Abstract
A naturally occurring atrazine-resistant cyanobacterial isolate, strain SG2, was isolated from an atrazine-containing wastewater treatment system at the Syngenta atrazine production facility in St. Gabriel, La. Strain SG2 was resistant to 1,000 μg of atrazine per ml but showed relatively low resistance to diuron [3-(3,4-dichlorophenyl)-1,1-dimethyl urea]. Analyses of 16S ribosomal DNA indicated that strain SG2 falls into the Synechocystis/Pleurocapsa/Microcystis group. Photosynthetically driven oxygen evolution in strain SG2 was only slightly inhibited (about 10%) by 2,000 μg of atrazine per ml, whereas in the control strain Synechocystis 6803, oxygen evolution was inhibited 90% by 1,000 μg of atrazine per ml. No atrazine accretion, mineralization, or metabolites were detected when strain SG2 was grown with [14C]atrazine. Strain SG2 contained three copies of the psbA gene, which encodes the D1 protein of the photosystem II reaction center. Nucleotide sequence analyses indicated that the psbA2 and psbA3 genes encoded predicted proteins with the same amino acid sequence. However, the psbA1 gene product contained five extra amino acids, which were not found in PsbA proteins from five other cyanobacteria. Moreover, the PsbA1 protein from strain SG2 had an additional 13 amino acid changes compared to the PsbA2/PsbA3 proteins and contained 10 amino acid alterations compared to conserved residues found in other cyanobacteria. Reverse transcriptase PCR analysis indicated that the psbA1 gene and the psbA2/psbA3 gene(s) were expressed in photosynthetically grown cells in the presence of atrazine. These results suggest that strong selection pressure conferred by the continual input of atrazine has contributed to the evolution of a herbicide-resistant, yet photosynthetically efficient, psbA gene in a cyanobacterium.
Triazine and phenylurea herbicides have been shown to inhibit photosystem II (PSII) in many plant species, algae, and cyanobacteria. The structure and function of PSII in the cyanobacteria are similar to those found in higher plants (25), and PSII-dependent electron transport in the cyanobacteria is similarly inhibited by these herbicides (19). Consequently, due to their ease of growth and single-cell nature, cyanobacteria have been extensively used as model organisms for studying photosynthesis at the molecular level and understanding the interaction of herbicides with their respective target sites.
The photosynthetic apparatus in cyanobacteria consists of thylakoid membrane-bound protein assemblies composed of a number of polypeptide components, including the 32-kDa D1 polypeptide (31) encoded by the psbA gene. The D1 protein is a subunit of the PSII core complex (47), and this protein, also known as the QB binding protein, is the main target for phenylurea, triazine, and phenolic-group herbicides (2, 37, 40). These herbicides compete with plastoquinone for binding and inhibit photosynthesis by blocking electron transfer from quinone acceptors QA to QB (52, 56).
The chromosomes of a number of cyanobacteria contain a family of psbA genes (9, 20), and three copies of this gene, psbA1, psbA2, and psbA3, have been identified in the genomes of Synechococcus sp. strain 7942 (21) and Synechocystis sp. strain 6803 (26). In Anacystis nidulans only one psbA gene is required for photoautotrophic growth (21). While the multiple copies of psbA genes are highly conserved in many different oxygenic phototrophic organisms (13), they are not identical (9).
A majority of cyanobacterial mutants resistant to atrazine (2-chloro-4-ethylamino-6-isopropylamino-1,3,5-s-triazine) anddiuron [3-(3,4-dichlorophenyl)-1,1-dimethylurea] (DCMU)have been shown to have one or more amino acid substitutions in the D1 protein. Herbicide resistance in higher plants, algae, and cyanobacteria can be conferred by a change at amino acid position 264 in the D1 protein (1, 10, 15, 20, 23, 37, 47). In Chlamydomonas reinhardtii and Synechococcus sp. strain PCC 7942, a change from Ser-264 to Gly-264 results in herbicide resistance (14, 20), while in higher plants, herbicide resistance has been shown to be due to an amino acid change from S264 to G264 (23). In C. reinhardtii, changes in amino acid residues 219, 251, 255, 256, and 275 also result in herbicide resistance (14, 28, 43). Although amino acid substitutions at position 264 in the D1 protein from plants result in herbicide resistance (22), these plants often display reduced photosynthetic efficiency and productivity (50). In contrast, it has been shown that amino acid substitutions at other positions of the D1 protein in several cyanobacteria also cause various levels of herbicide resistance, and in some cases there is no significant impact on photosynthetic productivity (2, 10, 17, 20, 24, 36, 37, 38, 50). As a result of this important distinction, cyanobacteria are very amenable to the study of structure-function relationships of the D1 protein and the interaction of triazine and phenylurea herbicides with PSII.
In this report, we describe the isolation and characterization of a cyanobacterial strain, SG2, from an atrazine-containing wastewater treatment system at the Syngenta atrazine production facility in St. Gabriel, La. While previous studies have used herbicide-resistant mutants to probe the participation of certain amino acids in the herbicide binding domain of the D1 protein (2, 10, 17, 20, 24, 36, 37, 38), here we investigated the D1 protein from a naturally occurring atrazine-resistant wild-type cyanobacterial strain from an atrazine-impacted environment. Results of our studies indicate that the psbA1 gene from cyanobacterial strain SG2 is strikingly different in primary amino acid sequence from those found in standard laboratory strains. Strain SG2 shows nearly normal photosynthetically driven oxygen evolution when expressing psbA1 in the presence of 2,000 μg of atrazine ml−1, suggesting that this strain has evolutionarily adapted to relatively strong selection pressure present at an atrazine production facility.
MATERIALS AND METHODS
Sampling site and isolation of cyanobacteria.
Cyanobacterial strain SG2 was isolated from a phototrophic consortium found in the atrazine biodegradation tanks at the Syngenta atrazine production facility in St. Gabriel, La. The consortium grew as a ring of photosynthetic material adhering to the sides of the 850,000 gal biological abatement system (BAS) tank at the liquid-air interfacial area. The mixed liquor within the tank contains atrazine, alkylamines, ammonia, and approximately 1% NaCl (Li-Ta Lien, personal communication). Microbial consortia were collected from eight different sites around the periphery of the tank and placed in 250-ml screw-cap flasks containing 20 ml of BG-11 medium (4) prior to transport to the lab. Field material (cyanobacterial consortia) consisted of a thin green-colored layer overlaying a thicker brown-colored mat. Field materials were cut into small pieces and transferred separately to wide-mouth screw-cap bottles containing 0.85% NaCl. After thorough agitation using a vortex mixer (30 min) and settling (10 min), aggregates were decimally diluted in 0.85% NaCl. Dilutions (0.1 ml) were plated on BG-11 agar plates and incubated at 25°C under cool-white fluorescent light (60 μmol of photons m−2 s−1) for 2 weeks. Four types of colonies (pink, white, yellow, and green) were obtained from the original field material. Single green-colored colonies were picked and repeatedly streaked (8 to 10 times) onto citric-acid-free BG-11 agar medium until pure cultures were obtained. A pure culture was obtained after about 20 weeks of repeated restreaking and was designated as cyanobacterial strain SG2. The purity of the cultures was ascertained by phase-contrast and bright-field microscopy and by plating onto Luria-Bertani medium (45), Plate Count (Difco) agar medium, and BG-11 agar medium supplemented with Casamino Acids (0.02 to 0.05%, wt/vol) and glucose (0.5%, wt/vol). The plates were incubated in the dark for 2 weeks at 25°C and examined for bacterial growth.
