Abstract
Computer vision adjunctive technology for oral lesion diagnoses has been developed to detect and identify Oral Potentially Malignant Disorders (OPMDs) and non-OPMDs. The early detection of OPMDs can reduce the risk of oral cancer development, improving the survival rate of the patients. This study aims to evaluate the computer vision technique in the white oral lesion domain within the scope of photographic images. Deep learning techniques for the classification of Convolution Neural Networks (CNNs) and transformer neural networks, and one-stage models of YOLOv7 and YOLOv8 were utilized to classify and detect five classes of OPMDs and non-OPMDs oral white lesions including oral leukoplakia, oral lichen planus, pseudomembranous candidiasis, oral ulcers covered with pseudomembrane and other white benign oral lesions. From the evaluation results of classification, the IFormerBase model achieves overperformance compared to CNN models with accuracy, precision, and F1 score of more than 80% on the test set. The best model for object detection is YOLOv7 with 84.5% mean Average Precision (mAP) at Intersection over Union (IoU) threshold of 0.3 and 74.5% at IoU of 0.5 on the test set. Object detection results reveal promising automatic oral lesion identification, which can be further developed to enhance the lesion screening system.
Supplementary Information
The online version contains supplementary material available at 10.1038/s41598-025-14450-w.
Keywords: Oral white lesions, Deep learning, Classification, Object detection, Cancer
Subject terms: Oral cancer, Biomedical engineering, Software
Introduction
Oral lesions are abnormalities that appear on oral mucosa and can be caused by various factors such as infections, injuries, immune alteration or underlying medical conditions. They can be classified according to the apparent colors of the lesion into white, red, red and white, and pigmented oral lesions1. Among these lesions, oral white lesions are common lesions that dentists frequently encounter. The diagnosis of oral white lesions can range from benign reactive conditions to more severe dysplastic and cancerous ones. Although certain characteristic features can aid in differentiating these lesions, the presence of similar patterns of the lesions can sometimes complicate the diagnosis. Oral white lesions are categorized either as acquired or congenital2. There are two types of acquired oral white lesions: those that can be scraped off and those that are not responsive to scraping. Five oral white lesions were examined in this study, two of which were scrapeable and three of which were not. These lesions consist of oral leukoplakia, Oral Lichen Planus (OLP), pseudomembranous candidiasis, oral ulcers covered with pseudomembrane, and other white benign oral lesions encompassing linea alba buccalis. The appearance of a white lesion can be used to identify the type of lesion. The classic appearance of pseudomembranous candidiasis is the curdled milk-like lesion that is scattered throughout the oral cavity and can be wiped off3. Considering oral ulcerative lesions, an epithelial defect in the ulcerative process is typically covered by a pseudomembrane made up of necrotic cells and fibrin. This condition is often observed in aphthous ulcers, erythema multiforme, and various other ulcerative disorders of the oral cavity. The fibrin clot’s color ranges from white to dirty yellow-white or grayish white4. Oral leukoplakia has been defined as a white patch or plaque that cannot be attributed to any clinically or histologically definite lesion5. Oral leukoplakia appears as an irreversible, non-scrapable, slightly elevated white plaque with a potentially wrinkled and leathery texture. These lesions are categorized into homogenous and non-homogenous types. The homogenous type is characterized by a smooth, white surface with well-defined edges. Treatment involves identifying and eliminating potential causes and performing an excisional biopsy. This condition requires close monitoring because it is considered a potentially malignant disorder. OLP has various clinical manifestations including papular, reticular, plaque-like, bullous, erythematous, and ulcerative features6. Fine white lines or striae, also called Wickham’s striae, comprise the reticular feature of OLP. Systemic and topical steroids are the main treatment of OLP. OLP is considered as a potentially malignant disorder, therefore, precise and frequent follow up should be advocated. Linea alba buccalis appears as a raised white line running along the plane where the upper and lower teeth occlude. It is considered normal, however, in some individuals with the parafunctional habit, the white line can be prominent. The fact that these five lesions are more common and that various forms of oral white lesions necessitate distinct treatments, some of them also require prompt identification and careful observation. General dentists’ diagnoses and healthcare professionals’ manual inspection and identification of oral white lesions can be subjective and time-consuming. The difference in appearance of a lesion brings about the question of the ability of deep learning to recognize and identify the type of oral lesion. Therefore, automated computer systems are attractive for identifying the type of oral lesion. Consequently, research has shown that the application of automated image analysis techniques for the classification and object detection of oral lesions can greatly enhance and expedite the diagnostic procedure.
Applying deep learning to medical diagnosis demonstrates the effectiveness in forecasting the outcomes, which is significantly boosted when compared to the reliance on a visual diagnostic technique7–9. Classification and object detection are techniques in machine learning and computer vision that allow a model to make predictions about an input. The classification model is trained to assign a label or class to an input sample. The goal of classification is an accurate prediction of an unseen input based on the features it implements. Object detection model is trained to locate objects within an image and classify them. This process typically involves training a model to detect an object by learning to classify regions within an image and then drawing a bounding box around the detected object10. Many published research papers reveal the successful classification and object detection of oral lesions as shown in Table 1. The classification models were achieved by the Convolutional Neural Network (CNN) architectures including VGG, ResNet, AlexNet, MobileNet, InceptionNet, SqueezeNet, Swin-S, etc. with adjustable parameters such as the layers for screening the oral lesion based on the photographic and histopathological images. The detection models have been performed by several state-of-the-art models such as Faster-RCNN, Mask R-CNN, and YOLO. These demonstrate further advancements in the automated classification and detection of Oral Potentially Malignant Disorders (OPMDs), which are promising strategies to enhance diagnostic efficiency.
Table 1.
