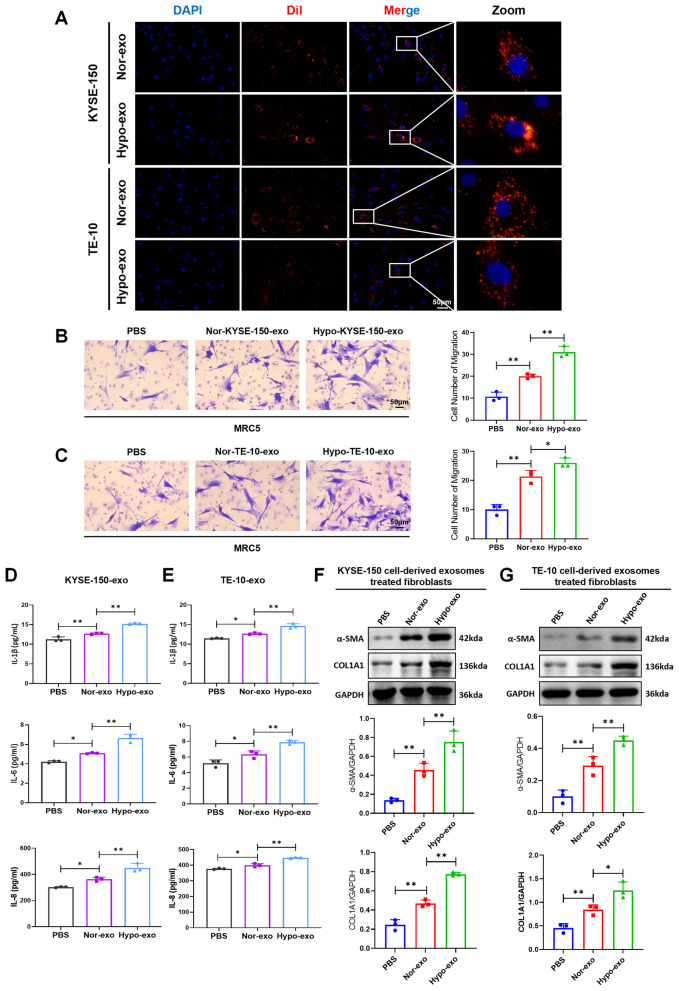
To investigate the regulatory effects of hypoxia-induced exosomes derived from ESCC on fibroblasts, we initially examined whether fibroblasts internalize exosomes secreted by ESCC cells. Normal fibroblasts (NFs) MRC5 cells were exposed to Dil fluorescent-labeled exosomes derived from ESCC cells. Microscopic imaging confirmed that the exosomes were internalized by the fibroblasts (Fig. 2A). For clarity and consistency, we hereafter refer to fibroblasts activated in vitro by ESCC-derived exosomes as CAFs, not primary CAFs isolated from tumors. The activation of NFs into CAFs was characterized by accelerated migration and an altered secretory profile [7]. Utilizing a transwell assay, we found that hypoxic exosomes significantly enhanced the migratory capacity of the fibroblasts compared to those exposed to normoxic exosomes (Fig. 2B, C). We then assessed the secretion of specific inflammatory cytokines via ELISA, which revealed an upregulation of IL-1β, IL-6, and IL-8 induced by hypoxic exosomes (Fig. 2D, E). Consistently, hypoxic exosomes enhanced α-SMA and COL1A1 expression in fibroblasts rather than normoxic exosomes (Fig. 2F, G). Taken together, hypoxic exosomes have the potential to transform NFs into CAFs.
Fig. 2.
Hypoxic exosomes induce the activation of CAFs. A Fluorescence images captured exosomes were internalized by fibroblast MRC5 cell line. B, C Transwell assay detected the migration of the treated fibroblasts. D, E ELISA assay examined the secretion of IL-1β, IL-6, and IL-8 in the treated fibroblasts. F, G Western blot detected the protein expression of α-SMA and COL1A1 in KYSE-150 and TE-10 cells-derived exosomes treated fibroblasts. (PBS: fibroblasts treated with PBS, Nor-exo: fibroblasts co-cultured with normoxic exosomes from TE-10 cells, Hypo-exo: fibroblasts treated with hypoxic exosomes from TE-10 cells.) Data represent mean ± S.D. of three independent experiments. *P < 0.05, **P < 0.01
circNRIP1 is enriched in hypoxic exosomes
To identify key circRNAs within exosomes derived from hypoxic ESCC cells, we initially utilized the exoRBaseV2 database to investigate which circRNAs can be enriched by ESCC cell-derived exosomes. Our analysis revealed 318 circRNAs exhibiting expression across 6 ESCC patient plasma exosomes. We first screened for circRNAs with both median and mean expression levels > 1000 in exosomes from 6 ESCC patient plasma samples, obtaining 52 candidate circRNAs. We visualized the top 35 of these circRNAs through a heatmap (Fig. 3A), with the complete expression profiles of all 52 circRNAs provided in supplementary materials (Supplementary Table S1). Subsequently, we intersected these 52 circRNAs with 19 previously reported ESCC-upregulated circRNAs [22], yielding a single overlapping circRNA, circNRIP1 (annotated as exo_circ_38881 in the exoRBaseV2 database), which was upregulated in ESCC and enriched within ESCC patient plasma exosomes (Fig. 3B). Meanwhile, circNRIP1 (exo_circ_38881) is alternatively cataloged as hsa_circ_0004771 in the circBase database. In addition, recent studies have reported that circNRIP1 contributes to the progression of ESCC [23], gastric cancer [24], and cervical cancer [25]. However, research on its role in exosomes and fibroblasts remains limited. Therefore, we focused on the possible role of exosomal circNRIP1 as novel players of the ESCC progression. circNRIP1 is located on chromosome 21 and formed by the back-splicing of exon 2 and exon3 of the NRIP1 gene (Fig. 3C).
