Abstract
Sixteen open-top chambers (diameter, 3.66 m) were established in a scrub oak habitat in central Florida where vegetation was removed in a planned burn prior to chamber installation. Eight control chambers have been continuously exposed to ambient air and eight have been continuously exposed to elevated CO2 at twice-ambient concentration (∼700 ppm) for 5 years. Soil cores were collected from each chamber to examine the influence of elevated atmospheric CO2 on the fungal community in different soil fractions. Each soil sample was physically fractionated into bulk soil, rhizosphere soil, and roots for separate analyses. Changes in relative fungal biomass were estimated by the ergosterol technique. In the bulk soil and root fractions, a significantly increased level of ergosterol was detected in the elevated CO2 treatments relative to ambient controls. Fungal community composition was determined by terminal-restriction fragment length polymorphism (T-RFLP) analysis of the internal transcribed spacer (ITS) region. The specificities of different ITS primer sets were evaluated against plant and fungal species isolated from the experimental site. Changes in community composition were assessed by principal component analyses of T-RFLP profiles resolved by capillary electrophoresis. Fungal species richness, defined by the total number of terminal restriction fragments, was not significantly affected by either CO2 treatment or soil fraction.
During the last decade a number of studies have been conducted to assess the influence of elevated atmospheric CO2 on aboveground plant community diversity and physiology as well as on belowground root dynamics, root exudates (6, 8, 22, 56, 60, 66), and nutrient availability (27, 29). In most cases these investigations document an impact of elevated CO2 upon plant growth and water requirements, but the reaction of the belowground microbial community has been less clear-cut. Some studies have reported no effect of elevated CO2 (45, 67), while others have reported increases in microbial activity (50), in microbial biomass (9, 49), or in the number of mycorrhizal infections (45). Few studies have attempted to directly assess changes in the composition and richness of the microbial community as a whole in response to elevated CO2. Among these studies, few have documented changes in either the richness or evenness of microbial community composition. For example, Griffith et al. (23) used community DNA hybridization and percent G+C base profiling to detect changes in rhizosphere microbial communities of wheat. They found only insignificant differences between treatments. Zak et al. (67), using phospholipid fatty acid analysis, did not find any significant changes in microbial community composition caused by elevated CO2 in soil beneath the bigtooth aspen (Populus grandidentata). Both Griffith et al. (23) and Zak et al. (67) used relatively broad-scale profiling techniques that are only able to detect changes at the genus level or higher. Montealegre et al. (40) on the other hand, using PCR-based fingerprinting of genomic DNA, reported that Rhizobium leguminosarum isolated from white clover grown at elevated CO2 concentrations were genetically different from isolates obtained from plants grown at ambient CO2 concentrations. Taken in aggregate, these results suggest that microbial responses to elevated CO2 concentrations may be multifactorial, difficult to measure, and easily confounded by other factors. Measures of microbial community composition may have been influenced by the experimental system used to elevate CO2 (45), the specific types of plants examined (51), the sensitivity of the microbial community profiling method (23, 67), or the diversity of microbial types present within the community (40). Molecular methods are not immune to sampling bias and technology limitations that may obscure the subtle signal of microbial compositional shifts amidst the noise of community heterogeneity. It has therefore been problematic to clearly separate CO2 treatment effects from artifacts of either experimental design or sampling.
Terminal-restriction fragment length polymorphism (T-RFLP) analysis is a direct DNA profiling method that has been used extensively to assess microbial community structure in habitats whose community compositions are both complex and diverse (11, 12, 35, 38, 46). The method has been targeted to multiple scales of phylogenetic resolution from species to domain level and is applicable to bacteria as well as archaea and eukarya (for a review, see reference 31). The primary molecular target of T-RFLP community profiling has been small-subunit rRNA (i.e., 16S and 18S ribosomal DNA [rDNA] genes), although functional gene applications have been developed for the mer (4), amoA (26), nosZ (52), and nirS (3) genes among bacteria. Multiple PCR primers that target the fungal rDNA subunits and intergenic spacer regions of diverse taxonomic groups have been reported for the strain typing of fungal isolates (58, 65). Only recently, however, have primers targeting small- and large-subunit rDNA or ribosomal intergenic spacer regions been developed for the specific amplification of fungal rDNA sequences directly from environmental and clinical samples (2, 28, 39, 57, 59). Here we describe the application of T-RFLP fungal community profiling of direct environmental extracts with primers targeting the internal transcribed spacer (ITS) regions of ascomycetes and basidiomycetes, or of basidiomycetes only, in order to detect changes in fungal community composition in response to elevated atmospheric CO2.
A second aim was to examine the effect of elevated CO2 on the quantity and partitioning of fungal biomass belowground as determined from the ergosterol content in the soil and root fractions, respectively. Ergosterol has been extensively used to quantify fungal biomass in soil (10, 21, 53, 64) and mycorrhizal systems (14, 44). As a specific measure of fungal biomass, ergosterol has several advantages. It is relatively easy to extract (13, 47, 55) and is in many cases the dominant fungal sterol (34, 61). Ergosterol is not exclusively found in fungi, but several workers have shown that it is a more sensitive and reliable indicator of viable fungal biomass than other biochemical molecules (7, 36, 42, 64).
There are, however, methodological limitations to this measure. Foremost among these is the large variation in ergosterol content among different fungal species (34, 41, 42) and the presence of ergosterol in a few nonfungal eukaryotes. Some green algae, for example, have been shown to produce ergosterol in small amounts, complicating the interpretation of this measure in aquatic environments (7, 43). In addition, changes in ergosterol content may primarily reflect changes in species composition rather than changes in total biomass in some environments (1, 47)—a problem exacerbated by the paucity of fungal species and environments examined so far (see, e.g., reference 34, 62, and 63). Although these problems limit interpretation of ergosterol as a quantitative measure of total fungal biomass across environments, the ergosterol assay remains the single most reliable relative indicator of fungal biomass in natural biological systems (7, 15, 21, 42, 55, 64).