To test for nitrogen (N) fixation ability, a single colony of strain SG2 was used to reinoculate fresh citric-acid-free BG-11 liquid medium containing 0, 0.75, or 1.5 g of NaNO3 liter−1. Media were supplemented with atrazine (30 μg ml−1) to test for growth in the presence of this herbicide. Cultures were grown photosynthetically for 3 weeks at 25°C and examined for growth microscopically. Additional growth studies were done to determine whether strain SG2 uses N-isopropylamine or ethylamine (present in the BAS) as a sole source of nitrogen for growth. Cells were grown in citric-acid-free BG-11 liquid medium containing 7.5 mM N-isopropylamine or 7.5 mM ethylamine in sealed serum bottles. Cells were incubated in the dark at 25°C for 2 weeks, and growth was determined by visual inspection of the serum bottles.
Morphology and size determinations.
Cell morphology was determined by using bright-field illumination and a Zeiss Universal microscope equipped with a 100× Neofluar objective. Cell sizes were determined using eyepiece and stage reticules (linear 5 mm, 0.05-mm division). Values presented are the means of 10 independent size measurements of duplicate cultures of strain SG2.
16S rDNA sequencing.
Cells were prepared in 100 ml of BG-11 culture medium and grown photosynthetically until the optical density at 730 nm reached 1 to 2. Cells were harvested by centrifugation at 5,000 × g for 10 min. DNAs from the cyanobacterial isolates were isolated as previously described (44). The following two sets of primer pairs were used to selectively amplify 16S ribosomal DNA (rDNA) gene segments from cyanobacterial isolates by PCR: CYA71 (forward primer [5′-GGTGAGTAACGCGTGAGAAT-3′])-CYA771R (reverse primer [5′-GGTCGATACAATCTACGCCT-3′]) and CYA634 (forward primer [5′-GGAAGAACATCGGTGGCGAA-3′])-CYA1281R (reverse primer [5′-CGGCGATTACTAGCGATTCC-3′]). The primers were selected from conserved regions of 16S rDNA sequences by alignment to the following 10 different cyanobacterial strains (GenBank accession numbers are in parentheses following strain names): Synechococcus sp. strain PCC 7002 (AJ000716), Synechococcus sp. strain PCC 7003 (AB015059), Synechococcus sp. strain PCC 7117 (AB15060), Synechococcus sp. strain PCC 73109 (AB015061), Cyanothece sp. strain PCC 7424 (AJ000715), Spirulina subsalsa M223 (AB003166), Merismopedia glauca B14481 (X94705), Prochloron didemni (X63141), Synechocystis sp. strain PCC 6803 (D90916), and Anabaena lemmermannii NIVACYA 281/1 (Z82797). Amplification was performed using an MJ Research PTC-100 thermal cycler (Waltham, Mass.) under the following conditions: 5 min at 94°C, followed by 30 cycles of 30 s at 95°C, 1 min at 55°C, and 1 min at 72°C, with a final extension period of 2 min at 72°C for CYA71 and CYA771R. Amplifications using CYA634 and CYA1281R were performed by using the same step cycle program described above, except that the annealing temperature was 60°C. PCR products were purified using a Qiagen QIAquick PCR column purification kit (Valencia, Calif.) and were sequenced at the University of Minnesota Advanced Genetic Sequencing Center (St. Paul, Minn.). The resulting DNA sequences were aligned and compared with entries in GenBank [Synechococcus sp. strain PCC6301 (accession number X03538), Merismopedia glauca B1448-1 (X94705), Prochloron didemni (X63141), Anabaena lemmermannii 281/1 (Z82797), Anabaena cylindrica PCC 7122 (AF247592), Nostoc muscorum PCC 7120 (X59559), Spirulina sp. strain PCC 6313 (X75045), Cyanothece aeruginosa 258/2 (Z82775), Oscillatoria rosea M-220 (AB003164), Plectonema boryanum UTEX 485 (AF132793), Oscillatoria williamsii PCC 7105 (X58359), Microcystis aeruginosa TAC71 (AB012332), Gloeothece membranacea PCC 6501 (X78680), Chamaesiphon subglobus PCC 7430 (RDP Chms.sglbs), and Pleurocapsa sp. strain PCC 7516 (X78681)] using the BLAST (Genome Sequencing Center BLAST Server [http://www.ncbi.nlm.nih.gov/BLAST/]) algorithm, and the 16S rDNA sequence was aligned to entries in the Ribosomal Database using the Ribosomal Database Project (RDP) server (Michigan State University, Ribosomal Database Project II, [http://www.cme.msu.edu/RDP/html/index.html]).
Determination of atrazine and diuron resistance.
Resistance of strain SG2 to atrazine and diuron was determined, in triplicate, by measuring growth in citric-acid-free BG-11 medium with several concentrations of atrazine and diuron. The concentrations of atrazine tested were 0, 30, 100, 500, and 1,000 μg ml−1, and the concentrations of diuron were 0, 5, 15, and 30 μg ml−1. Due to the presence of suspended herbicide at concentrations that interfere with optical density measurements (i.e., greater than 30 μg ml−1), cell growth was determined by direct cell counting using a Petroff-Hausser counting chamber. At least 10 fields, each containing 8 to 80 cells, were counted for each determination.
Oxygen evolution assays.
The steady-state rate of electron transport in PSII was measured at 25°C using a Clark-type oxygen electrode (YSI 4004; Yellow Springs Instrument Co., Yellow Springs, Ohio) and saturating actinic white light, at an intensity of 2,000 μmol of photons m−2 s−1, as described previously (36). Measurements were performed with cells (approximately 10 μg of chlorophyll ml−1) in BG-11 medium containing 1 mM potassium ferric cyanide and 0.5 mM 2,6-dichlorobenzoquinone (DCBQ). Herbicides were added to obtain a final concentration of 25, 1,000, or 2,000 μg of atrazine per ml or 5 μg of diuron per ml. Values are the means of triplicate determinations.
Determination of atrazine mineralization potential and atrazine metabolites.
To investigate the ability of strain SG2 to mineralize atrazine, cyanobacterial strain SG2 cultures were grown in citric-acid-free BG-11 medium in 250-ml biometer Erlenmeyer flasks. Triplicate cultures containing 30 μg of cold atrazine per ml were augmented with 10 μg of [ring-U-14C]atrazine (specific activity, 6.5 × 10−3 μCi mol−1) per ml. Each flask was equipped with a CO2 trap containing 2 ml of 2 M NaOH. Flasks were opened at each sampling time, and all of the NaOH in the trap and 1.5 ml of culture were removed. Prior to resealing and further incubation, the NaOH in each trap was replaced with fresh solution. The amount of 14CO2 evolved was determined with a Packard 1500 liquid scintillation counter (LSC). Aliquots of growth medium were centrifuged for 10 min at 5,000 × g, and 1 ml of the supernatant was removed for scintillation counting. The remainder of each aliquot was filtered through a 0.45-μm cellulose nitrate membrane (Whatman, Maidstone, United Kingdom) to collect cells for determination of [14C]atrazine uptake. Filters were washed five times with 5 ml of 0.85% saline to remove residual [14C]atrazine adsorbed to the exterior of cells or onto filters. Filters were placed in scintillation vials containing 10 ml of Ecolume (National Diagnostics, Atlanta, Ga.), incubated overnight in the dark at 25°C, and counted using the LSC. The atrazine-degrading Pseudomonas strain ADP served as a positive control, and heat-killed SG2 cells, supplemented with [14C]atrazine, and strain SG2 without [14C]atrazine served as negative controls.
