Abstract
Objectives
To develop a data harmonization framework for neonatal hypoxic-ischemic encephalopathy (HIE) studies and demonstrate its suitability for prognostic biomarker development.
Materials and Methods
Variables were first categorized by chronological stages and then by medical topics. We created a dictionary to harmonize variable names and value coding. We began by merging comprehensive data from 2 landmark nationwide therapeutic hypothermia for HIE trials (2008-2016, 21 sites) in the Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD) Neonatal Research Network (NRN). The 2 datasets differ in available variables, variable naming and coding, necessitating harmonization. To demonstrate the utility of this data harmonization framework, we computed the distributions of variables and ranked them by the strength of associations with 18- to 22-month outcomes. Associations were measured using Pearson’s correlation analysis. Outcomes were defined as (a) a 5-class variable: survivors with normal, mild, moderate, severe disability, or death and (b) the Bayley-III Scales.
Results
We created a dictionary of 1181 variables on 532 patients across 5 chronologic categories and 60 medical subcategories. The distribution of major predictive and outcome variables, and the variables strongly associated with neurodevelopmental outcomes at 18-22 months were presented. The modified Sarnat scores at the Post-intervention and NICU-discharge stage, and the NRN pattern of MRI injury score showed strong associations with outcome variables.
Conclusion
We designed a data harmonization framework specifically for HIE. Our initial effort in merging 2 iconic clinical trials shows strong predictor-outcome associations, allowing subsequent development of advanced prognostic biomarkers of neonatal HIE.
Keywords: hypoxic-ischemic encephalopathy, data harmonization framework, common data elements, biomarker development
Background and significance
Hypoxic-ischemic encephalopathy (HIE) is a brain condition occurring in 1-5 out of every 1000 term-born neonates, characterized by insufficient blood flow and oxygen to the brain, potentially resulting in lifelong brain functional disability.1–4 In 2022, the Consortium Of MRI Biomarkers In Neonatal Encephalopathy (COMBINE) was established by a group of neonatologists, pediatric neurologists, pediatric neuroradiologists, along with computer and data scientists. Its mission is to develop artificial intelligence (AI)-driven prognostic clinical and imaging biomarkers that can predict, during the neonatal stage, whether a specific HIE patient is at risk of developing adverse neurologic outcomes by 2 years of age. Although the introduction of therapeutic hypothermia in 2005 for moderate or severe HIE has reduced morbidity and mortality in high-income countries,5–8 the 2-year adverse outcome, defined as moderate/severe neurodevelopmental disability or death, continues to negatively impact around one third of moderate or severe HIE patients.9,10 An accurate prognostic biomarker could serve as the basis for identifying high-risk patients, evaluating treatment effects early, and expediting therapeutic innovations in targeted sub-cohorts, toward eventually improving neurodevelopmental outcomes in HIE.11–14
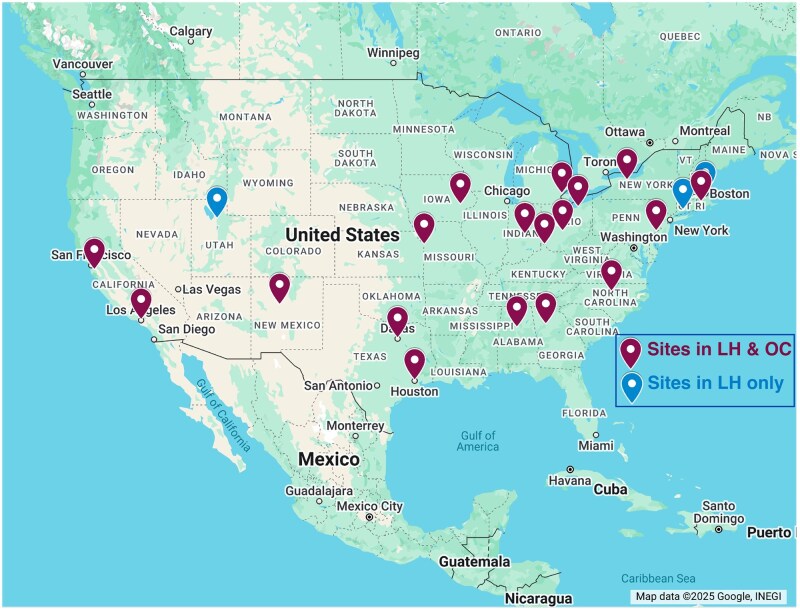
One fundamental step in COMBINE is to amass a large-scale and comprehensive database for AI-driven biomarker development. Most HIE biomarker studies to date have used data from only dozens or a few hundreds of patients, with clinical variables often considered in isolation. The lack of comprehensive large-scale diverse data hinders the accuracy and application of AI approaches for biomarker development. The Neonatal Research Network (NRN) within the Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD) has authorized the COMBINE consortium to use existing data from 2 large-scale nationwide HIE trials. The primary goal of the Late Hypothermia (LH, 2008-2016, NCT00614744, 21 sites) trial was to compare whether the initiation of cooling within 6-24 hours was neuroprotective compared to normothermia,15 while the Optimizing Cooling (OC, 2010-2014, NCT01192776, 18 sites) trial examined whether longer or deeper cooling was neuroprotective.10 A total of 532 patients were enrolled from 21 hospitals in the United States (see sites in Figure 1). Each trial collected over 1000 variables, covering pregnancy, delivery, maternal and infant demographics, treatment, infant neuroimaging, follow-up outcome measurements, and other comprehensive information.
Figure 1.
The 21 sites in the LH and 18 sites in the OC trials. Locations are based on Google Maps.
Merging comprehensive and diverse data into a coherent database is, however, not a trivial task.16,17 Challenges stem from at least 3 aspects: (a) variable availability: some variables are present in only one trial; (b) variable naming: the same variable may have different names across trials when deposited into a central data repository (eg, the variable for maternal race was named MRACE in the LH dataset but RACE in the OC dataset); and (c) coding of variable values: the values for the same variable may differ among different trials (eg, sex coded as M/F in one dataset and 0/1 in another).
This paper presents our benchmark work in the COMBINE consortium to merge and harmonize existing data from these 2 NICHD NRN trials on HIE. The Methods section details (1) the data harmonization framework that creates the dictionary of common data elements (CDE) to harmonize the 2 HIE datasets. The Results section covers (2) the characteristics of the merged database and the dictionary of HIE variables; and (3) results from mass-univariate analysis demonstrating the utility of the merged database.
Methods
Fortier et al. noted that, starting with defining the questions and objectives, at least 6 key steps are required to disseminate and preserve final harmonization products.16 Cheng et al, citing Stuckenschmidt, highlighted 3 essential components: file format, categorization of variables, and variable naming and value coding.17,18 The F.A.I.R. principles, denoting findability, accessibility, interoperability, and reusability, are widely recognized as foundational guidelines for data management and exchange.19 Our development of the data harmonization framework aligns with these 3 guidelines.
Harmonization framework
Variable categorization
In our data harmonization framework, we categorized the variables into 5 categories, reflecting the progression of HIE care: (1) Pre-intervention—screening and baseline information before starting the intervention, such as maternal demographics information, pregnancy history, labor and delivery details, and infant’s condition at birth; (2) Intervention—repeated measurements related to various body systems during the intervention, such as temperature, cardiovascular function, and blood values; (3) Post-intervention—measurements after the intervention, such as continuous blood measurements, brain MRI, and neurological exams; (4) Neonatal Intensive Care Unit (NICU) Discharge—records of clinical diagnosis at death or discharge; and (5) Follow-Up—study-specific measurements at the follow-up visit and interim medical history. Categories 1-4 occur during the neonatal period. In addition to this chronological categorization, we further subdivided the variables based on specific medical topics within each category.
Names and coding of variables
To ensure clarity, we used English-comprehensive naming conventions for the variables. Optionally, for variables with the same meaning across different categories, prefixes for categories may be added for further distinction. To precisely describe a variable, additional details may be necessary, such as the measurement unit for measurement variables or whether the screening criteria are inclusive or exclusive. This information is added as suffixes to the variable names.
Data types of variables
