Abstract
Programmed cell death (PCD) is indispensable for eukaryotic development. In animals, PCD is executed by the caspase family of cysteine proteases. Plants do not have close homologues of caspases but possess a phylogenetically distant family of cysteine proteases named metacaspases. The cellular function of metacaspases in PCD is unknown. Here we show that during plant embryogenesis, metacaspase mcII-Pa translocates from the cytoplasm to nuclei in terminally differentiated cells that are destined for elimination, where it colocalizes with the nuclear pore complex and chromatin, causing nuclear envelope disassembly and DNA fragmentation. The cell-death function of mcII-Pa relies on its cysteine-dependent arginine-specific proteolytic activity. Accordingly, mutation of catalytic cysteine abrogates the proteolytic activity of mcII-Pa and blocks nuclear degradation. These results establish metacaspase as an executioner of PCD during embryo patterning and provide a functional link between PCD and embryogenesis in plants. Although mcII-Pa and metazoan caspases have different substrate specificity, they serve a common function during development, demonstrating the evolutionary parallelism of PCD pathways in plants and animals.
Keywords: embryo suspensor, metacaspase, nuclear degradation
Programmed cell death (PCD) is indispensable for normal embryo development both in animals and in plants, where temporary, surplus, or aberrantly formed tissues and organs are removed for correct pattern formation (1, 2). The key morphogenetic event in plant embryogenesis is formation of the apicalbasal pattern via establishment of the proliferating embryo proper (apical) and the terminally differentiated suspensor (basal). Developmental programs of the embryo proper and the suspensor are closely coordinated, and imbalance causes embryonic defects or lethality (2-4). While the embryo proper gives rise to the plant, the suspensor functions during a brief period as a conduit of growth factors to the developing embryo and is subsequently eliminated by PCD (2). The terminal differentiation of the embryo suspensor is the earliest manifestation of cellular suicide in plant ontogenesis. However, the molecular mechanisms that regulate PCD in plant embryos are unknown.
The nucleus is the major target of cell degradation machinery during PCD. Nuclear degradation processes encompass chromatin events (i.e., chromatin condensation and DNA fragmentation) and nuclear envelope events (i.e., lobing of the nuclear surface and disassembly of nuclear pore complex) that occur simultaneously in the same cell (2, 5). The structural organization of plant and animal nuclei is conserved (6), explaining why the morphological pattern of nuclear degradation is also conserved (2). However, the molecular composition of plant and animal nuclear envelopes is not conserved (6), implying that different molecular mechanisms are responsible for nuclear envelope events during PCD in plants.
In animals, nuclear degradation during PCD is executed by a caspase family of cysteine proteases (5, 7). Plants do not have direct homologues of caspases but possess distantly related proteins called metacaspases that are also found in protozoa and fungi and, together with the paracaspases (found in Dictyostelium and metazoans), are suggested to be the ancestors of metazoan caspases (8-10). Despite conservation of the caspase-specific catalytic diad of histidine and cysteine in metacaspases, paracaspases, and canonical caspases, their overall sequence similarity is very low. The molecular roles of paracaspases have recently been shown to be unrelated to cell-death control (11, 12). In contrast, metacaspases are involved in PCD, because over- and underexpression of metacaspases affect the level of PCD in yeast cells (13, 14) and in plant embryos (15). The functional role of metacaspases in the PCD pathways is unknown.
Previously, we have reported that activation of protease(s) cleaving the caspase substrate Val-Glu-Ile-Asp (VEIDase activity) is essential for PCD and embryogenesis in the gymnosperm Norway spruce (Picea abies) (16). Silencing of P. abies metacaspase gene mcII-Pa inhibited VEIDase activity, suppressed PCD in the embryos, and blocked suspensor differentiation (15). Here we show that mcII-Pa is not VEIDase, because active mcII-Pa does not possess aspartate-specific proteolytic activity typical for animal caspases but prefers substrates containing arginine as the C-terminal amino acid. The proteolytic activity of mcII-Pa is paramount for the terminal differentiation and PCD of the suspensor. Immunolocalization analyses and functional assays show that mcII-Pa accumulates in the nuclei of the suspensor cells and is directly involved in the execution of nuclear disassembly.
Materials and Methods
Embryogenesis System. A normal developing embryogenic cell line 95.88.22 of P. abies, mcII-Pa-silenced clones derived from that line (15), and developmentally arrested line 88.1 (17, 18) were stored in liquid nitrogen and thawed 3 months before the experiments. After thawing, the lines were treated with auxin and cytokinin in liquid medium to promote cell proliferation followed by withdrawal of these growth factors (19). The latter treatment induced normal embryo development in line 95.88.22, but not in the silenced clones, and in line 88.1, which continued to proliferate without growth factors (15, 18).
