Abstract
Peptides are promising building blocks of designer materials with wide-ranging applications. These materials are stabilized by directional hydrogen-bonding patterns, giving rise to one-dimensional or two-dimensional assembly. It remains a challenge to mimic biology’s context-adaptive and flexible structures. Here we introduce minimalistic tripeptide sequences that form highly soluble dynamic ensembles through multivalent side-chain interactions. We observe these supramolecular dispersions undergo drying-induced sequential liquid–liquid phase separation followed by solidification, resulting in the formation of films of stiff, densely packed and porous peptide microparticles that can be instantaneously redispersed upon the re-introduction of water. Air-drying of peptide dispersions in the presence of proteins or small-molecule payloads results in efficient encapsulation and the retention of protein stability after redispersion, showing promise for the emulsification, encapsulation, protection and storage of biomacromolecules. The mechanism resembles the protective strategies in natural systems during desiccation, which rely on liquid–liquid phase separation to survive extreme conditions.
Subject terms: Self-assembly, Self-assembly
Here the authors design tripeptides that form dynamic soluble dispersions and undergo phase separation upon drying to assemble into porous particles. This evaporation-driven emulsification can be harnessed to encapsulate and stabilize biomolecules.
Main
All life forms share conserved sets of building blocks. Biomolecular covalent and non-covalent interactions dictate the rich structural design space, supporting wide-ranging functions, including recognition, self-assembly, catalysis and shapeshifting. Beyond their critical roles in the chemistry of life, these structures provide inspiration for supramolecular materials1–4 and systems5 for various technological and biotechnological applications6–8.
Peptide materials usually contain patterns of backbone hydrogen bonding derived from protein secondary structures9–13, typically giving rise to ordered one-dimensional (1D)14, two-dimensional (2D)15,16 or three-dimensional (3D)17 assemblies. These hydrogen-bond patterns may be further stabilized by aromatic stacking18,19, giving rise to architectures that are remarkably stiff and stable20. This is exemplified by ‘dry’ diphenylalanine (FF) zippers21, where the chiral organization of side chains provides additional control over assembly structures22. We previously established that aromatic tripeptides with polar groups, such as lysine–tyrosine–phenylalanine (KYF), retain strong directional self-assembly tendencies complemented with favourable solvent interactions, leading to hydrogelation23.
Besides ordered structures, the notion of non-directional assembly is also common in biology, for example, in liquid condensates, where it typically leads to spherical morphologies dictated by surface tension24,25. In these systems, side-chain interactions dictate the assembly, complementing flexible backbone interactions26. Short peptide motifs that display liquid–liquid phase separation27, including through flexible linkers separating aromatic dipeptides to disrupt directionality28, as well as the formation of unstructured solid glassy states29,30 have been reported.
Beyond structures with varying instrinsic directionalities through backbone or side-chain interactions, biology frequently employs architectures that display structural agility in response to external factors. This may be achieved through dynamic selection and adjustment from the complex side-chain interaction space that is available5. This concept has been demonstrated in synthetic systems, where the adaptive versatility of biomimetic random heteropolymer sequences with flexible backbones and multiple (randomized) functional side chains have been shown to adaptively decorate and wrap proteins31 or to recapitulate the dynamic features of protein ensembles in biofluids32.
Here we introduce sequence designs of tripeptides that are not isotropic through backbone hydrogen-bonding interactions but instead form dynamic ensembles through multiple possible combinations of weak and reversible interactions, by combining polar, hydrogen-bonding aromatic and basic residues (tryptophan (W), tyrosine (Y) and lysine (K); Fig. 1a). We show that these peptides form aqueous dispersions (Fig. 1b) with high aqueous solubility and unique context-adaptive properties, including dehydration-induced liquid–liquid phase separation and the formation of mechanically stiff particles with the ability to wrap, encapsulate, stabilize and protect proteins, while retaining instantaneous dispersibility upon rehydration.
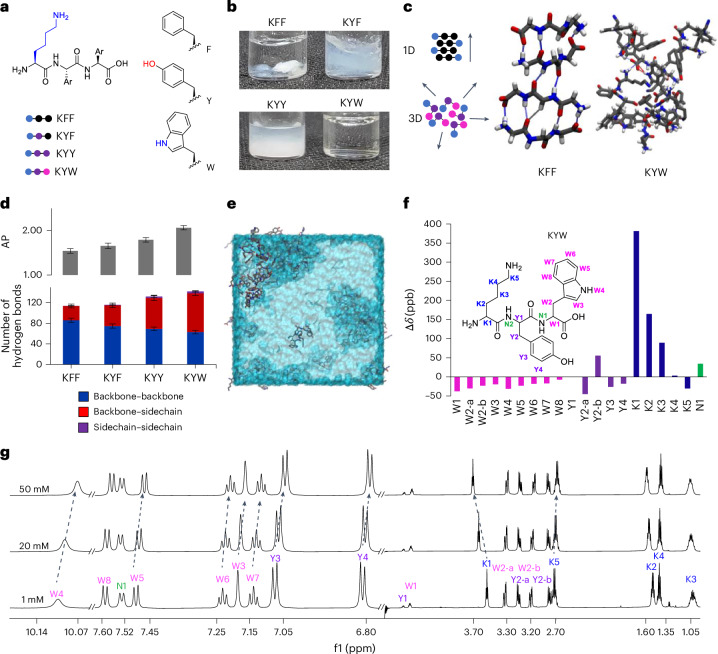
Fig. 1. Directionality and dispersibility in peptide self-assembly.
a, Chemical structures of tripeptides KFF, KYF, KYY and KYW. b, Macroscopic images of KFF, KYF, KYY and KYW peptides at 20 mM in phosphate buffer (PB) at pH 7.5. c, Selected snapshots of simulations for KFF (only backbone shown for clarity) and KYW, with predominant hydrogen-bond interactions highlighted. d, Sequence-dependent AP = SASAinitial/SASAequil (top) and distribution of hydrogen-bond interactions (bottom) between simulated peptides from MD trajectories for KFF, KYF, KYY and KYW. The backbone and side chain indicate the directionality of the self-assembly. Average and s.d. error bars were measured from three replicate simulations using 250 data points (the last 50 ns) of each analysis set. e, Snapshot from MD trajectory of KYW showing assembly inside a water box. f, Chemical shift perturbation (Δδ in parts per billion) for KYW between 50 mM and 1 mM concentrations. This figure highlights the evolving chemical environment for each KYW’s protons (colour-coded accordingly) during concentration-driven assembly. Downfield shifts are represented as positive values and upfield shifts as negative values, reflecting the strengthening or weakening of hydrogen bonding and π-type interactions, respectively. Each bar represents the average of three independent experiments, with s.d. error bars from three replicates. g, The 1H NMR spectra of KYW at concentrations of 1 mM, 20 mM and 50 mM in 99:1 D2O/H2O with 100 mM Na/PB at pH 7.5, illustrating concentration-dependent shifts in f1 (frequencies) that reveal key interactions in the formation of dynamic soluble ensembles.
Results
Directionality and dispersibility in peptide self-assembly
Tripeptide assembly is favoured by sequences of two aromatic amino acids complemented with a charged residue23,33. Aromatic amino acids (Y and W; Fig. 1a,c) add a diversity of interactions beyond backbone hydrogen-bond directionality, including side-chain π-stacking, hydrogen-bonding and dipole–dipole interactions, importantly, including those with water34. Lysine–phenylalanine–phenylalanine (KFF), KYF and lysine–tyrosine–tyrosine (KYY) undergo gelation and phase separation, but lysine–tyrosine–tryptophan (KYW) remarkably retains the appearance of a clear solution up to 20 mM in phosphate buffer23 (Fig. 1b). Notably, KYW assembly is strongly dependent on the ionic context and on forming fibres and gels in unbuffered conditions23,35.
We used atomistic molecular dynamics (MD) simulations to understand differences in assembly and hydrogen-bond interactions in KFF, KYF, KYY and KYW (Fig. 1d and Supplementary Fig. 11). First, exchanging F → Y → W leads to an increase in the aggregation propensity (AP; Fig. 1d and Supplementary Fig. 12). We analysed the contribution of the backbone–backbone (as a proxy for 1D self-assembly) and side-chain hydrogen bonds (Fig. 1d and Supplementary Figs. 13–16). Consistent with the increased AP, we observed a concomitant increase in the total number of hydrogen bonds in F → Y → W variants, with decreased backbone and increased side-chain hydrogen bonds resulting in non-directional assembly for KYW (Fig. 1d).
We investigated the concentration-dependent behaviour of the KYW peptide using extensive 1H nuclear magnetic resonance (NMR) analyses at 1, 20 and 50 mM in pH 7.5 phosphate buffer (Fig. 1f–g and Supplementary Fig. 17 and accompanying text). Across this range, KYW showed no line broadening or signal loss, indicating that it remains highly dynamic and fully soluble even at higher concentrations. The assembly process is driven primarily by π-type hydrophobic interactions, side-chain hydrogen bonding and electrostatic interactions, consistent with a dynamic, non-aggregating state.
The relative contributions of side chains and backbone interactions when substituting F → Y → W are also evident in the circular dichroism spectra (Supplementary Figs. 18–20). At a concentration of 5 mM, which is just above the critical aggregation for KYF and KYY (Supplementary Fig. 20a,b), the spectra show a carbonyl π → π* transition peak at 203 nm (Supplementary Figs. 18, 19a,b and 20a,b), suggesting backbone–backbone interactions, while the transitions of aromatic side chains may also contribute22. For KYW, this transition is less intense, while the appearance of the 229 nm positive signal that increases with concentration (Supplementary Figs. 18 and 19c) confirms the dominating influence of tryptophan side-chain interactions8,36.
Sequence dependence in tripeptide dispersions
We previously demonstrated differential assembly of tripeptide sequence isomers driven by side-chain-induced conformation selection33. We speculated that in a system stabilized by a multitude of possible side-chain and water interactions, these would not be defined by one dominant conformation but instead represented by a rich ensemble of conformers. We studied the six K/Y/W sequence isomers to determine the sequence-dependent self-assembly (Fig. 2a and Supplementary Figs. 21–26) and found lower variation in AP values (Fig. 2a and Supplementary Fig. 21) and similar ratios of side-chain/backbone interactions (Fig. 2a and Supplementary Figs. 22–25), consistent with the formation of non-directional ensembles. We also analysed the solvent exposure of tryptophan, measured through the solvent-accessible surface area (SASA; Fig. 2a and Supplementary Fig. 26) of the indole side chain. This was found to be sequence dependent, with tryptophan–tyrosine–lysine (WYK) and tryptophan–lysine–tyrosine (WKY) being the most contrasting in terms of AP and tryptophan solvent exposure (Fig. 2b).
