Abstract
The environmental distribution of Dehalococcoides group organisms and their association with chloroethene-contaminated sites were examined. Samples from 24 chloroethene-dechlorinating sites scattered throughout North America and Europe were tested for the presence of members of the Dehalococcoides group by using a PCR assay developed to detect Dehalococcoides 16S rRNA gene (rDNA) sequences. Sequences identified by sequence analysis as sequences of members of the Dehalococcoides group were detected at 21 sites. Full dechlorination of chloroethenes to ethene occurred at these sites. Dehalococcoides sequences were not detected in samples from three sites at which partial dechlorination of chloroethenes occurred, where dechlorination appeared to stop at 1,2-cis-dichloroethene. Phylogenetic analysis of the 16S rDNA amplicons confirmed that Dehalococcoides sequences formed a unique 16S rDNA group. These 16S rDNA sequences were divided into three subgroups based on specific base substitution patterns in variable regions 2 and 6 of the Dehalococcoides 16S rDNA sequence. Analyses also demonstrated that specific base substitution patterns were signature patterns. The specific base substitutions distinguished the three sequence subgroups phylogenetically. These results demonstrated that members of the Dehalococcoides group are widely distributed in nature and can be found in a variety of geological formations and in different climatic zones. Furthermore, the association of these organisms with full dechlorination of chloroethenes suggests that they are promising candidates for engineered bioremediation and may be important contributors to natural attenuation of chloroethenes.
The chloroethenes tetrachloroethene (PCE) and trichloroethene (TCE) are commonly used organic solvents and degreasing agents. As a result of past disposal practices and spills, chloroethenes are now widely distributed in the environment and are found in many sediments, soils, groundwater aquifers, and subsurface environments throughout the world (13, 21, 28). Traditional approaches to groundwater remediation practices, such as pump and treat methods, have been shown to be ineffective and costly when applied to chlorinated solvent plumes (2a, 2c, 23). Therefore, there is a need for microbe-based remediation approaches that could provide an inexpensive way to clean up chlorinated solvent contamination.
Chloroethene solvents were previously believed to be resistant to degradation by microorganisms. There now is rapidly accumulating laboratory and field evidence that microorganisms can transform chloroethenes to nontoxic products under a variety of environmental conditions (3, 8-10, 12, 15, 20, 22, 27, 28, 30, 31, 35). Maymó-Gatell et al. have isolated an organism from an anaerobic dechlorinating laboratory culture inoculated with municipal sewage sludge, which fully dechlorinates chlorinated ethenes (25, 26). This organism, Dehalococcoides ethenogenes strain 195, was shown to meet its energy needs, which are essential for growth, by a process known as dehalorespiration. In this process, chloroethenes function as electron acceptors and hydrogen is used as an electron donor (26). Further characterization of the pathway has shown that PCE is stoichiometrically converted to vinyl chloride (VC) in a zero-order process in which there is little accumulation of TCE and 1,2-cis-dichloroethene (cDCE) intermediates (25, 26). The organism further metabolizes VC to ethene, but by a slower first-order process in which PCE is depleted before dechlorination of VC begins (25, 26). Strain 195 can also use 1,1-DCE and 1,2-dichloroethane as electron acceptors (25, 26).
To date, D. ethenogenes strain 195 is the only known isolated organism capable of fully dechlorinating PCE and the other chloroethenes to ethene (1, 23, 26). Other Dehalococcoides strains have been found to be associated with enriched mixed cultures that anaerobically dechlorinate chloroethenes. These organisms, strains FL2 (23) and DCEH2 (GenBank accession number AJ249262), were detected in cultures as 16S rRNA sequences by either a PCR assay or community analysis. Analysis of their 16S rRNA gene (rDNA) sequences showed that both sequences were phylogenetically related to the D. ethenogenes strain 195 sequence (23; GenBank accession number AJ249262). Löffler et al. (23) used a two-step nested primer PCR approach to detect the Dehalococcoides sequences. In the nested assay a bacterial universal primer set is used first, and then a second PCR in which a Dehalococcoides group primer set is used is performed. Löffler et al. also used this approach to detect Dehalococcoides populations in river sediments in Michigan and a chloroethene dissolved plume near Jacksonville, Fla.
Recently, a second organism, strain CBDB1, was isolated and was reported to be phylogenetically related to D. ethenogenes strain 195, as determined by 16S rDNA sequence analysis (1). Like strain 195, CBDB1 meets its energy needs by a dehalorespiratory process in which hydrogen is used as an electron donor, but it uses the higher substituted chlorinated benzenes as electron acceptors. Strain CBDB1 stoichiometrically dechlorinates 1,2,3-trichlorobenzene (1,2,3-TCB), 1,2,4-TCB, 1,2,3,4-tetrachlorobenzene (1,2,3,4-TeCB), 1,2,3,5-TeCB, or 1,2,4,5-TeCB to dichlorobenzenes or 1,3,5-TCB (1). When tested, a CBDB1 culture was unable to dechlorinate PCE or abiotically converted TCE [reduced by Ti(III) in the buffer used in the experiment] to cDCE, VC, and ethene (1). VC and cDCE, however, were not tested in the dechlorination experiments. This analysis demonstrated that not all members of the genus Dehalococcoides use the higher chlorinated ethenes as electron acceptors.
Notwithstanding the promise of laboratory and environmental studies, little is known about the prevalence of Dehalococcoides strains in the environment, their geographical distribution, or their functional role in dechlorination processes. If Dehalococcoides organisms are capable of widespread survival in the environment, their favorable metabolic activity suggests that they could be promising candidates for engineered bioremediation and bioaugmentation and important contributors to natural attenuation of chloroethenes or the higher chlorinated benzenes.
The goals of this study were to evaluate how widely distributed Dehalococcoides strains are in the environment and to determine their association with dechlorination at chloroethene-contaminated sites. A final goal was to study the role of Dehalococcoides in chloroethene dechlorination.
MATERIALS AND METHODS
Chemicals.
High-performance liquid chromatography grade TCE and methanol were obtained from Fisher Scientific, Inc. (Springfield, N.J.). PCE was obtained from J. T. Baker (Phillipsburg, N.J.). VC and 1,2-trans-DCE were obtained from Aldrich Chemical Co. (Milwaukee, Wis.). 1,2-cDCE was obtained from Supelco Co. (Bellefonte, Pa.). Helium, argon, mixed gases (5% H2, 10% CO2, 85% N2), ethene ethane, and methane were purchased from Keen Compressed Gas Co. (Wilmington, Del.). All of the other chemicals used were reagent grade or better. All reagents and the Taq enzyme used for PCR and sequencing were obtained from Perkin-Elmer (Norwalk, Conn.). All materials used for growth and maintenance of microcosms were obtained from Aldrich Chemicals, Difco Laboratories (Detroit, Mich.), or Sigma Chemical Company (St. Louis, Mo.) unless otherwise specified.
Procedure for soil sampling.
Two procedures were used to obtain soil samples from test sites. Some soil core samples were obtained by split spoon sampling at depths ranging from 3 to 25 m, depending on the depth of the aquifer tested. The cores were shipped on ice in stainless steel cylinders or in sterile glass jars to the laboratory. Other soil core samples were obtained by using a Geoprobe system (Geoprobe Systems, Salina, Kans.). A coring tube was pounded to a depth of 3 to 15 m and retrieved. A sterile acrylic sleeve was then placed in the tube. The tube was reinserted into the ground and driven in an additional 0.7 m. The acrylic sleeve was then removed, capped, sealed with duct tape, and stored on ice for transport to the laboratory. Upon arrival all samples were stored in an anaerobic glove bag (chamber) (Coy Laboratory Products Inc., Ann Arbor, Mich.), whose atmosphere consisted of 5% H2, 10% CO2, and 85% N2. Sites where soil samples were taken are shown in Table 1.
TABLE 1.