Data types are important in data analysis. In addition to the typical Boolean, numerical, date, time, and text, there are several ordinal and nominal types for categorical variables that require standardized coding. The coding for these categorical variables was also designed to be English-comprehensive. For ordinal variables, in addition to standardized coding, we had implemented a mechanism that allows for different ranking strategies based on specific contexts.
Missing variables: union-based approach
Some variables may not be included from certain datasets. We took the “union” of all the variables from 2 datasets. The advantage was to keep all variables as long as they existed in at least one dataset. This better prepared our dictionary of variables when future datasets are to be merged. The disadvantage, though, was the existence of missing values if a variable does not exist in a dataset. This is addressed as described next.
Missing values: keep the original data
Missing values were anticipated following the merging of datasets, either due to their absence in the original sources or because certain variables were not present in all datasets after harmonization. To preserve the integrity of the original data, we did not infer the missing values in either scenario. As a result, all patient-level data remained unaltered and reflected actual records. The imputation of missing data was deferred to the data analysis stage.
Derived variables to reconcile different data types
Some variables recorded as different data types might be eligible for inference from one another. In addition to preserving the original data without inference, we created derived variables, where appropriate, and to the best of our knowledge, to infer values that could be further harmonized. For example, the 5-minute Apgar scores were recorded as numerical variables in both the LH and the OC trials. However, only the OC trial included an additional variable indicating whether the 5-minute Apgar score was less or equal to 5. In addition to retaining the OC-specific variable, Apgar5minLessEq5, we created a derived variable, Apgar5minLessEq5_deriv, inferred from the numerical 5-minute Apgar scores.
Repeated variables
Some variables were collected repeatedly over time. One type of such time-series variables was measured at regular time intervals, such as temperature being measured at 15, 60, and 1440 minutes after starting of the intervention. We included the common unit of time as the postfix of the indexing variable, such as temperatureTimeSlot_min. Another type of repeated variables recorded the natural incidences of events, such as serious adverse events. These events occurred randomly, and the indices could only represent the order of the occurrence. We used “number” as the postfix of the indexing variables, such as adverseEventNumber. In spreadsheet presentation, the repeated variables were flattened by appending the indices as the postfix of the variable, such as skinTemperature_C_15min, skinTemperature_C_1440min, and SAEOther_1.
Mass-univariate analysis to show data utility
To demonstrate the utility of our data harmonization framework, we used Pearson’s correlation coefficient analysis to explore the relationships between predictor and outcome variables. In this study, adapted from the LH and OC studies, we defined the outcome variables as the 5-level disability level or death (normal, mild, moderate, severe, and death), and the Bayley-III Composite Cognitive Scales of survivors at 18-22 months of age. Our analysis focused on variables collected at the Pre-intervention, Post-intervention, NICU Discharge, and Derived Data stages. Conventionally, the significance threshold is assigned as P-value < 0.05. However, considering the inclusion of over 1000 variables in the framework, we applied Bonferroni correction and assigned the significance threshold as P-value < 10−5. For the variables meeting this threshold, we ranked the variables based on their corresponding coefficient of determination R2 in Pearson’s correlation coefficient analysis.
Data privacy
Data privacy is always a major concern when dealing with medical data. Besides the usual already-anonymized site names and subject names replaced with anonymized identifiers, the privacy of the medical practitioners should be protected as well. We manually reviewed the text and removed the names and initials of the medical practitioners. Furthermore, information about date and time can be sensitive to privacy issues. The date and time is transformed to the days since birth, with precision to the hour. Only the year and month are revealed in the birth date.
Versioning
We recognize that different studies have varying aims. It is foreseeable that other studies may include variables not present in our framework, requiring expanding the current variables. Moreover, the medical subcategories may also need to be extended to account for the uniqueness of these new variables. In addition, the coding of existing ordinal and nominal variables, such as antibiotics, may need expansion as well to accommodate new values in the future. Lastly, it is possible that the chronological categories may be different for specific needs in the future. We use semantic versioning for different versions of the data harmonization framework.20
Compliance to the F.A.I.R. principles
The F.A.I.R. principles stand for findability, accessibility, interoperability, and reusability.19 Our data harmonization framework explicitly addresses each principle as follows: (a) Findability: We assigned persistent identifiers in metadata and data dictionary, and publicly release them in GitHub version control and Supplementary Material; (b) Accessibility: All metadata, data dictionary, and software are retrievable through open GitHub repositories using standard HTTPS protocols without authentication barriers; (c) Interoperability: Our framework employed standardized naming conventions and controlled vocabularies for categorical variables, facilitating integration with other HIE datasets. The hierarchical organization by chronological stages and medical topics creates a formal, shared knowledge representation structure; and (d) Reusability: We described variables with detailed attributes in our data dictionary (Supplementary Material). All released materials are provided under permissive licensing (CC BY 4.0 for results and MIT license for software). The comprehensive documentation of our harmonization process establishes detailed provenance. To comply with both the F.A.I.R. principles and the intellectual property of the original datasets, we adopted the “separation of the Datasets, Software, and Results” strategy. This strategy is described in the Data Availability Section and Code Availability sections.
Results
Variable categorization and dictionary construction
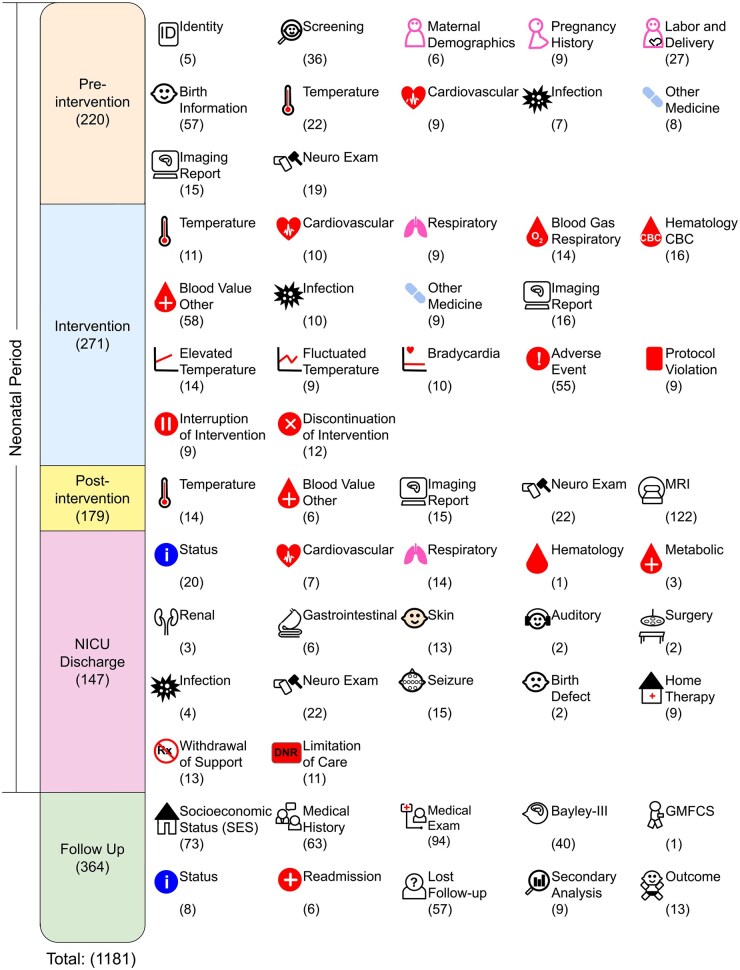
Figure 2 illustrates the categorization of variables in our data harmonization framework, grouped into 5 major categories during the clinical course and 60 subcategories based on medical topics from the LH and OC trials. The 5 major categories, along with their respective color codes, are as follows: (in the neonatal period) Pre-intervention: light peach, Intervention: light blue, Post-intervention: yellow, and Neonatal Intensive Care Unit (NICU) Discharge: pink; and (after the neonatal period) Follow-Up: green. A total of 1181 variables remain after excluding those with conceptually similar content.
Figure 2.
The categorization of variables in the LH and OC trials. The left-most column represents the 5 color-coded categories based on the clinical course. The 60 subcategories are presented with icons and descriptions. The numbers in parentheses represent the number of variables in each category and subcategory. There are 1181 variables in total. MRI, magnetic resonance imaging; GMFCS, Gross Motor Function Classification System.
As depicted in Figure 3, our framework includes 956 variables from the LH trial and 1091 variables from the OC trial, and 14 derived variables, such as derived total modified Sarnat scales and averaged MRI NRN pattern of injury scores. After taking a union of these variables, our merged database contains a total of 1181 variables.
Figure 3.