Fig. 3.
circNRIP1 is enriched in hypoxic exosomes. A Heat map displayed the top expressed 35 exosomal circRNAs in ESCC from the exoRBaseV2 database. B The Venn diagram took the overlap of 52 exosomal circRNAs and upregulated circRNAs in ESCC tissues compared to normal tissues. C Genomic location of circNRIP1. D PCR of gDNA and cDNA using divergent and convergent primers of circNRIP1 in TE-10 cell line. E circNRIP1 was amplified using a divergent primer, followed by Sanger sequencing of the back-splicing junction. F, G Real-time PCR detected the expression of circNRIP1 in normoxic and hypoxic KYSE-150 and TE-10 cells. (H, I) Real-time PCR detected the expression of circNRIP1 in normoxic and hypoxic exosomes from ESCC cells. Data represent mean ± S.D. of three independent experiments. *P < 0.05, **P < 0.01
Next, we designed convergent primers facing inward to amplify linear transcripts and divergent primers facing outward to amplify circular transcripts, with cDNA and gDNA as the templates. Agarose gel electrophoresis showed that the convergent primer amplified bands with cDNA and gDNA of both groups, whereas the divergent primer only amplified bands using cDNA templates (Fig. 3D). We further confirmed the presence of the back-splicing junction of circNRIP1 via Sanger sequencing of the amplification products of divergent primers on cDNA (Fig. 3E). To further elucidate the enrichment of circNRIP1 in hypoxic exosomes from ESCC cells, we used real-time PCR to detect the expression of circNRIP1 in hypoxic treated KYSE-150 and TE-10 cells and their exosomes. The results indicated that hypoxia had no significant effect on the expression of circNRIP1 in KYSE-150 cells (Fig. 3F, G). However, under hypoxic conditions, a marked elevation in circNRIP1 expression was observed in exosomes from both KYSE-150 and TE-10 cell lines (Fig. 3H, I), suggesting a role for hypoxia in modulating circNRIP1 abundance in ESCC-derived exosomes. The above results indicated that circNRIP1 is enriched in hypoxic exosomes and exhibits a circular structure.
Hypoxic exosomes induce CAF activation via transferring circNRIP1
To investigate the potential of hypoxia-induced exosomes in activating CAFs by transporting circNRIP1, we first measured the expression of circNRIP1 in fibroblasts treated with PBS, normoxic exosomes, and hypoxic exosomes derived from ESCC cells. Our findings revealed that the incubation of hypoxic exosomes robustly enhanced circNRIP1 expression within fibroblasts, surpassing that of their normoxic group (Fig. 4A). This enhancement suggested that hypoxic exosomes facilitated the internalization of circNRIP1 into fibroblasts, consequently elevating intracellular circNRIP1 levels in fibroblasts. Additionally, a rescue assay was executed to substantiate the contribution of circNRIP1-enriched exosomes from hypoxic ESCC cells to CAF activation. Hypoxic exosomes were harvested following the in vitro knockdown of circNRIP1 in ESCC cells (Fig. 4B). The transwell assay showed that the treatment with hypoxic exosomes significantly increased fibroblast migration in comparison to the treatment of normoxic exosomes. However, a notable reduction in the migratory capacity of CAFs was observed when treated with circNRIP1-depleted hypoxic exosomes, as compared to the hypoxic exosomes-NC group (Fig. 4C, D). These results suggest that the loss of circNRIP1 reverses the promoting effect of hypoxic exosomes on CAF activation. Subsequently, we employed ELISA and western blot to further confirm the effect of exosomal circNRIP1 on activating CAFs. The outcomes demonstrated that the loss of circNRIP1 led to a noticeable reduction in the levels of pro-inflammatory cytokines IL-1β, IL-6, and IL-8 (Fig. 4E), as well as in the expression of α-SMA and collagen types COL1A1 and COL3A1 (Fig. 4F-I). Real-time PCR revealed that circNRIP1 deprivation reversed the effect of hypoxic exosomes on elevated mRNA levels of COL1A1 and COL3A1 in TE-10 cells (Fig. 4J, K). Collectively, these findings indicate that hypoxic exosomes contributed to CAF activation via circNRIP1.
Fig. 4.
Hypoxic exosomes induce CAF activation via transferring circNRIP1. A PCR detected the expression of circNRIP1 in CAFs treated with exosomes derived from ESCC cells under normoxia or hypoxic conditions. B PCR detected the expression of circNRIP1 in TE-10 cells with lentiviral transfection. C, D Representative images and quantitative analysis of fibroblast migration based on transwell assays. E ELISA assay detected pro-inflammatory cytokines IL-1β, IL-6, and IL-8 secreted from fibroblasts. F-I Western blot detected the protein expression of pro-inflammatory cytokines α-SMA, COL1A1 and COL3A1 within fibroblasts. J, K PCR detected the relative mRNA expression of COL1A1 and COL3A1 within fibroblasts. (Nor-exo: fibroblasts co-cultured with normoxic exosomes from TE-10 cells, Hypo-exo-NC: fibroblasts treated with hypoxic exosomes from TE-10 cells transfected with control shRNA, Hypo-exo-shcircNRIP: fibroblasts treated with hypoxic exosomes from TE-10 cells transfected with circNRIP1 shRNA). Data represent mean ± S.D. of three independent experiments. *P < 0.05, **P < 0.01
Hypoxic exosomes circNRIP1-mediated CAFs promote ESCC cell invasion and migration
Given that CAFs possess the capability to secrete a diverse array of cytokines, growth factors, and extracellular matrix proteins, which contribute to cancer malignancy, we detected the migration and invasion of TE-10 cells using the conditioned medium (CM) from exosome-treated CAFs (Fig. 5A). Relative to the CM of normal fibroblasts, the medium induced by hypoxic exosomes substantially enhanced the migratory and invasive capacities of ESCC cells. Nonetheless, the depletion of circNRIP1 in hypoxic exosomes clearly abrogated the activating impact of the hypoxic exosome-enriched medium on cancer cell motility (Fig. 5B-D). Through these findings, we highlighted that hypoxic exosome-mediated CAFs promoted the migration and invasion of ESCC cells relying on circNRIP1.
