ABSTRACT
Novel strategies to control diseases caused by Xylella fastidiosa that affect important crops like grapevine, olive, almond, and citrus are necessary to prevent its establishment and spread in several countries. Target-oriented functional peptides toward the pathogen are potential new compounds, and peptides directed to lipopolysaccharides (LPS) are suitable candidates. In the present work, 36 peptides, including previously reported sequences and newly synthesized lipopeptides, were studied for their bactericidal and LPS neutralization activities, as well as for their hemolytic and phytotoxic activities. In a first stage, a simple and rapid method for detecting peptide-LPS interactions based on a chromogenic endotoxin assay was developed. A linear relationship was observed between LPS neutralization and the bactericidal activity. A strong LPS interaction was related to a high bactericidal activity, whereas peptides with slight or low interaction with LPS had low bactericidal activity. A selected group of lipopeptides that strongly interacted with the LPS was highly bactericidal with moderate-to-low hemolytic and phytotoxic activity (BP389, BP473, and BP475). These lipopeptides incorporate a butanoyl group at the side chain of a lysine residue and an L- or D-phenylalanine at position 4. The interaction and cell lytic effect between BP473 and X. fastidiosa cells was confirmed by confocal microscopy experiments. These lipopeptides that target key components of the outer bacterial cell membrane are good candidates to mitigate infections by X. fastidiosa in plant hosts.
IMPORTANCE
Xylella fastidiosa is a gram-negative bacterium that affects crops of economic importance such as grapevine, olive, almond, and citrus. The lack of efficient methods to control the diseases caused by this pathogen prompts the research of novel strategies. Since lipopolysaccharides (LPS) are a major virulence factor of X. fastidiosa, the development of new compounds that target LPS constitutes a promising approach. We identified cationic lipopeptides with a strong LPS interaction, high bactericidal activity, and low toxicity. These lipopeptides can be considered good candidates to control X. fastidiosa infections.
KEYWORDS: antimicrobial peptides, lipopeptides, LPS neutralization, plant pathogens, lytic effect
INTRODUCTION
Xylella fastidiosa is a xylem-limited gram-negative plant pathogen that infects a wide range of plants. This bacterium causes significant economic losses by threatening the agricultural production of several crops worldwide and provoking high disease management costs and surveillance expenses (1–4). It infects several hundred plant species, including grapevine, olive, almond, and citrus (1, 5–7), and transmission occurs via insect vectors from Aphrophodidae or Cicadellidae families (8, 9). Following vector transmission, X. fastidiosa proliferates in the xylem, which induces its occlusion, causing a reduction of the hydraulic conductivity and resulting in severe symptoms, such as drought in the plant hosts (10, 11).
Currently, the main strategies to manage diseases caused by X. fastidiosa rely on quarantine and eradication measures as well as on vector control (12). Other methods are focused on the use of chemical compounds aimed to control the pathogen population in infected plants (13, 14) or to stimulate plant defense responses, but these strategies are not sufficiently effective (15). A promising approach to control plant pathogenic bacteria and fungi, including X. fastidiosa, is the use of natural peptides or synthetic analogs (16, 17). Within this field, the focus has nowadays shifted from the search for lytic peptides that destabilize bacterial membranes to a more refined strategy aimed at specific targets crucial for the pathogenicity of the bacteria (18).
In this context, our group has identified peptides not only with bactericidal activity against X. fastidiosa but also with antibiofilm activity or with the capacity to inhibit its motility or to stimulate the host plant defense responses (18–22). These sequences belong to several families of antimicrobial peptides developed in our group during the last two decades, with activity against other gram-negative plant pathogenic bacteria. These families include linear undecapeptides (CECMEL11 library) (23, 24), lipopeptides, peptide conjugates derived from the CECMEL11 library (25, 26), cyclic decapeptides (27), and KSL-W derivatives (28). An example is BP178, a multifunctional peptide with high bactericidal activity against X. fastidiosa and the capacity to stimulate plant defenses (18, 19). In addition, one of the best peptides against this pathogen is 1036, which displays a dual activity (bactericidal and antibiofilm) (20). We have also found that BP100 stands out for its capacity to inhibit twitching motility of X. fastidiosa (22). Moreover, FV7 activates plant defense mechanisms of host plants of X. fastidiosa (21).
Targeting lipopolysaccharides (LPS) is a promising strategy to control plant pathogens. LPS, also called endotoxins, are part of the outer membrane of gram-negative bacteria and cover around 75% of the cell surface (29). They play a crucial role in the interaction between plant pathogens and their hosts and are essential for the virulence of plant pathogenic bacteria (30, 31). It has been described that in X. fastidiosa, the O-antigen from LPS delays the plant’s innate immune recognition, allowing the pathogen to evade the initial immune response (32). LPS also have an important contribution to cell-to-cell aggregation and play a crucial role in host colonization. Therefore, LPS constitute a major virulence factor in X. fastidiosa (29).
There have already been reported peptides with the capacity to interact with the LPS of bacteria. For instance, YW12D, LBP-14, Lf(28–34), and BPI(84–99) share the capacity to neutralize LPS and, in mammals, are able to reduce inflammation caused by gram-negative bacteria (33–38). Peptides such as colistin and other polymyxins, a family of amphipathic peptides with a cyclic peptide core and a fatty acid tail, target LPS and facilitate cell lysis (39–42).
These findings prompted us to screen the antimicrobial peptides developed in our group for their capacity to neutralize LPS in X. fastidiosa. We selected peptides from the different families and evaluated their interaction with LPS as well as for their bactericidal activity against X. fastidiosa and their toxicity. A methodology to extract and quantify the LPS from X. fastidiosa was also developed together with an in vitro assay useful to identify peptides able to interact with LPS. In addition, the relationship between bactericidal activity and LPS neutralization was studied, and the best peptides were selected by their biological profile. Thus, the final goal of this work was to identify peptides able to neutralize LPS in X. fastidiosa as a suitable strategy to provide new lead compounds useful to control this pathogen.
MATERIALS AND METHODS
Synthesis of peptides
Peptides were synthesized manually on solid phase using polypropylene syringes of either 2 or 5 mL, which were equipped with a porous polyethylene disk. Removal of solvents and soluble reagents was accomplished through the application of suction. Solid-phase synthesis was performed following the standard 9-fluorenylmethoxycarbonyl (Fmoc)/tert-butyl (tBu) strategy. A Fmoc-Rink-ChemMatrix resin (0.69 mmol/g) or a Fmoc-Rink-MBHA resin (0.56 mmol/g) was employed as a solid support. The Fmoc-Rink-ChemMatrix resin was chosen for the synthesis of peptides containing more than 11 residues. Peptide elongation was performed through sequential steps of Fmoc removal and coupling of the corresponding amino acid, as previously described (43). In the case of lipopeptides, the fatty acid chain was incorporated as previously described (26). Fluorescently labeled peptides were prepared incorporating 5(6)-carboxyfluorescein (CF) at the N-terminus (BP473-CF) or at the side chain of a lysine residue (BP473-K[CF]), which was coupled as Fmoc-Lys(Mtt)-OH. Upon completion of the peptide sequence, each resulting peptidyl resin was treated with trifluoroacetic acid (TFA)/H2O/triisopropylsilane (TIS) (95:2.5:2.5). Peptidyl resins that contained tryptophan and/or arginine were subjected to treatment with TFA/H2O/TIS/thioanisole/1,2-ethandithiol/phenol (81.5:5:1:5:2.5:5). Following TFA evaporation and diethyl ether extraction, the crude peptides were purified through reverse-phase column chromatography, lyophilized, analyzed by high performance liquid chromatography (HPLC), and characterized by mass spectrometry (Table S1).
Bacterial strains and growth conditions
Experiments were conducted within officially authorized laboratories that adhered to biosafety level II + under containment conditions as prescribed by the European and Mediterranean Plant Protection Organization (44) and the European Union (45). X. fastidiosa subsp. fastidiosa IVIA 5387.2 (ST1) isolated from almond in Mallorca (Spain) was used within the experiments (19).
The strain was stored in Pierce disease broth (PD2) (46) supplemented with glycerol at 30% and maintained at −80°C. When needed, the strain was cultured in buffered charcoal yeast extract (BCYE) agar plates (47) at 28°C for 7 days. Colonies were scrapped, and cells were cultured in fresh BCYE at 28°C for 7 additional days. In liquid cultures, PD3 was used (46). Cell suspensions were prepared in sterile succinate-citrate phosphate buffer for bactericidal experiments and in sterile phosphate-buffered saline (PBS) for LPS extractions. Suspensions were adjusted to 108 cfu/mL (OD600 ~ 0.32) (20).
