Abstract
High-resolution fMRI provides a window into the brain’s mesoscale organization. Yet, higher spatial resolution increases scan times, to compensate for the low signal and contrast-to-noise ratio. This work introduces a deep learning-based 3D super-resolution (SR) method for fMRI. By incorporating a resolution-agnostic image augmentation framework, our method adapts to varying voxel sizes without retraining. We apply this innovative technique to localize fine-scale motion-selective sites in the early visual areas. Detection of these sites typically requires ≤ 1mm isotropic data, whereas here, we visualize them based on lower resolution (2-3mm isotropic) fMRI data. Remarkably, the super-resolved fMRI is able to recover high-frequency detail of the interdigitated organization of these sites (relative to the color-selective sites), even with training data sourced from different subjects and experimental paradigms – including non-visual resting-state fMRI, underscoring its robustness and versatility. Quantitative and qualitative results indicate that our method has the potential to enhance the spatial resolution of fMRI, leading to a drastic reduction in acquisition time.
1. INTRODUCTION
High-resolution fMRI collected with ultra-high field scanners (e.g., 7T), is attracting increasing interest because of its capability to non-invasively visualize the mesoscale functional organization of the human brain. In the study of the visual cortex [9], this method enables the mapping of ocular dominance columns in the striate and motion/color/stereo-selective columns in the extrastriate visual cortex [13, 24, 3]. However, the improvement in spatial resolution demands increased signal averaging to counteract the decline in signal-to-noise ratio (SNR) intrinsic to smaller voxels [18]. Consequently, this necessitates extended scanning time, which can pose limitations in clinical and research settings due to constraints on scan time and subject comfort [19].
Super-resolution (SR) methods [4] that can enhance the local structure of lower resolution data acquired more quickly with reduced SNR, may offer a compelling avenue to circumvent the limitations in fMRI. Especially, deep learning-based methods [11] can interpolate fine details within the image [23], potentially reducing the necessity for prolonged scanning sessions while still achieving high-resolution fMRI data. Thus, they hold significant promise for elucidating the neural substrate with greater precision without proportionately extending the scanning time frame.
Deep learning for SR of structural brain MRI has been widely explored in the last few years [25, 8, 16]. One common strategy is to learn a non-linear interpolation mapping between the low-resolution and high-resolution images with deep convolutional neural networks. Typically, low-resolution images are obtained by downsampling the high-resolution ones and adding realistic artifacts (e.g., noise [20]). Combined with large datasets, this strategy enables the learning of powerful image priors, parameterized by deep neural networks, by minimizing the difference between the generated SR image and the high-resolution reference image.
In the context of high-resolution fMRI, Kornprobst et al. [10] made one of the earliest attempts to formulate fMRI SR as a convex optimization problem based on 2D local patches. Recent works [23, 14] employ 2D slices for training convolutional neural networks and validate their methods using fixed downsampling factors (which limits applicability in practice). In the 7T setting, we argue that 3D spatial information is crucial to provide stronger supervision signals than 2D slices. Furthermore, there is a lack of rigorous validation of how SR performs in cross-task and cross-subject settings, and how super-resolved images could assist fMRI analysis when considering the temporal information.
Here we propose the first method for 3D SR of 7T fMRI with deep learning to robustly improve its spatial resolution. Specifically, our approach is adaptive to varied resolutions and cross paradigms of tasks without retraining, benefiting from a domain-randomization-based data simulation strategy.
2. METHODS
Our SR approach operates on single 3D frames to be compatible with fMRI data with varied temporal resolutions. It includes two main components: 1) domain-randomizationbased data simulation, and 2) Optimization of deep neural networks using a compound loss function as shown in Fig. 1.
Fig. 1:

Training overview. We parse the 4D (3D+t) fMRI data to single-time-point 3D frames. We use a function to perform random affine and non-linear transformations, and random contrast adjustment. We then downgrade the augmented HR frames by another function to perform downsampling to random lower resolutions (including a low-pass antialias filter), adding noise, and linear interpolation to the original size.