Distributions of the variables in the LH and OC trials.
Data type plays a crucial role in data analysis. Table 1 shows the number of variables for each data type. In addition to common types such as Boolean, number, date, time, or text, various ordinal and nominal types are available for harmonizing categorical variables.
Table 1.
Distributions of different data types.
| Data type | Count | Example | Description |
|---|---|---|---|
| Boolean | 401 | Apgar5minLessEq5 | 5-minute Apgar score ≤ 5 |
| number | 207 | birthWeight_g | birth weight |
| ordinal | 202 | disabilityLevelDeath | disability level or death |
| nominal | 119 | anticonvulsants | anticonvulsants |
| text | 69 | screenComment | screening comments |
| date | 123 | birthDate | birth date |
| time | 60 | birthTime | birth time |
| Total | 1181 |
Table 2 presents excerpts from our constructed data dictionary in our data harmonization framework. The full table is in Supplementary Material S1, S2, and S4. Variable names that are different between the LH and OC datasets (the last 2 columns in Table 1) are harmonized into the same variable name in this dictionary (the second column in the table).
Table 2.
Excerpts of the data dictionary.
| Subcategory | Harmonized variable | Data type | Description | |
|---|---|---|---|---|

|
Screening | birthWeightLessEq1800g_e | Boolean | Exclusion: birth weight ≤ 1800 g |
| first60MinAnyBloodGasPHLessEq7_i | Boolean | Inclusion: pH ≤ 7 in any blood gas (cord, postnatal) within the first 60 minutes | ||

|
Pregnancy history | Gravida | Number | Gravida |
| antepartumHemorrhage | Boolean | Antepartum hemorrhage | ||

|
Labor and delivery | laborOnsetDate | Date | Date of labor onset |
| laborOnsetTime | Time | Time of labor onset | ||
| deliveryMode | Nominal | Final mode of delivery | ||

|
Birth | birthweight_g | Number | Birth weight |
| birthHeadCircumference_cm | Number | Birth head circumference | ||

|
Temperature | temperatureTimeSlot_min | Number | Time slot of the measurement (unit: minute) |
| esophagealTemperature_C | Number | Esophageal temperatures | ||

|
Infection | positiveCultureNumber | Number | Infection incidence number |
| positiveCultureDate | Date | Date of positive culture | ||
| positiveCultureTime | Time | Time of positive culture | ||

|
Neuro exam | post_NeuroExamPosture | Number | Posture in neuro exam in Post-intervention |
| post_NeuroExamMoro | Number | Moro reflex in neuro exam in Post-intervention | ||

|
Imaging report | headSonogramResult1 | Nominal | The first result in the head sonogram report |
| headCTResult1 | Nominal | The first result in the head computer tomography report | ||
| brainMRIResult1 | Nominal | The first result in the brain MRI report | ||

|
MRI | MRIAvailable | Boolean | MRI available |
| MRINRNPatternOfInjury | Ordinal | NRN pattern of injury score | ||

|
Status | status | Nominal | Status of the infant |
| deathCause | Nominal | Cause of death | ||

|
Bayley-III | BayleyIIIInEnglish | Boolean | Bayley–III exam was conducted in English |
| BayleyIIICognitiveComposite | Number | Bayley–III Composite Cognitive Scale | ||

|
Outcome | moderateSevereDisabilityOrDeath | Boolean | Moderate severe disability or death |
| disabilityLevelDeath | Ordinal | Disability level or death (normal, mild, moderate, severe, death) | ||
| Subcategory | Harmonized variable | Original LH variables | Original OC variables | |
|---|---|---|---|---|

|
Screening | birthWeightLessEq1800g_e | LH2WGHT | OC2WGHT |
| first60MinAnyBloodGasPHLessEq7_i | LH2PH | OC2PH | ||

|
Pregnancy history | gravida | LH4GRAV | OC4FRAV |
| antepartumHemorrhage | LH4HMRG | OC4ANTE | ||

|
Labor and delivery | laborOnsetDate | LH4LBDT | N/A |
| laborOnsetTime | LH4LBTM | N/A | ||
| deliveryMode | LH4MODE | OC4MODE | ||

|
Birth | birthweight_g | LH5BTWGT | OC5BWHGT |
| birthHeadCircumference_cm | LH5HC | OC5HCIRC | ||

|
Temperature | temperatureTimeSlot_min | L6ATMPRD | OC6TINTV |
| esophagealTemperature_C | L6AESPHT | OC6TESOT | ||

|
Infection | positiveCultureNumber | OC9IPCNU | |
| positiveCultureDate | L6FPADT | OC9IDATE | ||
| positiveCultureTime | L6FPATM | OC9ITIME | ||

|
Neuro exam | post_NeuroExamPosture | LH11PO_1 | OC11CPOS |
| post_NeuroExamMoro | LH11MR_1 | OC11CPRM | ||

|
Imaging report | headSonogramResult1 | LH9HSREA | OC12HSRA |
| headCTResult1 | LH9HCREA | OC12HCRA | ||
| brainMRIResult1 | LH9BMREA | OC12BMRA | ||

|
MRI | MRIAvailable | LM1AVAI | OM1MRIA |
| MRINRNPatternOfInjury | LM3PTINJ | OM3PATINJ | ||

|
Status | Status | LH12STAT | OC13STAT |
| deathCause | LH12DTCA | OC13COD | ||

|
Bayley-III | BayleyIIIInEnglish | NF9ABSEN | NF9ABSEN |
| BayleyIIICognitiveComposite | NF9ABSCC | NF9ABSCC | ||