Extraction of LPS from X. fastidiosa
Extraction of LPS from X. fastidiosa subsp. fastidiosa IVIA 5387.2 was performed using a lipopolysaccharide Isolation Kit (MAK339, Sigma-Aldrich, United States) following the manufacturer’s recommendations. Briefly, two tubes of 5 mL of an X. fastidiosa suspension with OD600 ≥ 0.6 were centrifuged two times. Then, lysis buffer was added; lysate was sonicated with a Sonifier S450D (Branson, United States), incubated on ice, and centrifuged. Proteinase K was added, and the tube was heated at 60°C. Finally, the lysate was centrifuged, and the lysate was transferred to a fresh tube and stored at −20°C.
Purity of LPS was evaluated on a Criterion Precast gel 16.5% Tris-Tricine/Peptide 1.0 mm (Bio-Rad, United States).
Coomassie blue G-stained LPS bands were viewed using the Image Lab Software (6.1) included in the Chemidoc XRS + System (Bio-Rad, USA).
LPS-peptide neutralization
LPS-peptide neutralization assays were set up and performed using the Pierce Chromogenic Endotoxin Quant Kit (Thermo Fisher Scientific, United States) in 96-well plates following the manufacturer’s recommendations. Briefly, standard curves were prepared and validated with a lyophilized Escherichia coli (O111:B4) endotoxin standard (from 0.1 to 1 EU/mL), and the optical density at 405 nm (OD405) was measured using Varioskan Flash (Thermo Fisher Scientific, United States). The degree of interaction was measured by the ratio (R) between OD405 of treatments (ODT) and OD405 of the non-treated control (ODNTC). R was calculated following the equation: R = ([ODT − ODB]/[ODCNT − OB]), where ODB is OD405 of endotoxin-free water. Each sample was conducted in triplicate.
To identify the best reference peptide to be used in the assays, peptides with previously described LPS interaction (Table 1) were mixed at a final concentration of 25 µM with E. coli LPS at 1 EU/mL or 0.5 EU/mL, diluted in endotoxin-free water, and tested as described previously.
TABLE 1.
Sequences and references of the selected peptides
| Code | Sequencea | Reference |
|---|---|---|
| Linear undecapeptides (CECMEL11) | ||
| BP013 | FKLFKKILKVL-NH2 | 48 |
| BP015 | KKLFKKILKVL-NH2 | 48 |
| BP016 | KKLFKKILKKL-NH2 | 48 |
| BP022 | Ac-LKLFKKILKVL-NH2 | 48 |
| BP076 | KKLFKKILKFL-NH2 | 48 |
| BP100 | KKLFKKILKYL-NH2 | 23 |
| Linear undecapeptides derived from BP100 containing a D-amino acid | ||
| BP143 | KKLfKKILKYL-NH2 | 24 |
| BP144 | KKlFKKILKYL-NH2 | 24 |
| BP145 | KkLFKKILKYL-NH2 | 24 |
| BP146 | KKLFKkILKYL-NH2 | 24 |
| Lipopeptides derived from BP100 | ||
| BP375 | Ac-KKLFKKIK(COC5H11)KYL-NH2 | 25 |
| BP377 | Ac-KKLFKKILKK(COC5H11)L-NH2 | 25 |
| BP387 | Ac‐KKLFKKIK(COC3H7)KYL‐NH2 | 25 |
| BP389 | Ac-KKLFKKILKK(COC3H7)L-NH2 | 25 |
| BP393 | Ac-KK(COC11H23)LFKKILKYL-NH2 | 25 |
| BP473 | Ac-KKLfKK(COC3H7)ILKYL-NH2 | 26 |
| BP474 | Ac-KKLfKKIK(COC3H7)KYL-NH2 | 26 |
| BP475 | Ac-KKLfKKILKK(COC3H7)L-NH2 | 26 |
| BP485 | C3H7CO-KKLfKKILKYL-NH2 | 26 |
| BP490 | Ac-KKLfKKIK(COC11H23)KYL-NH2 | 26 |
| BP494 | Ac-KKLfKKK(COC5H11)LKYL-NH2 | 26 |
| BP495 | Ac-KKLfKKILKYK(COC5H11)-NH2 | 26 |
| BP496 | Ac-KKLfKKILKYK(COC3H7)-NH2 | 26 |
| BP498 | Ac-KKLfK(COC3H7)KILKYL-NH2 | 26 |
| BP499 | Ac-KKLfKKILK(COC3H7)YL-NH2 | 26 |
| BP500 | Ac-KKK(COC11H23)fKKILKYL-NH2 | 26 |
| BP501 | Ac-KKLHKKILKK(COC3H7)L-NH2 | This work |
| BP545 | Ac-K(COC3H7)KLfKKILKYL-NH2 | This work |
| BP546 | Ac-KKK(COC3H7)fKKKLKYL-NH2 | This work |
| BP547 | Ac-KKLk(COC3H7)KKILIYL-NH2 | This work |
| BP548 | Ac-KKLfKKK(COC3H7)LKYL-NH2 | This work |
| BP549 | Ac-KKK(COC3H7)fKKILKYL-NH2 | This work |
| BP550 | Ac-KKLk(COC3H7)KKILKYL-NH2 | This work |
| Cyclic decapeptide (CYCLO10) | ||
| BPC098W | c(LLKKKWKKLQ) | 27 |
| Peptide analog of KSL-W | ||
| BP442 | KKVVFWVKFk-NH2 | 28 |
| Peptide conjugate derived from BP100 | ||
| BP178 | KKLFKKILKYLAGPAGIGKFLHSAKKDEL-OH | 49 |
| Peptides described with LPS-neutralizing activity | ||
| BPI(84-99) | NIKISGKWKAQKRFLK-NH2 | 33, 34 |
| LBP-14 | RVQGRWKVRASFFK-NH2 | 35, 36 |
| Lf (28–34) | RKVRGPP-NH2 | 37, 38 |
| YW12D | YVKLWRMIKFIR-NH2 | 38 |
| CF-labeled BP473 analogs | ||
| BP473-CF | CF-KKLfKK(COC3H7)ILKYL-NH2 | This work |
| BP473-K(CF) | Ac-K(CF)KKLfKK(COC3H7)ILKYL-NH2 | This work |
Lowercase letters indicate a D-amino acid; Ac = acetyl.
To adjust peptide concentration for the LPS-peptide assays, YW12D peptide (used as reference) at different final concentrations (50, 100, 150, and 200 µM) was mixed with E. coli LPS at 0.5 EU/mL. Upon selection of the desired concentration, selected peptides were mixed at a final concentration of 150 µM with E. coli LPS at 0.5 EU/mL and tested. To ensure the peptides do not shield the OD405 signal through some polar or similar residues, most of them were also evaluated without LPS.
For comparison purposes with E. coli LPS, some peptides were tested at a final concentration of 150 µM using LPS extracted from X. fastidiosa subsp. fastidiosa IVIA 5387.2. In this case, the LPS concentration was adjusted following kit recommendations by preparing serial dilutions and comparing to the standard curve made with E. coli (O111:B4) endotoxin. Each sample was conducted in triplicate.
Bactericidal activity
The bactericidal activity of peptides was assessed by a contact test coupled with viable PCR (v-qPCR) as previously described (50). The v-qPCR permits the quantification of viable cells, discarding dead or non-viable cells, thanks to PMA, a nucleic acid binding dye. Sensitivity and amplification efficiency of the v-qPCR and standard curves were previously evaluated and set up for the studied strain (20).
Peptides were solubilized in sterile Milli-Q water to a stock concentration of 1 mM and filter sterilized through a 0.22 µM pore size filter. Briefly, the bactericidal activity of the selected peptides was determined by a 3 h contact test at room temperature with a 107 cfu/mL X. fastidiosa suspension. PMA was added, and samples were incubated and processed following previously described protocols (50). The reduction of viability was obtained as described previously (20). Peptides were tested generally at a final concentration of 50 µM. Highly active peptides (those causing a reduction in viability ≥2 logs) were further tested at 12.5 and 3.1 µM to better characterize their bactericidal activity. BP377, BP393, and BP473 were tested at 3.1, 6.25, 12.5, 25, and 50 µM to study their dose-response effect. All treatments were performed in triplicate.
Phytotoxicity
Peptides were evaluated for their phytotoxicity as previously described (49). Tobacco plants (Nicotiana tabacum) were grown from seed in a glasshouse between 20 and 30 days. A total of 100 µL of each peptide at 150 µM was infiltrated into the mesophylls of fully expanded tobacco leaves. For each peptide and concentration, at least three leaves randomly distributed along distinct plants were similarly infiltrated. For the purposes of control, infiltrations with water (negative control) or with the highly phytotoxic melittin peptide (positive control) were performed at the same concentration. After 48 h, the phytotoxicity was quantified as the lesion diameter.