Characteristics and data for culture, soil, and groundwater samples
| Location | Site | Sample matrixa | Dechlorination end productb | Dehalococcoides detectedc | 16S rDNA sequencedd | Phylogenetic subgroupe | GenBank accession no. |
|---|---|---|---|---|---|---|---|
| Characterized microcosm cultures | |||||||
| Largo, Fla. (Pinellas) | DOE sitef | SA | Ethene | + | Yes | Pinellas | AF388548 |
| Victoria, Tex. | Industrial site | SA | Ethene | + | Yes | Victoria | AF388550 |
| Southern Ontario, Canada | Industrial site | SCA | Ethene | + | Yes | Pinellas | AF388540 |
| Cornell | AF388539 | ||||||
| Soil samples used for microcosm cultures in this study | |||||||
| Beaumont, Tex.g | Industrial site | SP | Ethene | + | Yes | Victoria | AF388533 |
| Cornell | AF388531 | ||||||
| Pinellas | AF388532 | ||||||
| DeLisle, Miss. | Industrial site | SA | Ethene | + | Yes | Victoria | AF388536 |
| Dover, Del. | Dover AFB | LFSA | Ethene | + | Yes | Victoria | AF388542 |
| Dover, Del. | Dover AFB | SA | cDCE | −h | N/Ai | N/A | |
| Dover, Del.j | Dover AFB | SABP | Ethene | + | Yes | Pinellas | AF388535 |
| Newark, Ohio | Industrial site | SA | Ethene | + | Yes | Cornell | AF388547 |
| Pompton Lakes, N.J. | Industrial site | SA | Ethene | + | Yes | Victoria | AF388549 |
| San Antonio, Tex. | Kelly AFB | LFC | Ethene | + | Yes | Cornell | AF388541 |
| Winfield, Kans. | Industrial site | SA | Ethene | + | Yes | Victoria | AF388538 |
| Groundwater samples | |||||||
| Amsterdam, Holland | Industrial site | SA | Ethene | + | Yes | Pinellas | AF388530 |
| Central Holland | Industrial site | SA | Ethene | + | Yes | Pinellas | AF388534 |
| Cheshire, northern England | Industrial site | SA | Ethene | + | No | Pinellas | |
| Dordrecht, Holland | Industrial site | SAP | Ethene | + | No | Pinellas | |
| Kent, Wash. | Industrial site | SA | Ethene | + | No | Pinellas | |
| Lorenz, Calif. | Industrial site | SA | Ethene | + | Yes | Pinellas | AF388543 |
| Niagara Falls, N.Y., site 1 | Industrial site | FB | Ethene | + | Yes | Pinellas | AF388545 |
| Cornell | AF388544 | ||||||
| Niagara Falls, N.Y., site 2 | Industrial site | LFFB | Ethene | + | No | Pinellas | |
| Niagara Falls, N.Y., site 3 | Defense plant | FB | Ethene | + | Yes | Pinellas | AF388546 |
| Sacramento, Calif. | Defense plant | SA | cDCE | −k | N/A | N/A | |
| San Antonio, Tex. | Kelly AFB | SA | cDCE | − | N/A | N/A | |
| San Antonio, Tex.j | Kelly AFB | SABP | Ethene | + | Yes | Pinellas | AF388537 |
Soil or earth matrices of the chloroethene-contaminated sites. SA, sandy aquifer; SCA, sandy clay aquifer; SP, sludge pond; LFSA, landfill on a sandy aquifer; SABP, sandy aquifer bioaugmentation pilot area; LFC, landfill on clay soil; SAP, a sandy aquifer interlayered with peat; FB, fractured bedrock; LFFB, landfill on fractured bedrock.
The terminal end products detected by chemical analysis, as reported by site stewards.
+, PCR with primers specific for Dehalococcoides yielded the desired PCR product; −, no product obtained.
The presence of complete Dehalococcoides 16S rDNA sequences was verified by sequencing both strands (Yes). For some samples, the presence of the sequences was not verified for both strands (No).
Dehalococcoides phylogenetic subgroup.
DOE, Department of Energy.
The sample consisted of sludge from the bottom of a pond.
A visible band was not obtained as a product with the DHC PCR assay.
N/A, there was no PCR product to sequence and analyze.
Samples were taken from the bioaugmentation pilot experimental area at this site.
A visible band was not obtained as a product with the DHC PCR assay. A nested primer approach was used, in which universal primers were used for the initial amplification and then DHC PCR primers were used. A PCR product was also not obtained in the second DHC PCR assay.
Procedure for obtaining groundwater samples.
Groundwater samples were taken from chloroethene-contaminated sites that included a variety of aquifer matrices, including sand, peat, clay, and fractured bedrock (Table 1). Plastic sampling bottles that were filled to the top with 250 to 1,000 ml of groundwater from monitoring wells and were double bagged were shipped to the lab on ice. Upon arrival, the samples were either stored overnight at 4°C or immediately centrifuged at 9,000 × g by using a GSA or G-3 rotor and an RC5B Sorvall Superspeed centrifuge. The resulting cell-soil pellets were resuspended in 2 ml of 1× PBS (10 mM sodium phosphate, 150 mM sodium chloride; pH 7.6) and either stored at −20°C or extracted to obtain community DNA. Sites where groundwater samples were taken are shown in Table 1.
Microcosm preparation.
Microcosms were prepared in 250-ml Wheaton bottles (Wheaton Co., Millville, N.J.) in the Coy anaerobic chamber. Microcosms were prepared with a minimal salt medium, BTZ-3 medium (4.3 g of NH4Cl per liter, 50 g of KH2PO4 per liter, 20 g of MgCl·6H2O per liter, 1 g of CaCl2·2H2O per liter, 50 mM HEPES) (10 ml of mineral solution per liter, 5 ml of resazurin per liter) (31). Microcosms were amended with 0.05% yeast extract. For each experiment, duplicate microcosms were prepared for the following conditions: killed control (live soil autoclaved for 1 h on two consecutive days), live soil, and live soil plus 0.05% yeast extract. Each microcosm contained 20% (wt/vol) soil and 80% BTZ-3 medium. The microcosm bottles were filled with little or no headspace. They were then closed with Teflon-lined septa and crimp sealed with aluminum seals (Wheaton Co.). Resazurin was used as a visual indicator of low-redox-potential conditions. Microcosms were amended with 5 ppm of PCE or 5 ppm of TCE from stock solutions that were saturated with water. KB1, a dechlorinating culture obtained from Elizabeth Edwards (University of Toronto), was transferred into fresh microcosm minimal salt medium, and the culture was amended with 10 ppm of TCE and 100 ppm of methanol. The microcosms were incubated on their sides in the anaerobic chamber in the dark at room temperature (22°C). Samples were analyzed the next day (time zero) and then twice a week for 180 days for dechlorination of PCE or TCE and formation of cDCE, VC, or ethene. Cell growth was determined either by visually observing an increase in the turbidity of the microcosm medium or by microscopic examination.
Analysis of chloroethenes and ethene.
Chloroethene and ethene contents were determined by using a Hewlett-Packard 5890 Series II gas chromatograph fitted with a ChromPack CP-PoraPLOT Q-HT fused silica WCOT column (length, 25 m; inside diameter, 0.53 mm; df (film thickness), 20 μm (catalog no. 7559); CP 7559; Varian USA, Walnut Creek, Calif.) and a flame ionization detector. The injector temperature was set at 200°C, and the detector temperature was set at 250°C. The oven temperature was programmed as follows: 45°C, increased to 180°C at a rate of 30°C/min, 180°C for 8 min, increased to 195°C at a rate of 25°C/min, 195°C for 4 min, increased to 250°C at a rate of 25°C/min, and 250°C for 10 min. The carrier gas was helium at a flow rate of 10 ml/min. Aqueous samples (5 ml) were dispensed into 10-ml headspace vials (HP 5182-0838), which were sealed with Teflon-coated butyl septa with aluminum crimp caps (HP 5183-4480). The headspace vials were placed in the vial tray of a Hewlett-Packard headspace autosampler (HP 7694) for subsequent automated injection of the headspace (gas phase) samples into the gas chromatograph. The headspace sampler was programmed to heat each sample at 80°C for 13 min prior to injection. Calibration was performed with custom-made external standards purchased from Ultra Scientific. The concentrated stock solutions were diluted in water to make standard solutions for calibration. This method, which is a modification of the method described in ChromPack Application Note 1268-GC Rev. 3 (ChromPack; Varian USA) (Robert J. West, Dow Chemical, personal communication), detected the chloroethenes and ethene.
DNA extraction procedure.
Nucleic acids were extracted from the microcosm cultures or groundwater samples by a bead mill homogenization procedure, using a FastDNA SPIN kit for soil (Q·biogene, Carlsbad, Calif.) and FastPrep FP120 (Savant/Bio 101 Holbrook, N.Y.), which was designed to isolate genomic DNA from all cell types. For groundwater, 1 ml of a pellet resuspended in 1× PBS was used. For soil microcosm cultures, a 10-ml sample was centrifuged to obtain a pellet, which was resuspended in 500 μl of the culture medium. The community DNA was isolated from both types of samples by using the silica matrix system of a FastDNA SPIN kit as recommended by the manufacturer. The community DNA in a sample was eluted by adding 80 μl of sterile, deionized water and then mixing the matrix and water together with a pipette tip and centrifuging the preparation for 1 min at 14,000 × g. The flowthrough product was transferred to a 1.5-ml screw-cap tube and stored at −20°C until it was needed for the PCR assay.
Design of Dehalococcoides group primers for the PCR assay.