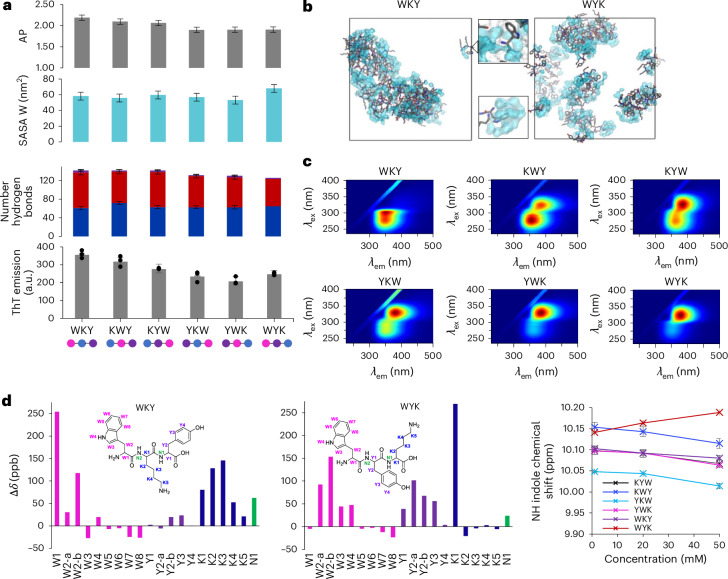
Fig. 2. Sequence dependence in tripeptide dispersions.
a, From top to bottom, the AP score, tryptophan SASA and hydrogen-bonding interaction distribution analyses for K/Y/W sequence isomer MD trajectories. Averages and s.d. error bars were measured from three replicate simulations using 250 data points (last 50 ns) of each analysis set. Below is the ThT fluorescence emission intensity (50 µM) in the presence of peptide at 20 mM. Averages and s.d. error bars from three replicates. Data represent mean ± s.d. (n = 3). b, Snapshots for WKY and WYK showing non-directional self-assembly resulting in differential tryptophan environments and water binding (shown in blue) with zoomed-in areas showing a dry (top) and water-bound (bottom) tryptophan. c, The 3D fluorescence spectra showing the differential polarity and water interactions of tryptophan residues in solution. λem, emission wavelength. d, Chemical shift perturbation (Δδ in parts per billion) for WKY and WYK between concentrations of 50 mM and 1 mM. Each plot corresponds to a different sequence, with downfield shifts represented as positive values and upfield shifts as negative values. Each bar represents the average of three independent experiments, with s.d. error bars from three replicates. e, Concentration-dependent chemical shifts of the indole NH proton in K/Y/W sequence isomers, highlighting variations in the chemical environment and intermolecular interactions as concentration increases (1, 20 and 50 mM). Averages and s.d. error bars from triplicates.
Tryptophan exposure in peptide ensembles could be experimentally verified by measuring the tryptophan emission, which is widely used as a probe to detect the tryptophans’ environment and solvent exposure in proteins, with redshifted emissions indicating an increase in local polarity and blueshifts indicating hydrophobic environments37–39. These ensembles (20 mM) did not show evidence of phase separation when observed by optical microscope (shown for KYW in Fig. 1b). We measured the emission spectra of all sequence isomers in soluble dispersions (Fig. 2c) and found that emission maxima (excitation wavelength λex = 280 nm) range from 357 nm to 370 nm, with the most redshifted emission corresponding to the most hydrated tryptophan (WYK) and the most blueshifted emission corresponding to the most hydrophobic tryptophan environment (WKY; Fig. 2b). The 3D excitation/emission spectra of contrasting sequences (Fig. 2c) differed by the appearance of a new band in WYK’s spectrum at λex = 318 nm, which is also prominent in tyrosine–lysine–tryptophan (YKW) and tyrosine–tryptophan–lysine (YWK). We also found that the intensity and absence/presence of the band is concentration dependent, suggesting its origin from an emissive state of a supramolecular arrangement (Supplementary Fig. 27).
Concentration-dependent NMR studies on all K/Y/W sequence isomers revealed that while all maintain high dynamicity and solubility up to 50 mM, π-type, cation–π, hydrogen-bonding and electrostatic interactions contribute across all isomers, and the nature and strength of intermolecular interactions vary with sequence (Fig. 2d, Extended Data Fig. 1 and Supplementary Fig. 28 and accompanying text).
Extended Data Fig. 1. Chemical shift for each proton in various peptide sequence isomers.
Chemical shift perturbation (Δδ in ppb) between 50 mM and 1 mM concentrations for each proton in various peptide sequence isomers. Each plot corresponds to a different sequence, with downfield shifts represented as positive values and upfield shifts as negative values. Each bar represents the average of three independent experiments, with s.d. error bars from 3 replicates.
The local chemical environment of tryptophan residues was probed by following the NH (W4) and CH (W3) indole protons in the 1H NMR spectra of the six K/Y/W peptide solutions at variable concentrations and showed contrasting concentration-dependent chemical shifts of tryptophan protons (Fig. 2e and Supplementary Fig. 29). The indole-ring protons in WYK shift downfield as the concentration increases, indicating their positioning in an electron-poor environment and the formation of a polar network and hydrogen bonding, while (minimal) upfield shifts were observed for WKY, in line with the formation of π-stacking-driven aggregation. The other four sequences exhibited moderate upfield shifts, less pronounced than those of WKY, in agreement with the MD simulation data (Fig. 2a).
To quantify the degree of assembly across peptide sequences, we monitored changes in thioflavin T (ThT) emission upon co-incubation with 20 mM peptide solutions (Fig. 2a), revealing enhanced fluorescence consistent with the formation of dynamic hydrophobic pockets. This sequence-dependent ThT response, aligned with calculated AP scores and supported by NMR chemical shifts, confirms the presence of soluble dispersions featuring dynamic hydrophobic domains.
To further probe dynamic ensembles in solution, we conducted fluorescence spectroscopy across increasing concentrations (2–20 mM), observing progressive tryptophan quenching (Supplementary Fig. 30). Concentration-dependent diffusion changes, analysed by 1H diffusion-ordered spectroscopy (DOSY) NMR, were performed at 1, 20 and 50 mM (Supplementary Fig. 31), revealing decreased diffusion with increasing concentration. The peptide-to-water diffusion ratio suggests small soluble aggregates, consistent with unbroadened 1H NMR signals. The 1H–1H nuclear Overhauser effect spectroscopy (NOESY) NMR spectra for all six K/Y/W tripeptides highlight water interactions, predominantly at W4 and W3 (Supplementary Fig. 32). WYK and YWK exhibit enhanced water contacts at positions W2 and W1, consistent with higher SASA values (Fig. 2a), while WKY also shows a W1–water correlation that aligns with the 1H NMR downfield shift (Fig. 2d and Supplementary Fig. 28 and accompanying text), likely due to its N-terminal location.
Evaporation-driven assembly
Upon investigating the fluorescence of KYW peptide assemblies using confocal microscopy, we observed a remarkable phase separation behaviour triggered by localized evaporation during drying. Given the (competing) role of water in modulating interactions in these dynamic ensembles, in particular those involving tryptophan40, the observation led us to study the assembly during gradual removal of water through the evaporation of sessile droplets41–43 (Fig. 3a).
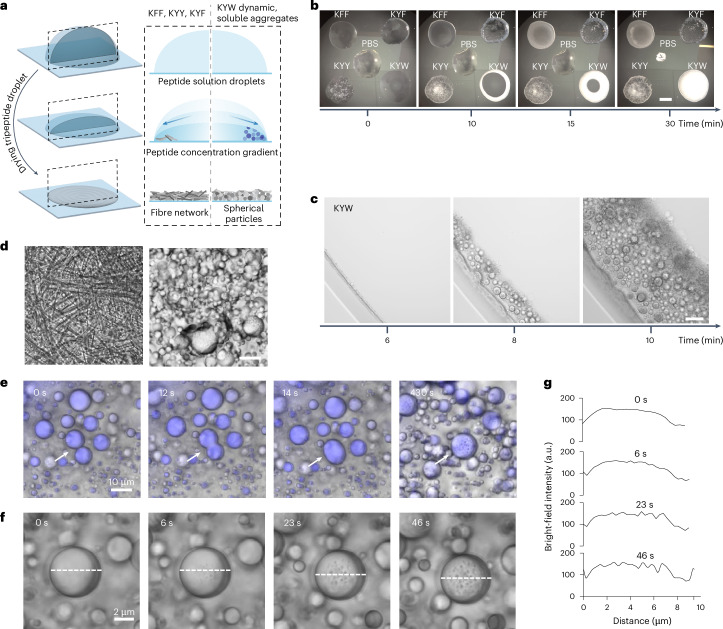
Fig. 3. Evaporation-driven assembly.
a, Representation of evaporation-driven self-assembly in sessile droplets. b, Evaporation patterns over time, with the differential assembly of KYW clearly visible. Scale bar, 1 mm. PBS, phosphate-buffered saline. c, Microscopy time course for evaporation-driven assembly of KYW. Scale bar, 50 µm. d, Bright-field microscopy images showing 1D assembly for KYY (left) and the formation of spherical particles upon drying for KYW (right). Scale bar, 10 µm. e, The dynamic droplet fusion process of WKY is recorded by time-lapse confocal microscopy. From 0 s to 14 s, a transient fusion of two liquid–liquid phase separation droplets is seen. By 430 s, the rigidification of the droplets and appearance of pores on the surface are seen. f,g, Time-lapse confocal analysis (f) revealed the onset of pore formation upon solidification. The moment that the droplet stationized was set as 0 s. The spectrum of bright-field images across the particle (white dashed lines) during the solidification process were profiled (g) for quantitative visualization.
Macroscopically, when observing tripeptide assembly in drying droplets starting from 20 mM peptide dispersions, we observed contrasting behaviours: a strongly scattering opaque ring upon drying for KYW, in contrast to KFF, KYF and KYY, which lead to heterogeneous semi-transparent structures (Fig. 3b and Supplementary Videos 1 and 2). When observed by microscopy (Fig. 3d and Supplementary Fig. 33), the differential morphologies (1D fibres versus 3D spherical objects) are evident (Supplemental Fig. 34 showing KFF and KYF). Over time, KYW and its five sequence isomers all display a remarkable drying-induced phase separation behaviour, giving rise to the formation of films composed of densely packed spherical particles (Fig. 3d, Supplementary Video 3 and Extended Data Fig. 2a,b), contrasting with KFF, KYF and KYY, which lead to 1D structures resembling previous observations for the evaporation-driven assembly of peptide derivatives43 (Supplementary Fig. 34, Extended Data Fig. 2a and Supplementary Videos 4–6). We propose that the multitude of possible interactions and the flexibility of the backbone enable the K/Y/W peptides to more effectively reconfigure and adapt to the newly formed interfaces that form upon water evaporation. The observed directionality (in the case of KFF, KYF, KYY) or lack of directionality (in the case of K/Y/W) initiates at the molecular level and clearly translates hierarchically to macroscopic scales. We observe complex drying patterns related to the solvent and solute gradients in the drying of droplets, with phase separation occurring at the drying front due to concentration gradients inside the drying droplet. Streaks observed during drying are a consequence of complex Marangoni flows in these drying droplets (Extended Data Fig. 2c–f and Supplementary Video 7).
Extended Data Fig. 2. Spherical particles upon drying.
a.Bright field microscopy images showing formation of spherical particles upon drying for KYW, KWY, YKW, YWK, WKY, and WYK. Scale bar = 10 µm; b. Bright field microscopy images of the time course of KWY, WKY, WYK, YWK and YKW evaporation process; c-f. Streaks formation during the evaporation-driven peptide solution drying. The moving tracks of the KYW droplets during the drying process were reconstructed using Imaris; d-e. based on the timelapsing confocal images. (a). The sum of the tracks in (e) shows the streaks formed at the edge as observed in (f). Scale bar, 50 µm in c, d, e and 500 µm in f. Color scale bar in (e) represents time.
Based on the previously reported interfacial supramolecular aggregation of tripeptides in oil–water emulsions35, we hypothesized that the ensembles would lead to interfacial aggregation and a reduction in surface tension. We measured the dynamic tension profile via the pendant drop method44 of the six K/Y/W tripeptides, which have contrasting tryptophan solvent exposures and environments. The dynamic surface tension indicates that the peptide sequence dictates the surface activity, with a general trend observed in which less solvent-exposed tryptophan (in WKY) gives rise to higher surface activity (Supplementary Fig. 35), demonstrating that the peptide ensembles dynamically interact at the air–water interface.
Time-dependent cryo-electron microscopy (cryo-EM) images illustrate the nanoscale structural transformations from amorphous aggregates to spherical droplets to porous droplets observed at the early stages of assembly (Extended Data Fig. 3). Dynamic droplet fusion and subsequent solidification is captured by time-lapse confocal microscopy (Fig. 3e, Supplementary Fig. 36 and Supplementary Video 8). Time-dependent confocal analysis revealed the onset of pore formation upon solidification (Fig. 3f,g). Time-dependent fluorescence spectroscopy during the drying-induced assembly gave rise to a decreased emission due to concentration-induced quenching, followed by a sharp fluorescence increase upon liquid–liquid phase separation (5–10 min), and subsequent quenching during droplet solidification (Supplementary Fig. 37).