MATERIALS AND METHODS
Study site.
An experiment to monitor the long-term effects of elevated atmospheric CO2 in a nutrient-limited coastal scrub oak forest was initiated and funded by the U.S. Department of Energy in 1995 at the John F. Kennedy Space Center in Florida as part of a collaborative research project between the National Aeronautics and Space Administration and the Smithsonian Institution (27). The site was cleared of scrub vegetation in a controlled burn in 1995 prior to installation of 16 open-top chambers, of which 8 were supplied with ambient air and 8 were supplied with ambient air plus 350 μl of CO2 liter of air−1 (29).
Sampling.
A single soil core (diameter, 5 cm; mean depth, 10 cm) was taken from each of the chambers and separated into three fractions: bulk soil, rhizosphere soil, and roots. The soil fractions were obtained by passing freshly collected sample through a 2-mm-pore-size sieve. The sieved soil was named the bulk soil. The soil attached to roots (rhizosphere) was washed off with sterile deionized water and collected in a sterile bucket. The resulting soil slurry was centrifuged at 500 × g for 20 min to separate the water from the rhizosphere soil.
From each of the three subsamples (bulk soil, rhizosphere soil, and roots), approximately 5 g (wet weight) was used to determine dry weight after drying at 80°C for 24 h and ash content (ash generated at 600°C for 6 h). The soil moisture content was 6.5% ± 2.7% (wt/wt) (mean ± standard deviation) and the soil pH was in the range of 4.9 to 5.5, with no difference between CO2 treatments.
Extractions.
The bulk and rhizosphere soils were divided into subsamples for ergosterol and DNA analyses, respectively. The scarce amount of roots in the samples yielded only sufficient material for dry weight and ergosterol determinations. Bulk and rhizosphere soil samples for DNA extraction were stored at −80°C until analysis, while ergosterol was extracted immediately using the procedure of Pennanen et al. (48).
DNA was extracted from 0.25-g soil fractions using the UltraClean soil DNA isolation kit (Mo Bio Laboratories, Solana Beach, Calif.). The manufacturer's protocol was followed, with the modifications suggested by Clement and Kitts (5). The yield was determined using the Pico-green assay (Molecular Probes Inc., Portland, Oreg.) according to the manufacturer's protocol in a Dynex MFX microplate fluorometer with serial dilutions of each extract. Bulk soil yielded 1.8 to 12.8 ng of DNA/μl, and rhizosphere soil yielded 3.3 to 45.9 ng of DNA/μl.
PCR was performed in 50-μl reaction volumes using fluorescently labeled forward (6-FAM) and reverse (HEX) oligonucleotide primers (synthesized by Operon Technologies, Alameda, Calif.) targeting the ITS region of the rDNA operon. Different primer sets were designed to be specific for different groups of fungi as follows: ITS 1F-ITS 4, specific for higher fungi; ITS 1F-ITS 4B, specific for basidiomycetes (18); and ITS 1F-ITS 4A, specific for ascomycetes (33). Finally we tested the combination ITS 1-ITS 4B at increased annealing temperature (60°C) in order to increase specificity for basidiomycetes. Final concentrations in the PCRs were: 2.0 mM MgCl2, 1× buffer (Applied Biosystems [Foster City, Calif.] buffer II), a 200 μM concentration of each deoxynucleoside triphosphate, a 1.0 μM concentration of each primer, 0.4 μg of bovine serum albumin/μl, and 1.25 U of AmpliTaq DNA polymerase (Applied Biosystems). The thermocycler reaction conditions were as follows: 5-min initial denaturation at 94°C followed by 35 cycles of 0.5 min at 94°C, 2 min of annealing using different temperatures for the different primer sets (as indicated in Table 1), and a 3-min extension at 72°C. The final extension was 5 min at 72°C. Reaction yield and target specificity were determined by agarose gel electrophoresis. The PCR products were cleaned using a QIAquick PCR purification kit (Qiagen, Hilden, Germany), digested with different restriction endonucleases (HinfI, TaqI, and MseI at 5 U per reaction according to manufacturer's instructions), cleaned with a QIAquick nucleotide removal kit (Qiagen), denatured at 95°C for 10 min, and separated by capillary electrophoresis on an ABI 310 genetic analyzer in GeneScan mode (Applied Biosystems).
TABLE 1.