The presence of atrazine metabolites in photosynthetically grown strain SG2 was determined by analyzing [14C]atrazine-amended culture medium 20 days after inoculation. Cultures were centrifuged at 10,000 × g for 5 min to remove cells, and the supernatant was filtered through a 0.2-μm filter. High-performance liquid chromatography analyses were done as described by Boundy-Mills et al. (5). One-milliliter fractions were collected each min for 45 min, and the radioactivity in fractions was measured using the LSC after the addition of 10 ml of Ecolume. Scintillation vials were allowed to sit in the dark at 25°C for 8 to 12 h prior to counting.
DNA methods.
Restriction endonuclease digestions, gel electrophoresis, and Southern hybridizations were done according to Sambrook et al. (45). Total genomic DNA from strain SG2 was digested to completion with HincII and DraI, and restriction fragments were separated by horizontal gel electrophoresis using 0.8% agarose gels. DNA was blotted to Nytran membrane (Schleicher & Schuell, Keene, N.H.) and hybridized, under conditions of high stringency, to a 0.7-kb psbA2 gene probe from Synechocystis strain 6803. The psbA2 gene probe was obtained from plasmid pALTER-1:psbA2 (obtained from Bridgette Barry, University of Minnesota) by using PCR and the 5′-GAAGCCGCTTCCTTAGAT-3′ and 5′-RCGYTCGTGCATTACTTC-3′ primers. The PCR master mix contained 1× PCR buffer, 0.2 mM each deoxynucleoside triphosphate, 1.5 mM MgCl2, 0.5 μM concentrations of each primer, and 2.5 U of Taq DNA polymerase. Autoclaved double-distilled water was added to provide a total reaction volume of 100 μl. PCR conditions were 98°C for 5 min and 35 cycles of 95°C for 1 min, 55°C for 1 min, and 72°C for 1 min. There was a final extension period of 5 min at 72°C. The gel-purified 0.7-kb PCR product was labeled with [α-32P]dCTP using the Rediprime random primer labeling kit (Amersham Life Science, Arlington Heights, Ill.). Bands hybridizing to the psbA2 gene probe were excised, eluted from 0.8% agarose gels, and amplified by PCR using the same primer pair as above. PCR fragments were sequenced by using a PRISM Ready Reaction DyeDeoxy terminator cycle sequencing kit (Perkin-Elmer Corp., Norwalk, Conn.) and sequenced at the University of Minnesota Advanced Genetic Sequencing Center. Searches of protein and nucleic acid sequence data banks were done on the National Center for Biotechnology Information website (http://www.ncbi.nlm.nih.gov) using the BLAST network service. Amino acid alignment was performed on the Institut National de la Recherche Agronomique website (http://www.toulouse.inra.fr/multalin/multalin.html) using Multalin version 5.3.3.
The presence of atzA in cyanobacterial cultures was determined by PCR using atzA primers (5′-CCATGTGAACCAGATCCT-3′ and 5′-TGAAGCGTCCAC ATTACC-3′) (11). DNA from Pseudomonas strain ADP (30) served as the positive control for PCR studies.
Expression of psbA genes by RT-PCR.
Cells were grown in 10 ml of BG11 medium containing 1,000 μg of atrazine per ml, harvested by centrifugation at 5,000 × g for 10 min, and resuspended in 100 μl of water. Total RNA was extracted using the FastRNA Kit-Blue Protocol (BIO101 Inc., Vista, Calif.) according to the manufacturer's instructions. RNA was treated with RQ1 RNase-free DNase (Promega, Madison, Wis.) according to the manufacturer's instructions. Reverse transcription (RT) and subsequent amplification of target cDNA were performed with a GeneAmp 7 RNA PCR kit (Perkin-Elmer Roche, Branchburg, N.J.) following the manufacturer's instructions. The RT reaction mixtures contained 1 μl of DNase-treated mRNA. Reverse transcription was carried in a PTC-100 thermal cycler using the following PCR conditions: 10 min at 25°C, followed by 1 cycle of 15 min at 42°C, 5 min at 99°C, and 5 min at 51°C. Target cDNA was amplified using the same primers used for the preparation of the psbA hybridization probe described above. RT-PCR products were visualized by gel electrophoresis using 0.8% agarose gels with Tris-borate-EDTA buffer (Sigma Chemical Co., St. Louis, Mo.) and purified with a Qiagen QIAquick gel extraction kit (Chatsworth, Calif.) according to the manufacturer's directions. Purified PCR products were sequenced and analyzed as described above.
Expression of the psbA1 gene was evaluated by reverse transcription of mRNA, and the subsequent amplification of target cDNA, by using the primer pair 5′-TGCGTCCTTGGATCT-3′ and 5′-CTGTTGCATTCACC-3′. This primer pair was designed from the psbA1 gene region containing the five extra amino acids. PCR conditions were 95°C for 2 min, followed by 35 cycles of 95°C for 15 s, 58°C for 15 s, and 72°C for 30 s, with a final extension period of 7 min at 72°C. Purified PCR products were cloned using the pGEM-T Easy vector system (Promega) according to the manufacturer's instructions. Clones were sequenced by using M13 primers and analyzed as described above.
Chemicals.
Authentic atrazine (2-chloro-4-ethylamino-6-isopropylamino-1,3,5-s-triazine) was obtained from Syngenta Crop Protection, Greensboro, N.C. Diuron [3-(3,4-dichlorophenyl)-1,1-dimethylurea] (98% purity) and [ring-U-14C]atrazine (5 to 25 mCi/mmol; approximately 95% radiochemical purity) were purchased from Sigma Chemical Co.
RESULTS AND DISCUSSION
Cyanobacterial strain SG2 and analysis of atrazine resistance.
A pure culture of a cyanobacterial isolate was obtained from the atrazine biodegradation tank at the Syngenta atrazine production facility by repeated dilution plating and streaking. The streaking procedure was repeated 8 to 10 times until a single, pure, atrazine-resistant cyanobacterial culture was obtained. This pure culture was subsequently designated as cyanobacterial strain SG2. The bacterium was initially present as part of a phototrophic consortium that was adhering to the circumference of the BAS biodegradation tank at the liquid-air interfacial area. Since the effluent wastewater in the BAS tank contains a continual supply of atrazine, ammonia, and alkylamines, it was unexpected to find this cyanobacterium growing under these conditions.
Cyanobacterial strain SG2 produces cylindrical-oval cells that reproduce by binary transverse fission in one plane. Cells occurred singly, in pairs, or in short chains and were devoid of sheaths and heterocysts. Single cells of strain SG2 grown in BG-11 liquid medium were 7 to 9 μm in width and 12 to 13 μm in length, with an oval cell shape. The strain grew in citric-acid-free BG-11 medium with 0.75 or 1.5 g liter−1 of NaNO3 as an N source. However, strain SG2 failed to grow in the same medium without added N or to reduce acetylene and did not grow with atrazine (30 μg ml−1), ethylamine, or N-isopropylamine as the sole N source. Since Syngenta's atrazine BAS tank frequently contains large amounts of ammonia as a byproduct of the production of atrazine and other triazine herbicides and as the final product of atrazine biodegradation, this compound is likely providing the N source for the growth of this bacterium in this environment.
