Abstract
CRISPR/Cas9 gene editing is widely used to manipulate gene expression and integrate transgenes into specific target sites, making it a powerful tool for recombinant protein expression. In this study, we generated a single donor cassette for CRISPR/Cas9-mediated knock-in at either the αAmy3 or αAmy8 locus in rice cells. The transgene was inserted downstream of the promoter and first exon of the rice αAmy3 or αAmy8 genes, which are highly expressed under sugar-starved conditions in rice suspension cultures. We constructed a simple vector with the homologous intron sequences of the αAmy3 and αAmy8, along with rice codon-optimized recombinant receptor binding domain (rcRBD) of the SARS-CoV-2 spike protein, a functional domain responsible for binding to the angiotensin-converting enzyme 2 (ACE2) receptor on host cells. Using this construct, rcRBD was successfully integrated into the intron 1 of either the αAmy3 or αAmy8 genes. As a result, rcRBD expression was driven by endogenous αAmy3 or αAmy8 promoter-signal peptide. Under the control of αAmy3-signal peptide, rcRBD proteins was detected in both the soluble cellular protein fraction and culture medium, whereas expression driven by the αAmy8 promoter-signal peptide was exclusively detected in the culture medium of rice suspension cells. The highest secreted protein yield of rcRBD in the rice culture medium under the control of αAmy8 endogenous promoter reached 20.7 mg/L, demonstrating a production efficiency comparable to that driven by the endogenous αAmy3 promoter.
Supplementary Information
The online version contains supplementary material available at 10.1007/s00253-025-13549-4.
Keywords: αAmy3 promoter, αAmy8 promoter, CRISPR/Cas9, Intron, Rice suspension cells, Single donor cassette
Key points
Targeted integration at the αAmy3 or αAmy8 locus in rice cells was achieved using a single donor cassette.
rcRBD was driven by sugar-starvation inducible endogenous rice promoters.
Secreted rcRBD yield reached 20.7 mg/L in rice suspension cells.
Supplementary Information
The online version contains supplementary material available at 10.1007/s00253-025-13549-4.
Introduction
In recent years, plants have been extensively studied as platforms for recombinant protein production due to several advantages of their expression systems, including large-scale production capacity, low cultivation costs, minimal risk of human pathogen contamination, and the ability to perform post-translational modifications (Ghaderi et al. 2012; Ma et al.2003; Obembe et al. 2011). To date, over 100 recombinant pharmaceutical proteins, antibodies, enzymes, and vaccines have been successfully produced in plants (Francisco et al. 1997; Staub et al. 2000). Some of these proteins have reached the clinical trial stage (Neutralizing 2009; Richter et al. 2000) and have been commercialized (Magnusdottir et al. 2013; Zimran et al. 2018). In 2012, the Food and Drug Administration (FDA) approved Elelyso, a recombinant enzyme produced in carrot cell culture, for the treatment of Gaucher’s disease (Tekoah et al. 2015). ZMapp, a combination of three monoclonal antibodies made from transgenic tobacco, successfully treated Ebola virus infections when administered to Rhesus macaques (Qiu et al. 2014). In 2022, Canada approved the world’s first plant-based COVID-19 vaccine, Medicago’s coronavirus virus-like particle (CoVLP) vaccine, marking a significant milestone in plant-derived therapeutics (Hager et al. 2022; Ward et al. 2021). Additionally, phase I clinical trials in the USA and Japan have demonstrated that the oral vaccine MucoRice-cholera toxin B (CTB) line 51 A safely induces CTB-specific antibodies capable of neutralizing cholera toxin (Yuki et al. 2022). Moreover, a dose-escalation phase I trial at Chiba University Hospital, expected to conclude in 2026, aims to evaluate the safety, tolerability, and efficacy of MucoRice-CTB line 19 A, an investigational oral cholera vaccine formulated as rice flour (Yuki et al. 2024). These advancements highlight the growing potential of plant-based recombinant protein production systems, demonstrating their ability to compete effectively with conventional recombinant protein platforms.
α-Amylases are well-known enzymes that hydrolyze starch, and their gene expression is highly induced by sugar starvation and gibberellin (GA) during cereal seed germination (Akazawa and Hara-NiShimura 1985; Beck and Ziegler 1989; Lu et al.1998). The rice genome contains at least ten α-amylase genes, among which αAmy3/RAmy3D, αAmy7/RAmy1A, and αAmy8/RAmy3E are predominantly expressed during seed germination and in suspension-cultured cells (Yu et al.1992). The expression of αAmy3 and αAmy8 is dramatically upregulated in sugar-starved rice suspension-cultured cells, whereas other α-amylase genes show only slightly increased or no changes in expression (Huang et al.1993). The αAmy3 promoter-signal peptide-based system for recombinant protein production in rice suspension culture cells is a well-established plant-based platform (Macharoen et al.2020). By fusing a target protein with the signal peptide, the recombinant protein is secreted directly into the culture medium (Huang et al.2015), allowing for simple and cost-effective harvesting and purification. Several recombinant proteins have been successfully produced using the αAmy3 promoter in rice suspension cells (Hong et al. 2008; Huang et al.2005; Jung 2022; Macharoen et al. 2021; Van Giap et al.2019). To further enhance recombinant protein yield, various strategies have been developed, including culture medium optimization (Liu et al.Kim et al. 2014; 2018), rice codon optimization (Bastami and Hosseini 2024; Nguyen et al. 2024), and genetic engineering of the rice cell host (Sinaga et al. 2021). In contrast, the αAmy8 promoter is less frequently reported for recombinant protein production. However, it has been used to drive recombinant protein accumulation in both rice suspension cells and aerobic rice seedlings (Chiang et al. 2005; Su et al. 2012). Furthermore, modification of the αAmy8 promoter by adding a G box and duplicating a TA box resulted in a 28.4-fold increase in hEGF protein accumulation in hypoxic seedlings compared to the wild-type αAmy8 promoter (Wu et al. 2014).
The CRISPR/Cas9 system is widely recognized for its ability to edit genes at specific target sites across various cell types and organisms (Sander and Joung 2014). This system comprises two key components: the guide RNA (gRNA) and the CRISPR-associated protein 9 (Cas9). The gRNA consists of a CRISPR RNA (crRNA), which contains a 20-nucleotide sequence complementary to the target DNA, and a trans-activating CRISPR RNA (tracrRNA), which serves as a scaffold for Cas9 nuclease (DeltcHeva et al. 2011). Once the Cas9 nuclease recognizes the Protospacer Adjacent Motif (PAM) site, double-stranded breaks (DSBs) occur three base pairs upstream of the PAM site (Liu et al.2019b). These DSBs are subsequently repaired by two mechanisms: non-homologous end joining (NHEJ) and homology-directed repair (HDR) pathways. In the NHEJ pathway, DNA fragments are rejoined, often resulting in insertion or deletion (indels) mutations. In contrast, the HDR pathway relies on an exogenous donor template and is commonly used for gene replacement and knock-in applications (Dillingham and Kowalczykowski 2008; Schiml et al.2014; Shukla et al. 2009). Recent studies have demonstrated the feasibility of achieving gene knock-in in plants using the CRISPR/Cas9 system via the NHEJ repair pathway (Pang et al.2020; Weinthal et al.2013). Additionally, multiplex CRISPR/Cas9 approaches enable simultaneous gene knock-in at multiple genomic loci. Wang et al. (2021) demonstrated that a multiplex CRISPR/Cas9 system, combined with two large donor DNA fragments, enabled the knock-in of human immunoglobulin heavy (Ig H) and light (Ig L) chain genes into two native mouse loci, IgH and IgK (Wang et al.2021). Moreover, the same system was used to simultaneously insert two antimicrobial peptide transgenes at multiple loci in catfish (Wang et al.2024). However, in both cases, each donor DNA was specific to only one target site, limiting the flexibility of donor usage across multiple loci.
Previously, we developed a modified rice suspension cell platform for recombinant protein production by using the endogenous αAmy3 promoter and signal peptide via the CRISPR/Cas9 knock-in system (Nguyen et al.2022). An artificial 3′ splicing site was fused upstream of the recombinant gene, allowing protein expression to be controlled by the native αAmy3 promoter and signal peptide. Using this platform, green fluorescent protein (GFP) (Nguyen et al. 2022) and the mature form of human bone morphogenetic protein 2 (rhBMP2m) (Nguyen et al. 2024) were successfully produced. The highest yield of rhBMP2 reached 1.03% of the total soluble protein in the homozygous rhBMP2 knock-in cell line.
Although sugar starvation enhances the expression of αAmy8 and αAmy3, these genes are active in different tissues and developmental stages of rice. The goal of this study was to develop a single donor cassette for CRISPR/Cas9-mediated insertion of a transgene into intron 1 of αAmy8 and/or αAmy3 loci, enabling both promoters to drive recombinant gene expression in the rice suspension cells. The receptor-binding domain (RBD) of the SARS-CoV-2 spike protein of is a functional domain responsible for binding to the angiotensin-converting enzyme 2 (ACE2) receptor on the human cell membrane (Lan et al. 2020). The RBD-ACE2 interaction is critical for viral entry, making RBD an attractive target for vaccine development (Dickey et al. 2022; Montgomerie et al. 2023; Yang et al. 2020). We designed a single donor cassette flanked by αAmy8 and αAmy3 intron 1 target sites on both ends of the rice codon-optimized RBD gene (rcRBD), allowing excision and insertion into either target site via CRISPR/Cas9. The rcRBD DNA fragment was successfully integrated into intron 1 of αAmy8 or αAmy3, and expression was driven by the corresponding endogenous promoter and signal peptide. The highest yield of rcRBD protein secreted into the culture medium, under the control of the αAmy8 promoter, reached 20.7 mg/L.