Extended Data Fig. 3. Cryo-EM micrographs of early stages of drying-induced assembly.
a. Drop of 2 µl WYK solution was applied onto the TEM grid and went through evaporation under ambient conditions for indicated time and proceeded with cryo-EM. Cryo-EM images illustrate the structural transformations from random aggregates and short fibers to spherical assemblies. Scale bar = 500 nm; b. Cryo-EM micrographs of different resolutions captured at 5’03” show the process of evaporation-driven porous droplet formation, and the temporal information could be decoded from spatial information; c. Different stages of the assembly process are captured. 1. early stage formation of nanosized spherical assembly; 2-4. nano-sized air pockets inside spherical peptide assemblies; 5-7. larger spheres with multiple pores; 8. fusion of larger spheres with multiple pores. Scale bar = 200 nm.
Upon analysis of the tryptophan emission in the dried films at 5% relative humidity, we noted a general loss of emission from the 318 nm band in the 3D excitation/emission spectra, with spectra of the dried films now suggesting a redistribution of conformations (Supplementary Figs. 38 and 39). The original variable tryptophan exposure was partly recovered by temporarily exposing the solid film to high humidity (95% relative humidity). These data highlight the adaptive nature of the assemblies, in which tryptophan partitioning reversibly changes depending on the water competing via weak multivalent interactions.
Formation of buoyant droplets and porous particles
Upon microscopy analysis of the dried particulate films, we observed the expected spherical objects, but also porous, disc-shaped particles of varying sizes with flat surfaces (Fig. 4a). These structures are found at the top of the sample (Fig. 4b,c and Extended Data Fig. 4), suggesting that they are formed at the air–water interface. The observation suggests that, in contrast to typical dense coacervate droplets that form as a sediment, the structures studied here are low-density, buoyant condensates. The observation of flattened 2D hemispherical objects suggests that these particles are initially liquid, and due to their low density, they rise to the interface, where they deform, flatten and solidify. This optical observation (Supplementary Fig. 41) is confirmed using focused ion beam scanning electron microscopy (FIB-SEM; Fig. 4c), which shows a part-circular cross-section. The pore size distribution ranged from 30–500 nm (Supplementary Fig. 40).
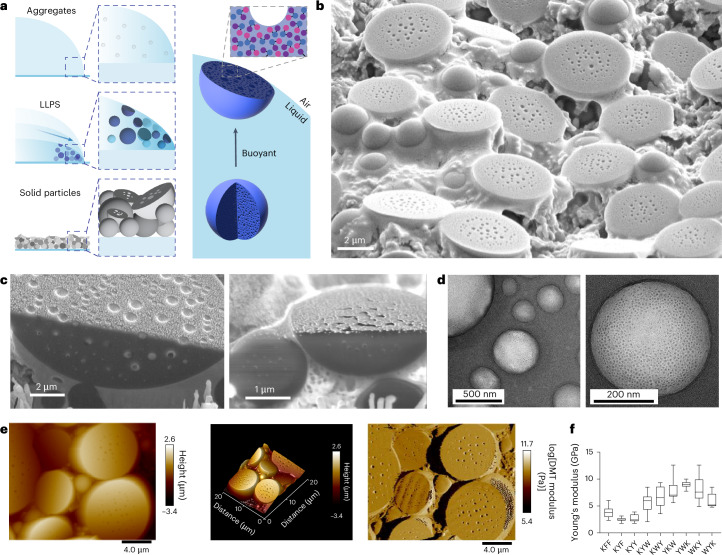
Fig. 4. Formation of buoyant droplets and porous particles.
a, Schematic representation of evaporation-driven assembly of buoyant liquid droplets that settle at the droplet interface and, upon drying, form half-dome-shaped particles. LLPS, liquid–liquid phase separation. b, SEM image showing a mixture of porous spheres and half-dome particles at the interface and also showing surface pores. c, FIB-SEM analysis confirming the half-dome shape and revealing the porous structure inside the dried particles. Note that the fine granular structures on the surface are from sputter-coated gold. d, TEM analysis of glutaraldehyde-crosslinked dissociated KWY particles reveals that non-surface particles are highly porous spheres. e, AFM analysis showing porous half-dome particles. f, The plot of Young’s modulus showing the stiffness of fibres or particles formed by the indicated peptides. A total of 15–36 particles or fibres were profiled for each sequence. In the box plots, the centre line shows the median, the box edges delineate the first and third quartiles and the whiskers show the range of values.
Extended Data Fig. 4. SEM and AFM micrographs of KFF, KYF, KYY, KYW, KWY, YKW, YWK, WKY, WYK.
The first column shows SEM images of all nine tripeptides. KFF, KYF and KYY show similar linear fiber-like structures. The other six tripeptides show sphere structures of various sizes, each of such spheres containing hollow bubbles. The second, third, and forth columns show the height, 3D topography, and stiffness (Young’s Modulus) AFM images.
We propose that the observed buoyancy is related to the formation of gas bubbles inside liquid droplets (Fig. 4a). This may be a consequence of a loss of solvating water from the peptide structures, water that is replaced with air and leads to micro-cavitation, resulting in a porous structure upon dehydration (Fig. 4a). Notably, degassing of the peptide dispersion prior to evaporation-driven assembly leads to smaller particles of more spherical morphology (Fig. 5e). This observation indicates that the buoyant nature of the droplets relates to their gas content, and it can be regulated. Non-surface particles are smaller and spherical, indicating fusion and flattening at the air–water interface (Fig. 4b,c). Transmission electron microscopy (TEM) analysis of glutaraldehyde-crosslinked KWY particles reveals the porous and spherical structure of the non-surface particles (Fig. 4d). The hemispherical porous structure was further confirmed by atomic force microscopy (AFM; Fig. 4e and Extended Data Fig. 4). A Young’s modulus from Derjaguin-Muller-Toporov (DMT) model fitting analysis demonstrated that these particles, formed by all sequence isomers, are mechanically stiff (6 GPa) due to the efficient packing (Fig. 4f), in line with previously reported FF derivatives20,21.
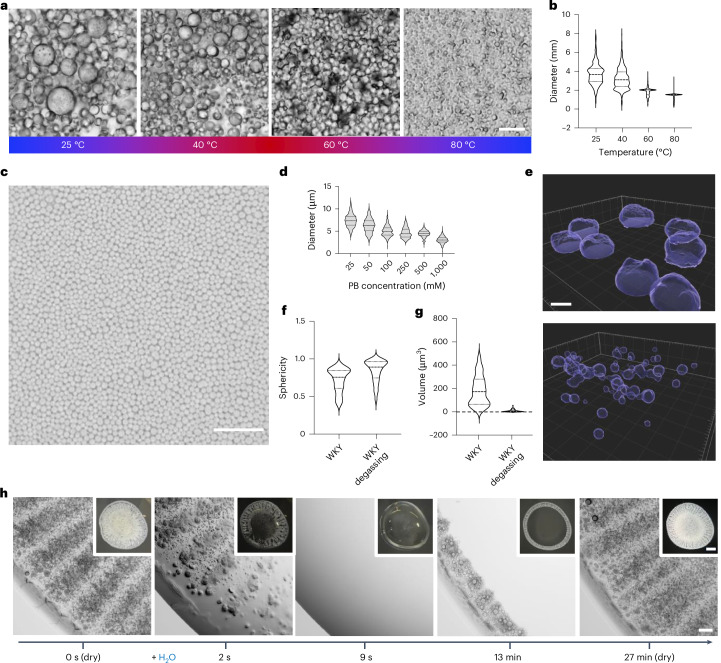
Fig. 5. Size control, shape control and reversibility.
a,b, Temperature dictates the size distribution of the dried peptide particles. KWY shows a reduction in particle size and more homogeneous size distribution as temperature increases during evaporation. Scale bar, 10 µm. A total of 368–1,456 particles at each temperature were analysed. c, The 20 mM WKY peptide was dissolved in Na/PB at concentrations of 25, 50, 100, 250, 500 and 1,000 mM. A bright-field image of dried WKY in 500 mM PB is shown. Scale bar, 50 µm. d, Plot of 296–662 Imaris-3D-rendered particles at each concentration demonstrates the reduction in particle size and increase in homogeneity with the increase of salt concentration. e–g, The peptide solution was degassed, followed by the evaporation assay; the example of WKY is shown (e). Imaris 3D rendering demonstrated a reduction in particle size and increase in sphericity, as plotted in f and g, respectively. Scale bar, 5 µm. h, Fully reversible formation of porous peptide particles upon addition of water and re-evaporation. Representative time-lapse images of KWY are shown. Scale bar, 50 µm. The macroscopic images of 5 µl peptide solution drops at each stage are shown in the insets. Scale bar in the insert, 500 µm. In the violin plots, the horizontal lines show the median as well as the 25th and 75th percentiles.
Size control, shape control and reversibility
We first investigated whether the size distribution of the particles formed could be controlled. Given the important role of water evaporation in this process, we reasoned that by simply increasing the temperature, evaporation would occur faster and consequently, the formation of solid particles would be accelerated. We observed that temperature increase leads to more homogeneous and smaller particles (Fig. 5a,b and Extended Data Fig. 5a,b). The combination of reduced buoyancy, due to lower gas content at elevated temperature, with the acceleration of the phase transition and solidification yields particles that are monodisperse (Extended Data Fig. 5c,d and Supplementary Fig. 41). Another approach to control sphericity was through an increase of the buffer concentration, which results in smaller and monodisperse particles (Fig. 5c), observed through an accelerated phase change and solidification (Fig. 5d and Extended Data Fig. 5e).
Extended Data Fig. 5. Control of particle size and shape.
a. The effect of temperature on the size of porous peptide particles formed by tripeptides, and KWY was shown. For Imaris 3D Rendering, traceable blue fluorescence signals excited by 405 nm were captured by confocal coupling with bright-field imaging. Spot rendering was employed to analyze the diameter of the particle; b. while the actual shape was revealed by surface rendering in high transparency mode; c-d. The Sphericity of approximately 200 particles at 40 °C and 1000 particles at 80 °C were profiled, indicating that at high temperature, droplets are not given sufficient time to accumulate at the interface and consequently remain rounded. In violin plot, the horizontal lines display the median as well as 25th and 75th percentiles.); e. Buffer concentration regulates the dynamics of tripeptide particle formation. 20 mM WKY peptide was dissolved in phosphate buffer at the concentrations of 25, 50, 100, 250, 500, 1000 mM. Imaris 3D rendering of the dried particles at each concentration demonstrated reduction in particle size and increase in homogeneity with the increase of salt concentration. Color scale bar represents size, scale bar = 10 µm; f. The peptide solution was degassed followed by evaporation assay, and the example of WKY is shown. Time-lapse confocal imaging of bright field and 405 nm excitation demonstrated reduction in particle size. Scale bar = 50 µm; g. Further AFM analysis of WKY shows fewer and smaller pores in the dried particles of the degassed sample.
As an alternative method to control particle size, we removed the air from the solution through sonication under a vacuum, followed by storage under argon gas, and examined the drying process in a degassed solution under argon. Live confocal time-lapse imaging of the particle-forming process revealed smaller droplets (Extended Data Fig. 5f). Further Imaris 3D reconstruction and analysis demonstrated a 20-fold reduction in particle volume and increase in sphericity in the degassed samples compared with a prevalence of disc-shaped particles in non-degassed samples (Fig. 5e–g and Supplementary Videos 9 and 10). Consistent with the role of air in pore formation, degassed peptide particles show fewer and smaller pores (Extended Data Fig. 5g).