Hemolytic activity
The hemolytic activity of peptides was evaluated by determining hemoglobin release from erythrocyte suspensions of horse blood (5% [vol/vol]; Thermo Fisher Scientific, Spain) as previously described (23). In brief, the peptides were dissolved in a TRIS buffer and subsequently mixed with purified horse erythrocytes that were diluted 10-fold. The peptides were tested at final concentrations of 50 and 250 µM. The hemolysis percentage (H) was determined using the mathematical equation: H = 100 × ([Op − Ob]/[Om − Ob]), where Op is the density for a given compound concentration, Ob is for the buffer, and Om is for the melittin-positive control.
Confocal microscopy
Cell suspensions of X. fastidiosa at 107 cfu/mL were treated with CF and CF-labeled peptides and incubated at different times (30 s, 5 min, 10 min, and 30 min). Then, adapted from previously described protocols (51–53), paraformaldehyde was added for fixation, reaching a final concentration of 2%, incubated at room temperature for 20 min followed by two washes with PBS and stored at 4°C until cell observation.
Confocal microscopy was performed with a NIKON Ti Eclipse motorized inverted microscope with a NIKON A1R confocal module (NIKON, Japan; Research Technical Services from the University of Girona). Images were processed and compared with the software NIS elements v4.10 (NIKON, Japan).
Data analysis
Statistical significance in the assays comparing techniques for detecting peptide-LPS interactions was assessed using a two-way analysis of variance (ANOVA), considering treatments (peptides) and techniques (LPS from X. fastidiosa or E. coli) as the two factors. The significance of the effect of peptides in LPS neutralization and their bactericidal activity against X. fastidiosa was evaluated by one-way ANOVA. Means were separated according to Duncan’s test at a p-value of <0.05 (IBM Statistics for Windows, Version 29.0 released in 2023 by IBM Corp, Armonk NY, United States).
The relationship between the LPS neutralization capacity and bactericidal activity was analyzed using a simple linear regression model and was fitted with MATLAB (MathWorks, Natick, MA). The software was used to calculate the regression line parameters, the coefficient of determination (R2), and the p-value for the test of whether the slope differs significantly from zero.
In lipopeptides, for every variable studied (bactericidal activity, LPS neutralization, hemolysis, and effect on tobacco plants), a threshold was arbitrarily determined to separate peptides in different biological activity profiles: 2.0 LogN0/N for bactericidal activity, 0.25 ratio for LPS neutralization, 10% for hemolytic activity, and 7 mm for the effect on tobacco plants.
For dose-response assays, the half-maximal effective concentration (EC50) of the peptides was calculated using EC50 calculator with the two-parameter model (ATT Bioquest, Pleasanton, USA).
RESULTS
Selection and synthesis of peptides
The 36 peptides selected for this study are shown in Table 1. They belong to the following peptide families: (i) linear undecapeptides from CECMEL11 library; (ii) linear undecapeptides derived from BP100 containing a D-amino acid; (iii) lipopeptides derived from BP100, including seven sequences not previously described; (iv) a cyclic decapeptide from the CYCLO10 library; (v) a peptide analog of KSL-W; (vi) a peptide conjugate derived from BP100. In addition, four previously reported peptides from other sources, with the capacity to neutralize LPS, were also selected as positive controls. Furthermore, lipopeptide BP473 was fluorescently labeled at the N-terminus (BP473-CF) or at the side chain of an additional Lys incorporated at the N-terminus (BP473-K[CF]). All peptides were manually synthesized and purified, being obtained in HPLC purities >90% except for BP015, BP143, BP144, and BP146 (purities >80%). Peptide identity was confirmed by high-resolution mass spectrometry (Table S1).
Activity of peptides on LPS neutralization
The Pierce Chromogenic Endotoxin Quant Kit was first tested to ensure the detection of free LPS and the achievement of a reduction in the ratio of free LPS in the presence of LPS-neutralizing peptides. To set up the LPS neutralization assay, a standard curve was validated following the manufacturer’s recommendations, obtaining the equation OD405 = 1.039X – 0.034, where X is LPS concentration expressed as EU/mL, and with R2 = 0.9924.
Peptides with previously described LPS neutralization activity (YW12D, LBP-14, Lf[28–34), and BPI[84–99]) were tested with E. coli 0111:B4 LPS at 0.5 and 1 EU/mL (Fig. S1). A lower OD405 ratio between peptide-treated samples and NTC was observed at 0.5 EU/mL, and this concentration was selected for the next assays. The peptide that showed the lowest OD405 ratio was YW12D, which was chosen for the next experiment to determine in more detail the optimal peptide concentration to perform the LPS neutralization assays. Thus, YW12D was evaluated at concentrations ranging from 25 to 200 µM (Fig. S2). It was considered that the OD405 ratio obtained at 150 µM was adequate to carry out the LPS neutralization assays for screening the 36 selected peptides (Table 1).
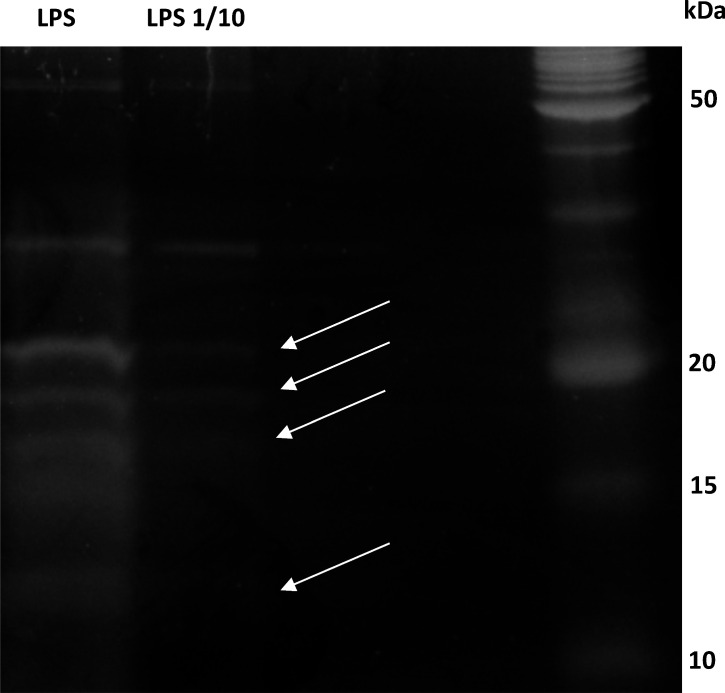
Next, LPS from X. fastidiosa subsp. fastidiosa IVIA 5387.2 were extracted and partially purified by electrophoresis (Fig. 1). Bands corresponding to LPS were identified. The resulting extracts were highly concentrated, and their dilution was required. These extracts were quantified using the standard curve obtained from the E. coli assays. An equivalent LPS concentration of 0.4 EU/mL for a 10−7 dilution of the extracts was found as the most appropriate for the assays.
Fig 1.
Electrophoresis gel showing extracts of LPS from X. fastidiosa subsp. fastidiosa. Direct LPS extract (lane 1), a 1/10 dilution (lane 2), and the molecular marker (lane 3). White arrows correspond to expected bands with LPS weights according to the extraction kit.
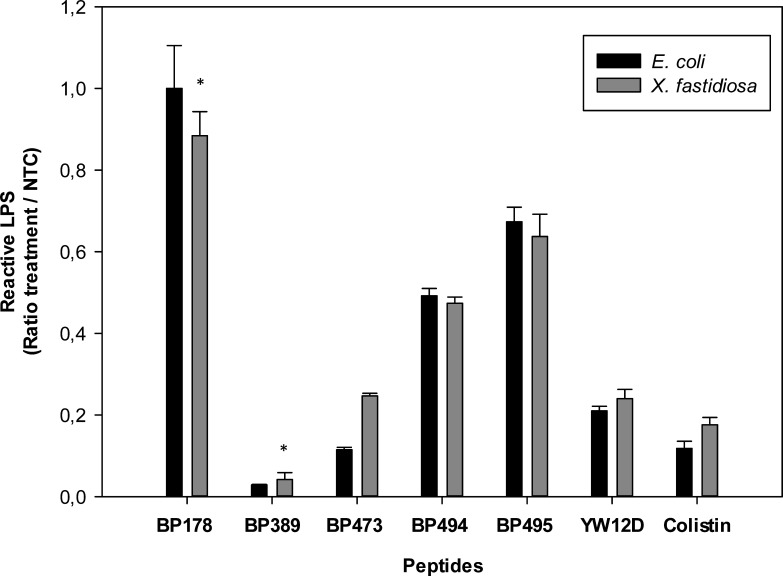
Once the conditions of the test were set up, the LPS neutralization capacity of seven peptides (colistin, YW12D, BP178, BP389, BP473, BP494, and BP495) was analyzed with LPS from both E. coli 0111:B4 and X. fastidiosa subsp. fastidiosa IVIA 5387.2 (Fig. 2). Colistin and YW12D were used as LPS-neutralization reference peptides.
Fig 2.