Recombinant McII-Pa. For biochemistry and autoprocessing analyses, purified recombinant mcII-Pa and the catalytically inactive mutant C139A of mcII-Pa were produced in Escherichia coli (details are available in Supporting Methods, which is published as supporting information on the PNAS web site).
EnzymeActivityAssays.Allfluorogenicamino-4-methylcoumarin-conjugated peptidic substrates were obtained from Bachem and inhibitors from Roche Biosciences, except for caspase inhibitors (Peptide Institute, Osaka) and H-Glu-Gly-Arg-chloromethyl ketone (H-ERG-cmk; Bachem). Proteolytic activity of recombinant mcII-Pa and P. abies cell extracts (16) were assayed in 150-μl reaction mixtures containing 0.3 μg of recombinant protein or 22.5 μg of total protein, respectively, and 50 μM individual substrate in the optimized assay buffer: 100 mM Hepes, pH 7.0/50 mM CaCl2/0.1% 3-[(3-cholamidopropyl)dimethylammonio]-1-propanesulfonate/5 mM DTT (Fig. 7, which is published as supporting information on the PNAS web site). First-order kinetics of substrate cleavage was measured as described (16). For each independent experiment, at least three assays were performed, each assay containing triplicate reactions.
Polyclonal Antiserum and Immunoblotting. The mcII-Pa recombinant protein was dialyzed against PBS overnight. In each immunization, 50-100 μg of recombinant protein was used per mouse, mixed 1:1 with Freund's incomplete adjuvant. Three boosts were performed after 2-week intervals. Serum was collected 10 days after the final boost and stored at -80°C. The immunoblotting procedure is described in Supporting Methods.
Immunofluorescence Microscopy. Embryos from line 95.88.22 were collected after 7 days following withdrawal of growth factors (18, 19). For visualization of both nuclear DNA fragmentation and mcII-Pa, whole-embryo samples were prepared as described (18), with the following order of treatments: (i) fixation with 3.7% paraformaldehyde, (ii) incubation with a TUNEL reaction mixture containing TMR red-labeled dUTP and terminal deoxynucleotidyltransferase (TdT) (Roche Biosciences), (iii) treatment with cell wall-digesting enzymes, (iv) probing with anti-mcII-Pa (dilution 1:1,000) and then with anti-mouse FITC conjugates (dilution 1:250; Sigma-Aldrich), and (v) staining with DAPI. To ensure that TUNEL does not affect the binding of anti-mcII-Pa, two additional groups of embryos were prepared for visualization of either DNA fragmentation or mcII-Pa. Application of TMR red labeled-dUTP without TdT and omission of anti-mcII-Pa were used as negative controls. Approximately 500 embryos were examined by using a Leica DMRE epifluorescence microscope and 100 embryos with a Bio-Rad Radian 2000 confocal microscope. The preimmune serum produced no signal at dilution 1:1,000. When anti-mcII-Pa antibody was preincubated with recombinant mcII-Pa before application onto the embryos, no specific signal was detected (Fig. 8, which is published as supporting information on the PNAS web site). Additionally, 100 control embryos were stained with TUNEL, with antibody against MAP-65 (18), and with DAPI and examined by using confocal microscopy. These embryos did not show colocalization of microtubule-associated protein, MAP-65 (18), with the TUNEL-positive nuclei.
Immunogold Transmission Electron Microscopy. Embryos were prepared for immunogold labeling as described (19). The detailed procedure is available in Supporting Methods.
Cell-Free System. Nuclei were isolated from protoplasts prepared from 3-day-old proliferating cell line 88.1, as described (19). Protoplasts were separated from undigested cell clumps by filtration through 80-μm nylon mesh and collected by centrifugation at 90 × g for 5 min. Pelleted protoplasts were washed three times by repeated centrifugation in a solution containing 5 mM CaCl2 and 0.4 M mannitol (pH 5.8). The washed protoplasts were transferred to 1 ml of prechilled lysis buffer containing 100 mM Hepes (pH 7.0), 80 mM KCl, 0.1 mM EDTA, 5 mM DTT, 1 mM spermidine, and 1 mM spermine in a 50-ml Dounce homogenizer and left for 30 min on ice. After addition of 30 ml of fresh lysis buffer to the swollen protoplasts in the homogenizer, the protoplasts were broken by 50 strokes to release nuclei. The homogenate was then layered 1:2 onto a 30% sucrose solution prepared in the lysis buffer and centrifuged at 500 × g for 15 min at 4°C. The pellet with intact nuclei was resuspended in 2 ml of washing solution containing 100 mM Hepes (pH 7.0), 2 mM MgSO4, and 5 mM DTT and centrifuged at 250 × g for 3 min at 4°C. The pellet containing purified nuclei was then diluted with the fresh washing solution to a concentration of 6 × 106 nuclei ml-1.