Next we probed the reversibility of the process. Upon a re-introduction of a 5 µl droplet of water on top of a dried film, the structures immediately redispersed, as shown in Fig. 5h. When this droplet was left to dry again, the particle film reappeared after a few minutes, demonstrating that the process is fully reversible (Supplementary Video 11). These observations strongly contrast with those previously reported for peptide materials, where the formations of 1D and 2D structures are highly stable due to a large enthalpic gain from cooperative hydrogen bonds, which are not readily reversed20,21. Peptide ensembles with multivalent, weak and dynamic interactions offer more reversible assemblies, as water solvation can compete with the energetics of the interaction space5.
Drying-induced encapsulation and protection
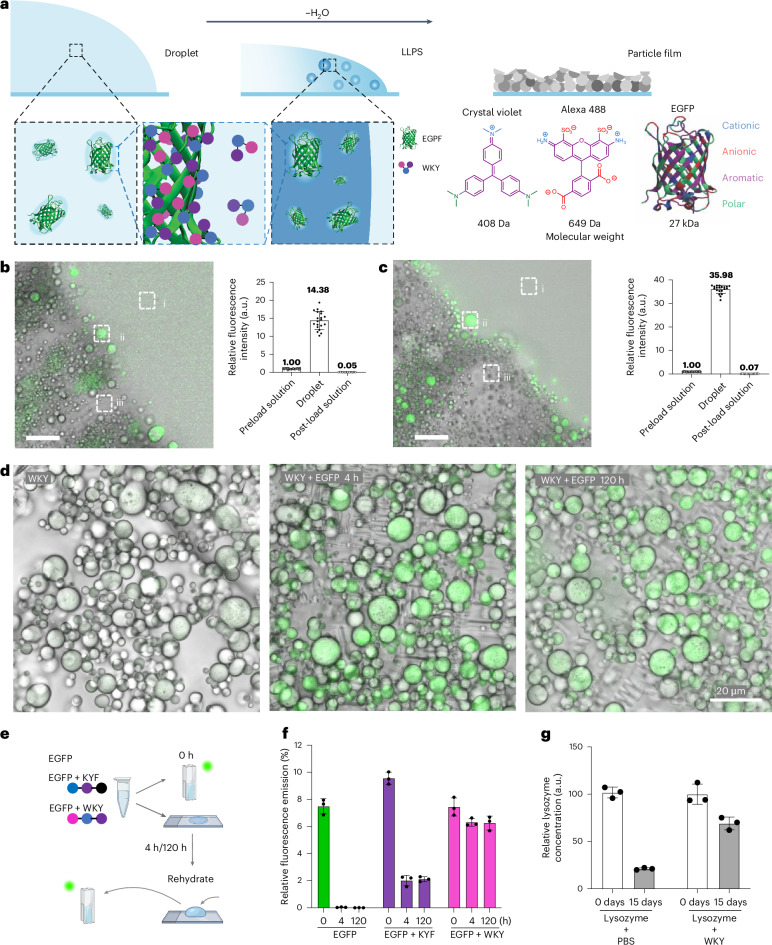
We propose that the spontaneous formation of tightly packed particles upon air-drying could be useful in encapsulation, protection and drying strategies, inspired by phase-separation protection mechanisms seen in biological systems45 (Fig. 6a). We first used the small organic fluorophore Alexa 488, which has been previously demonstrated to non-covalently conjugate through the sulfonate group and lysine side chains46. Upon introduction of the dye, it is encapsulated during KWY droplet formation and solidification (Fig. 6b and Supplementary Video 12). Analysis of the fluorescence intensity of the preload mixture of peptides and dye, the formed liquid droplets and the post-load solution revealed a highly efficient enrichment of the dye inside the particles. Similar efficient encapsulation was obtained for crystal violet, a positively charged hydrophobic dye (Extended Data Fig. 6).
Fig. 6. Drying-induced encapsulation and protection.
a, Proposed mechanism for payload uptake involving dynamic complexation of payload molecules through dynamic side-chain interactions, followed by the molecules being surrounded to form droplets upon liquid–liquid phase separation. b, Confocal live imaging (left) of KWY peptide particles incorporating Alexa 488 shows efficient enrichment of the dye in peptide droplets. Scale bar, 20 µm. Fluorescence intensity analysis (right) of the preload solution (i), the droplet (ii) and the post-load solution (iii) at the frontier of emulsification. A total of 20 regions of interest of each group from six time-lapse frames were quantified, and the average intensity of the preload solution was set as 100 arbitrarily. Data represent mean ± s.d. (n = 20). Error bars represent s.d. c, Encapsulation assay of the EGFP protein by the WKY peptide was performed as described in b. Data represent mean ± s.d. (n = 20). d, Confocal and bright-field imaging of dried peptide particles reveal that the green fluorescence signals of EGFP remain in WKY peptide particles under ambient conditions for five days after solidification. e,f, Experiment to test the storage and redispersion ability of WKY with a schematic representation (e) of the experiment and the fluorescence emission at 510 nm (λex = 488 nm) of the corresponding solution (EGFP, EGFP + KYF and EGFP + WKY) over time (f). Data represent mean ± s.d. (n = 3). Error bars represent s.d. from three replicate experiments. g, The amount of reactive lysozyme in the dried particles with PBS or WKY at day 0 and day 15 under ambient conditions were measured by enzyme-linked immunosorbent assay (ELISA). Data represent mean ± s.d. (n = 3).
Extended Data Fig. 6. Dye encapsulation.
a-b Tryptophan fluorescence spectra of WKY (20 mM in 100 mM PB pH 7.5) in the absence and presence of dye at increasing concentrations (10 µM, 100 µM, 500 µM): (a) Alexa 488, (b) Crystal Violet. c. Confocal live imaging of WKY peptide particles incorporating crystal violet shows substantial enrichment of the dye in peptide droplets, and the dried particles post-encapsulation were shown in d. Scale bar = 50 µm. e. Fluorescence intensity analysis of the pre-load solution, the peptide droplet, and the post-load solution at the frontier of emulsification. A total of 20 regions of interest of each group from 6 time-lapse frames were quantified, and the average intensity of pre-load solution was set as 1 arbitrarily. Error bar represents s.d.
Next, inspired by the ability of dynamic interacting block copolymers with random distributions of side-chain functionalities to stabilize proteins through an adaptive matching of side-chain interactions to complementary patches on the protein surface31, we investigated the ability to encapsulate and stabilize enhanced green fluorescent protein (EGFP). We observed efficient encapsulation in WKY (Fig. 6c, Supplementary Fig. 42a,b and Supplementary Video 13). By comparing the encapsulation efficiency across the six sequence isomers, we conclude that both the Alexa 488 and EGFP encapsulation efficiencies are comparable for all peptide sequence isomers studied (Extended Data Fig. 7).
Extended Data Fig. 7. Uniform Alexa 488 and EGFP encapsulation across all six K/Y/W sequence isomers.
The six sequence isomers of KYW, KWY, YKW, YWK, WKY, WYK demonstrated comparable encapsulation efficiency for both Alexa 488 dye; (a) and EGFP protein (c) analyzed using confocal microscopy. Scale bar = 10 µm. The quantification of green fluorescence intensity in 10 dried particles of Alexa 488 (b) and EGFP (d) were plotted. In box plots, the centre line shows the median, box edges delineate first and third quartiles, and whiskers show the range of values.
As EGFP fluorescence requires a folded conformation, we evaluated the preservation of structure in the dried state (Fig. 6d). After drying and subsequent rehydrating, the EGFP solution loses its fluorescence, whereas the WKY + EGFP sample shows retention of fluorescence emission after drying and storage in dried form for five days, followed by instantaneous re-dissolution (Fig. 6e,f). The control KYF + EGFP (fibrous assembly)23 reveals a decrease in the fluorescence (Fig. 6e,f and Supplementary Fig. 42c), suggesting a level of stabilization through protein–peptide interactions.
These results highlight that the drying peptide ensembles provide an environment where proteins are stabilized, including in the dried state, possibly through mechanisms analogous to dynamic interfacial complexation via the self-organization of side chains with a protein surface45 (Fig. 6a). Using either a desiccator or lyophilizer, we found comparable EGFP fluorescence recovery, indicating that further dehydration does not impair dry-state protein preservation (Supplementary Fig. 43). Under degassed conditions, we observed that the encapsulation of EGFP inside the particles was comparable to that at ambient conditions and does not depend on the microporosity (Supplementary Fig. 44).
The encapsulation mechanism in this system is distinct from conventional partitioning into preformed condensates. Here, small molecules or proteins first interact with dynamic peptide ensembles in solution through non-covalent side-chain interactions—particularly electrostatic and π-related contacts involving the lysine, tyrosine and tryptophan residues. These interactions result in complexation with the payload in solution, before phase separation resembling previously described dynamic surface-stabilization approaches32, enabling efficient engulfment of the payload during droplet formation.
A series of experiments support this mechanism: tryptophan fluorescence quenching confirmed the binding, before the liquid–liquid phase separation, of both positively and negatively charged aromatic dyes (crystal violet and Alexa 488; Extended Data Fig. 6a,b), and confocal microscopy demonstrated efficient encapsulation regardless of charge (Extended Data Fig. 6c–e). GFP variants with varied net charge47,48 revealed that a higher charge enhances encapsulation (Extended Data Fig. 8a–d), reinforcing the role of electrostatics and π-interactions. Furthermore, simultaneous encapsulation of multiple fluorescent proteins of differing charge and colour49 preserved the fluorescence over time, highlighting the system versatility (Extended Data Fig. 8e,f). Finally, functional lysozyme encapsulation and the retention of active enzyme after 15 days in the dried state confirmed both the stability and bioactivity preservation (Fig. 6g and Supplementary Fig. 45). The drying-induced assembly process was evaluated for potential scalability by heating, vacuum (Extended Data Fig. 9a) or producing small droplets for rapid drying, resulting in comparable particle formation and EGFP encapsulation within these sprayed micro-droplets (Extended Data Fig. 9b–d and Supplementary Video 14). These data suggest that K/Y/W isomer tripeptides offer potential for encapsulation, drying, storage and the redispersion of biomolecules.
Extended Data Fig. 8. Proteins encapsulation.
Encapsulation of GFP with varied net charge. a. Engineered GFP proteins with net charge of −30, −18, 0, and +18 were pre-mixed with WKY solution individually followed by evaporation assay. b. The green fluorescence intensity of around 150 dried particles were profiled using confocal microscopy. c. The fluorescence intensity of the starting WKY + GFP mixture solution, which accounts for the variation in intrinsic fluorescence among this array of GFP proteins, were also measured and were used to normalize the raw green fluorescence intensity measured in b; d. The adjusted fluorescence intensity indicates greater encapsulation capacity with GFPs of higher charge, either positive or negative; e. Simultaneous encapsulation by tripeptide. Three fluorescent proteins of different colors and charges: miRFP682 (-6) (magenta), mScarlet-l (+5) (red) and GFP (0) (green) were pre-mixed with WKY solution together followed by evaporation assay; f. The fluorescence intensity of 30 dried particles were profiled using confocal microscopy with excitation at 633 nm, 561 nm, 488 nm laser respectively on both day 0 and day 6 under ambient conditions. Data represent mean ± s.d.
Extended Data Fig. 9. Scale up assays.
a. The volume of 20 µl WKY peptide solution was used and three different approaches were tested for scale up assays. While heating up the solution or vacuum condition could accelerate the drying process noticeably, the spray assay achieves the highest efficiency; b. Scheme of HXT M5 sprayer setup to spray large volume (20 µl) of WKY peptide solution into small drops onto microscope slides. Scale bar = 1 mm; c. Tested 80 µl/min spray rate achieved the best retention of micro-drop morphology, resulting in variable droplet sizes from tens to hundreds of microns. Scale bar = 50 µm; d. Further inspection by confocal microscopy demonstrated comparable particle formation with smaller size within sprayed micro-droplets, and (e) revealed efficient encapsulation of EGFP in this system. Scale bar = 10 µm.