Fig. 5.
circNRIP1-mediated CAFs promote ESCC cell invasion and migration. A The schematic drawing of transwell detecting hypoxic exosomal circNRIP1 activated CAF promoting the TE-10 migration: MRC5-derived fibroblasts were treated with hypoxic exosomes from ESCC cells to induce CAF activation. After 24 h, the conditioned medium from the activated CAFs was collected and used to treat TE-10 cells, whose migration was then assessed by the transwell assay. B-D Representative images and quantitative analysis of fibroblast migration based on transwell assays. Hypo-exo ( ±): Fibroblasts treated with ( +) or without (-) hypoxia-conditioned exosomes derived from TE-10 cells. shcircNRIP1 ( ±): Fibroblasts receiving exosomes from TE-10 cells with ( +) or without (-) circNRIP1 knockdown. (blank: TE-10 co-cultured with PBS, Fibroblast CM: TE-10 co-cultured with the CM of normal fibroblasts, Hypo-exo-NC: TE-10 co-cultured with the CM of fibroblasts adding control shRNA exosomes, Hypo-exo-shcircNRIP1: TE-10 co-cultured with the CM of fibroblasts adding shcircNRIP1 exosomes). Data represent mean ± S.D. of three independent experiments. **P < 0.01
circNRIP1 activates CAFs by binding to TRMT6
Next, we delved deeper into the molecular mechanism underlying the circNRIP1-mediated activation of CAFs. While existing research has established circNRIP1 as miRNA sponges, the RNA binding protein of circNRIP1 has yet to be fully identified. To uncover proteins intricately linked to circNRIP1 and likely contributors to oncogenic processes, we performed bioinformatics predictions using catRAPID omics v2.1 (Supplementary Fig. 1A-C). Go enrichment analysis revealed that most circNRIP1-binding proteins have necessary domains and motifs for ‘RNA binding’ (Supplementary Fig. 1D). Of these RNA-binding proteins, TRMT6 has been implicated in promoting tumorigenesis across various cancer types [26–28], prompting us to concentrate our investigations on TRMT6. Analysis result indicated the physical interaction between circNRIP1 and transfer RNA (tRNA) m(1)A58 ‘writer’ proteins TRMT6 (Fig. 6A). Subsequent western blot validation in the RNA pull-down products (Fig. 6B, C) revealed that TRMT6 is a protein targeted by circNRIP1 in fibroblasts.
Fig. 6.
circNRIP1 activates CAFs by binding to TRMT6. A Graphical representation of the molecular docking between circNRIP1 and the TRMT6 protein using HDock. B, C Western blot was conducted to validate the circNRIP1 pull-down products in fibroblasts. D The expressions of TRMT6 were determined by western blot and real-time PCR. E, F Transwell assays detected the migration of CAFs. G ELISA assays detected the secretion of IL-1β and TGF-β in CAFs. H Western blot detected the expression of α-SMA and COL1A1 in CAFs. (NC: fibroblasts without treatment, shcircNRIP1: fibroblasts co-cultured with shcircNRIP1 exosomes, TRMT6-OE: TRMT6-overexpressing fibroblasts, shcircNRIP1 + TRMT6-OE: TRMT6-overexpressing fibroblasts co-cultured with shcircNRIP1 exosomes.) Data represent mean ± S.D. of three independent experiments. *P < 0.05, **P < 0.01, ****P < 0.0001
To verify that circNRIP1 regulates CAF activation by targeting TRMT6, a rescue experiment was undertaken using fibroblasts treated with exosomes carrying circNRIP1 knockdown and subsequent overexpression of TRMT6. The increased expression of TRMT6 was confirmed in treated fibroblasts by RT-PCR as well as western blot (Fig. 6D). Transwell assays revealed a significant reduction in migration in the shcircNRIP1 group compared to the negative control. In addition, overexpression of TRMT6 in CAFs enhanced the motility of the CAFs and allowed for restoration of CAF phenotype under shcircNRIP1 treatment (Fig. 6E, F). We further measured the expression of soluble factors produced by CAFs utilizing ELISA and western blot. Consistent with the observed migratory behaviors, circNRIP1 loss inhibited the secretion of IL-1β and TGF-β (Fig. 6G) and the protein expression of α-SMA and COL1A1 (Fig. 6H). The addition of TRMT6 elevated inflammatory cytokine and matrix protein expression compared to the negative control and yet notably rescued the effect of circNRIP1 knockdown in CAFs.
Furthermore, to clarify that the CAF activation defect was specifically due to the loss of TRMT6, we performed rescue experiments. First, MRC5-derived fibroblasts were infected with shNC or shTRMT6 lentiviruses (Supplementary Fig. 2A), followed by treatment with hypoxic exosomes derived from TE-10 cells. Transwell migration assays revealed that TRMT6 knockdown significantly reduced fibroblast motility compared to the PBS-treated shNC control group (Supplementary Fig. 2B, C). Notably, the defect of TRMT6 reversed the increased migration of CAFs upon treatment with hypoxia-induced exosomes. Consistent with these findings, ELISA analysis of fibroblast supernatants showed that TRMT6 depletion led to a significant decrease in CAF-associated secretory factors, including IL-1β and TGF-β (Supplementary Fig. 2D). In particular, the loss of TRMT6 reversed the enhancing effect of hypoxia-induced exosome treatment on the levels of both cytokines. Next, we examined the expression of CAF-related markers in fibroblasts by immunoblotting (Supplementary Fig. 2E). Loss of TRMT6 protein was directly responsible for the reduction in α-SMA, COL1A1, and COL3A1 levels and abrogated their expression by hypoxic exosome addition. Taken together, these findings support a causal role for circNRIP1 in activating the CAF phenotype by targeting TRMT6. The schematic overview displayed the mechanism in this study (Fig. 7).
Fig. 7.
