Abstract
The in vitro activity of BMS-284756 against 602 Staphylococcus aureus isolates, including 152 that were both methicillin and ciprofloxacin resistant (MIC ≥4 μg/ml), was determined. For ciprofloxacin-susceptible and nonsusceptible isolates, the MICs at which 50% of organisms were inhibited were 0.015 and 2 μg/ml and the MICs at which 90% of organisms were inhibited were 0.03 and 4 μg/ml, respectively.
Over the last 2 decades, the increasing incidence of methicillin-resistant Staphylococcus aureus (MRSA) has caused significant clinical concern worldwide. MRSA is also associated with resistance to several commonly used antimicrobial agents, including the macrolides, lincosamides, quinolones, trimethoprim-sulfamethoxazole, and aminoglycosides. There is therefore a need to evaluate the efficacy of new antimicrobial agents that may be useful for the treatment of multiresistant staphylococcal infections. In this study, we determined the activity of BMS-284756 against clinical isolates of S. aureus, including methicillin- and ciprofloxacin-resistant strains.
Fluoroquinolones exert their bactericidal activity by interacting with two type II topoisomerases, DNA gyrase (subunits encoded by gyrA and gyrB) and topoisomerase IV (subunits encoded by grlA and grlB for S. aureus and termed parC and parE for other organisms) (6). Resistance to quinolones in S. aureus arises primarily from mutations in the quinolone resistance-determining region (QRDR) of topoisomerase IV and/or DNA gyrase (2, 5). However, the multidrug efflux pump NorA has also been shown to contribute to quinolone resistance (7).
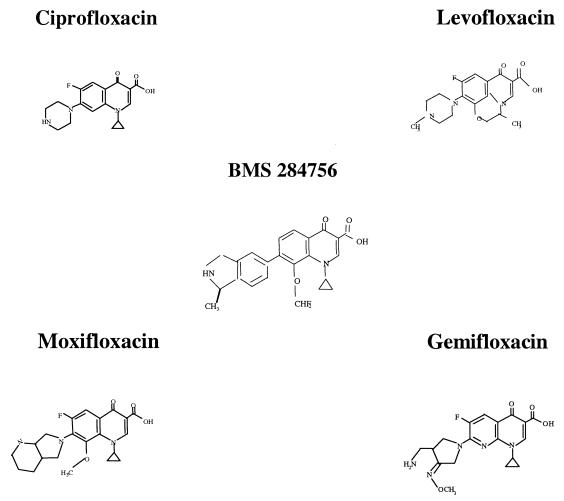
BMS-284756 is a novel des-fluoro(6) quinolone, which differs from recently approved quinolones in that it lacks a fluorine at the C-6 position (Fig. 1). It has been found to have excellent activity against gram-positive bacteria, including staphylococci (3). In contrast to ciprofloxacin, DNA gyrase is the primary target of BMS-284756 in S. aureus (2).
FIG. 1.
Chemical structures of quinolone compounds.
This study determined the activity of BMS-284756 against S. aureus isolates collected from hospitals across Canada and the United States prior to 1999. The QRDRs were analyzed in 28 isolates for which BMS 284756 MICs ranged from 0.007 to 64 μg/ml in order to assess the effect of existing QRDR mutations on the activity of BMS-284756.
A total of 602 geographically and genetically diverse clinical isolates of S. aureus were used in this study. These included 86 MRSA and 6 methicillin-sensitive S. aureus (MSSA) isolates selected from the New York City Public Health Research Institute's culture collection and represented more than 29 different protein A genotypes (B. N. Kreiswirth, unpublished results). A total of 382 MSSA and 128 MRSA strains were obtained from the Canadian Bacterial Surveillance Network during 1997 and 1998, the details of which are described elsewhere (1). Isolates were screened for methicillin resistance as per current National Committee for Clinical Laboratory Standards (NCCLS) guidelines (8). S. aureus ATCC 29213 and ATCC 43300 were used for quality control purposes.
The MIC of each antimicrobial agent was determined for all staphylococci by broth microdilution methods in accordance with NCCLS guidelines (8). The antimicrobial agents were supplied by their respective manufacturers or purchased from Sigma (St. Louis, Mo.). To classify the S. aureus isolates as either MRSA or MSSA, isolates for which oxacillin MICs indicated borderline resistance were confirmed as MRSA by demonstrating the presence of the PBP 2′ protein by using the Denka Seiken monoclonal antibody kit (Med-Ox, Ottawa, Ontario, Canada).
Isolates were randomly selected for QRDR analysis from each BMS-284756 MIC cluster (MIC range, 0.007 to 64 μg/ml). The QRDRs of grlA, grlB, gyrA, and gyrB were amplified by PCR. Briefly, a loop of overnight growth on Columbia nutrient agar supplemented with 5% sheep blood was suspended in 100 μl of lysis buffer (100 mM NaCl, 10 mM Tris-HCl [pH 8.3], 1 mM EDTA, 1% Triton X-100) and boiled for 10 min; 10 μl of the supernatant was used as a DNA template in a 50-μl reaction volume. Primers were based on published sequences (4, 10). Amplification products were purified with either the QIAquick PCR purification kit (Qiagen Inc., Mississauga, Ontario, Canada) or the Concert rapid PCR purification kit (Life Technologies, Burlington, Ontario, Canada).