Conclusions
We demonstrate the unique phase behaviour of tripeptides that form non-directional soluble dispersions covering a rich interaction space, leading to context-adaptive assembly. Through air-drying, these K/Y/W tripeptides lead to the spontaneous formation of high-porosity solids upon water evaporation that are instantaneously redispersed. The observed assembly and phase behaviour is governed by side-chain interactions, in contrast to the rigid-backbone hydrogen-bond topologies that typically dictate directional architectures in self-assembling molecules. We demonstrate that the six sequence isomers of K/W/Y form soluble ensembles with the sequence dictating the preferred interactions, but upon drying-induced assembly, all sequences lead to particle films with comparable emission in the dried state, suggesting that regardless of sequence, tryptophan has comparable stabilizing interactions. The phase separation process enables the encapsulation of small-molecule and protein payloads, and they are rapidly and fully reversible upon the re-introduction of water. We propose that this peptide-based evaporation-driven emulsification and encapsulation approach, without mechanical energy input, may find applications in the storage, formulation and desiccation protection of both biomolecules and biomacromolecules. Tryptophan’s rich interaction potential makes it a good candidate for adaptive and interfacial assembly5,8, building on its roles in proteins that are enabled by its unique heteroaromatic nature, such as its specific interfacial functions, for example, near lipid bilayers and ion channels50,51. We note a remarkable similarity with the protective mechanism of the natural adaptations in organisms such as tardigrades and plant seeds, which rely on liquid–liquid phase separation and vitrification of disordered proteins to preserve biomolecular function45,52.
Methods
All reagents and solvents were purchased from commercial suppliers and used without further purification. Peptides were either synthesized in‑house or purchased from GenScript and used after trifluoroacetic acid removal.
MD simulations
All-atom tripeptide (PDB) structures (KFF, KYF, KYY, KYW, KWY, WKY, WYK, YKW and YWK) were generated using Avogadro (ref. 53) or PyMOL v.2.5.0. These were then processed using the GROMACS tool gmx pdb2gmx to generate a topology file in the OPLS-AA/L all-atom force field54 (2001 amino acid dihedrals), and the TIP3P (ref. 55) explicit water model was chosen to simulate the solvent. The ionizable peptide side chains and termini were set to their respective charged states (NH3+ for Lys and N termini, and COO− for C termini), and 60 molecules of these tripeptides were inserted in an 8 × 8 × 8 nm3 box using a random seed with the tool gmx insert-molecules, giving an effective peptide concentration of ~200 mM. This box was then solvated using the tool gmx solvate by using a pre-equilibrated solvent box (SPC216) as a reference. Sixty solvent molecules were then randomly replaced by 60 Cl− ions to neutralize the net positive charge from the cationic tripeptides using the tool gmx genion. This neutralized box was subjected to steepest-descent energy minimization to remove packing conflicts. Short (1 ns) NVT (constant number of particles, volume and temperature) (temperature T = 298 K) and NPT (constant number of particles, pressure and temperature) (T = 298 K and pressure P = 1 bar) equilibrations were carried out while imposing position restraints on the peptides to allow the water to equilibrate. Self-assembly production MD simulations in the NPT ensemble were then carried out for 1,000 ns in triplicate for all sequences with no restraints on positions with the GROMACS v.2022.x package56 using the leap-frog integrator (time step = 2 fs). LINCS (ref. 57) was used to constrain bonds, and the particle mesh Ewald method was used for long-range electrostatics with a Coulomb and van der Waals cut-off of 1.0 nm each. Temperature coupling was achieved using the modified Berendsen thermostat58, whereas isotropic pressure coupling was maintained using the Parrinello–Rahman barostat59. The trajectory was saved every 100.0 ps or 0.1 ns, giving 10,000 frames per 1,000 ns simulation. The last 50 ns or 500 frames were used for statistical analysis of hydrogen bonding, SASA and AP.
The number of hydrogen bonds were measured in peptide–peptide interactions (total) as well as backbone–backbone, backbone/side-chain and side-chain/side-chain interactions using the tool gmx hbond with the donor–acceptor distance cut-off set to 0.35 nm and the hydrogen–donor–acceptor angle cut-off set to 30°. SASA was measured for the total peptides and the tryptophan side chain specifically using the tool gmx sasa with a solvent probe radius of 0.14 nm. Running AP was computed by a custom Python script dividing the initial SASA by the SASA at the current frame: AP = SASAinitial/SASAtime. Visual Molecular Dynamics (VMD; ref. 60) was used to visualize the trajectories and render the water surface around the tryptophan side chain within the 3 Å first hydration shell.
Peptide solutions and aggregates
Peptide solutions (and aggregates) were prepared by dissolving peptides in 100 mM sodium PB at pH 8 to a final peptide concentration of 20 mM. The pH was adjusted to 7.5 using 0.5 M HCl solution. The stock solution was heated for 5 min at 70 °C and left to cool for 1 h at room temperature (22–25 °C) to remove kinetically trapped aggregates. For lower concentrations (from 0.5 mM to 10 mM), the stock solutions were first heated and then left to cool to room temperature and the required amount collected. The diluted solutions were then heated for 5 min at 70 °C and aged for 1 h at room temperature.
Peptide solutions for NMR
Peptide solutions (and aggregates) were prepared by dissolving peptides in 100 mM sodium PB at pH 8 to final peptide concentrations of 1 mM, 20 mM and 50 mM in a 90:10 (v/v) H2O/D2O ratio. The pH was adjusted to 7.5 using 0.5 M HCl solution. The stock solution was heated for 5 min at 70 °C and left to cool for 1 h at room temperature (22–25 °C) to remove kinetically trapped aggregates. For lower concentrations (from 0.5 mM to 10 mM), the stock solutions were first heated and then left to cool to room temperature and the required amount collected. The diluted solutions were then heated for 5 min at 70 °C and aged for 1 h at room temperature.
NMR
The 1H and 19F NMR experiments were performed on a Bruker AVANCE HD III 700.20 MHz spectrometer and 282.2 MHz in the frequencies of 1H and 19F, respectively. The NMR experiments were performed at 298.2 K on a 5 mm QCI-F cryoprobe and on a multinuclear Broadband Fluorine Observe (BBFO) probe with z gradients. Typical 1D 19F NMR acquisition parameters were as follows: 8.0 μs pulse length (60° flip angle), 100 ppm sweep width (65,789 Hz) centred at −80 ppm, 0.50 s acquisition time, 2 s relaxation delay time and 256 scans (11 min). The 1D 19F NMR spectra were processed with 5 Hz exponential line broadening and visualized using TopSpin v.3.6.4. The 1D 1H NMR spectra were acquired using excitation sculpting (zgesgp) for water suppression, 32,768 points (time domain) and a recycling delay (relaxation delay (d1)) of 1.5 s. The 2D NMR experiments (1H–1H correlation spectroscopy (COSY), 1H–1H NOESY, 13C/1H heteronuclear single quantum coherence (HSQC) and 13C/1H heteronuclear multiple bond correlation (HMBC)) were acquired at 298.2 K using the same spectrometer and cryoprobe. The number of scans was set according to sample concentration. For the 13C/1H HSQC experiments (the hsqcetgpprsisp2.2 pulse program), the 1JCH was adjusted to 145 Hz, and the 13C/1H HMBC experiments (the hmbcgplpndprqf pulse program) were performed using an nJCH of 6 Hz; both pulse sequences used presaturation for water suppression. The 1H diffusion (DOSY) experiments were performed using the stimulated echo pulse sequence, stebpgp1s19. The following parameters were used for all peptides and at all concentrations: diffusion time (Δ) was set to 80 ms, the little delta (δ) was set to 2.4 ms, a linear ramp of 16 points varying from 1.019 to 47.906 G cm−1 was used and a d1 of 1.5 s was used. The water diffusion coefficient at the different peptide concentrations was measured using the stimulated echo pulse sequence without water suppression, stebpgp1s. The diffusion time (Δ) was set to 20 ms, the little delta (δ) was set to 2.0 ms, a linear ramp of 16 points varying from 1.019 to 42.809 G cm−1, and a d1 of 2 s was used.
Evaporation-driven assembly in sessile droplets
Evaporation-driven assembly was performed by depositing 5 µl droplets of peptide solution (prepared as per previous section) on a glass no. 1.5 coverslip and leaving it to evaporate at room temperature (22–25 °C) at around 40–50% humidity. For the imaging of the final products of peptides after evaporation, the 5 µl droplets were left to dry at the ambient conditions indicated above for about 30 min. The coverslips with dried peptide residue were subsequently used for analysis by confocal microscopy, AFM, FIB-SEM or TEM, with sample preparation and imaging occurring within 1 h. For live imaging, the droplets were first left to evaporate at ambient conditions for about 10 min and then moved to the confocal stage for time-lapse imaging under laser excitation. We note that while this approach may elevate the temperature of samples by a few degrees, we observe similar behaviour in the absence of laser irradiation.
The 3D spectroscopy
Fluorescence emission and excitation spectra were measured on a JASCO FP-8500 spectrofluorometer with emitted light measured orthogonally to the excitation light sourced from a xenon lamp in quartz cells with a path length of 1.5 mm. The spectra were measured on 50 µl peptide solutions, using a scanning speed of 200 nm min−1 with an emission bandwidth of 5 nm, a 1 nm data collection interval, a 1 s response time and medium sensitivity. The 3D spectra were recorded at λex = 230 to 450 nm and λem = 230 to 600 nm.
Fluorescence spectroscopy with EGFP
Emission spectra were recorded on Jasco FP-6500 fluorometer, using a 1.5 mm black quartz cuvette. The spectra were measured with a bandwidth of 2.5 nm and response time of 0.1 s, with collection at 500 nm min−1 and 0.1 nm intervals. Some 15 µl of each sample was placed on the cuvette and excited at 488 nm. The emission spectra were recorded from 490 to 580 nm. The dry samples were resolubilized with 15 µl distilled water to evaluate their fluorescence in solution. The experiment was replicated with three individual samples.
Degassing
Degassing was carried out by placing the peptide sample in a small Eppendorf vial and placing this in a septum vial. The sample was then placed on ice and put under vacuum for 5–10 min while being sonicated and subsequently refilled with argon. This was repeated five times. Alternative degassing by a 20 min gentle sonication in a Bioruptor (30 s on and 30 s off, 20 cycles) at 4 °C achieved a similar degassing effect.
Bright-field macroscopy imaging
Some 5 µl droplets of peptides were deposited onto a glass coverslip and allowed to dry at room temperature. To assess the drying time and patterns at the macroscopic level, videos coupled with microscopy were taken of the drying process. Bright-field videos of KFF, KYF, KYY, KYW and PBS were taken using a Zeiss Stemi 580 dissecting microscope outfitted with a smartphone adaptor (Celestron NexYZ three-axis universal smartphone adaptor for a telescope, purchased from Amazon.com) at ×2.5 physical zoom and ×2 digital zoom to allow for recording with a smartphone (iPhone 13, 12MP camera system). Side-view video of KYW was taken by a wireless digital microscope at ×75 zoom (STPCTOU wireless digital microscope, 2MP camera, 1080P, purchased from Amazon.com) connected to a Dell Precision Tower 5810 computer, using the microscope’s companion recording software, Inskam. Post-processing of the videos to create time-lapses was performed in Adobe Premier Pro.
Confocal laser scanning microscopy
Confocal imaging was performed using a Zeiss LSM880 Airyscan instrument. Live images were taken, focusing at the bottom left edge of the 5 µl drying peptide droplet sample at ×10 or ×20 magnification. Time series were taken with a scan speed of 7 or 8, a line average of 2 and a frame size of 1,024 × 1,024 or 512 × 512 pixels under the objective EC Plan Neofluor ×10 (0.3 numerical aperture) or Plan Apo ×20 (0.8 numerical aperture). Time-lapse videos were captured continuously without interval and exported as .avi files at 10–25 frames per second. Processing of the videos to create time-lapses was performed in Adobe Premier Pro.