16S rDNA sequence analysis.
Cyanobacteria have been recognized as one of the largest subgroups of gram-negative prokaryotes (42); however, their taxonomy is in a state of flux and is being revised (12, 18, 39, 53, 54). The taxonomic identity of strain SG2 was analyzed by using sequence analysis of 16S rDNA (18, 53, 54). Maximum-likelihood analysis performed on 16S rDNA sequences from strain SG2 and 15 cyanobacteria in the RDP database indicated that strain SG2 falls into the Synechocystis/Pleurocapsa/Microcystis group (data not shown) as described by Turner et al. (53). The exact genus status of this cyanobacterial isolate could not be determined, because the taxonomy of this group is poorly defined. However, of the 15 organisms used for 16S rDNA sequence alignments, strain SG2 was most closely related to Microcystis aeruginosa (nucleotide identity, 83%), Prochloron didemni (nucleotide identity, 86%), Pleurocapsa sp. strain PCC7516 (nucleotide identity, 85%), and Gloeothece membranacea (nucleotide identity, 85%).
Herbicide resistance.
The resistance of cyanobacterial strain SG2 to atrazine and diuron was determined by examining growth rates and O2 evolution in the presence of several concentrations of these two herbicides. Since insoluble atrazine at a concentration greater than about 28 μg ml−1 interferes with optical density measurements, growth was observed for 15 days by using the direct counting method. Results in Fig. 1A show that strain SG2 was markedly resistant to a high concentration of atrazine (up to 1,000 μg ml−1). Strain SG2 showed no significant differences in growth rate in the presence of 30, 100, 500, or 1,000 μg of atrazine ml−1; the mean maximum-growth rate was about 0.057 h−1 regardless of the atrazine concentration. This result is in contrast to those reported by Dalla-Chiesa et al. (10) and Ohad and Hirschberg (38), who reported that while several mutations in the D1 protein of Synechocystis sp. strain PCC 6803 resulted in slight atrazine resistance, the mutant strains nonetheless failed to adequately grow under photoautotrophic conditions due to impairment of PSII.
FIG. 1.
Growth of cyanobacterial strain SG2 in the presence of several concentrations of (A) atrazine (symbols: •, 0 μg/ml; ○, 30 μg/ml; ▾, 100 μg/ml; ▿, 500 μg/ml; ▪, 1,000 μg/ml) and (B) diuron (DCMU) ( symbols: •, 0 μg/ml; ▪, 5 μg/ml; ▴, 15 μg/ml; ⧫, 30 μg/ml [off scale]). Values are the means of triplicate samples, and error bars represent standard errors of the means.
The influence of the herbicides on net photosynthesis by strain SG2 was investigated by measuring oxygen evolution. Results in Table 1 show that O2 evolution by strain SG2 in the presence of 25, 1,000, or 2,000 μg of atrazine per ml was about 90% of that found in the atrazine-free control culture. In contrast, Synechocystis sp. strain 6803 (a control strain) evolved about 11% as much O2 at 1,000 μg of atrazine per ml as the amount evolved by the atrazine-free control culture. While the solubility of atrazine in water is only 28 μg ml−1 at 20°C, increasing the concentration above the solubility limit shows increasing inhibition of oxygen evolution when tested with other cyanobacterial strains (Table 2.) Taken together, our results show that strain SG2 was resistant to high concentrations of atrazine and that the herbicide only marginally affected photosynthetic ability and had no effect on growth rate. This is in contrast to results reported by Nurusaka et al. (36) from a study using several herbicide-resistant mutants of Synechocystis sp. strain PCC 6803, which grew slower under photosynthetic growth conditions and evolved 70% less O2 than the control strain grown under herbicide-free conditions. Interestingly, Dalla-Chiesa and coworkers (10) reported that while a serine 264-to-proline 264 mutation in the D1 protein of Synechocystis sp. strain PCC 6803 allowed the strain to grow photoautotrophically and to be slightly resistant to atrazine, oxygen evolution was only 60% of that of the wild-type control strain.
TABLE 1.
Oxygen evolution by atrazine-resistant cyanobacterium strain SG2
| Strain | Herbicide
|
% O2 evolutiona | |
|---|---|---|---|
| Type | Concn (μg ml−1) | ||
| Synechocystis sp. strain 6803 | None | 100 ± 3.7 | |
| Atrazine | 25 | 13 ± 3.7 | |
| Atrazine | 1,000 | 11 ± 3.7 | |
| Diuron | 5 | 13 ± 0.0 | |
| Cyanobacterial strain SG2 | None | 100 ± 3.5 | |
| Atrazine | 25 | 88 ± 7.0 | |
| Atrazine | 1,000 | 90 ± 3.5 | |
| Atrazine | 2,000 | 90 ± 3.5 | |
| Diuron | 5 | 25 ± 6.1 | |
One hundred percent corresponds to 260 and 270 μmol of O2 evolved (mg of chlorophyll h−1) for Synechocystis sp. strain 6803 and cyanobacterial strain SG2, respectively. Values are the means of three replicates ± standard errors.
TABLE 2.
Amino acid substitutions in the psbA genes of several herbicide-resistant cyanobacteria and relatedness to psbA1 in cyanobacterial strain SG2
| Organisma | Amino acid change
|
Resistance (μM)b
|
Reference or source | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 211 | 218 | 219 | 248 | 251 | 254 | 255 | 259 | 260 | 261 | 262 | 263 | 264 | 265 | 266 | 267 | 271 | 275 | 295 | 312 | Atrazine | DCMU | ||
| Synechocystis PCC 6803 WT | F | L | V | I | A | Y | F | I | F | Q | Y | A | S | F | N | N | L | L | F | R | 0.45 | 0.08 | 38 |
| SG2 | V | G | R | T | N | E | I I L G V N | A | M | H | 9,259.2c | 21.4d | This study | ||||||||||
| Synchco. 7942 | A | 3.0 | 3.0 | 20 | |||||||||||||||||||
| Synchco. 7942 | L | A | 0.45 | 45.0 | 24 | ||||||||||||||||||
| Syncyst. 6714 | A | 210 | 6.5 | 2 | |||||||||||||||||||
| L | A | 3 | 78.0 | ||||||||||||||||||||
| S | V | 300 | 0.26 | ||||||||||||||||||||
| Syncyst. 6803 | V | 0.9 | 0.24 | 37 | |||||||||||||||||||
| M | 1.4 | 0.08 | |||||||||||||||||||||
| A | 0.9 | 0.08 | 38 | ||||||||||||||||||||
| F | 0.45 | 0.08 | |||||||||||||||||||||
| Synchco. 7942 | Y | 7.5 | 0.045 | ||||||||||||||||||||
| G | 600 | 0.30 | |||||||||||||||||||||
| A | 18.0 | 3.0 | |||||||||||||||||||||
| Y | A | 60.0 | 4.8 | ||||||||||||||||||||
| Y | G | 69.0 | 0.09 | ||||||||||||||||||||
| L | A | 0.39 | 45.0 | ||||||||||||||||||||
| Syncyst. 6803 | P | 1,600 | 6.0 | 10 | |||||||||||||||||||
| P | 8,000 | 0.10 | |||||||||||||||||||||
| Syncyst. 6803 | Y | >1,000 | 1.5 | 36 | |||||||||||||||||||
| C | >1,000 | 3.0 | |||||||||||||||||||||
| S | 6.0 | 1.0 | |||||||||||||||||||||
| T | >1,000 | 42.8 | |||||||||||||||||||||
| F | 44.6 | 641.7 | |||||||||||||||||||||
| L | C | >1,000 | >1,000 | ||||||||||||||||||||
| F | 1.7 | 4.5 | |||||||||||||||||||||
Abbreviations: Synchco., Synechococcus mutant; Syncyst., Synechocystis mutant; WT, wild type.