The PCR primers (Table 2) were designed by using unique sequences of consensus variable and hypervariable regions in the 16S rDNA sequence of D. ethenogenes (E. R. Hendrickson and R. C. Ebersole, 2000, PCT: WO 0063443-A32; GenBank accession number AF004928) and Dehalococcoides group sequences found in the Victoria culture and the Pinellas culture (this study). The Dehalococcoides sequences were aligned with 16S rDNA sequences of 106 organisms by using MegAlign (6, 37) (DNAstar, Madison, Wis.) or Pileup (Wisconsin Package version 10.0; Genetics Computer Group, Madison, Wis.). This set of sequences represented most major domains, families, and genera in the kingdoms Bacteria and Archaea. The sequences that were unique to the Dehalococcoides group were identified by using the alignment report method of MegAlign. Candidate primer sequences were manually selected from the unique sequences. The uniqueness of these sequences was examined further by comparing the candidate sequences to the GenBank ribosomal sequence database sequences by using Blast (2) or FastA (32). The sequences were analyzed by using Ribosomal Database Project II and its on-line Probe Match program (http://www.cme.msu.edu/RDP/cgis/probe_match.cgi?su=SSU, RDPII; Michigan State University, East Lansing) (24). The primer sequences selected for this study are shown in Table 2 (Hendrickson and Ebersole, PCT: WO 0063443-A32). The primers were tested with the PCR assay by using either community DNA, which was isolated from Pinellas and Victoria microcosm cultures, or a plasmid containing a cloned Victoria Dehalococcoides 16S rDNA sequence.
TABLE 2.
Designations, positions, and sequences of primers used in this study
| Primer | bp coordinatesa | Primer sequence | Product size (bp) |
|---|---|---|---|
| DHC primer sets | |||
| DHC primer set 1 | |||
| Fp DHC 1 | 1-17 | 5"GATGAACGCTAGCGGCG3" | 692 |
| Rp DHC 692 | 692-673 | 5"TCAGTGACAACCTAGAAAAC3" | |
| DHC primer set 2 | |||
| Fp DHC 1 | 1-17 | 5"GATGAACGCTAGCGGCG3" | 1,212 |
| Rp DHC 1212 | 1220-1199 | 5"GGATTAGCTCCAGTTCACACTG3" | |
| DHC primer set 3 | |||
| Fp DHC 1 | 1-17 | 5"GATGAACGCTAGCGGCG3" | 1,377 |
| Rp DHC 1377 | 1385-1366 | 5"GGTTGGCACATCGACTTCAA3" | |
| DHC primer set 4 | |||
| Fp DHC 385 | 385-405 | 5"GGGTTGTAAACCTCTTTTCAC3" | 421 |
| Rp DHC 806 | 806-787 | 5"GTTAGCTTCGGCACAGAGAG3" | |
| DHC primer set 5 | |||
| Fp DHC 587 | 587-611 | 5"GGACTAGAGTACAGCAGGAGAAAAC3" | 503 |
| Rp DHC 1090 | 1090-1072 | 5"GGCAGTCTCGCTAGAAAAT3" | |
| DHC primer set 6 | |||
| Fp DHC 774 | 774-791 | 5"GGGAGTATCGACCCTCTC3" | 438 |
| Rp DHC 1212 | 1220-1199 | 5"GGATTAGCTCCAGTTCACACTG3" | |
| DHC primer set 7 | |||
| Fp DHC 946 | 946-963 | 5"AGTGAACCGAAAGGGAAA3" | 266 |
| Rp DHC 1212 | 1220-1199 | 5"GGATTAGCTCCAGTTCACACTG3" | |
| Universal bacterial primers | |||
| Universal primer set 1 | |||
| 517F | 517-532 | 5"GCCAGCAGCCGCGGTA3" | 889 |
| 1406R | 1406-1392 | 5"ACGGGCGGTGTGTAC3" | |
| Universal primer set 2 | |||
| 8F | 8-27 | 5"AGAGTTTGATCCTGGCTCAG3" | 1,502 |
| 1492R | 1510-1492 | 5"GGTTACCTTGTTACGACTT3" |
For the DHC primer sets the coordinates are the D. ethenogenes strain 195 16S rDNA base position coordinates, and for the universal bacterial primers the coordinates are the E. coli 16S rDNA base position coordinates.
PCR procedure used for both the DHC PCR assay and the community diversity analysis.
The 16S RNA gene sequences were amplified by using one of the seven Dehalococcoides 16S rDNA group primer sets for the Dehalococcoides PCR (DHC PCR) assay or one of the universal Bacteria primer sets shown in Table 2. All PCR amplifications were performed with a Perkin-Elmer GeneAmp 9600 thermal cycler and a GeneAmp PCR kit (P-E Applied Biosystems, Norwalk, Conn.). The PCR mixtures contained 1× PCR buffer (10 mM Tris-HCl [pH 8.3], 50 mM KCl), 1.5 mM MgCl2, each deoxynucleoside triphosphate at a concentration of 10 μM, 20 pmol of each primer (a forward primer and a reverse primer [Table 2]), 2.5 U of Taq polymerase, and 1 μl of the community DNA or 1 μl of a 1:10-diluted sample in a final reaction volume of 50 μl. The following PCR thermocycling program was used: 2 min of denaturation at 95°C, followed by either 30 or 40 cycles of 1 min at 94°C, 1 min at 55°C, and 1 min at 72°C and finally cooling at 4°C. Eight microliters of each PCR product was visualized on a 2% agarose gel (SeaKem GTG; FMC BioProducts, Rockland, Maine) that was stained with ethidium bromide (0.5 μg/ml). The HinfI fragments of a molecular weight marker obtained from Gibco BRL (Gaithersburg, Md.), a 1-kb ladder obtained from Gibco BRL, or molecular weight marker VIII (Boehringer GmbH, Mannheim, Germany) was used as the molecular size marker. For the community diversity analysis we used universal Bacteria primer set 1 (Table 2) and the PCR conditions described above. In some assays, when a product was not produced by the initial PCR performed with the DHC PCR primer sets, a nested primer approach was used (23). The initial PCR was performed with universal Bacteria primer set 2 (Table 2) by using the PCR conditions described above. DHC PCR primer sets were then used in a nested PCR; 2-μl portions of the amplified products from the initial PCR were used as the templates, and the PCR conditions described above were used. To measure the product yield in a PCR for a dose-response assay or semiquantification of 16S rDNA PCR product bands, images of the ethidium bromide-stained gels were obtained by using an Eagle Eye II still video system (Stratagene, La Jolla, Calif.). The DNA band intensities in the electronic images were analyzed by using NIH Image 1.62 (written by Wayne Rasband of the National Institutes of Health; available at http://rsb.info.nih.gov/nih-image/download.html).
Analysis of Dehalococcoides group sequences.
When the Dehalococcoides sequences were amplified for community analysis or by the DHC PCR, the products were cloned and sequenced. The community diversity sequences were amplified from community DNA obtained from the dechlorinating cultures by using forward primer 517F and reverse primer 1406R, which resulted in fragments that were approximately 889 bp long. The Dehalococcoides group 16S rDNA sequences were amplified from sample community DNA by using DHC PCR primer sets 2 (Fp DHC 1 and Rp DHC 1212) and 3 (Fp DHC 1 and Rp DHC 1385) (Table 2), which produced 1,212- and 1,377-bp fragments, respectively. The PCR products obtained in each case were cloned by using the PCR dA/T cloning system (Invitrogen, Inc., San Diego, Calif.). Plasmids containing cloned 16S rDNA sequences were isolated by using the QIAvac system (Qiagen, Inc., Valencia, Calif.) according to the manufacturer's recommended procedure. The cloned amplicons were sequenced by using the fluorescent Dye-Deoxy method of Perkin-Elmer and a model 377 DNA sequencer (Applied Biosystems, Perkin-Elmer, Foster City, Calif.). Both strands of each clone insert were completely sequenced. The sequences were assembled and edited by using the Seqman II program (version 4.0; DNAstar, Inc.) (36). The rDNA sequences were analyzed to verify that the amplicon was Dehalococcoides 16S rDNA by using either the Pileup and Neighbor-Joining tree programs in the Wisconsin Package (version 10.0; Genetics Computer Group) or the Clustal alignment and phylogenetic tree functions of the MegAlign program in the DNAstar package. Unique sequences were verified by using Blast (2) or FastA (32). For sequence analysis, default values of the software were used unless otherwise specified.
Phylogenetic analysis of 16S rRNA sequences.
Sequences were aligned by using the ClustalW program through the San Diego Supercomputer Center's Biology Workbench website (http://workbench.sdsc.edu/). Subsequent analyses were carried out by using programs of the Phylip package (14). Aligned bacterial sequences (1,310 bases) were bootstrapped by using the Seqboot program. Phylogenetic trees were generated from 100 data sets by using DNAml (DNA maximum likelihood). A consensus tree was constructed by using the Consense program. Plots were generated by using Drawgram. Sequence RFLP17 (33) was used to root the phylogenetic tree.
Nucleotide sequence accession numbers.
Twenty-one Dehalococcoides rDNA sequences have been deposited in the GenBank database under accession numbers AF388530 to AF388550 (2b).
RESULTS AND DISCUSSION
Development of the DHC PCR assay.