Specificities of primer sets used in this study
| Species | Specificity of primer pair at indicated annealing tempa
|
|||||
|---|---|---|---|---|---|---|
| Euk F-Euk R, 58°C | ITS 1-ITS 4, 55°C | ITS 1F-ITS 4, 50°C | ITS 1F-ITS 4A, 62°C | ITS 1F-ITS 4B, 55°C | ITS 1-ITS 4B, 60°C | |
| Plants | ||||||
| Quercus myrtifolia Willd. | ++ | ++ | − | − | + | − |
| Q. geminata Small | ++ | ++ | − | − | + | − |
| Q. chapmanii Sarg. | ++ | ++ | − | − | (+) | − |
| Serenoa repens (W. Bartram) Small | + | + | − | − | (+) | − |
| Befaria racemosa Vent. | ++ | − | − | − | − | − |
| Lyonia ferruginea (Walt.) Nutt. | ++ | − | − | − | − | − |
| Vaccinium myrsinites Lam. | ++ | − | − | − | − | − |
| Myrica cerifera L. | ++ | + | − | − | − | − |
| Galactia elliottii Nuttall. | ++ | + | − | − | − | − |
| Gaylussacia dumosa (Andr.) A. Gray | ++ | − | − | − | − | − |
| Aristida stricta Michx. | ++ | − | − | − | − | − |
| Ascomycetes | ||||||
| Alternaria alternata (Fr.) Keissler | ++ | ++ | ++ | − | − | |
| Acremonium strictum W. Gams | ++ | ++ | ++ | (+) | − | |
| Acremonium sp. | ++ | ++ | − | + | (+) | |
| Aspergillus sp. | ++ | ++ | ++ | + | − | |
| Cladosporium cladosporioides (Fres.) de Vries | ++ | ++ | ++ | − | − | |
| Exophiala sp. | ++ | ++ | ++ | − | − | |
| Fusarium sp. | ++ | ++ | ++ | + | − | |
| Geotrichum sp. | ++ | ++ | ++ | (+) | − | |
| Penicillium purpurogenum Stoll | ++ | ++ | ++ | (+) | − | |
| Penicillium sp. strain I | ++ | ++ | − | (+) | − | |
| Penicillium sp. strain II | ++ | ++ | ++ | + | − | |
| Scopulariopsis brevicaulis (Sacc.) Bain. | − | + | − | − | − | |
| Trichoderma viride Pers. ex Gray | ++ | ++ | ++ | (+) | − | |
| Basidiomycetes | ||||||
| Agaricus sp. | ++ | ++ | + | ++ | ++ | |
| Amanita citrina (Schaeff) Roques | ++ | ++ | ++ | ++ | ++ | |
| Cortinarius sp. I | ++ | ++ | (+) | ++ | ++ | |
| Cortinarius sp. II | ++ | ++ | − | ++ | ++ | |
| Croogumphus vinicolor (Peck) O. K. Miller, Jr. | ++ | ++ | − | ++ | ++ | |
| Entoloma sp. | ++ | ++ | − | ++ | ++ | |
| Laccaria laccata (Scop.:Fr.) Cooke | ++ | ++ | − | ++ | ++ | |
| Lactarius paradoxus Beards. & Burl. | ++ | ++ | − | ++ | ++ | |
| Lactarius subisabellinus var. murrilianus | ++ | ++ | (+) | ++ | ++ | |
| Pluteus cervinus (Schaef.) Kumm | ++ | ++ | − | ++ | ++ | |
| Russula sp. (fragiloides?) | ++ | ++ | − | ++ | ++ | |
| Suillus castanellus? (Peck) Smith & Thiers | ++ | ++ | − | ++ | ++ | |
The temperatures indicate the annealing temperatures used with the different primer sets. Symbols: −, no amplification; (+), weak amplification; +, good amplification; ++, strong amplification.
In addition to collecting these soil samples, we also collected soil for isolation of fungi by enrichment plating according to the procedure described by Klamer and Søchting (32). The soil particles were plated on malt extract agar and carboxymethyl cellulose agar (17) and incubated at room temperature. Fungi growing on the plates were identified morphologically at least to the genus level. Mushrooms found on the surface in the area were also collected during the autumn of 2000. Because of a severe drought, mushrooms were collected in a wider area than the study site. DNA was extracted and analyzed using the same methods as described above with the sole exception being that fungal tissue from fruiting bodies was extracted using bead-beating lysis instead of heat.
The fungal DNA extracted from the pure cultures was used to optimize PCR parameters, to test the specificity of primer sets, and to select appropriate restriction enzymes for use in the community analysis.
To further test the specificity of the primer sets, DNAs from leaves from 11 plant species from the site (Table 1) constituting more than 90% of the aboveground biomass (54) were extracted and amplified using the technique described above (yield, 0.7 to 2.7 ng of DNA/μl). The DNAs from three oak species and Serenoa repens were extracted using DNeasy (Qiagen) for plant DNA extraction, since the Mo Bio kit did not yield amplifiable DNA for these species. Prior to extraction the leaves were surface sterilized using the following procedure: 1 min in 95% ethanol, 3 min in 4% sodium hypochlorite solution, 30 s in 95% ethanol, 5 min of washing in sterile water. To determine if the plant DNA was suitable for amplification, PCR was conducted using a universal eukaryotic primer set: Euk SSU_f-Euk SSU_r (37) (Table 1).
Principal component analyses (PCA) were performed using SPSS 10 software. Only terminal restriction fragments (TRFs) that were present in at least four samples were included. Each individual TRF was scored as present or absent and analyzed as binary data.
RESULTS
Fungal biomass.
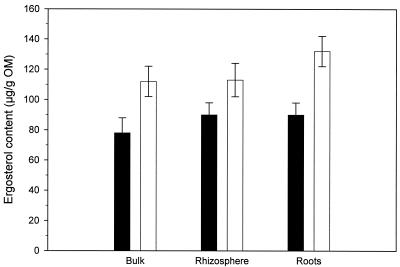
The amount of ergosterol calculated on the basis of organic matter (OM) found in the three soil compartments is shown in Fig. 1. In the bulk soil and root fractions, the increase in ergosterol in the elevated CO2 chambers was significant (two-tailed t test, P = 0.01 and 0.05, respectively [n = 8]) compared to the ambient chambers. There was no difference in ergosterol content between the soil fractions for either ambient or elevated chambers.
FIG. 1.
Influence of elevated atmospheric CO2 concentrations on ergosterol content in three different fractions of soil (n = 8). Closed bars, ambient CO2 level; open bars, elevated CO2 level. Error bars represent one standard error.
Fungal community analysis.