Materials and methods
Plant materials
Rice calli were induced from immature seeds of Oryza sativa L. cultivar Tainung67 (TNG67), obtained from the Taiwan Agricultural Research Institute, Ministry of Agriculture, Taichung, Taiwan. The immature seeds were dehulled, sterilized with 2.4% (v/v) sodium hypochlorite, washed thoroughly with sterile water, and placed on CIM-I medium (NiShimura et al. 2006). They were then kept in a growth chamber at 28 °C to generate healthy calli. For rice suspension cells, compact and globular calli with an intense yellow color were transferred into a 125-mL flask containing 25 mL of MS liquid medium (MuraShige and Skoog 1962) supplemented with 10 µM 2,4-dichlorophenoxyacetic acid (2,4-D) and 3% sucrose. The cells were maintained on an orbital shaker at 110 rpm in a dark culture room at 28 °C and subcultured every week in fresh MS medium with 2,4-D.
Plasmid construction
The αAmy8,3 master donor vector was constructed in pET-28a backbone following the method described in Nguyen et al. 2022), with modification. The final construct incorporated the αAmy8 E8 site, αAmy3 C3 site, AscI site, NotI site, Nos terminator DNA region (NosT), αAmy3 C3 site, and αAmy8 E8 site (Supplementary Fig. S1) to enable specific target of αAmy8 and αAmy3 genes. The recombinant receptor binding domain (rcRBD), optimized for rice codons (Supplementary Fig. S2), was synthesized and cloned into the αAmy8,3 master donor vector using AscI and NotI to generate the αAmy8,3-rcRBD donor cassette. To construct the pH-Cas9-αAmy3sgRNA and pH-Cas9-αAmy8sgRNA vectors, 20 base pair spacers for αAmy3 and αAmy8 were synthesized with BsaI overhangs and then cloned into the CRISPR vector as previously reported (Ma et al. 2015).
Plant transformation
The donor plasmid αAmy8,3-rcRBD was mixed with pH-Cas9-αAmy3sgRNA and pH-Cas9-αAmy8sgRNA in a 3:1:1 molar ratio before bombardment. One-month-old immature embryo-derived calli were transferred to CIM-II medium containing 0.4% phytagel and bombarded with the plasmid mixture using the PDS-1000/He system (Bio-Rad, Hercules, CA, USA) at 1100 psi, as described in a previous study (Liu et al. 2014). Three days after bombardment, the calli were moved to N6 selection medium containing 50 mg L−1 hygromycin B.
PCR-based genotype analysis
Genomic DNA was extracted from calli using a previously reported method (Huang et al. 2005). Briefly, rice samples were ground with liquid nitrogen using a mortar and pestle, and genomic DNA was extracted using an extraction buffer containing 100 mM Tris(hydroxymethyl)aminomethane hydrochloride (Tris–HCl), pH 8.0, 50 mM ethylenediaminetetraacetic acid (EDTA, pH 8.0), 100 mM sodium chloride (NaCl), 1% sodium dodecyl sulfate (SDS), and 1% β-mercaptoethanol. Subsequently, 1 μg of each genomic DNA sample was amplified via PCR with various gene-specific primers (Supplemental Table S1). Finally, the resulting PCR products were separated by agarose gel electrophoresis.
RT-PCR
For RT-PCR and quantitative real-time polymerase chain reaction (qRT-PCR) analysis, total RNA was isolated using the TRIzol reagent (Invitrogen, Carlsbad, CA, USA). To remove possible DNA contamination, the isolated total RNA was treated with RNase-free DNase I (NEB, Ipswich, MA, USA). Two and a half micrograms of total RNA were reverse transcribed using ReverTra Ace® reverse transcriptase (Toyobo, Osaka, Japan) with oligo-dT primers. A tenfold dilution of the first-strand cDNA product was amplified using the specific primer sets mentioned above. The expression of ACT1 was employed as a reference gene for normalization.
Western blot analysis
The culture medium was collected and centrifuged at 18,000 × g at 4 °C for 20 min to remove cell debris and harvest cultured-medium proteins. Total soluble cellular protein was isolated as previously described (Huang et al. 2005). The concentration of the total protein was determined using the Bio-Rad Protein Assay reagent (Bio-Rad, Hercules, CA, USA). Thirty microliters of cultured medium or 100 µg of total soluble proteins were separated by 15% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) after being denatured for 5 min at 100 °C and then transferred onto a polyvinylidene difluoride (PVDF) membrane (Merck, Darmstadt. Germany). The rabbit anti-SARS-CoV-2 spike-receptor-binding domain (RBD) region antibody (Merck, Darmstadt. Germany) was used as the primary antibody, and horseradish peroxidase (HRP)-conjugated anti-rabbit IgG was used as the secondary antibody to detect rcRBD protein. The protein signal was obtained by chemiluminescence using ECL Prime Western Blot Detection Reagent (Merck, Darmstadt. Germany). The commercial recombinant 2019-nCoV spike RBD protein with a His tag (Merck, Darmstadt. Germany) was used as a positive control.
Enzyme-linked immunosorbent assay (ELISA)
The concentration of rcRBD in the cultured medium was determined by ELISA. A 60 µL sample was coated onto 96-well microtiter plates and incubated overnight at 4 °C. Commercial recombinant 2019-nCoV spike RBD protein was used as the standard. The rabbit anti-SARS-CoV-2 spike RBD region antibodies (Merck, Darmstadt. Germany) was added to the wells and incubated for 2 h at room temperature. Goat HRP-conjugated anti-rabbit IgG antibodies were then added for the detection of the rabbit IgG antibodies and incubated at room temperature for 2 h. A substrate of 3,3′,5,5′-tetramethylbenzidine (TMB) solution (Abcam, Cambridge, UK) was added to the wells and incubated for 30 min in the dark, followed by the addition of 2 M H₂SO₄. The optical density at 450 nm of each well was recorded using an AMR-100 ELISA Reader (Allsheng, Hangzhou, China).
Results
Strategy for inserting a recombinant gene into two targets by a single donor
The CRISPR/Cas9 system has demonstrated the capability to perform gene knock-ins across various organisms including rice. By providing a gene donor in the form of double-stranded DNA (dsDNA) or single-stranded DNA (ssDNA), the desired gene can be knocked into a target genome site via the CRISPR/Cas9 system through non-homologous end joining (NHEJ) or homology-directed repair (HDR) mechanisms (Han et al. 2023; He et al. 2016; Li et al. 2017; SawatSubaShi et al. 2018). In recent years, the development of multiplex target sites using the CRISPR/Cas9 system has allowed for the production of multiple single guide RNAs (sgRNAs) from a single transcript (Lowder et al. 2017; McCarty et al. 2020; Xie and Yang 2019). Wang et al. (2024) reported a multi-sgRNA-based multi-gene editing (msMGE) system that combines with four linear dsDNA donors to generate transgenic fish bearing two copies of two transgenes at four target loci (Wang et al. 2024). To overcome the limitation of one donor to one target for multiple knock-ins, we developed a single donor cassette to target two sites: intron 1 of the αAmy8 and/or αAmy3 genes.
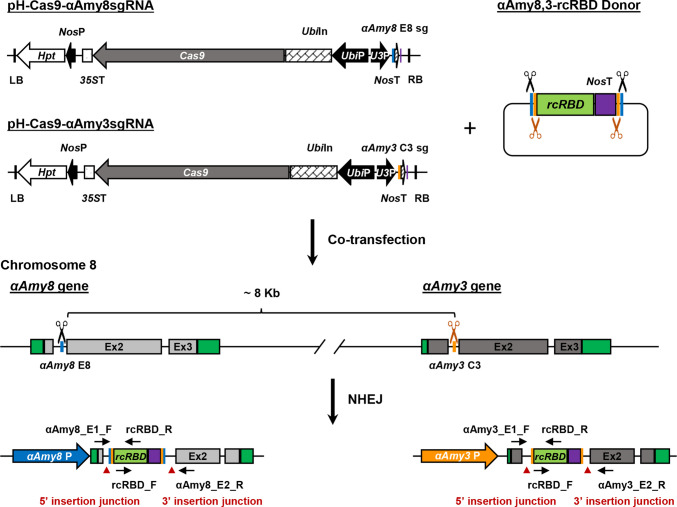
The αAmy8 (Os08g0473600) and αAmy3 (Os08g0473900) genes are located on chromosome 8 of the rice genome. Both genes consist of three exons separated by two introns, with each exon 1 encoding a signal peptide (SP) (Mitsui et al.1996). In a previous study, we established a CRISPR knock-in system targeting the αAmy3 intron 1 for recombinant protein production controlled by the endogenous αAmy3 promoter and signal peptide. Since the intron can be spliced during mRNA maturation, generating mutations via DNA knock-in at the intron site may not affect recombinant protein production. Using this system, GFP (Nguyen et al. 2022) and human mature BMP2 (Nguyen et al. 2024) were produced in rice suspension cell system. Here, we selected the intron 1 sites of αAmy8 and αAmy3 for our single donor CRISPR/Cas9 knock-in system. We generated a donor plasmid, αAmy8,3-rcRBD, consisting of αAmy8 E8 site, αAmy3 C3 site, rcRBD cDNA, Nos terminator DNA region (NosT), αAmy3 C3 site, and αAmy8 E8 site (Fig. 1) to facilitate DNA integration at the C3 and E8 site via the NHEJ repair mechanism. To create a chimeric intron when rcRBD DNA was inserted into either αAmy3 intron 1 or αAmy8 intron 1, a 3′ splice site was added upstream of the rcRBD cDNA (Supplementary Fig. S1).
Fig. 1.