Cell extract was prepared from 7-day-old proliferating cell line 95.88.22 in the assay buffer containing 100 mM Hepes (pH 7.0), 50 mM CaCl2, 2 mM MgSO4, and 5 mM DTT, as described (16). Protein concentration was adjusted to 1 mg·ml-1. Aliquots of 50 μl of cell extract (or assay buffer) mixed with 10-μl aliquots of purified nuclei (6 × 104 nuclei) were incubated with or without recombinant mcII-Pa (0.5 μg) or its inhibitor H-EGR-cmk (final concentration in the assay, 20 μM) for 150 min at 25°C. Nuclei were then fixed with 2% paraformaldehyde in PBS (pH 7.4) on polylysine-coated slides for 1 h at 42°C, labeled with TUNEL diluted 1:1 in TUNEL dilution buffer (Roche Biosciences) and stained with DAPI, as reported (19). The frequencies of TUNEL-positive and morphologically aberrant nuclei were determined by scoring >1,000 nuclei with at least three replicates for every treatment. The whole cell-free system experiment was repeated twice.
Results
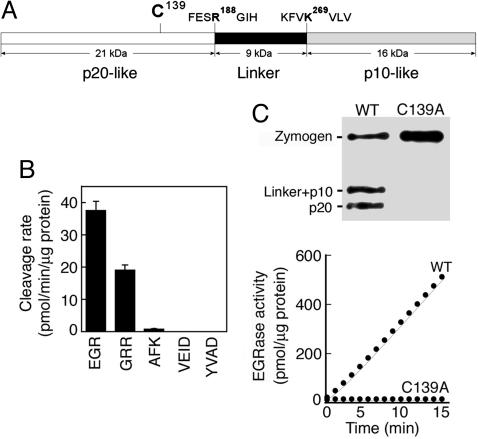
McII-Pa and Caspases Have Different Substrate Specificity. Bacterially expressed recombinant mcII-Pa zymogen was autocatalytically cleaved at two sites after: (i) Arg-188 separating a p20 caspase-like subunit and a type II metacaspase-specific linker region, and (ii) Lys-269 separating the linker region and p10 caspase-like subunit (Fig. 1A). This cleavage pattern indicated arginine- and/or lysine-specific proteolytic activity of mcII-Pa, which was confirmed by using a fluorometric peptide cleavage assay (Fig. 1B). Although recombinant mcII-Pa cleaved peptides with both arginine and lysine at the P1 position, the cleavage rate was 25- to 50-fold higher for arginine- than for lysine-containing peptides (Fig. 1B). None of the caspase-specific substrates (i.e., containing aspartic acid at P1) were cleaved by mcII-Pa (Fig. 1B and data not shown), indicating that mcII-Pa, like four recently reported Arabidopsis metacaspases (14, 20), has different substrate specificity than animal caspases. Like caspases, mcII-Pa shows maximal proteolytic activity at neutral pH but in contrast was 10 times more sensitive to Zn2+ than the reported caspases (21) (Fig. 7 B and C). Although caspase activity does not depend on Ca2+ (21), mcII-Pa requires millimolar Ca2+ concentration (optimum at 50 mM) for activation (Fig. 7A). Proteolytic activity of mcII-Pa was insensitive to caspase-specific inhibitors but was readily inhibited by leupeptin, Nα-tosyl-l-lysine-cmk, and mcII-Pa substrate-mimetic inhibitor H-EGR-cmk (Fig. 7D). Mutation of the catalytic Cys-139 abrogated autoprocessing of zymogen, as shown by Western blotting with antibody to mcII-Pa (Fig. 1C). The C139A mutant failed to cleave Boc-EGR-amido-4-methylcoumarin, the preferred substrate of WT mcII-Pa (Fig. 1C). Therefore, mcII-Pa is a processive arginine-specific cysteine-dependent protease.
Fig. 1.
McII-Pa is a processive cysteine protease with arginine-specific enzymatic activity. (A) McII-Pa zymogen consists of p20 caspase-like subunit with catalytic Cys-139, a linker region, and p10 caspase-like subunit. Autoprocessing of mcII-Pa in E. coli occurs by autocleavage after Arg-188 and Lys-269 between the domains. (B) Substrate specificity of recombinant mcII-Pa tested on five different substrates containing Arg, Lys, or Asp at the P1 position. (C) Western blotting demonstrating that WT recombinant mcII-Pa (lane WT), but not catalytic Cys-139 mutant (lane C139A), undergoes autoprocessing. The graph shows that Cys-139 mutant is proteolytically inactive.
