Abstract
T4 polynucleotide kinase (Pnk), in addition to being an invaluable research tool, exemplifies a family of bifunctional enzymes with 5′-kinase and 3′-phosphatase activities that play key roles in RNA and DNA repair. T4 Pnk is a homotetramer composed of a C-terminal phosphatase domain and an N-terminal kinase domain. The 2.0 Å crystal structure of the isolated kinase domain highlights a tunnel-like active site through the heart of the enzyme, with an entrance on the 5′ OH acceptor side that can accommodate a single-stranded polynucleotide. The active site is composed of essential side chains that coordinate the β phosphate of the NTP donor and the 3′ phosphate of the 5′ OH acceptor, plus a putative general acid that activates the 5′ OH. The structure rationalizes the different specificities of T4 and eukaryotic Pnk and suggests a model for the assembly of the tetramer.
Keywords: bacteriophage T4/3′ phosphatase/polynucleotide kinase/RNA repair
Introduction
Polynucleotide kinase (Pnk) was discovered by the Richardson and Hurwitz laboratories in T4 and T2 bacteriophage-infected Escherichia coli (Richardson, 1965; Novogrodsky and Hurwitz, 1966; Novogrodsky et al., 1966). The use of bacteriophage T4 Pnk to label 5′ DNA or RNA ends with 32P was instrumental in the development of methods for the analysis of nucleic acid structure, molecular cloning and nucleic acid sequencing. Although the historical importance of T4 Pnk in the recombinant DNA revolution is well known, it is less widely appreciated that T4 Pnk spearheads a pathway of ‘RNA repair’ in vivo. During T4 infection, Pnk participates in an elaborate pathogen–host dynamic whereby the bacterium attempts to thwart T4 protein synthesis by inducing site-specific breakage of host-cell tRNAs, to which the phage responds by repairing the broken tRNAs using Pnk and a phage-encoded RNA ligase (Amitsur et al., 1987). T4 Pnk catalyzes two enzymatic reactions in this pathway: (i) the transfer of the γ phosphate from ATP to the 5′ OH terminus of RNA; and (ii) the hydrolytic removal of a 3′ PO4 terminus from RNA (Richardson, 1965; Novogrodsky and Hurwitz, 1966; Novogrodsky et al., 1966; Becker and Hurwitz, 1967; Cameron and Uhlenbeck, 1977). Eukaryotic tRNA splicing entails a similar series of reactions (Abelson et al., 1998).
It is now clear that T4 Pnk exemplifies a growing family of repair enzymes that heal broken termini in RNA or DNA by converting 3′ PO4/5′ OH ends into 3′ OH/5′ PO4 ends, which are then suitable for sealing by RNA or DNA ligases. In fission yeast and metazoans, Pnk orthologs with specificity for phosphorylation of 5′ OH DNA termini function in the repair of DNA damage induced by oxidation, radiation and topoisomerase I poisons (Jilani et al., 1999a; Karimi-Busheri et al., 1999; Whitehouse et al., 2001; Meijer et al., 2002). No structural information is available for any member of the Pnk enzyme family. Here we investigate the functional organization of T4 Pnk and report the atomic structure of the N-terminal 5′-kinase domain at 2.0 Å resolution.
Results and discussion
Pnk functional domains
T4 Pnk is a homotetramer of a 301 amino acid polypeptide (Panet et al., 1973; Midgley and Murray, 1985; Wang and Shuman, 2001). Despite its wide use in nucleic acid biochemistry, the structure and mechanism of this bifunctional enzyme remain largely uncharted. Early studies by Soltis and Uhlenbeck (1982a,b) implicated the N-terminus of Pnk in the 5′-kinase reaction and the C-terminus in the 3′-phosphatase function. Recently, we employed site-directed mutagenesis to localize essential constituents of the active sites for the 5′-kinase and 3′-phosphatase activities (Wang and Shuman, 2001, 2002). Amino acids required for catalysis of the 5′-kinase reaction (Lys15, Ser16, Asp35, Arg38, Asp85 and Arg126) map to the N-terminal half, whereas residues essential for the 3′-phosphatase function (Asp165, Asp167, Arg176, Arg213, Asp254 and Asp278) cluster in the C-terminal half (Figure 1A). These results suggest the existence of a domain boundary located between Arg126 and Asp165.
Fig. 1. Functional domains of T4 Pnk. (A) Domain organization of T4 Pnk. The N-terminal 5′-kinase and C-terminal 3′-phosphatase domains are depicted as horizontal bars. The Walker A-box motif near the N-terminus of the kinase domain is in black. Basic and acidic side chains essential for the kinase (K15, S16, D35, R38, D85 and R126) and phosphatase (D165, D167, R176, R213, D254 and D278) activities are highlighted. (B) Aliquots (3 µg) of the Ni–agarose preparations of full-length wild-type (WT) Pnk and the indicated deletion mutants were analyzed by SDS–PAGE. Polypeptides were visualized by staining with Coomassie Blue dye. The positions and sizes (in kDa) of marker proteins are indicated on the left. (C) Phosphate release from 3′ dTMP is plotted as a function of input WT (closed cirles), NΔ138 (open circles), NΔ148 (squares) and NΔ158 (triangles) protein. (D) Pnk(149–301) [NΔ148] was sedimented in a glycerol gradient together with BSA, ovalbumin and cytochrome c. Aliquots (20 µl) of the odd numbered fractions were analyzed by SDS–PAGE. A Coomassie Blue-stained gel is shown. The direction of sedimentation is from the right (top of gradient) to the left (bottom).