Schematic diagram of the recombinant receptor-binding domain (rcRBD) of the SARS-CoV-2 spike protein, inserted into αAmy8 and/or αAmy3 intron 1 using the CRISPR/Cas9 system. Two CRISPR plasmids were constructed, each comprising a Cas9 expression cassette (Cas9) driven by the Ubiquitin promoter (Bui) and a hygromycin resistance gene (Hat) under the control of the Nos promoter (NosP). One plasmid carried the αAmy8 E8 sgRNA (sg), while the other contained the αAmy3 E3 sgRNA. The donor plasmid includes rcRBD flanked by E8 and C3 sgRNA target sites, which consists of the intron 3′ splicing site (3′ ss), rcRBD coding region, and Nos terminator (NosT). Upon co-bombardment of the CRISPR and donor plasmids, the 3′ss-rcRBD-NosT cassette is integrated into either the αAmy8 or αAmy3 intron 1, resulting in expression units driven by the respective endogenous αAmy8 or αAmy3 promoters and signal peptides. LB, left border; RB, right border; 35ST, 35S CaMV terminator; UbiIn, Ubiquitin intron (in plasmid); E1, Exon 1; Ex2, Exon 2; E3, Exon 3 (in genomic DNA)
Generation of recombinant receptor-binding domain DNA insertions into intron 1 of rice α-amylase genes
To evaluate the ability of a single donor cassette to mediate DNA knock-ins at two rice genomic sites, the αAmy8,3-rcRBD donor plasmid was co-transformed with the CRISPR plasmids pH-Cas9-αAmy3sgRNA and pH-Cas9-αAmy8sgRNA into rice calli using particle bombardment at 3:1:1 molar ratio. Following Cas9-mediated recognition of the target sequences, double-stranded breaks were introduced, releasing the rcRBD donor DNAs at either the αAmy8 E8 or αAmy3 C3 site. This enabled site-specific integration of the rcRBD-NosT into intron 1 of either αAmy8 or αAmy3. PCR screening identified eight calli containing potential αAmy8SP-rcRBD and five calli containing αAmy3SP-rcRBD inserts which further subjected to the protein blot (Supplementary Fig. S3). To assess the insertion of rcRBD cDNA into the introns of αAmy8 or αAmy3 in hygromycin-resistant calli, PCR-based genotyping was performed using site-specific primers (Fig. 2A). Two putative αAmy3SP-rcRBD lines (L1 and L42) and four αAmy8SP-rcRBD lines (L6, L32, L52, and L85) were further analyzed by 5′-junction sequencing to confirm integration sites (Fig. 2B). In L1 and L42, the rcRBD DNA fragments were located downstream of αAmy3 intron 1 DNA sequence, and the absence of the αAmy8 E8 site suggested that rcRBD was excised from the αAmy8,3-rcRBD plasmid through the αAmy3 C3 site (Fig. 2B, Supplementary Fig. S4). In contrast, since the αAmy8 E8 site precedes the αAmy3 C3 site upstream of rcRBD in the donor plasmid, C3 sequences remained detectable within the insertions at the αAmy8 locus (Fig. 2B, Supplementary Fig. S5). Additionally, knock-in lines were subjected to 3′-junction analysis (Supplementary Fig. S4 and S5). These results indicate that the rcRBD DNA was integrated into intron 1 of either αAmy3 or αAmy8 in a forward orientation in all independent lines. To our knowledge, this is the first report of recombinant DNA insertion at two distinct genomic loci using a single donor cassette, highlighting potential of this strategy for multi-site DNA integration when combined with system such as msMGE.
Fig. 2.
Generation of rcRBD knock-in rice cells. A PCR-based genotyping of rcRBD knock-in rice callus cell lines. Specific primer sets were used to amplify Cas9, rcRBD, αAmy3-rcRBD, αAmy8-rcRBD, and ACT1 as a reference gene. B DNA sequencing analysis of two αAmy3-rcRBD and four αAmy8-rcRBD CRISPR-mediated knock-in rice cell lines. PCR-amplified DNA fragments were subjected to Sanger sequencing to examine their 5′-junction sequences. Dark grey characters represent rice genomic DNA sequences upstream of the C3 sites, while light grey for upstream and downstream of the E8 site; blue and orange characters indicate E8 and C3 site sequences, respectively, with underlines indicating protospacer adjacent motif (PAM) sites. Red characters represent the artificial 3′ splice site. Green characters indicate the AscI restriction enzyme adaptor, and light green “ATG” represents the start codon of the rcRBD sequence. Purple characters represent NosT sequences; gold characters indicate the inserted sequences. Negative (−) and positive (+) signs denote nucleotides deleted and inserted at the intron target sites, respectively. Dots (…) indicate sequence continuation, and the double arrowhead (< + >) in gold represents a 361-nt insertion. Wild-type (WT) sequences were used as references for alignment with the knock-in rice cell line sequences, with introns shown in lowercase and exons in uppercase
Stable efficiency of recombinant DNA knock-in at intron 1 of αAmy3 or αAmy8 by single donor strategy
To evaluate the knock-in efficiency of a single donor cassette at two genomic target sites, three additional rice transformation events were performed using particle bombardment. Results from four independent examinations revealed that the frequency of rcRBD DNA insertion into αAmy3 intron 1 in the forward direction ranged from 6.5 to 11.1%, consistent with previous observations (Nguyen et al. 2022). In contrast, the frequency of forward-direction rcRBD DNA insertion at αAmy8 intron 1 ranged from 26.1 to 31.1% (Table 1). These results indicate that recombinant DNA is more likely to be inserted at αAmy8 than at αAmy3 (Table 1). Notably, among a total of 77 knock-in calli analyzed, no line exhibited dual insertion event of rcRBD at both αAmy8 and αAmy3 introns in the forward direction, indicating the lower rate of sequential Cas9 procession.
Table 1.
Efficiency of the rcRBD DNA cassette inserted into αAmy3 intron 1 or αAmy8 intron 1
| No. of calli | Hygromycin resistant calli | rcRBD inserted at αAmy3 intron 1 | rcRBD inserted at αAmy8 intron 1 | |
|---|---|---|---|---|
| Bombardment I | 155 | 45 (29.0%) | 5 (11.1%) | 14 (31.1%) |
| Bombardment II | 180 | 52 (28.8%) | 4 (7.7%) | 15 (28.8%) |
| Bombardment III | 150 | 46 (30.6%) | 3 (6.5%) | 13 (28.3%) |
| Bombardment IV | 200 | 65 (32.5%) | 6 (9.2%) | 17 (26.2%) |
| Total | 685 | 208 (30.4%) | 18 (8.6%) | 59 (28.4%) |
Endogenous α-amylases promoters control the expression of recombinant rcRBD DNA in knock-in suspension cells
During NHEJ repair in a population of cells, the frequency spectrum of indels that form depends on the target’s local DNA sequence (van Overbeek et al. 2016). These indels, which are usually less than 50 bp, can be reliably predicted from DNA sequences (Allen et al. 2019; CHen et al. 2019). The frequent occurrence of single-base-pair insertions, derived from sequences near the cleavage site, has been specifically associated with Cas9-induced double-strand breaks featuring a one-nucleotide 5′ overhang (Jones et al. 2021; Shi et al. 2019). Nevertheless, high size indels also happen and have an impact on Cas9 mutagenesis’s overall effectiveness (Kosicki et al. 2022; Longo et al. 2024).
Given these considerations, we selected transgenic rice callus lines with minimal indel mutations at the rcRBD insertion sites for further study one line each for insertion into αAmy3 and αAmy8. For the αAmy3 insertion, the rcRBD-42 line, which contained a 3-nucleotide deletion, was chosen over rcRBD-1, which had a 361-nucleotide insertion. For the αAmy8 insertion, the rcRBD-32 line was selected due to its relatively small 18-nucleotide deletion, the smallest among the αAmy8SP-rcRBD lines (Fig. 2B).