Summary of related works.
| Author | Title | Summarized description |
|---|---|---|
| Welikala R. A., et al. (2020)11 |
Automated detection and classification of oral lesions using deep learning for early detection of oral cancer |
Dataset: The binary class of lesion and no-lesion, referral and non-referral, and the multiclass including the categories of high risk and low risk of OPMD were presented. Originality: The automated detection and classification for the early detection of oral cancer was conducted by ResNet101 for classification and Faster-RCNN for object detection. Result: The achievement of image classification was F1 score of 87.07% for identification of images that contained lesions and 78.30% for the identification of images that required referral. Object detection achieved an F1 score of 41.18% for the detection of lesions that required referral. |
| Lin H., et al. (2021)12 | Automatic detection of oral cancer in smartphone-based images using deep learning for early diagnosis |
Dataset: The multiclass dataset of oral lesions including aphthous ulcer, low-risk OPMD, high-risk OPMD, and cancer was used. Oral lesion images were collected from four different smartphone types. Originality: Three CNN architectures of VGG16, ResNet50, and DenseNet169 were implemented and compared with their proposed method (HRNet). Result: The performance of the proposed method (HRNet) achieved a sensitivity of 83.0%, specificity of 96.6%, precision of 84.3%, and F1 score of 83.6% on 455 test images. |
| Tanriver G., et al. (2021)13 | Automated detection and classification of oral lesions using deep learning to detect oral potentially malignant disorders |
Dataset: Photographic images of 3 classes of oral lesions, including benign, OPMD, and carcinoma, were used to construct the models. Originality: Classification of different CNNs, segmentation of U-net backbone, Mask R-CNN with ResNet backbones, and object detection of YOLOv5 were implemented. Result: Test results of U-Net with EfficientNet-b7 applied test-time augmentation (TTA) achieved Dice test of 0.929. Mask-RCNN results on test set with ResNet backbones achieved approximate AP50 at 80% and Mask AP50 in the range of 70–80%. YOLOv5l achieved 0.953 on AP50 with TTA applied. The results of classification of 4 different CNN models were reported in precision, recall, and F1 score, which were similarly obtained in the range of 80–90% across all of the implemented CNN models. |
| Warin K., et al. (2021)14 | Automatic classification and detection of oral cancer in photographic images using deep learning algorithms |
Dataset: The binary class of oral photographic images including 350 images of oral squamous cell carcinoma and 350 images of normal oral mucosa was used to construct the models. Originality: The CNN architecture of DenseNet121 for classification and Faster R-CNN for object detection were conducted with the balanced binary class of datasets for oral cancer screening. Result: The classification accuracy of DenseNet121 model achieved a precision of 99%, a recall of 100%, an F1 score of 99%, a sensitivity of 98.75%, a specificity of 100%, and an area under the receiver operating characteristic curve of 99%. The detection accuracy of a Faster R-CNN model achieved a precision of 76.67%, a recall of 82.14%, an F1 score of 79.31%, and an area under the precision-recall curve of 0.79. |
| Warin K., et al. (2022)15 | AI-based analysis of oral lesions using novel deep convolutional neural networks for early detection of oral cancer |
Dataset: The dataset is oral photographic images of oral potentially malignant disorders (OPMDs) and oral squamous cell carcinoma (OSCC). Originality: The classification models were created by DenseNet-169, ResNet-101, SqueezeNet, and Swin-S. The object detection models were created by Faster R-CNN, YOLOv5, RetinaNet, and CenterNet2. The performance of trained models was assessed additionally by comparing oral and maxillofacial surgeons and general practitioners. Result: DenseNet-196 yielded the best multiclass image classification performance with AUC of 1.00 and 0.98 on OSCC and OPMD, respectively. The AUC of the best multiclass CNN-base object detection models, Faster R-CNN, was 0.88 and 0.64 on OSCC and OPMDs, respectively. In comparison with experts and general practitioners, CNN-based models showed comparable diagnostic performances to expert level in classifying OSCC and OPMDs on oral photographic images. |
| Das M., et al. (2023)16 | Automatic detection of oral squamous cell carcinoma from histopathological images of oral mucosa using deep convolutional neural network |
Dataset: Binary class of histopathological images including cancerous and non-cancerous leions was used. The images were preprocessed by the Gaussian filter. Originality: The author proposed 10-layer CNN architecture to construct the model and compared the performance with 7 CNN architectures of VGG16, VGG19, AlexNet, ResNet50, ResNet101, Mobile Net, and Inception Net for classification. Result: The proposed 10-layer CNN model outperformed the other comparative models with the highest accuracy of 0.9782. AlexNet, ResNet50, ResNet101, Mobile Net, and Inception Net achieved an accuracy of 0.88, 0.91, 0.89, 0.93, and 0.92, respectively. VGG16 and VGG19 yielded the lowest performance with an accuracy of 0.74 and 0.71, respectively. |
From the literature review, the recent research points to the classification and object detection for the early stage of cancer development of OPMDs, but no research focuses on the multiclass of oral white lesions in the defined classes of correlated OPMDs and certain non-OPMDs. Furthermore, the focused identification of clinically correlated well-defined OPMDs and non-OPMDs contributes a more comprehensive and targeted approach to early detection and diagnosis, which fills the identified research gap on the multiclass of oral white lesions. In this study, we aim to design and develop an automated classification and detection system for screening the type of oral white lesions. This system will classify multiple types of lesions, including OPMDs such as oral leukoplakia and OLP, and non-OPMDs like pseudomembranous candidiasis, oral ulcers covered with pseudomembrane, and other white benign oral lesions, resulting in a total of five classes in the dataset. The performance of each classification and object detection pipeline was examined and compared. This study achieves the purposes through the classification of the CNN and transformer neural network architectures and the object detection on the upgraded versions of YOLOv7 and YOLOv8. The system is expected to assist general clinicians in identifying white OPMDs and benign lesions in the oral cavity, ultimately contributing to improving patient survival rates.
Materials and methods
Dataset preparation
This study was approved by the Ethical Committee of the Faculty of Dentistry/Faculty of Pharmacy, Mahidol University COA.NO.MU-DT/PY-IRB 2021/092.2010, which was in full compliance with International Guidelines for Human Research Protection including the Helsinki Declaration, the Belmont Report, CIOMS Guideline, and the International Conference on Harmonization in Good Clinical Practice. The date of ethics approval was October 20, 2021. Since the images used in this study were intraoral images, participant identification was not applicable. Informed consent in Thai language was obtained from all participants in the study.