|
Outcome | moderateSevereDisabilityOrDeath | disab_die | disab_die |
| disabilityLevelDeath | N/A | N/A | ||
Subcategories in the leftmost column are examples from Figure 2. The original LH/OC variable names in the rightmost 2 columns were harmonized in the second left column. MRI, magnetic resonance imaging; N/A: not available; NRN, Neonatal Research Network.
Characteristics of the merged database
Figure 4 presents the distributions of 20 variables in the merged database. In the context of prognostic biomarker development, the selected 20 variables include variables on basic demographics and birth conditions, clinician’s evaluation of severity, and outcome measures.
Figure 4.
Distributions of 20 variables in the merged database from the LH and OC trials.
Utility of the merged database
Tables 3 and 4 highlight the results of the mass-univariate analyses. A total of 341 variables from Pre-intervention, Post-intervention, and NICU Discharge were analyzed using Pearson’s correlation coefficient analysis. These results demonstrate the utility of the merged database for the future development of prognostic biomarkers. For clarity, variables with definitions too similar to the listed ones, such as the scores from the subcategories of the modified Sarnat grading scales or specific regions described in the MRI reports, were excluded. The full results of all the 341 variables are in Supplementary Material S3. The color coding indicates the corresponding categories of the variables in Figure 2.
Table 3.
Association with disability level or death (normal, mild, moderate, severe, and death).
| Category | Variable | r | R 2 | P-value |
|---|---|---|---|---|
| DC | dischargeTotalModifiedSarnatScore | 0.722 | 0.522 | <10−74 |
| DC | dischargeFullNippleFeed | −0.585 | 0.342 | <10−31 |
| Post | MRINRNPatternOfInjuryAvg | 0.568 | 0.322 | <10−37 |
| Post | post_TotalModifiedSarnatScore | 0.557 | 0.310 | <10−35 |
| Pre | pre_TotalModifiedSarnatScore | 0.455 | 0.207 | <10−25 |
| Pre | encephalopathyLevel | 0.422 | 0.178 | <10−22 |
| DC | homeTherapyStatus | 0.391 | 0.153 | <10-11 |
| DC | dischargeAnticonvulsants | 0.375 | 0.141 | <10−16 |
| Pre | Apgar5min | −0.352 | 0.124 | <10−15 |
| Pre | Apgar10min | −0.346 | 0.120 | <10−12 |
| DC | dischargeHomeTherapyGastrostomyTubeFeed | 0.343 | 0.118 | <10-7 |
| DC | dischargeHearingTestNormal | −0.314 | 0.099 | <10-10 |
| Pre | firstPostnatalBloodGasPH | −0.303 | 0.092 | <10−10 |
| Post | post_BloodValueASTSGOT_UPerL | 0.297 | 0.088 | <10-5 |
| Pre | at10MinChestCompression | 0.289 | 0.084 | <10−8 |
| DC | dischargeEEGAbnormalBackgroundActivity | 0.285 | 0.081 | <10-8 |
| Pre | Apgar1min | −0.283 | 0.080 | <10−9 |
| Post | post_BloodValueALTSGPT_UPerL | 0.283 | 0.080 | <10-5 |
| DC | dischargeSeizure | 0.275 | 0.076 | <10−9 |
| DC | dischargeVentilator_day | 0.258 | 0.066 | <10−8 |
The abbreviations in the Category column represent the different chronological categories of the variables in Figure 2 (Pre: Pre-intervention, Post: Post-intervention, DC: NICU discharge). r: Pearson’s correlation coefficient; R2: coefficient of determination; P-value: P-value from each univariate Pearson’s correlation coefficient analysis. The P-value was adjusted using the Bonferroni correction, with the significance threshold set at P-value <10−5.
Table 4.
Association with the Bayley-III Composite Cognitive Scale.
| Category | Variable | r | R 2 | P-value |
|---|---|---|---|---|
| DC | dischargeHomeTherapyGastrostomyTubeFeed | −0.364 | 0.133 | <10−7 |
| Post | MRINRNPatternOfInjuryAvg | −0.342 | 0.117 | <10−11 |
| DC | dischargeTubeFeedingDuration_day | −0.338 | 0.114 | <10−11 |
| DC | dischargeTotalModifiedSarnatScore | −0.333 | 0.111 | <10−11 |
| Post | post_TotalModifiedSarnatScore | −0.297 | 0.088 | <10−8 |
| DC | homeTherapyStatus | −0.283 | 0.080 | <10-6 |
| DC | dischargeFullNippleFeed | 0.269 | 0.073 | <10−5 |
The color coding and heading are the same as Table 3.
As shown in Table 3, the total modified Sarnat grading scales from all 3 stages, MRI NRN Pattern of Injury derived from the Post-intervention stage, Apgar scores from the Pre-intervention stage, and several other variables were significantly associated with the primary outcome, defined as 5-level disability level or death (normal, mild, moderate, severe, and death).
Additionally, Table 4 displays correlations between the variables and the numerical Bayley Scales of Infant Development, Third Edition—Composite Cognitive Scale (Bayley–III Cognitive) (see Supplementary Material S3 for Bayley–III Language and Motor scales).21 The total modified Sarnat grading scales in the Post-intervention and NICU-discharge stages, the MRI NRN Pattern of Injury derived from Post-intervention stage, and feeding difficulty variables were among the top-ranked factors associated with Bayley–III Cognitive scales.
Discussion
The goal of COMBINE is to develop artificial intelligence (AI)-driven prognostic clinical and imaging biomarkers that can predict, during the neonatal stage, whether a specific HIE patient is at risk of developing adverse neurological outcomes by 2 years of age. Large-scale, comprehensive, diverse, and representative data are essential for developing quantitative, objective, and accurate prognostic biomarkers for HIE. When multiple trials or studies exist, a natural extension, as presented in this paper, is to harmonize and merge existing datasets. This approach facilitates the creation of a unified database with a larger sample size, covering not only more sites but also more studies, more comprehensive data elements, and increasing the diversity and heterogeneity of the data. This harmonized database can also enhance the generalizability of results to real-world applications.
Selecting representative datasets is an essential step before starting the construction of a data harmonization framework.16 The LH and OC trials are the follow-ups to the landmark trial that established therapeutic hypothermia as the current norm for HIE treatment.5 They aimed to further explore the potential of therapeutic hypothermia for a broader group of neonatal HIE patients who cannot be treated within the typical 6 hours of life (LH), as well as to optimize the intervention parameters (OC). As illustrated in Figure 1, the 21 sites reflect the geographic diversity of the population across the United States in these 2 studies. Furthermore, the 60 medical subcategories, encompassing a total of 1181 variables across the 5 major chronological categories, demonstrate the comprehensive considerations involved in conducting the HIE-related clinical trials, including the rigorous design of the trials, data definitions, and the training of neuro examiners, psychologists, and psychometricians. The representativeness and comprehensiveness of the LH and OC trials make them the ideal datasets to create the proposed harmonization framework.
An important component of our data harmonization framework is the creation of a dictionary of variables, as presented in Table 2. We recognized that while some variables may exist in only one trial, they remain crucial due to the uniqueness of each clinical study. Therefore, we retained a union of the variables from both trials. We resolved the differences in variable names by creating standardized data element names in this dictionary, followed by standardizing the coding of each variable across trials. This effort aligns with similar dedications to merge and harmonize heterogeneous datasets, such as medication in the intensive care units,22 cancer,23 stroke,24 pediatric sepsis,25 and SARS-CoV-2.26,27 The creation of common data elements (CDE) or standardized dictionaries for various diseases is also a key NIH effort.28
The harmonized dictionary of HIE variables we created in this paper can be used in at least 2 scenarios. First, it can serve as a reference to merge additional existing HIE datasets, further expanding our unified database. Potential large-scale datasets include the BONBID-HIE dataset (237 patients from 2 sites and 20+ variables),29 the HEAL-HIE data (500 patients from 17 sites),30 etc. These datasets may offer variables not present in the LH and OC trials, so the dictionary will be continuously expanded beyond the current 1181 variables as new datasets are merged. In addition, the dictionary of HIE variables can guide the future design of HIE clinical trials. By using the same variable names, definitions, and coding conventions in the design phase, newly acquired data can be easily merged into the dynamically expanding database.