Sequences for designing the PCR primers used in this study were obtained from D. ethenogenes strain 195 and Dehalococcoides sequences isolated from two chloroethene-dechlorinating cultures. Both of these cultures completely dechlorinated PCE to ethene. One of them was derived from the Department of Energy's Pinellas site near Largo, Fla. (12, 17). The other was developed with soil samples taken from an industrial site near Victoria, Tex. (2a, 9, 18, 22, 31, 34, 39) (Table 1 and Fig. 1). Community rDNA was amplified by PCR performed with universal Bacteria primer set 1 (Table 2) and community DNA from both cultures. The amplified rDNA products were cloned to form community libraries and analyzed. Dehalococcoides group sequences were found in the libraries of both cultures. The 16S rDNA sequences from the Pinellas and Victoria cultures were designated DHC-pin and DHC-vic, respectively. Dehalospirillum multivorans and Dehalobacter restrictus, which are known to dechlorinate PCE and TCE to cDCE, were also found in the Pinellas culture (20, 29, 35).
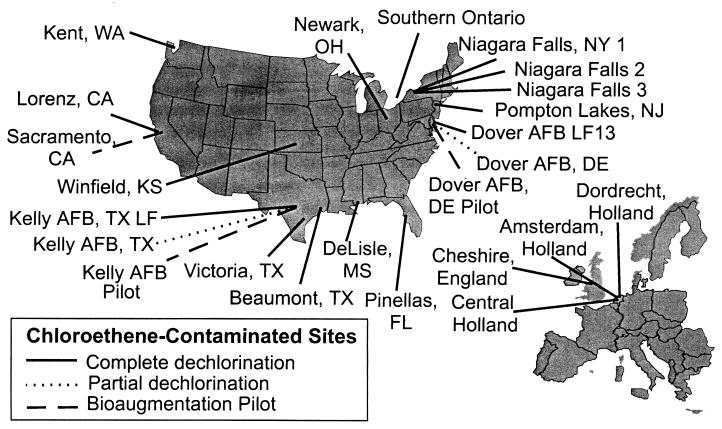
FIG. 1.
Locations of the chloroethene-contaminated sites in North America and Europe that were tested for Dehalococcoides group 16S rDNA sequences.
The DHC-vic, DHC-pin, and D. ethenogenes strain 195 sequences were analyzed to determine unique sequences that could be used to design primers specific for Dehalococcoides 16S rDNA. The unique sequences were determined by aligning the three rDNA sequences with 106 selected GenBank prokaryote 16S rDNA sequences which represented a wide range of phylogenetically distinct members of the Eubacteria and Archaea. Analysis of the aligned sequences showed that Dehalococcoides 16S rDNA had unique sequences in nine variable regions, including eight hypervariable regions. The structure of the Dehalococcoides rDNA sequences was found to be similar to the structure of other prokaryote 16S rDNA sequences (4, 7, 38). Seven DHC PCR primer sets (Table 2) were designed for these unique sequences.
The DHC PCR primer sets and the PCR assay were evaluated for specificity and amplification of Dehalococcoides 16S rDNA sequences. This was accomplished by testing microbial community DNA from three types of mixed microbial communities. These DNAs included DNA from fully dechlorinating cultures, DNA from partially dechlorinating cultures, and DNA from a nondechlorinating culture. In these tests the specificity of the DHC PCR primer sets was tested with DNA from diverse populations of organisms.
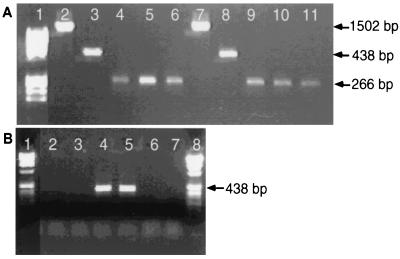
The DHC PCR primer sets were first evaluated with the community DNA from the fully dechlorinating cultures obtained from Pinellas, Fla., and Victoria, Tex. The DHC PCR assays performed with community DNA from the Pinellas and Victoria cultures yielded amplified products of the anticipated size for each DHC PCR primer set used. An example of the PCR results is shown in Fig. 2A; specifically, a 438-bp product (Fp DHC 774 and Rp DHC 1212) and a 266-bp product (Fp DHC 946 and Rp DHC 1212) were obtained.
FIG. 2.
Detection of Dehalococcoides group 16S rDNA sequences in chloroethene-dechlorinating cultures. (A) Direct PCR of fully dechlorinating cultures (Victoria and Pinellas cultures). Lane 1, molecular weight markers (φX174 HinfI fragments); lanes 2 through 6, Victoria community DNA (lane 2, universal primers 8F and 1492R [PCR control]; lane 3, primers Fp DHC 774 and Rp DHC 1212; lanes 4 through 6, primers Fp DHC 946 and Rp DHC 1212); lanes 7 through 11, Pinellas community DNA (lane 7, universal primers 8F and 1492R [PCR control]; lane 8, primers Fp DHC 774 and Rp DHC 1212; lanes 9 through 11, primers Fp DHC 946 and Rp DHC 1212). (B) Direct PCR with a partially dechlorinating microcosm from Dover AFB. Lanes 1 and 8, molecular weight marker VIII (Boehringer); lanes 2 and 3, Dover microcosm community DNA; lanes 4 and 5, Victoria cloned 16S rDNA sequence (PCR positive control); lanes 6 and 7, no-template PCR negative control.
The identities of the DHC PCR products were verified by sequencing. The sequences amplified from both cultures exhibited 98 to 99% identity with the 16S rDNA sequences of D. ethenogenes strain 195 and strains FL2 and CBDB1 (1, 23, 26). The set of sequences from the Pinellas culture exhibited 100% identity with the Pinellas sequence (DHC-pin) and 98% identity with the Victoria sequence (DHC-vic). The Victoria rDNA sequence set exhibited 100% identity with DHC-vic and 98% identity with DHC-pin. DHC PCR-amplified products did not exhibit identity with other bacteria in either community.
Experiments were also conducted to ascertain if Dehalococcoides rDNA sequences could be detected in samples from a site at which PCE or TCE is partially dechlorinated and dechlorination stops at cDCE. The Dover Air Force Base (AFB) aquifer (Dover, Del.) was found to be an aerobic oxidized environment in which TCE is partially dechlorinated to cDCE. Four microcosms were inoculated with soil samples obtained from a Dover AFB chloroethene-contaminated aquifer. Similar to field observations, each of the four Dover AFB microcosms only partially dechlorinated TCE to cDCE.
Community DNA was extracted from the Dover microcosms and used to test the seven DHC PCR primer sets (Table 2) in PCR assays to determine if some members of the genus Dehalococcoides could be associated with partially dechlorinating cultures. An example of the DHC PCR assay response to samples from partially dechlorinating cultures is shown in Fig. 2B. The DHC PCR primer set which included Fp DHC 774 and Rp DHC 1212 did not produce the anticipated 438-bp product. None of the DHC PCR primer sets produced amplified products. Therefore, we concluded that Dehalococcoides rDNA sequences were not present in these partially dechlorinating microcosms.
The DHC PCR primer sets were also tested with DNA from two nondechlorinating cultures, host Escherichia coli DNA used for community analysis (TOP10; Invitrogen) and community DNA from a bioreactor sludge culture that degrades picric acid and 2,4-dinitrophenol. PCR products were not generated with the DNA from these samples (data not shown).
The seven DHC PCR primer sets produced amplified products of the expected sizes and with the expected sequence identities when we used DNA from mixed cultures that exhibited full dechlorinating activity or plasmid DNA containing the cloned DHC-vic sequence. The DHC PCR assay did not produce a positive response with bacterial DNA from other types of communities. Therefore, we concluded that the DHC PCR primer sets are selective for Dehalococcoides sequences.
Sensitivity of the DHC PCR assay.
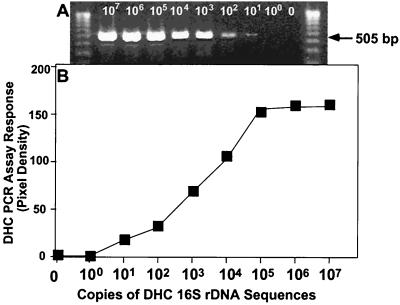
The Dehalococcoides organisms used in this study have not been isolated; they have been detected and identified on the basis of 16S rDNA sequences obtained from mixed cultures. To evaluate the DHC PCR assay's sensitivity, known concentrations of the plasmid containing the cloned DHC-vic rDNA sequence were used to determine the detection limit. The plasmid was serially diluted and then tested with the DHC PCR assay by using one of the DHC PCR primer sets, Fp DHC 587 and Rp DHC 1090. An example of the results obtained is shown in Fig. 3, which shows that there was a DHC PCR dose-response relationship. The detection limit when this primer set was used was 10 to 100 copies of the gene per reaction mixture. The assay sensitivities when the other six DHC PCR primer sets were used ranged from 10 to 1,000 copies per reaction mixture.
FIG. 3.