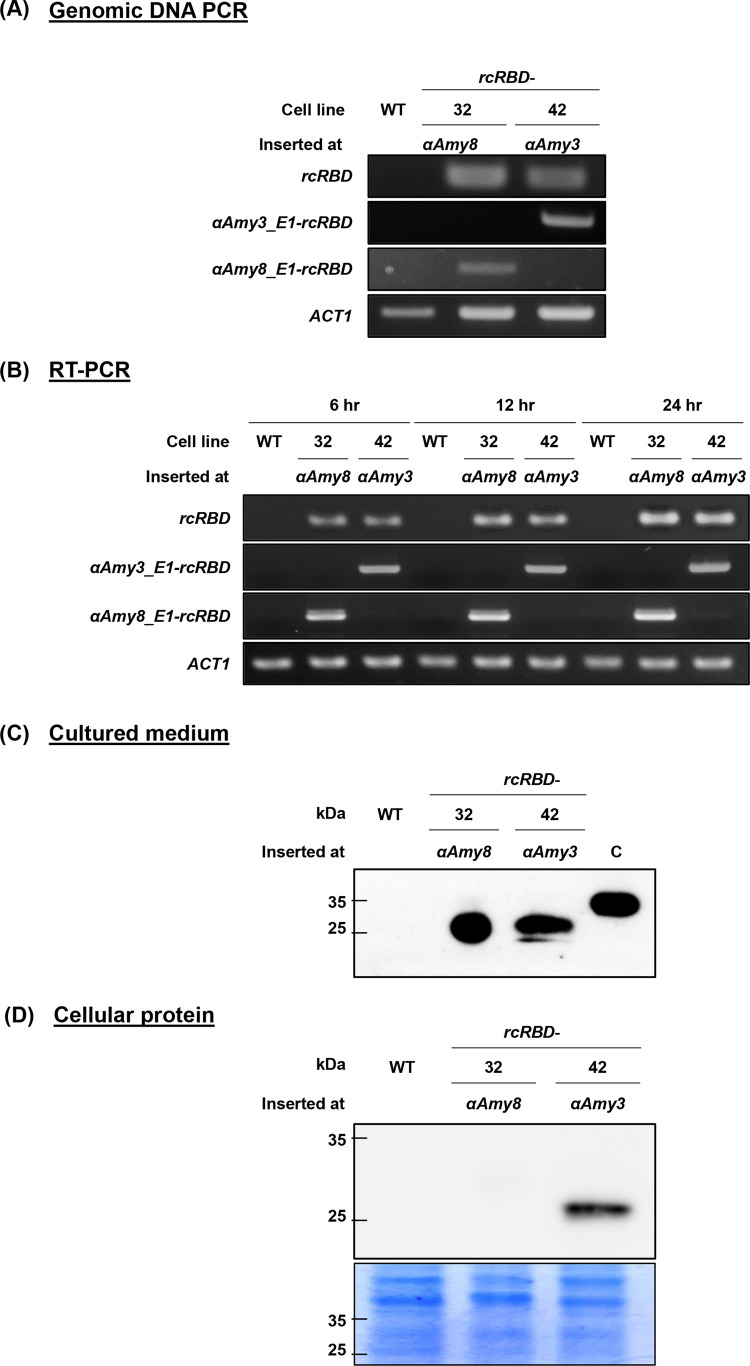
The endogenous αAmy3 and αAmy8 promoters are known to be activated by sugar depletion in cultured rice suspension cells (Chan et al.1994; Lu et al. 1998). It has also been shown that the endogenous αAmy3 promoter can drive the expression of recombinant genes inserted into αAmy3 intron 1 (Nguyen et al. 2024). Using the selected knock-in lines, we established rice suspension cell cultures to examine whether the rcRBD gene was transcriptionally regulated by the endogenous αAmy3 and αAmy8 promoters. Genomic DNA PCR with either αAmy8_E1_F and rcRBD_R or αAmy3_E1_F and rcRBD_R primer sets confirmed the presence of αAmy8-rcRBD and αAmy3-rcRBD insertions in the suspension cell lines, respectively (Fig. 3A). Next, 2% (v/v) of the cells were incubated in sucrose-containing medium for 3 days, followed by transfer to sucrose-free medium for 6, 12, and 24 h. Total RNAs were extracted from sugar-starved cells and subjected to RT-PCR analysis. All sugar-starved cells had rcRBD mRNA expression, as seen in Fig. 3B, and these levels rose even more while the sugar-starved treatment period. Among these lines, rcRBD expression was slightly higher in rcRBD-42 line than in rcRBD-32 line after 6 h of sugar starvation. At 12 and 24 h of treatment, however, rcRBD was shown to be higher in rcRBD-32 line than in rcRBD-42 line (Fig. 3B, Supplementary Fig. S6).
Fig. 3.
Characterizations of rcRBD knock-in rice cell lines. A Genotype analysis of rcRBD knock-in suspension cell lines. Suspension cell lines were cultured from two rcRBD knock-in cell lines, including rcRBD insertion into αAmy8 intron 1 (rcRBD-32) and αAmy3 intron 1 (rcRBD-42). PCR-based genotyping was performed using specific primers for rcRBD, αAmy3_E1-rcRBD, αAmy8_E1-rcRBD, and ACT1 to confirm the rcRBD-integration in these cell lines. B Detection of rcRBD mRNA in knock-in rice suspension cell lines. Total RNA was isolated from wild-type (WT), rcRBD-32, and rcRBD-42 suspension cell lines after 6, 12, and 24 h of culture in sugar-starved (-S) medium. RT-PCR analysis was conducted using specific primers for rcRBD, αAmy3_E1-rcRBD, αAmy8_E1-rcRBD, and ACT1. C, D Detection of rcRBD protein in knock-in rice suspension cells. The secreted protein from culture medium (C) and total soluble protein (D) were extracted from WT, rcRBD-32, and rcRBD-42 cells after 8 days of sugar starvation. A total of 7.5 μl of rcRBD32 culture medium protein and 30 μl of rcRBD-42 culture medium protein were loaded in (C). The same amount 80 ug of total soluble protein from each cell lines were loaded in (D). Western blot analysis was performed using RBD antiserum. Recombinant SARS-CoV-2 spike RBD protein produced from HEK293 cells served as a positive control. Molecular weight markers (kDa) are shown on the left side of the figure. Coomassie blue was used as loading control
Both endogenous signal peptides of the αAmy3 and αAmy8 direct recombinant rcRBD protein into the culture medium from the CRISPR-mediated knock-in rice cells
To analyze recombinant rcRBD protein production and secretion in these selected lines, 34% (v/v) cells of wild type (WT), rcRBD-32, and rcRBD-42 lines were incubated in sucrose-free medium for 8 days. The total proteins from the culture medium and the total cellular proteins were subjected to immunoblotting analysis with anti-SARS-COV-2 spike RBD region antibodies. Fifty nanograms of commercial recombinant 2019-nCoV spike RBD protein produced from HEK293 cells was used as a positive control. The 26 kDa rcRBD proteins were detected in the culture medium of both rcRBD-32 and rcRBD-42 lines (Fig. 3C), indicating that the signal peptides of both αAmy3 and αAmy8 direct rcRBD protein secretion. Additionally, rcRBD proteins were detected in the culture medium of four and seven rcRBD knock-in T0 calli at αAmy3 and αAmy8 introns, respectively (Supplementary Fig. S3), further supporting this finding. Recombinant rcRBD proteins were also detected in the cytosolic protein of rcRBD-42 line but not in rcRBD-32 line (Fig. 3D).
Production profiling of recombinant rcRBD protein in CRISPR-mediated knock-in rice cells
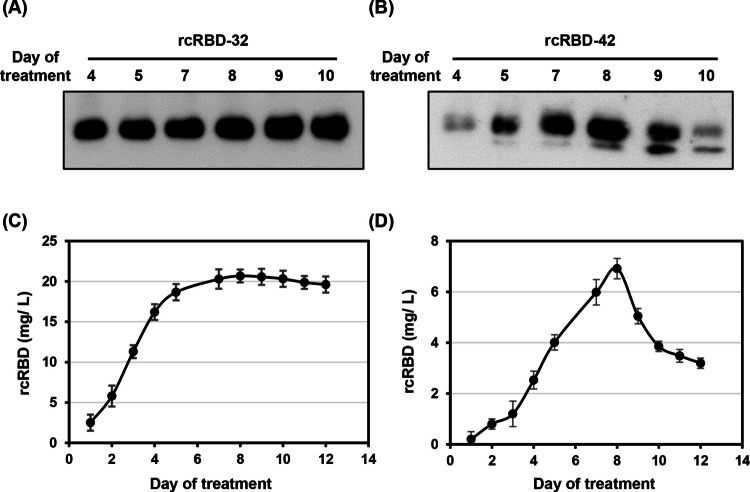
To determine the optimal production of rcRBD protein in these CRISPR-mediated knock-in rice cell lines, 20% (v/v) cells of the rcRBD-32 and rcRBD-42 lines were incubated in sucrose-free medium for 4 to 10 days. The abundance of rcRBD during sugar starvation was detected via western blotting using anti-SARS-COV-2 spike RBD region antibodies. In the rcRBD-32 line, rcRBD proteins were detected starting from day 4, with levels continuously increasing over the following days (Fig. 4A). In the rcRBD-42 line, rcRBD protein was also detected from day 4, continuously increasing until day 8, after which it gradually decreased over time (Fig. 4B). The yield of rcRBD in the culture medium of the rcRBD-32 and rcRBD-42 lines was measured using ELISA. The maximal yield was detected on day 8 after sugar depletion in both lines, reaching 20.7 mg/L for rcRBD-32 and 7 mg/L for rcRBD-42 (Fig. 4C and D), suggesting the expressed rcRBD in the rcRBD-42 line was distributed intra- and extracellularly, reducing its yield in the culture medium compared to the rcRBD-32 line.
Fig. 4.
Profiling of secreted recombinant rcRBD production in knock-in cell line. A–B Suspension cells from rcRBD-32 (A) and rcRBD-42 (B) cell lines were cultured in a 4 mL of sugar-free MS medium. The culture medium was collected from days 4 to 10, and equal amounts of secreted medium protein from each sample were subjected to western blot analysis using RBD antiserum. C–D The rcRBD productivity in the culture medium of rice suspension cultures. The rcRBD protein levels in rcRBD-32 (C) and rcRBD-42 (D) cell lines were determined by ELISA, relative to a standard using a serial dilution of RBD protein derived from HEK293 cells. Error bars represents the standard deviation (SD) from triplicate cultures
The antigen–antibody interaction of rice-produced rcRBD proteins detected by COVID-19 self-test kit
To test whether rice cell-produced rcRBD proteins can potentially be used as a vaccine for protecting humans from COVID-19 infection, the antigen–antibody interaction ability of recombinant rcRBD proteins isolated from the culture medium of the rcRBD-32 and rcRBD-42 lines was evaluated using the COVID-19 Antigen Detection Kit. The antibody binding signal was obtained for rice cell-produced rcRBD from both rcRBD-32 and rcRBD-42 cell lines (Fig. 5). This result indicates that the rice cell-produced rcRBD protein can be recognized by COVID-19 antibodies.
Fig. 5.