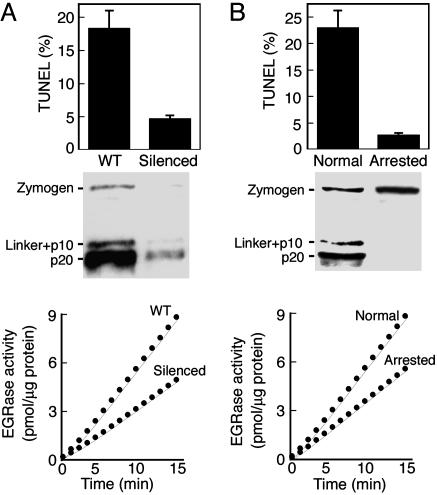
Proteolytic Activity of McII-Pa Is Essential for Embryonic Pattern Formation. We assessed whether proteolytic activity of the metacaspase is important for regulating developmental PCD using P. abies somatic embryogenesis. Somatic embryo development resembles zygotic embryogenesis and can also be synchronized by growth factors enabling systematic observations of embryo patterning (3, 17). Silencing of the metacaspase gene in this system resulted in increased cell proliferative activity and lack of suspensor differentiation (15). The magnitude of PCD was estimated by calculating the percentage of cells with fragmented nuclear DNA. This showed that PCD was suppressed in the silenced clones by ≈4-fold (Fig. 2A). Concomitantly, arginine-specific proteolytic activity on Boc-EGR-amido-4-methylcoumarin was reduced but not abolished in mcII-Pa-silenced clones, indicating that other arginine-specific proteases were not affected by gene silencing (Fig. 2 A).
Fig. 2.
Cell-death function of mcII-Pa relies on its proteolytic activity. (A) Western blotting of WT (WT) and mcII-Pa silenced lines probed with anti-mcII-Pa shows reduction of mcII-Pa level in the silenced line. TUNEL of WT and silenced lines shows suppression of PCD in the silenced lines (Top). The EGRase proteolytic activity is reduced in the silenced lines (Bottom). Data are representative of three silenced clones. (B) A developmentally arrested cell line unable to form the embryos exhibits a reduced level of TUNEL (Top), lack of mcII-Pa processing (Western blot), and a reduced level of EGRase activity (Bottom). Error bars indicate SEM.
We then determined whether constitutive PCD deficiency is also associated with reduced mcII-Pa activity. For this, we used the developmentally arrested P. abies cell line, which is unable to differentiate suspensors and hence represents a phenotypic twin of the mcII-Pa-silenced clones (17, 18). McII-Pa was expressed in the developmentally arrested line but was not processed (Fig. 2B). This line also showed decreased EGRase proteolytic activity (cleavage of Boc-EGR-amido-4-methylcoumarin) and a 10-fold decrease of the TUNEL frequency compared with the normal line (Fig. 2B). Together, these data demonstrate that the cell-death function of metacaspase during embryogenesis relies on its proteolytic activity that in turn requires processing of mcII-Pa zymogen.
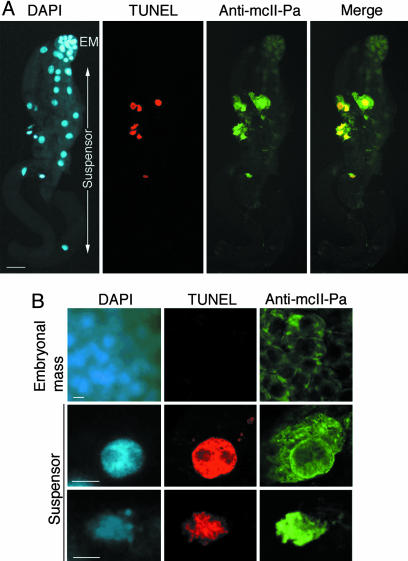
McII-Pa Accumulates in the Nuclei of Dying Cells. In all gymnosperms, the suspensor is composed of several layers of terminally differentiated cells, originating from asymmetric cell divisions in the embryonal mass (functionally similar to the embryo proper of angiosperms) (2, 22). Suspensor cells do not divide but instead become committed to PCD as soon as they are produced. Although the cells in the upper layer of the suspensor (i.e., adjacent to the embryonal mass) are in the commitment phase of PCD, the cells in the lower layers exhibit a gradient of successive stages of autophagy-mediated cell dismantling toward the basal end of the suspensor where hollow walled cell corpses are located (2, 18). Thus, successive cell-death processes can be observed simultaneously in a single embryo. Moreover the position of the cell within the embryo can be used as a marker of the stage of cell death. The intracellular localization of mcII-Pa protein was correlated with PCD progression in the embryos by epifluorescence microscopy by using a mcII-Pa-specific antibody. To visualize the nuclei undergoing DNA fragmentation, the embryos were also stained with TUNEL and DAPI. McII-Pa was detected in all embryonic cells, with a sharp increase of the signal in the TUNEL-positive cells in the suspensor (Fig. 3A). Further analysis of embryos using confocal laser-scanning microscopy led to the identification of two distinct mcII-Pa immunolocalization patterns correlated with the degree of nuclear degradation in the suspensor (Fig. 3B). Cells with fragmented DNA but still possessing an intact nuclear envelope accumulated mcII-Pa at the nuclear periphery and in the perinuclear cytoplasm, whereas cells with a degraded nuclear envelope showed intense labeling inside the nucleus (Fig. 3B).