Effect of selected peptides at 150 µM on LPS neutralization. Bars correspond to assays using lyophilized LPS from E. coli 0111:B4 (black bars) or X. fastidiosa subsp. fastidiosa IVIA 5387.2 (gray bars) both at 0.5 EU/mL. Values are the means of three replicates, and error bars represent the confidence interval α = 0.05. Significant differences between techniques within each treatment according to a two-way ANOVA are indicated by asterisks (P < 0.05).
In the two-way ANOVA assessing the effect of treatment and bacterial LPS (X. fastidiosa or E. coli) on LPS neutralization, the main effect of treatment was highly significant (F = 1,020.253, P < 0.001), confirming that different peptides have clearly distinct LPS-binding capacities. The main effect of bacterium LPS was not significant (F = 0.000, P = 0.990), suggesting that, overall, the two bacterial LPS systems responded similarly to the treatments. However, the interaction effect between treatment and bacterial LPS was statistically significant (F = 15.382, P < 0.001), indicating that in some cases, the response to a given peptide depended on the bacterial LPS system used.
To explore this further, we performed univariate analyses of simple effects (per treatment) to compare responses between the two bacterial LPS models. These analyses showed that only two treatments (BP473 and BP178) presented statistically significant differences between bacterial LPS systems (p < 0.001). For the remaining treatments, no significant differences were observed (p > 0.05). This indicates that, in most cases, the two models produce comparable LPS neutralization profiles. Despite the differences observed in the interaction effect, both techniques (X. fastidiosa and E. coli) are able to discriminate peptides with very high, high, moderate, and low LPS-binding capacity. Therefore, it was concluded that peptide-LPS neutralization assays performed with LPS extracted from X. fastidiosa were equivalent to those carried out using the commercial kit containing E. coli LPS. Thus, the next experiments were run with this kit.
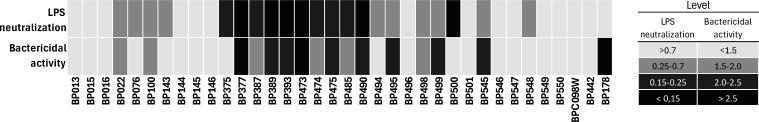
The LPS interaction of the 36 antimicrobial peptides and the four reference peptides with previously described LPS interaction was analyzed with the above kit (Table S2). Peptides were classified into four different groups (Fig. 3). The first group included six lipopeptides (BP377, BP389, BP393, BP473, BP490, and BP500) with a very high LPS neutralization level (a ratio <0.15). The second group comprised five lipopeptides (BP375, BP387, BP474, BP475, and BP485) with a high activity (a ratio between 0.15 and 0.25). The third group included 10 peptides with a moderate activity (a ratio between 0.25 and 0.7), and the rest of the peptides showed low LPS neutralization (a ratio higher than 0.7). Peptides previously described with LPS neutralization activity exhibited moderate to high levels of LPS interaction (a ratio between 0.21 and 0.30).
Fig 3.
LPS neutralization and bactericidal activity of the 36 peptides screened.
Bactericidal activity of peptides
Bactericidal activity of the 36 peptides was determined at a concentration of 50 µM. The peptides were classified into four statistically different groups (Fig. 3; Table S2). Peptide BP178 and lipopeptides BP377 and BP473 showed very high bactericidal activity (>2.5 log), and lipopeptides BP389, BP393, BP475, BP490, BP495, BP499, and BP545 displayed high bactericidal activity (between 2.0 and 2.5 log). Six peptides exhibited a moderate activity (between 1.5 and 2.0 log), and the rest of the sequences were low active (<1.5 log). The most active peptides were assayed at 12.5 and 3.1 µM, and notably, BP178, BP473, BP490, and BP499 showed high activity at 12.5 µM.
The bactericidal activity of the CF-labeled BP473 analogs (BP473-CF and BP473-K[CF]) was studied at 25 µM (Table S2). BP473-K(CF) was the most active analog (1.78 ± 0.053 log) and was selected to perform the confocal microscopy assays.
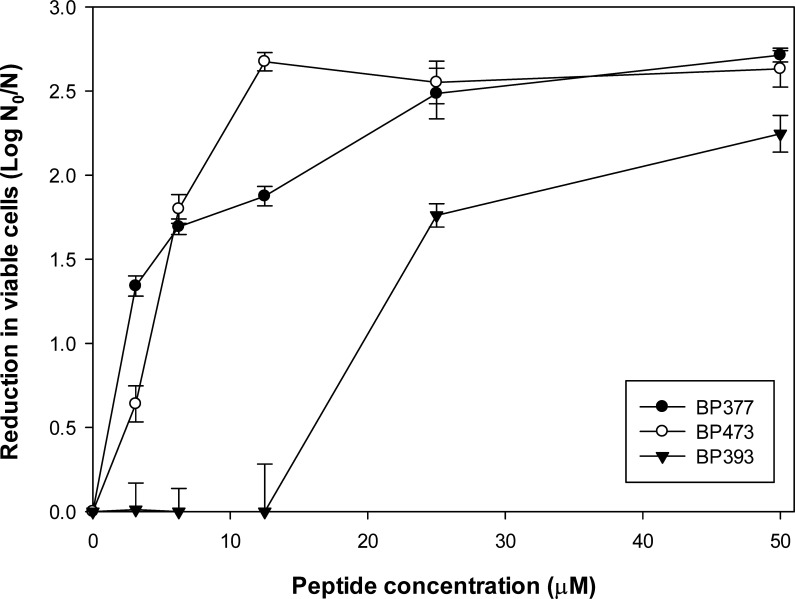
Three selected peptides with a high LPS-neutralizing level (BP377, BP393, and BP473) were studied for bactericidal activity in more detail at concentrations ranging from 3.1 to 50 µM, indicating a saturation relationship (Fig. 4). The median effective concentration (EC50) for the peptides was low for BP377 and BP473 (3.67 µM ± 0.23 and 4.75 µM ± 0.06, respectively), indicating high activity, compared to BP393 (20.11 µM ± 1.16).
Fig 4.
Dose-response relationship between the reduction in viability of X. fastidiosa subsp. fastidiosa IVIA 5387.2 and peptide concentration after 3-hour contact with selected peptides. N0 is 107 cfu. Values are the means of three replicates, and error bars represent confidence interval (α = 0.05).
Relationship between bactericidal activity and LPS neutralization
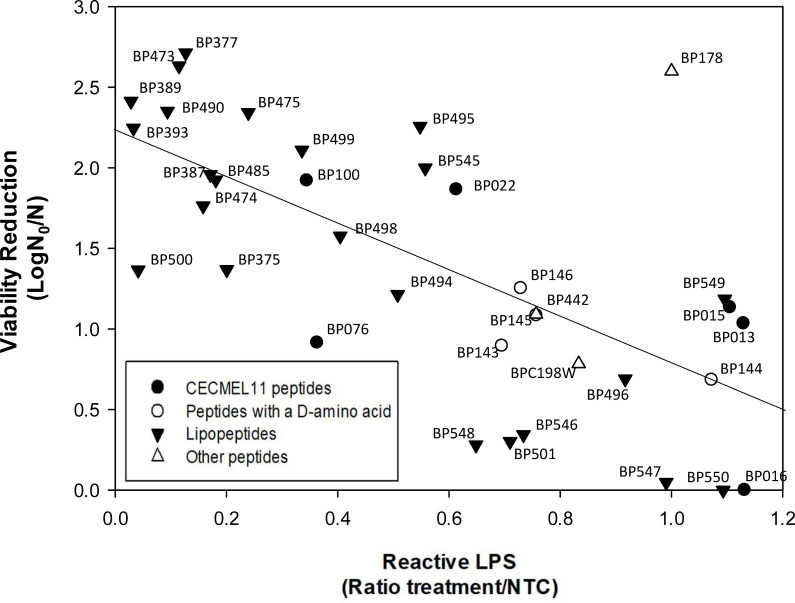
Among the 36 peptides studied, a moderate linear relationship (R2 = 0.582, P < 0.01) was observed between the bactericidal activity and the capacity to neutralize LPS (Fig. 5), with lipopeptides showing high LPS neutralization capacity generally exhibiting higher bactericidal activity. Particularly, the best sequences with both activities were lipopeptides BP377, BP389, BP393, BP473, and BP490, which displayed viability reduction levels >2 log and a ratio of reactive LPS <0.2.
Fig 5.
Relationship between bactericidal activity of peptides against X. fastidiosa and LPS neutralization. The regression equation is y = −1.63X + 2.26; R2 = 0.582 and P-value = 1.57 × 10−7. BP178 has been excluded from the regression as an outlier.