Values represent the concentrations of herbicide that inhibit the oxygen evolution rate by 50%, except as noted.
Value represents the concentration of herbicide that inhibit the oxygen evolution rate by 10%.
Value represents the concentration of herbicide that inhibit the oxygen evolution rate by 80%.
In contrast, strain SG2 had relatively low resistance to diuron (<15 μg ml−1) (Fig. 1B); strain SG2 was completely inhibited at concentrations of diuron exceeding 30 μg ml−1, and cell death occurred within 1 day. The maximum growth rates of strain SG2 in 0, 5, and 15 μg of diuron ml−1 were 0.053, 0.023, and 0.013 h−1, respectively. However, while strain SG2 grew in the presence of a low concentration of diuron (5 to 15 μg ml−1), oxygen evolution was inhibited 75% by 5 μg of this herbicide ml−1 (Table 1). Synechocystis sp. strain 6803 similarly evolved only 13% as much O2 in the presence of 5 μg of diuron per ml as the amount evolved by the control medium. Results from the analysis of D1 mutants of several cyanobacteria and Chlamydomonas strains indicated that there is not a direct correspondence between atrazine resistance and resistance to diuron. Moreover, this relationship differs considerably among mutations in the psbA genes from different species of cyanobacteria (1, 10, 17, 38).
Sequence of psbA gene family.
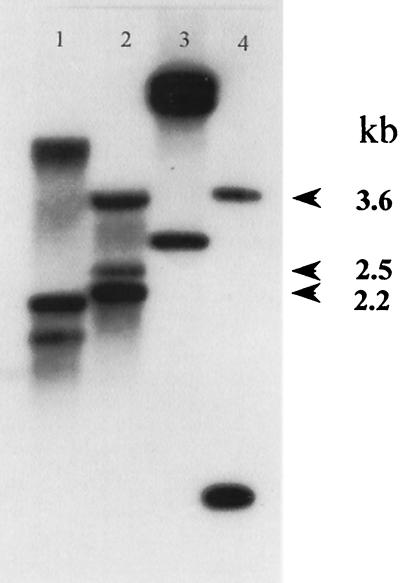
Southern hybridization analyses done using a psbA gene probe and HincII-digested total genomic DNA from strain SG2 indicated that this cyanobacterial isolate contained three psbA genes (psbA1, psbA2, and psbA3), located on 3.6-, 2.5-, and 2.2-kb HincII fragments, respectively (Fig. 2). To confirm the presence of the three psbA genes on these DNA fragments, approximately 0.7 kb of nucleotide sequence was determined on both strands of each HincII fragment. The results of this analysis confirmed that strain SG2 contains three copies of the psbA gene. The sequences of the psbA1 gene and the psbA2/psbA3 gene(s) from cyanobacterial strain SG2 have been deposited in GenBank under accession numbers AF402783 and AF402784, respectively. It should be noted that the chromosomes of a number of other cyanobacteria also have been shown to contain a family of psbA genes (9, 20, 21, 26).
FIG. 2.
Southern hybridization of total genomic DNA from cyanobacterial strain SG2 and Synechocystis sp. strain 6803 digested with DraI and HincII to a 0.7-kb psbA2 gene PCR product from Synechocystis sp. strain 6803. Lane 1, strain SG2 DNA digested with DraI; lane 2, strain SG2 DNA digested with HincII; lane 3, strain 6803 DNA digested with DraI; lane 4, strain 6803 DNA digested with HincII. Values in margin are in kilobases.
The partial deduced amino acid sequences for the psbA1, psbA2, and psbA3 genes from cyanobacterial strain SG2 are shown in Fig. 3A. Alignment of deduced amino acid sequences of the psbA gene family from strain SG2 revealed that the PsbA1 amino acid sequence differs from those of PsbA2 and PsbA3. Specifically, PsbA1 contains a five-amino-acid insertion (Ile, Leu, Gly, Val, and Asn), beginning at amino acid position 266, and has 13 amino acid differences relative to PsbA2 and PsbA3 (Fig. 3A). The psbA2 and psbA3 genes had identical nucleotide and deduced amino acid sequences. Moreover, relative to conserved regions of psbA2 from several standard cyanobacterial strains [Synechocystis sp. strain PCC 6803 (accession number X13547), Microcystis aeruginosa (D84228), Synechococcus sp. strain PCC6301 (X04617), Anabaena sp. strain PCC 7210 (U21331), and Cyanothece sp. strain ATCC 51142 (U39610)], PsbA1 from strain SG2 has 10 amino acid alterations and 5 extra amino acids (Fig. 3B). These results show that the cyanobacterial strain SG2 contains a unique psbA1 gene that differs significantly from the psbA2 and psbA3 genes in strain SG2 and those present in other cyanobacteria. The psbA genes from cyanobacterium strain SG2 showed 82 to 95% identity to psbA genes from Synechocystis sp. strain PCC 6803, Microcystis aeruginosa K-81, Synechococcus sp. strain PCC 6301, Anabaena sp. strain PCC 7210, and Cyanothece sp. strain ATCC 51142. These results and those of several other studies (1, 6, 21, 23, 25, 35, 55) indicate that multiple copies of psbA genes appear to be the norm in cyanobacteria and that these genes are differentially expressed in different strains.
FIG. 3.
Amino acid sequence alignments of the psbA gene family from cyanobacterial strain SG2 (A) and the psbA1 gene from cyanobacterial strain SG2 to other cyanobacteria (B). Amino acids not showing variation and preceding amino acid position 214 are not shown. Boxes represent amino acids showing variation. Abbreviations: SG2, cyanobacterial strain SG2; SyntPCC6803, Synechocystis sp. strain PCC 6803 (accession number X13547); Microcystis, Microcystis aeruginosa K-81 (D84228); SyncPCC6301, Synechococcus sp. strain PCC6301 (X04617); Anabaena, Anabaena sp. strain PCC 7210 (U21331); Cyanothece, Cyanothece sp. strain ATCC 51442 (U39610).
The relationship between amino acid substitutions and herbicide resistance has been most studied in a region of PsbA between amino acid positions 211 and 275 (2, 10, 17, 20, 24, 36, 37, 38). In several studies it has been shown that modification of one or two amino acids at different positions of PsbA2 results in resistance to atrazine and diuron (2, 3, 10, 17, 24, 36, 37, 38, 46, 48, 50). However, only mutations in psbA1 of Synechocystis sp. strain 6714 result in resistance to diuron, atrazine, and metribuzin (6). One of the most common mutations shown to alter herbicide resistance in cyanobacteria is a mutation of serine 264 to alanine (Table 2). Similarly, in algae and higher plants, mutation of serine 264 to glycine, threonine, or glutamic acid results in herbicide resistance (1, 19, 37, 38, 45).