Problem definition:
The objective is to learn a mapping from a 3D low-resolution (LR) domain (with unknown voxel size) to a 3D high-resolution (HR) domain (with fixed voxel size, in practice 1x1x1mm). Let denote an LR image space and an HR image space. In practice, will have a voxel size that is common in low-resolution fMRI studies (e.g., any between 2mm and 3mm isotropic resolution).
Data generation:
Since obtaining both LR and HR images from the same subjects is impractical, we introduce LR domain characteristics in a generative fashion, using a model-based approach that simulates LR from HR by downsampling to a random resolution (voxel sizes uniformly sampled between 1 and 3.5mm in each direction), random rotation, brightness adjustment, and adding Gaussian noise [22]. In summary, we aim to learn a direct resolution-agnostic interpolation function from the LR to the HR domain .
Loss function:
To minimize the difference between the super-resolved image and the reference HR image, we propose a compound loss considering voxel-wise reconstruction accuracy and local structural similarity:
| (1) |
where . measures the structural similarity of two images [21]. is a weighting parameter of the loss terms and is empirically set to 0.2 [2] in all experiments.
Network architecture:
We employ a 3D fully convolutional neural network architecture inspired by [20] (SR Network in Fig. 1). The network comprises ten dense layers connected in a residual fashion and allows for an improved flow of information and gradients with a global residual connection – crucial for learning fine details in the SR task.
Model inference:
After learning the interpolation function from HR fMRI data, we apply the trained model to 4D fMRI data on a frame-by-frame basis. We then concatenate the SR frames in the temporal dimension to yield the super-resolved 4D fMRI.
Structural and functional MRI analysis:
For each subject, inflated and flattened (i.e., structural) cortical surfaces are reconstructed based on their high-resolution anatomical data [5]. Details of fMRI pre-processing steps, including motion correction, functional-to-structural data registration [7], columnar smoothing [1], and GLM modeling [15] are described in Kennedy et al. (2023) [9].
Analysis of regions of interest:
For each participant, regions of interest (ROIs) encompassing the visual areas V2, V3, and V3A were delineated based on the subject’s own retinotopic mapping, as detailed previously [9]. To enhance the sensitivity of all analyses, data from both the left and right hemispheres were combined. No hemispheric data were omitted, and all vertices within each defined ROI were incorporated into the analyses.
3. EXPERIMENTS AND RESULTS
3.1. Image Acquisition
Participants:
The study complied with NIH guidelines and had Massachusetts General Hospital IRB approval. Nine participants aged 23-44 years, all with normal or corrected-to-normal vision, standard color vision, and normal stereo vision, enrolled. They underwent multiple scans in a 7T ultra-high field scanner (Siemens Healthcare, Germany) for functional tests and were provided with written informed consent.
Imaging parameters:
Scans were obtained using a 7T whole-body scanner and a 32-channel head coil. Voxel dimensions were nominally 1.0 mm, isotropic. Functional images were acquired using single-shot gradient-echo EPI with the following parameters: TR=3 s, TE=28 ms, flip angle=78°, matrix=192×192, BW=1184 Hz/pix, echo-spacing=1 ms, 7/8 phase partial Fourier, FOV=192×192 mm, 44 oblique-coronal slices, acceleration factor R=4 with GRAPPA reconstruction.
Training set:
In four subjects, we measured fMRI activity in separate scan sessions under (a) resting-state conditions with eyes closed, and (b) as subjects view random dot stereograms perceived as motion in depth vs. in the frontoparallel plane. Details on stimuli variations were in our previous work [9].
Test set:
In five subjects, we collected fMRI data as they were presented with (a) moving vs. stationary concentric rings and (b) color-varying vs. luminance-varying stimuli. These contrasts helped localize motion- and color-selective sites [6].
3.2. Experimental setup
Two SR networks were trained on the high-resolution fMRI data acquired from stereo stimuli and resting state tasks and were applied to motion stimuli from different subjects. This allowed us to validate our approach in a cross-task, cross-subject fashion rigorously. Low-resolution images were generated by downsampling the 1mm high-resolution images to 2 and 3mm isotropic. The focus of the evaluation sought to map motion-selective sites instead of obtaining high-quality 3D frames that could be commonly quantified by quantitative metrics such as PSNR.