Antigen–antibody recognition of rice cell secreted rcRBD. The culture medium was collected on day 8 from sugar-starved WT, rcRBD-32, and rcRBD-42 suspension cells. An equal total volume (100 uL) of cultured medium from each cell lines was used to test antigen–antibody recognition using the COVID-19 Antigen Detection Kit (Celltrion DiaTrust™, Humasis Co, Gyeonggi-do, Republic of Korea). A positive result is indicated by the presence of two lines, one at the control (C) region and one at the test (T) region. A negative result is indicated by a single line at the C region
Discussion
Potential influence of chromatin structure on insertion efficiency
In our result, we observed a higher frequency of forward rcRBD insertions at αAmy8 intron 1 compared to αAmy3 intron 1 (Table 1). This could be partially attributed to the relative genomic positions of these two genes. On chromosome 8, αAmy8 is located upstream of αAmy3 (Fig. 1), and co-expression of αAmy8 E8sgRNA-Cas9 and αAmy3 C3sgRNA-Cas9 may result in DNA double-strand breaks (DSBs) at either site, or simultaneously at both sites (Supplementary Fig. S7). When both DSBs occur, a large genomic deletion (~ 8 kb) spanning from αAmy8 intron to αAmy3 intron 1 region may occur. In such cases, only forward insertion of rcRBD at αAmy8 intron 1 is detectable, as insertion at αAmy3 becomes undetectable due to the loss of its promoter and 5′ regulatory sequences (Supplementary Fig. S7).
In addition to genomic positioning, chromatin structure at the target site is a critical factor influencing the efficiency of CRISPR/Cas9-mediated genome editing (Liu et al.Jensen et al. 2017; 2019a). The accessibility of the Cas9 enzyme to its target DNA sequence is largely dependent on chromatin conformation. Prior studies have shown that Cas9 binding and cleavage are hindered in densely packed chromatin regions, such as those enriched with histones or DNA methylation (Liu et al. 2019a). Conversely, Cas9 activity is more efficient in regions of open, euchromatic chromatin. Plant studies have further supported the role of chromatin characteristics in modulating CRISPR/Cas9 targeting and binding efficiency. Heterochromatic regions generally exhibit lower chromatin accessibility, leading to reduced targeting and cleavage efficiency compared to euchromatic regions (Weiss et al. 2022). Furthermore, DNA repair following Cas9-induced cleavage—particularly via the NHEJ pathway—is more efficient in euchromatic regions, where the DNA repair machinery has greater access to the break site (ScHep et al. 2021). Taken together, both genomic context and local chromatin structure likely contribute to the observed difference in knock-in efficiency between the αAmy8 and αAmy3 loci.
The relationship between gene expression and chromatin structure has long been recognized in molecular biology (Felsenfeld et al. 1996; Yang and Seto 2003). Euchromatin, characterized by a more open and accessible chromatin configuration, is typically associated with actively transcribed genes, as it allows transcriptional machinery to access the DNA. In contrast, heterochromatin represents a more condensed chromatin state that limits DNA accessibility and is generally linked to transcriptional repression (TaMaru 2010). Under sucrose containing conditions during bombardment-mediated transformation, sugar suppression of αAmy8 is not as stringent, and residual expression has been observed compared to αAmy3 (CHen et al. 2002; Umemura et al. 1998), suggesting that αAmy8 may reside in a more open chromatin state. In contrast, the tight sugar suppression of αAmy3 expression implies that it may be located in a more compact chromatin region. Thus, the gene editing efficiency via the CRISPR/Cas9 and NHEJ pathway might be expected to be higher at αAmy8 than at αAmy3 under sucrose containing culture condition. However, there is currently insufficient evidence to clearly distinguish the chromatin structures of αAmy8 and αAmy3. Further studies are necessary to fully understand how sucrose modulates chromatin structure and transcriptional activity of αAmy3 and αAmy8, and how these factors impact CRISPR/Cas9-mediated gene editing efficiency.
Differential rcRBD expression in αAmy8 and αAmy3 knock-in lines
In our study, the αAmy8 knock-in line rcRBD-32 had higher rcRBD expression than αAmy3 knock-in line rcRBD-42 after 12 and 24 h of sugar starvation (Fig. 3B, Supplementary Fig. S6). This observation contrasts with previous reports, which found that sugar starvation induced αAmy3 expression more strongly than αAmy8 (Huang et al. 1993; Yu et al. 1992). We hypothesize that this unexpected expression pattern may be related to intron-mediated enhancement (IME), a phenomenon in which specific introns enhance gene expression. Although introns are typically removed during mRNA splicing, numerous studies have demonstrated their critical roles in regulating gene expression and RNA stability in plants, including in O. sativa (Morello and Breviario 2008). IME has been widely documented as a mechanism through which certain introns, especially the first intron, enhance transcription (Back and WaltHer 2021; Laxa 2017). The first discovery of this effect was made in maize, where inclusion of introns significantly enhanced gene expression in cultured cells (Callis et al. 1987). More recently, introduction of an intron into a codon-optimized TmDGAT1 gene led to a sevenfold increase in transcript accumulation across five transgenic leaf samples (Cao et al. 2023). In another case, splicing of the first intron of Sh1 was shown to be essential for transcript enhancement, potentially by recruiting cellular trafficking machinery through transcript modifications involving a T-rich motif (CLancy and Hannah 2002). Although the exact molecular mechanisms underlying IME remain to be fully elucidated, it is believed that introns may influence gene expression both before and after transcription (Gallegos and Rose 2019; Parra et al. 2011; Rose et al. 2011; SaMadder et al. 2008). IME-associated sequence elements are frequently enriched near the 5′ end of the first intron, particularly within the 5′-UTR and proximal coding regions (Parra et al. 2011). To our knowledge, no studies have yet examined whether intron 1 of αAmy8 and αAmy3 contains active IME elements. Nonetheless, our findings suggest that differences in rcRBD expression between the two knock-in lines may, in part, be influenced by intronic regulatory effects. These results confirm that the inserted rcRBD gene is transcriptionally regulated by the endogenous αAmy3 and αAmy8 promoters, and open new avenues for investigating IME in rice gene expression systems.
Differential rcRBD localization and molecular weight in αAmy8 and αAmy3 knock-in lines
In our study, the recombinant rcRBD protein was successfully produced and was able to be secreted into culture media in both rcRBD-42 and rcRBD-32 lines. Chen et al. (2004) reported that the signal peptides (SP) of αAmy3 and αAmy8 could direct a cargo protein to localize simultaneously in dual locations: the cell wall/extracellular compartment and plastids (CHen et al. 2004). Furthermore, the endogenous αAmy3 SP has been shown to direct a cargo protein, GFP, into the culture medium and the amyloplasts (Nguyen et al. 2022). Our results are consistent with the observation that the αAmy3 SP directs dual localization of recombinant protein. Interestingly, our study implies that the αAmy8 signal peptide is more efficient at secreting rcRBD protein than the αAmy3 signal peptide. Therefore, a larger quantity of rcRBD protein was directed to the culture medium by the αAmy8 SP rather than to the amyloplasts.
We also noted that the molecular weight of rcRBD was smaller compared to its mammalian counterpart, which is around 38 kDa, as indicated by western blot analysis (Fig. 3C). Two N- and two O-glycosylation sites have been identified on the SARS-COV-2 spike RBD protein that is fully glycosylated when expressed in a heterologous expression system (Antonopoulos et al. 2021; Watanabe et al. 2020). Recombinant RBD produced by Expi293 cells has a molecular weight of approximately 35 kDa (Garcia-Cordero et al. 2021). A similar molecular weight was also found in recombinant RBD generated from HEK293T cells, which was around 36 kDa (Arias-Arias et al. 2023). However, plant-produced RBD has a smaller molecular weight, around 25 kDa in BY-2 cells and 20 to 30 kDa in Medicago cells (Rebelo et al. 2022). The different molecular weights of the rcRBD protein observed in different organisms are due to distinct glycol-forms (Berndt et al. 2021; Mardanova et al. 2022; Poodts et al. 2022; Rebelo et al. 2022).
Production of rcRBD protein in αAmy8 and αAmy3 insertion lines
Besides rice, recombinant RBD has been reported to be expressed in various expression systems, including bacteria, yeast, insect cells, mammalian cells, and plants. In Nicotiana benthamiana, the highest RBD protein yield was reported to be 117 ± 41 to 63 ± 10 μg/g fresh leaves (Shin et al. 2021). Additionally, RBD protein produced by Medicago suspension cells accumulated to 0.5–2.8 mg/L (Ferreira et al. 2023). Our current work surpasses previous plant systems, demonstrating that the use of the endogenous αAmy8 promoter and signal peptide can yield high levels of rcRBD protein. However, in the rcRBD-42 line, rcRBD protein was targeted to both intra- and extracellular locations, resulting in lower rcRBD protein production in the culture medium compared to the rcRBD-32 line.
In addition, higher levels of protein degradation were found in the rcRBD-42 line which probably links to RNA stability. The 3′ untranslated region (3′ UTR) of messenger RNA (mRNA) molecules is a crucial component of this regulatory environment since it has been demonstrated to have a major impact on RNA stability and, consequently, protein expression levels. A wide variety of RNA-binding proteins can bind to the AU-rich regions found in the 3′ UTR, either promoting or inhibiting RNA decay (García-Mauriño et al. 2017). In particular, some RNA-binding proteins can bind to the 3′ UTR and prevent mRNA from degrading, increasing the transcript’s stability and longevity and eventually raising the amount of the produced protein (Jonkhout et al. 2017). In response to the availability of sugar, the 3′ UTR of αAmy3 acts to regulate the stability of mRNA (CHan and Yu 1998). Certain regions of the αAmy3 3′ UTR have been shown to function as destabilizing components when carbohydrates like sucrose are present. When carbohydrates are plentiful, these sequences mediate the quick breakdown of αAmy3 mRNA, which lowers the amounts of α-amylase protein. On the other hand, when sugar is depleted, the mRNA stabilizes and increases the production of proteins (CHan and Yu 1998). Instead of using the 3′ UTR of endogenous αAmy3 or αAmy8, the Nos terminator is used in this work to indicate the end of rcRBD gene expression, which may affect protein production in rcRBD-42 (Fig. 4). Nevertheless, it is still unclear how the 3′ UTR of αAmy8 and RNA stability are related.