Fig. 3.
McII-Pa colocalizes with nuclei containing fragmented DNA in the embryo-suspensor cells. (A) Epifluorescence microscopy of DAPI, TUNEL, and anti-mcII-Pa labeling in the whole embryo. A representative example of one of ≈500 embryos analyzed. EM, embryonal mass. (B) Confocal microscopy of embryonal mass cells (Top), and suspensor cells with morphologically intact (Middle), and degraded (Bottom) nuclear envelope. A representative example of 1 of the 100 embryos analyzed. [Bars, 100 (A) and 10 μm(B).]
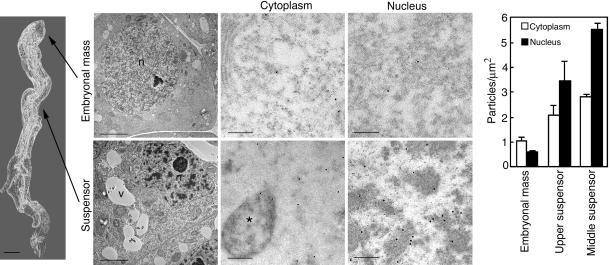
To assess whether mcII-Pa translocates from the cytoplasm to the nuclei during PCD, the subcellular distribution of mcII-Pa was analyzed by using immunogold labeling and transmission electron microscopy. The density of gold particles was determined in the cytoplasm and nuclei of proliferating cells located in the embryonal mass, cells at the commitment phase of PCD in the upper zone of the suspensor, and cells at the execution phase of PCD in the middle zone of the suspensor (18) (Fig. 4). The cells at the more advanced stages of cell dismantling located in the basal part of the suspensor were not analyzed, because they contained very little cytoplasm, with the majority of cell volume occupied by the vacuoles (18, 19). In parallel with a steady increase of the mean density of gold particles in the cytoplasm and nuclei toward the middle zone of the suspensor, the nucleus-to-cytoplasm ratio of labeling changed dramatically, from 0.6 in the embryonal mass to 1.7 and 2.0 in the upper and middle zones of the suspensor, respectively (Fig. 4). In the cytoplasm of the suspensor cells, labeling was concentrated around the nuclear envelope, whereas virtually all gold particles in the nuclei were associated with patches of electron-dense chromatin (Fig. 4) and the disassembling nuclear pore complex (19) (Fig. 5). No labeling was detected in the autophagic vacuoles, typical organelles of the autophagic PCD pathway responsible for cell content autodestruction (1, 2, 19). This result is consistent with observations that mcII-Pa requires neutral or near-neutral pH for catalysis, whereas the vacuole content is acidic (Fig. 7B). The combined immunolocalization data demonstrate that during PCD in the embryo suspensor, mcII-Pa translocates from the cytoplasm into the nuclei.
Fig. 4.
McII-Pa translocates from the cytoplasm into the nucleus during PCD in the embryo suspensor. Phase-contrast image of the representative embryo (Left; Bar, 100 μm) analyzed by immunogold electron microscopy with antibody to mcII-Pa (Center). Upper Center corresponds to the embryonal mass cells. Lower Center corresponds to the cells from the middle zone of the suspensor. Left Center shows cell ultrastructure (Bars, 5 μm), whereas Right Center shows immunogold labeling of the cytoplasm and nucleus, respectively (Bars, 200 nm). Gold particles are associated with the condensed chromatin in the suspensor nucleus. Graph (Right) shows density of the gold particles in the cytoplasm and nuclei of the embryonal mass and upper and middle zones of the suspensor. Error bars indicate SEM. n, nucleus; v, vacuole; asterisk indicates autophagosome.