As shown in Table 1, none of the plant species amplified with the primer set specific for higher fungi (ITS 1F-ITS 4). The annealing temperature was lowered to 50°C with this primer set since some of the ascomycetes failed to amplify at 55°C in pure culture. The three oak species and approximately two-thirds of the ascomycetes did, however, amplify with the basidiomycete-specific primer set (ITS 1F-ITS 4B), although the amplifications were weaker than those for the basidiomycetes. This led us to test the ITS 1-ITS 4B combination, since these primers have higher predicted melting temperatures (64.5 and 66.3°C, respectively). This allowed us to increase the annealing temperature to 60°C and thereby increase the specificity while maintaining amplification of all the tested fungi extracted from pure culture. As a result of these primer evaluations, the ITS 1F-ITS 4 primer set was used as a specific primer set for the fungal community (ascomycetes and basidiomycetes), and the ITS 1-ITS 4B primer set was used for basidiomycetes. We also tested the primer set ITS 1F-ITS 4A, designed to be specific for ascomycetes (33), but we were not able to obtain satisfactory specificity with this primer set and its application for community analyses was terminated (Table 1).
In both the bulk and rhizosphere soils the increase in atmospheric CO2 did not result in significant changes (t test) in the total richness of either the fungal community or the basidiomycetes, as evaluated by the total number of TRFs (Table 2). Only few TRFs very common for both bulk and rhizosphere soils, e.g., for TaqI TRFs 1 to 9 (average, 3.1) TRFs were common for the two soil fractions from the 16 samples. In the rhizosphere soil there was no difference in species richness between the fungal community and the basidiomycetes evaluated with HinfI. When evaluated with TaqI, on the other hand, the number of TRFs was significantly lower for the basidiomycete fraction compared to the whole fungal community for both the ambient and elevated CO2 treatment (t test, P = 0.04 and 0.003, respectively).
TABLE 2.
Species richness evaluated by number of terminal restriction fragments
| CO2 concn | Mean no. of terminal restriction fragments (SE) in:
|
|||||
|---|---|---|---|---|---|---|
| Bulk soil fungal community
|
Rhizosphere soil
|
|||||
| Fungal community
|
Basidiomycetes
|
|||||
| Hinf I | Taq I | Hinf I | Taq I | Hinf I | TaqI | |
| Ambient | 25 (2.6) | 29 (3.4) | 23 (4.7) | 29 (4.2) | 21 (2.4) | 21 (2.8) |
| Elevated | 23 (2.8) | 25 (3.3) | 20 (2.2) | 35 (2.1) | 19 (2.1) | 24 (2.4) |
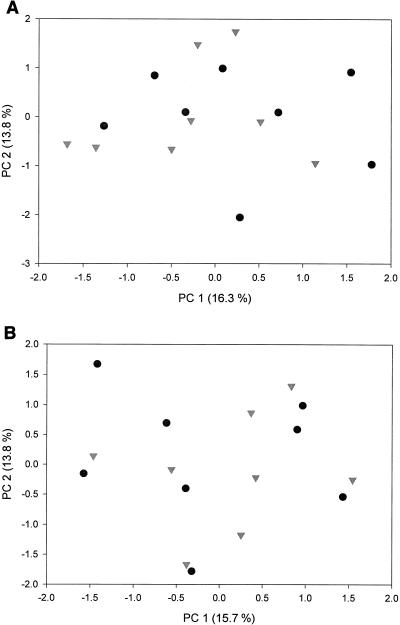
No impact of the CO2 treatment was observed in the fungal community in either the bulk or the rhizosphere soil as evaluated from the PCA (Fig. 2). There was a large variation between samples for both treatments.
FIG. 2.
PCA showing the influence of elevated atmospheric CO2 concentrations on the fungal community in soil. Symbols: •, ambient CO2 level; ▾, elevated CO2 level. (A) Bulk soil; (B) rhizosphere soil.
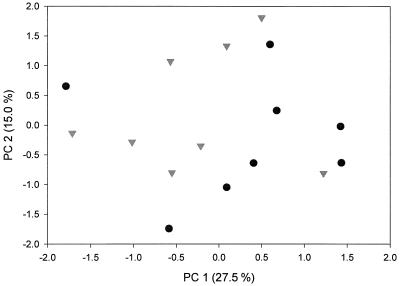
A PCA of the basidiomycetes in the rhizosphere soil is shown in Fig. 3. Although there is a large variation, there is a grouping of the ambient samples to the right and the elevated CO2 to the left, indicating that there may have been a shift in the composition of the community of Basidiomycetes caused by the elevated CO2. When tested using a Mantel test (Mantel-Struct 1.0) on all the samples, this change was not significant (P = 0.646), but when the two outliers in Fig. 3 were excluded, a significant treatment effect was found (P = 0.040). No treatment effect was observed for the basidiomycetes in the bulk soil (data not shown).
FIG. 3.
PCA showing the influence of elevated atmospheric CO2 concentrations on the basidiomycetes in the rhizosphere soil. Symbols: •, ambient CO2 level; ▾, elevated CO2 level.
DISCUSSION
Fungal biomass increased in each of the three soil fractions in response to the CO2 treatment (Fig. 1), although this increase was only significant for the bulk soil and root fractions. The increased ergosterol content can be interpreted as a secondary response to elevated atmospheric CO2 in this way: in the elevated CO2 treatments, the tree species Quercus myrtifolia, Q. geminata, and Q. chapmanii, which comprise more than 60% of the aboveground biomass (54), allocate more carbon to the ectomycorrhiza in the root fraction. The carbon is transferred from the roots to the extraradical mycelium via the Hartig net, resulting in an increase in fungal biomass in the root and the two soil fractions.