This study is a retrospective analysis using an archive of images taken between September 1, 2016, and September 1, 2021. The dataset was collected by 2 oral medicine specialists from the Faculty of Dentistry, Mahidol University. The oral lesions have been pathologically confirmed. Pseudomembranous candidiasis was confirmed using microbiological method (candida culture). The images of the oral cavity at the site of the lesion were taken using two smartphones. The dataset was generated into five oral white lesions, including leukoplakia, OLP, pseudomembranous candidiasis, ulcers covered with pseudomembrane, and other white benign oral lesions. The number of images of each class is 450 images of leukoplakia (from 152 participants), 273 images of OLP (from 133 participants), 238 images of pseudomembranous candidiasis (from 119 participants), 335 images of ulcer covered with pseudomembrane (from 115 participants), and 283 images of other white benign oral lesions (from 141 participants). The examples of image datasets of each class are shown in Fig. 1.
Fig. 1.
Examples of image dataset of each class; (a) Leukoplakia, (b) Lichen planus, (c) Pseudomembranous candidiasis, (d) Other white benign oral lesions, and (e) Ulcers covered with pseudomembrane.
The image augmentation was performed to increase the number of images and for the adjustable balance dataset and reduce the bias. Image augmentation techniques include rotation, shear, zoom, horizontal flip, and brightness. The number of augmented datasets for classification became 1000 images per class and 607 images per class for object detection. For classification, the dataset was resized to 600 × 600 pixels. The dataset was split into training and validation ratio of 80:20 for each class. The images dataset for object detection was not resized, and the training set and validation set ratio was split at 90:10 for each class. The unseen test set of 40 images for each class was initially separated from the training and validation set, which was prepared for evaluation of classification and object detection models.
The classification model construction
Deep learning models of the CNN known as keras applications are provided alongside pre-trained weights called transfer learning17. These models can be applied to feature extraction, prediction, and fine-tuning. In this research, the pre-trained CNN models were selected to compare the performance for classification, namely DenseNet12118, DenseNet20118, Xception19, and ResNet5020. Moreover, transformer neural networks were also implemented to compare the performance for classification, including Contextual Transformer Networks for Visual Recognition (CoTNet)21,22, Dual Attention Vision Transformers (DaViT)23, Inception Transformer (IFormer)24, and Fast Pretraining Distillation for Small Vision Transformers (TinyViT)25. These transformer models are applied from keras cv attention models library from python. All models were trained on NVIDIA RTX 2060 Graphics Processing Unit (GPU).
The original oral lesion images of 5 classes were resized to 600 × 600 pixels. These resized datasets were used to train those 8 models with input image dimensions of 100 × 100 pixels. Moreover, the post-training was carried out by defining the same hyperparameters and other values as optimizers of Adam, loss of categorical cross entropy, and learning rate of 0.0001. Callback functions are also provided in this training, including early stopping, model checkpoint, and reduceLRonPlateau. For the early stopping, the defined parameters include monitor of validation loss, patience of 100, and mode of min. For the model checkpoint, the name of model was defined by monitoring of validation loss, mode of min, and save_best_only of True for saving the best model from training that there is the lowest validation loss. For reduceLRonPlateau, we defined parameters including monitor of validation loss, factor of 0.1, patience of 7, min_delta of 10−4, and mode of min. All 8 models were trained for up to 300 epochs, but each was stopped at the different number of iterations due to the early stopping function, which was implemented to prevent overfitting and excessive training. After training was completed, the best model was saved and then loaded again to evaluate its performance on the test set.
The object detection model construction
A computer vision technique called object detection involves locating and identifying objects in an image. Object localization involves drawing a bounding box around the object to reveal its precise location, and object classification refers to identifying the category or class of the object within an image. YOLO architecture is part of a class of one-stage object detectors that integrates localization and classification tasks in a single network and, as a result, operates very quickly due to its straightforward design. In this study, state-of-the-art algorithms for object detection, YOLOv726 and YOLOv827 were performed to detect oral lesions. YOLOv7 is available in two versions, including YOLOv7 and YOLOv7x with the default test size equal to 640 × 640 pixels. YOLOv8 is also available in five versions, including YOLOv8n, YOLOv8s, YOLOv8m, YOLOv8l, and YOLOv8x. In order to avoid overfitting, pre-trained weights on the COCO dataset were used to initialize all four iterations of the model, which were then tested for their performance on the lesion detection task. Four YOLO models were trained with 100 epochs. All models were trained on NVIDIA RTX 2060 (GPU), which is the same as the classification task. The annotation of the lesions for ground truth bounding box creation and labeling was done manually using the LabelImg program.
Performance evaluation
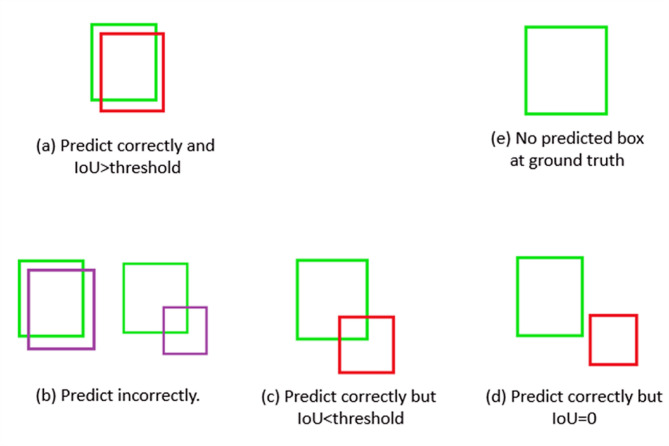
To evaluate the model’s performance, we utilized standard image classification metrics including precision, recall, F1 score, Receiver-Operating Characteristic (ROC) curve, and Area Under the Curve (AUC) score28. Moreover, Cohen’s Kappa score was calculated to measure the level of agreement between the predicted and actual classifications. The quality model’s prediction performance was analyzed using the Matthews Correlation Coefficient (MCC) score. The MCC ranges from − 1 to 1, where values near 1 indicate excellent prediction, reflecting a strong positive correlation between the predicted values and the actual labels29. For the object detection task, the performance of oral lesion localization and classification was evaluated by both validation and test datasets. The statistical analysis for object detection includes Intersection over Union (IoU), Average Precision (AP), Average Recall (AR), mean Average Precision (mAP), and the optimal Localization Recall Precision (oLRP)30. In the context of our study, green boxes are the ground truth. Red boxes are predicted correctly. Purple boxes are predicted incorrectly as other classes. The definitions of predicted outcomes are shown in Fig. 2. We evaluated the models using IoU and oLRP at thresholds of 0.3 and 0.5. The AR was assessed across two ranges of AR at 0.3–0.95 and AR at 0.5–0.95, providing a comprehensive evaluation of model performance across different detection thresholds.