Biological profile of lipopeptides
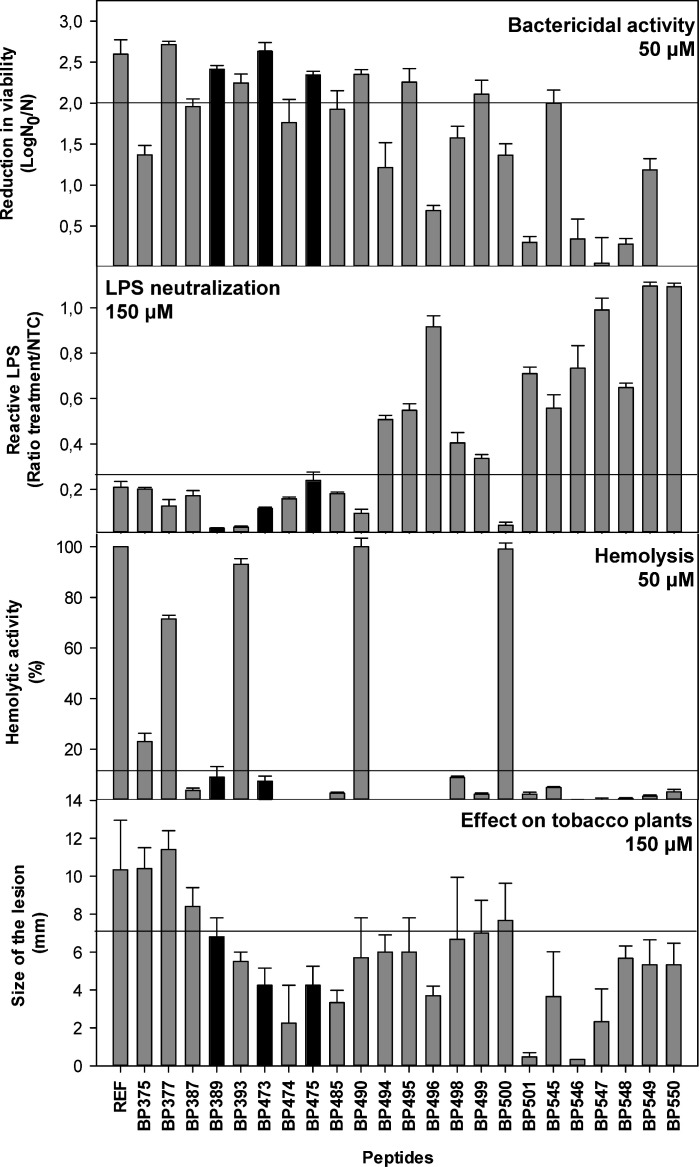
The biological activity profile of the 23 lipopeptides was further studied because most of them showed a good relationship between bactericidal activity and LPS neutralization. They were screened for their hemolytic activity and phytotoxicity (Fig. 6). Eighteen lipopeptides showed a hemolytic activity ≤10% at 50 µM (Table S3). Nineteen lipopeptides showed lesions ≤7 mm at 150 µM when infiltrated into the mesophylls of tobacco leaves (Table S3).
Fig 6.
Biological activity of lipopeptides. Bactericidal activity against X. fastidiosa, LPS neutralization, hemolysis, and effect on tobacco leaves. Values are the means of three replicates, and error bars represent the confidence interval (α = 0.05). REF indicates reference compounds: BP178 at 25 µM for bactericidal activity, YW12D for LPS neutralization, and melittin for hemolysis and effect on tobacco leaves. Horizontal lines represent arbitrarily defined thresholds used to select peptides based on their biological activity profiles. Black bars highlight peptides with high bactericidal activity, high LPS neutralization, and low toxicity.
The biological profile of the 23 lipopeptides was analyzed by comparing their bactericidal activity, LPS neutralization, hemolytic activity, and leaf infiltration effect (Fig. 6). To classify these lipopeptides in groups, the following arbitrary threshold values for each of these activities were established: a reduction of viability ≥2 log (high bactericidal activity), a ratio of reactive LPS ≤0.25 (high LPS neutralization), ≤10% hemolysis (low hemolytic activity), and diameter of leave lesion ≤7 mm (low phytotoxicity). According to these threshold values, five groups were defined: (i) lipopeptides with a high LPS neutralization, high bactericidal activity, and low toxicity (BP389, BP473, and BP475); (ii) lipopeptides with high bactericidal and LPS activity but highly toxic (BP377, BP393, and BP490); (iii) lipopeptides with high LPS neutralization but low bactericidal activity (BP375, BP387, BP474, BP485, and BP500); (iv) lipopeptides with high bactericidal activity but low LPS neutralization (BP495 and BP499); and (v) lipopeptides with low bactericidal and LPS activity and low toxicity (BP494, BP496, BP498, BP501, BP545, BP546, BP547, BP548, BP549, and BP550). The best biological profile was considered for the peptides BP389, BP473, and BP475.
Interaction between BP473-K(CF) and X. fastidiosa
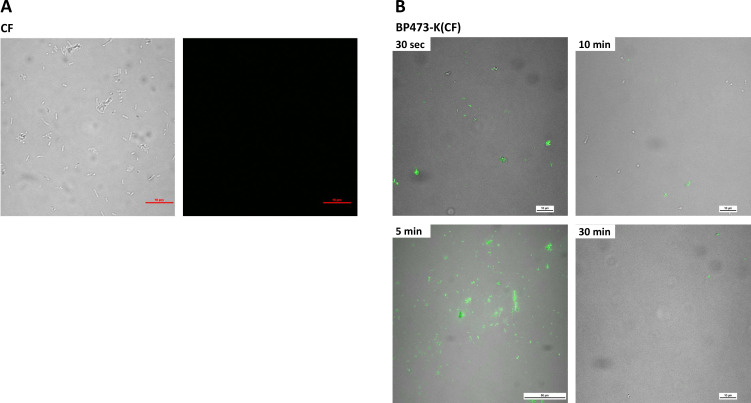
The interaction between the labeled BP473-K(CF) and X. fastidiosa was studied using a contact test, and fixed samples were observed with confocal microscopy (Fig. 7). This experiment was also performed using CF alone for comparison purposes. After 30 s, cells treated with BP473-K(CF) at 3.1 µM showed a fluorescence signal, and no fluorescence was observed in cells treated with CF. After 5 min, fluorescence was detected for most of the cells treated with BP473-K(CF), indicating a peptide-cell interaction. At longer contact times, there was a decrease in fluorescence, probably due to the lytic effect of the peptide.
Fig 7.
Microscopy images after a contact test with CF or BP473-K(CF) and X. fastidiosa. (A) On the right, confocal fluorescence microscopy channel (excitation at 492 nm and emission at 517 nm), and on the left, a merge of confocal and interference contrast channels. Image at 30 s contact test with CF and X. fastidiosa. Treatments were performed at 3.1 µM. (B) Merge of confocal and interference contrast channels. Images after different contact times with BP473-K(CF). Treatments were performed at 5 µM.
DISCUSSION
The use of peptides to control X. fastidiosa is a research topic that has begun to be explored in recent years, and several peptides with bactericidal effects against this bacterium have been identified (19, 20, 54). Among them, BP178 is highlighted because, in addition to regulating various plant defense mechanisms, it also possesses bactericidal properties due to a lytic effect (19). The present work identified peptides able to interact with LPS of X. fastidiosa, which are a major virulence factor of this bacterium. These peptides were selected from previous libraries developed in our group with activity against other gram-negative plant pathogenic bacteria (24–26, 28, 48, 55).
The study of the interaction between peptides and LPS requires the use of a reliable method. Several techniques have previously been used to study the capacity of peptides to neutralize LPS, including surface plasmon resonance (56) or the detection of tumor necrosis factor-alpha in human monocytes using enzyme-linked immunosorbent assay techniques (57). In the present work, we used a Limulus amebocyte lysate assay because it is a simple and rapid method to detect LPS-protein interaction (58), thus permitting the screening of a large collection of peptides. This method was assayed and validated in the present study following the manufacturer’s recommendations with peptides YW12D, LBP-14, Lf (28–34), and BPI(84–99), which have been reported to reduce LPS-induced inflammation in mammal cells both in vitro and in vivo (33, 34, 59, 60).
We observed that the LPS fraction obtained from X. fastidiosa subsp. fastidiosa IVIA 5387.2 cells was not pure enough in contrast to the commercial E. coli (O111) LPS. Additional stages of purification, like the hot-phenol method followed by dialysis, would provide LPS of greater purity (32, 61, 62), but they were not attempted in the present work. Fortunately, when the functionality of the X. fastidiosa extracts was checked, it was concluded that performing peptide-LPS neutralization assays with commercial E. coli endotoxins is equivalent to performing the test with X. fastidiosa LPS.