To test this hypothesis, we engineered a series of N-terminal Pnk truncation mutants, NΔ138, NΔ148 and NΔ158, and produced the recombinant proteins in bacteria as His10-tagged fusions. The full-length and truncated Pnk derivatives were purified by Ni–agarose chromatography (Figure 1B). Phosphatase activity was measured by the release of inorganic phosphate from deoxythymidine 3′ monophosphate. The specific activities of NΔ138 and NΔ148 were virtually identical to that of full-length Pnk, whereas the NΔ158 mutant was 19% as active (Figure 1C). These data indicate that the C-terminal half of Pnk (amino acids 149–301) comprises an autonomous 3′-phosphatase domain with full catalytic activity. The native size of the NΔ148 protein was investigated by glycerol gradient sedimentation. Marker proteins bovine serum albumin (BSA; 66 kDa), ovalbumin (45 kDa) and cytochrome c (13 kDa) were included as internal standards. The NΔ148 polypeptide, with a calculated mass of 20 kDa, sedimented a discrete peak at a position coincident with ovalbumin, which indicated that the NΔ148 protein is a homodimer (Figure 1D). The 3′-phosphatase activity profile coincided with the sedimentation profile of the NΔ148 polypeptide (data not shown). The NΔ138 derivative also sedimented as a homodimer (data not shown). We conclude that the C-terminal phosphatase domain suffices for the formation of a stable dimer, and we infer that the N-terminal domain is required for the transition from dimer to tetramer.
We showed previously that the N-terminal segment of Pnk from amino acids 1–181 (corresponding to a chymotrypsin-resistant fragment characterized by limited proteolysis) retained 5′-kinase activity, albeit at an order of magnitude lower specific activity than full-length Pnk. The Pnk(1–181) protein sedimented as a monomer in a glycerol gradient (Wang and Shuman, 2001). In light of the above results defining the proximal boundary of the phosphatase domain near position 149, we constructed a shorter N-terminal derivative, Pnk(1–147), and purified the His-tagged recombinant protein from bacteria. The 5′-kinase activity of Pnk(1–147) was virtually identical to that of Pnk(1–181) and glycerol gradient sedimentation indicated that Pnk(1–147) was monomeric (data not shown). These results indicate that the 5′-kinase and 3′-phosphatase domains of Pnk can be physically separated at a domain boundary, but also suggest that optimal 5′-kinase activity may depend on Pnk oligomerization.
Crystal structure of the kinase domain
We crystallized the N-terminal kinase domain Pnk(1–181) and solved the structure by single isomorphous replacement with anomalous scattering using diffraction data from native and Hg-derivatized crystals. The refined model at 2.0 Å resolution had a crystallographic R-factor of 21.0% and an Rfree of 23.3% with excellent stereochemistry (Table I). The structure comprised a continuous polypeptide from Pnk residues 1 to 152. The N-terminal His-tag and the C-terminal segment from amino acids 153 to 181 (i.e. the proximal segment of the phosphatase domain) were disordered and had no interpretable electron density.
Table I. Summary of crystallographic analysis.
| Data collection | Native (high) | 2 mM HgAc |
|---|---|---|
| dMin/λ (Å) | 20–2.0/0.9790 | 20–2.25/1.5418 |
| No. of sites | — | 1 |
| Rsyma (%) overall (outer shell) | 5.1 (22.3) | 5.5 (22.3) |
| Coverage (%) overall (outer shell) | 89.2 (77.7) | 79.4 (49.4) |
| I/σ (I) overall (outer shell) | 15.3 (4.3) | 22.4 (2.1) |
| Reflections (total/unique) | 36111/12892 | 31576/14574 |
| Phasing statistics | ||
| MFIDb (20–2.25 Å) (%) | 20.7 (26.7) | |
| Overall phasing powerc (centric/acentric) | 0.75/0.81 | |
| Mean FOMd (centric/acentric) | 0.26/0.27 | |
| Mean FOM after RESOLVE (20–2.0 Å) (centric/acentric) | 0.55/0.44 | |
| Refinement | ||
| Resolution range (Å) | 20–2.0 | |
| Reflections > 0.0σ | 13199 | |
| Total atoms/water/sulfate | 1404/142/20 | |
| R/Rfreee | 0.210/0.231 | |
| R.m.s.d. bond (Å)/angles (°) | 0.006/1.000 | |
| R.m.s.d. B (Å2) (main chain/side chain) | 0.968/1.291 |
aRsym = Σ|I – <I>|/ΣI, where I is the observed intensity and <I> the average intensity.
bMFID (mean fractional isomorphous difference) = Σ||Fph| – |Fp||/Σ|Fp|, where Fp is the protein structure factor amplitude and |Fph| is the heavy-atom derivative structure factor amplitude.
cPhasing power = r.m.s. (|Fh|/E, where |Fh| is the heavy-atom structure factor amplitude and E is the residual lack of closure error).
dMean FOM = combined figure of merit.
eR = R based on 95% of the data used in refinement. Rfree = R based on 5% of the data withheld for the cross-validation test.
Numbers in parentheses indicate last outer shell statistics.
Statistics for derivative data treat Bijvöet pairs independently.