Fig. 2.
Definition of true positive, false positive, and false negative. Green boxes are ground truth. Red boxes are predicted correctly. (a) is true positive, (b), (c), and (d) are false positive, (e) is false negative.
Results
Image classification
The loss, accuracy, precision, recall, and F1 score of validation and test sets for evaluated CNN and transformers models are reported in Table 2. All trained models offer validation accuracy, precision, and recall of more than 90% with small loss. The learning curve contains the accuracy and loss of the training-validation set of all 8 trained models as depicted in Supplementary Fig. S1 online. All trained models effectively learn the tasks, displaying improved accuracy and reduced loss, with stability approaching 1 for accuracy and 0 for loss. The resulting learning curves indicate that the validation and unseen test set outcomes reflect the models’ optimal performance, with no overfitting between the validation and unseen test sets. The results of confusion matrices of the unseen test set of all 8 trained models are depicted in Supplementary Fig. S2 online. The outcomes of True Positive (TP), True Negative (TN), False Positive (FP), and False Negative (FN) in confusion matrix were mathematically calculated to obtain evaluation metrics in Table 2.
Table 2.
The classification results of validation and test sets.
| Models | Validation | Unseen Test | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Loss | Accuracy | Precision | Recall | Accuracy | Precision | F1 score | Cohen’s Kappa | MCC | |
| DenseNet121 | 0.0223 | 0.994 | 0.994 | 0.993 | 0.780 | 0.798 | 0.780 | 0.775 | 0.776 |
| DenseNet201 | 0.0202 | 0.995 | 0.995 | 0.994 | 0.785 | 0.809 | 0.788 | 0.731 | 0.735 |
| Xception | 0.0290 | 0.990 | 0.992 | 0.989 | 0.770 | 0.791 | 0.769 | 0.712 | 0.717 |
| ResNet50 | 0.0340 | 0.990 | 0.992 | 0.988 | 0.790 | 0.795 | 0.784 | 0.737 | 0.740 |
| DaViT_T | 0.0108 | 0.996 | 0.996 | 0.996 | 0.840 | 0.846 | 0.840 | 0.800 | 0.801 |
| CotNetSE152D | 0.0166 | 0.993 | 0.993 | 0.993 | 0.835 | 0.852 | 0.833 | 0.793 | 0.798 |
| IFormerBase | 0.0096 | 0.996 | 0.997 | 0.996 | 0.860 | 0.874 | 0.861 | 0.825 | 0.827 |
| TinyViT_21M | 0.0331 | 0.994 | 0.995 | 0.992 | 0.830 | 0.848 | 0.833 | 0.787 | 0.790 |
The transformer model of IFormerBase correctly predicts the positive and negative classes of TP and TN and less incorrectly predicts the positive and negative classes of FP and FN compared to other trained models giving better performance results (Fig. 3). However, testing the trained model by the unseen test set, the models predict lower results of less than 90% expectation. The highest performance is accepted from IFormerBase model of more than 86% of all metrics. The Cohen’s Kappa and MCC score metrics for all classification models were measured on the unseen test, with results ranging from 0.7 to 0.8. These scores indicate a high level of agreement between the model’s predictions and the true labels, demonstrating a strong positive correlation29.
Fig. 3.
Results of confusion matrices of unseen test on IFormerBase model.
The ROC curve and AUC score of the validation set of all trained models are depicted in Supplementary Fig. S3 online. The results are predictable to achieve above 0.9 of AUC score of all models, which is a high accuracy for confidence interval to distinguish the positive and negative classes31. The IFormerBase model demonstrated the best performance, as indicated by its results across various metrics. The results of ROC curve and AUC score of IFormerBase are presented in Fig. 4.
Fig. 4.
Validation set results of ROC curve and AUC score of IFormerBase.
Object detection
Different versions of YOLOv7 and YOLOv8 were evaluated for the five-class lesion detection task. Model hyperparameters were optimized based on the performance of the validation set for all models. Validation results of each model are reported in Table 3.
Table 3.
Multiclass image detection results on the validation set.
| Detection models | Class | P | R | mAP@0.5 | mAP@0.5:0.95 |
|---|---|---|---|---|---|
| YOLOv7 | All | 0.717 | 0.687 | 0.673 | 0.363 |
| Leukoplakia | 0.734 | 0.789 | 0.768 | 0.465 | |
| Lichen planus | 0.668 | 0.732 | 0.651 | 0.338 | |
| Candidiasis | 0.486 | 0.587 | 0.469 | 0.217 | |
| Other white | 0.838 | 0.626 | 0.72 | 0.406 | |
| Ulcer | 0.858 | 0.701 | 0.759 | 0.39 | |
| YOLOv7x | All | 0.694 | 0.676 | 0.641 | 0.351 |
| Leukoplakia | 0.687 | 0.789 | 0.737 | 0.45 | |
| Lichen planus | 0.675 | 0.701 | 0.587 | 0.303 | |
| Candidiasis | 0.496 | 0.586 | 0.465 | 0.215 | |
| Other white | 0.758 | 0.654 | 0.68 | 0.388 | |
| Ulcer | 0.855 | 0.647 | 0.734 | 0.4 | |
| YOLOv8n | All | 0.563 | 0.532 | 0.55 | 0.303 |
| Leukoplakia | 0.573 | 0.617 | 0.627 | 0.397 | |
| Lichen planus | 0.495 | 0.584 | 0.583 | 0.331 | |
| Candidiasis | 0.528 | 0.523 | 0.504 | 0.247 | |
| Other white | 0.675 | 0.542 | 0.596 | 0.32 | |
| Ulcer | 0.543 | 0.394 | 0.439 | 0.218 | |
| YOLOv8x | All | 0.569 | 0.454 | 0.474 | 0.258 |
| Leukoplakia | 0.609 | 0.617 | 0.625 | 0.416 | |
| Lichen planus | 0.486 | 0.403 | 0.412 | 0.186 | |
| Candidiasis | 0.541 | 0.494 | 0.477 | 0.226 | |
| Other white | 0.601 | 0.467 | 0.5 | 0.297 | |
| Ulcer | 0.611 | 0.292 | 0.356 | 0.166 |
The metrics were computed for the mAP over IoU thresholds at 0.5 (mAP@0.5) and from 0.5 to 0.95 (mAP@0.5:0.95). YOLOv7 performs the best, as shown by the Precision-Recall (PR) curve plots and mAP of the validation set presented in Fig. 5.