An increase in root length densities was also observed in the elevated CO2 treatment compared to the ambient treatment, 14.2 versus 8.7 mm cm−2, respectively (8). This result cannot explain the increased amount of ergosterol in the root fraction, since the ergosterol concentration was measured per unit root. The range found in this study, 80 to 130 μg/g of OM, falls within the range reported by Frostegård and Bååth (16). They measured ergosterol content in a series of Spruce and Beech forests, and found concentrations in the range of 35 to 218 μg/g of OM. In contrast to the findings in this study, however, Wiemken et al. (66) found increased fungal biomass only in a nutrient-rich calcareous soil and no effect in a nutrient-poor siliceous soil. They used the amount of the phospholipid fatty acid, 18:2ω6,9, to estimate the fungal biomass in a Beech-Spruce model system exposed to elevated atmospheric CO2.
With the reported combinations of ITS primers, it was possible to amplify higher fungi and basidiomycetes, respectively, without amplification of plant nucleic acids. The combination of ITS 1-ITS 4B with an elevated annealing temperature of 60°C yielded a higher degree of specificity for basidiomycetes in this study than the ITS 1F-ITS 4B primer set. The ITS 1-ITS 4B primer combination remains to be tested on a larger collection of different basidiomycete species in order to verify whether this primer set actually can amplify all the basidiomycetes found in this ecosystem. Glen et al. (20) tested three different primer sets targeting the ITS region and found a high specificity for basidiomycetes with one of them. However, none of the primer sets were able to amplify all 94 of the basidiomycetes tested.
The species richness, evaluated as the total number of TRFs was not influenced by the CO2 treatment, and the richness seemed to be identical in the bulk and rhizosphere soil, respectively (Table 2). Relative species richness was expected to decrease in the basidiomycete community compared to the whole fungal community, as the basidiomycetes only constitute a fraction of the whole fungal community. In the rhizosphere soil the species richness evaluated with HinfI was about 21 for both the whole fungal community and the basidiomycete community, while a significant reduction in richness was observed with TaqI. The pattern obtained with TaqI was confirmed with another tetrameric restriction enzyme, MseI (data not shown). The reason why the results from HinfI differed from those observed with TaqI and MseI is not known, but this difference emphasizes the importance of using more than one restriction enzyme when evaluating microbial community profiles from environmental samples.
PCA of both the bulk and rhizosphere soil fraction fungal communities failed to reveal any significant CO2 treatment effect (Fig. 2), but any signal may have been obscured by the large variation between samples. Mycorrhizal fungi are known to be nonrandomly distributed (i.e., spatially clustered) in soil systems. Gardes and Bruns (19) have reported that the majority of mycorrhizal fungal species occurred in less than 10% of the soil samples taken. In a recent review by Horton and Bruns (25), it was noted that the most abundant types of fungal biomass occurred in only one or two soil samples, suggesting a highly heterogeneous distribution. Furthermore, canopy cover has been shown to impact the amount, if not also the type, of ectomycorrhizal fungi associated with oak (68). Changing the sample locations only slightly would change the perception of species presence and estimates of richness considerably. In this study, we have eight replicate samples from the two treatments, which should yield sufficient sample density to detect major changes in fungal community composition resulting from the CO2 treatments. It is obvious, however, that additional spatial and temporal sampling intervals will be required in future studies to dampen the noise inherent in patchy ecosystems and augment any possible signal of community compositional changes arising from elevated CO2 treatments.
Despite a large variation among replicate samples in the rhizosphere soil PCA plot, the data support a change in the basidiomycete community as a consequence of the elevated atmospheric CO2 (Fig. 3). Other unaccounted factors may, however, play a role as well. For example, there is an indication of a change in the composition of oak species, with Q. geminata being less abundant in the plots with elevated CO2 levels. Changes in the composition of the basidiomycete community may therefore be a secondary effect arising from the altered oak species composition, as it is likely that a large proportion of the basidiomycetes are ectomycorrhizal fungi associated with particular plant species. There is other evidence, however, that at least part of the ectomycorrhizal fungi are able to colonize a variety of tree species (24, 30), which could counteract the importance of changes in the aboveground plant species composition. Any change in either the oak species composition or their physiology would result in changes in root physiology and the rhizosphere that would affect the associated ectomycorrhiza and fungi. Fungi may be critical reservoirs of carbon in systems with elevated atmospheric CO2 since they constitute the highest amount of microbial biomass in the soil system and are intimately involved in the flux of carbon channeled directly from the aboveground biomass and indirectly from the atmosphere. Given the rather limited knowledge of fungal diversity and ecosystem function in elevated CO2 environments, it is unclear what potential impact any changes in fungal composition would have on community stability at either a functional diversity or richness level.
We conclude that 5 years of continuous treatment with elevated atmospheric CO2 concentrations has resulted in a higher fungal biomass in the soil. Neither fungal community composition nor richness, however, was demonstrably affected by the elevated CO2 treatment. The change in the composition of basidiomycetes is believed to be a secondary effect reflecting changes in either the plant species composition (e.g., oaks) or a shift in the ectomycorrhizal community on oak roots arising from changes in carbon flux.
Acknowledgments
This project was supported by a NASA Technology Development Grant from Kennedy Space Center through the University of Central Florida (M.K.).
We are grateful for the support by S. and G. Sheine in collecting and identifying basidiomycetes. We thank G. Hymus and P. Schmalzer for their help with identification of the plant species.