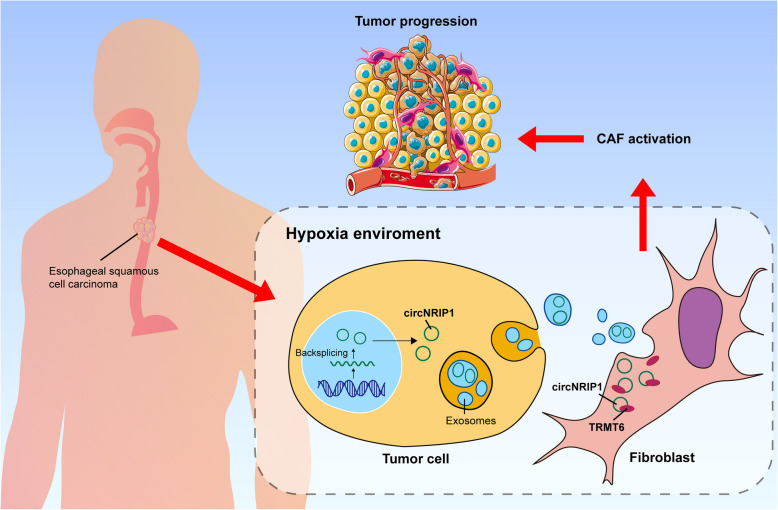
Schematic representation of how hypoxia-derived exosomes from ESCC cells activate CAFs and promote tumor progression by delivering circNRIP1 to target TRMT6
Discussion
ESCC is a heterocellular disease characterized by the collaboration of several different cell types through established interactions, which collectively form a functional TME [29, 30]. This disease exhibits a notably higher incidence in the East Asian population and is associated with a poor prognosis [31]. In this study, we found that hypoxia-derived exosomes from ESCC cells not only increase in quantity but also activate CAFs through circNRIP1 delivery, finally promoting ESCC invasion. Additionally, we demonstrated that circNRIP1 interacts with the TRMT6 protein, thereby mediating the activation of CAFs. Our research elucidates the mechanisms and roles of hypoxic exosomes in the progression of ESCC, contributing novel insights into tumor progression and opening new avenues for treatment.
Environmental stress is a critical factor contributing to the challenges associated with curing ESCC. Low oxygen tension, a common characteristic of the TME, modifies both homotypic and heterotypic interactions between tumor cells and stromal cells, thereby inducing tumorigenesis through various pathways [32, 33]. Exosomes, which are abundant extracellular vesicles within the ESCC TME, facilitate intercellular communication that plays a significant role in the development and aggressiveness of cancer [34]. Hypoxic exosomal circRNA ZNF609 is internalized by endothelial cells, promoting VEGFA-induced angiogenesis, a process essential for forming a pre-metastatic niche in ESCC [35]. In our study, we observed an increase in the release of exosomes from hypoxic ESCC cells, indicating enhanced intercellular communication. Recent studies have indicated that exosomes derived from tumor cells can promote the activation of CAFs. Tumor-derived exosomes induce CAFs to shape an inflammatory tumor microenvironment in head and neck squamous cell carcinoma [36]. Freag et al. found that inhibiting exosome biogenesis and release from lung cancer cells can block the differentiation of lung fibroblasts into CAFs [37]. Our study revealed that ESCC-derived hypoxic exosomes enhance fibroblast migration and their transformation into CAFs, thereby promoting migration and invasion of ESCC cells.
circRNAs are highly abundant and exhibit tissue specificity within the TME of ESCC, and their aberrant expression in exosomes can regulate the intercellular communication [38] and malignant behavior [39]. We identified circNRIP1 as a highly expressed circRNA in both ESCC exosomes and cells, with particular upregulation in hypoxic exosomes. circNRIP1, broadly expressed and implicated in promoting proliferation, metastasis, and inhibiting apoptosis and chemotherapy resistance, is a significant marker across various tumors [40]. Z. Shi et al. found that BMSC-EVs facilitate the delivery of circNRIP1 to osteosarcoma cells, where it outcompetes miR-532-3p for binding, thereby reducing miR-532-3p’s inhibitory effect on AKT3 and promoting tumorigenesis [41], which proposed that circNRIP1 contributes to ESCC progression through its interactions in the TME. Previous studies reported that circNRIP1 knockdown effectively suppressed ECM-related protein expression in keloid-derived fibroblasts [42]. Similarly, we found that exosomal circNRIP1 upregulates α-SMA and Collagen I, two key markers of ECM synthesis [43]. In addition, excessive ECM remodeling and cytokine secretion are hallmarks of CAFs [44]. Our findings confirm that exosomal circNRIP1 increased the secretion of inflammatory cytokines (IL-1β and TGF-β) and ECM-related proteins α-SMA and Collagen I (encoded by COL1A1 gene). Therefore, our results demonstrate that exosomal circNRIP1 directly activates CAFs by inducing inflammatory cytokine secretion and matrix remodeling. Activated CAFs promoted tumor invasion, angiogenesis, and growth by ECM remodeling [45]. Meanwhile, CAFs interact with tumor-infiltrating immune cells by secreting various cytokines, growth factors, and chemokines, thereby shaping an immunosuppressive TME that enables cancer cells to evade immune surveillance [46]. Previous studies have shown that exosomal miRNA-activated CAFs from tumor cells promote hepatocellular carcinoma metastasis via inflammatory cytokines [47]. Knockdown of circNRIP1 in gastric cancer cells significantly reduces the expression of mesenchymal markers while upregulating the epithelial marker E-cadherin in organoid models, suggesting its critical role in epithelial-mesenchymal transition and tumor extracellular matrix remodeling [24]. Our findings confirm that exosomal circNRIP1 activates CAFs by enhancing the secretion of inflammatory cytokines (IL-1β and TGF-β) and ECM-related proteins (α-SMA and Collagen I). Based on this evidence, we speculate that exosome-activated CAFs may promote ESCC progression through ECM remodeling and inflammatory factor secretion. However, further in vivo validation is required to confirm these findings. Our findings suggested that circNRIP1 depletion can significantly reverse the effect of hypoxic exosomes from ESCC cells on CAF activation. CAF activation leads to the collective invasion of ESCC cells by remodeling the extracellular matrix [10]. In vivo studies demonstrate that therapeutic interventions targeting MFGE8 secreted by CAFs or its receptor significantly suppress ESCC proliferation and metastasis [11]. We observed that culturing ESCC cells with a conditioned medium from hypoxic exosome-induced CAFs enhanced their migration and invasion, effects that were abrogated by circNRIP1 depletion.