DHC PCR sensitivity assay. (A) PCR dose-response assay of serially diluted Victoria Dehalococcoides 16S rDNA plasmid DNA, performed with Dehalococcoides group primers Fp DHC 586 and Rp DHC 1090. The two outside gel lanes are molecular weight markers (Gibco BRL 1-kb ladder); the eight left interior lanes, PCR performed on serial diluted plasmid copies (107 to 100) per 50-μl reaction. The rightmost interior lane is a PCR control with no plasmid template (0). (B) PCR dose-response assay in ethidium bromide-stained gels: graph of the pixel densities in gel panel A for the PCR products, as determined by the NIH Image 1.62 program, and the numbers of copies of cloned Victoria Dehalococcoides 16S rDNA sequences in 50-μl PCR mixtures.
Detection of Dehalococcoides in soil.
Soil samples were collected from eight chloroethene-contaminated sites. The locations used for testing were chosen because they were geographically distributed locations that included a wide range of climatic zones (from subtropical to temperate zones) and types of dechlorination. Each site was found to be an anaerobic, reduced environment where PCE or TCE was dechlorinated to cDCE, VC, and ethene (Table 1). Site stewards verified that there was dechlorination activity, as determined by chemical analysis. The sites included a PCE-contaminated industrial site in DeLisle, Miss.; a site with chloroethene-contaminated sludge from an industrial area in Beaumont, Tex.; and three PCE- and TCE-contaminated industrial sites located in Pompton Lakes, N.J., Winfield, Kans., and Newark, Ohio. Other locations included a TCE-contaminated aquifer at Dover AFB in Dover, Del., and two PCE- and TCE-contaminated landfills located in Dover, Del. (Dover AFB), and San Antonio, Tex. (Kelly AFB). At Dover AFB, soil samples were obtained from the bioaugmentation pilot area, where the Pinellas culture had been injected into the subsurface for bioaugmentation of TCE degradation to ethene (12; E. R. Hendrickson, M. A. Elberson, J. A. Tabinowski, and D. E. Ellis, Second Int. Conf. Remed. Chlorinated Recalcitrant Compounds, abstr. E10, 2000). An additional dechlorinating culture, KB1, was also included in the study; this culture was developed from groundwater and soil obtained from an industrial site in southern Ontario (11; E. Edwards, E. Cox, S. Dworatzek, M. McMaster, and D. Major, Goldschmidt 2000, Int. Conf. Geochem. J. Conf. Abstr. 5:367, 2000).
All the soil samples collected from the eight sites were used to inoculate microcosms in order to verify their abilities to dechlorinate PCE or TCE to ethene. The KB1 culture and the eight enriched microcosms dechlorinated PCE or TCE to ethene and displayed the characteristic sequential production of TCE, cDCE, VC, and ethene.
Each of the nine fully dechlorinating microcosms was tested for the presence of Dehalococcoides 16S rDNA sequences. Community DNA was extracted from the microcosm cultures and tested for the presence of Dehalococcoides 16S rDNA by using the seven DHC PCR primer sets. Samples from all nine cultures yielded expected PCR products. The sequences from the nine cultures exhibited 97.7 to 100% identity with Dehalococcoides sequences in the databases and with DHC-pin and DHC-vic. The sequence from the Dover bioaugmentation pilot area exhibited 100% identity with DHC-pin. This was not surprising since the Pinellas culture was used to bioaugment the aquifer (12). The results of the DHC PCR assay for the nine sites are summarized in Table 1.
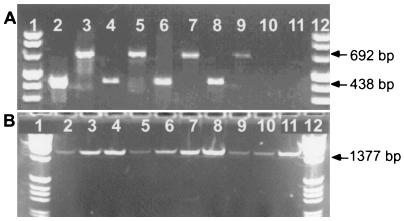
An example of the PCR products generated from the fully dechlorinating microcosms is shown in Fig. 4A. The gel data illustrate that DHC PCR products were formed with DNA from each of four fully dechlorinating microcosms started with soil samples obtained from DeLisle, Miss., Pompton Lakes, N.J., Victoria, Tex., and the Dover AFB bioaugmented pilot area. Specifically, the PCR assays performed with the DHC PCR primers produced amplified products of the anticipated sizes, including a 438-bp product (Fp DHC 774 and Rp DHC 1212) and a 692-bp product (Fp DHC 1 and Rp DHC 692). The Victoria culture was used as a PCR assay control in this experiment; in previous tests it had been shown to produce amplified products that were identified as Dehalococcoides 16S rDNA sequences.
FIG. 4.
Detection of Dehalococcoides 16S rDNA sequences in samples from PCE- or TCE-contaminated sites at which chloroethenes are fully dechlorinated to ethene. (A) Test performed with community DNA from fully dechlorinating microcosms started with soil from contaminated sites inoculated with soil from chloroethene-contaminated sites. The assay was performed with DHC PCR primers Fp DHC 774 and Rp DHC 1212 (lanes 2, 4, 6, 8, and 10) and with DHC PCR primers Fp DHC 1 and Rp DHC 692 (lanes 3, 5, 7, 9, and 11). Lanes 1 and 12, molecular weight marker VIII (Boehringer); lanes 2 and 3, DeLisle microcosm; lanes 4 and 5, Dover AFB bioaugmentation pilot microcosm; lanes 6 and 7, Pompton Lakes microcosm; lanes 8 and 9, Victoria culture (PCR positive control); lanes 10 and 11, no template (PCR negative control). (B) PCR assay results for groundwater samples collected from several sites in the United States and Europe. The assay was performed with DHC PCR primers Fp DHC 1 and Rp DHC 1385. Lanes 1 and 12, molecular weight markers (1-kb ladder; Gibco BRL); lane 2, sample A from Dordrecht, The Netherlands, industrial site; lane 3, sample B from Dordrecht, The Netherlands, industrial site; lane 4, Kelly AFB clay landfill; lane 5, sample A from Lorenz, Calif., industrial site; lane 6, sample B from Lorenz, Calif., industrial site; lane 7, Niagara Falls, N.Y., industrial landfill site 2; lane 8, Niagara Falls, N.Y., industrial site 1; lane 9, Amsterdam, The Netherlands, industrial site; lane 10, central Netherlands industrial site; lane 11, Niagara Falls, N.Y., site 3 (a defense plant).
The DHC PCR assay demonstrated that Dehalococcoides 16S rDNA sequences were present in microcosm cultures that fully dechlorinated chloroethenes to ethene. The data are consistent with previously reported dechlorination findings (23, 25, 26), and together, the findings demonstrate that the Dehalococcoides sequences are associated with full dechlorination of chloroethenes.
Correlation of DHC PCR assay response with chloroethene dechlorination.
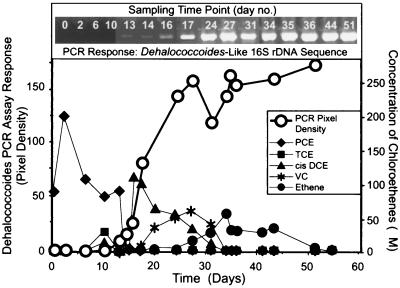
To determine if Dehalococcoides appears to be functionally associated with specific steps in the dechlorination processes, PCR product formation and the disappearance of dechlorination products were monitored over time. In this experiment, some of the nine dechlorinating microcosms were simultaneously monitored for PCE or TCE dechlorination products and for the appearance of Dehalococcoides 16S rDNA sequences. Duplicate samples were taken from each microcosm; one sample was used for the DHC PCR assay, and the other was used to measure the formation of dechlorination products by gas chromatography and to evaluate the extent of PCE dechlorination. Figure 5 shows the results obtained with a microcosm inoculated with soil from an industrial site in DeLisle, Miss. In this microcosm, the concentrations of DHC PCR products increased as PCE and TCE disappeared and the concentrations of dechlorination intermediates (cDCE and VC) increased. The PCR product formation curve resembled a growth curve. Each Dehalococcoides cell should have approximately the same number of rRNA gene sequences in its chromosomal DNA. Thus, the increase in the DHC PCR assay response indicated that the number of Dehalococcoides rDNA sequences had increased. These data imply that there was cell growth that occurred parallel to the metabolic dechlorination activity, resulting in the formation of Dehalococcoides rDNA, which appeared to be associated with assimilation and dechlorination of both cDCE and VC and production of ethene. The data are consistent with the hypothesis that Dehalococcoides participated in the later stages of dechlorination in the DeLisle microcosm. Alternatively, since the PCR response plateaued when cDCE disappeared, it is difficult to conclude that Dehalococcoides plays a direct role in degradation of VC. Maymó-Gatell et al. (25, 26) obtained data that implied that D. ethenogenes strain 195 uses VC as a cometabolite. Thus, Dehalococcoides may assist other organisms in completing dechlorination of VC to ethene. In the control microcosm that was not amended with PCE there was not an increase in the level of Dehalococcoides rDNA sequences over time, which supports the hypothesis that Dehalococcoides is involved in chloroethene-dechlorinating processes (data not shown).
FIG. 5.
Monitoring the relationship of dechlorinating activity (PCE dechlorination product formation) to ethene and the change in the concentration of Dehalococcoides 16S rDNA in the DeLisle microcosm. The pixel densities of the PCR products as determined by the NIH Image 1.62 program were plotted against time.