In view of the highly conserved amino acid sequence of PsbA among different species of cyanobacteria (23), the amino acid composition of PsbA1 from strain SG2 is remarkable in that it contains a relatively large number of amino acid substitutions and additions. The alteration of D1 protein in strain SG2 most likely changes the binding affinity of the atrazine molecule in the herbicide binding niche, which has been defined as consisting of the amino acids between Phe211 and Leu275 (36). While crystal structure data are available for the L subunit of the photosynthetic reaction centers from Rhodopseudomonas viridis (29, 34) and Rhodobacter sphaeroides (7), none are available for the D1 protein of cyanobacteria. However, the L subunit and the D1 proteins from higher plants, algae, and cyanobacteria show striking sequence similarities that allow for structure-function comparisons between these proteins (29, 33). In the D1 protein, the Ser264 corresponds to SerL223 from Rhodopseudomonas viridis (33, 49). SerL223, and by analogy Ser264, has been shown to facilitate binding of atrazine andterbutryn (2-t-butylamino-4-ethylamino-6-methylthio-s-tri-azine) by hydrogen bonding to the N of the ethylamino side chain of these herbicides (29, 33, 49). This has been confirmed by X-ray crystallographic data (29) and modeling studies (51). Mutations in Ser264 and SerL223 result in resistance to triazine herbicides due to loss of a hydrogen-bonding acceptor (29). In cyanobacterial strain SG2, serine 264 is present as glutamate 264 (Table 2), and a change in this residue most likely contributes to atrazine resistance. In addition, the alterations in 13 other amino acids and the insertion of 5 additional amino acids in the PsbA1 of strain SG2 most likely alters the structure and binding affinity of the D1 protein for atrazine, thereby allowing this bacterium to grow in the presence of large concentrations of herbicide.
psbA gene expression.
To determine whether all three psbA genes were expressed in strain SG2, we isolated total RNA from cells grown in the presence of atrazine and performed RT-PCR. The resulting cDNA was amplified by using the psbA primer pair, and the 0.7-kb RT-PCR product was sequenced. Sequencing results showed that the RT-PCR product was identical to those of the the psbA2 and psbA3 genes in strain SG2 (data not shown). This sequence is similar to those of psbA genes found in other cyanobacteria (21, 32, 41).
To determine whether transcripts from the psbA1 gene were also present, total mRNA was amplified by PCR using a primer pair specific for the psbA1 gene. A 0.4-kb band was observed from the RT-PCR (data not shown). To further verify that the 0.4-kb RT-PCR fragment contained psbA1, we cloned the RT-PCR product into the pGEM-T Easy vector and sequenced the entire fragment. The sequence generated from the cDNA clone was 100% identical to the sequence of psbA1 derived from the genomic DNA copy of the gene in strain SG2. These results indicated that the psbA1 gene is expressed when strain SG2 is grown in the presence of atrazine.
Taken together, these results indicate that all three copies of psbA are expressed in cyanobacterial strain SG2. This result is in agreement with those of Golden et al. (21), who reported that all three psbA genes are expressed in Anacystis nidulans strain R2.
Lack of amplification of the atzA gene from strain SG2.
Atrazine, its metabolites, and several related s-triazine compounds are catabolized by several soil bacteria (8, 16, 27) and by Pseudomonas sp. strain ADP to carbon dioxide and ammonia (30). In Pseudomonas sp. strain ADP and several other gram-negative bacteria, the atzA gene encodes atrazine chlorohydrolase, which transforms atrazine to hydroxyatrazine. Homologs of atrazine chlorohydrolase from four different atrazine-degrading bacteria—Alcaligenes sp. strain SG1, Ralstonia sp. strain M91-3, Agrobacterium sp. strain J14a, and isolate 38/38—were found to have greater than 99% sequence identity to the atzA gene from Pseudomonas sp. strain ADP (11). To determine if cyanobacterial strain SG2 had atrazine degradation genes similar to those found in other bacteria, total chromosomal DNA from strain SG2 was used as a template for PCRs using atzA primers. No PCR product was generated using this primer pair, and total genomic DNA from strain SG2 failed to hybridize to an atzA gene probe under stringent and nonstringent hybridization conditions (data not shown), indicating that this bacterium does not contain a homolog to that catabolic gene.
[14C]atrazine metabolism.
[ring-U-14C]atrazine was used to determine whether strain SG2 metabolized atrazine. [14C]atrazine degradation was determined by measuring the radioactivity in the growth medium and the NaOH trap from biometer flasks. The results of this study showed that no 14CO2 was present in the NaOH trap when strain SG2 was used as the inoculum. In contrast, after 72 h of growth, about 90% of the label was recovered as 14CO2 for the positive control strain Pseudomonas sp. strain ADP (data not shown). Moreover, cultures of strain SG2 grown in the presence of atrazine did not show any atrazine metabolites as determined by high-performance liquid chromatography analysis, and no radioactivity was associated with cells of strain SG2, indicating that the cells did not accumulate [14C]atrazine. These results indicate that strain SG2 does not have the catabolic machinery needed for atrazine degradation and that atrazine resistance is not conferred by metabolism of this herbicide.
In summary, we report here the isolation and characterization of a naturally occurring cyanobacterial strain, SG2, which is highly resistant to large concentrations of atrazine. In contrast to what has been previously reported for atrazine-resistant mutants of several cyanobacteria, strain SG2 displays normal growth and oxygen evolution rates in the presence of >1,000 μg of atrazine per ml. Consequently, it appears that strain SG2 has evolved, under the strong selection pressure imparted by high concentrations of atrazine, an efficient photosystem that can overcome the damaging influence of a potent photosystem II inhibitor. Atrazine is an inhibitor of photosynthetic electron transport and functions by binding to a specific site of the D1 protein (the herbicide binding niche), thereby inhibiting electron flow from QA to QB (52). While previous studies using random and site-directed mutagenesis approaches have identified several amino acid substitutions of the D1 protein (mapping between amino acid positions 211 and 275) that impart some degree of atrazine resistance and presumably alter atrazine binding, these approaches have identified only some of the possible substitutions. Diversity in the D1 proteins of atrazine-resistant cyanobacteria can best be explored by examining variations present in natural populations of herbicide-resistant cyanobacteria. This is clearly shown here by our analysis of the D1 proteins encoded by the three psbA genes in strain SG2. The psbA1 gene in this bacterium contains a heretofore unreported insertion of five amino acids in the psbA1 gene. Moreover, since the psbA2 and psbA3 genes of strain SG2 are similar to each other and to those of other cyanobacteria, our results suggest that the novel psbA1 gene most likely allows for growth in the presence of elevated atrazine. This conclusion is further supported by the fact that the psbA1 gene is expressed in this bacterium and atrazine resistance is not due to herbicide degradation. Taken together, these results contribute to our understanding of atrazine resistance among cyanobacteria and the structure-function of the photosystem II D1 protein.
Acknowledgments
This work was supported in part by a grant from Syngenta Crop Protection, Greensboro, N.C.; by a grant from the University of Minnesota Agricultural Experiment Station (to M.J.S.); by a scholarship from the Anandhamahidol Foundation of Thailand (to K.S.); and by a postdoctoral fellowship from BARD (to N.S.).