Currently, there is no widely accepted standard dictionary for HIE clinical studies. A parallel work by Newborn Brain Society Guidelines and Publications Committee found that only 4 variables (birth weight, gestational age, 1- and 5-minute Apgar scores) were common across 22 HIE registries, with inconsistent naming and coding, highlighting the urgent need for harmonization.4 Our dictionary (Figure 2, Table 2, and Supplementary Material S1, S2, and S4) aligns with the National Institute of Health’s Common Data Elements (CDE) Initiative, which supports data standardization to facilitate cross-study comparison and data sharing.28 Integrating into the NIH CDE Repository will promote the interoperability and broad adoption of the framework. In addition, several frameworks and applications, such as the Observational Medical Outcomes Partnership (OMOP) Common Data Model (CDM) by Observational Health Data Sciences and Informatics (OHDSI),31 can be integrated with electronic healthcare records (EHR) systems, such as Epic’s analytic database storing as a Microsoft SQL database (Clarity) or a cloud-based database (Snowflake),32–34 to extract EHR and transform them into harmonized variables for further utilization of the integrated data. It is anticipated that our data harmonization framework will yield real-world impact through collaboration with relevant stakeholders and seamless integration with their applications.
In the literature, there are studies exploring the ability of clinical variables to predict HIE outcomes. Broadly speaking, our results (Figure 4; Tables 3 and 4) are consistent with previous reports on neuroimaging scores,1,11,13,35–38 Sarnat grading scales,39–46 and Apgar scores at 5 and 10 minutes after birth,47 in that the associations between these variables and the primary outcome (disability level or death) are significant.
We envision several directions for future exploration. First, other HIE trials may include additional variables, such as fine-grained electrocardiogram (ECG) and electroencephalogram (EEG) monitoring, different timing of assessments or different expert interpretations of neuroimaging. We will continuously expand our dictionary as we merge new datasets into our database. In addition, we conducted only a basic mass-univariate analysis, as the primary goal of this paper is not to perform an extensive analysis but to present the utility of the merged database. Our ultimate objective in the COMBINE consortium is to develop multivariate AI analyses that can thoroughly examine the combinatorial effects of over 1000 variables and enhance the predictive value for each individual patient. Furthermore, our data harmonization framework has the potential to extend beyond biomarker development. It can also function as a knowledge base or support an HIE-specific chatbot to further enhance clinical training and practice.
Conclusion
In conclusion, we have established a data harmonization framework tailored for HIE-related projects. Utilizing this framework, we successfully combined data from 2 HIE clinical trials from the NICHD NRN into a cohesive database. The resulting dictionary, which contains 1181 variables, serves as a foundation for integrating additional existing HIE datasets and recommending common data elements for future HIE trials or studies. This framework and the merged database will facilitate the subsequent development of prognostic HIE biomarkers.
Supplementary Material
Acknowledgments
The authors express immense gratitude to Dr Henry A. Feldman from Boston Children’s Hospital for his profound expertise in statistics and his patience during their discussions.
The COMBINE consortium includes the following sites and researchers. Investigators were arranged by affiliations at the time of OC and LH trials, to acknowledge data collection in these trials, with additional parentheses of their current affiliations.
- Boston Children’s Hospital (P. Ellen Grant, MD, MSc; Yangming Ou, PhD; Janet S. Soul, MD);
- Brown University and Women & Infants Hospital of Rhode Island (Abbot R. Laptook, MD);
- Case Western Reserve University and Rainbow Babies and Children’s Hospital (Michele C. Walsh, MD);
- Children’s Hospital of Philadelphia (Eric C. Eichenwald, MD);
- Children’s Mercy Hospital Kansas City (William E. Truog, MD);
- Cincinnati Children’s Hospital Medical Center (Stephanie L. Merhar, MD; Brenda L.B. Poindexter, MD, MS [now at Emory University and Children’s Healthcare of Atlanta]; Lili He, PhD);
- Duke University (C. Michael Cotten, MD);
- Emory University (Shannon E.G. Hamrick, MD [now at Food and Drug Administration]);
- Indiana University Medical Center (Gregory M. Sokol, MD);
- Massachusetts General Hospital (Sara V. Bates, MD);
- Nationwide Children’s Hospital, Abigail Wexner Research Institute at Nationwide Children’s Hospital, and The Ohio State University College of Medicine (Pablo J. Sanchez, MD; Edward G. Shepherd, MD; Mai-Lan Ho, MD [now at the University of Missouri Health Care]);
- RTI International (Jeanette O. Auman);
- Stanford University (Krisa P. Van Meurs, MD; Susan R. Hintz, MD; Sonia L. Bonifacio, MD);
- Tufts Medical Center (Ivan D. Frantz, III, MD);
- University of Alabama at Birmingham (Namasivayam Ambalavanan, MD);
- University of Iowa (Edward F. Bell, MD; Patrick McNamara, MD);
- University of New Mexico (Kristi Watterberg, MD);
- University of Rochester (Carl T. D’Angio, MD);
- University of Texas Health Science Center at Houston (Jon E. Tyson, MD; Charles E. Green, PhD);
- University of Texas Southwestern Medical Center at Dallas (Lina F. Chalak, MD);
- University of Utah (Bradley A. Yoder, MD);
- Wayne State University (Seetha Shankaran, MD; Sanjay Chawla, MD).
The National Institutes of Health, the Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD), and the National Center for Advancing Translational Sciences provided grant support for the Neonatal Research Network’s Optimizing Cooling and Late Hypothermia Trials through cooperative agreements. While NICHD staff had input into the study design, conduct, analysis, and manuscript drafting, the comments and views of the authors do not necessarily represent the views of NICHD, the National Institutes of Health, the Department of Health and Human Services, or the US Government.
Participating NRN sites collected data and transmitted it to RTI International, the data coordinating center for the network, which stored, managed, and analyzed the data for this study. On behalf of the NRN, RTI International had full access to all data in the study, and with the NRN Center Principal Investigators, takes responsibility for the integrity of the data and accuracy of the data analysis.
We are indebted to our medical and nursing colleagues and the infants and their parents who agreed to take part in this study. The following investigators, in addition to those listed as authors, participated in this study:
NRN Steering Committee Chairs: Michael S. Caplan, MD, University of Chicago, Pritzker School of Medicine (2006-2011); Richard A. Polin, MD, Division of Neonatology, College of Physicians and Surgeons, Columbia University (2011-2023).
Alpert Medical School of Brown University and Women & Infants Hospital of Rhode Island (U10 HD27904): Martin Keszler, MD; William Oh, MD; Betty R. Vohr, MD; Angelita M. Hensman, PhD, RNC-NIC; Barbara Alksninis, RNC, PNP; Kristin Basso, MaT, RN; Carmena Bishop; Joseph Bliss, MD, PhD; Robert T. Burke, MD, MPH; William Cashore, MD; Melinda Caskey, MD; Dan Gingras, RRT; Nicholas Guerina, MD, PhD; Katharine Johnson, MD; Mary Lenore Keszler, MD; Andrea M. Knoll; Theresa M. Leach, MEd, CAES; Martha R. Leonard, BA, BS; Emilee Little, RN, BSN; Elizabeth C. McGowan, MD; Leslie T. McKinley, MS, RD; Hussnain Mirza, MD; Birju A. Shah, MD, MPH; Ross Sommers, MD; Bonnie E. Stephens, MD; Suzy Ventura; Elisa Vieira, RN, BSN; Victoria E. Watson, MS, CAS.
Case Western Reserve University, Rainbow Babies & Children’s Hospital (U10 HD21364, M01 RR80): Anna Maria Hibbs, MD; Deanne E. Wilson-Costello, MD; Michele C. Walsh, MD, MS; Elizabeth Roth, PhD; Nancy S. Newman, RN; Monika Bhola, MD; Bonnie S. Siner, RN; Eileen K. Stork, MD; Gulgun Yalcinkaya, MD.
Children’s Mercy Hospital and University of Missouri Kansas City School of Medicine (U10 HD68284): William E. Truog, MD; Eugenia K. Pallotto, MD, MSCE; Howard W. Kilbride, MD; Cheri Gauldin, RN, BSN, CCRC; Anne Holmes, RN, MSN, MBA-HCM, CCRC; Kathy Johnson, RN, CCRC; Allison Knutson, BSN, RNC-NIC.
Cincinnati Children’s Hospital Medical Center, University of Cincinnati Medical Center, and Good Samaritan Hospital (U10 HD27853, UL1 TR77): Stephanie Merhar, MD, MS; Kurt Schibler, MD; Brenda B. Poindexter, MD, MS; Suhas G. Kallapur, MD; Teresa L. Gratton, PA; Cathy Grisby, BSN, CCRC; Barbara Alexander, RN; Estelle E. Fischer, HSA, MBA; Jody Hessling, MSN, RN; Lenora D. Jackson, CRC; Jennifer Jennings, RN, BSN; Kristin Kirker, CRC; Greg Muthig, BA; Sandra Wuertz, RN, BSN, CLC; Kimberly Yolton, PhD.
Duke University School of Medicine, Duke University Hospital, and University of North Carolina at Chapel Hill (U10 HD40492, UL1 TR1117): Ronald N. Goldberg, MD; C. Michael Cotten, MD; Ricki F. Goldstein, MD; William F. Malcolm, MD; Joanne Propst, RN, JD; Patricia L. Ashley, MD, PhD; Kimberley A. Fisher, PhD, FNP-BC, IBCLC; Sandra Grimes, RN, BSN; Kathryn E. Gustafson, PhD; Melody B. Lohmeyer, RN, MSN; Deesha Mago-Shah, MD; Mollie Warren, MD; Matthew M. Laughon, MD, MPH; Carl L. Bose, MD; Janice Bernhardt, MS, RN; Cynthia L. Clark, RN; Diane D. Warner, MD, MPH; Janice K. Wereszcsak, CPNP-AC/PC.
Emory University, Children’s Healthcare of Atlanta, Grady Memorial Hospital, and Emory University Hospital Midtown (U10 HD27851, UL1 TR454): Barbara J. Stoll, MD; David P. Carlton, MD; Ira Adams-Chapman, MD (deceased); Yvonne Loggins, RN; Ellen C. Hale, RN, BS, CCRC; Diane I. Bottcher, MSN, RN; Sheena L. Carter, PhD; Shannon E.G. Hamrick, MD; Colleen Mackie, BS, RT; Maureen Mulligan LaRossa, RN; Lynn C. Wineski, RN, MS.