The activity of peptides on LPS neutralization revealed that the presence of a fatty acid chain in the cationic amphipathic peptide favors LPS-peptide interaction. In fact, among the 36 sequences tested, those with the highest capacity to neutralize LPS were cationic lipopeptides. Notably, six of these lipopeptides (BP377, BP389, BP393, BP473, BP490, and BP500) showed a ratio of reactive LPS close to zero. Similar to colistin and other polymyxin antibiotics (41, 42), these results suggested that there is an affinity between the positively charged moiety of peptides and the negatively charged phosphate groups of the lipid A, component of LPS. The hydrophobicity of the fatty acid chain might facilitate the interaction of the peptide sequence with the LPS. However, no correlation between the length or the position of the fatty acid chain and this activity could be found. In addition, the presence of a D-Phe at position 4 did not influence the interaction of the lipopeptide with LPS.
Among the six lipopeptides with a very high LPS neutralization level, five sequences (BP377, BP389, BP393, BP473, and BP490) also exhibited a high bactericidal activity against X. fastidiosa. Notably, BP473 showed a reduction of cell viability >2.5 log at 12.5 µM. In addition, the bactericidal activity of this lipopeptide followed a dose-dependent relationship with a saturation pattern typical of bactericidal peptides (63). Confocal microscopy experiments confirmed the interaction and cell lytic effect of BP473 because most X. fastidiosa cells showed fluorescence when treated with fluorescently labeled BP473 at short contact test times, whereas a marked decrease in fluorescence was observed at longer times.
The biological profile of these lipopeptides indicates that the fatty acid chain may have a role in LPS neutralization and, in consequence, in the bactericidal activity against X. fastidiosa. It is known that the incorporation of a fatty acid into a peptide sequence clearly modifies its hydrophobicity, which facilitates its anchoring to the membranes of gram-negative bacteria, causing their disruption (64–66). Accordingly, in the case of colistin, after the electrostatic interaction with lipid A and subsequent insertion of the fatty acid chain, membrane disruption occurs (41, 42).
Interestingly, peptide BP178 did not fit into the general relationship between bactericidal activity and LPS interaction and did not interact with LPS. This finding confirms that there are lytic peptides with high bactericidal activity that may act by different mechanisms to disturb the bacterial cell membrane and that are independent of LPS interaction, such as the case of the undecapeptide BP100 (67) or the cyclic decapeptide BPC194 (68).
Lipopeptides that exhibited the best biological activity profile in terms of high LPS neutralization, high bactericidal activity, and low toxicity were BP389, BP473, and BP475 (Fig. 8). These peptides incorporate a butanoyl chain at positions 10, 6, and 10, respectively. In addition, BP473 and BP475 contain a D-Phe at position 4. Interestingly, these lipopeptides also stood up for their high antibacterial activity against other gram-negative plant pathogenic bacteria, including Erwinia amylovora, Pseudomonas syringae pv. syringae, P. syringae pv. actinidiae, Xanthomonas arboricola pv. pruni, X. fragariae, and X. axonopodis pv. vesicatoria (25, 26). Furthermore, these three lipopeptides were low hemolytic and phytotoxic (<10% hemolysis at 50 µM, leave lesion diameter ≤7 mm at 150 µM).
Fig 8.
General sequence of lipopeptides with the best biological activity profile studied in this work: BP389 (Phe4, Lys6, and Lys10[COC3H7]), BP473 (D-Phe4, Lys6[COC3H7], and Tyr10), and BP475 (D-Phe4, Lys6, and Lys10[COC3H7]). Lowercase letters correspond to a D-amino acid. Asterisk corresponds to a lysine incorporating a butanoyl group. Ac stands for acetyl.
In summary, a collection of lipopeptides with the capacity to neutralize LPS was identified. Notably, those with the highest LPS neutralization were also highly active against X. fastidiosa and low toxic. Therefore, these compounds constitute promising candidates to mitigate infections caused by X. fastidiosa in host plants. Studies to decipher structural-activity relationships and their mechanism of action are ongoing.
ACKNOWLEDGMENTS
The authors acknowledge Research Technical Services from the University of Girona (Girona, Spain) for the mass spectrometry analysis and for confocal microscopy. We are thankful to Gemma Roselló and Héctor Saravia for participating in various tasks during the preparation and maintenance of the plant material.
This work was supported by grants from the European Commission BeXyl (grant 101060593) and from the Spain Ministerio de Ciencia, Innovación y Universidades (MCIU; TED2021-130110B-C43; PID2022-140040OB-C21 and C22/MICIU/AEI/10.13039/501100011033/FEDER, UE). P.C.-F. was the recipient of a research grant from Spain MCIU (Ref. FPU20/07152).
P.C.-F.: main experiments and manuscript writing. L.M. and J.L.: experiments collaboration. E.M., M.P., and L.F.: research design, funding, laboratory facilities, and writing.
Contributor Information
Lidia Feliu, Email: lidia.feliu@udg.edu.
Emilio Montesinos, Email: emilio.montesinos@udg.edu.
Gladys Alexandre, The University of Tennessee Knoxville, Knoxville, Tennessee, USA.
SUPPLEMENTAL MATERIAL
The following material is available online at https://doi.org/10.1128/aem.00734-25.
Analysis and characterization data of peptides; biological activity of peptides; and HPLC, ESI-MS, and HRMS of purified lipopeptides.
ASM does not own the copyrights to Supplemental Material that may be linked to, or accessed through, an article. The authors have granted ASM a non-exclusive, world-wide license to publish the Supplemental Material files. Please contact the corresponding author directly for reuse.
REFERENCES
- 1. Purcell A. 2013. Paradigms: examples from the bacterium Xylella fastidiosa. Annu Rev Phytopathol 51:339–356. doi: 10.1146/annurev-phyto-082712-102325 [DOI] [PubMed] [Google Scholar]
- 2. Baker R, Bragard C, Caffier D, Candresse T, Gilioli G. 2015. Scientific opinion on the risks to plant health posed by Xylella fastidiosa in the EU territory, with the identification and evaluation of risk reduction options. EFSA Journal 13:3989. doi: 10.2903/j.efsa.2015.3989 [DOI] [Google Scholar]
- 3. Olmo D, Nieto A, Borràs D, Montesinos M, Adrover F, Pascual A, Gost PA, Quetglas B, Urbano A, García J de D, Velasco-Amo MP, Olivares-García C, Beidas O, Juan A, Marco-Noales E, Gomila M, Rita J, Moralejo E, Landa BB. 2021. Landscape epidemiology of Xylella fastidiosa in the Balearic islands. Agronomy 11:473. doi: 10.3390/agronomy11030473 [DOI] [Google Scholar]
- 4. Burbank LP. 2022. Threat of Xylella fastidiosa and options for mitigation in infected plants. CABI Rev 17:021. doi: 10.1079/cabireviews202217021 [DOI] [Google Scholar]
- 5. Trkulja V, Tomić A, Iličić R, Nožinić M, Milovanović TP. 2022. Xylella fastidiosa in Europe: from the introduction to the current status. Plant Pathol J 38:551–571. doi: 10.5423/PPJ.RW.09.2022.0127 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Gibin D, Pasinato L, Delbianco A, European Food Safety Authority (EFSA) . 2023. Update of the Xylella spp. host plant database – systematic literature search up to 31 December 2022. EFS2 21:e08061. doi: 10.2903/j.efsa.2023.8061 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Lago C, Giménez-Romero À, Morente M, Matías MA, Moreno A, Fereres A. 2023. Degree-day-based model to predict egg hatching of Philaenus spumarius (Hemiptera: Aphrophoridae), the main vector of Xylella fastidiosa in Europe. Environ Entomol 52:350–359. doi: 10.1093/ee/nvad013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Marcus IM, White D, Backus EA, Walker SL, Roper MC. 2022. Fluid dynamic simulations at the interface of the blue-green sharpshooter functional foregut and grapevine xylem sap with implications for transmission of Xylella fastidiosa. PLoS One 17:e0265762. doi: 10.1371/journal.pone.0265762 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Boukhris-Bouhachem S, Souissi R, Abou Kubaa R, El Moujabber M, Gnezdilov V. 2023. Aphrophoridae as potential vectors of Xylella fastidiosa in Tunisia. Insects 14:119. doi: 10.3390/insects14020119 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Sun Q, Sun Y, Walker MA, Labavitch JM. 2013. Vascular occlusions in grapevines with Pierce’s disease make disease symptom development worse. Plant Physiol 161:1529–1541. doi: 10.1104/pp.112.208157 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Giménez-Romero À, Moralejo E, Matías MA. 2023. A compartmental model for Xylella fastidiosa diseases with explicit vector seasonal dynamics. Phytopathology 113:1686–1696. doi: 10.1094/PHYTO-11-22-0428-V [DOI] [PubMed] [Google Scholar]