REFERENCES
- 1.Bermingham, S., L. Maltby, and R. C. Cooke. 1995. A critical assessment of the validity of ergosterol as an indicator of fungal biomass. Mycol. Res. 99:479-484. [Google Scholar]
- 2.Borneman, J., and R. J. Hartin. 2000. PCR primers that amplify fungal rRNA genes from environmental samples. Appl. Environ. Microbiol. 66:4356-4360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Braker, G., H. L. Ayala-del-Rio, A. H. Devol, A. Fesefeldt, and J. M. Tiedje. 2001. Community structure of denitrifiers, bacteria, and archaea along redox gradients in Pacific Northwest marine sediments by terminal restriction fragment length polymorphism analysis of amplified nitrite reductase (nirS) and 16S rRNA genes. Appl. Environ. Microbiol. 67:1893-1901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bruce, K. D. 1997. Analysis of the mer gene subclass within bacterial communities in soils and sediments resolved by fluorescent-PCR restriction fragment length polymorphism profiling. Appl. Environ. Microbiol. 63:4914-4919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Clement, B. G., and C. L. Kitts. 2000. Isolating PCR-quality DNA from human feces with a soil DNA kit. BioTechniques 28:640-645. [DOI] [PubMed] [Google Scholar]
- 6.Curtis, P. S., D. R. Zak, K. S. Pregitzer, and J. A. Teeri. 1994. Above and below ground response of Populus grandidentata to elevated atmospheric CO2 and soil N availability. Plant Soil 165:45-51. [Google Scholar]
- 7.Davis, M. W., and R. T. Lamar. 1992. Evaluation of methods to extract ergosterol for quantification of soil fungal biomass. Soil Biol. Biochem. 24:189-198. [Google Scholar]
- 8.Day, F. P., E. P. Weber, C. R. Hinkle, and B. G. Drake. 2000. Effects of elevated CO2 on fine root length and distribution in an oak-palmetto scrub ecosystem in central Florida. Global Change Biol. 2:143-148. [Google Scholar]
- 9.Diaz, S., J. P. Grime, J. Harris, and E. Mcpherson. 1993. Evidence for a feedback mechanism limiting plant response to elevated carbon dioxide. Nature 364:616-617. [Google Scholar]
- 10.Djajakirana, G., and R. G. Joergensen. 1996. Changes in soil organic matter, microbial biomass C and ergosterol under a fairy ring of Marasmius oreades. Pedobiololgy 40:498-504. [Google Scholar]
- 11.Dollhopf, S. L., S. A. Hashsham, F. B. Dazzo, R. F. Hickey, C. S. Criddle, and J. M. Tiedje. 2001. The impact of fermentative organisms on carbon flow in methanogenic systems under constant low-substrate conditions. Appl. Microbiol. Biotechnol. 56:531-538. [DOI] [PubMed] [Google Scholar]
- 12.Dunbar, J., L. O. Ticknor, and C. R. Kuske. 2000. Assessment of microbial diversity in four Southwestern United States soils by 16S rRNA gene terminal restriction fragment analysis. Appl. Environ. Microbiol. 66:2943-2950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Eash, N. S., P. D. Stahl, T. B. Parkin, and D. L. Karlen. 1996. A simplified method for extraction of ergosterol from soil. Soil Sci. Soc. Am. J. 60:468-471. [Google Scholar]
- 14.Ek, H., M. Sjögren, K. Arnebrant, and B. E. Söderström. 1994. Extramatrical mycelial growth, biomass allocation and nitrogen uptake in ectomycorrhizal systems in response to collembolan grazing. Appl. Soil Ecol. 1:155-169. [Google Scholar]
- 15.Frankland, J. C. 1990. Ecological methods of observing and quantifying soil fungi. Trans. Mycol. Soc. Jpn. 31:89-101. [Google Scholar]
- 16.Frostegård, Å., and E. Bååth. 1996. The use of phospholipid fatty acid analysis to estimate bacterial and fungal biomass in soil. Biol. Fertil. Soils 22:59-65. [Google Scholar]
- 17.Gams, W., H. A. van der Aa, A. J. van der Plaats-Niterink, R. A. Samson, and J. A. Stalpers. 1987. CBS course of mycology. Centraalbureau voor Schimmelcultures, Baarn, The Netherlands.
- 18.Gardes, M., and T. D. Bruns. 1993. ITS primers with enhanced specificity for basidiomycetes: application to the identification of mycorrhizae and rusts. Mol. Ecol. 2:113-118. [DOI] [PubMed] [Google Scholar]
- 19.Gardes, M., and T. D. Bruns. 1996. Community structure of ectomycorrhizal fungi in a Pinus muricata forest: above- and below-ground views. Can. J. Bot. 74:1572-1583. [Google Scholar]
- 20.Glen, M., I. C. Tommerup, N. L. Bougher, and P. A. O'Brien. 2001. Specificity, sensitivity and discrimination of primers for PCR-RFLP of larger basidiomycetes and their applicability to identification of ectomycorrhizal fungi in Eucalyptus forests and plantations. Mycol. Res. 105:138-149. [Google Scholar]
- 21.Grant, W. D., and A. W. West. 1986. Measurement of ergosterol, diaminopimelic acid and glucosamine in soil: evaluation as indicators of microbial biomass. J. Microbiol. Methods 6:47-53. [Google Scholar]