Fig. 5.
The PR-curve plots of four detection models: (a) YOLOv7 model, (b) YOLOv7x model, (c) YOLOv8n model, and (d) YOLOv8x model.
The AP of the unseen test sets is reported in Table 4. We conducted trial evaluations of the models at IoU thresholds of 0.3 and 0.5. It was found that at an IoU of 0.3, the models could satisfactorily localize lesion areas, achieving a mAP of 0.845, which is higher than the 0.745 mAP obtained at an IoU of 0.5 using YOLOv7. The Optimal Localization Recall Precision (oLRP), Average Recall (AR), and the number of bounding box predictions for each model are presented to evaluate model performance. These metrics consider localization errors, recall, and precision, providing a more accurate assessment of true detections and false detections30.
Table 4.
Multiclass image detection results of the unseen test set.
| Models | IoU threshold 0.3 | IoU threshold 0.5 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AP | mAP | AP | mAP | |||||||||
| Leukoplakia | Lichen planus | Candidiasis | Other white | Ulcer | Leukoplakia | Lichen planus | Candidiasis | Other white | Ulcer | |||
| YOLOv7 | 0.895 | 0.697 | 0.771 | 0.961 | 0.899 | 0.845 | 0.798 | 0.504 | 0.662 | 0.941 | 0.818 | 0.745 |
| YOLOv7x | 0.920 | 0.675 | 0.726 | 0.968 | 0.778 | 0.813 | 0.820 | 0.566 | 0.627 | 0.936 | 0.633 | 0.717 |
| YOLOv8n | 0.780 | 0.612 | 0.844 | 0.824 | 0.861 | 0.784 | 0.684 | 0.454 | 0.732 | 0.824 | 0.829 | 0.705 |
| YOLOv8x | 0.803 | 0.535 | 0.824 | 0.838 | 0.868 | 0.774 | 0.711 | 0.275 | 0.715 | 0.795 | 0.841 | 0.668 |
The high AR values achieved by YOLOv7 and YOLOv7x across multiple classes, as shown in Table 5, indicate strong object detection capabilities, minimizing missed detections. The results confirm that the models generate an optimal number of bounding boxes, providing insights into the number of detections per class while effectively capturing relevant objects. The oLRP for all models has been computed using an optimal threshold selection approach, ensuring an accurate evaluation of detection quality, with values ranging between 0.1 and 0.3. Among the evaluated models, YOLOv7 demonstrated the best performance, as evidenced in Tables 4 and 5.
Table 5.
Evaluation metrics for object detection: average recall, bounding box predictions, and oLRP.
| Model | IoU threshold | The quantity of predicted boxes per class | AR | oLRP | |||||
|---|---|---|---|---|---|---|---|---|---|
| Leukoplakia | Lichen planus | Candidiasis | Other white | Ulcer | Total | ||||
| YOLOv7 | 0.3 | 84 | 109 | 264 | 49 | 73 | 579 | 0.949 | 0.169 |
| 0.5 | 84 | 109 | 264 | 49 | 73 | 579 | 0.931 | 0.187 | |
| YOLOv7x | 0.3 | 80 | 96 | 184 | 53 | 78 | 491 | 0.942 | 0.172 |
| 0.5 | 80 | 96 | 184 | 53 | 78 | 491 | 0.926 | 0.189 | |
| YOLOv8 | 0.3 | 53 | 34 | 106 | 31 | 46 | 270 | 0.631 | 0.217 |
| 0.5 | 53 | 34 | 106 | 31 | 46 | 270 | 0.582 | 0.244 | |
| YOLOv8x | 0.3 | 60 | 32 | 125 | 34 | 47 | 298 | 0.633 | 0.227 |
| 0.5 | 60 | 32 | 125 | 34 | 42 | 293 | 0.583 | 0.253 | |
An example of YOLOv7’s predicted results for each of the five classes with a confidence level greater than 0.5 is shown in Fig. 6. The post-trained model of YOLOv7 could localize lesions by drawing the predicted bounding boxes overlapping the ground truth area precisely. Moreover, the model could satisfactorily classify each class of oral white lesions.
Fig. 6.
Example of images detected by YOLOv7. Green box is the ground truth. The predicted boxes are: (a) Leukoplakia (grey), (b) Lichen planus (red), (c) Pseudomembranous candidiasis (purple), (d) Other white benign oral lesions (orange), and (e) Ulcer covered with pseudomembrane (blue).
Discussion
Some types of OPMDs, including leukoplakia and OLP, carry the highest risk of developing into oral cancer. While pseudomembranous candidiasis, ulcers covered with pseudomembrane, and other white benign oral lesions have very low risk of developing into oral cancer. Properly categorizing oral lesions is crucial in clinical practice to increase patients’ survival rate from early diagnosis. The different types of CNN and transformer neural network models were evaluated for the multiclass classification of oral white lesions based on the risk of cancerous transformation. Among the 8 models, IFormerBase transformer model showed the highest accuracy of 0.86, precision of 0.874, and F1 score of 0.861 followed by DaViT_T transformer model, which achieved an accuracy of 0.84, precision of 0.846, and F1 score of 0.840. The state-of-the-art transformer models with complex architectures outperformed among the other models evaluated for the classification task. Misclassification of the IFormerBase model was observed as the leukoplakia class had the lowest precision due to the FP of six OLPs, two candidiasis, one other white, and four ulcers were misclassified as shown in Fig. 3. OLP had the lowest recall due to the FN of six images were predicted as leukoplakia, and the other 4 were mispredicted. However, this misclassification of OPL as leukoplakia may not pose a significance since both types of lesions are OPMDs, which detected either should be further diagnosed by medical specialists or pathological diagnosis immediately. On the other hand, incorrectly labeling benign lesions (candidiasis, other white, and ulcer) as leukoplakia or OPL may result in referrals to oral cancer experts, which increases the workload of the clinical staff. However, the recall rate for leukoplakia was rather good, which is reassuring for the critical leukoplakia identification for oral cancer screening. Nevertheless, improving prediction performance still requires ensuring reliability. The performance of multiclass classification is usually lower than that of binary classification, as reported in some studies10,13,32. The performance metrics could reach 100% by DenseNet121 for Oral Squamous Cell Carcinoma (OSCC) and normal oral mucosa13. Large amounts of qualitative medical images are an important key to achieve the goal of high-performance assistant diagnosis33. In addition, the development of deep learning models on purpose for object localization with classification can improve the confidence of automated medical image diagnosis.