Conclusions
This study demonstrated the effective use of a single donor cassette in the CRISPR/Cas9 system to insert recombinant genes into two specific sites in the rice genome, targeting intron 1 of the αAmy3 or αAmy8 genes. Results showed higher efficiency of DNA insertion at the αAmy8 site compared to the αAmy3 site. The endogenous αAmy3 and αAmy8 promoters successfully controlled rcRBD gene expression, and both signal peptides facilitated rcRBD protein secretion into the culture medium, with the αAmy8 signal peptide being more efficient. Optimal rcRBD protein production was achieved on day 8 of sugar depletion, with yields of 20.7 mg/L, surpassing previous plant expression systems. Additionally, rice cell-produced rcRBD proteins could interact with the COVID-19 antibodies and have the potential to be a vaccine candidate.
Supplementary Information
Below is the link to the electronic supplementary material.
Acknowledgements
This study was supported by grants from the National Science and Technology Council of the Republic of China (Grant No. 111-2313-B-155-001-MY3 to LFH and 112-2313-B-008-002-MY3 to CAL). We thank Professors Shaw-Jye Wu (SJW) and Ching-Hui Yeh (CHY) from the Department of Life Sciences, National Central University—SJW for taking over CAL’s grant, and CHY for serving as DSS’s advisor after CAL’s passing in July 2024—and for continuing to financially support DSS through CAL’s original funding.
Author contribution
LFH and CAL designed the research and contributed to manuscript writing. CAL was DSS’s advisor and passed away in July 2024. Following his passing, DSS and CKH revised the manuscript in collaboration with LFH. DSS and PYH performed the experiments and prepared the manuscript. The remaining four authors have read and approved the final version of the manuscript.
Funding
This study was supported by grants from the National Science and Technology Council of the Republic of China (Grant No. 111–2313-B-155–001-MY3 to LFH and 112–2313-B-008–002-MY3 to CAL).
Data availability
No datasets were generated or analysed during the current study.
Declarations
Ethical approval
This study did not involve any human participants or animal subjects. All experiments were conducted using rice (O. sativa) cells. Therefore, ethical approval was not required for this research.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Chung-An Lu, Email: chungan@cc.ncu.edu.tw.
Li-Fen Huang, Email: hlf326@saturn.yzu.edu.tw.
References
- Akazawa T, Hara-Nishimura I (1985) Topographic aspects of biosynthesis, extracellular secretion, and intracellular storage of proteins in plant cells. Annu Rev Plant Physiol 36(1):441–472 [Google Scholar]
- Allen F, Crepaldi L, Alsinet C, Strong AJ, Kleshchevnikov V, De Angeli P, Páleníková P, Khodak A, Kiselev V, Kosicki M (2019) Predicting the mutations generated by repair of Cas9-induced double-strand breaks. Nat Biotechnol 37(1):64–72 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Antonopoulos A, Broome S, Sharov V, Ziegenfuss C, Easton RL, Panico M, Dell A, Morris HR, Haslam SM (2021) Site-specific characterization of SARS-CoV-2 spike glycoprotein receptor-binding domain. Glycobiology 31(3):181–187 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arias-Arias JL, Molina-Castro SE, Monturiol-Gross L, Lomonte B, Corrales-Aguilar E (2023) Stable production of recombinant SARS-CoV-2 receptor-binding domain in mammalian cells with co-expression of a fluorescent reporter and its validation as antigenic target for COVID-19 serology testing. Biotechnology Reports 37:e00780 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Back G, Walther D (2021) Identification of cis-regulatory motifs in first introns and the prediction of intron-mediated enhancement of gene expression in Arabidopsis thaliana. BMC Genomics 22(1):390 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bastami M, Hosseini R (2024) The codon optimised gene produces an active human basic fibroblastic growth factor in rice cell suspension culture. Growth Factors 42(4):171–187 [DOI] [PubMed] [Google Scholar]
- Beck E, Ziegler P (1989) Biosynthesis and degradation of starch in higher plants. Annu Rev Plant Biol 40(1):95–117 [Google Scholar]
- Berndt AJ, Smalley TN, Ren B, Simkovsky R, Badary A, Sproles AE, Fields FJ, Torres-Tiji Y, Heredia V, Mayfield SP (2021) Recombinant production of a functional SARS-CoV-2 spike receptor binding domain in the green algae Chlamydomonas reinhardtii. PLoS ONE 16(11):e0257089 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Callis J, Fromm M, Walbot V (1987) Introns increase gene expression in cultured maize cells. Genes Dev 1(10):1183–1200 [DOI] [PubMed] [Google Scholar]
- Cao VD, Luo G, Korynta S, Liu H, Liang Y, Shanklin J, Altpeter F (2023) Intron-mediated enhancement of DIACYLGLYCEROL ACYLTRANSFERASE 1 expression in energycane promotes a step change for lipid accumulation in vegetative tissues. Biotechnology for Biofuels and Bioproducts 16(1):153 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chan M-T, Yu S-M (1998) The 3′ untranslated region of a rice α-amylase gene functions as a sugar-dependent mRNA stability determinant. Proc Natl Acad Sci 95(11):6543–6547 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chan M-T, Chao Y-C, Yu S-M (1994) Novel gene expression system for plant cells based on induction of alpha-amylase promoter by carbohydrate starvation. J Biol Chem 269(26):17635–17641 [PubMed] [Google Scholar]
- Chen P-W, Lu C-A, Yu T-S, Tseng T-H, Wang C-S, Yu S-M (2002) Rice α-amylase transcriptional enhancers direct multiple mode regulation of promoters in transgenic rice. J Biol Chem 277(16):13641–13649 [DOI] [PubMed] [Google Scholar]
- Chen M-H, Huang L-F, Li H-m, Chen Y-R, Yu S-M (2004) Signal peptide-dependent targeting of a rice α-amylase and cargo proteins to plastids and extracellular compartments of plant cells. Plant Physiol 135(3):1367–1377 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen W, McKenna A, Schreiber J, Haeussler M, Yin Y, Agarwal V, Noble WS, Shendure J (2019) Massively parallel profiling and predictive modeling of the outcomes of CRISPR/Cas9-mediated double-strand break repair. Nucleic Acids Res 47(15):7989–8003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiang C-M, Yeh F-S, Huang L-F, Tseng T-H, Chung M-C, Wang C-S, Lur H-S, Shaw J-F, Yu S-M (2005) Expression of a bi-functional and thermostable amylopullulanase in transgenic rice seeds leads to autohydrolysis and altered composition of starch. Mol Breeding 15:125–143 [Google Scholar]
- Clancy M, Hannah LC (2002) Splicing of the maize Sh1 first intron is essential for enhancement of gene expression, and a T-rich motif increases expression without affecting splicing. Plant Physiol 130(2):918–929 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deltcheva E, Chylinski K, Sharma CM, Gonzales K, Chao Y, Pirzada ZA, Eckert MR, Vogel J, Charpentier E (2011) CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471(7340):602–607 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dickey TH, Tang WK, Butler B, Ouahes T, Orr-Gonzalez S, Salinas ND, Lambert LE, Tolia NH (2022) Design of the SARS-CoV-2 RBD vaccine antigen improves neutralizing antibody response. Sci Adv 8(37):eabq8276 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dillingham MS, Kowalczykowski SC (2008) RecBCD enzyme and the repair of double-stranded DNA breaks. Microbiol Mol Biol Rev 72(4):642–671 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Felsenfeld G, Boyes J, Chung J, Clark D, Studitsky V (1996) Chromatin structure and gene expression. Proc Natl Acad Sci 93(18):9384–9388 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferreira AC, Rebelo BA, Abranches R (2023) A simplified protocol for Agrobacterium-mediated transformation of cell suspension cultures of the model species Medicago truncatula A17. Plant Cell, Tissue Organ Cul (PCTOC) 153(3):669–675 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Francisco JA, Gawlak SL, Miller M, Bathe J, Russell D, Chace D, Mixan B, Zhao L, Fell HP, Siegall CB (1997) Expression and characterization of bryodin 1 and a bryodin 1-based single-chain immunotoxin from tobacco cell culture. Bioconjug Chem 8(5):708–713 [DOI] [PubMed] [Google Scholar]
- Gallegos JE, Rose AB (2019) An intron-derived motif strongly increases gene expression from transcribed sequences through a splicing independent mechanism in Arabidopsis thaliana. Sci Rep 9(1):13777 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garcia-Cordero J, Mendoza-Ramirez J, Fernandez-Benavides D, Roa-Velazquez D, Filisola-Villasenor J, Martinez-Frias SP, Sanchez-Salguero ES, Miguel-Rodriguez CE, Maravillas Montero JL, Torres-Ruiz JJ (2021) Recombinant protein expression and purification of N and RBD of SARS from mammalian cells and their potential applications. Diagnostics 11(10):1808 [DOI] [PMC free article] [PubMed] [Google Scholar]