We thank Brian Kinkle for help with phylogenetic analyses, Li-Ta Lien for help with collecting samples, Mervyn de Souza for helpful discussions, and Bridgette Barry, Lorraine Anderson, and Idelisa Ayala for the psbA2 gene from Synechocystis sp. strain 6803 and for assistance with the O2 evolution experiments.
REFERENCES
- 1.Ajlani, G., D. Kirilovsky, M. Picaud, and C. Astier. 1989. Molecular analysis of psbA mutations responsible for various herbicide resistance phenotypes in Synechocystis 6714. Plant Mol. Biol. 13:469-479. [DOI] [PubMed] [Google Scholar]
- 2.Ajlani, G., I. Meyer, C. Vernotte, and C. Astier. 1989. Mutation in phenol-type herbicide resistance maps within the psbA gene in Synechocystis 6714. FEBS Lett. 246:207-210. [DOI] [PubMed] [Google Scholar]
- 3.Alfonso, M., J. J. Pueyo, K. Gaddour, A. L. Etienne, D. Kirilovsky, and R. Picorel. 1996. Induced new mutation of D1 serine-268 in soybean photosynthetic cell cultures produced atrazine resistance, increased stability of S2QB- and S3QB- states, and increased sensitivity to light stress. Plant Physiol. 112:1499-1508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Allen, M. M. 1968. Simple conditions for growth of unicellular blue-green algae on plates. J. Phycol. 4:1-4. [DOI] [PubMed] [Google Scholar]
- 5.Boundy-Mills, K. L., M. L. de Souza, R. T. Mandelbaum, L. P. Wackett, and M. J. Sadowsky. 1997. The atzB gene of Pseudomonas sp. strain ADP encodes the second enzyme of a novel atrazine degradation pathway. Appl. Environ. Microbiol. 63:916-923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bouyoub, A., C. Vernotte, and C. Astier. 1993. Functional analysis of the two homologous psbA gene copies in Synechocystis PCC 6714 and PCC 6803. Plant Mol. Biol. 21:249-258. [DOI] [PubMed] [Google Scholar]
- 7.Chang, C.-H., O. El-Kabbani, D. Tiede, J. Norris, and M. Schiffer. 1991. Structure of the membrane-bound protein photosynthetic reaction center from Rhodobacter sphaeroides. Biochemistry 30:5353-5360. [DOI] [PubMed] [Google Scholar]
- 8.Cook, A. M. 1987. Biodegradation of s-triazine xenobiotics. FEMS Microbiol. Rev. 46:93-116. [Google Scholar]
- 9.Curtis, S. E., and R. Haselkorn. 1984. Isolation, sequence and expression of two members of the 32 kd thylakoid membrane protein gene family from the cyanobacterium Anabaena 7120. Plant Mol. Biol. 3:249-258. [DOI] [PubMed] [Google Scholar]
- 10.Dalla-Chiesa, M., G. Friso, Z. Deák, I. Vass, J. Barber, and P. J. Nixon. 1997. Reduced turnover of the D1 polypeptide and photoactivation of electron transfer in novel herbicide resistant mutants of Synechocystis sp. PCC 6803. Eur. J. Biochem. 248:731-740. [DOI] [PubMed] [Google Scholar]
- 11.de Souza, M. L., J. Seffernick, B. Martinez, M. J. Sadowsky, and L. P. Wackett. 1998. The atrazine catabolism genes atzABC are widespread and highly conserved. J. Bacteriol. 180:1951-1954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Douglas, S. E., and N. Carr. 1988. Examination of genetic relatedness of marine Synechococcus spp. by using restriction fragment length polymorphism. Appl. Environ. Microbiol. 54:3071-3078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Erickson, J. M., P. Delepelair, and J.-D. Rochaix. 1985. Analysis of genes encoding two photosystem II proteins of the 30-34 kD size class, p. 53-65. In K. Steinback, S. Bonitz, C. J. Arntzen, and L. Bogorad (ed.), Molecular biology of the photosynthetic apparatus. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
- 14.Erickson, J. M., M. Rahire, J. D. Rochaix, and L. Mets. 1985. Herbicide resistance and cross resistance: changes at three distinct sites in the herbicide-binding protein. Science 228:204-207. [DOI] [PubMed] [Google Scholar]
- 15.Erickson, J. M., M. Rahire, P. Bennoun, P. Delepelaire, B. Diner, and J. D. Rochaix. 1984. Herbicide resistance in Chlamydomonas reinhardtii results from a mutation in chloroplast gene for the 32-kilodalton protein of photosystem II. Proc. Natl. Acad. Sci. USA 81:3617-3621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Erickson, L. E., and K. H. Lee. 1989. Degradation of atrazine and related s-triazines. Crit. Rev. Environ. Control 19:1-4. [Google Scholar]
- 17.Gingrich, J. C., J. S. Buzby, V. L. Stirewalt, and D. A. Bryant. 1988. Genetic analysis of two new mutations resulting in herbicide resistance in the cyanobacterium Synechococcus sp. PCC 7002. Photosynth. Res. 16:83-99. [DOI] [PubMed] [Google Scholar]
- 18.Giovannoni, S. J., T. B. Britschgi, C. L. Moyer, and K. G. Field. 1990. Genetic diversity in Sargasso Sea bacterioplankton. Nature 345:60-63. [DOI] [PubMed] [Google Scholar]
- 19.Golden, S. S., and L. A. Sherman. 1984. Biochemical and biophysical characterization of herbicide-resistant mutants of the unicellular cyanobacterium, Anacystis nidulans R2. Biochim. Biophys. Acta 764:239-246. [Google Scholar]
- 20.Golden, S. S., and R. Haselkorn. 1985. Mutation to herbicide resistance maps within the psbA gene of Anacystis nidulans R2. Science 229:1104-1107. [DOI] [PubMed] [Google Scholar]
- 21.Golden, S. S., J. Brusslan, and R. Haselkorn. 1986. Expression of a family of psbA genes encoding a photosystem II polypeptide in the cyanobacterium Anacystis nidulans R2. EMBO J. 5:2789-2798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hirschberg, J., A. Ben-Yehuda, I. Pecker, and N. Ohad. 1987. Mutations resistant to photosystem II herbicides, p. 357-366. In D. Von Wettstein and N. H. Chua (ed.), Plant molecular biology. Plenum, New York, N.Y.
- 23.Hirschberg, J., and L. McIntosh. 1983. Molecular basis of herbicide resistance in Amaranthus hybridus. Science 222:1346-1348. [DOI] [PubMed] [Google Scholar]
- 24.Hirschberg, J., N. Ohad, I. Pecker, and A. Rahat. 1987. Isolation and characterization of herbicide resistant mutants in the cyanobacterium Synechococcus R2. Z. Naturforsch. 42C:758-761. [Google Scholar]
- 25.Ho, K. K., and D. W. Krogmann. 1982. Photosynthesis, p. 191-214. In N. G. Carr and B. A. Whitton (ed.), The biology of cyanobacteria. University of California Press, Berkeley, Calif.