The results of the experiments described above, in conjunction with the DHC PCR assay response of the previously described soil cultures obtained from chloroethene-dechlorinating sites, support the conclusion that Dehalococcoides 16S rDNA sequences are associated with full dechlorination of chloroethenes.
Testing for Dehalococcoides in environmental groundwater.
Soil sample collection and microcosm culture studies are expensive and time-consuming. Using groundwater samples as test samples would be easier, less expensive, and more practical if they proved to be useful sources of Dehalococcoides rDNA sequences that could be correlated with dechlorination activity.
To determine if groundwater can be used for testing for Dehalococcoides, 10 groundwater samples were collected from very diverse chloroethene-contaminated sites. These samples represented different geographical locations and a variety of climatic zones. The groundwater samples came from anaerobic and reductive aquifers in North America, England, and The Netherlands. The sites also comprised a variety of geological soils and earth matrices, including sand, clay, and fractured bedrock (Table 1 and Fig. 1). The sites at which full dechlorination of chloroethenes occurred were shown to contain the dechlorination products VC and ethene. Again, stewards verified by chemical analysis that dechlorination occurred at the sites.
Community DNA extracted from groundwater samples was tested for Dehalococcoides 16S rDNA sequences. Each sample was tested with the seven DHC PCR primer sets listed in Table 2. Groundwater samples from all 10 sites yielded the anticipated amplification products of the correct sizes in the DHC PCR assay (Table 1). Following PCR, the identities of the amplified products were verified by sequencing. The rDNA sequences found at the 10 groundwater sites exhibited between 97.7 and 99.7% identity with the 16S rDNA sequences of strains 195, DCEH2, FL2, and CBDB1 and with the microcosm Dehalococcoides 16S rDNA sequences.
Figure 4B shows the results of a DHC PCR assay performed with groundwater samples. The PCR assays were performed with community DNA extracted from 10 groundwater samples taken from eight sites. The PCR assay performed with DHC PCR primers Fp DHC 1 and Rp DHC 1385 produced the anticipated 1,377-bp product for each sample.
Groundwater samples from partially dechlorinating sites.
Groundwater samples were taken from two sites that had aquifer characteristics similar to those of the Dover AFB aquifer (i.e., an aerobic, oxidized environment) (Table 1 and Fig. 1) and were shown by chemical analysis to partially dechlorinate PCE to cDCE. Site stewards provided evidence that partial dechlorination occurred. Microcosm studies of the sites were conducted to verify that partial dechlorination occurred under the anaerobic, reduced conditions needed for full dechlorination.
The first set of groundwater samples was obtained from Kelly AFB in San Antonio, Tex. Microcosms prepared with samples from this site were amended with methanol, lactate, ethanol, or acetate, and they did not completely transform PCE or TCE to ethene and only partially dechlorinated chloroethenes to cDCE (Elizabeth Edwards, personal communication). The second set of groundwater samples was obtained from an industrial site in Sacramento, Calif. Microcosms prepared with samples from this site were also amended with methanol, lactate, ethanol, or acetate, and they did not completely transform PCE; the dechlorination process stopped at cDCE (Michaye McMaster, Geosyntec, Guelph, Canada, personal communication). Both sets of results confirmed that chloroethenes are partially dechlorinated at these sites.
DHC PCR analysis of the community DNA extracted from both types of groundwater samples did not yield PCR products (data not shown). To increase the sensitivity of the PCR assay, the samples were assayed by using the nested primer approach, in which a universal bacterial 16S rDNA primer set (primers 8F and 1492R) was used in an initial amplification reaction (23). The amplified product from the first PCR was amplified with a nested DHC PCR primer set (primers Fp DHC 385 and Rp DHC 806) to detect the presence of Dehalococcoides sequences. The nested approach did not yield PCR products (data not shown). Based on the results of the groundwater PCR analysis, site, and microcosm data, we assumed that Dehalococcoides was not present either in the subsurface of the Kelly AFB aquifer or at the Sacramento industrial site.
The DHC PCR assay results for the groundwater samples were similar to the results of the dechlorinating microcosm tests. Dehalococcoides rDNA sequences were detected only in samples that were linked to full dechlorination of chloroethenes. Linking the detection of Dehalococcoides 16S rDNA sequences to full dechlorination of chloroethenes provides a key indicator for the presence of favorable dechlorination conditions that support growth and maintenance of dehalorespiring and dechlorinating bacteria.
Analysis of Dehalococcoides 16S rDNA sequences.
Sequence analysis of DHC PCR assay amplicons from the dechlorinating microcosms confirmed that the assay products were Dehalococcoides 16S rDNA sequences. A total of 21 different complete partial sequences were obtained from 17 of the sites that tested positive for Dehalococcoides sequences (Table 1). The sequences were generated from a 1,377-bp PCR fragment produced by using the DHC PCR primer set comprising primers Fp DHC 1 and Rp DHC 1385. Analysis of the sequences revealed identity to Dehalococcoides sequences in the sequence databases. Comparisons of the sequences showed that the levels of identity with D. ethenogenes strain 195, DCEH2, FL2, and CBDB1 sequences ranged from 97.7 to 99.9%. The differences between the rDNA base sequences ranged from 1 to 25 bases (1, 23, 26). The analysis also confirmed that the Dehalococcoides rDNA sequences in this study and those in the databases form a unique 16S rDNA phylogenetic group. Collectively, they are referred to as Dehalococcoides sequences. Phylogenetically, Dehalococcoides clusters with the green nonsulfur bacteria and is loosely affiliated with sequences from noncultured bacteria obtained from environmental samples (1, 5, 16, 26, 33).
Representative 16S rDNA sequences from this study were used to determine if there were specific base differences in the Dehalococcoides sequences that were signature differences and could be used to divide the Dehalococcoides group. The representative rDNA sequences from Pompton Lakes (DHC-plk), Victoria (DHC-vic), DeLisle (DHC-dll), and Pinellas (DHC-pin) were included in this comparison. Computer analysis of these four rDNA sequences aligned with the strain 195, CBDB1, FL2, and DCEH2 sequences revealed unique base substitutions at specific base positions in variable regions 2 and 6, as shown in Fig. 6.
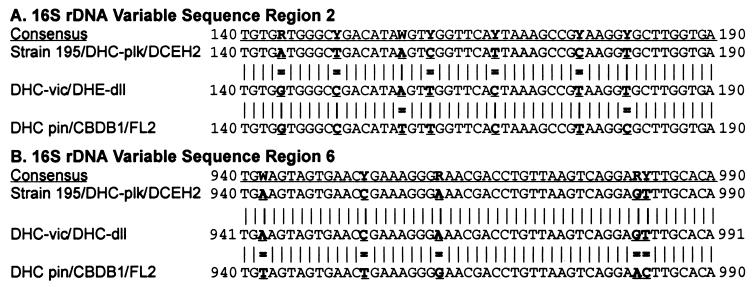
FIG. 6.
Sequences of Dehalococcoides 16S rDNA variable regions 2 and 6, showing signature base differences that define the Cornell, Victoria, and Pinellas phylogenetic branches. The strain 195 (Cornell branch) sequence was the sequence to which the other Dehalococcoides 16S rDNA sequences were compared. An equal sign indicates a mismatch between a 16S rDNA sequence and the Cornell sequence at a signature base position. R = A or G; Y = C or T; and W = A or T.
The first set of base substitutions was in variable region 2 of the Dehalococcoides rDNA sequence, between Dehalococcoides bp 144 and 181 (Fig. 6A). The bases that were different from the bases in the strain 195 sequence were at bp 144, 150, 160, 167, and 176 in the DHC-vic, DHC-dll, DHC-pin, FL2, and CBDB1 sequences (Fig. 6A). Two additional base substitutions occurred in the DHC-pin, CBDB1, and FL2 sequences at Dehalococcoides bp 157 and 181 (Fig. 6A). The second set of bases that were different from the bases in the strain 195 sequence was in variable region 6 between Dehalococcoides bp 944 and 985. The unique base changes were found in the DHC-pin, CBDB1, and FL2 sequences at bp 944, 955, 963, 984, and 985 (Fig. 6B). The base changes in variable region 6 were not found in the other four sequences examined, the DCEH2, DHC-plk, DHC-vic, and DHC-dll sequences.