Fig. 5.
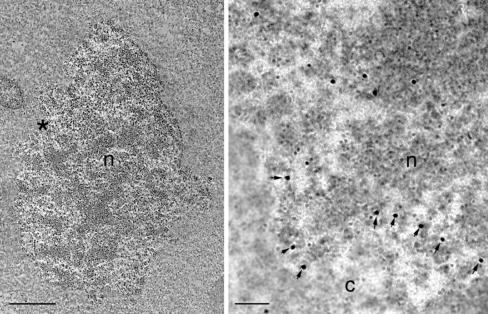
Immunogold labeling of mcII-Pa at the sites of nuclear pore complex disassembly in the embryo-suspensor cells. (Left) Lobed nucleus typical for the early stages of nuclear degradation. Asterisk denotes the area shown enlarged (Right). The gold particles marked by arrows are associated with nuclear pore complex disassembly. c, cytoplasm; n, nucleus. [Bars, 5 μm (Left) and 100 nm (Right).]
Proteolytically Active McII-Pa Induces Nuclear Degradation. Given clear evidence that mcII-Pa is important for PCD in vivo (Fig. 2) and specific intranuclear accumulation of mcII-Pa in dying cells (Figs. 3, 4, 5), we examined whether metacaspase can directly affect nuclear integrity. For this, a cell-free system was developed where nuclei from the PCD-deficient cell line (Fig. 2B) were coincubated with cell extracts prepared from a normal embryogenic cell line. The frequency of TUNEL-positive nuclei was not affected by incubation with the cell extract alone but increased dramatically after addition of recombinant WT mcII-Pa (Fig. 6A). A peptidic inhibitor for mcII-Pa, H-EGR-cmk (Fig. 7D), significantly diminished the effect of mcII-Pa on DNA fragmentation. Addition of a catalytically inactive mutant form of mcII-Pa had no effect on DNA fragmentation (Fig. 6A).
Fig. 6.
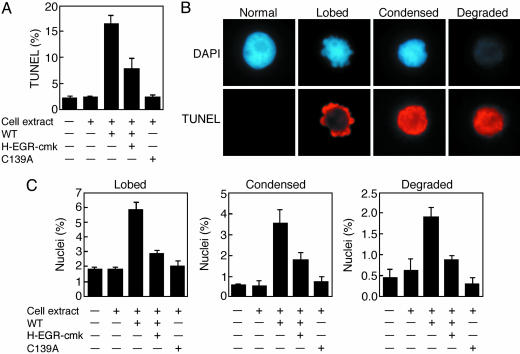
McII-Pa induces nuclear degradation in the cell-free system. (A) Frequency of TUNEL-positive nuclei before and after treatment with cell extract, recombinant WT (WT) mcII-Pa in the absence or presence of H-EGR-cmk, and catalytically inactive mutant mcII-Pa (C139A), as indicated below each bar. (B) Representative examples of normal nuclei and three stages of nuclear degradation after staining with DAPI and TUNEL. (C) Frequency of nuclei at each stage of degradation in the cell-free system experiments conducted as described in A. Error bars indicate SEM.
We observed that nuclear DNA fragmentation in the cell-free system was generally accompanied by stereotyped changes of nuclear morphology. These changes occurred in three major stages distinguished by DAPI staining: (i) lobing of the nuclear envelope, (ii) condensation of chromatin, and (iii) complete nuclear degradation (Fig. 6B). This pathway resembles the process of nuclear disassembly previously described for embryo-suspensor cells in vivo, although chromatin condensation is less pronounced in the latter case (19). Addition of active mcII-Pa to the cell-free system induced a several-fold increase of the frequency of nuclei at each of the three degradation stages (Fig. 6C). This effect was greatly diminished or abrogated in the presence of H-EGR-cmk (Fig. 6C). As expected, mutant protein was ineffective in inducing nuclear changes (Fig. 6C).
Discussion
Our findings establish that metacaspase mcII-Pa is directly involved in the pathway leading to nuclear degradation, which is a key event of most, if not all, eukaryotic cell-death programs (23). The high frequency of immunogold labeling of mcII-Pa at sites of nuclear pore complex disassembly and in electron-dense chromatin patches (Figs. 4 and 5) suggests that metacaspase, like effector caspases 3 and 6, may cleave nuclear structural proteins, leading to disassembly of the nuclear envelope, and thereby possibly facilitates chromatin degradation by nucleases (24, 25). The molecular composition of plant and animal nuclei is not conserved (6); thus, it is not surprising that metacaspases have different substrate specificity from caspases responsible for nuclear degradation in animals. However, despite the differences in primary structure and substrate specificity of metacaspases and caspases, they serve common cellular functions as executioners of PCD, demonstrating evolutionary parallelism of the cell-death pathways in plants and animals.