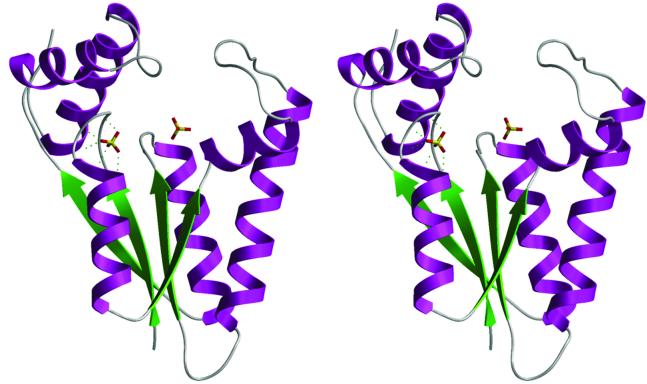
The kinase domain consists of a central four-stranded parallel β-sheet (strand order β2–β3–β1–β4) flanked by three α-helices on each side. In the view shown in Figure 2, helices α1, α5 and α6 are located on the left side of the β-sheet, whereas α2, α3 and α4 are on the right side. The kinase contains a consensus Walker A-box motif (9GxxGxGKS16) located between the first β-strand and the first α-helix. The A-box of Pnk forms a classical P-loop structure found in many nucleotide-dependent phosphotransferases. Two sulfate ions bound to Pnk, one to the P-loop and a second to α2 and the loop connecting β3 to α4, demarcate the NTP and phosphate-acceptor binding sites, respectively (see below). The kinase can be divided into two sub-domains. One module, composed of the β-sheet and helices α1, α3, α4 and α5, forms a globular pedestal supporting the two sulfates (Figure 2). A second bipartite module, composed of α5, α6 and the interconnecting loop on one side plus the α2–α3 loop on the other side, comprises a lid-like structure above the sulfate-binding sites (Figure 2). A surface view shows that the two components of the lid meet at the center and form a tunnel through the heart of the protein (Figure 3).
Fig. 2. Structure of the 5′-kinase domain. A stereo ribbon image is shown with the central β-sheet in green and flanking α helices in purple. Two enzyme-bound sulfates coordinated at the active site tunnel are shown as a stick models. Coordination of one sulfate within the oxyanion hole formed by the P-loop is depicted by dashed lines.
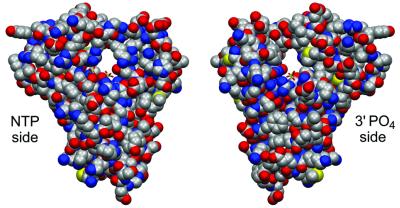
Fig. 3. Active site tunnel of Pnk. Space-filling images of the protein in CPK coloring highlight a see-through tunnel in T4 Pnk. (Left) A view looking into the tunnel from the NTP phosphate-donor side of the active site. (Right) A view from the phosphate-acceptor side. The sulfates corresponding to the β phosphate of the NTP and the 3′ phosphate of the terminal nucleotide of the 5′ OH acceptor are shown in the tunnel.
The kinase monomer of the asymmetric unit makes contact with another monomer across a crystallographic 2-fold axis. The protein–protein interface buries a surface area of 2400 Å2 (1200 Å2 per protomer) and is formed by α1, β2, α2, α3 and the α2–α3 loop (see Figure 5). The molecular contacts at the interface are predominantly hydrophobic, but also include hydrogen bonds between main chains (from the Asn31 amide to the Ile41 carbonyl), from side chain to main chain (Asn31 amide to Ser40 carbonyl) and from side chain to side chain side (Asp36 Oδ1 to His44Nε; Asp36 Oδ2 to His44Nε and Ser40 Oγ).
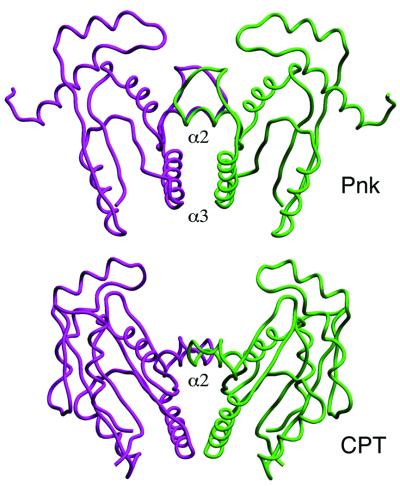
Fig. 5. Crystallographic dimer interfaces in Pnk and CPT. The backbone traces of the Pnk and CPT dimers are shown with one protomer in purple and the other protomer in green. The dimers were superimposed by a least-squares fit in program O and the images were offset vertically in SETOR.
Comparison with other phosphotransferases
A search of the protein structure database using DALI (Holm and Sander, 1993) found significant similarities between Pnk and other P-loop-containing enzymes. The two best matches were to the kinase domain of 6-phos phofructo-2-kinase/fructose-2,6-bisphosphatase (PFK-2) [Protein Data Bank (PDB) code 1bif; z score 14.0; r.m.s.d. of 2.5 Å over 140 Cα positions of equivalence to Pnk] and to chloramphenicol phosphotransferase (CPT) (PDB code 1qhs; z score 12.2; r.m.s.d. of 2.7 Å over 134 Cα positions of equivalence to Pnk) (Hasemann et al., 1996; Izard and Ellis, 2000). The next set of matches were to adenylate kinase (Adk) (PDB code 1zin; z score 11.7; r.m.s.d. of 2.5 Å over 131 Cα positions of equivalence to Pnk) and shikimate kinase (PDB code 1shk; z score 8.9; r.m.s.d. of 2.9 Å over 120 Cα positions of equivalence to Pnk) (Berry and Phillips, 1998; Krell et al., 1998). Overall structural similarity between Pnk and Adk had been predicted based on mutational results for Pnk (Wang and Shuman, 2002). Adk catalyzes phosphoryl transfer from ATP to AMP to form two molecules of ADP. The Adk reaction is superficially similar to that of Pnk insofar as it entails nucleophilic attack on the γ phosphorus of ATP and is dependent on a divalent cation cofactor. However, in the Adk reaction, the attacking nucleophile is a 5′ phosphate oxygen of AMP, whereas in the Pnk reaction it is the 5′ OH of the nucleoside sugar. Thus, Adk and Pnk are chemically distinct, entailing the interconversion of phosphoanhydride linkages in the case of Adk and the transformation of a phosphoanhydride into a phosphomonoester in the Pnk reaction. Although the structural similarity of Pnk to PFK-2, CPT and shikimate kinase had not been appreciated previously, this finding is of potential mechanistic significance given that the reactions catalyzed by Pnk, PFK-2, CPT and shikimate kinase all involve phosphoryl transfer to an alcohol moiety on the phosphate acceptor.