The potential of YOLOv7 for early detection of oral cancer was confirmed and reported by Hsu Y., et al., 202434. The lesions were categorized into three classes according to referral grades: benign (green), potentially malignant (yellow), and malignant (red). The YOLOv7 models, particularly the YOLOv7-E6, demonstrated high precision and recall across all lesion categories. The YOLOv7 backbone, implemented in the YOLOv7-D6 model, demonstrated excellent performance in identifying malignant lesions, achieving precision, recall, and F1 scores of approximately 0.7. Additionally, the AP@0.5 was 0.758, and the AP@0.5:0.95 was 0.4533. Comparing the object detection in this study, YOLOv7 model achieves over other models, with mAP@0.5 of 0.673 and mAP@0.5:0.95 of 0.363 on the validation set, as shown in Table 3. From results for OPMDs of leukoplakia and OLP, the YOLOv7 also showed notable precision, recall, and mAP@0.5 results approximated to 0.7. For OLP, the precision and mAP@0.5 were in the range of 0.6–0.7. When mAP@0.5:0.95 was investigated, the values were 0.465 for leukoplakia and 0.338 for OLP. For both results of mAP@0.5 and mAP@0.5:0.95 on YOLOv7, leukoplakia shows the highest predicted performance among 5 classes. Fortunately, oral leukoplakia appears as a slightly elevated white plaque with a potentially wrinkled and leathery texture. This condition requires close monitoring because it is considered a potentially malignant disorder.
The performance of the state-of-the-art YOLOv7 and YOLOv8 architecture was additionally evaluated for the detection and classification of oral white lesions by the unseen test set. The unseen test was performed on a different dataset, separate from the one used for training the models to ensure generalization to real-world scenarios. The validation thresholds were set at 0.3 and 0.5 on the unseen test, as shown in Table 4. The YOLOv7 yields the best performance results with mAP@0.3 of 0.845 and mAP@0.5 of 0.745. Table 5 presents an analysis of FP and FN in relation to AR and oLRP. The results indicate that YOLOv7 remains the best-performing model, particularly for applications requiring a wider range of object detection. However, the upgraded versions of YOLOv8n and YOLOv8x did not improve performance. The mAP of both YOLOv8n and YOLOv8x was lower than that of YOLOv7 in both versions. Furthermore, we have operated upgraded versions of YOLOv9, YOLOv10, YOLOv11, and YOLOv12. However, the results were unsatisfactory as the upgraded version did not improve the performance as expected. The mAP@0.5 was 0.684, 0.599, 0.584, and 0.598 for YOLOv9, YOLOv10, YOLOv11x, and YOLOv12x, respectively. These results indicate that the object detection models can identify the target area of the lesion and correctly classify it satisfactorily, and meet the purpose of OPMDs identification. Therefore, automated detection of oral lesions by object detection is promising to help the general dentist identify the OPMD cases. However, applying computer vision requires improvement to obtain more confidence in usability.
The strength of this study lies in its novelty, as no previous research has explored different types of oral white lesions. Since each lesion exhibits a unique pattern, deep learning models could assist clinicians in distinguishing between them and enable faster, more accurate diagnoses. Furthermore, eight CNN and transformer models were used for classification, while four YOLO architectures were employed for object detection.
Despite these strengths, certain limitations remain. Applying deep learning for automated oral lesion identification in real-world scenarios requires a variety of datasets to improve the model’s generalizability. This study scopes on a single dataset, which was obtained from a single source, in which our study focuses on data collected from the Faculty of Dentistry, Mahidol University, following ethical guidelines and approvals from the Institutional Review Board (IRB). Only two smartphones were used for capturing the images, and only two oral medicine specialists participated in the image collection process. This limitation hinders the generalization of the models to datasets from new sources. Moreover, external validation with other resources is crucial for the development of the practical application. However, to ensure that the model did not overfit the training dataset, its evaluation was conducted using an unseen dataset that the models had not encountered during the training process. Therefore, the unseen test in this study could basically overcome the limitation of the single dataset and confirm the reliability of prediction results. Interestingly, understanding the internal workings and decision-making process of deep learning models, interpretability, and explainability are valuable areas of future study. Integrating an attention heat map that visualizes the area the classification model uses to make a decision helps users understand how the model reaches its decisions, thereby fostering the trust essential for medical reliability, and effectively collaborating with the model35–37. The Grad-CAM technique for CNNs has been successfully applied to the DenseNet201 model. In cases of correct predictions, the method accurately highlighted the crucial regions of the lesion, while in cases of incorrect predictions, the highlighted areas deviated from the true lesion area. These results demonstrate the value of Explainable AI (XAI) in revealing model decision processes, supporting its application as a promising approach to enhance the transparency and reliability of deep learning models in a medical context. The YOLOv7 model spent 14.5 h training and up to 21 h for YOLOv7x. This demonstrates that users can update the input dataset and retrain the model within 24 h, which is reasonably efficient and supports the model’s reproducibility in the real-world clinical setting. For the next phase of the research, the study will include a larger group of external Institutes of dentists and a variety of smartphones and cameras to increase the diversity and quantity of the collected images for further improvement in the generalizability of the model and practical application. The image dataset will not only be limited to OPMDs, but oral cancer images will also be included. The smartphone-based application is being developed to provide an accessible and practical tool for oral lesion detection. Transfer learning is a viable approach for adapting the model to new environments. Future research will explore fine-tuning strategies to leverage knowledge from our current dataset and improve performance on external datasets.