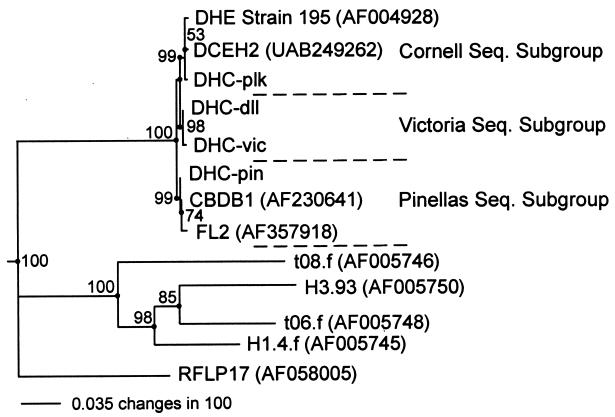
The consistency of the specific base substitutions among the sequences analyzed suggested that the base positions might be signature positions. To test this premise, the Dehalococcoides sequences were phylogenetically analyzed, which resulted in the 16S rDNA phylogenetic tree shown in Fig. 7. Analysis of the tree data indicated that the specific base substitutions contributed to the formation of three phylogenetic subbranches in the Dehalococcoides branch of the phylogenetic tree. The results of a bootstrap analysis of the data during formation of the phylogenic tree supported the robustness of the tree branches. The bootstrap values for the subgroup branch points ranged from 99 to 100. The sequences of rDNA with base substitutions in regions 2 and 6 that were identical to the D. ethenogenes strain 195 sequence formed a phylogenetic subbranch designated the Cornell group. In addition, the rDNA sequences that had the same base substitutions and were affiliated on the same phylogenetic subbranch with the DHC-vic sequence and the DHC-pin sequence were designated members of the Victoria group and the Pinellas group, respectively. Each sequence subgroup was named for the site of origin of the rDNA that was the first rDNA sequenced for the group. The Cornell group's name was derived from the place where strain 195 was first isolated and described (26).
FIG. 7.
16S rDNA phylogenetic tree showing phylogenetic relationships based on the base sequence differences in regions 2 and 6 for the Dehalococcoides group, as well as the compositions of three sequence subgroups (Cornell, Victoria, and Pinellas sequence subgroups). The positions of the loosely affiliated 16S rDNA from uncultured environmental samples are also shown. The RFLP17 sequence (33) was used to root the phylogenetic tree. Bootstrap values are indicated at the branch points. DHE, D. ethenogenes.
The 16S rDNA of this study which was completely sequenced and identified belonged to one of the three Dehalococcoides phylogenetic subgroups listed in Table 1. Generally, samples from individual sites had one unique Dehalococcoides 16S rDNA sequence that could be classified into one of the three sequence subgroups. However, samples from three sites contained rDNA sequences belonging to more than one phylogenetic subgroup. The microcosm culture from an industrial site in southern Ontario, KB1, contained one sequence belonging to the Pinellas group and another sequence belonging to the Cornell group. The groundwater sample from industrial site 1 in Niagara Falls, N.Y., also contained two sequences, one belonging to the Pinellas group and the other belonging to the Cornell group. The microcosm culture from an industrial site in Beaumont, Tex., contained three unique Dehalococcoides rDNA sequences. One sequence was classified in the Cornell group, another sequence was classified in the Pinellas group, and the third sequence was classified in the Victoria group.
The significance of the fact that we found three Dehalococcoides phylogenetic subgroups at one site and not at another site is not known. Each subgroup or even its members may play a different transformation role in dechlorination of a variety of chlorinated solvents. The genus Dehalococcoides is a group of organisms that exhibit 16S rDNA phylogenetic relationships and have the ability to perform dehalorespiration. However, the targeted chlorinated electron acceptor may be different for some members of the group. For example, CBDB1, a member of the Pinellas group, does not dechlorinate PCE or TCE. Unlike the other Dehalococcoides strains in this study and the databases, it uses a different dehalorespiration process; it prefers to use TeCB and TCB as electron acceptors. In light of these findings, should the name D. ethenogenes be reserved for organisms that dechlorinate PCE and TCE to ethene? It is speculative to predict what environmental or dechlorination conditions contribute to selection of a Dehalococcoides subgroup or, in some cases, support a synergistic relationship between members of different subgroups. Further investigation is required to establish the role and significance of the different members of the genus Dehalococcoides in dechlorination in the field. It also remains to be seen if there are other organisms in the environment that can catalyze the conversion of PCE and TCE to ethene, specifically transforming cDCE or VC to ethene.
The DHC PCR assay is easy to implement, and when it is used in combination with microcosm and field data, it provides a new tool for evaluating the potential for dechlorination activity and for tracking changes in dechlorinating organisms in populations. The sequence data corroborate the finding that the Dehalococcoides 16S rDNAs in this study and in the databases are detected by the primers of the DHC PCR assay. Furthermore, we speculate that a positive PCR response indicates that there are or at one time were favorable subsurface conditions that supported the growth and dechlorinating activity of Dehalococcoides group organisms. The DHC PCR assay may be used to characterize chloroethene-contaminated sites for spatial distribution of Dehalococcoides sequences in order to correlate the location of the sequences with dechlorination activity and products. Other potential applications of the PCR assay may be to assess contaminated sites as candidates for engineered bioremediation and to monitor natural attenuation or bioaugmentation projects (19; Hendrickson et al., Second Int. Conf. Remed. Chlorinated Recalcitrant Compounds).
In summary, the findings of this study are consistent with the hypothesis that Dehalococcoides sequences (organisms) are widely distributed in the environment and are associated with fully dechlorinating processes. Members of the genus Dehalococcoides were shown to survive in a wide range of geographical locations, in a wide range of geological matrices, and in a variety of climatic zones. Furthermore, a functional relationship was observed between an increase in the amount of Dehalococcoides 16S rDNA and the formation of the dechlorination products cDCE and VC, and ethene dechlorination was demonstrated in dechlorinating microcosms. Thus, members of the genus Dehalococcoides may be important contributors to natural dechlorination processes that occur in the environment. Nevertheless, members of the Dehalococcoides subgroups have been shown to be associated with full chloroethene dechlorination activity and appear to be associated with specific sites. Furthermore, as the role and significance of the different Dehalococcoides organisms are discovered, information gained from PCR assays and sequence analysis may provide an opportunity to optimize dechlorinating community structure in order to deal with different environmental conditions and different chloroethene compositions at contaminated sites.
Acknowledgments
We thank DuPont's Corporate Remediation Group (CRG), Philip A. Palmer, and Hugh J. Campbell for their support of this study. Special thanks are extended to CRG members John E. Vidumsky, Edward J. Lutz, Ron J. Buchanan, and Alfred A. Biehle and also to J. Martin Odom of Central Research and Development, who provided samples, suggestions, and guidance for this project. Also, we thank members of the Remediation Technologies Development Forum who also provided samples, including Martin J. Bell, John W. Davis, Mark Harkness, David Major, Michaye McMaster, and Phil Morgan. We also extend our appreciation to Elizabeth A. Edwards, Sandra Dworatzek (University of Toronto), Perry McCarty, Alfred Spormann (Stanford University), Michael Lee (Terra Systems, Inc., Wilmington, Del.), and Leo Lehmicke (Exponent Co., Bellevue, Wash.) for providing samples and suggestions. We also thank Ania Stelfox (University of Toronto) for her help with the bootstrap analysis of the Dehalococcoides sequences. E. Erin Mack, Michelle M. Thomson, and Lois W. Buonamici are gratefully acknowledged for critically reading the manuscript. The DNA sequences of DHC PCR primers are the subject matter of US Patent Application USSN 09/548,998 (Hendrickson and Ebersole, U.S. Patent Office, 1999; PCT: WO 0063443-A3L, 2000).
REFERENCES
- 1.Adrian, L., U. Szewzyk, J. Wecke, and H. Görisch. 2000. Bacterial dehalorespiration with chlorinated benzenes. Nature 408:580-583. [DOI] [PubMed] [Google Scholar]
- 2.Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. (Suppl.) 215:403-410. [DOI] [PubMed] [Google Scholar]
- 2a.Beeman, R. E., J. E. Howell, S. H. Shoemaker, E. A. Salazar, and J. R. Buttram. 1994. A field evaluation on in situ microbial reductive dehalogenation by biotransformation of chlorinated ethylenes, p. 14-27. In R. E. Hinchee, A. Leeson, L. Semprini, and S. K. Ong (ed.), Bioremediation of chlorinated and polycyclic aromatic hydrocarbon compounds. Lewis Publishers, Boca Raton, Fla.