Although we chose to focus on circNRIP1, we acknowledge that exosomes carry numerous proteins and different types of RNA, and the other abundant exosomal circRNAs may also play roles in the tumorigenesis. For example, we identified exo_circ_27945 (hsa_circ_0000437) and exo_circ_27945 (hsa_circ_0000896) in plasma-derived exosomes from ESCC patients. hsa_circ_0000437 promoted lymphangiogenesis and lymph node metastasis, when it was enriched in gastric cancer-secreted exosomes and transferred into lymphatic endothelial cells [48]. hsa_circ_0000896 (circFARSA) facilitated the malignant phenotype of NSCLC cells by elevating B7H3 expression through sponging miR-15a-5p [49]. These circRNAs may work synergistically with circNRIP1 to support the activation of CAF, but exosomal circNRIP1 remains an important factor for CAF activation under hypoxia conditions. Beyond circRNAs, exosomes carry proteins and miRNAs reported to activate CAFs. For instance, bladder cancer-derived exosomes deliver TGF-β to fibroblasts, activating SMAD signaling and CAF differentiation [50]. HSPC111 in cancer exosomes reprograms CAFs to facilitate colorectal cancer liver metastasis [51]. Similarly, CRC exosomal miR-146a-5p and miR-155-5p promote CAF activation via JAK2-STAT3/NF-κB pathways [52]. These findings imply that exosomal proteins and miRNAs may synergize with circNRIP1 in CAF activation. Furthermore, we have only demonstrated that hypoxic exosomal circNRIP1 activates CAFs in vitro and that the induced fibroblasts further promote cancer cell migration and invasion in vitro, the role of hypoxic exosomal circNRIP1 in vivo remains to be further verified. However, previous studies provide partial support that hypoxic exosomal circNRIP1 may promote tumor growth in vivo. Specifically, it has been shown that circNRIP1 is highly expressed in ESCC patient tissues and can promote ESCC tumor progression [23]. Activated CAFs have been reported to promote tumor growth [53], including the growth and dissemination of ESCC cells [54]. Moreover, hypoxia-induced fibroblasts have been shown to promote the carcinogenesis of ESCC [55]. These data suggest that hypoxic exosome circNRIP1 may promote ESCC tumor growth and metastasis in vivo by activating CAFs. We acknowledge that the “hypoxic exosome circNRIP1-CAF-tumor growth” signaling axis requires validation in animal models or patient-derived tissues, representing a limitation of this study. Future investigations will employ in vivo models through co-injection of ESCC cells with fibroblasts or orthotopic xenografts under hypoxic or normoxic conditions to verify whether circNRIP1-enriched exosomes indeed promote CAF activation and tumor growth, thereby elucidating the underlying mechanisms.
Evidence suggests that circRNAs participate in various cancer biological processes by regulation at the transcriptional level, serving as protein and miRNA sponges, translation into functional peptides, and mediation in cell signaling [56]. circNRIP1 can regulate the activity of ERK1/2, PI3K/AKT, and AKT/mTOR signaling pathways [40]. Most studies involving circNRIP1 have been confined to its role as microRNA sponges [24, 57]. Besides, circNRIP1 contributed to thoracic aortic dissection progression by promoting CXCL5 mRNA stability via competitively binding to m6A ‘reader’ protein IGF2BP1 [58]. Our research has discovered that circNRIP1 is capable of binding to the protein TRMT6. TRMT6, a tRNA methyltransferase, is involved in post-translational modifications through N1-methyladenosine (m1A) modification. In bladder cancer, TRMT6 regulates the gene-silencing activity and the unfolded protein response [27]. TRMT6 also drives liver tumourigenesis by elevating the m1A methylation in tRNAs to regulate translation, which in turn triggers cholesterol synthesis [26]. Our findings confirmed that TRMT6 overexpression significantly upregulates two key ECM markers (Collagen I and α-SMA) and inflammatory cytokines (IL-1β and TGF-β), all of which are established drivers of CAF activation. Conversely, TRMT6 knockdown is responsible for the defect of pro-inflammatory cytokines. Importantly, TRMT6 overexpression rescues the inhibitory effects of exosomal circNRIP1 knockdown on CAF-related protein expression, while TRMT6 knockdown abolishes the hypoxic exosome-induced CAF activation. These results strongly suggest that hypoxic exosomal circNRIP1 activates CAFs by binding TRMT6 to induce Collagen I, α-SMA, IL-1β and TGF-β expression. While m1A modification has been reported to participate in nearly all RNA biological processes [59], the precise mechanism by which TRMT6 regulates these targets remains to be elucidated. Specifically, it remains unknown whether TRMT6 directly mediates m1A modification of COL1A1, α-SMA, IL-1β and TGF-β RNAs to affect their protein expression, or indirectly modulates their expression through other signaling pathways. This represents an important area for future investigation in our ongoing research.
However, certain limitations of this study should be acknowledged. First, our findings are based on experiments using two ESCC cell lines and an in vitro fibroblast model, which cannot fully capture the heterogeneity of ESCC tumors or completely examine the correlation between exosomal circNRIP1 levels and intratumoral CAF marker expression. Future investigations should incorporate additional ESCC models or patient-derived samples. These studies should include blocking circNRIP1 in exosomes and injecting them into orthotopic tumor models, as well as examining circNRIP1 levels in circulating exosomes from ESCC patients and CAF marker expression in matched tumor tissues. These important studies will be critical for establishing the clinical relevance of targeting exosomal circNRIP1 as a potential therapeutic strategy. In addition, while the human embryonic lung fibroblast MRC5 cell line has been widely used to investigate tumor-stromal interactions in hepatocellular carcinoma and ESCC [47, 60, 61], its biological relevance to in vivo conditions remains uncertain. We acknowledge that the “hypoxic exosomal circNRIP1-CAF-tumor growth” axis needs to be confirmed in animal models or patient-derived tissues, which is a limitation of our study. In the future, we will confirm that circNRIP1-enriched exosomes indeed promote CAF activation and tumor growth or metastasis by co-injecting ESCC cells and fibroblasts in in vivo models or using orthotopic xenografts under hypoxic and normoxic conditions to further explore the underlying mechanisms. Consequently, the generalizability of exosomal circNRIP1-mediated CAF activation across all ESCC contexts requires further validation. Furthermore, the precise molecular cascade linking circNRIP1-TRMT6 binding to CAF activation warrants deeper investigation, including potential crosstalk with TRMT6’s established roles in tsRNA regulation.