An alignment of the primary structures of Pnk and CPT (Figure 4) reveals 20 positions of side-chain identity plus 14 positions of side-chain similarity. The secondary structures (Figure 4) and folding topologies (Figure 5) are conserved, with the exception of the segment between helices α2 and α3, which contains an antiparallel β-hairpin in CPT that has no counterpart in Pnk. This unique segment of CPT includes constituents of the chloramphenicol binding pocket (Izard and Ellis, 2000), which would not be relevant to Pnk. A salient feature of the structural comparison is that five of the side chains that are essential for the kinase activity of Pnk (Lys15, Ser16, Asp35, Asp85 and Arg126) are identical and occupy equivalent positions in the structure of CPT (Figure 4).
Fig. 4. Structural similarity between Pnk and CPT. The secondary structures of T4 Pnk (pnk) and Streptomyces venezuelae CPT (cpt) are shown above and below their respective amino acid sequences. The crystal structures were aligned by DALI. Gaps in the alignment are indicated by dashes. The essential side chains of Pnk are highlighted in shaded boxes, which include the equivalent positions of CPT when conserved.
Higher order similarities between Pnk and CPT are revealed by the conservation of a predominantly hydrophobic dimer interface across their respective crystallographic 2-fold axes (Figure 5). Both interaction surfaces involve contributions from the equivalent secondary structure elements, particularly α2 and α3 and their adjacent loops (Figure 5). The CPT dimer superimposed on the Pnk dimer with a root-mean-square deviation (r.m.s.d.) of 3.7 Å over 268 Cα positions of equivalence to dimeric Pnk. CPT is a dimer under physiological solution conditions; Izard and Ellis (2000) proposed that the CPT subunit interface that superimposes on Pnk represents the physiological homodimerization surface. We draw similar inferences for the Pnk kinase domain.
Pnk active site
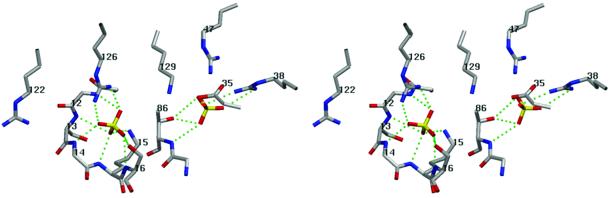
Two well-ordered sulfate ions were coordinated in the interior of the Pnk tertiary structure (Figure 2). The first sulfate engages in an extensive set of contacts with the backbone amide nitrogens of the P-loop and with the side chains of A-box residues Lys15 and Ser16 (Figure 6). Lys15 and Ser16 are both essential for 5′-kinase activity (Wang and Shuman, 2001). The network of sulfate to P-loop interactions in the Pnk crystal typifies contacts made to the nucleotide β phosphate in the structures of numerous other P-loop-containing phosphotransferases; therefore, we surmise that this sulfate demarcates the NTP binding site of Pnk. The sulfate mimicking the β phosphate is also engaged in a bidentate interaction with the terminal guanidinium nitrogens of Arg126. Arg126 is strictly essential for 5′-kinase activity and it cannot be functionally replaced by lysine, highlighting the importance of the bidentate interaction with the NTP substrate (Wang and Shuman, 2002). Arg126 is situated at the distal end of α5 and is part of the lid subdomain (formed by helices α5 and α6 and the intervening loop) that covers the NTP binding site. The equivalent Arg side chains in CPT and Adk make multiple contacts with the phosphates of ATP (Dreusicke et al., 1988; Müller and Schulz, 1992; Abele and Schulz, 1995; Berry and Phillips, 1998; Izard and Ellis, 2000). Arg126 of Pnk is situated within a motif, 122RxxxR126, that is conserved in the structures of CPT and Adk enzymes (Figure 4). The proximal arginine stacks on the adenine base of ATP in the CPT and Adk structures via a cation–π interaction. Superposition of the Pnk and CPT/ATPγS structures suggests that Arg122 of Pnk would also interact with the base of the NTP substrate.
Fig. 6. Pnk active site. Stereo view highlighting the interactions of two sulfates bound at the active site. The P-loop main-chain amides and side chains Lys15, Ser16 and Arg126 coordinate a sulfate corresponding to the β phosphate of the NTP substrate. A second sulfate corresponding to the 3′ phosphate of the 3′ NMP substrate is coordinated to the NH and Oγ of Thr86 and the side chain of Arg38. Contacts are denoted by dashed lines.
The second sulfate ion is situated 8.5 Å from the first sulfate at a position in the tertiary structure that corresponds to the phosphate acceptor sites of other P-loop-containing phosphotransferases. The second sulfate is coordinated to the main-chain amide nitrogen and the side-chain Oγ of Thr86 (in the β3–α4 loop) and to both terminal guanidinium nitrogens of Arg38 (in α2) (Figure 6). The minimal phosphate acceptor for the T4 kinase reaction is a nucleoside 3′ monophosphate. Indeed, the 3′ phosphate moiety is essential for substrate recognition, i.e. Pnk is unable to transfer the γ phosphate from ATP to nucleosides (Novogrodsky et al., 1966). Therefore, we infer that the contacts between Pnk and the second sulfate ion mimic its interactions with the 3′ PO4 of the terminal nucleotide of the acceptor substrate. This model is supported by mutagenesis results showing that the Arg38 side chain is essential for phosphorylation of 3′ CMP and that its catalytic function cannot be fulfilled by lysine (Wang and Shuman, 2002), thus underscoring the importance of the bidentate contact seen in the crystal structure. Mutational analysis indicates that the hydrogen bond of the sulfate (3′ phosphate) to the Thr86 Oγ is not functionally important (Wang and Shuman, 2001). Yet, the contact to the main-chain amide of Thr86 may be essential, insofar as replacement of the neighboring Asp85 side chain by alanine resulted in a loss of function. Asp85 (in β3) forms a bidentate hydrogen bond with the main-chain amide of Arg34 (α2) and the side-chain amide nitrogen of Gln64 (α4) and may thereby tether the Thr86-NH in the correct position to bind the substrate. An aspartate is conserved at the equivalent position (Asp93) of CPT (Figure 4).