Conclusion
This study proposes to evaluate the potential of deep learning for multiclass oral white lesions, including OPMDs; leukoplakia, OLP, and non-OPMDs; pseudomembranous candidiasis, ulcers covered with pseudomembrane, and other white benign oral lesions by comparing the performance of the 8 models for classification and 4 models for object detection. For the classification task, the transformer model of IFormerBase outperforms compared to CNN models. For the object detection task, the post-trained model of YOLOv7 showed satisfactory detection and classification of all 5 classes. YOLOv7 model outperforms at the highest mAP@0.3 of 0.845 and mAP@0.5 of 0.745. For practical applications, the YOLOv7 model with an IoU threshold of 0.3 demonstrates optimal performance for detecting and screening various types of oral white lesions. Consequently, this model can be considered to create an application for early detection in the future. Furthermore, external validation is essential to enhance the model’s generalization for real-world applications. To achieve this, we are actively collaborating with larger groups of Institutes to obtain a more diverse dataset. This is anticipated to be an innovative diagnostic tool to help general practitioners and general dentists preliminarily screen the lesion and enhance expert-level decision-making in the oral cancer screening program.
Supplementary Information
Below is the link to the electronic supplementary material.
Acknowledgements
This research was conducted as part of the research project titled ‘Classification and identification of oral precancerous lesion by deep learning technology’ (grant number RE-KRIS/FF68/63) by King Mongkut’s Institute of Technology Ladkrabang (KMITL), with funding support from National Science, Research and Innovation Fund (NSRF).
Author contributions
The study was conceptualized and designed by S.P.K, B.T, S.P, and T.T. Data collection was done by S.P.K and D.R. Data analysis and experiment were performed by K.P, S.P, and T.T. The manuscript drafting and revision were carried out by K.P, S.P.K, and T.T. All authors made contributions to the article and approved the published version of the manuscript.
Data availability
The datasets used and/or analyzed during the current study are not publicly available due to the confidentiality of the participants but are available from the corresponding author on reasonable request.
Declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the Ethical Committee of the Faculty of Dentistry/Faculty of Pharmacy, Mahidol University COA.NO.MU-DT/PY-IRB 2021/092.2010, which was in full compliance with International Guidelines for Human Research Protection including the Helsinki Declaration, the Belmont Report, CIOMS Guideline, and the International Conference on Harmonization in Good Clinical Practice. The date of ethics approval was 20 October 2021. Since the images used in this study were intraoral images, participant identification was not applicable. Informed consent in Thai language was obtained from all the participants in the study.
Footnotes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Suvit Poomrittigul, Email: suvit@it.kmitl.ac.th.
Treesukon Treebupachatsakul, Email: treesukon.tr@kmitl.ac.th.
References
- 1.Zahid, E., Bhatti, O., Zahid, M. A. & Stubbs, M. Overview of common oral lesions. Malays Fam Physician. 17 (3), 9–21. 10.51866/rv.37 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Mortazavi, H. et al. Oral white lesions: an updated clinical diagnostic decision tree. Dent. J. (Basel). 7 (1), 15. 10.3390/dj7010015 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Hellstein, J. W. & Marek, C. L. Candidiasis: red and white manifestations in the oral cavity. Head Neck Pathol.13 (1), 25–32. 10.1007/s12105-019-01004-6 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Philipone, E. M. & Peters, S. M. Ulcerative and inflammatory lesions of the oral mucosa. Oral Maxillofac. Surg. Clin. North. Am.35 (2), 219–226. 10.1016/j.coms.2022.10.001 (2023). [DOI] [PubMed] [Google Scholar]
- 5.van der Waal, I. Oral leukoplakia, the ongoing discussion on definition and terminology. Med. Oral Patol. Oral Cir. Bucal. 20 (6), e685–e692. 10.4317/medoral.21007 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Chiang, C. P. et al. Oral lichen planus - Differential diagnoses, serum autoantibodies, hematinic deficiencies, and management. J. Formos. Med. Assoc.117 (9), 756–765. 10.1016/j.jfma.2018.01.021 (2018). [DOI] [PubMed] [Google Scholar]
- 7.Adate, A., Arya, D., Shaha, A. & Tripathy, B. Impact of deep neural learning on artificial intelligence research. In: Bhattacharyya, S., Snasel, V., Ella Hassanien, A., Saha, S., Tripathy, B. (eds) Deep Learning: Research and Applications, Berlin, Boston: De Gruyter. 69–84 (2020). 10.1515/9783110670905-004
- 8.Marks, R. The epidemiology of non-melanoma skin cancer: who, why and what can we do about it. J. Dermatol.22 (11), 853–857. 10.1111/j.1346-8138.1995.tb03935.x (1995). [DOI] [PubMed] [Google Scholar]
- 9.Ozsunkar, P. S. et al. Detecting white spot lesions on post-orthodontic oral photographs using deep learning based on the YOLOv5x algorithm: a pilot study. BMC Oral Health. 24, 490. 10.1186/s12903-024-04262-1 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Géron, A. Hands-on Machine Learning with scikit-learn, Keras, and Tensorflow: Concepts, Tools, and Techniques To Build Intelligent Systems 2nd edn (O’Reilly Media, Inc, 2019).