- 2b.Benson, D. A., I. Karsch-Mizrachi, D. J. Lipman, J. Ostell, B. A. Rapp, and D. L. Wheeler. 2000. GenBank. Nucleic Acids Res. 28:15-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2c.Biswas, N. 1992. Model for predicting PCE desorption from contaminated soils. Water Environ. Res. 64:170. [Google Scholar]
- 3.Bolesch, D., R. Nielsen, and J. Keasling. 1997. Complete reductive dechlorination of trichloroethene by a groundwater microbial consortium. Ann. N. Y. Acad. Sci. 829:97-102. [DOI] [PubMed] [Google Scholar]
- 4.Böttenger, E. C. 1996. Approaches for the identification of microorganisms. ASM News 62:247-250. [Google Scholar]
- 5.Chandler, D. P., F. J. Brockman, T. J. Bailey, and J. K. Fredrickson. 1998. Phylogenetic diversity of Archaea and Bacteria in a deep subsurface paleosol. Microb. Ecol. 36:37-50. [DOI] [PubMed] [Google Scholar]
- 6.Clewley, J. P., and C. Arnold. 1997. MEGALIGN. The multiple alignment module of LASERGENE. Methods Mol. Biol. 70:119-129. [PubMed] [Google Scholar]
- 7.Dams, E., L. Hendriks, Y. Van de Peer, J. M. Neefs, G. Smits, I. Vandenbempt, and R. De Wachter. 1988. Compilation of small subunit RNA sequences. Nucleic Acids Res. 16(Suppl.):r87-r173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.de Bruin, W. P., M. J. J. Kotterman, M. A. Posthumus, G. Schraa, and A. J. B. Zehnder. 1992. Complete biological reductive transformation of tetrachloroethylene to ethane. Appl. Environ. Microbiol. 58:1996-2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.DeWeerd, K. A., W. P. Flanagan, M. J. Brennan, J. M. Principe, and J. L. Spivack. 1998. Biodegradation of trichloroethylene and dichloroethylene in contaminated soil and groundwater. Bioremed. J. 2:29-42. [Google Scholar]
- 10.DiStefano, T. D., J. M. Gossett, and S. H. Zinder. 1991. Reductive dechlorination of high concentrations of tetrachloroethene to ethene by an anaerobic enrichment culture in the absence of methanogenesis. Appl. Environ. Microbiol. 57:2287-2292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Edwards, E. A., and E. E. Cox. 1997. Field and laboratory studies of sequential anaerobic-aerobic chlorinated solvent biodegradation, p. 261-265. In B. C. Alleman and A. Leeson (ed.), In situ and on-site bioremediation, vol. 3. Fourth International Symposium on In Situ and On-Site Biodegradation. Battelle Press, Columbus, Ohio. [Google Scholar]
- 12.Ellis, D. E., E. J. Lutz, J. M. Odom, R. J. Buchanan, Jr., M. D. Lee, C. L. Bartlett, M. R. Harkness, and K. A. DeWeerd. 2000. Bioaugmentation for accelerated in situ anaerobic bioremediation. Environ. Sci. Technol. 34:2254-2260. [Google Scholar]
- 13.Ensley, B. D. 1991. Biochemical diversity of trichloroethylene metabolism. Annu. Rev. Microbiol. 45:283-299. [DOI] [PubMed] [Google Scholar]
- 14.Felsenstein, J. 1989. PHYLIP--phylogeny inference package (version 3.5). Cladistics 5:164-166. [Google Scholar]
- 15.Freedman, D. L., and J. M. Gossett. 1989. Biological reductive dechlorination of tetrachloroethylene and trichloroethylene to ethylene under methanogenic conditions. Appl. Environ. Microbiol. 55:2144-2151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Godon, J.-J., E. Zumstein, P. Dabert, F. Habouzit, and R. Moletta. 1997. Molecular microbial diversity of an anaerobic digestor as determined by small-subunit rDNA sequence analysis. Appl. Environ. Microbiol. 63:2802-2813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Harkness, M. R., A. A. Bracco, M. J. Brennan, Jr., K. A. DeWeerd, and J. L. Spivack. 1999. Use of bioaugmentation to stimulate complete reductive dechlorination of trichloroethene in Dover soil columns. Environ. Sci. Technol. 33:1100-1109. [Google Scholar]
- 18.Haston, Z. C., and P. L. McCarty. 1999. Chlorinated ethene half-velocity coefficients (Ks) for reductive dehalogenation. Environ. Sci. Technol. 33:223-226. [Google Scholar]
- 19.Hendrickson, E. R., M. G. Starr, M. A. Elberson, J. A. Tabinowski, E. E. Mack, M. L. McMaster, and D. E. Ellis. 2001. Using a molecular approach to monitor a bioaugmentation pilot, p. 43-52. In A. Leeson, B. C. Alleman, P Alverez, and V. S. Magar (ed.), Sixth International Symposium on In Situ and On-Site Bioremediation. Battelle Press, Columbus, Ohio.
- 20.Holliger, C., D. Hahn, H. Harmsen, W. Ludwig, W. Schumacher, B. Tindall, F. Vazquez, N. Weiss, and A. J. B. Zehnder. 1998. Dehalobacter restrictus gen. nov. and sp. nov., a strictly anaerobic bacterium that reductively dechlorinates tetra- and trichloroethene in anaerobic respiration. Arch. Microbiol. 169:313-321. [DOI] [PubMed] [Google Scholar]
- 21.Hopkins, G. D., L. Semprini, and P. L. McCarty. 1993. Microcosms and in situ field studies of enhanced biotransformation of trichloroethylene by phenol-utilizing organisms. Appl. Environ. Microbiol. 59:2277-2285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lee, M. D., J. M. Odom, and R. J. Buchanan, Jr. 1998. New perspectives on microbial dehalogenation of chlorinated solvents: insights from the field. Annu. Rev. Microbiol. 52:423-452. [DOI] [PubMed] [Google Scholar]
- 23.Löffler, F. E., Q. Sun, J. Li, and J. M. Tiedje. 2000. 16S rRNA gene-based detection of tetrachloroethene-dechlorinating Desulfuromonas and Dehalococcoides species. Appl. Environ. Microbiol. 66:1369-1374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Maidak, B. L., J. R. Cole, T. G. Lilburn, C. T. Parker, Jr., P. R. Saxman, J. M. Stredwick, G. M. Garrity, B. Li, G. J. Olsen, S. Pramanik, T. M. Schmidt, and J. M. Tiedje. 2000. The RDP (Ribosomal Database Project) continues. Nucleic Acids Res. 28:173-174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Maymó-Gatell, X., T. Anguish, J. M. Gossett, and S. H. Zinder. 1999. Reductive dechlorination of chlorinated ethenes and 1,2-dichloroethane by “Dehalococcoides ethenogenes” 195. Appl. Environ. Microbiol. 65:3108-3113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Maymó-Gatell, X., Y. T. Chien, J. M. Gossett, and S. H. Zinder. 1997. Isolation of a bacterium that reductively dechlorinates tetrachloroethene to ethene. Science 276:1568-1571. [DOI] [PubMed] [Google Scholar]
- 27.Maymó-Gatell, X., V. Tandoi, J. M. Gossett, and S. H. Zinder. 1995. Characterization of an H2-utilizing enrichment culture that reductively dechlorinates tetrachloroethene to vinyl chloride and ethene in the absence of methanogenesis and acetogenesis. Appl. Environ. Microbiol. 61:3928-3933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.McCarty, P. L. 1997. Breathing with chlorinated solvents. Science 276:1521-1522. [DOI] [PubMed] [Google Scholar]
- 29.Neumann, A., H. Scholz-Muramatsu, and G. Diekert. 1994. Tetrachloroethene metabolism of Dehalospirillum multivorans. Arch. Microbiol. 162:295-301. [DOI] [PubMed] [Google Scholar]
- 30.Nielsen, R., and J. Keasling. 1999. Reductive dechlorination of chlorinated ethene DNAPLs by a culture enriched from contaminated groundwater. Biotechnol. Bioeng. 62:160-165. [DOI] [PubMed] [Google Scholar]
- 31.Odom, J. M., J. A. Tabinowski, M. D. Lee, and B. Z. Fathepure. 1995. Anaerobic biodegradation of chlorinated solvents: comparative laboratory study of aquifer microcosms, p. 17-24. In R. E. Hinchee, A. Leeson, and L. Semprini (ed.), Bioremediation of chlorinated solvents, vol. 1. Battelle Press, Columbus, Ohio. [Google Scholar]
- 32.Pearson, W. R., and D. J. Lipman. 1988. Improved tools for biological sequence comparison. Proc. Natl. Acad. Sci. USA 85:2444-2448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Pulliam Holoman, T. R., M. A. Elberson, L. A. Cutter, H. D. May, and K. R. Sowers. 1998. Characterization of a defined 2,3,5,6-tetrachlorobiphenyl-ortho-dechlorinating microbial community by comparative sequence analysis of genes coding for 16S rRNA. Appl. Environ. Microbiol. 64:3359-3367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Rosner, B. M., P. L. McCarty, and A. M. Spormann. 1997. In vitro studies on reductive vinyl chloride dehalogenation by an anaerobic mixed culture. Appl. Environ. Microbiol. 63:4139-4144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Scholz-Muramatsu, H., A. Neumann, M. Messmer, E. Moore, and G. Diekert. 1995. Isolation and characterization of Dehalospirillum multivorans gen. nov., sp. nov., a tetrachloroethene-utilizing, strictly anaerobic bacterium. Arch. Microbiol. 163:48-56. [Google Scholar]
- 36.Swindell, S. R., and T. N. Plasterer. 1997. SeqMan. Contig assembly. Methods Mol. Biol. 70:75-89. [PubMed] [Google Scholar]
- 37.Thompson, J. D., D. G. Higgins, and T. J. Gibson. 1994. Clustal W--improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22:4673-4680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Woese, C. R. 1987. Bacterial evolution. Microbiol. Rev. 51:221-271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yang, Y., and P. L. McCarty. 1998. Competition for hydrogen within a chlorinated solvent dehalogenating anaerobic culture. Environ. Sci. Technol. 32:3591-3592. [Google Scholar]