Structural insights into the kinase catalytic mechanism
The stereochemical course of the 5′-kinase reaction of T4 Pnk entails inversion of configuration of the transferred phosphate, indicative of an in-line mechanism in which the polynucleotide 5′ OH directly attacks the γ phosphorus of ATP (Jarvest and Lowe, 1981). This scheme implies a means to activate the 5′ OH nucleophile, presumably by abstraction of its proton. Although the Pnk structure does not include a nucleoside, we can infer its location on the side of the 3′ phosphate (sulfate 2) that faces toward the NTP binding site. Asp35, which is essential for catalysis, is located on this side of the sulfate and is a likely candidate to coordinate the 5′ OH and activate its attack on the γ phosphorus. The complete loss of activity when Asp35 is replaced by Asn is consistent with its proposed function as a general base catalyst. The Asp35 equivalent is conserved in the α2 helix of CPT (Figure 4), where it is hydrogen-bonded to the C3-hydroxyl group of chloramphenicol to which the phosphate is transferred. Izard and Ellis (2000) suggested that this Asp could function as a general base catalyst in the CPT reaction. An Asp is present at the equivalent position in α2 of shikimate kinase, where it is poised to coordinate the reactive C3-hydroxyl of shikimate (Krell et al., 1998). A general base catalyst in α2 appears to be a conserved feature of P-loop kinases that transfer phosphate to either a primary (Pnk and CPT) or secondary (shikimate kinase) alcohol.
We presume that the T4 Pnk reaction is promoted by contacts between the enzyme and the γ phosphate of ATP that serve to stabilize a pentacoordinate transition state. The essential Lys15 and Arg126 side chains are poised for such a role. In addition, the Pnk crystal structure shows that Lys129 and Arg47 project downward from the lid toward the active site (Figure 6). To evaluate whether Lys129 or Arg47 contribute to Pnk activity, we replaced them individually with alanine and purified the recombinant full-length K129A and R47A proteins from bacteria. The specific activity of the K129A and R47A mutants in phosphorylating 3′ CMP were 51 and 77% of the activity of wild-type Pnk, respectively (data not shown). Thus, neither Lys129 nor Arg47 is important for catalysis.
The lid domain overhanging the substrate binding sites encloses a tunnel through the enzyme (Figure 3). We envision that entrance to the tunnel on the P-loop side admits the NTP phosphate donor and controls release of the NDP reaction product. The close contacts inferred between the nucleotide and both the P-loop and the overhanging lid suggest conformational transitions, whereby the lid is raised to allow ATP binding, closed over the bound ATP to promote phosphoryl transfer, and then raised for product release. The tunnel opening on the opposite face of the protein, extending from the 3′-phosphate site (sulfate 2) out to the surface, is lined with basic side chains (Arg92, Lys51 and Lys54) and affords a plausible path for ingress of the 5′ end of a polynucleotide to the T4 kinase active site.
Clues to T4 Pnk quaternary structure
The biochemical and crystallographic results presented here engender a model for how the 5′-kinase and 3′-phosphatase enzymatic domains might be arranged in the native T4 Pnk homotetramer (Figure 7). The C-terminal half of the Pnk protomer, comprising an autonomous phosphatase catalytic domain (‘P’ in Figure 7), forms a stable homodimer in solution at low protein concentrations in the absence of its substrate. Thus, Pnk tetramers may be assembled from two pre-formed homodimers via inter-dimer contacts between the N-terminal kinase domains (‘K’ in Figure 7), presumably similar to those observed across the symmetry axis in the crystal of the isolated kinase domain. Although glycerol gradient experiments had suggested a monomeric structure for the isolated kinase domain, the sedimentation analysis was performed at much lower protein concentration and under very different solute conditions than those employed for crystallization of the kinase domain. Reports in the literature of the native size of intact Pnk under different conditions of analysis suggest an equilibrium exists between tetramer and dimer states (Panet et al., 1973; Cameron and Uhlenbeck, 1977; Lillehaug, 1977; Soltis and Uhlenbeck, 1982a; Wang and Shuman, 2001), with the tetrameric form being stabilized by high ionic strength and by the binding of the NTP and 3′ NMP substrates of the kinase reaction (Lillehaug, 1977). The high ionic strength of the kinase domain crystallization buffer and the occupancy of both substrate sites by sulfate may have stabilized a homodimer of the isolated kinase domain in the crystal, whereas the interactions of the N-terminal domain in standard purification buffers would require prior dimerization of the phosphatase domains.

Fig. 7. A model of T4 Pnk quaternary structure. We speculate that native Pnk homotetramer is assembled via high-affinity interactions between the C-terminal phosphatase domains (P) to form Pnk homodimers that are then converted into tetramers via lower-affinity interactions between the N-terminal kinase domains (K). The tetramer is depicted in a square planar arrangement for the sake of clarity.