- 12. Sicard A, Zeilinger AR, Vanhove M, Schartel TE, Beal DJ, Daugherty MP, Almeida RPP. 2018. Xylella fastidiosa: insights into an emerging plant pathogen. Annu Rev Phytopathol 56:181–202. doi: 10.1146/annurev-phyto-080417-045849 [DOI] [PubMed] [Google Scholar]
- 13. Amanifar N, Taghavi M, Salehi M. 2016. Xylella fastidiosa from almond in Iran: overwinter recovery and effects of antibiotics. Phytopathol Mediterr 55:337–345. doi: 10.14601/Phytopathol_Mediterr-17682 [DOI] [Google Scholar]
- 14. Scortichini M, Chen J, Caroli M, Dalessandro G, Pucci N, Modesti V, L’Aurora A, Petriccione M, Zampella L, Mastrobuoni F, Migoni D, Del Coco L, Girelli CR, Piacente F, Cristella N, Marangi P, Laddomada F, Cesare M, Cesari G, Fanizzi FP, Loreti S. 2018. A zinc, copper and citric acid biocomplex shows promise for control of Xylella fastidiosa subsp. pauca in olive trees in Apulia region (southern Italy). Phytopathol Mediterr 57:48–72. doi: 10.14601/Phytopathol_Mediterr-21985 [DOI] [Google Scholar]
- 15. Bragard C, Dehnen‐Schmutz K, Di Serio F, Gonthier P, Jacques M, Jaques Miret JA, Justesen AF, MacLeod A, Magnusson CS, Milonas P, et al. 2019. Effectiveness of in planta control measures for Xylella fastidiosa. EFS2 17:5666. doi: 10.2903/j.efsa.2019.5666 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Badosa E, Planas M, Feliu L, Montesinos L, Bonaterra A, Montesinos E. 2022. Synthetic peptides against plant pathogenic bacteria. Microorganisms 10:1784. doi: 10.3390/microorganisms10091784 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Montesinos E. 2023. Functional peptides for plant disease control. Annu Rev Phytopathol 61:301–324. doi: 10.1146/annurev-phyto-021722-034312 [DOI] [PubMed] [Google Scholar]
- 18. Moll L, Baró A, Montesinos L, Badosa E, Bonaterra A, Montesinos E. 2022. Induction of defense responses and protection of almond plants against Xylella fastidiosa by endotherapy with a bifunctional peptide. Phytopathology 112:1907–1916. doi: 10.1094/PHYTO-12-21-0525-R [DOI] [PubMed] [Google Scholar]
- 19. Baró A, Mora I, Montesinos L, Montesinos E. 2020. Differential susceptibility of Xylella fastidiosa strains to synthetic bactericidal peptides. Phytopathology 110:1018–1026. doi: 10.1094/PHYTO-12-19-0477-R [DOI] [PubMed] [Google Scholar]
- 20. Moll L, Badosa E, Planas M, Feliu L, Montesinos E, Bonaterra A. 2021. Antimicrobial peptides with antibiofilm activity against Xylella fastidiosa. Front Microbiol 12:753874. doi: 10.3389/fmicb.2021.753874 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Moll L, Giralt N, Planas M, Feliu L, Montesinos E, Bonaterra A, Badosa E. 2024. Prunus dulcis response to novel defense elicitor peptides and control of Xylella fastidiosa infections. Plant Cell Rep 43:190. doi: 10.1007/s00299-024-03276-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Moll L, Badosa E, De La Fuente L, Montesinos E, Planas M, Bonaterra A, Feliu L. 2025. Mitigation of almond leaf scorch by a peptide that inhibits the motility of Xylella fastidiosa. Plant Dis 109:327–340. doi: 10.1094/PDIS-07-24-1414-RE [DOI] [PubMed] [Google Scholar]
- 23. Badosa E, Ferre R, Planas M, Feliu L, Besalú E, Cabrefiga J, Bardají E, Montesinos E. 2007. A library of linear undecapeptides with bactericidal activity against phytopathogenic bacteria. Peptides 28:2276–2285. doi: 10.1016/j.peptides.2007.09.010 [DOI] [PubMed] [Google Scholar]
- 24. Güell I, Cabrefiga J, Badosa E, Ferre R, Talleda M, Bardají E, Planas M, Feliu L, Montesinos E. 2011. Improvement of the efficacy of linear undecapeptides against plant-pathogenic bacteria by incorporation of D-amino acids. Appl Environ Microbiol 77:2667–2675. doi: 10.1128/AEM.02759-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Oliveras À, Baró A, Montesinos L, Badosa E, Montesinos E, Feliu L, Planas M. 2018. Antimicrobial activity of linear lipopeptides derived from BP100 towards plant pathogens. PLoS One 13:e0201571. doi: 10.1371/journal.pone.0201571 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Oliveras À, Moll L, Riesco-Llach G, Tolosa-Canudas A, Gil-Caballero S, Badosa E, Bonaterra A, Montesinos E, Planas M, Feliu L. 2021. D-Amino acid-containing lipopeptides derived from the lead peptide BP100 with activity against plant pathogens. Int J Mol Sci 22:6631. doi: 10.3390/ijms22126631 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Camó C, Torné M, Besalú E, Rosés C, Cirac AD, Moiset G, Badosa E, Bardají E, Montesinos E, Planas M, Feliu L. 2017. Tryptophan-containing cyclic decapeptides with activity against plant pathogenic bacteria. Molecules 22:1817. doi: 10.3390/molecules22111817 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Camó C, Bonaterra A, Badosa E, Baró A, Montesinos L, Montesinos E, Planas M, Feliu L. 2019. Antimicrobial peptide KSL-W and analogues: promising agents to control plant diseases. Peptides 112:85–95. doi: 10.1016/j.peptides.2018.11.009 [DOI] [PubMed] [Google Scholar]
- 29. Clifford JC, Rapicavoli JN, Roper MC. 2013. A rhamnose-rich O-antigen mediates adhesion, virulence, and host colonization for the xylem-limited phytopathogen Xylella fastidiosa. Mol Plant Microbe Interact 26:676–685. doi: 10.1094/MPMI-12-12-0283-R [DOI] [PubMed] [Google Scholar]
- 30. Newman MA, Dow JM, Daniels MJ. 2001. Bacterial lipopolysaccharides and plant-pathogen interactions. Eur J Plant Pathol 107:95–102. doi: 10.1023/A:1008738817551 [DOI] [Google Scholar]
- 31. Bertani B, Ruiz N. 2018. Function and biogenesis of lipopolysaccharides. EcoSal Plus 8:1. doi: 10.1128/ecosalplus.ESP-0001-2018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Rapicavoli JN, Blanco-Ulate B, Muszyński A, Figueroa-Balderas R, Morales-Cruz A, Azadi P, Dobruchowska JM, Castro C, Cantu D, Roper MC. 2018. Lipopolysaccharide O-antigen delays plant innate immune recognition of Xylella fastidiosa. Nat Commun 9:390. doi: 10.1038/s41467-018-02861-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Bhunia A, Chua GL, Domadia PN, Warshakoon H, Cromer JR, David SA, Bhattacharjya S. 2008. Interactions of a designed peptide with lipopolysaccharide: Bound conformation and anti-endotoxic activity. Biochem Biophys Res Commun 369:853–857. doi: 10.1016/j.bbrc.2008.02.105 [DOI] [PubMed] [Google Scholar]
- 34. Zhang L, Wei X, Zhang R, Petitte JN, Si D, Li Z, Cheng J, Du M. 2019. Design and development of a novel peptide for treating intestinal inflammation. Front Immunol 10:1841. doi: 10.3389/fimmu.2019.01841 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Wang Z, Liu X, Mao R, Hao Y, Yang N, Wang X, Li Z, Wang X, Wang J, Da Teng . 2020. Development of chimeric peptides to facilitate the neutralisation of lipopolysaccharides during bactericidal targeting of multidrug-resistant Escherichia coli. Commun Biol 3:41. doi: 10.1038/s42003-020-0761-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Pristovsek P, Simcic S, Wraber B, Urleb U. 2005. Structure of a synthetic fragment of the lipopolysaccharide (LPS) binding protein when bound to LPS and design of a peptidic LPS inhibitor. J Med Chem 48:7911–7914. doi: 10.1021/jm050762a [DOI] [PubMed] [Google Scholar]
- 37. Elass-Rochard E, Roseanu A, Legrand D, Trif M, Salmon V, Motas C, Montreuil J, Spik G. 1995. Lactoferrin-lipopolysaccharide interaction: involvement of the 28-34 loop region of human lactoferrin in the high-affinity binding to Escherichia coli 055B5 lipopolysaccharide. Biochem J 312 (Pt 3):839–845. doi: 10.1042/bj3120839 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Kim H, Jang JH, Kim SC, Cho JH. 2016. Enhancement of the antimicrobial activity and selectivity of GNU7 against Gram-negative bacteria by fusion with LPS-targeting peptide. Peptides 82:60–66. doi: 10.1016/j.peptides.2016.05.010 [DOI] [PubMed] [Google Scholar]