3.3. Qualitative Results of 3D Frames
As Fig. 2 demonstrates, the gray-white matter segregation and the overall shape of sulci/gyri are notably clearer in the super-resolved images compared to the downsampled 2mm and 3mm isotropic resolution images driving the SR model. Our model effectively recovers image details across various resolutions without retraining. This enhancement is also evident in functionally mapping the motion-selective sites based on their stronger response to moving versus stationary stimuli (Fig. 3A). Specifically, these fine-scale sites, mostly missing in maps based on lower spatial resolution (Fig. 3B-C), are recovered in the super-resolved images (Fig. 3D-E), even when using a model trained with resting-state data (3E).
Fig. 2:

Image quality enhancement by applying the SR method to low-resolution images. Gray-white matter segregation is more apparent in super-resolved images compared to downsampled images.
Fig. 3:
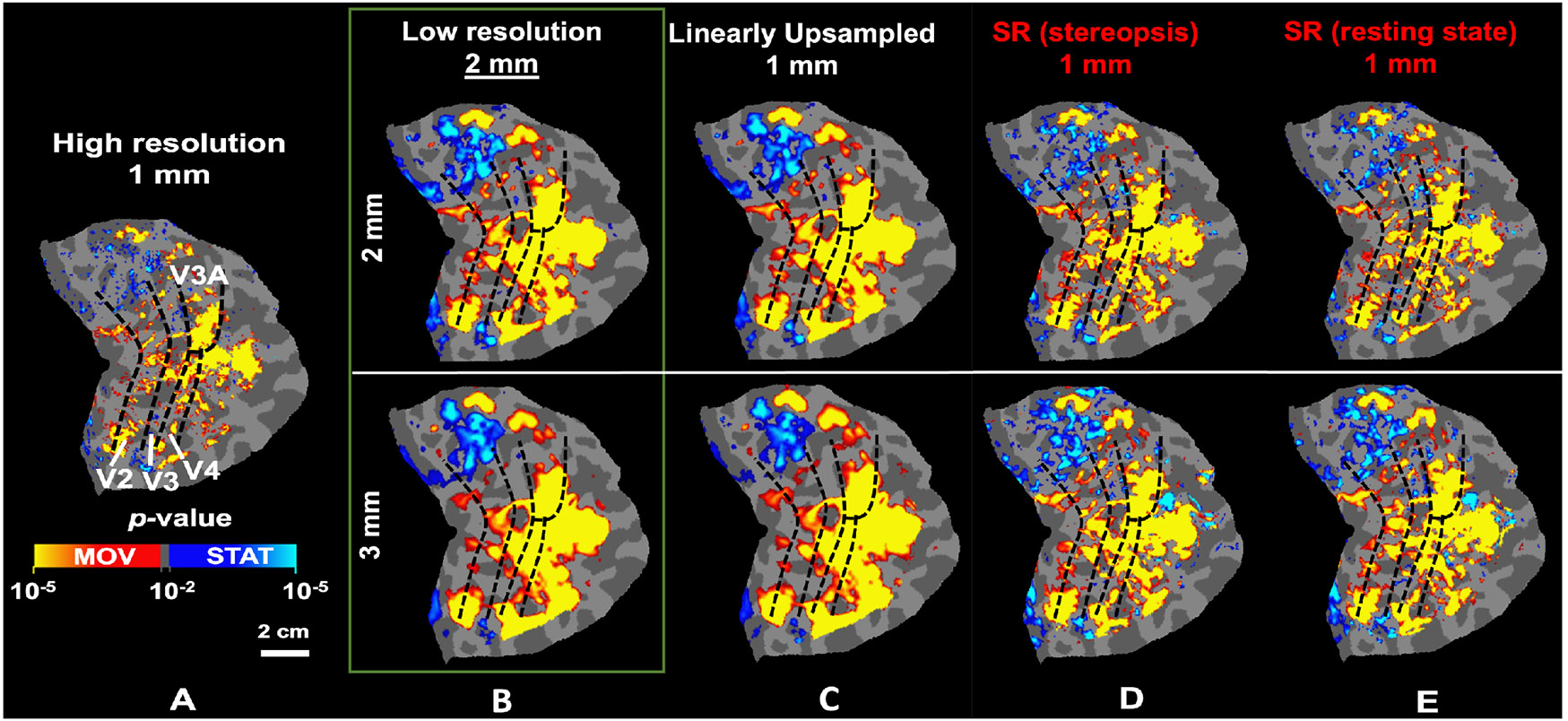
The application of the SR method improves the localization of motion-selective activity maps from low-resolution fMRI. A) Localization of motion-selective sites across V2, V3, V3A, and V4 based on the original high-resolution fMRI. B-C) Localization of the same sites based on downsampled data. Fine-scale sites are either absent or fused, causing overestimation in the size of the selective sites. D-E) Localization of motion-selective sites based on super-resolved images. The fine-scale motion-selective sites are mostly recovered in these maps. In all panels, dashed black lines indicate the borders of visual areas, defined retinotopically.