Structural insights into substrate specificity and the relationship between T4 and eukaryotic Pnk
Mammals and fission yeast contain a bifunctional Pnk enzyme with 5′-kinase and 3′-phosphatase activities analogous to those of T4 Pnk. However, whereas T4 Pnk phosphorylates 5′ OH ends of RNA, DNA and nucleo side 3′ monophosphates, the eukaryotic Pnk enzymes are specific for 5′ OH DNA substrates. The primary structures of the kinase components of the T4 and eukaryotic Pnks are similar and four of the five amino acids that are essential for catalysis and comprise the active site in the T4 Pnk crystal structure (Lys15, Ser16, Asp35 and Arg126) are conserved in human and Schizosaccharomyces pombe Pnks (Jilani et al., 1999a; Karimi-Busheri et al., 1999; Meijer et al., 2002). Thus, we surmise that T4 and eukaryotic Pnk kinase domains are likely to have similar folds with highly conserved binding sites for the NTP phosphate donor and a similar mechanism of catalysis involving an essential aspartate side chain (equivalent to Asp35 in T4 Pnk). The striking difference is that none of the eukaryotic Pnk proteins contains a basic side chain at the position corresponding to Arg38 of T4 Pnk, an essential residue that makes a bidentate contact with the 3′ phosphate of the terminal nucleotide of the phosphate acceptor. The existence of a dedicated electrostatic interaction with the 3′ phosphate almost certainly accounts for the ability of T4 Pnk to phosphorylate 3′ NMPs, whereas the lack of an equivalent arginine in the mammalian Pnk suffices to explain its failure to utilize 3′ dNMP as a substrate (Jilani et al., 1999b). Indeed, mammalian Pnk requires that the phosphate acceptor have a minimum chain length of approximately eight deoxynucleotides for the kinase reaction to occur (Karimi-Busheri and Weinfeld, 1997).
Other fundamental differences in the phosphate acceptor specificities of T4 versus mammalian Pnks are clarified by the structure of the T4 domain. Although T4 Pnk is widely used to label DNA 5′ ends in vitro, it is actually quite selective in phosphorylating DNAs with 5′ single-strand extensions; T4 Pnk is poorly active or inactive on blunt duplex 5′ OH termini or 5′ OH termini that are recessed within duplex regions or at the junction of a 3′ single-strand tail (Jilani et al., 1999b). The crystal structure of the T4 kinase domain indicates that the tunnel aperture leading to the 3′ phosphate binding site of the phosphate acceptor (Figure 3) is too narrow to allow facile ingress of duplex nucleic acid, but can readily accommodate a single-stranded polynucleotide. The physiological substrates of T4 Pnk in vivo are broken tRNAs incised 5′ to the wobble base of the anticodon loop (Amitsur et al., 1987). The resulting 5′ OH end at the break is free to enter the T4 kinase active site as a five nucleotide single-strand extension coming off the anticodon stem. Unlike T4 Pnk, the mammalian kinase preferentially phosphorylates 5′ ends recessed within duplex DNA structures (Karimi-Busheri and Weinfeld, 1997), implying that the phosphate acceptor site of the mammalian Pnk is fundamentally different from that of the phage T4 enzyme.
The most obvious structural differences between the T4 and eukaryotic Pnks are that (i) the H2N-kinase/phosphatase-COOH domain order of T4 Pnk is inverted in the eukaryotic enzymes as H2N-phosphatase/kinase-COOH (Jilani et al., 1999a; Karimi-Busheri et al., 1999; Meijer et al., 2002) and (ii) T4 Pnk is a 140 kDa homotetramer, whereas the human Pnk is a 60 kDa monomer (Mani et al., 2001). The evolutionary forces shaping the divergence in both domain order and quaternary structure in ‘RNA repair’ versus ‘DNA repair’ Pnks are unclear and their elucidation will hinge on obtaining additional atomic structures of the native enzymes from diverse sources.
Materials and methods
Crystallization and data collection
Recombinant His-Pnk(1–181) was purified from soluble E.coli extracts by Ni–agarose chromatography as described previously (Wang and Shuman, 2001), except that Triton X-100 was omitted from the lysis and column buffers. Crystals of His-Pnk(1–181) were grown at 22°C by the hanging drop vapor diffusion method. The protein sample (7 mg/ml in 50 mM Tris–HCl pH 7.5, 0.2 M NaCl, 0.3 M imidazole, 10% glycerol) was mixed with an equal volume of the reservoir buffer containing 100 mM of sodium acetate pH 4.6, 5 mM DTT, 16% PEG-2000 monomethyl ether and 0.4 M ammonium sulfate. Crystals were grown over 2 days to 1 week. Prior to diffraction, the crystals were cryo-preserved in reservoir buffer containing 30% glycerol and then flash-frozen in liquid nitrogen. X-ray diffraction data for a native crystal were collected at the National Synchrotron Light Source (Brookhaven, NY) at beamline X9A using an MAR CCD detector. Diffraction data for a mercury acetate derivative were collected at a laboratory copper Kα source (Rigaku RU200) equipped with a confocal Osmic multilayer system and a Raxis-IV imaging plate detector system. Data were integrated, scaled and merged using DENZO and SCALEPACK (Otwinowski and Minor, 1997). Pnk(1–181) crystallized in space group C2221 (a = 65.3 Å, b = 91.8 Å, c = 67.9 Å; α, β and γ = 90°).
Structure determination and refinement
The Pnk structure was solved by single isomorphous replacement with anomalous scattering. Phases (2.0Å) were calculated with SOLVE/RESOLVE (Terwilliger and Berendzen, 1999) and the CCP4 suite (Collaborative Computational Project, 1994). The model was built manually into the electron density map using the program O (Jones et al., 1991). Refinement was performed with CNS (Brunger et al., 1998). The refined model at 2.0 Å resolution (Rfree = 23.3, R = 21.0) consists of Pnk residues 1–152, 142 waters and four sulfates. No electron density was observed for the N-terminal His-tag and for amino acids 153–181. All of the amino acids are found within the most favored or allowed regions in the Ramachandran plot. Refinement statistics are listed in Table I. The coordinates are deposited in the Protein Data Bank under code 1LY1. Images of the Pnk structure were prepared with SETOR (Evans, 1993).