DNA sequencing was performed by ABI Prism Big Dye terminator cycle sequencing with the ABI 377 automated sequencer (PE Applied Biosystems, Mississauga, Ontario, Canada). Nucleotide and protein sequence comparisons were performed with the multiple alignment sequence function of Vector NTI Suite software (InforMax Inc., Bethesda, Md.). The GenBank accession numbers for wild-type sequences used for comparison purposes were D67075 for grlA and grlB, D10489 for gyrA, and D10489 for gyrB.
The results of in vitro susceptibility testing are summarized in Table 1. The prevalence of ciprofloxacin resistance (MIC ≥ 4 μg/ml) was 3% (13 of 388) in MSSA and 71% (152 of 214) in MRSA. BMS-284756 had excellent activity against ciprofloxacin-susceptible isolates, with MICs at which 50% of isolates were inhibited of 0.015 μg/ml and MICs at which 90% of isolates were inhibited of 0.03 μg/ml. However, for 71 (41%) of the non-ciprofloxacin-susceptible strains, MICs were >2 μg/ml, the breakpoint used in the BMS-284756 clinical trials (3).
TABLE 1.
Distribution frequencies of the MICs for S. aureus isolates with different methicillin and ciprofloxacin susceptibilities
| Antimicrobial | Susceptibility category (n)a | No. of isolates for which the MIC (μg/ml) was:
|
||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ≤0.007 | 0.015 | 0.03 | 0.06 | 0.12 | 0.25 | 0.5 | 1 | 2 | 4 | 8 | 16 | 64 | ||
| Ciprofloxacin | MS (388) | 333b | 37 | 5 | 1 | 3 | 9c | |||||||
| MR (214) | 52b | 8 | 2 | 1 | 6 | 145c | ||||||||
| BMS-283756 | CS (430) | 43b | 275 | 97 | 15 | |||||||||
| CI (7) | 2 | 4 | 1 | |||||||||||
| CR (165) | 3 | 3 | 0 | 0 | 1 | 5 | 16 | 66 | 69 | 1 | 0 | 1 | ||
MS, methicillin sensitive; MR, methicillin resistant; CS, ciprofloxacin susceptible; CI, ciprofloxacin intermediate; CR, ciprofloxacin resistant.
The number of isolates was less than or equal to this value.
The number of isolates was greater than or equal to this value.
For the purposes of this study, we assumed that a BMS-284756 MIC for S. aureus indicating susceptibility was 0.015 μg/ml (Table 1) and a ciprofloxacin MIC indicating susceptibility was 0.12 μg/ml (11). Twenty-one of the 28 isolates examined for QRDR mutations had at least one amino acid substitution in either GrlA, GrlB, or GyrA. No mutations in gyrB were noted for any of the isolates examined. The QRDR mutations identified are summarized in Table 2. None of the 11 isolates for which ciprofloxacin MICs were 0.12 to 0.25 μg/ml had amino acid substitutions in GrlA or GyrA hot spots (Ser-80 or Glu-84 and Ser-84 or Glu-88, respectively.) Of the 14 isolates for which BMS-284756 MICs were 0.007 to 0.03 μg/ml, four had Ser-80-Phe (Tyr) GrlA substitutions. The failure of such mutations to increase the BMS-284756 MIC supports the suggestion by Discotto et al. (2) that DNA gyrase is the primary target for this compound in S. aureus. There were two isolates for which BMS-284756 MICs were fourfold greater than the BMS-284756 susceptibility MIC but for which ciprofloxacin MICs were the same as the ciprofloxacin susceptibility MIC. These data support those of Roychoudhury et al. (9), who found that the relative MICs of nonfluorinated and fluorinated quinolones do not always correlate. As Roychoudhury suggested, this may be because certain resistance mechanisms are effective against one group of quinolones and not the other. Furthermore, it is likely that the mechanisms referred to are not quinolone specific, considering that these isolates have never been exposed to the nonfluorinated compounds. Isolates of S. aureus for which BMS-284756 MICs were 0.25 to 4 μg/ml had a grlA and a gyrA hot spot mutation (Table 2). Although 15 of the isolates analyzed had an amino acid substitution in GrlB, their association with resistance remains unclear (Table 2).
TABLE 2.