3.4. Quantitative Results of fMRI Analysis
To quantitatively measure the performance of the SR method, we measure the correlation between the original motion-selectivity map (Fig. 3A) and the recovered maps for each ROI. These measurements are normalized relative to the correlation between the original color-selectivity map (not shown here) and the recovered motion-selectivity maps. The recovered motion-selective maps are more correlated with the original motion-selectivity than the color-selectivity map. Consistently, as shown in Fig. 4, the SR methods enhance the correlation between the original and recovered (compared to LR) maps across all tested ROIs. Moreover, the consistency of observations across five test subjects underscores the robustness of our SR model across subjects and stimuli.
Fig. 4:

Consistency between original motion-selectivity maps and those from downsampled, super-resolved images are shown in two panels for 2mm and 3mm resolution datasets, respectively. The “consistency index” represents the correlation between regenerated and original motion-selectivity maps, subtracted by the correlation with an independent high-resolution color-selectivity map. (VS=visual stimuli, RS=resting state.)
4. SUMMARY AND CONCLUSION
We have demonstrated that deep learning can effectively super-resolve ultra-high-field fMRI images from lower spatial resolutions, ultimately enhancing downstream functional maps – even across subjects and stimuli. The consistency of observations across five test subjects further highlights the robustness of our SR model. Moving forward, our research will focus on training with a larger cohort of subjects to further refine and validate this technique. Additionally, we plan to extend our exploration to include data collected inherently at lower spatial resolutions and field strengths (e.g. 3T) and compare with other image super-resolution methods such as generative adversarial works [12] and diffusion models [17].
ACKNOWLEDGEMENT
Funded by NIH (1RF1MH123195, R01EY030434, R01AG070-988, R01EB031114, UM1MH130981, RF1AG080371), Jack Satter Foundation and Swiss Postdoc Mobility Fellowship.
6. REFERENCES
- [1].Blazejewska AI, Fischl B, Wald LL, and Polimeni JR. Intracortical smoothing of small-voxel fMRI data can provide increased detection power without spatial resolution losses compared to conventional large-voxel fMRI data. Neuroimage, 189:601–614, 2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [2].Chen R, Tang X, Zhao Y, Shen Z, Zhang M, Shen Y, Li T, Chung CHY, Zhang L, Wang J, et al. Single-frame deep-learning super-resolution microscopy for intracellular dynamics imaging. Nature Communications, 14(1):2854, 2023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [3].Cheng K, Waggoner RA, and Tanaka K. Human ocular dominance columns as revealed by high-field functional magnetic resonance imaging. Neuron, 32(2):359–374, 2001. [DOI] [PubMed] [Google Scholar]
- [4].Dong C, Loy CC, He K, and Tang X. Image superresolution using deep convolutional networks. IEEE Trans Pat Anal Mach Intel, 38(2):295–307, 2015. [DOI] [PubMed] [Google Scholar]
- [5].Fischl B. Freesurfer. Neuroimage, 62:774–781, 2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [6].Goddard E and Mullen KT. fMRI representational similarity analysis reveals graded preferences for chromatic and achromatic stimulus contrast across human visual cortex. NeuroImage, 215:116780, 2020. [DOI] [PubMed] [Google Scholar]