- 22.Griffiths, B. S., K. Ritz, R. D. Bardgett, R. Cook, S. Christensen, F. Ekelund, S. J. Sorensen, E. Bååth, J. Bloem, P. C. d. Ruiter, J. Dolfing, and B. Nicolardot. 2000. Ecosystem response of pasture communities to fumigation-induced microbial diversity reductions: an examination of the biodiversity-ecosystem function relationship. Oikos 90:279-294. [Google Scholar]
- 23.Griffiths, B. S., K. Ritz, N. Ebblewhite, E. Paterson, and K. Killham. 1998. Ryegrass rhizosphere microbial community structure under elevated carbon dioxide concentrations, with observations on wheat rhizosphere. Soil Biol. Biochem. 30:315-321. [Google Scholar]
- 24.Horton, T. R., and T. D. Bruns. 1998. Multiple-host fungi are the most frequent and abundant ectomycorrhizal types in a mixed stand of Douglas fir (Pseudotsuga menziesii) and bishop pine (Pinus muricata). New Phytol. 139:331-339. [Google Scholar]
- 25.Horton, T. R., and T. D. Bruns. 2001. The molecular revolution in ectomycorrhizal ecology: peeking into the black-box. Mol. Ecol. 10:1855-1871. [DOI] [PubMed] [Google Scholar]
- 26.Horz, H.-P., J.-H. Rotthauwe, T. Lukow, and W. Liesack. 2000. Identification of major subgroups of ammonia-oxidizing bacteria in environmental samples by T-RFLP analysis of amoA PCR products. J. Microbiol. Methods 39:197-204. [DOI] [PubMed] [Google Scholar]
- 27.Hungate, B. A., P. Dijkstra, D. W. Johnson, C. R. Hinkle, and B. G. Drake. 1999. Elevated CO2 increases nitrogen fixation and decreases soil nitrogen mineralization in Florida scrub oak. Global Change Biol. 5:781-789. [Google Scholar]
- 28.Jackson, C. J., R. C. Barton, and E. G. Evans. 1999. Species identification and strain differentiation of dermatophyte fungi by analysis of ribosomal-DNA intergenic spacer regions. J. Clin. Microbiol. 37:931-936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Johnson, D. W., B. A. Hungate, P. Dijkstra, G. Hymus, and B. G. Drake. 2001. Effects of elevated carbon dioxide on soils in a Florida scrub oak ecosystem. J. Environ. Qual. 30:501-507. [DOI] [PubMed] [Google Scholar]
- 30.Kernaghan, G. 2001. Ectomycorrhiza fungi at tree line in the Canadian Rockies. II. Identification of ectomycorrhizae by anatomy and PCR. Mycorrhiza 10:217-229. [Google Scholar]
- 31.Kitts, C. L. 2001. Terminal restriction fragment patterns: a tool for comparing microbial communities and assessing community dynamics. Curr. Issues Intest. Microbiol. 2:17-25. [PubMed] [Google Scholar]
- 32.Klamer, M., and U. S¢chting. 1998. The fungi in a controlled compost system—with special emphasis on thermophilic fungi. Acta Hort. 469:405-413. [Google Scholar]
- 33.Larena, I., O. Salazar, V. Gonzalez, M. C. Julian, and V. Rubio. 1999. Design of a primer for ribosomal DNA internal transcribed spacer with enhanced specificity for ascomycetes. J. Bio/Technology 75:187-194. [DOI] [PubMed] [Google Scholar]
- 34.Lösel, D. M. 1988. Fungal lipids, p. 699-806. In C. Ratledge and S. G. Wilkinson (ed.), Microbial lipids. Academic Press, London, United Kingdom.
- 35.Lukow, T., P. F. Dunfield, and W. Liesack. 2000. Use of the T-RFLP technique to assess spatial and temporal changes in the bacterial community structure within an agricultural soil planted with transgenic and non-transgenic potato plants. FEMS Microbiol. Ecol. 32:241-247. [DOI] [PubMed] [Google Scholar]
- 36.Matcham, S. E., B. R. Jordan, and D. A. Wood. 1985. Estimation of fungal biomass in a solid substrate by three independent methods. Appl. Microbiol. Biotechnol. 21:108-112. [Google Scholar]
- 37.Medlin, L., H. J. Elwood, S. Stickel, and M. L. Sogin. 1988. The characterization of enzymatically amplified eukariotic 16S-like rRNA-coding regions. Gene 71:491-499. [DOI] [PubMed] [Google Scholar]
- 38.Moeseneder, M. M., J. M. Arrieta, G. Muyzer, C. Winter, and G. J. Herndl. 1999. Optimization of terminal-restriction fragment length polymorphism analysis for complex marine bacterioplankton communities and comparison with denaturing gradient gel electrophoresis. Appl. Environ. Microbiol. 65:3518-3525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Mohlenhoff, P., P. Muller, A. A. Gorbushina, and K. Petersen. 2001. Molecular approach to the characterisation of fungal communities: methods for DNA extraction, PCR amplification and DGGE analysis of painted art objects. FEMS Microbiol. Lett. 195:169-173. [DOI] [PubMed] [Google Scholar]
- 40.Montealegre, C. M., C. van Kessel, J. M. Blumenthal, H.-G. Hur, U. A. Hartwig, and M. J. Sadowsky. 2000. Elevated atmospheric CO2 alters microbial structure in a pasture ecosystem. Global Change Biol. 6:475-482. [Google Scholar]
- 41.Nemec, T., K. Jernejc, and A. Cimerman. 1997. Sterols and fatty acids of different Aspergillus species. FEMS Microbiol. Lett. 149:201-205. [Google Scholar]
- 42.Newell, S. Y. 1992. Estimating fungal biomass and productivity in decomposing litter, p. 521-562. In G. C. Carroll and D. T. Wicklow (ed.), The fungal community—its organization and role in the ecosytem. Marcel Dekker, Inc., New York, N.Y.