- García-Mauriño SM, Rivero-Rodríguez F, Velázquez-Cruz A, Hernández-Vellisca M, Díaz-Quintana A, De la Rosa MA, Díaz-Moreno I (2017) RNA binding protein regulation and cross-talk in the control of AU-rich mRNA fate. Front Mol Biosci 4:71 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghaderi D, Zhang M, Hurtado-Ziola N, Varki A (2012) Production platforms for biotherapeutic glycoproteins Occurrence, impact, and challenges of non-human sialylation. Biotechnol Genet Eng Rev 28(1):147–176 [DOI] [PubMed] [Google Scholar]
- Hager KJ, Pérez Marc G, Gobeil P, Diaz RS, Heizer G, Llapur C, Makarkov AI, Vasconcellos E, Pillet S, Riera F (2022) Efficacy and safety of a recombinant plant-based adjuvanted Covid-19 vaccine. N Engl J Med 386(22):2084–2096 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han W, Li Z, Guo Y, He K, Li W, Xu C, Ge L, He M, Yin X, Zhou J (2023) Efficient precise integration of large DNA sequences with 3′-overhang dsDNA donors using CRISPR/Cas9. Proc Natl Acad Sci 120(22):e2221127120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- He X, Tan C, Wang F, Wang Y, Zhou R, Cui D, You W, Zhao H, Ren J, Feng B (2016) Knock-in of large reporter genes in human cells via CRISPR/Cas9-induced homology-dependent and independent DNA repair. Nucleic Acids Res 44(9):e85–e85 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hong S-Y, Lee T-S, Kim J, Jung J-H, Choi C-W, Kim T-G, Kwon T-H, Jang Y-S, Yang M-S (2008) Tumor targeting of humanized fragment antibody secreted from transgenic rice cell suspension culture. Plant Mol Biol 68:413–422 [DOI] [PubMed] [Google Scholar]
- Huang N, Chandler J, Thomas BR, Koizumi N, Rodriguez RL (1993) Metabolic regulation of α-amylase gene expression in transgenic cell cultures of rice (Oryza sativa L.). Plant Mol Biol 23:737–747 [DOI] [PubMed] [Google Scholar]
- Huang L-F, Liu Y-K, Lu C-A, Hsieh S-L, Yu S-M (2005) Production of human serum albumin by sugar starvation induced promoter and rice cell culture. Transgenic Res 14:569–581 [DOI] [PubMed] [Google Scholar]
- Huang L-F, Tan C-C, Yeh J-F, Liu H-Y, Liu Y-K, Ho S-L, Lu C-A (2015) Efficient secretion of recombinant proteins from rice suspension-cultured cells modulated by the choice of signal peptide. PLoS ONE 10(10):e0140812 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jensen KT, Fløe L, Petersen TS, Huang J, Xu F, Bolund L, Luo Y, Lin L (2017) Chromatin accessibility and guide sequence secondary structure affect CRISPR-Cas9 gene editing efficiency. FEBS Lett 591(13):1892–1901 [DOI] [PubMed] [Google Scholar]
- Jones SK, Hawkins JA, Johnson NV, Jung C, Hu K, Rybarski JR, Chen JS, Doudna JA, Press WH, Finkelstein IJ (2021) Massively parallel kinetic profiling of natural and engineered CRISPR nucleases. Biophys J 120(3):138a [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jonkhout N, Tran J, Smith MA, Schonrock N, Mattick JS, Novoa EM (2017) The RNA modification landscape in human disease. RNA 23(12):1754–1769 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jung J-W (2022) Production of recombinant human acid α-glucosidase with mannosidic N-glycans in α-mannosidase I mutant rice cell suspension culture. Plant Biotechnology Reports 16(3):333–342 [Google Scholar]
- Kim N-S, Yu H-Y, Chung N-D, Kwon T-H, Yang M-S (2014) High-level production of recombinant trypsin in transgenic rice cell culture through utilization of an alternative carbon source and recycling system. Enzyme Microb Technol 63:21–27 [DOI] [PubMed] [Google Scholar]
- Kosicki M, Allen F, Steward F, Tomberg K, Pan Y, Bradley A (2022) Cas9-induced large deletions and small indels are controlled in a convergent fashion. Nat Commun 13(1):3422 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lan J, Ge J, Yu J, Shan S, Zhou H, Fan S, Zhang Q, Shi X, Wang Q, Zhang L (2020) Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581(7807):215–220 [DOI] [PubMed] [Google Scholar]
- Laxa M (2017) Intron-mediated enhancement: a tool for heterologous gene expression in plants? Front Plant Sci 7:1977 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H, Beckman KA, Pessino V, Huang B, Weissman JS, Leonetti MD (2017) Design and specificity of long ssDNA donors for CRISPR-based knock-in. BioRxiv:178905. 10.1101/178905
- Liu G, Campbell BC, Godwin ID (2014) Sorghum genetic transformation by particle bombardment. Cereal Genom: methods proto 1099:219–234. 10.1007/978-1-62703-715-0_18 [DOI] [PubMed] [Google Scholar]
- Liu Y-K, Lu C-W, Chang J-Y, Lu C-F, Tan C-C, Huang L-F (2018) Optimization of the culture medium for recombinant protein production under the control of the αAmy3 promoter in a rice suspension-cultured cell expression system. Plant Cell. Tissue and Organ Culture (PCTOC) 132:383–391 [Google Scholar]
- Liu G, Yin K, Zhang Q, Gao C, Qiu J-L (2019) Modulating chromatin accessibility by transactivation and targeting proximal dsgRNAs enhances Cas9 editing efficiency in vivo. Genome Biol 20:1–11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu M, Rehman S, Tang X, Gu K, Fan Q, Chen D, Ma W (2019) Methodologies for improving HDR efficiency. Front Genet 9:691 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Longo GM, Sayols S, Kotini AG, Heinen S, Möckel MM, Beli P, Roukos V (2024) Linking CRISPR–Cas9 double-strand break profiles to gene editing precision with BreakTag. Nat Biotechnol 43:608–622 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowder L, Malzahn A, Qi Y (2017) Rapid construction of multiplexed CRISPR-Cas9 systems for plant genome editing. Plant Pattern Recog Receptors: Methods Protocols 1578:291–307. 10.1007/978-1-4939-6859-6_25 [DOI] [PubMed] [Google Scholar]
- Lu C-A, Lim E-K, Yu S-M (1998) Sugar response sequence in the promoter of a rice α-amylase gene serves as a transcriptional enhancer. J Biol Chem 273(17):10120–10131 [DOI] [PubMed] [Google Scholar]
- Ma JK, Drake PM, Christou P (2003) The production of recombinant pharmaceutical proteins in plants. Nat Rev Genet 4(10):794–805 [DOI] [PubMed] [Google Scholar]
- Ma X, Zhang Q, Zhu Q, Liu W, Chen Y, Qiu R, Wang B, Yang Z, Li H, Lin Y (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8(8):1274–1284 [DOI] [PubMed] [Google Scholar]
- Macharoen K, McDonald KA, Nandi S (2020) A method to simplify bioreactor processing for recombinant protein production in rice cell suspension cultures. MethodsX 7:101139 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Macharoen K, Du M, Jung S, McDonald KA, Nandi S (2021) Production of recombinant butyrylcholinesterase from transgenic rice cell suspension cultures in a pilot-scale bioreactor. Biotechnol Bioeng 118(4):1431–1443 [DOI] [PubMed] [Google Scholar]
- Magnusdottir A, Vidarsson H, Björnsson JM, Örvar BL (2013) Barley grains for the production of endotoxin-free growth factors. Trends Biotechnol 31(10):572–580 [DOI] [PubMed] [Google Scholar]
- Mardanova ES, Kotlyarov RY, Ravin NV (2022) Rapid transient expression of receptor-binding domain of SARS-CoV-2 and the conserved M2e peptide of Influenza A virus linked to flagellin in Nicotiana benthamiana plants using self-replicating viral vector. Plants 11(24):3425 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCarty NS, Graham AE, Studená L, Ledesma-Amaro R (2020) Multiplexed CRISPR technologies for gene editing and transcriptional regulation. Nat Commun 11(1):1281 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mitsui T, Yamaguchi J, Akazawa T (1996) Physicochemical and serological characterization of rice a-amylase lsoforms and identification of their corresponding genes. Plant Physiol 110(4):1395–404. 10.1104/pp.110.4.1395 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Montgomerie I, Bird TW, Palmer OR, Mason NC, Pankhurst TE, Lawley B, Hernández LC, Harfoot R, Authier-Hall A, Anderson DE (2023) Incorporation of SARS-CoV-2 spike NTD to RBD protein vaccine improves immunity against viral variants. Iscience 26(4):106256. 10.1016/j.isci.2023.106256 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morello L, Breviario D (2008) Plant spliceosomal introns: not only cut and paste. Curr Genomics 9(4):227–238 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiologia plantarum 15(3):473–497. 10.1111/j.1399-3054.1962.tb08052.x [Google Scholar]
- Nochi T, Yuki Y, Katakai Y, Shibata H, Tokuhara D, Mejima M, Kurokawa S, Takahashi Y, Nakanishi U, Ono F (2009) A rice-based oral cholera vaccine induces macaque-specific systemic neutralizing antibodies but does not influence pre-existing intestinal immunity. The Journal of Immunology 183(10):6538–6544 [DOI] [PubMed] [Google Scholar]
- Nguyen TM, Lu C-A, Huang L-F (2022) Applications of CRISPR/Cas9 in a rice protein expression system via an intron-targeted insertion approach. Plant Sci 315:111132 [DOI] [PubMed] [Google Scholar]
- Nguyen TM, Wu P-Y, Chang C-H, Huang L-F (2024) High-yield BMP2 expression in rice cells via CRISPR and endogenous αAmy3 promoter. Appl Microbiol Biotechnol 108(1):206 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishimura A, Aichi I, Matsuoka M (2006) A protocol for Agrobacterium-mediated transformation in rice. Nat Protoc 1(6):2796–2802 [DOI] [PubMed] [Google Scholar]
- Obembe OO, Popoola JO, Leelavathi S, Reddy SV (2011) Advances in plant molecular farming. Biotechnol Adv 29(2):210–222 [DOI] [PubMed] [Google Scholar]
- Pang J, Zhou J, Yang D (2020) Knock-in at GluA1 locus improves recombinant human serum albumin expression in rice grain. J Biotechnol 321:87–95 [DOI] [PubMed] [Google Scholar]