Eunice Kennedy Shriver National Institute of Child Health and Human Development: Rosemary D. Higgins, MD; Stephanie Wilson Archer, MA.
Indiana University, University Hospital, Methodist Hospital, Riley Hospital for Children at Indiana University Health, and Eskenazi Health (U10 HD27856, UL1 TR6): Gregory M. Sokol, MD; Brenda B. Poindexter, MD, MS; Anna M. Dusick, MD (deceased); Lu-Ann Papile, MD; Heidi M. Harmon, MD, MS; Dianne E. Herron, RN, CCRC; Jessica Bissey, PsyD, HSPP; Lon G. Bohnke, MS; Ann B. Cook, MS; Susan Gunn, NNP-BC, CCRC; Abbey C. Hines, PsyD; Darlene Kardatzke, MD (deceased); Carolyn Lytle, MD, MPH; Heike M. Minnich, PsyD, HSPP; Leslie Richard, RN; Lucy C. Smiley, CCRC; Leslie Dawn Wilson, BSN, CCRC.
McGovern Medical School at The University of Texas Health Science Center at Houston, Children’s Memorial Hermann Hospital (U10 HD21373): Jon E. Tyson, MD, MPH; Amir M. Khan, MD; Kathleen A. Kennedy, MD, MPH; Andrea F. Duncan, MD, MSClinRes; Georgia E. McDavid, RN; Elizabeth Allain, PhD; Julie Arldt-McAlister, MSN, APRN; Katrina Burson, RN, BSN; Allison G. Dempsey, PhD; Patricia W. Evans, MD; Carmen Garcia, RN, BSN; Charles Green, PhD; Margarita Jimenez, MD, MPH; Janice John, CPNP; Patrick M. Jones, MD, MA; M. Layne Lillie, RN, BSN; Terri Major-Kincade, MD, MPH; Karen Martin, RN; Sara C. Martin, RN, BSN; Shannon McKee, EdS; Claudia Pedroza, PhD; Patti L. Pierce Tate, RCP; Kimberly Rennie, PhD; Shawna Rodgers, RNC-NIC, BSN; Saba Khan Siddiki, MD; Daniel K. Sperry, RN; Sharon L. Wright, MT (ASCP).
Nationwide Children’s Hospital and The Ohio State University Wexner Medical Center (U10 HD68278): Pablo J. Sánchez, MD; Leif D. Nelin, MD; Jonathan Slaughter, MD, MPH; Sudarshan R. Jadcherla, MD; Patricia Luzader, RN; Roopali Bapat, MD; Thomas Bartman, MD; Elizabeth Bonachea, MD; Louis G. Chicoine, MD; Bronte Clifford; Marliese Dion Nist, BSN; Erin Ferns; Christine A. Fortney, PhD, RN; Jennifer Fuller, MS, RNC; Ish Gulati, MD; Julie Gutentag, BSN; Krista Haines, MD; Brandon Hart, MD; Michael Hokenson, MD; Marissa E. Jones, RN, MBA; Sarah McGregor, BSN, RNC; Nehal A. Parikh, MD; Elizabeth Ann Rodgers, BSN; Ruth Seabrook, MD; Tiffany Sharp; Edward G. Shepherd, MD; Jodi A. Ulloa, MSN, APRN NNP-BC; Jon Wispe, MD; Tara Wolfe, BSN; L. Yossef, MD; Nahla Zaghoul, MD.
RTI International (U10 HD36790): Abhik Das, PhD; Marie G. Gantz, PhD; Dennis Wallace, PhD; Kristin M. Zaterka-Baxter, RN, BSN, CCRP; Margaret M. Crawford, BS, CCRP; Jenna Gabrio, BS, CCRP; Breda Munoz, PhD; Jamie E. Newman, PhD, MPH; Carolyn M. Petrie Huitema, MS, CCRP; James W. Pickett II, BS.
Stanford University and Lucile Packard Children’s Hospital (U10 HD27880, M01 RR70, UL1 TR93): Valerie Y. Chock, MD, MS Epi; Krisa P. Van Meurs, MD; David K. Stevenson, MD; Susan R. Hintz, MD, MS Epi; M. Bethany Ball, BS, CCRC; Elizabeth F. Bruno, PhD; Alexis S. Davis, MD, MS Epi; Maria Elena DeAnda, PhD; Anne M. DeBattista, RN, PNP, PhD; Lynne C. Huffman, MD; Casey E. Krueger, PhD; Melinda S. Proud, RCP; Nicholas H. St John, PhD; Hali E. Weiss, MD.
Tufts Medical Center, Floating Hospital for Children (U10 HD53119, M01 RR54): Ivan D. Frantz III, MD; John M. Fiascone, MD; Elisabeth C. McGowan, MD; Brenda L. MacKinnon, RNC; Ana Brussa, MS, OTR/L; Anne Furey, MPH; Brian Gilchrist, MD; Juliette C. Madan, MD, MS; Ellen Nylen, RN, BSN; Cecelia Sibley, PT, MHA.
University of Alabama at Birmingham Health System and Children’s Hospital of Alabama (U10 HD34216, M01 RR32): Waldemar A. Carlo, MD; Namasivayam Ambalavanan, MD; Myriam Peralta-Carcelen, MD, MPH; Monica V. Collins, RN, BSN, MaEd; Shirley S. Cosby, RN, BSN; Vivien A. Phillips, RN, BSN; Richard V. Rector, PhD; Sally Whitley, MA, OTR-L, FAOTA.
University of California—Los Angeles, Mattel Children’s Hospital, Santa Monica Hospital, Los Robles Hospital and Medical Center, and Olive View Medical Center (U10 HD68270): Uday Devaskar, MD; Meena Garg, MD; Isabell B. Purdy, PhD, CPNP; Teresa Chanlaw, MPH; Rachel Geller, RN, BSN.
University of Iowa and Mercy Medical Center (U10 HD53109, UL1 TR442): Tarah T. Colaizy, MD, MPH; Edward F. Bell, MD; Jane E. Brumbaugh, MD; Michael J. Acarregui, MD, MBA; Karen J. Johnson, RN, BSN; Vipinchandra Bhavsar, MB, BS; John M. Dagle, MD, PhD; Diane L. Eastman, RN, CPNP, MA; Jonathan M. Klein, MD; Nancy J. Krutzfield, RN, MA; Claire A. Lindauer, RN; Julie B. Lindower, MD, MPH; Steven J. McElroy, MD; Lauritz R. Meyer, MD; Glenda K. Rabe, MD, MME; Robert D. Roghair, MD; Jeffrey L. Segar, MD; Jacky R. Walker, RN; Dan L. Ellsbury, MD; Donia B. Campbell, RNC-NIC; Cary R. Murphy, MD.
University of New Mexico Health Sciences Center (U10 HD53089, UL1 TR41): Kristi L. Watterberg, MD; Robin K. Ohls, MD; Janell Fuller, MD; Jean R. Lowe, PhD; Conra Backstrom Lacy, RN; Sandra Sundquist Beauman, MSN, RNC; Andrea F. Duncan, MD, MSClin.
University of Pennsylvania, Hospital of the University of Pennsylvania, Pennsylvania Hospital, and Children’s Hospital of Philadelphia (U10 HD68244): Sara B. DeMauro, MD, MSCE; Eric C. Eichenwald, MD; Barbara Schmidt, MD, MSc; Haresh Kirpalani, MB, MSc; Antoinette Mancini, RN, BSN, CCRC; Soraya Abbasi, MD; Judy C. Bernbaum, MD; Aasma S. Chaudhary, BS, RRT; Dara M. Cucinotta, RN; Kevin C. Dysart, MD; Marsha Gerdes, PhD; Hallam Hurt, MD.
University of Rochester Medical Center, Golisano Children’s Hospital, and the State University New York at Buffalo Women’s and Children’s Hospital of Buffalo (U10 HD68263, UL1 TR42): Carl T. D’Angio, MD; Ronnie Guillet, MD, PhD; Gary J. Myers, MD; Melissa Bowman, MSN; Patrick Conway, MS; Osman Farooq, MD; Rosemary L. Jensen; Nirupama Laroia, MD; Joan Merzbach, LMSW; Ann Marie Scorsone, MS; Holly I.M. Wadkins, MA; Kelley Yost, PhD; Anne Marie Reynolds, MD, MPH; Satyan Lakshminrusimha, MD; Ashley Williams, MS, Ed; Stephanie Guilford, BS; Michael G. Sacilowski, MAT; Karen Wynn, NNP, RN; William Zorn, PhD; Michele Hartley-McAndrew, MD; Constance Orme; Cait Fallone, MA; Kyle Binion, BS.
University of Texas Southwestern Medical Center at Dallas, Parkland Health & Hospital System, and Children’s Medical Center Dallas (U10 HD40689, M01 RR633): Myra Wyckoff, MD; Pablo J. Sánchez, MD; Luc P. Brion, MD; Roy J. Heyne, MD; Diana M. Vasil, MSN, BSN, RNC-NIC; Sally S. Adams, MS, RN, CPNP; Lijun Chen, PhD, RN; Alicia Guzman; Elizabeth T. Heyne, MS, MA, PA-C, PsyD; Lizette E. Lee, RN; Melissa H. Leps, RN; Linda A. Madden, BSN, RN, CPNP; Nancy A. Miller, RN; Janet S. Morgan, RN; Emma Ramon, RNC-NIC, RN, BSN; Catherine Twell Boatman, MS, CIMI.
University of Utah Medical Center, Intermountain Medical Center, LDS Hospital, and Primary Children’s Medical Center (U10 HD53124): Bradley A. Yoder, MD; Roger G. Faix, MD; Sarah Winter, MD; Shawna Baker, RN; Karie Bird, RN, BSN; Anna Bodnar, MD; Jill Burnett, RNC, BSN; Cynthia Spencer, RNC; R. Edison Steele, RN; Mike Steffen, PhD; Karena Strong, RN, BSN; Kimberlee Weaver-Lewis, RN, BSN; Karen Osborne, RN, BSN, CCRC; Karen Zanetti, RN; Laura Cole Bledsoe, RN.
Wayne State University, University of Michigan, Hutzel Women’s Hospital, Children’s Hospital of Michigan and Mott Children’s Hospital (U10 HD21385): Girija Natarajan, MD; Beena G. Sood, MD, MS; Athina Pappas, MD; Rebecca Bara, RN, BSN; Monika Bajaj, MD; Sanjay Chawla, MD; Lilia C. De Jesus, MD; Melissa February MD; Prashant Agarwal MD; Laura A. Goldston, MA; Eunice Hinz Woldt, RN, MSN; Mary E. Johnson, RN, BSN; John Barks, MD; Mary Christensen, RT; Stephanie A. Wiggins, MS; Martha Carlson, MD; Diane F. White, RRT, CCRP.
Yale-New Haven Children’s Hospital (U10 HD27871, UL1 TR142): Richard A. Ehrenkranz, MD (deceased); Matthew Bizzarro, MD; Monica Konstantino, RN, BSN; Nancy Close, PhD; JoAnn Poulsen, RN; Elaine Romano, MSN; Janet Taft, RN, BSN.
Contributor Information
Chuan-Heng Hsiao, Fetal-Neonatal Neuroimaging Developmental Science Center, Division of Newborn Medicine, Department of Pediatrics, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, United States.
Anna N Foster, Fetal-Neonatal Neuroimaging Developmental Science Center, Division of Newborn Medicine, Department of Pediatrics, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, United States.
Scott A McDonald, RTI International, Research Triangle Park, NC 27709, United States.
Rutvi Vyas, Fetal-Neonatal Neuroimaging Developmental Science Center, Division of Newborn Medicine, Department of Pediatrics, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, United States.
Aseelah Ashraf, Fetal-Neonatal Neuroimaging Developmental Science Center, Division of Newborn Medicine, Department of Pediatrics, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, United States.
Rina Bao, Fetal-Neonatal Neuroimaging Developmental Science Center, Division of Newborn Medicine, Department of Pediatrics, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, United States.
Lena Tran, Fetal-Neonatal Neuroimaging Developmental Science Center, Division of Newborn Medicine, Department of Pediatrics, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, United States.
Ankush Kesri, Fetal-Neonatal Neuroimaging Developmental Science Center, Division of Newborn Medicine, Department of Pediatrics, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, United States.