For the confocal imaging of dried peptide samples, 5 µl of each peptide sample was deposited onto a no. 1.5 coverslip and allowed to dry at the ambient conditions indicated above or on a heated plate at a designated temperature, and the complete dried samples were subjected to confocal imaging. Transmitted light coupled with a 561 nm laser at 0.2% laser power and a T-PMT detector module was used to capture the bright-field imaging under a Plan Apo ×20, 0.8 numerical aperture objective. A 405 nm laser at 1.0% laser power was used to detect the weak but traceable blue fluorescence (414–472 nm) of the peptide aggregates, and the blue fluorescence signal was used for Imaris 3D rendering.
Encapsulation of dyes and fluorescent proteins
For the encapsulation of the Alexa 488 experiment, 1 mM aqueous stock solution of Alexa 488 dye (Thermo Fisher) was added such that the final concentration of the dye was 10 µM. For the encapsulation of the crystal violet experiment, 1 mM aqueous stock solution of crystal violet dye (Thermo Scientific) was mixed with 20 mM WKY peptide solution to reach the final concentration of 10 µM. For the encapsulation experiment of native EGFP (charge –7), lyophilized recombinant EGFP was purchased from ProSpec (catalogue no. PRO-1606) and reconstituted into 100 µg ml−1 stock solution. The stock solution was mixed with 20 mM WKY or KYF or 10 mM PBS solution to a final concentration of 10 µg ml−1. For encapsulation of the varying charged GFP proteins, a 2 mg ml−1 aqueous stock solution of GFP with net charge of −30, −18, 0 or +18 was mixed with 20 mM WKY solution to a final concentration of 10 µg ml−1. For the simultaneous encapsulation of multiple proteins, 2 mg ml−1 aqueous stock solutions of miRFP682 (−6), mScarlet (+5) and GFP (0) were mixed with 20 mM WKY solution to a final concentration of 10 µg ml−1 each.
All the encapsulation assays were conducted using 5 µl of the mixture dropped onto a coverslip. For Alexa 488 and EGFP protein encapsulation imaging, a 488 nm laser at 0.5% and 0.3% laser power was used, respectively, to detect green fluorescence emission at 502–561 nm, together with transmitted light to detect the peptide aggregate structure. For crystal violet imaging, an excitation wavelength of 561 nm at 0.2% laser power was used to detect red fluorescence emission at 575–631 nm. For various charged GFP protein encapsulations, a 488 nm laser at 0.2% laser power and an emission of 502–561 nm were used. For the multicolour fluorescent protein encapsulation experiment, a 488 nm laser at 0.2% laser power was used to detect GFP (0) emission at 502–561 nm; a 561 nm laser at 0.2% laser power was used to detect red fluorescent protein mScarlet (+5) emission at 575–631 nm; and a 633 nm laser at 0.5% laser power was used to detect near infrared fluorescent protein miRFP682 (−6) emission at 638–677 nm. All the fluorescence imaging was captured together with transmitted light to detect the peptide aggregate structure.
Lysozyme encapsulation assay and preservation assay
For the lysozyme–Alexa 488 encapsulation assay, a 100 µg ml−1 aqueous stock solution of lysozyme–Alexa 488 was mixed with 20 mM WKY or PBS solution to a final concentration of 10 µg ml−1. For confocal fluorescence imaging, 5 µl of each mixture was dropped onto a coverslip to dry through evaporation at ambient conditions. The dried particles were examined using a confocal microscope with excitation at 488 nm by a laser and green fluorescence emission of 502–561 nm.
For the lysozyme preservation experiment, three 5 µl droplets of a mixture of 20 mM WKY plus 10 µg ml−1 lysozyme were dropped onto a coverslip for an evaporation assay at ambient conditions. The dried particles were rehydrated using 5 µl double-distilled H2O on day 0 and day 15. The rehydrated solution from 3 dried droplets were combined into a measurable volume of 15 µl as one replicate to proceed with concentration measurement using a Chicken Lysozyme ELISA Kit (NOVUS Biologicals, catalogue no. NBP2-60088) following manufacturer instructions. A microplate reader (SpectraMax 190) was used to measure absorbance at 450 nm, with the correction wavelength set at 540 nm. A four-parameter logistic standard curve was generated by plotting the mean absorbance for each standard on the x axis against the concentration on the y axis using GraphPad Prism. The data were linearized by plotting the log of the lysozyme concentrations versus the log of the optical density, and the best fit line was determined by regression analysis. The reading of optical density values of each sample was matched to this standard curve to obtain the corresponding lysozyme concentration. All experiments were carried out in three replicates.
Scale-up assays including heating, vacuum and spray
For scaling up the evaporation by heat, 20 µl of WKY peptide solution was dropped onto a coverslip, which was placed on a heated plate at 35 °C. For scaling up the evaporation by vacuum, a coverslip with 20 µl of 20 mM WKY peptide solution was placed in a desiccator connected to a vacuum. The start of the drying was seen by the forming of an opaque ring at the edge of the drop, and the completion of the drying was determined by the closing of the ring structure to form an opaque film. The duration of each process under each condition was recorded.
For scaling up the evaporation by spraying, 20 µl of 20 mM WKY peptide solution was injected into an HTX M5 sprayer (HTX Tech) through a Microfluidics Programmable Syringe Pump (NE-1002X). Different flow rates of 40, 60, 80 and 160 µl min−1 were tested. The same spray parameters were used for all tested flow rates: a velocity of 1,300 mm min−1, track spacing of 2 mm, N2 pressure of 10 psi and nozzle height of 40 mm. The sprayer was operated for 15 s to complete one spray cycle. Spraying at the flow rate of 80 µl min−1 achieved the best retention of the micro-drop morphology.
To analyse the encapsulation efficiency of the spray system, 20 µl of 20 mM WKY peptide solution with 10 µg ml−1 EGFP was injected into the HXT M5 and sprayed at 80 µl min−1. The enrichment of GFP fluorescence in the dried particles within each micro-drop was examined using confocal microscopy, as described above.
The z stack and Imaris rendering
Confocal z-stack images were obtained at ×20 magnification with ×4 zoom. The optical sections were taken at the optimal interval suggested by a Zeiss Zen Black 8.0 with 50% overlap. The thickness of the sets of z stacks ranged from 10 µm to 15 µm. The z-stack images were exported as .czi files and then converted to .ims files using ImarisConverter. Analysis was done on Imaris v.10.0.0 (Bitplane) and MatLab v.8.0 (Mathworks) on a Dell Precision 7920 Workstation with a 12 GB NVIDIA 4070Ti graphics card. The 3D reconstruction was performed on z-stack images using the ‘Spots’ and ‘Surfaces’ functions in Imaris with a surface detail of 0.4 µm and automatic thresholding using local contrast in transparency mode. Objects were statistically coded by diameter, with a range of 0.5 µm to 8 µm. MATLAB XTension was used to control camera angles in animated videos. Approximately 200–1,000 particles were analysed in each of the conditions. Dots were not marked due to large sample size (n > 10). Quantification of object diameter, volume or sphericity was generated in the Imaris statistics tab, and GraphPad Prism was used to generate box plots. The moving tracks of the liquid–liquid phase separation droplets were reconstructed using the ‘Object Tracking’ function of Imaris, and the sum of the tracks at the end of the movement was presented with the time being pseudo colour-coded.
Cryo-EM
TEM grids were purchased from Ted Pella (product no. 01895-F) with a lacey carbon support film. All grids were treated for 35 s in Fischione Nanoclean 1070 (70% power) with a mixture of argon (75%) and oxygen (25%). Cryo-EM grids were prepared in Vitrobot Mark IV (Thermo Fisher Scientific) at 21 °C with the following settings: relative humidity, 100%; wait time, 0 s; blot time, 3 s; and blot force, 4. A 2 µl of sample solution was gently pipetted onto a freshly glow-discharged lacey carbon grid and put through evaporation under ambient conditions for the indicated periods of time before being blotted with filter paper and plunged into liquid ethane that was precooled by liquid nitrogen. The cryo-EM grids were then transferred to and stored in liquid nitrogen. The cryo-EM grid was transferred in liquid nitrogen into a Gatan 626 cryo-specimen holder that was then inserted into the microscope stage. The specimen temperature was maintained at about −170 °C during data collection. Cryo-EM imaging was performed in low dose mode of a Titan Halo TEM instrument (Thermo Fisher Scientific) operating at 300 kV, and images were recorded on a CETA camera (Thermo Fisher Scientific).
FIB-SEM
The 5 µl peptide samples were first applied onto glass slides and air dried. They were then mounted onto SEM pin stubs using double-sided carbon tape. Gold was sputter-coated on the sample surface to a thickness of 10 nm using a Leica EM ACE600 coater for better electrical conductivity. These samples were then imaged in a Thermo Scientific (FEI) Helios NanoLab 660 FIB-SEM instrument with a high tension (voltage) of 5 kV and a current of 25 pA using an Everhart–Thornley detector and a through lens detector. A FIB was used with Ga+ ions at 30 kV and 0.79 nA to mill cross sections at a scale of tens of micrometres. The stage was tilted at 52° for FIB milling and cross-section imaging.
AFM
Height images and mechanical property mapping were simultaneously carried out in PeakForce QNM (quantitative nanoscale mechanical) tapping mode using a Bruker Multimode 8 AFM. The optical lever sensitivity of the AFM cantilevers (RTESPA-525, Bruker) was calibrated on a sapphire surface. The cantilever spring constant was calibrated by Bruker Thermal Tune software. Prior to measuring tripeptide samples, a flat polystyrene surface of 3.5 GPa Young’s modulus was used to confirm the Young’s modulus reading of the AFM set-up. The measurement from 5–10 solid porous particles or fibres of each peptide was plotted.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Online content
Any methods, additional references, Nature Portfolio reporting summaries, source data, extended data, supplementary information, acknowledgements, peer review information; details of author contributions and competing interests; and statements of data and code availability are available at 10.1038/s41563-025-02300-z.
Supplementary information
Supplementary figures.
Data sources for supplementary figures. The file contains the datasets for ‘Directionality and dispersibility in peptide self-assembly’, ‘Sequence dependence in tripeptide dispersions’ and ‘Evaporation-driven assembly’.
Macroscopic top view of tripeptides KFF, KYF, KYY and KYW drying. Top view of 5 µl of 20 mM KFF, KYF, KYY and KYW drying on a coverslip at ambient conditions, captured by a Zeiss Stemi 580 dissecting microscope outfitted with a smartphone adaptor. A 5 µl drop of PBS was used as a control. The video starts 15 min after the solution is dotted onto the slide. The video is played at ×100 speed.
Macroscopic side view of KYW drying. Side view of 5 µl of 20 mM KYW drying, captured by a STPCTOU wireless digital microscope. The video starts 5 min after the solution was dotted. The video is played at ×50 speed.
KYW drying. Time-lapse confocal imaging of 5 µl of 20 mM KYW drying on a coverslip, captured by a ×20 objective with ×2 digital zoom of a Zeiss LSM880 confocal microscope. Note the phase separation during the drying process. The video is played at ×15 speed.
KFF drying. Time-lapse confocal imaging of 5 µl of 20 mM KFF drying on a coverslip, captured by a ×20 objective of a Zeiss LSM880 confocal microscope. The video is played at ×30 speed.
KYF drying. Time-lapse confocal imaging of 5 µl of 20 mM KYF drying on a coverslip, captured by a ×20 objective of a Zeiss LSM880 confocal microscope. The video is played at ×40 speed.
KYY drying. Time-lapse confocal imaging of 5 µl of 20 mM KYY drying on a coverslip at ambient conditions, captured by a ×20 objective of a Zeiss LSM880 confocal microscope. The video is played at ×125 speed.