- Parra G, Bradnam K, Rose AB, Korf I (2011) Comparative and functional analysis of intron-mediated enhancement signals reveals conserved features among plants. Nucleic Acids Res 39(13):5328–5337 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poodts J, Smith I, Birenbaum JM, Rodriguez MS, Montero L, Wolman FJ, Marfía JI, Valdez SN, Alonso LG, Targovnik AM (2022) Improved expression of SARS-CoV-2 spike RBD using the insect cell-baculovirus system. Viruses 14(12):2794 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qiu X, Wong G, Audet J, Bello A, Fernando L, Alimonti JB, Fausther-Bovendo H, Wei H, Aviles J, Hiatt E (2014) Reversion of advanced Ebola virus disease in nonhuman primates with ZMapp. Nature 514(7520):47–53 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rebelo BA, Folgado A, Ferreira AC, Abranches R (2022) Production of the SARS-CoV-2 spike protein and its receptor binding domain in plant cell suspension cultures. Front Plant Sci 13:995429 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richter LJ, Thanavala Y, Arntzen CJ, Mason HS (2000) Production of hepatitis B surface antigen in transgenic plants for oral immunization. Nat Biotechnol 18(11):1167–1171 [DOI] [PubMed] [Google Scholar]
- Rose AB, Emami S, Bradnam K, Korf I (2011) Evidence for a DNA-based mechanism of intron-mediated enhancement. Front Plant Sci 2:98 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Samadder P, Sivamani E, Lu J, Li X, Qu R (2008) Transcriptional and post-transcriptional enhancement of gene expression by the 5′ UTR intron of rice rubi3 gene in transgenic rice cells. Mol Genet Genomics 279:429–439 [DOI] [PubMed] [Google Scholar]
- Sander JD, Joung JK (2014) CRISPR-Cas systems for editing, regulating and targeting genomes. Nat Biotechnol 32(4):347–355 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sawatsubashi S, Joko Y, Fukumoto S, Matsumoto T, Sugano SS (2018) Development of versatile non-homologous end joining-based knock-in module for genome editing. Sci Rep 8(1):593 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schep R, Brinkman EK, Leemans C, Vergara X, van der Weide RH, Morris B, van Schaik T, Manzo SG, Peric-Hupkes D, van den Berg J (2021) Impact of chromatin context on Cas9-induced DNA double-strand break repair pathway balance. Molecular cell 81(10):2216-2230. e10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schiml S, Fauser F, Puchta H (2014) The CRISPR/C as system can be used as nuclease for in planta gene targeting and as paired nickases for directed mutagenesis in Arabidopsis resulting in heritable progeny. Plant J 80(6):1139–1150 [DOI] [PubMed] [Google Scholar]
- Shi X, Shou J, Mehryar MM, Li J, Wang L, Zhang M, Huang H, Sun X, Wu Q (2019) Cas9 has no exonuclease activity resulting in staggered cleavage with overhangs and predictable di-and tri-nucleotide CRISPR insertions without template donor. Cell Discovery 5(1):53 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shin Y-J, König-Beihammer J, Vavra U, Schwestka J, Kienzl NF, Klausberger M, Laurent E, Grünwald-Gruber C, Vierlinger K, Hofner M (2021) N-glycosylation of the SARS-CoV-2 receptor binding domain is important for functional expression in plants. Front Plant Sci 12:689104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shukla VK, Doyon Y, Miller JC, DeKelver RC, Moehle EA, Worden SE, Mitchell JC, Arnold NL, Gopalan S, Meng X (2009) Precise genome modification in the crop species Zea mays using zinc-finger nucleases. Nature 459(7245):437–441 [DOI] [PubMed] [Google Scholar]
- Sinaga DS, Ho S-L, Lu C-A, Yu S-M, Huang L-F (2021) Knockdown expression of a MYB-related transcription factor gene, OsMYBS2, enhances production of recombinant proteins in rice suspension cells. Plant Methods 17:1–10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Staub JM, Garcia B, Graves J, Hajdukiewicz PT, Hunter P, Nehra N, Paradkar V, Schlittler M, Carroll JA, Spatola L (2000) High-yield production of a human therapeutic protein in tobacco chloroplasts. Nat Biotechnol 18(3):333–338 [DOI] [PubMed] [Google Scholar]
- Su C-F, Kuo I-C, Chen P-W, Huang C-H, Seow SV, Chua KY, Yu S-M (2012) Characterization of an immunomodulatory Der p 2-FIP-fve fusion protein produced in transformed rice suspension cell culture. Transgenic Res 21:177–192 [DOI] [PubMed] [Google Scholar]
- Tamaru H (2010) Confining euchromatin/heterochromatin territory: jumonji crosses the line. Genes Dev 24(14):1465–1478 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tekoah Y, Shulman A, Kizhner T, Ruderfer I, Fux L, Nataf Y, Bartfeld D, Ariel T, Gingis-Velitski S, Hanania U (2015) Large-scale production of pharmaceutical proteins in plant cell culture—the protalix experience. Plant Biotechnol J 13(8):1199–1208 [DOI] [PubMed] [Google Scholar]
- Umemura T-a, Perata P, Futsuhara Y, Yamaguchi J (1998) Sugar sensing and α-amylase gene repression in rice embryos. Planta 204:420–428 [DOI] [PubMed] [Google Scholar]
- Van Giap D, Jung J-W, Kim N-S (2019) Production of functional recombinant cyclic citrullinated peptide monoclonal antibody in transgenic rice cell suspension culture. Transgenic Res 28:177–188 [DOI] [PubMed] [Google Scholar]
- van Overbeek M, Capurso D, Carter MM, Thompson MS, Frias E, Russ C, Reece-Hoyes JS, Nye C, Gradia S, Vidal B (2016) DNA repair profiling reveals nonrandom outcomes at Cas9-mediated breaks. Mol Cell 63(4):633–646 [DOI] [PubMed] [Google Scholar]
- Wang X, Ray R, Kratochvil S, Melzi E, Lin YC, Giguere S, Xu L, Warner J, Cheon D, Liguori A (2021) Multiplexed CRISPR/CAS9-mediated engineering of pre-clinical mouse models bearing native human B cell receptors. EMBO J 40(2):e105926 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang J, Torres IM, Shang M, Al-Armanazi J, Dilawar H, Hettiarachchi DU, Paladines-Parrales A, Chambers B, Pottle K, Soman M (2024) One-step knock-in of two antimicrobial peptide transgenes at multiple loci of catfish by CRISPR/Cas9-mediated multiplex genome engineering. Int J Biol Macromol 260:129384 [DOI] [PubMed] [Google Scholar]
- Ward BJ, Gobeil P, Séguin A, Atkins J, Boulay I, Charbonneau P-Y, Couture M, D’Aoust M-A, Dhaliwall J, Finkle C (2021) Phase 1 randomized trial of a plant-derived virus-like particle vaccine for COVID-19. Nat Med 27(6):1071–1078 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Watanabe Y, Allen JD, Wrapp D, McLellan JS, Crispin M (2020) Site-specific glycan analysis of the SARS-CoV-2 spike. Science 369(6501):330–333 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weinthal DM, Taylor RA, Tzfira T (2013) Nonhomologous end joining-mediated gene replacement in plant cells. Plant Physiol 162(1):390–400 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss T, Crisp PA, Rai KM, Song M, Springer NM, Zhang F (2022) Epigenetic features drastically impact CRISPR–Cas9 efficacy in plants. Plant Physiol 190(2):1153–1164 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu C-S, Kuo W-T, Chang C-Y, Kuo J-Y, Tsai Y-T, Yu S-M, Wu H-T, Chen P-W (2014) The modified rice αAmy8 promoter confers high-level foreign gene expression in a novel hypoxia-inducible expression system in transgenic rice seedlings. Plant Mol Biol 85:147–161 [DOI] [PubMed] [Google Scholar]
- Xie K, Yang Y (2019) A multiplexed CRISPR/Cas9 editing system based on the endogenous tRNA processing. Plant Genome Edit with CRISPR Sys: Methods and Protocols 1917:63–73. 10.1007/978-1-4939-8991-1_5 [DOI] [PubMed] [Google Scholar]
- Yang X-J, Seto E (2003) Collaborative spirit of histone deacetylases in regulating chromatin structure and gene expression. Curr Opin Genet Dev 13(2):143–153 [DOI] [PubMed] [Google Scholar]
- Yang J, Wang W, Chen Z, Lu S, Yang F, Bi Z, Bao L, Mo F, Li X, Huang Y (2020) A vaccine targeting the RBD of the S protein of SARS-CoV-2 induces protective immunity. Nature 586(7830):572–577 [DOI] [PubMed] [Google Scholar]
- Yu SM, Tzou WS, Lo WS, Kuo YH, Lee HT, WU R (1992) Regulation of α-amylase-encoding gene expression in germinating seeds and cultured cells of rice. Gene 122(2):247–253 [DOI] [PubMed] [Google Scholar]
- Yuki Y, Nojima M, Kashima K, Sugiura K, Maruyama S, Kurokawa S, Yamanoue T, Nakahashi-Ouchida R, Nakajima H, Hiraizumi T (2022) Oral MucoRice-CTB vaccine is safe and immunogenic in healthy US adults. Vaccine 40(24):3372–3379 [DOI] [PubMed] [Google Scholar]
- Yuki Y, Kurokawa S, Sugiura K, Kashima K, Maruyama S, Yamanoue T, Honma A, Mejima M, Takeyama N, Kuroda M (2024) MucoRice-CTB line 19A, a new marker-free transgenic rice-based cholera vaccine produced in an LED-based hydroponic system. Front Plant Sci 15:1342662 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zimran A, Gonzalez-Rodriguez DE, Abrahamov A, Cooper PA, Varughese S, Giraldo P, Petakov M, Tan ES, Chertkoff R (2018) Long-term safety and efficacy of taliglucerase alfa in pediatric Gaucher disease patients who were treatment-naive or previously treated with imiglucerase. Blood Cells Mol Dis 68:163–172 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
No datasets were generated or analysed during the current study.