Pnk mutants
N-terminal deletion mutants Pnk(139–301) [NΔ138], Pnk(149–301) [NΔ148] and Pnk(159–301) [NΔ158] were constructed by PCR amplification with mutagenic sense-strand primers that introduced an NdeI restriction site and a methionine codon in lieu of the codon for 138, 148 or 158. Alanine mutations were introduced into the full-length Pnk gene by using the two-stage PCR-based overlap extension method. The PCR products were digested with NdeI and BamHI and then inserted into pET16b. The inserts were sequenced completely to confirm the desired mutations and to exclude the acquisition of unwanted changes. The pET-PnkΔ and pET-Pnk-Ala plasmids were introduced into E.coli BL21(DE3)-CodonPlus and the truncated or Ala-substituted Pnk were purified from soluble bacterial lysates as described for the wild-type Pnk (Wang and Shuman, 2001). Protein concentrations were determined by using the Bio-Rad dye reagent with BSA as the standard.
5′-kinase assay
Reactions mixtures (10 µl) containing 70 mM Tris–HCl pH 7.6, 10 mM MgCl2, 5 mM DTT, 25 µM [γ-32P]ATP, 1 mM 3′ CMP (Sigma) and Pnk as specified were incubated for 20 min at 37°C. Aliquots of the mixtures were applied to a polyethyleneimine-cellulose TLC plate, which was developed with 1 M formic acid and 0.5 M LiCl. The [γ-32P]ATP substrate and [α-32P]pCp product were visualized and quantitated by scanning the gel with a Fujix BAS2500 phosphoimager.
3′-phosphatase assay
Reaction mixtures (25 µl) containing 100 mM imidazole pH 6.0, 10 mM MgCl2, 10 mM β-mercaptoethanol, 0.1 mg/ml BSA, 1.6 mM 3′ dTMP (Sigma) and Pnk as specified were incubated for 20 min at 37°C. The reactions were quenched by adding 75 µl of cold water and 1 ml of malachite green reagent (BIOMOL Research Laboratories). Phosphate release was determined by measuring A620 and extrapolating the value to a phosphate standard curve.
Velocity sedimentation
Aliquots (40 µg) of the PnkΔ preparations were mixed with BSA (30 µg), ovalbumin (30 µg) and cytochrome c (30 µg) and the mixtures were applied to 4.8-ml 15–30% glycerol gradients containing 50 mM Tris–HCl pH 8.0, 0.2 M NaCl, 1 mM EDTA, 2.5 mM DTT and 0.1% Triton X-100. The gradients were centrifuged in a SW50 rotor at 50 000 r.p.m. for 24 h at 4°C. Fractions were collected from the bottoms of the tubes.
Acknowledgments
Acknowledgements
We thank the staff of beamline X9A at the National Synchrotron Light Source. This research was supported by NIH grant GM42498 (to S.S.) and a Young Investigator Award from the Arnold and Mabel Beckman Foundation (to C.D.L.).
References
- Abele U. and Schulz,G.E. (1995) High-resolution structures of adenylate kinase from yeast liganded with inhibitor Ap5A, showing the pathway of phosphoryl transfer. Protein Sci., 4, 1262–1271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abelson J., Trotta,C.R. and Li,H. (1998) tRNA splicing. J. Biol. Chem., 273, 12685–12688. [DOI] [PubMed] [Google Scholar]
- Amitsur M., Levitz,R. and Kaufman,G. (1987) Bacteriophage T4 anticodon nuclease, polynucleotide kinase, and RNA ligase reprocess the host lysine tRNA. EMBO J., 6, 2499–2503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Becker A. and Hurwitz,J. (1967) The enzymatic cleavage of phosphate termini from polynucleotides. J. Biol. Chem., 242, 936–950. [PubMed] [Google Scholar]
- Berry M.B. and Phillips,G.N. (1998) Crystal structures of Bacillus stearothermophilus adenylate kinase with bound Ap5A, Mg2+ Ap5A, and Mn2+ Ap5A reveal an intermediate lid position and six coordinate octahedral geometry for bound Mg2+ and Mn2+. Proteins, 32, 276–288. [DOI] [PubMed] [Google Scholar]
- Brunger A.T. et al. (1998) Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D, 54, 905–921. [DOI] [PubMed] [Google Scholar]
- Cameron V. and Uhlenbeck,O.C. (1977) 3′-Phosphatase activity in T4 polynucleotide kinase. Biochemistry, 16, 5120–5126. [DOI] [PubMed] [Google Scholar]
- Collaborative Computational Project (1994) The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D, 50, 760–763. [DOI] [PubMed] [Google Scholar]
- Dreusicke D., Karplus,P.A. and Schulz,G.E. (1988) Refined structure of porcine cytosolic adenylate kinase at 2.1 Å resolution. J. Mol. Biol., 199, 359–371. [DOI] [PubMed] [Google Scholar]
- Evans S.V. (1993) SETOR: hardware-lighted three-dimensional solid model representations of macromolecules. J. Molec. Graph., 11, 134–138. [DOI] [PubMed] [Google Scholar]
- Hasemann C.A., Istvan,E.S., Uyeda,K. and Deisenhofer,J. (1996) The crystal structure of the bifunctional enzyme 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase reveals distinct domains homologies. Structure, 4, 1017–1029. [DOI] [PubMed] [Google Scholar]
- Holm L. and Sander,C. (1993) Protein structure comparison by alignment of distant matrices. J. Mol. Biol., 233, 123–138. [DOI] [PubMed] [Google Scholar]