- 39. Allende D, McIntosh TJ. 2003. Lipopolysaccharides in bacterial membranes act like cholesterol in eukaryotic plasma membranes in providing protection against melittin-induced bilayer lysis. Biochemistry 42:1101–1108. doi: 10.1021/bi026932s [DOI] [PubMed] [Google Scholar]
- 40. Sabnis A, Hagart KLH, Klöckner A, Becce M, Evans LE, Furniss RCD, Mavridou DA, Murphy R, Stevens MM, Davies JC, Larrouy-Maumus GJ, Clarke TB, Edwards AM. 2021. Colistin kills bacteria by targeting lipopolysaccharide in the cytoplasmic membrane. Elife 10:e65836. doi: 10.7554/eLife.65836 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Novović K, Jovčić B. 2023. Colistin resistance in Acinetobacter baumannii: molecular mechanisms and epidemiology. Antibiotics (Basel) 12:516. doi: 10.3390/antibiotics12030516 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Padhy I, Dwibedy SK, Mohapatra SS. 2024. A molecular overview of the polymyxin-LPS interaction in the context of its mode of action and resistance development. Microbiol Res 283:127679. doi: 10.1016/j.micres.2024.127679 [DOI] [PubMed] [Google Scholar]
- 43. Caravaca-Fuentes P, Camó C, Oliveras À, Baró A, Francés J, Badosa E, Planas M, Feliu L, Montesinos E, Bonaterra A. 2021. A bifunctional peptide conjugate that controls infections of Erwinia amylovora in pear plants. Molecules 26:3426. doi: 10.3390/molecules26113426 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Intentional import of organisms that are plant pests or potential plant pests. 2006. EPPO Bulletin 36:191–194. doi: 10.1111/j.1365-2338.2006.00908.x [DOI] [Google Scholar]
- 45. Jeger M, Caffier D, Candresse T, Chatzivassiliou E, Dehnen‐Schmutz K, Gilioli G, Grégoire J, Jaques Miret JA, MacLeod A, Navajas Navarro M, et al. 2018. Updated pest categorisation of Xylella fastidiosa. EFSA J 16:e0537. doi: 10.2903/j.efsa.2018.5357 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Davis MJ, French WJ, Schaad NW. 1981. Axenic culture of the bacteria associated with phony disease of peach and plum leaf scald. Curr Microbiol 6:309–314. doi: 10.1007/BF01566883 [DOI] [Google Scholar]
- 47. Wells JM, Raju BC, Nyland G, Lowe SK. 1981. Medium for isolation and growth of bacteria associated with plum leaf scald and phony peach diseases. Appl Environ Microbiol 42:357–363. doi: 10.1128/aem.42.2.357-363.1981 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Ferre R, Badosa E, Feliu L, Planas M, Montesinos E, Bardají E. 2006. Inhibition of plant-pathogenic bacteria by short synthetic cecropin A-melittin hybrid peptides. Appl Environ Microbiol 72:3302–3308. doi: 10.1128/AEM.72.5.3302-3308.2006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Badosa E, Moiset G, Montesinos L, Talleda M, Bardají E, Feliu L, Planas M, Montesinos E. 2013. Derivatives of the antimicrobial peptide BP100 for expression in plant systems. PLoS One 8:e85515. doi: 10.1371/journal.pone.0085515 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Baró A, Badosa E, Montesinos L, Feliu L, Planas M, Montesinos E, Bonaterra A. 2020. Screening and identification of BP100 peptide conjugates active against Xylella fastidiosa using a viability-qPCR method. BMC Microbiol 20:229. doi: 10.1186/s12866-020-01915-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Levin PA. 2002. 6 Light microscopy techniques for bacterial cell biology. Methods Microbiol 31:115–132. doi: 10.1016/s0580-9517(02)31007-9 [DOI] [Google Scholar]
- 52. University of Toronto’s Flow Cytometry Facility . 2025. Fixing cells with paraformaldehyde. Available from: https://flowcytometry.utoronto.ca/
- 53. Zhu L, Rajendram M, Huang KC. 2021. Effects of fixation on bacterial cellular dimensions and integrity. iScience 24:102348. doi: 10.1016/j.isci.2021.102348 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. El Handi K, Sabri M, Valentini F, De Stradis A, Achbani EH, Hafidi M, El Moujabber M, Elbeaino T. 2022. Exploring active peptides with antimicrobial activity in planta against Xylella fastidiosa. Biology (Basel) 11:1685. doi: 10.3390/biology11111685 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Montesinos L, Bundó M, Badosa E, San Segundo B, Coca M, Montesinos E. 2017. Production of BP178, a derivative of the synthetic antibacterial peptide BP100, in the rice seed endosperm. BMC Plant Biol 17:63. doi: 10.1186/s12870-017-1011-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Shi J, Zhao Y, Wang Y, Gao W, Ding J, Li P, Hu L, Shao F. 2014. Inflammatory caspases are innate immune receptors for intracellular LPS. Nature 514:187–192. doi: 10.1038/nature13683 [DOI] [PubMed] [Google Scholar]
- 57. Martínez-Sernández V, Orbegozo-Medina RA, Romarís F, Paniagua E, Ubeira FM. 2016. Usefulness of ELISA methods for assessing LPS interactions with proteins and peptides. PLoS ONE 11:e0156530. doi: 10.1371/journal.pone.0156530 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Brandenburg K, Garidel P, Andra J, Jürgens G, Müller M, Blume A, Koch MHJ, Levin J. 2003. Cross-linked hemoglobin converts endotoxically inactive pentaacyl endotoxins into a physiologically active conformation. J Biol Chem 278:47660–47669. doi: 10.1074/jbc.M304743200 [DOI] [PubMed] [Google Scholar]
- 59. Elass-Rochard E, Legrand D, Salmon V, Roseanu A, Trif M, Tobias PS, Mazurier J, Spik G. 1998. Lactoferrin inhibits the endotoxin interaction with CD14 by competition with the lipopolysaccharide-binding protein. Infect Immun 66:486–491. doi: 10.1128/IAI.66.2.486-491.1998 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Wu X, Qian G, Zhao Y, Xu D. 2005. LBP inhibitory peptide reduces endotoxin-induced macrophage activation and mortality. Inflamm Res 54:451–457. doi: 10.1007/s00011-005-1378-1 [DOI] [PubMed] [Google Scholar]
- 61. Rezania S, Amirmozaffari N, Tabarraei B, Jeddi-Tehrani M, Zarei O, Alizadeh R, Masjedian F, Zarnani AH. 2011. Extraction, purification and characterization of lipopolysaccharide from Escherichia coli and Salmonella typhi. Avicenna J Med Biotechnol 3:3–9. [PMC free article] [PubMed] [Google Scholar]
- 62. Al-AAlim AM, Al-Iedani AA, Hamad MA. 2022. Extraction and purification of lipopolysaccharide from Escherichia coli (local Isolate) and study its pyrogenic activity. IJVS 36:45–51. doi: 10.33899/ijvs.2021.128963.1614 [DOI] [Google Scholar]
- 63. Rautenbach M, Gerstner GD, Vlok NM, Kulenkampff J, Westerhoff HV. 2006. Analyses of dose-response curves to compare the antimicrobial activity of model cationic α-helical peptides highlights the necessity for a minimum of two activity parameters. Anal Biochem 350:81–90. doi: 10.1016/j.ab.2005.11.027 [DOI] [PubMed] [Google Scholar]
- 64. Schmidtchen A, Pasupuleti M, Malmsten M. 2014. Effect of hydrophobic modifications in antimicrobial peptides. Adv Colloid Interface Sci 205:265–274. doi: 10.1016/j.cis.2013.06.009 [DOI] [PubMed] [Google Scholar]
- 65. Malmsten M. 2015. Interactions of antimicrobial peptides with bacterial membranes and membrane components. CTMC 16:16–24. doi: 10.2174/1568026615666150703121518 [DOI] [PubMed] [Google Scholar]
- 66. Gagat P, Ostrówka M, Duda-Madej A, Mackiewicz P. 2024. Enhancing antimicrobial peptide activity through modifications of charge, hydrophobicity, and structure. Int J Mol Sci 25:10821. doi: 10.3390/ijms251910821 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Riesco-Llach G, Oliveras À, Gil-Caballero S, Badosa E, Bonaterra A, Montesinos E, Feixas F, Planas M, Feliu L. 2025. Structure-activity relationship of peptide conjugates derived from BP100 and insights into their interactions with lipid membranes by NMR and MD simulations. J Biomol Struct Dyn:1–18. doi: 10.1080/07391102.2025.2458328 [DOI] [PubMed] [Google Scholar]
- 68. Moiset G, Cirac AD, Stuart MCA, Marrink SJ, Sengupta D, Poolman B. 2013. Dual action of BPC194: a membrane active peptide killing bacterial cells. PLoS ONE 8:e61541. doi: 10.1371/journal.pone.0061541 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Analysis and characterization data of peptides; biological activity of peptides; and HPLC, ESI-MS, and HRMS of purified lipopeptides.