In summary, this study revealed that hypoxic conditions induce the enrichment of circNRIP1 in ESCC-derived exosomes. Hypoxic exosomal circNRIP1 activated CAFs, potentially through targeting TRMT6, which further promotes the migration and invasion of ESCC cells. By elucidating the tumor-stroma crosstalk mediated by hypoxic exosomal circNRIP1, our research contributes to a deeper understanding of the molecular mechanisms driving ESCC progression in oxygen-deprived environments.
Supplementary Information
Supplementary Material 2: Supplementary Fig. 1 Predicting the binding protein of circNRIP1. (A) Presence of RNA − binding domains among circNRIP1-binding protein. (B) Most detected RNA − binding domains. (C) Most detected RNA − binding motifs among circNRIP1-binding protein. (D) GO enrichment analysis of circNRIP1-binding protein.
Supplementary Material 3: Supplementary Fig. 2 Rescue experiments confirm the essential role of TRMT6 in CAF activation. (A) The expression of TRMT6 were determined by western blot. (B, C) Transwell assays detected the migration of CAFs. (D) ELISA assays detected the secretion of IL-1β and TGF-β in CAFs. (E) Western blot detected the expression of α-SMA and COL1A1 in CAFs. (PBS + shNC: mimic control-transfected fibroblasts treated with PBS, PBS + shTRMT6: TRMT6-silienced fibroblasts treated with PBS, Hypo-exo + shNC: mimic control-transfected fibroblasts co-cultured with shcircNRIP1 exosomes, Hypo-exo + shTRMT6: TRMT6-silienced fibroblasts co-cultured with shcircNRIP1 exosomes.) Data represent mean ± S.D. of three independent experiments. *P < 0.05, **P < 0.01.
Acknowledgements
Not applicable.
Authors’ contributions
Guan’en Qiao, Shuanli Xin, and Changjuan Li were responsible for overall data analysis and wrote the manuscript. Bing Meng, Lixia Zhang, Ruiqing Zhang, and Huihui Qi performed the validation and investigation. Meng Wang, Wenjuan Zhang, and Junjie Shi were responsible for the statistical analysis. Guan’en Qiao and Shuanli Xin were responsible for the study design and revised the manuscript. All the authors read and approved the final form of the manuscript.
Funding
This study was supported by Hebei Natural Science Foundation (No. H2022109014).
Data availability
The datasets used during the present study are available from the corresponding author upon reasonable request.
Declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Guan’en Qiao, Email: qiaoguanen@qq.com.
Shuanli Xin, Email: xhdsdyyy1968@126.com.
References
- 1.Morgan E, et al. The global landscape of esophageal squamous cell carcinoma and esophageal adenocarcinoma incidence and mortality in 2020 and projections to 2040: New estimates from GLOBOCAN 2020. J Gastroenterol. 2022;163(3):649–658.e2. [DOI] [PubMed]
- 2.Thrift AP. Global burden and epidemiology of Barrett oesophagus and oesophageal cancer. J Nat Rev Gastroenterol Hepatol. 2021;18(6). [DOI] [PubMed]
- 3.Shi Y, Ge X, Ju M, Zhang Y, Di X, Liang L. “Circulating Tumor Cells in Esophageal Squamous Cell Carcinoma - Mini Review”, (in eng). Cancer Manag Res. 2021;13:8355–65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Xiaoqi M, et al. Crosstalk between cancer-associated fibroblasts and immune cells in the tumor microenvironment: new findings and future perspectives. J Mol Cancer. 2021;20(1):131–131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Quail DF and Joyce JA. Microenvironmental regulation of tumor progression and metastasis. Nat Med. 2013;19(11):1423–37. [DOI] [PMC free article] [PubMed]
- 6.Biffi G, Tuveson DA. “Diversity and Biology of Cancer-Associated Fibroblasts”, (in eng). Physiol Rev. 2021;101(1):147–76. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Caligiuri G, Tuveson DA. “Activated fibroblasts in cancer: Perspectives and challenges”, (in eng). Cancer Cell. 2023;41(3):434–49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Yang X, et al. “Collagen 1-mediated CXCL1 secretion in tumor cells activates fibroblasts to promote radioresistance of esophageal cancer”, (in eng). Cell Rep. 2023;42(10): 113270. [DOI] [PubMed] [Google Scholar]
- 9.Duan Y, et al. “Hypoxia-induced miR-5100 promotes exosome-mediated activation of cancer-associated fibroblasts and metastasis of head and neck squamous cell carcinoma”, (in eng). Cell Death Dis. 2024;15(3):215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Qiu L, Yue J, Ding L, Yin Z, Zhang K, Zhang H. “Cancer-associated fibroblasts: An emerging target against esophageal squamous cell carcinoma”, (in eng). Cancer Lett. 2022;546: 215860. [DOI] [PubMed] [Google Scholar]
- 11.Beilei L, et al. Targeting MFGE8 secreted by cancer-associated fibroblasts blocks angiogenesis and metastasis in esophageal squamous cell carcinoma. Proc Natl Acad Sci U S A. 2023;120(42):e2307914120–e2307914120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Zhang S, Li D, Wang H, Liu B, Du F and Wang Q, CAFs-derived Exosomal miR-889–3p Might Repress M1 Macrophage Polarization to Boost ESCC Development by Regulating STAT1. Cell Biochem Biophys. 2025;83(1):633–46. [DOI] [PubMed]
- 13.Xueman C, Erwei S. Turning foes to friends: targeting cancer-associated fibroblasts. Nat Rev Drug Discov. 2019;18(2):99–115. [DOI] [PubMed] [Google Scholar]
- 14.van Niel G, D’Angelo G, Raposo G. Shedding light on the cell biology of extracellular vesicles. Nat Rev Mol Cell Biol. 2018;19(4):213–28. [DOI] [PubMed] [Google Scholar]