Imaris four-dimensional (4D) rendering of KYW drying, showing streaks. The moving tracks of the KYW droplets during the drying process were 4D rendered using Imaris based on the time-lapse confocal images to show the streak formation. The accumulated tracks are shown at the end of the video. Colour scale bar represents time. The video is played at ×14 speed.
Fusion and solidification of WKY droplets. Time-lapse confocal imaging of 20 µl of 20 mM WKY drying on a coverslip at ambient conditions, captured by a ×20 objective of a Zeiss LSM880 confocal microscope. The dynamic droplet fusion process and the transition from liquid droplets to solid particles are shown. The video is played at ×12.5 speed up to 95 s and at ×37.5 speed afterward.
WKY drying. Time-lapse confocal imaging of 5 µl of 20 mM WKY drying on a coverslip, captured by a ×20 objective of a Zeiss LSM880 confocal microscope. The video is played at ×60 speed.
Degassed WKY drying. Time-lapse confocal imaging of 5 µl of 20 mM degassed WKY drying on a coverslip, captured by a ×20 objective of a Zeiss LSM880 confocal microscope. Note the smaller size of the droplets compared with the non-degassed sample in Supplementary Video 9. The video is played at ×60 speed.
Reversibility of tripeptide KWY drying and rehydration. Time-lapse confocal imaging of 5 µl of 20 mM KWY drying, being rehydrated with 5 µl of H2O and re-drying on a coverslip, captured by a ×10 objective of a Zeiss LSM880 confocal microscope. The video is played at ×60 speed.
Encapsulation of Alexa 488 by KWY and reversibility. Time-lapse confocal imaging of 5 µl of 20 mM KWY mixed with 10 µM Alexa 488, drying, being rehydrated with 5 µl of H2O and re-drying on a coverslip, excited by a 488 nm laser and captured by a ×20 objective of a Zeiss LSM880 confocal microscope. Note the enrichment of the dye in the droplets. The video plays at ×25 speed, except for a few key moments (187–210 s and 518–538 s) that play at ×2 speed and the period between rehydration and the second evaporation, which plays at ×100 speed.
Encapsulation of EGFP by WKY. Time-lapse confocal imaging of 5 µl of 20 mM KWY mixed with 10 µg ml−1 EGFP drying on a coverslip at ambient conditions, excited by a 488 nm laser and captured by a ×20 objective of a Zeiss LSM880 confocal microscope. Note the enrichment of EGFP in the droplets. The video is played at ×50 speed.
Spray assay of tripeptide drying and EGFP encapsulation. The 20 µl mixture solution of 20 mM WKY with 10 µg ml−1 EGFP was sprayed as micro-droplets onto microscope slides by an HXT M5 sprayer at a 80 µl min−1 spray rate. The drying process and encapsulation were examined under a confocal microscope immediately after the spray. The video is played at ×4.8 speed.
Source data
Statistical data for Fig. 1d,f.
Statistical data for Fig. 2a,c–e.
Statistical data for Fig. 3g.
Statistical data for Fig. 4f.
Statistical data for Fig. 5b,d,f,g.
Statistical data for Fig. 6b,c,f,g.
Statistical data for Extended Data Fig. 1.
Statistical data for Extended Data Fig. 5d.
Statistical data for Extended Data Fig. 6a,b.
Statistical data for Extended Data Fig. 7b.
Statistical data for Extended Data Fig. 8b–d,f.
Acknowledgements
R.V.U., D.R.D. and M.C. acknowledge funding from the Office of Naval Research for the Vannevar Bush Faculty Fellowship (grant no. N00014-21-1-2967). We thank the Air Force Office of Scientific Research for funding R.V.U., D.S. and S.K. (grant no. FA9550-21-1-0091). We also acknowledge partial support from the National Science Foundation (NSF) grant no. HRD-2112550 (Phase II CREST Center IDEALS). The TFA removal protocol was provided by G. Scott. The imaging, NMR and mass spectrometry experiments were carried out in the Live Imaging and Bioenergetics Core, Surface Science Core, Imaging Suite, NMR Core and Mass Spectrometry Core at Advanced Science Research Center (ASRC). Y.H. and T.W. are supported by a Faculty Research Award jointly funded by the Professional Staff Congress and the City University of New York (PSC-CUNY). M.A.C. and S.D. were supported by the Velay Women Scholarship, and K.V. was supported by the Sloan Foundation CUNY Summer Undergraduate Research Program (CSURP). T.-D.L. acknowledges support through the National Institutes of Health (NIH), 1S10OD030401-01A1. S.Y.A. and A.C.O. acknowledge support from the National Institutes of Health (NIH) under award R35GM138378. We also thank H. Kuntrapakam for valuable feedback on our NMR interpretation.
Extended data
Author contributions
R.V.U., Y.H., D.R.D., S.K., M.C. and L.X. conceived the project and designed the experiments. Y.H. led the imaging and functional assays. R.V.U. led the supramolecular design and characterization and oversaw the project. Y.H., M.T.-N. and D.S. first observed the phase behaviour; Y.H. and M.T.-N. conducted preliminary experiments, including microscopy, fluorescence and drying studies. D.R.D. and S.D. performed molecular simulations with input from M.M.; S.K., M.C. and P.C.-D.L. synthesized and purified peptides and prepared samples. S.K. and M.C. conducted circular dichroism and liquid chromatography–mass spectrometry analyses; P.C.-D.L. performed fluorescence measurements. Y.H., L.X., M.C., M.P.-F. and P.C.-D.L. performed the encapsulation experiments. L.X., P.C.-D.L., M.C., T.Z. and M.P.-F. investigated the salt and concentration effects. D.P., under the guidance of X.C., conducted the humidity-responsive fluorescence measurements. M.C., S.K. and D.C.F. performed and interpreted the NMR analysis. L.X., K.V., M.A.C. and T.Z. carried out the macroscopy imaging, heating and reversibility assays, and Imaris analysis. L.X. performed the scale-up experiments, confocal imaging and functional assays. S.Z. conducted the SEM analysis; T.W. and Y.H. performed the cryo-TEM; T.-D.L. performed the AFM. L.O.M. and N.S.A., under R.T.’s supervision, measured surface tension. C.W. and J.M. performed the TEM. A.C.O. and S.Y.A. provided protein variants and contributed to experimental design and data interpretation. L.X. quantified all imaging data. S.K., D.R.D., M.C., L.X., R.V.U. and Y.H. contributed to the mechanistic understanding and cowrote the manuscript. All authors discussed the results.
Peer review
Peer review information
Nature Materials thanks Tao Jiang and Junbai Li for their contribution to the peer review of this work.
Data availability
The data that support the findings of this study are available within the paper and its Supplementary Information and are available from the corresponding authors upon request. Source data are provided with this paper.
Competing interests
R.V.U., Y.H., M.C., L.X., D.R.D., S.K. and M.T.-N. have submitted a patent application related to the phase separation, encapsulation and preservation technology described in this work. R.V.U. and M.C. are cofounders of BioWraptor, a company with a potential interest in licensing this technology.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
These authors contributed equally: Dhwanit R. Dave, Salma Kassem, Maeva Coste, Lele Xu, Mona Tayarani-Najjaran.
Change history
8/16/2025
In the version of the article initially published, the surname of Darjan Podbevšek appeared incorrectly, and is now amended in the HTML and PDF versions of the article.
Contributor Information
Ye He, Email: yhe1@gc.cuny.edu.
Rein V. Ulijn, Email: rulijn@gc.cuny.edu
Extended data
is available for this paper at 10.1038/s41563-025-02300-z.
Supplementary information
The online version contains supplementary material available at 10.1038/s41563-025-02300-z.
References
- 1.Aida, T., Meijer, E. W. & Stupp, S. I. Functional supramolecular polymers. Science335, 813–818 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Zhang, S. Fabrication of novel biomaterials through molecular self-assembly. Nat. Biotechnol.21, 1171–1178 (2003). [DOI] [PubMed] [Google Scholar]
- 3.Ghadiri, M. R., Granja, J. R., Milligan, R. A., McRee, D. E. & Khazanovich, N. Correction: self-assembling organic nanotubes based on a cyclic peptide architecture. Nature366, 324–327 (1993). [DOI] [PubMed] [Google Scholar]
- 4.Freeman, R. et al. Reversible self-assembly of superstructured networks. Science362, 808–813 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kaygisiz, K. et al. Context dependence in assembly code for supramolecular peptide materials and systems. Nat. Rev. Mater.10, 449–472 (2025). [Google Scholar]
- 6.Du, X., Zhou, J., Shi, J. & Xu, B. Supramolecular hydrogelators and hydrogels: from soft matter to molecular biomaterials. Chem. Rev.115, 13165–13307 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Bianco, S. et al. Mechanical release of homogenous proteins from supramolecular gels. Nature631, 544–548 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Berisha, N. et al. Directed discovery of high-loading nanoaggregates enabled by drug-matched oligo-peptide excipients. Chem11, 102404 (2025). [Google Scholar]
- 9.Levin, A. et al. Biomimetic peptide self-assembly for functional materials. Nat. Rev. Chem.4, 615–634 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Woolfson, D. N. A brief history of de novo protein design: minimal, rational, and computational. J. Mol. Biol.433, 167160 (2021). [DOI] [PubMed] [Google Scholar]
- 11.DeGrado, W. F. & Korendovych, I. V. De novo protein design, a retrospective. Q. Rev. Biophys.53, e3 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hulgan, S. A. H. & Hartgerink, J. D. Recent advances in collagen mimetic peptide structure and design. Biomacromolecules23, 1475–1489 (2022). [DOI] [PubMed] [Google Scholar]
- 13.Omenetto, F. G. & Kaplan, D. L. New opportunities for an ancient material. Science329, 528–531 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Yuan, C. et al. Hierarchically oriented organization in supramolecular peptide crystals. Nat. Rev. Chem.3, 567–588 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gallego, L., Woods, J. F. & Rickhaus, M. Recent concepts for supramolecular 2D materials. Org. Mater.4, 137–145 (2022). [Google Scholar]
- 16.Magnotti, E. L. et al. Self-assembly of an α‑helical peptide into a crystalline two- dimensional nanoporous framework. J. Am. Chem. Soc.138, 16274–16282 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Ganatra, P. et al. Diverse proteomimetic frameworks via rational design of π-stacking peptide tectons. J. Am. Chem. Soc.146, 22236–22246 (2024). [DOI] [PubMed] [Google Scholar]
- 18.Reches, M. & Gazit, E. Casting metal nanowires within discrete self-assembled peptide nanotubes. Science300, 625–627 (2003). [DOI] [PubMed] [Google Scholar]
- 19.Chakraborty, P. et al. Co‐assembly between Fmoc diphenylalanine and diphenylalanine within a 3D fibrous viscous network confers atypical curvature and branching. Angew. Chem.132, 23939–23947 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Azuri, I., Adler-Abramovich, L., Gazit, E., Hod, O. & Kronik, L. Why are diphenylalanine-based peptide nanostructures so rigid? Insights from first principles calculations. J. Am. Chem. Soc.136, 963–969 (2014). [DOI] [PubMed] [Google Scholar]
- 21.Bera, S. et al. Rigid helical-like assemblies from a self-aggregating tripeptide. Nat. Mater.18, 503–509 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Garcia, AnaM. et al. Chirality effects on peptide self-assembly unraveled from molecules to materials. Chem4, 1862–1876 (2018). [Google Scholar]
- 23.Frederix, P. W. J. M. et al. Exploring the sequence space for (tri-)peptide self-assembly to design and discover new hydrogels. Nat. Chem.7, 30–37 (2015). [DOI] [PubMed] [Google Scholar]