Erfan Darzidehkalani, Fetal-Neonatal Neuroimaging Developmental Science Center, Division of Newborn Medicine, Department of Pediatrics, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, United States.
Matheus D Soldatelli, Fetal-Neonatal Neuroimaging Developmental Science Center, Division of Newborn Medicine, Department of Pediatrics, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, United States; Department of Radiology, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, United States.
Jeanette O Auman, RTI International, Research Triangle Park, NC 27709, United States.
Janet S Soul, Department of Neurology, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, United States.
Lina F Chalak, Division of Neonatal-Perinatal Medicine, Department of Pediatrics, Department of Psychiatry, University of Texas Southwestern Medical Center at Dallas, Dallas, TX 75390, United States.
C Michael Cotten, Department of Pediatrics, Duke University School of Medicine, Durham, NC 27710, United States.
Seetha Shankaran, Department of Neonatal-Perinatal Medicine, Wayne State University, Detroit, MI 48202, United States; Department of Pediatrics, University of Texas at Austin Dell Medical School, Austin, TX 78712, United States.
Abbot R Laptook, The Warren Alpert Medical School, Brown University, Providence, RI 02903, United States; Department of Pediatrics, Women and Infants Hospital of Rhode Island, Providence, RI 02905, United States.
P Ellen Grant, Fetal-Neonatal Neuroimaging Developmental Science Center, Division of Newborn Medicine, Department of Pediatrics, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, United States; Department of Radiology, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, United States.
Yangming Ou, Fetal-Neonatal Neuroimaging Developmental Science Center, Division of Newborn Medicine, Department of Pediatrics, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, United States; Department of Radiology, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115, United States.
Consortium Of MRI Biomarkers In Neonatal Encephalopathy (COMBINE):
P Ellen Grant, III, Yangming Ou, Janet S Soul, Abbot R Laptook, Michele C Walsh, Eric C Eichenwald, William E Truog, Stephanie L Merhar, Brenda L B Poindexter, Lili He, C Michael Cotten, Shannon E G Hamrick, Gregory M Sokol, Sara V Bates, Pablo J Sanchez, Edward G Shepherd, Mai-Lan Ho, Jeanette O Auman, Krisa P Van Meurs, Susan R Hintz, Sonia L Bonifacio, Ivan D Frantz, Namasivayam Ambalavanan, Edward F Bell, Patrick McNamara, Kristi Watterberg, Carl T D’Angio, Jon E Tyson, Charles E Green, Lina F Chalak, Bradley A Yoder, Seetha Shankaran, and Sanjay Chawla
Author contributions
Chuan-Heng Hsiao (Data curation, Formal analysis, Software, Validation, Visualization, Writing—original draft), Anna N. Foster (Data curation, Writing—review & editing), Scott A. McDonald (Data curation, Writing—review & editing), Rutvi Vyas (Data curation, Visualization, Writing—review & editing), Aseelah Ashraf (Data curation, Writing—review & editing), Rina Bao (Data curation, Writing—review & editing), Lena Tran (Data curation, Writing—review & editing), Ankush Kesri (Data curation, Writing—review & editing), Erfan Darzidehkalani (Data curation, Writing—review & editing), Matheus D. Soldatelli (Data curation, Writing—review & editing), Jeanette O. Auman (Data curation, Writing—review & editing), Janet S. Soul (Supervision, Writing—review & editing), Lina F. Chalak (Supervision, Writing—review & editing), C. Michael Cotten (Conceptualization, Funding acquisition, Methodology, Supervision, Writing—review & editing), Seetha Shankaran (Conceptualization, Funding acquisition, Methodology, Supervision, Writing—review & editing), Abbot Laptook (Conceptualization, Funding acquisition, Methodology, Supervision, Writing—review & editing), P. Ellen Grant (Conceptualization, Funding acquisition, Methodology, Supervision, Writing—review & editing), and Yangming Ou (Conceptualization, Funding acquisition, Methodology, Supervision, Writing—review & editing)
Supplemental material
Supplementary materials are available at JAMIA Open online.
Funding
This work was supported, in part, by the National Institute of Health by grant R61NS126792.
Conflicts of interest
All authors declare that they have no financial or non-financial competing interests in relation to this paper.
Data availability
The definition of the metadata and the data dictionary of the framework is maintained as spreadsheets and REDCap format and can be found in Supplementary Material S1, S2, and S4.48 With the separation of Datasets, Software, and Results, the Results, including the definition of the metadata and the data dictionary of the framework, and the statistical analysis results, are licensed under CC BY 4.0 and can be found at the public repository https://github.com/i3-research/COMBINE-harmonizer/tree/v1.1.0/results. The LH dataset and the OC dataset have been released to the public.49,50
Code availability
With the separation of Datasets, Software, and Results, the Software for this study is available on GitHub under the MIT License and can be accessed via this link https://www.github.com/i3-research/COMBINE-harmonizer/tree/v1.1.0.
References
- 1. Shankaran S. Therapeutic hypothermia for neonatal encephalopathy. Curr Opin Pediatr. 2015;27:152-157. 10.1097/MOP.0000000000000199 [DOI] [PubMed] [Google Scholar]
- 2. Namusoke H, Nannyonga MM, Ssebunya R, et al. Incidence and short term outcomes of neonates with hypoxic ischemic encephalopathy in a peri urban teaching hospital, Uganda: a prospective cohort study. Matern Health Neonatol Perinatol. 2018;4:6. 10.1186/s40748-018-0074-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Park J, Park SH, Kim C, et al. Growth and developmental outcomes of infants with hypoxic ischemic encephalopathy. Sci Rep. 2023;13:23100. 10.1038/s41598-023-50187-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Peeples ES, Mietzsch U, Molloy E, et al. ; Newborn Brain Society Guidelines and Publications Committee. Data collection variability across neonatal hypoxic-ischemic encephalopathy registries. J Pediatr. 2025;279:114476. 10.1016/j.jpeds.2025.114476 [DOI] [PubMed] [Google Scholar]
- 5. Shankaran S, Laptook AR, Ehrenkranz RA, et al. ; National Institute of Child Health and Human Development Neonatal Research Network. Whole-body hypothermia for neonates with hypoxic–ischemic encephalopathy. New Engl J Med. 2005;353:1574-1584. 10.1056/NEJMcps050929 [DOI] [PubMed] [Google Scholar]
- 6. Azzopardi DV, Strohm B, Edwards AD, et al. ; TOBY Study Group. Moderate hypothermia to treat perinatal asphyxial encephalopathy. New Engl J Med. 2009;361:1349-1358. 10.1056/NEJMoa0900854 [DOI] [PubMed] [Google Scholar]
- 7. Jacobs SE, Morley CJ, Inder TE, et al. ; Infant Cooling Evaluation Collaboration. Whole-body hypothermia for term and near-term newborns with hypoxic-ischemic encephalopathy. Arch Pediatr Adolesc Med. 2011;165:692-700. 10.1001/archpediatrics.2011.43 [DOI] [PubMed] [Google Scholar]
- 8. Groenendaal F, Casaer A, Dijkman KP, et al. Introduction of hypothermia for neonates with perinatal asphyxia in The Netherlands and Flanders. Neonatology. 2013;104:15-21. 10.1159/000348823 [DOI] [PubMed] [Google Scholar]
- 9. Douglas-Escobar M, Weiss MD. Hypoxic-ischemic encephalopathy: a review for the clinician. JAMA Pediatr. 2015;169:397-403. 10.1001/jamapediatrics.2014.3269 [DOI] [PubMed] [Google Scholar]
- 10. Shankaran S, Laptook AR, Pappas A, et al. ; Eunice Kennedy Shriver National Institute of Child Health and Human Development Neonatal Research Network. Effect of depth and duration of cooling on death or disability at age 18 months among neonates with hypoxic-ischemic encephalopathy—a randomized clinical trial. JAMA. 2017;318:57-67. 10.1001/jama.2017.7218 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Rutherford M, Ramenghi LA, Edwards AD, et al. Assessment of brain tissue injury after moderate hypothermia in neonates with hypoxic–ischaemic encephalopathy: a nested substudy of a randomised controlled trial. Lancet Neurol. 2010;9:39-45. 10.1016/S1474-4422(09)70295-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Shankaran S, Barnes PD, Hintz SR, et al. ; Eunice Kennedy Shriver National Institute of Child Health and Human Development Neonatal Research Network. Brain injury following trial of hypothermia for neonatal hypoxic–ischaemic encephalopathy. Arch Dis Child Fetal Neonatal Ed. 2012;97:F398-F404. 10.1136/archdischild-2011-301524 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Weeke LC, Groenendaal F, Mudigonda K, et al. A novel magnetic resonance imaging score predicts neurodevelopmental outcome after perinatal asphyxia and therapeutic hypothermia. J Pediatr. 2018;192:33-40.e2. 10.1016/j.jpeds.2017.09.043 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Weiss RJ, Bates SV, Song Y, et al. Mining multi-site clinical data to develop machine learning MRI biomarkers: application to neonatal hypoxic ischemic encephalopathy. J Transl Med. 2019;17:385. 10.1186/s12967-019-2119-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Laptook AR, Shankaran S, Tyson JE, et al. ; Eunice Kennedy Shriver National Institute of Child Health and Human Development Neonatal Research Network. Effect of therapeutic hypothermia initiated after 6 hours of age on death or disability among newborns with hypoxic-ischemic encephalopathy—a randomized clinical trial. JAMA. 2017;318:1550-1560. 10.1001/jama.2017.14972 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Fortier I, Raina P, Van den Heuvel ER, et al. Maelstrom research guidelines for rigorous retrospective data harmonization. Int J Epidemiol. 2017;46:103-105. 10.1093/ije/dyw075 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Cheng C, Messerschmidt L, Bravo I, et al. A general primer for data harmonization. Sci Data. 2024;11:152. 10.1038/s41597-024-02956-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Stuckenschmidt H. Ontology-Based Information Sharing in Weakly Structured Environments. PhD Dissertation. Vrije Universiteit Amsterdam; 2003.