- Izard T. and Ellis,J. (2000) The crystal structures of chloramphenicol phosphotransferases reveal a novel inactivation mechanism. EMBO J., 11, 2690–2700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jarvest R.L. and Lowe,G. (1981) The stereochemical course of the phosphoryl transfer reaction catalyzed by polynucleotide kinase (bacteriophage-T4-infected Escherichia coli B). Biochem. J., 199, 273–276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jilani A., Ramotar,D., Slack,C., Ong,C., Yang,X.M., Scherer,S.W. and Lasko,D.D. (1999a) Molecular cloning of the human gene, PNKP, encoding a polynucleotide kinase 3′-phosphatase and evidence for its role in repair of DNA strand breaks caused by oxidative damage. J. Biol. Chem., 274, 24176–24186. [DOI] [PubMed] [Google Scholar]
- Jilani A., Slack,C., Matheos,D., Zannis-Hadjopoulos,M. and Lasko,D.D. (1999b) Purification of a polynucleotide kinase from calf thymus, comparison of its 3′-phosphatase domain with T4 polynucleotide kinase, and investigation of its effect on DNA replication in vitro. J. Cell. Biochem., 73, 188–203. [PubMed] [Google Scholar]
- Jones T.A., Zou,J.Y., Cowan,S.W. and Kjeldgaard,M. (1991) Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A, 47, 110–118. [DOI] [PubMed] [Google Scholar]
- Karimi-Busheri F. and Weinfeld,M. (1997) Purification and substrate specificity of polydeoxynucleotide kinases isolated from calf thymus and rat liver. J. Cell. Biochem., 64, 258–272. [PubMed] [Google Scholar]
- Karimi-Busheri F. et al. (1999) Molecular characterization of a human DNA kinase. J. Biol. Chem., 274, 24187–24194. [DOI] [PubMed] [Google Scholar]
- Krell T., Coggins,J.R. and Lapthorn,A.J. (1998) The three-dimensional structure of shikimate kinase. J. Mol. Biol., 278, 983–997. [DOI] [PubMed] [Google Scholar]
- Lillehaug J.R. (1977) Physicochemical properties of T4 polynucleotide kinase. Eur. J. Biochem., 73, 499–506. [DOI] [PubMed] [Google Scholar]
- Mani R.S., Karimi-Busheri,F., Cass,C.E. and Weinfeld,M. (2001) Physical properties of human polynucleotide kinase: hydrodynamic and spectroscopic studies. Biochemistry, 40, 12967–12973. [DOI] [PubMed] [Google Scholar]
- Meijer M., Karimi-Busheri,F., Huang,T.Y., Weinfeld,M. and Young,D. (2002) Pnk1, a DNA kinase/phosphatase required for normal response to DNA damage by γ-radiation or camptothecin in Schizosaccharomyces pombe. J. Biol. Chem., 277, 4050–4055. [DOI] [PubMed] [Google Scholar]
- Midgley C.A. and Murray,N.E. (1985) T4 polynucleotide kinase; cloning of the gene (pseT) and amplification of its product. EMBO J., 4, 2695–2703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Müller C.W. and Schulz,G.E. (1992) Structure of the complex between adenylate kinase from Escherichia coli and the inhibitor Ap5A refined at 1.9 Å resolution: a model for a catalytic transition state. J. Mol. Biol., 224, 159–177. [DOI] [PubMed] [Google Scholar]
- Novogrodsky A. and Hurwitz,J. (1966) The enzymatic phosphorylation of ribonucleic acid and deoxyribonucleic acid: phosphorylation at 5′-hydroxyl termini. J. Biol. Chem., 241, 2923–2932. [PubMed] [Google Scholar]
- Novogrodsky A., Tal.,M., Traub,A. and Hurwitz,J. (1966) The enzymatic phosphorylation of ribonucleic acid and deoxyribonucleic acid: further properties of the 5′-hydroxyl polynucleotide kinase. J. Biol. Chem., 241, 2933–2943. [PubMed] [Google Scholar]
- Otwinowski Z. and Minor,W. (1997) Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol., 276, 307–326. [DOI] [PubMed] [Google Scholar]
- Panet A., van de Sande,J.H., Loewen,P.C., Khorana,H.G., Raae,A.J., Lillehaug,J.R. and Kleppe,K. (1973) Physical characterization and simultaneous purification of bacteriophage T4 induced polynucleotide kinase, polynucleotide ligase, and deoxyribonucleic acid polymerase. Biochemistry, 12, 5045–5050. [DOI] [PubMed] [Google Scholar]
- Richardson C.C. (1965) Phosphorylation of nucleic acid by an enzyme from T4 bacteriophage-infected Escherichia coli. Proc. Natl Acad. Sci. USA, 54, 158–165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soltis D.A. and Uhlenbeck,O.C. (1982a) Isolation and characterization of two mutant forms of T4 polynucleotide kinase. J. Biol. Chem., 257, 11332–11339. [PubMed] [Google Scholar]
- Soltis D.A. and Uhlenbeck,O.C. (1982b) Independent location of kinase and 3′-phosphatase activities on T4 polynucleotide kinase. J. Biol. Chem., 257, 11340–11345. [PubMed] [Google Scholar]
- Terwilliger T.C. and Berendzen,J. (1999) Automated MAD and MIR structure solution. Acta Crystallogr. D, 55, 849–861. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang L.K. and Shuman,S. (2001) Domain structure and mutational analysis of T4 polynucleotide kinase. J. Biol. Chem., 276, 26868–26874. [DOI] [PubMed] [Google Scholar]
- Wang L.K. and Shuman,S. (2002) Mutational analysis defines the 5′-kinase and 3′-phosphatase active sites of T4 polynucleotide kinase. Nucleic Acids Res., 30, 1073–1080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whitehouse C.J., Taylor,R.M., Thistlethwaite,A., Zhang,H.M., Karimi-Busheri,F., Lasko,D.D., Weinfeld,M. and Xaldecott,K.W. (2001) XRCC1 stimulates human polynucleotide kinase activity at damaged DNA termini and accelerates DNA single-strand break repair. Cell, 104, 107–117. [DOI] [PubMed] [Google Scholar]