- 24.Perry, S. L. Phase separation: bridging polymer physics and biology. Curr. Opin. Colloid Interface Sci.39, 86–97 (2019). [Google Scholar]
- 25.Abbas, M., Lipiński, W. P., Wang, J. & Spruijt, E. Peptide-based coacervates as biomimetic protocells. Chem. Soc. Rev.50, 3690–3705 (2021). [DOI] [PubMed] [Google Scholar]
- 26.Leshem, A. B. et al. Biomolecular condensates formed by designer minimalistic peptides. Nat. Commun.14, 421 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Gabryelczyk, B. et al. Hydrogen bond guidance and aromatic stacking drive liquid-liquid phase separation of intrinsically disordered histidine-rich peptides. Nat. Commun.10, 5465 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Abbas, M., Lipiński, W. P., Nakashima, K. K., Huck, W. T. S. & Spruijt, E. A short peptide synthon for liquid–liquid phase separation. Nat. Chem.13, 1046–1054 (2021). [DOI] [PubMed] [Google Scholar]
- 29.Xing, R., Yuan, C., Fan, W., Ren, X. & Yan, X. Biomolecular glass with amino acid and peptide nanoarchitectonics. Sci. Adv.9, eadd8105 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Finkelstein-Zuta, G. et al. A self-healing multispectral transparent adhesive peptide glass. Nature630, 368–374 (2024). [DOI] [PubMed] [Google Scholar]
- 31.Panganiban, B. et al. Random heteropolymers preserve protein function in foreign environments. Science359, 1239–1243 (2018). [DOI] [PubMed] [Google Scholar]
- 32.Ruan, Z. et al. Population-based heteropolymer design to mimic protein mixtures. Nature615, 251–258 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lampel, A. et al. Polymeric peptide pigments with sequence-encoded properties. Science356, 1064–1068 (2017). [DOI] [PubMed] [Google Scholar]
- 34.Khemaissa, S., Sagan, S. & Walrant, A. Tryptophan, an amino-acid endowed with unique properties and its many roles in membrane proteins. Crystals11, 1032 (2021). [Google Scholar]
- 35.Scott, G. G., McKnight, P. J., Tuttle, T. & Ulijn, R. V. Tripeptide emulsifiers. Adv. Mater.28, 1381–1386 (2016). [DOI] [PubMed] [Google Scholar]
- 36.Scheiner, S., Kar, T. & Pattanayak, J. Comparison of various types of hydrogen bonds involving aromatic amino acids. J. Am. Chem. Soc.124, 13257–13264 (2002). [DOI] [PubMed] [Google Scholar]
- 37.Vivian, J. T. & Callis, P. R. Mechanisms of tryptophan fluorescence shifts in proteins. Biophys. J.80, 2093–2109 (2001). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kwok, A. et al. A thermodynamic model for interpreting tryptophan excitation-energy-dependent fluorescence spectra provides insight into protein conformational sampling and stability. Front. Mol. Biosci.8, 778244 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Liu, H., Zhang, H. & Jin, B. Fluorescence of tryptophan in aqueous solution. Spectrochim. Acta A Mol. Biomol. Spectrosc.106, 54–59 (2013). [DOI] [PubMed] [Google Scholar]
- 40.Stollar, E. J. et al. Unconventional interactions between water and heterocyclic nitrogens in protein structures. Proteins57, 1–8 (2004). [DOI] [PubMed] [Google Scholar]
- 41.Moon, B. U. et al. Evaporation-driven water-in-water droplet formation. Langmuir36, 14333–14341 (2020). [DOI] [PubMed] [Google Scholar]
- 42.Shaw, M., Bella, A. & Ryadnov, M. G. CREIM: coffee ring effect imaging model for monitoring protein self-assembly in situ. J. Phys. Chem. Lett.8, 4846–4851 (2017). [DOI] [PubMed] [Google Scholar]
- 43.Cohen-Gerassi, D. et al. Phase transition and crystallization kinetics of a supramolecular system in a microfluidic platform. Chem. Mater.32, 8342–8349 (2020). [Google Scholar]
- 44.Maldonado-Valderrama, J., Wege, H. A., Rodríguez-Valverde, M. A., Gálvez-Ruiz, M. J. & Cabrerizo-Vílchez, M. A. Comparative study of adsorbed and spread β-casein monolayers at the water–air interface with the pendant drop technique. Langmuir19, 8436–8442 (2003). [Google Scholar]
- 45.Olgenblum, G. I., Hutcheson, B. O., Pielak, G. J. & Harries, D. Protecting proteins from desiccation stress using molecular glasses and gels. Chem. Rev.124, 5668–5694 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Kumar, M. et al. In situ, noncovalent labeling and stimulated emission depletion-based super-resolution imaging of supramolecular peptide nanostructures. ACS Nano14, 15056–15063 (2020). [DOI] [PubMed] [Google Scholar]
- 47.Kapelner, R. A. & Obermeyer, A. C. Ionic polypeptide tags for protein phase separation. Chem. Sci.10, 2700–2707 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Cummings, C. S. & Obermeyer, A. C. Phase separation behavior of supercharged proteins and polyelectrolytes. Biochemistry57, 314–323 (2018). [DOI] [PubMed] [Google Scholar]
- 49.Ahn, S. Y. & Obermeyer, A. C. Selectivity of complex coacervation in multiprotein mixtures. JACS Au4, 3800–3812 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Khemaissa, S., Walrant, A. & Sagan, S. Tryptophan, more than just an interfacial amino acid in the membrane activity of cationic cell-penetrating and antimicrobial peptides. Q. Rev. Biophys.55, e10 (2022). [DOI] [PubMed]
- 51.Andersen, O. S., Greathouse, D. V., Providence, L. L., Becker, M. D. & Koeppe, R. E. Importance of tryptophan dipoles for protein function: F-fluorination of tryptophans in gramicidin a channels. J. Am. Chem. Soc.120, 5142–5146 (1998). [Google Scholar]
- 52.Lim, S. et al. Tardigrade secretory proteins protect biological structures from desiccation. Commun. Biol.7, 633 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Hanwell, M. D. et al. Avogadro: an advanced semantic chemical editor, visualization, and analysis platform. J. Cheminformatics4, 17 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Jorgensen, W. L., Maxwell, D. S. & Tirado-Rives, J. Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. J. Am. Chem. Soc.118, 11225–11236 (1996). [Google Scholar]
- 55.Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys.79, 926–935 (1998). [Google Scholar]
- 56.Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX1–2, 19–25 (2015). [Google Scholar]
- 57.Hess, B. P-LINCS: a parallel linear constraint solver for molecular simulation. J. Chem. Theory Comput.4, 116–122 (2007). [DOI] [PubMed] [Google Scholar]
- 58.Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys.126, 014101 (2007). [DOI] [PubMed] [Google Scholar]
- 59.Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys.52, 7182–7190 (1998). [Google Scholar]
- 60.Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph.14, 33–38 (1996). [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary figures.
Data sources for supplementary figures. The file contains the datasets for ‘Directionality and dispersibility in peptide self-assembly’, ‘Sequence dependence in tripeptide dispersions’ and ‘Evaporation-driven assembly’.
Macroscopic top view of tripeptides KFF, KYF, KYY and KYW drying. Top view of 5 µl of 20 mM KFF, KYF, KYY and KYW drying on a coverslip at ambient conditions, captured by a Zeiss Stemi 580 dissecting microscope outfitted with a smartphone adaptor. A 5 µl drop of PBS was used as a control. The video starts 15 min after the solution is dotted onto the slide. The video is played at ×100 speed.
Macroscopic side view of KYW drying. Side view of 5 µl of 20 mM KYW drying, captured by a STPCTOU wireless digital microscope. The video starts 5 min after the solution was dotted. The video is played at ×50 speed.
KYW drying. Time-lapse confocal imaging of 5 µl of 20 mM KYW drying on a coverslip, captured by a ×20 objective with ×2 digital zoom of a Zeiss LSM880 confocal microscope. Note the phase separation during the drying process. The video is played at ×15 speed.
KFF drying. Time-lapse confocal imaging of 5 µl of 20 mM KFF drying on a coverslip, captured by a ×20 objective of a Zeiss LSM880 confocal microscope. The video is played at ×30 speed.
KYF drying. Time-lapse confocal imaging of 5 µl of 20 mM KYF drying on a coverslip, captured by a ×20 objective of a Zeiss LSM880 confocal microscope. The video is played at ×40 speed.
KYY drying. Time-lapse confocal imaging of 5 µl of 20 mM KYY drying on a coverslip at ambient conditions, captured by a ×20 objective of a Zeiss LSM880 confocal microscope. The video is played at ×125 speed.
Imaris four-dimensional (4D) rendering of KYW drying, showing streaks. The moving tracks of the KYW droplets during the drying process were 4D rendered using Imaris based on the time-lapse confocal images to show the streak formation. The accumulated tracks are shown at the end of the video. Colour scale bar represents time. The video is played at ×14 speed.
Fusion and solidification of WKY droplets. Time-lapse confocal imaging of 20 µl of 20 mM WKY drying on a coverslip at ambient conditions, captured by a ×20 objective of a Zeiss LSM880 confocal microscope. The dynamic droplet fusion process and the transition from liquid droplets to solid particles are shown. The video is played at ×12.5 speed up to 95 s and at ×37.5 speed afterward.
WKY drying. Time-lapse confocal imaging of 5 µl of 20 mM WKY drying on a coverslip, captured by a ×20 objective of a Zeiss LSM880 confocal microscope. The video is played at ×60 speed.
Degassed WKY drying. Time-lapse confocal imaging of 5 µl of 20 mM degassed WKY drying on a coverslip, captured by a ×20 objective of a Zeiss LSM880 confocal microscope. Note the smaller size of the droplets compared with the non-degassed sample in Supplementary Video 9. The video is played at ×60 speed.
Reversibility of tripeptide KWY drying and rehydration. Time-lapse confocal imaging of 5 µl of 20 mM KWY drying, being rehydrated with 5 µl of H2O and re-drying on a coverslip, captured by a ×10 objective of a Zeiss LSM880 confocal microscope. The video is played at ×60 speed.
Encapsulation of Alexa 488 by KWY and reversibility. Time-lapse confocal imaging of 5 µl of 20 mM KWY mixed with 10 µM Alexa 488, drying, being rehydrated with 5 µl of H2O and re-drying on a coverslip, excited by a 488 nm laser and captured by a ×20 objective of a Zeiss LSM880 confocal microscope. Note the enrichment of the dye in the droplets. The video plays at ×25 speed, except for a few key moments (187–210 s and 518–538 s) that play at ×2 speed and the period between rehydration and the second evaporation, which plays at ×100 speed.
Encapsulation of EGFP by WKY. Time-lapse confocal imaging of 5 µl of 20 mM KWY mixed with 10 µg ml−1 EGFP drying on a coverslip at ambient conditions, excited by a 488 nm laser and captured by a ×20 objective of a Zeiss LSM880 confocal microscope. Note the enrichment of EGFP in the droplets. The video is played at ×50 speed.
Spray assay of tripeptide drying and EGFP encapsulation. The 20 µl mixture solution of 20 mM WKY with 10 µg ml−1 EGFP was sprayed as micro-droplets onto microscope slides by an HXT M5 sprayer at a 80 µl min−1 spray rate. The drying process and encapsulation were examined under a confocal microscope immediately after the spray. The video is played at ×4.8 speed.
Statistical data for Fig. 1d,f.
Statistical data for Fig. 2a,c–e.
Statistical data for Fig. 3g.
Statistical data for Fig. 4f.
Statistical data for Fig. 5b,d,f,g.
Statistical data for Fig. 6b,c,f,g.
Statistical data for Extended Data Fig. 1.
Statistical data for Extended Data Fig. 5d.
Statistical data for Extended Data Fig. 6a,b.
Statistical data for Extended Data Fig. 7b.
Statistical data for Extended Data Fig. 8b–d,f.
Data Availability Statement
The data that support the findings of this study are available within the paper and its Supplementary Information and are available from the corresponding authors upon request. Source data are provided with this paper.