- 15.Ruixue B, Yunong L, Lingyan J, Yuehui Y, Lin Z, Minjie W. The hypoxia-driven crosstalk between tumor and tumor-associated macrophages: mechanisms and clinical treatment strategies. Mol Cancer. 2022;21(1):177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Jiang H, et al. “Hypoxia Induced Changes of Exosome Cargo and Subsequent Biological Effects”, (in eng). Front Immunol. 2022;13: 824188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Chuchu S, et al. Role of hypoxia-induced exosomes in tumor biology. Mol Cancer. 2018;17(1):120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jiewei L, et al. Hypoxia-induced exosomal circPDK1 promotes pancreatic cancer glycolysis via c-myc activation by modulating miR-628–3p/BPTF axis and degrading BIN1. J Hematol Oncol. 2022;15(1):128–128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Rui Z, et al. Exosomal circLPAR1 functions in colorectal cancer diagnosis and tumorigenesis through suppressing BRD4 via METTL3–eIF3h interaction. Mol Cancer. 2022;21(1):49–49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Bei P, et al. Identification of Serum Exosomal hsa-circ-0004771 as a Novel Diagnostic Biomarker of Colorectal Cancer. Front Genet. 2019;10:1096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Tianxia L, Hongya G, Jia L, Jiang W, Yueli Z. Curcumin inhibits esophageal squamous cell carcinoma progression through down-regulating the circNRIP1/miR-532–3p/AKT pathway. Environ Toxicol. 2023;38(11):2705–16. [DOI] [PubMed] [Google Scholar]
- 22.Zhang X, et al. “Circular RNAs and esophageal cancer”, (in eng). Cancer Cell Int. 2020;20:362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Luo T, Guan H, Liu J, Wang J, Zhang Y. “Curcumin inhibits esophageal squamous cell carcinoma progression through down-regulating the circNRIP1/miR-532-3p/AKT pathway”, (in eng). Environ Toxicol. 2023;38(11):2705–16. [DOI] [PubMed] [Google Scholar]
- 24.Zhang X, et al. “Circular RNA circNRIP1 acts as a microRNA-149-5p sponge to promote gastric cancer progression via the AKT1/mTOR pathway”, (in eng). Mol Cancer. 2019;18(1):20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Li X, et al. “Circular RNA circNRIP1 promotes migration and invasion in cervical cancer by sponging miR-629-3p and regulating the PTP4A1/ERK1/2 pathway”, (in eng). Cell Death Dis. 2020;11(5):399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Wang Y, et al. “N(1)-methyladenosine methylation in tRNA drives liver tumourigenesis by regulating cholesterol metabolism”, (in eng). Nat Commun. 2021;12(1):6314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Su Z, et al. “TRMT6/61A-dependent base methylation of tRNA-derived fragments regulates gene-silencing activity and the unfolded protein response in bladder cancer”, (in eng). Nat Commun. 2022;13(1):2165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Zhao K, et al. “Comprehensive analysis of m(6)A/m(5)C/m(1)A-related gene expression, immune infiltration, and sensitivity of antineoplastic drugs in glioma”, (in eng). Front Immunol. 2022;13: 955848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Saigí M, et al. Clinical relevance of histologic subtypes in locally advanced esophageal carcinoma treated with pre-operative chemoradiotherapy: Experience of a monographic oncologic centre. PLoS ONE. 2017;12(9):e0184737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Shaikha A, Mehreen A, Abdolrahman NS. Understanding cell-cell communication and signaling in the colorectal cancer microenvironment. Clin Transl Med. 2021;11(2):e308–e308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Han B, et al. Cancer incidence and mortality in China, 2022. J Natl Cancer Cent. 2024;4(1):47–53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Yingming Z, Yuanwei Z, Minghuan L, Jinming Y. Inhibition of hypoxia-inducible factor-1α by PX-478 as a potential targeted therapy in ESCC. J Clin Oncol. 2017;35(15):e14083–e14083. [Google Scholar]
- 33.Pouysségur J, Dayan F, Mazure NM. Hypoxia signalling in cancer and approaches to enforce tumour regression. Nature. 2006;441(5):437–43. [DOI] [PubMed] [Google Scholar]
- 34.Zhao L, et al. “Mechanisms of function and clinical potential of exosomes in esophageal squamous cell carcinoma”, (in eng). Cancer Lett. 2023;553:215993. [DOI] [PubMed] [Google Scholar]
- 35.Yu M, Jiahao W, Yimin W, Zhanzhao F, Lixin D, Jia L. Hypoxia induced exosomal Circ-ZNF609 promotes pre-metastatic niche formation and cancer progression via miR-150–5p/VEGFA and HuR/ZO-1 axes in esophageal squamous cell carcinoma. Cell Death Discov. 2024;10(1):133–133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ikko M, et al. Tumor-derived exosomes elicit cancer-associated fibroblasts shaping inflammatory tumor microenvironment in head and neck squamous cell carcinoma. Oral Oncol. 2023;136:106270–106270. [DOI] [PubMed] [Google Scholar]
- 37.Freag MS, Mohammed MT, Kulkarni A, Emam HE, Maremanda KP, Elzoghby AO. Modulating tumoral exosomes and fibroblast phenotype using nanoliposomes augments cancer immunotherapy. Sci Adv. 2024;10(9):eadk3074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Jinyi L, et al. The biology, function, and applications of exosomes in cancer. Acta Pharmaceutica Sinica B. 2021;11(9):2783–97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Bo T, Qingfeng Z, Kui L, Yun H. Exosomal circRNA FNDC3B promotes the progression of esophageal squamous cell carcinoma by sponging miR-490–5p and regulating thioredoxin reductase 1 expression. Bioengineered. 2022;13(5):13829–48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Du Z. “CircNRIP1: An emerging star in multiple cancers”, (in eng). Pathol Res Pract. 2023;241: 154281. [DOI] [PubMed] [Google Scholar]