- 43.Newell, S. Y., J. D. Miller, and R. D. Fallon. 1987. Ergosterol content of salt-marsh fungi: effect of growth conditions and mycelial age. Mycologia 79:688-695. [Google Scholar]
- 44.Olsson, P. A., E. Bååth, I. Jakobsen, and B. E. Söderström. 1996. Soil bacteria respond to presence of roots but not to mycelium of arbuscular mycorrhizal fungi. Soil Biol. Biochem. 28:463-470. [Google Scholar]
- 45.O'Neill, E. G., R. J. Luxmoore, and R. J. Norby. 1987. Increases in mycorrhizal colonization and seedling growth in Pinus echinata and Quercus alba in an enriched CO2 atmosphere. Can. J. For. Res. 17:878-883. [Google Scholar]
- 46.Osborn, A. M., E. R. B. Moore, and K. N. Timmis. 2000. An evaluation of thermal-restriction fragment length polymorphism (T-RFLP) analysis for the study of microbial community structure and dynamics. Environ. Microbiol. 2:39-50. [DOI] [PubMed] [Google Scholar]
- 47.Padgett, D. E., and M. H. Posey. 1993. An evaluation of the efficiencies of several ergosterol extraction techniques. Mycol. Res. 97:1476-1480. [Google Scholar]
- 48.Pennanen, T., H. Fritze, P. Vanhala, O. Kiikkila, S. Neuvonen, and E. Bååth. 1998. Structure of a microbial community in soil after prolonged addition of low levels of simulated acid rain. Appl. Environ. Microbiol. 64:2173-2180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Rice, C. W., F. O. Garcia, C. O. Hampton, and C. E. Owensby. 1994. Soil microbial response in tall grass prairie to elevated CO2. Plant Soil 165:67-74. [Google Scholar]
- 50.Rouhier, H., G. Billes, A. El Kohen, M. Mousseau, and P. Bottner. 1994. Effect of elevated CO2 on carbon and nitrogen distribution within a tree (Castanea sativa) soil system. Plant Soil 162:281-292. [Google Scholar]
- 51.Sadowsky, M. J., and M. Schortemeyer. 1997. Soil microbial responses to increased concentrations of atmospheric CO2. Global Change Biol. 3:217-224. [Google Scholar]
- 52.Scala, D., and L. Kerkhof. 2000. Horizontal heterogeneity of denitrifying bacterial communities in marine sediments by terminal restriction fragment length polymorphism analysis. Appl. Environ. Microbiol. 66:1980-1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Scheu, S., and D. Parkinson. 1994. Changes in bacterial and fungal biomass C, bacterial and fungal biovolume and ergosterol content after drying, remoistening and incubation of different layers of cool temperate forest soils. Soil Biol. Biochem. 26:1515-1525. [Google Scholar]
- 54.Schmalzer, P. A., and C. R. Hinkle. 1996. Biomass and nutrients in aboveground vegetation and soils of Florida oak-saw palmetto scrub. Castanea 61:168-193. [Google Scholar]
- 55.Schnürer, J. 1993. Comparison of methods for estimating the biomass of three food-borne fungi with different growth patterns. Appl. Environ. Microbiol. 59:552-555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Schortemeyer, M., P. Dijkstra, D. W. Johnson, and B. G. Drake. 2000. Effects of elevated atmospheric CO2 concentration on C and N pools and rhizosphere processes in a Florida scrub oak community. Global Change Biol. 6:383-391. [Google Scholar]
- 57.Smit, E., P. Leeflang, B. Glandorf, J. D. van Elsas, and K. Wernars. 1999. Analysis of fungal diversity in the wheat rhizosphere by sequencing of cloned PCR-amplified genes encoding 18S rRNA and temperature gradient gel electrophoresis. Appl. Environ. Microbiol. 65:2614-2621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Taylor, J. W., D. M. Geiser, A. Burt, and V. Koufopanou. 1999. The evolutionary biology and population genetics underlying fungal strain typing. Clin. Microbiol. Rev. 12:126-146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.van Elsas, J. D., G. F. Duarte, A. C. Keijzer-Wolters, and E. Smit. 2000. Analysis of the dynamics of fungal communities in soil via fungal-specific PCR of soil DNA followed by denaturing gradient gel electrophoresis. J. Microbiol. Methods 43:133-151. [DOI] [PubMed] [Google Scholar]
- 60.VAn Veen, J. A., E. Liljeroth, L. J. A. Lekkerkerk, and S. C. Van de Geijn. 1991. Carbon fluxes in plant-soil systems at elevated atmospheric CO2 levels. Ecol. Appl. 1:175-181. [DOI] [PubMed] [Google Scholar]
- 61.Weete, J. D. 1980. Lipid biochemistry of fungi and other organisms. Plenum Press, New York, N.Y.
- 62.Weete, J. D., and S. R. Gandhi. 1997. Sterols of the phylum Zygomycota: phylogenetic implications. Lipids 32:1309-1316. [DOI] [PubMed] [Google Scholar]
- 63.Weete, J. D., and S. R. Gandhi. 1999. Sterols and fatty acids of the Mortierellaceae: taxonomic implications. Mycologia 91:642-649. [Google Scholar]
- 64.West, A. W., W. D. Grant, and G. P. Sparling. 1987. Use of ergosterol, diaminopimelic acid and glucosamine contents of soils to monitor changes in microbial populations. Soil Biol. Biochem. 19:607-612. [Google Scholar]
- 65.White Thomas, J., T. D. Bruns, S. Lee, and J. W. Taylor. 1990. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics, p. 315-324. In M. A. Innis, D. H. Gelfand, J. J. Sninsky, and T. J. White (ed.), PCR protocols: a guide to methods and applications. Academic Press, Inc., San Diego, Calif..
- 66.Wiemken, V., E. Laczkó, K. Ineichen, and T. Boller. 2001. Effects of elevated carbon dioxide and nitrogen fertilization on mycorrhizal fine roots and the soil microbial community in Beech-Spruce ecosystems on siliceous and calcareous soil. Microb. Ecol. 42:126-135. [DOI] [PubMed] [Google Scholar]
- 67.Zak, D. R., D. B. Ringelberg, K. S. Pregitzer, D. L. Randlett, D. C. White, and P. S. Curtis. 1996. Soil microbial communities beneath Populus granddentata grown under elevated atmospheric CO2. Ecol. Appl. 6:257-262. [Google Scholar]
- 68.Zhou, M., T. L. Sharik, M. F. Jurgensen, and D. L. Richter. 1997. Ectomycorrhizal colonization of Quercus rubra seedlings in response to vegetation removals in oak and pine stands. For. Ecol. Manag. 93:91-99. [Google Scholar]