- [7].Greve DN and Fischl B. Accurate and robust brain image alignment using boundary-based registration. Neuroimage, 48(1):63–72, 2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [8].Iglesias JE, Billot B, Balbastre Y, Tabari A, et al. Joint super-resolution and synthesis of 1 mm isotropic MP-RAGE volumes from clinical MRI exams with scans of different orientation, resolution and contrast. Neuroimage, 237:118206, 2021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [9].Kennedy B, Bex P, Hunter D, and Nasr S. Two fine-scale channels for encoding motion and stereopsis within the human magnocellular stream. Progress in Neurobiology, 220:102374, 2023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [10].Kornprobst P, Peeters R, Nikolova M, Deriche R, et al. A superresolution framework for fMRI sequences and its impact on resulting activation maps. In MICCAI, pages 117–125, 2003. [Google Scholar]
- [11].LeCun Y, Bengio Y, and Hinton G. Deep learning. nature, 521(7553):436–444, 2015. [DOI] [PubMed] [Google Scholar]
- [12].Li H, Paetzold JC, Sekuboyina A, Kofler F, Zhang J, Kirschke JS, Wiestler B, and Menze B. Diamondgan: unified multi-modal generative adversarial networks for mri sequences synthesis. In Medical Image Computing and Computer Assisted Intervention–MICCAI 2019: 22nd International Conference, Shenzhen, China, October 13–17, 2019, Proceedings, Part IV 22, pages 795–803. Springer, 2019. [Google Scholar]
- [13].Nasr S, Polimeni JR, and Tootell RB. Interdigitated color-and disparity-selective columns within human visual cortical areas V2 and V3. Journal of Neuroscience, 36(6):1841–1857, 2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [14].Ota J, Umehara K, Kershaw J, Kishimoto R, et al. Super-resolution generative adversarial networks with static T2* WI-based subject-specific learning to improve spatial difference sensitivity in fMRI activation. Scientific Reports, 12(1):10319, 2022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [15].Penny W and Friston K. Mixtures of general linear models for functional neuroimaging. IEEE Trans Med Im, 22(4):504–514, 2003. [DOI] [PubMed] [Google Scholar]
- [16].Rudie J, Gleason T, Barkovich M, Wilson DM, et al. Clinical assessment of deep learning–based superresolution for 3D volumetric brain MRI. Radiology:AI, 4(2), 2022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [17].Song Y, Sohl-Dickstein J, Kingma DP, Kumar A, Ermon S, and Poole B. Score-based generative modeling through stochastic differential equations. arXiv preprint arXiv:2011.13456, 2020. [Google Scholar]
- [18].Triantafyllou C, Hoge RD, Krueger G, Wiggins CJ, et al. Comparison of physiological noise at 1.5 T, 3 T and 7 T and optimization of fMRI acquisition parameters. Neuroimage, 26(1):243–250, 2005. [DOI] [PubMed] [Google Scholar]
- [19].Vu AT, Jamison K, Glasser MF, Smith SM, et al. Tradeoffs in pushing the spatial resolution of fmri for the 7T Human Connectome Project. Neuroimage, 154:23–32, 2017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [20].Wang X, Yu K, Wu S, Gu J, et al. Esrgan: Enhanced super-resolution generative adversarial networks. In ECCV workshops, 2018. [Google Scholar]
- [21].Wang Z, Bovik AC, Sheikh HR, and Simoncelli EP. Image quality assessment: from error visibility to structural similarity. IEEE Trans Im Proc, 13(4):600–612, 2004. [DOI] [PubMed] [Google Scholar]
- [22].Wang Z, Chen J, and Hoi SC. Deep learning for image super-resolution: a survey. arxiv:1902.06068, 2019. [DOI] [PubMed] [Google Scholar]
- [23].Wang Z, Chen J, and Hoi SC. Deep learning for image super-resolution: A survey. IEEE Trans Pat Anal Mach Intel, 43(10):3365–3387, 2020. [DOI] [PubMed] [Google Scholar]
- [24].Yacoub E, Shmuel A, Logothetis N, and Uğurbil K. Robust detection of ocular dominance columns in humans using hahn spin echo BOLD functional MRI at 7 Tesla. Neuroimage, 37(4):1161–1177, 2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [25].Zhao X, Zhang Y, Zhang T, and Zou X. Channel splitting network for single MR image super-resolution. IEEE Trans Im Proc, 28(11):5649–5662, 2019. [DOI] [PubMed] [Google Scholar]