- 41.Shi Z, Wang K, Xing Y, Yang X. “CircNRIP1 Encapsulated by Bone Marrow Mesenchymal Stem Cell-Derived Extracellular Vesicles Aggravates Osteosarcoma by Modulating the miR-532-3p/AKT3/PI3K/AKT Axis”, (in eng). Front Oncol. 2021;11: 658139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Wang B, Yin H, Zhang H, Wang T. circNRIP1 facilitates keloid progression via FXR1-mediated upregulation of miR-503-3p and miR-503-5p. Int J Mol Med. 2021;47(5):70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Cao W, Feng Y. “LncRNA XIST promotes extracellular matrix synthesis, proliferation and migration by targeting miR-29b-3p/COL1A1 in human skin fibroblasts after thermal injury”, (in eng). Biol Res. 2019;52(1):52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Barbazan J, et al. “Cancer-associated fibroblasts actively compress cancer cells and modulate mechanotransduction”, (in eng). Nat Commun. 2023;14(1):6966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Jia H, Chen X, Zhang L, Chen M. “Cancer associated fibroblasts in cancer development and therapy”, (in eng). J Hematol Oncol. 2025;18(1):36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Mao X, et al. “Crosstalk between cancer-associated fibroblasts and immune cells in the tumor microenvironment: new findings and future perspectives”, (in eng). Mol Cancer. 2021;20(1):131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Fang T, et al. “Tumor-derived exosomal miR-1247-3p induces cancer-associated fibroblast activation to foster lung metastasis of liver cancer”, (in eng). Nat Commun. 2018;9(1):191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Shen X, et al. “Hsa_circ_0000437 promotes pathogenesis of gastric cancer and lymph node metastasis”, (in eng). Oncogene. 2022;41(42):4724–35. [DOI] [PubMed] [Google Scholar]
- 49.Nie J, et al. Circular RNA circFARSA promotes the tumorigenesis of non-small cell lung cancer by elevating B7H3 via sponging miR-15a-5p. Cell Cycle. 2022;21(24):2575–89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Ringuette Goulet C, Bernard G, Tremblay S, Chabaud S, Bolduc S, Pouliot F. Exosomes Induce Fibroblast Differentiation into Cancer-Associated Fibroblasts through TGFβ Signaling. Mol Cancer Res. 2018;16(7):1196–204. [DOI] [PubMed] [Google Scholar]
- 51.Zhang C, et al. “Cancer-derived exosomal HSPC111 promotes colorectal cancer liver metastasis by reprogramming lipid metabolism in cancer-associated fibroblasts”, (in eng). Cell Death Dis. 2022;13(1):57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Wang D, et al. “Exosomal miR-146a-5p and miR-155-5p promote CXCL12/CXCR7-induced metastasis of colorectal cancer by crosstalk with cancer-associated fibroblasts”, (in eng). Cell Death Dis. 2022;13(4):380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Peng Z, Tong Z, Ren Z, Ye M, Hu K. Cancer-associated fibroblasts and its derived exosomes: a new perspective for reshaping the tumor microenvironment. Mol Med. 2023;29(1):66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Yao L, Zhou C, Liu L, He J, Wang Y and Wang A, Cancer-associated fibroblasts promote growth and dissemination of esophageal squamous cell carcinoma cells by secreting WNT family member 5A. Mol Cell Biochem. 2025;480(6):3857–72. [DOI] [PubMed]
- 55.Ou Z, et al. “Hypoxia-Induced Senescent Fibroblasts Secrete IGF1 to Promote Cancer Stemness in Esophageal Squamous Cell Carcinoma”, (in eng). Can Res. 2025;85(6):1064–81. [DOI] [PubMed] [Google Scholar]
- 56.Ju C, et al. “Current advances and future perspectives on the functional roles and clinical implications of circular RNAs in esophageal squamous cell carcinoma: more influential than expected”, (in eng). Biomarker research. 2022;10(1):41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Gu L, et al. “circCYP24A1 facilitates esophageal squamous cell carcinoma progression through binding PKM2 to regulate NF-κB-induced CCL5 secretion”, (in eng). Mol Cancer. 2022;21(1):217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Cao X, Fang H, Zhou L. “CircNRIP1 promotes proliferation, migration and phenotypic switch of Ang II-induced HA-VSMCs by increasing CXCL5 mRNA stability via recruiting IGF2BP1”, (in eng). Autoimmunity. 2024;57(1):2304820. [DOI] [PubMed] [Google Scholar]
- 59.Li Y, Jin H, Li Q, Shi L, Mao Y, Zhao L. “The role of RNA methylation in tumor immunity and its potential in immunotherapy”, (in eng). Mol Cancer. 2024;23(1):130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Armstrong PC, et al. “Novel whole blood assay for phenotyping platelet reactivity in mice identifies ICAM-1 as a mediator of platelet-monocyte interaction”, (in eng). Blood. 2015;126(10):e11–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Dai X, et al. “Immunotherapy for targeting cancer stem cells in hepatocellular carcinoma”, (in eng). Theranostics. 2021;11(7):3489–501. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Material 2: Supplementary Fig. 1 Predicting the binding protein of circNRIP1. (A) Presence of RNA − binding domains among circNRIP1-binding protein. (B) Most detected RNA − binding domains. (C) Most detected RNA − binding motifs among circNRIP1-binding protein. (D) GO enrichment analysis of circNRIP1-binding protein.
Supplementary Material 3: Supplementary Fig. 2 Rescue experiments confirm the essential role of TRMT6 in CAF activation. (A) The expression of TRMT6 were determined by western blot. (B, C) Transwell assays detected the migration of CAFs. (D) ELISA assays detected the secretion of IL-1β and TGF-β in CAFs. (E) Western blot detected the expression of α-SMA and COL1A1 in CAFs. (PBS + shNC: mimic control-transfected fibroblasts treated with PBS, PBS + shTRMT6: TRMT6-silienced fibroblasts treated with PBS, Hypo-exo + shNC: mimic control-transfected fibroblasts co-cultured with shcircNRIP1 exosomes, Hypo-exo + shTRMT6: TRMT6-silienced fibroblasts co-cultured with shcircNRIP1 exosomes.) Data represent mean ± S.D. of three independent experiments. *P < 0.05, **P < 0.01.
Data Availability Statement
The datasets used during the present study are available from the corresponding author upon reasonable request.