- 11.Welikala, R. A. et al. Automated detection and classification of oral lesions using deep learning for early detection of oral cancer. IEEE Access.8, 132677–132693. 10.1109/ACCESS.2020.3010180 (2020). [Google Scholar]
- 12.Lin, H., Chen, H., Weng, L., Shao, J. & Lin, J. Automatic detection of oral cancer in smartphone-based images using deep learning for early diagnosis. J. Biomed. Opt.26 (8), 086007. 10.1117/1.JBO.26.8.086007 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Tanriver, G., Soluk Tekkesin, M. & Ergen, O. Automated detection and classification of oral lesions using deep learning to detect oral potentially malignant disorders. Cancers (Basel). 13 (11), 2766. 10.3390/cancers13112766 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Warin, K., Limprasert, W., Suebnukarn, S., Jinaporntham, S. & Jantana, P. Automatic classification and detection of oral cancer in photographic images using deep learning algorithms. J. Oral Pathol. Med.50 (9), 911–918. 10.1111/jop.13227 (2021). [DOI] [PubMed] [Google Scholar]
- 15.Warin, K. et al. AI-based analysis of oral lesions using novel deep convolutional neural networks for early detection of oral cancer. PLoS ONE. 17 (8), e0273508. 10.1371/journal.pone.0273508 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Das, M., Dash, R. & Mishra, S. K. Automatic detection of oral squamous cell carcinoma from histopathological images of oral mucosa using deep convolutional neural network. Int. J. Environ. Res. Public. Health. 20 (3), 2131. 10.3390/ijerph20032131 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Cullell-Dalmau, M., Noe, S., Otero-Vinas, M., Meic, I. & Manzo, C. Convolutional neural network for skin lesion classification: Understanding the fundamentals through hands-on learning. Front. Med. (Lausanne). 8, 644327. 10.3389/fmed.2021.644327 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Huang, G., Liu, Z., Van Der Maaten, L. & Weinberger, K. Q. (eds) Densely connected convolutional networks. Computer vision and pattern recognition. Preprint at (2017). 10.48550/arXiv.1608.06993
- 19.Chollet, F. Xception Deep learning with depthwise separable convolutions. 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 1800–1807 (2017). 10.1109/CVPR.2017.195
- 20.He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. Computer vision and pattern recognition. Preprint At.10.48550/arXiv.1512.03385 (2015). [Google Scholar]
- 21.Ali, A. M. et al. Vision Transformers in image restoration: A survey. Sens. (Basel). 23 (5), 2385. 10.3390/s23052385 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Li, Y., Yao, T., Pan, Y. & Mei, T. Contextual transformer networks for visual recognition. IEEE Trans. Pattern Anal. Mach. Intell.45 (2), 1489–1500. 10.1109/TPAMI.2022.3164083 (2022). [DOI] [PubMed] [Google Scholar]
- 23.Ding, M. et al. Dual attention vision transformers. Computer Vision (ECCV 2022), Lecture notes in computer science. 13684, 74–92 (2022). 10.1007/978-3-031-20053-3_5
- 24.Si, C. et al. Inception transformer. Computer vision and pattern recognition. Preprint At.10.48550/arXiv.2205.12956 (2022). [Google Scholar]
- 25.Wu, K. et al. Tinyvit: Fast pretraining distillation for small vision transformers. Computer Vision (ECCV 2022), Lecture notes in computer science. 13681, 68–85 (2022). 10.1007/978-3-031-19803-8_5
- 26.Wang, C. Y., Bochkovskiy, A. & Liao, H-Y-M. YOLOv7: Trainable bag-of-freebies sets new state-of-the-art for real-time object detectors. Proceedings of the IEEE/CVF conference on computer vision and pattern recognition (CVPR). Preprint at. Preprint at (2023). 10.48550/arXiv.2207.02696 (2023).
- 27.Reis, D., Kupec, J., Hong, J. & Daoudi, A. Real-time flying object detection with YOLOv8. Computer vision and pattern recognition. Preprint At.10.48550/arXiv.2305.09972 (2023). [Google Scholar]
- 28.Çorbacıoğlu, Ş. K. & Aksel, G. Receiver operating characteristic curve analysis in diagnostic accuracy studies: A guide to interpreting the area under the curve value. Turk. J. Emerg. Med.23 (4), 195–198. 10.4103/tjem.tjem_182_23 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Margherita, G., Enrico, B. & Giorgio, V. Metrics for multi-class classification: an overview. ArXiv10.48550/arXiv.2008.05756 (2020). abs/2008.05756. [Google Scholar]
- 30.Oksuz, K., Cam, B. C., Akbas, E. spsampsps Kalkan, S. Localization recall precision (LRP): A new performance metric for object detection. In: Ferrari, V., Hebert, M., Sminchisescu, C., Weiss, Y. (eds) Computer Vision – ECCV 2018. Lecture Notes in Computer Science, 11211, 521–537 (2018). 10.1007/978-3-030-01234-2_31
- 31.Padilla, R., Netto, S. L. & Da Silva, E. A. A survey on performance metrics for object-detection algorithms. 2020 Int. Conf. Syst. Signals Image Process. (IWSSIP). 10.1109/IWSSIP48289.2020.9145130 (2020). [Google Scholar]
- 32.Warin, K., Limprasert, W., Suebnukarn, S., Jinaporntham, S. & Jantana, P. Performance of deep convolutional neural network for classification and detection of oral potentially malignant disorders in photographic images. Int. J. Oral Maxillofac. Surg.51 (5), 699–704. 10.1016/j.ijom.2021.09.001 (2022). [DOI] [PubMed] [Google Scholar]
- 33.Fourcade, A. & Khonsari, R. H. Deep learning in medical image analysis: A third eye for Doctors. J. Stomatol. Oral Maxillofac. Surg.120 (4), 279–288. 10.1016/j.jormas.2019.06.002 (2019). [DOI] [PubMed] [Google Scholar]
- 34.Hsu, Y. et al. Oral mucosal lesions triage via YOLOv7 models. J. Formos. Med. Assoc.10.1016/j.jfma.2024.07.010 (2024). [DOI] [PubMed] [Google Scholar]
- 35.Preechakul, K., Sriswasdi, S., Kijsirikul, B. & Chuangsuwanich, E. Improved image classification explainability with high-accuracy heatmaps. iScience25 (3), 103933. 10.1016/j.isci.2022.103933 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Tjoa, E., Khok, H. J., Chouhan, T. & Guan, C. Enhancing the confidence of deep learning classifiers via interpretable saliency maps. Neurocomputing562, 126825. 10.1016/j.neucom.2023.126825 (2023). [Google Scholar]
- 37.Storås, A. M. et al. Usefulness of heat map explanations for Deep-Learning-Based electrocardiogram analysis. Diagnostics (Basel Switzerland). 13 (14), 2345. 10.3390/diagnostics13142345 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets used and/or analyzed during the current study are not publicly available due to the confidentiality of the participants but are available from the corresponding author on reasonable request.