- 26.Jansson, C., R. J. Debus, H. D. Osiewaez, M. Gurevitz, and L. McIntosh. 1987. Construction of an obligate photoheterotrophic mutant of the cyanobacterium Synechocystis 6803. Plant Physiol. 85:1021-1025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Jessee, J. A., R. E. Benoit, A. C. Hendricks, G. C. Allen, and J. L. Neal. 1983. Anaerobic degradation of cyanuric acid, cysteine, and atrazine by a facultatively anaerobic bacterium. Appl. Environ. Microbiol. 45:97-102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Johanningmeier, U. Bodner, and G. F. Wildner. 1987. A new mutation in the gene coding for the herbicide-binding protein in Chlamydomonas. FEBS Lett. 211:221-224. [Google Scholar]
- 29.Lancaster, C. R. D., and H. Michel. 1999. Refined crystal structure of reaction centres from Rhodopseudomonas viridis in complexes with the herbicide atrazine and the two chiral atrazine derivatives also lead to a new model of the bound carotenoid. J. Mol. Biol. 286:883-898. [DOI] [PubMed] [Google Scholar]
- 30.Mandelbaum, R. T., D. L. Allan, and L. P. Wackett. 1995. Isolation and characterization of Pseudomonas sp. that mineralizes the s-triazine herbicide atrazine. Appl. Environ. Microbiol. 61:1451-1457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Mathis, P. 1987. Primary reactions of photosynthesis: discussion of current issues, p. 151-160. In J. Biggins (ed.), Progress in photosynthesis research, vol. 1. Martinus Nijhoff Publishers, Dordrecht, The Netherlands. [Google Scholar]
- 32.Metz, J., P. Nixon., and B. Diner. 1990. Nucleotide sequence of the psbA3 gene from the cyanobacterium Synechocystis PCC 6803. Nucleic Acids Res. 18:6715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Michel, H., and J. Deisenhofer. 1988. Relevance of the phtosynthetic reaction center from purple bacteria to the structure of photosystem II. Biochemistry 27:1-7. [Google Scholar]
- 34.Michel, H., O. Epp, and J. Deisenhofer. 1986. Pigment-protein interactions in the photosynthetic reaction centre from Rhodopseudomonas viridis. EMBO J. 5:2445-2451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Mohamed, A., and C. Jansson. 1989. Influence of light on accumulation of photosynthesis-specific transcripts in the cyanobacterium Synechocystis 6803. Plant Mol. Biol. 13:693-700. [DOI] [PubMed] [Google Scholar]
- 36.Narusaka, Y., M. Narusaka, H. Kobayashi, and K. Satoh. 1998. The herbicide-resistant species of the cyanobacterial D1 protein obtained by thorough and random in vitro mutagenesis. Plant Cell Physiol. 39:620-626. [DOI] [PubMed] [Google Scholar]
- 37.Ohad, N., and J. Hirschberg. 1990. A similar structure of the herbicide-binding site in photosystem II of plants and cyanobacteria is demonstrated by site specific mutagenesis of the psbA gene. Photosynth. Res. 23:73-79. [DOI] [PubMed] [Google Scholar]
- 38.Ohad, N., and J. Hirschberg. 1992. Mutations in the D1 subunit of photosystem II distinguish between quinone and herbicide binding sites. Plant Cell 4:273-282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Palenik, B., and R. Haselkorn. 1992. Multiple evolutionary origins of prochlorophytes, the chlorophyll b-containing prokaryotes. Nature 355:265-267. [DOI] [PubMed] [Google Scholar]
- 40.Pfister, K., and C. J. Arntzen. 1979. The mode of action of photosystem II specific inhibitors in herbicide-resistant weed biotypes. Z. Naturforsch. 34:996-1009. [Google Scholar]
- 41.Ravnikar, P. D., R. Debus, J. Sevrinck, P. Saetaert, and L. McIntosh. 1989. Nucleotide sequence of a second psbA gene from the unicellular cyanobacterium Synechocystis 6803. Nucleic Acids Res. 17:3991-3997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Rippka, R., J. Deruelles, J. B. Waterbury, M. Herdman, and R. Y. Stanier. 1979. Generic assignments, strain histories and properties of pure cultures of cyanobacteria. J. Gen. Microbiol. 111:1-61. [Google Scholar]
- 43.Rochiax, J. D., and J. M. Erickson. 1988. Function and assembly of photosystem II: genetic and molecular analysis. Trends Biochem. Sci. 13:56-59. [DOI] [PubMed] [Google Scholar]
- 44.Sadowsky, M. J., R. E. Tully, P. B. Cregan, and H. H. Keyser. 1987. Genetic diversity in Bradyrhizobium japonicum serogroup 123 and its relation to genotype-specific nodulation of soybean. Appl. Environ. Microbiol. 53:2624-2630. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Press, Cold Spring Harbor, N.Y.
- 46.Sato, F., Y. Shigematsu, and Y. Yamada. 1988. Selection of an atrazine-resistant tobacco cell line having a mutant psbA gene. Mol. Gen. Genet. 214:358-360. [DOI] [PubMed] [Google Scholar]
- 47.Satoh, K., H. Y. Nakatani, K. E. Steinback, J. Watson, and C. J. Arntzen. 1983. Polypeptide composition of a photosystem II core complex presence of a herbicide-binding protein. Biochim. Biophys. Acta 724:142-150. [Google Scholar]
- 48.Schonfed, M., T. Yaacoby, A. Ben-Yehuda, and B. Rubin. 1986. Triazine resistance in Phalaris paradoxa: physiological and molecular analyses. Z. Naturforsch. Sect. C 42:779-782. [Google Scholar]
- 49.Sinning, I. 1992. Herbicide binding in the bacterial photosynthetic reaction center. Trends Biochem. Sci. 17:150-154. [DOI] [PubMed] [Google Scholar]
- 50.Smeda, R. J., P. M. Hasegawa, P. B. Goldsbrough, N. K. Singh, and S. C. Weller. 1993. A serine-to-threonine substitution in the triazine herbicide-binding protein in potato cells results in atrazine resistance without impairing productivity. Plant Physiol. 103:911-917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Sobolev, V., and M. Edelman. 1995. Modeling the quinone-B binding site of the photosystem-II reaction center using notions of complementarity and contact-surface between atoms. Proteins 21:214-225. [DOI] [PubMed] [Google Scholar]
- 52.Tischer, W., and H. Strotmann. 1977. Relationship between inhibitor binding by chloroplasts and inhibition of photosynthetic electron transport. Biochim. Biophys. Acta 460:113-125. [DOI] [PubMed] [Google Scholar]
- 53.Turner, S., K. M. Pryer, V. P. W. Miao, and J. D. Palmer. 1999. Investigating deep phylogenetic relationships among cyanobacteria and plastids by small subunit rRNA sequence analysis. J. Eukaryot. Microbiol. 46:327-338. [DOI] [PubMed] [Google Scholar]
- 54.Urbach, E., D. L. Robertson, and S. W. Chisholm. 1992. Multiple evolutionary origins of prochlorophytes within the cyanobacterial radiation. Nature 355:267-270. [DOI] [PubMed] [Google Scholar]
- 55.Vrba, J. M., and S. E. Curtis. 1990. Characterization of a four-member psbA gene family from the cyanobacterium Anabaena PCC 7120. Plant Mol. Biol. 14:81-92. [DOI] [PubMed] [Google Scholar]
- 56.Wraight, C. A. 1981. Oxidation-reduction physical chemistry of the acceptor quinone complex in bacterial photosynthetic reaction centers: evidence for a new model of herbicide activity. Israel J. Chem. 21:348-354. [Google Scholar]