- 19. Wilkinson MD, Dumontier M, Aalbersberg I, et al. The FAIR guiding principles for scientific data management and stewardship. Sci Data. 2016;3:160018. 10.1038/sdata.2016.18 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Preston-Werner T. Semantic versioning. Accessed September 25, 2024. https://semver.org/
- 21. Bayley N. Bayley Scales of Infant and Toddler Development. 3rd ed. San Antonio, TX: Harcourt Assessment; 2005. [Google Scholar]
- 22. Sikora A, Keats K, Murphy DJ, et al. A common data model for the standardization of intensive care unit medication features. JAMIA Open. 2024;7:ooae033. 10.1093/jamiaopen/ooae033 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Rolland B, Reid S, Stelling D, et al. Toward rigorous data harmonization in cancer epidemiology research: one approach. Am J Epidemiol. 2015;182:1033-1038. 10.1093/aje/kwv133 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Grinnon ST, Miller K, Marler JR, et al. National Institute of Neurological Disorders and stroke common data element project—approach and methods. Clin Trials. 2012;9:322-329. 10.1177/1740774512438980 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Mawji A, Li E, Chandna A, et al. Common data elements for predictors of pediatric sepsis: a framework to standardize data collection. PLoS One. 2021;16:e0253051. 10.1371/journal.pone.0253051 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Rinaldi E, Stellmach C, Rajkumar NMR, et al. Harmonization and standardization of data for a pan-European cohort on SARS-CoV-2 pandemic. NPJ Digit Med. 2022;5:75. 10.1038/s41746-022-00620-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Dolin G, Saitwal H, Bertodatti K, et al. Establishing data elements and exchange standards to support long COVID healthcare and research. JAMIA Open. 2024;7:ooae095. 10.1093/jamiaopen/ooae095 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Rubinstein YR, McInnes P. NIH/NCATS/GRDR® common data elements: a leading force for standardized data collection. Contemp Clin Trials. 2015;42:78-80. 10.1016/j.cct.2015.03.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Bao R, Song Y, Bates SV, et al. BOston Neonatal Brain Injury Data for Hypoxic Ischemic Encephalopathy (BONBID-HIE): I. MRI and lesion labeling. Sci Data. 2025;12:53. 10.1038/s41597-024-03986-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Wu YW, Comstock BA, Gonzalez FF, et al. ; HEAL Consortium. Trial of erythropoietin for hypoxic–ischemic encephalopathy in newborns. New Engl J Med. 2022;387:148-159. 10.1056/NEJMoa2119660 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. George H, Duke Jon D, Shah Nigam H, et al. Observational Health Data Sciences and Informatics (OHDSI): opportunities for observational researchers. In: Sarkar IN, Georgiou A, de Azevedo Marques PM, eds. MEDINFO 2015: eHealth-enabled Health. Studies in Health Technology and Informatic. Vol. 216. IOS Press; 2015;574-578. 10.3233/978-1-61499-564-7-574 [DOI] [PMC free article] [PubMed]
- 32. Epic Systems Corporation. Epic. Accessed May 6, 2025. https://www.epic.com/
- 33. Chishtie J, Sapiro N, Wiebe N, et al. Use of epic electronic health record system for health care research: scoping review. J Med Internet Res 2023; 25: e51003. 10.2196/51003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Snowflake Corporation. Snowflake. Accessed May 6, 2025. https://www.snowflake.com/
- 35. Barkovich AJ, Hajnal BL, Vigneron D, et al. Prediction of neuromotor outcome in perinatal asphyxia: evaluation of MR scoring systems. AJNR Am J Neuroradiol. 1998;19:143-149. [PMC free article] [PubMed] [Google Scholar]
- 36. Trivedi SB, Vesoulis ZA, Rao R, et al. A validated clinical MRI injury scoring system in neonatal hypoxic-ischemic encephalopathy. Pediatr Radiol. 2017;47:1491-1499. 10.1007/s00247-017-3893-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Al Amrani F, Marcovitz J, Sanon P-N, et al. Prediction of outcome in asphyxiated newborns treated with hypothermia: is a MRI scoring system described before the cooling era still useful? Eur J Paediatr Neurol. 2018;22:387-395. 10.1016/j.ejpn.2018.01.017 [DOI] [PubMed] [Google Scholar]
- 38. Wu YW, Monsell SE, Glass HC, et al. How well does neonatal neuroimaging correlate with neurodevelopmental outcomes in infants with hypoxic-ischemic encephalopathy? Pediatr Res. 2023;94:1018-1025. 10.1038/s41390-023-02510-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Sarnat HB, Sarnat MS. Neonatal encephalopathy following fetal distress: a clinical and electroencephalographic study. Arch Neurol. 1976;33:696-705. 10.1001/archneur.1976.00500100030012 [DOI] [PubMed] [Google Scholar]
- 40. Thompson C, Puterman A, Linley L, et al. The value of a scoring system for hypoxic ischaemic encephalopathy in predicting neurodevelopmental outcome. Acta Paediatr. 1997;86:757-761. 10.1111/j.1651-2227.1997.tb08581.x [DOI] [PubMed] [Google Scholar]
- 41. Perez JMR, Golombek SG, Sola A. Clinical hypoxic-ischemic encephalopathy score of the Iberoamerican Society of Neonatology (Siben): a new proposal for diagnosis and management. Rev Assoc Med Bras. 2017;63:64-69. 10.1590/1806-9282.63.01.64 [DOI] [PubMed] [Google Scholar]
- 42. Chalak LF, Adams-Huet B, Sant’Anna G. A total Sarnat score in mild hypoxic-ischemic encephalopathy can detect infants at higher risk of disability. J Pediatr. 2019;214:217-221.e1. 10.1016/j.jpeds.2019.06.026 [DOI] [PubMed] [Google Scholar]
- 43. Sarnat HB, Flores-Sarnat L, Fajardo C, et al. Sarnat grading scale for neonatal encephalopathy after 45 years: an update proposal. Pediatr Neurol. 2020;113:75-79. 10.1016/j.pediatrneurol.2020.08.014 [DOI] [PubMed] [Google Scholar]
- 44. Morales MM, Montaldo P, Ivain P, et al. Association of total Sarnat score with brain injury and neurodevelopmental outcomes after neonatal encephalopathy. Arch Dis Child Fetal Neonatal Ed. 2021;106:669-672. 10.1136/archdischild-2020-321164 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Walsh BH, Munster C, El-Shibiny H, et al. Comparison of numerical and standard Sarnat grading using the NICHD and SIBEN methods. J Perinatol. 2022;42:328-334. 10.1038/s41372-021-01180-w [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Mietzsch U, Kolnik SE, Wood TR, et al. ; HEAL Trial Study Group. Evolution of the Sarnat exam and association with 2-year outcomes in infants with moderate or severe hypoxic-ischaemic encephalopathy: a secondary analysis of the HEAL trial. Arch Dis Child Fetal Neonatal Ed. 2024;109:308-316. 10.1136/archdischild-2023-326102 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Natarajan G, Shankaran S, Laptook AR, et al. ; Extended Hypothermia Subcommittee of the Eunice Kennedy Shriver National Institute of Child Health and Human Development Neonatal Research Network. Apgar scores at 10 min and outcomes at 6–7 years following hypoxic-ischaemic encephalopathy. Arch Dis Child Fetal Neonatal Ed. 2013;98:F473-F479. 10.1136/archdischild-2013-303692 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Harris PA, Taylor R, Thielke R, et al. Research electronic data capture (REDCap)—a metadata-driven methodology and workflow process for providing translational research informatics support. J Biomed Inform. 2009;42:377-381. 10.1016/j.jbi.2008.08.010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Laptook A. Evaluation of systemic hypothermia initiated after 6 hours of age in infants ≥ 36 weeks gestation with hypoxic-ischemic encephalopathy: a Bayesian evaluation (Version 1) [Dataset]. 2020. 10.57982/hs6z-4j46 [DOI]
- 50. Shankaran S. Optimizing cooling strategies at <6 hours of age for neonatal hypoxic-ischemic encephalopathy (Version 1) [Dataset]. 2019. 10.57982/yjay-3487. [DOI]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The definition of the metadata and the data dictionary of the framework is maintained as spreadsheets and REDCap format and can be found in Supplementary Material S1, S2, and S4.48 With the separation of Datasets, Software, and Results, the Results, including the definition of the metadata and the data dictionary of the framework, and the statistical analysis results, are licensed under CC BY 4.0 and can be found at the public repository https://github.com/i3-research/COMBINE-harmonizer/tree/v1.1.0/results. The LH dataset and the OC dataset have been released to the public.49,50