McII-Pa is a nuclear-cytoplasmic, but not vacuolar, protease. There is growing evidence of the autophagic nature of PCD in plants, where “cell-dismantling” and “corpse-clearance” tasks are fulfilled simultaneously by growing vacuoles (1, 2, 26-28). Although the elimination of the embryo suspensor is a typical example of autophagic PCD (2), we have previously reported that nuclear degradation does not correlate morphologically with the progression of autophagic dismantling of the cytoplasm in the suspensor (19). This is consistent with the idea that execution of PCD in plants is controlled by two groups of enzymes having separate cellular localization (29). One group, including the vacuolar processing enzymes (30, 31), is accumulated and activated inside vacuoles, whereas the other group encompassing the metacaspase mcII-Pa have a nuclear-cytoplasmic localization. Silencing of mcII-Pa inhibited arginine-specific proteolytic activity (Fig. 2 A) and at the same time resulted in a decrease of VEIDase caspase-like activity (15). Recombinant mcII-Pa does not cleave caspase substrates, including the VEID sequence-containing substrate (Fig. 1B), indicating that VEIDase activity is caused by different protease(s). Pharmacological inhibition of VEIDase impaired the embryonic pattern formation, yielding a phenotype similar to mcII-Pa-silenced clones (16), suggesting that mcII-Pa lies upstream of yet unidentified protease(s) with VEIDase activity in the same cell-death pathway essential for plant embryogenesis.
The terminal differentiation of the embryo-suspensor is the earliest manifestation of PCD in plant ontogenesis, and failure to activate PCD in the suspensor results in severe developmental abnormalities, often leading to the death of the whole embryo (2). The identification of the role of the metacaspase in the suspensor PCD establishes a mechanistic link between PCD and embryo patterning and contributes to our understanding of the earliest stages of plant development.
Supplementary Material
Acknowledgments
We thank Sara von Arnold, Patrick Hussey, and Sten Orrenius for critical reading of the manuscript. This work was supported by the Swedish Foundation for International Cooperation in Research and Higher Education, the Carl Tryggers Foundation, the Spanish Ministry of Education, the Swedish Research Foundation, and the Royal Swedish Academy of Agriculture and Forestry.
Author contributions: P.V.B., M.F.S., and A.S. designed research; P.V.B., M.F.S., L.H.F., G.D., A.A.Z., S.R.-N., and A.S. performed research; G.D. and B.Z. contributed new reagents/analytic tools; P.V.B., M.F.S., L.H.F., and A.S. analyzed data; and P.V.B., B.Z., and A.S. wrote the paper.
Abbreviations: PCD, programmed cell death; cmk, chloromethyl ketone.
Data deposition: The sequence reported in this paper has been deposited in the EMBL database [accession no. AJ534970 (mcII-Pa)].
References
- 1.Baehrecke, E. (2002) Nat. Rev. Mol. Cell Biol. 3, 779-787. [DOI] [PubMed] [Google Scholar]
- 2.Bozhkov, P. V., Suarez, M. F. & Filonova, L. H. (2005) Curr. Top. Dev. Biol. 67, 135-179. [DOI] [PubMed] [Google Scholar]
- 3.Goldberg, R. B., de Paiva, G. & Yadegari, R. (1994) Science 266, 605-614. [DOI] [PubMed] [Google Scholar]
- 4.Lukowitz, W., Roeder, A., Parmenter, D. & Somerville, C. (2004) Cell 116, 109-119. [DOI] [PubMed] [Google Scholar]
- 5.Earnshaw, W. C. (1995) Curr. Opin. Cell Biol. 7, 337-343. [DOI] [PubMed] [Google Scholar]
- 6.Rose, A., Patel, S. & Meier, I. (2004) Planta 218, 237-336. [DOI] [PubMed] [Google Scholar]
- 7.Earnshaw, W. C., Martins, L. M. & Kaufmann, S. H. (1999) Annu. Rev. Biochem. 68, 383-424. [DOI] [PubMed] [Google Scholar]
- 8.Uren, A. G., O'Rourke, K., Aravind, L., Pisabarro, M. T., Seshagiri, S., Koonin, E. V. & Dixit, V. M. (2000) Mol. Cell 6, 961-967. [DOI] [PubMed] [Google Scholar]
- 9.Aravind, L. & Koonin, E. V. (2002) Proteins 46, 355-367. [DOI] [PubMed] [Google Scholar]
- 10.Koonin, E. V. & Aravind, L. (2002) Cell Death Differ. 9, 394-404. [DOI] [PubMed] [Google Scholar]
- 11.Ruefli-Brasse, A. A., French, D. M. & Dixit, V. M. (2003) Science 302, 1581-1584. [DOI] [PubMed] [Google Scholar]
- 12.Roisin-Bouffay, C., Luciani, M. F., Klein, G., Levraud, J. P., Adam, M. & Golstein, P. (2004) J. Biol. Chem. 279, 11489-11494. [DOI] [PubMed] [Google Scholar]
- 13.Madeo, F., Herker, E., Maldener, C., Wissing, S., Lächelt, S., Herlan, M., Fehr, M., Lauber, K., Sigrist, S. J., Wesselborg, S., et al. (2002) Mol. Cell 9, 911-917. [DOI] [PubMed] [Google Scholar]
- 14.Watanabe, N. & Lam, E. (2005) J. Biol. Chem. 280, 14691-14699. [DOI] [PubMed] [Google Scholar]
- 15.Suarez, M. F., Filonova, L. H., Smertenko, A., Savenkov, E. I., Clapham, D. H., von Arnold, S., Zhivotovsky, B. & Bozhkov, P. V. (2004) Curr. Biol. 14, R339-R340. [DOI] [PubMed] [Google Scholar]
- 16.Bozhkov, P. V., Filonova, L. H., Suarez, M. F., Helmersson, A., Smertenko, A. P., Zhivotovsky, B. & von Arnold, S. (2004) Cell Death Differ. 11, 175-182. [DOI] [PubMed] [Google Scholar]
- 17.Filonova, L. H., Bozhkov, P. V. & von Arnold, S. (2000) J. Exp. Bot. 51, 249-264. [DOI] [PubMed] [Google Scholar]
- 18.Smertenko, A. P., Bozhkov, P. V., Filonova, L. H., von Arnold, S. & Hussey, P. J. (2003) Plant J. 33, 813-824. [DOI] [PubMed] [Google Scholar]
- 19.Filonova, L. H., Bozhkov, P. V., Brukhin, V. B., Daniel, G., Zhivotovsky, B. & von Arnold, S. (2000) J. Cell Sci. 113, 4399-4411. [DOI] [PubMed] [Google Scholar]
- 20.Vercammen, D., van de Cotte, B., de Jaeger, G., Eeckhout, D., Casteels, P., Vandepoele, K., Vandenberghe, I., van Beeumen, J., Inze, D. & van Breusegem, F. (2004) J. Biol. Chem. 279, 45329-45336. [DOI] [PubMed] [Google Scholar]
- 21.Stennicke, H. R. & Salvesen, G. S. (1997) J. Biol. Chem. 272, 25719-25723. [DOI] [PubMed] [Google Scholar]
- 22.Singh, H. (1978) in Handbuch der Pflanzenanatomie, eds. Zimmerman, W., Carlquist, Z., Ozenda, P. & Wuff, H. D. (Gebrüder Bornträger, Berlin), pp. 187-241.
- 23.Clarke, P. G. H. (1990) Anat. Embryol. 181, 195-213. [DOI] [PubMed] [Google Scholar]
- 24.Lazebnik, Y. A., Takahashi, A., Moir, R. D., Goldman, R. D., Poirier, G. G., Kaufmann, S. H. & Earnshaw, W. C. (1995) Proc. Natl. Acad. Sci. USA 92, 9042-9046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Casiano, C. A., Martin, S. J., Green, D. R. & Tan, E. M. (1996) J. Exp. Med. 184, 765-770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Lockshin, R. A. & Zakeri, Z. (2002) Curr. Opin. Cell Biol. 14, 727-733. [DOI] [PubMed] [Google Scholar]
- 27.Zhivotovsky, B. (2002) Cell Death Differ. 9, 867-869. [DOI] [PubMed] [Google Scholar]
- 28.Lam, E. (2004) Nat. Rev. Mol. Cell Biol. 5, 305-315. [DOI] [PubMed] [Google Scholar]
- 29.Woltering, E. J. (2004) Trends Plant Sci. 9, 469-472. [DOI] [PubMed] [Google Scholar]
- 30.Hatsugai, N., Kuroyanagi, M., Yamada, K., Meshi, T., Tsuda, S., Kondo, M., Nishimura, M. & Hara-Nishimura, I. (2004) Science 305, 855-858. [DOI] [PubMed] [Google Scholar]
- 31.Rojo, E., Martin, R., Karter, C., Zouhar, J., Pan, S., Plotnikova, J., Jin, H., Paneque, M., Sanchez-Serrano, J. J., Baker, B., et al. (2004) Curr. Biol. 9, 1897-1906. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.