BMS-284756 MICs for S. aureus and associated QRDR amino acid substitutions in topoisomerase IV and DNA gyrase
| MIC (μg/ml)
|
No. of isolates | Amino acid substitutiona
|
|||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| GrlA
|
GyrA
|
GrlB
|
|||||||||
| BMSr 284756 | Ciprofloxacin | S80 | E84 | P144 | S84 | S85 | E88 | E422 | E471 | H478 | |
| 64 | 64 | 1 | F | K | L | P | |||||
| 8 | 256 | 1 | Y | G | L | ||||||
| 4 | 32-256 | 3 | F | L | D | ||||||
| 2 | 128 | 1 | F | L | D | ||||||
| 2 | 16 | 1 | K | L | D | ||||||
| 2 | 16 | 1 | F | L | |||||||
| 1 | 128 | 1 | F | L | K | ||||||
| 0.5 | 16 | 1 | F | S | K | D | |||||
| 0.5 | 8 | 1 | F | K | |||||||
| 0.25 | 8 | 1 | F | G | |||||||
| 0.06 | 0.12 | 1 | Y | ||||||||
| 0.03 | 2 | 1 | G | Y | |||||||
| 0.03 | 1 | 1 | F | S | D | ||||||
| 0.03 | 1 | 1 | F | ||||||||
| 0.03 | 0.25 | 1 | D | ||||||||
| 0.015 | 2 | 1 | Y | D | |||||||
| 0.015 | 1 | 1 | F | S | D | ||||||
| 0.007 | 0.12-0.25 | 2 | S | D | |||||||
Y, tyrosine; F, phenylalanine; K, lysine; G, glycine; P, proline; S, serine; D, aspartic acid; E, glutamic acid; L, leucine; H, histidine.
The one isolate for which the BMS-284756 MIC was 8 μg/ml and the ciprofloxacin MIC was 256 μg/ml had two grlA hot spot mutations and one gyrA hot spot mutation, whereas the isolate for which the BMS-284756 MIC was 64 μg/ml and the ciprofloxacin MIC was 64 μg/ml had two mutations at each hot spot. Although our data suggest that in S. aureus, grlA mutations precede gyrA mutations in the development of ciprofloxacin resistance, it is less clear as to the advantage of double mutations in grlA and gyrA when taking into consideration that these were selected for at a time when they were most likely exposed only to ciprofloxacin. Since most types of antibiotic resistance impose a cost on the fitness of the organism, it is tempting to speculate that these have been selected as fitness-compensating mutations and not resistance-conferring mutations. Regardless, these additional mutations significantly affect the susceptibility of S. aureus to the in vitro activity of BMS-284756.
Acknowledgments
This work was supported by a grant from the Canadian Bacterial Diseases Network. D.J.B. is a recipient of a Medical Research Council of Canada postdoctoral research fellowship.
REFERENCES
- 1.Chen, D. K., A. McGeer, J. C. de Azavedo, D. E. Low, et al. 1999. Decreased susceptibility of Streptococcus pneumoniae to fluoroquinolones in Canada. N. Engl. J. Med. 341:233-239. [DOI] [PubMed] [Google Scholar]
- 2.Discotto, L. F., L. E. Lawrence, K. L. Denbleyker, and J. F. Barrett. 2001. Staphylococcus aureus mutants selected by BMS-284756. Antimicrob. Agents Chemother. 45:3273-3275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Fung-Tomc, J. C., B. Minassian, B. Kolek, E. Huczko, L. Aleksunes, T. Stickle, T. Washo, E. Gradelski, L. Valera, and D. P. Bonner. 2000. Antibacterial spectrum of a novel des-fluoro(6) quinolone, BMS-284756. Antimicrob. Agents Chemother. 44:3351-3356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Gootz, T. D., R. P. Zaniewski, S. L. Haskell, F. S. Kaczmarek, and A. E. Maurice. 1999. Activities of trovafloxacin compared with those of other fluoroquinolones against purified topoisomerases and gyrA and grlA mutants of Staphylococcus aureus. Antimicrob. Agents Chemother. 43:1845-1855. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Hooper, D. C. 1999. Mechanisms of fluoroquinolone resistance. Drug Resist. Update 2:38-55. [DOI] [PubMed] [Google Scholar]
- 6.Hooper, D. C. 2001. Mechanisms of action of antimicrobials: focus on fluoroquinolones. Clin. Infect. Dis. 32:S9-S15. [DOI] [PubMed]
- 7.Kaatz, G. W., S. M. Seo, and C. A. Ruble. 1993. Efflux-mediated fluoroquinolone resistance in Staphylococcus aureus. Antimicrob. Agents Chemother. 37:1086-1094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.National Committee for Clinical Laboratory Standards. 2000. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically. Approved standard M7-A5. NCCLS document M100-S10/M7. National Committee for Clinical Laboratory Standards, Wayne, Pa.
- 9.Roychoudhury, S., C. E. Catrenich, E. J. McIntosh, H. D. McKeever, K. M. Makin, P. M. Koenigs, and B. Ledoussal. 2001. Quinolone resistance in staphylococci: activities of new nonfluorinated quinolones against molecular targets in whole cells and clinical isolates. Antimicrob. Agents Chemother. 45:1115-1120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Schmitz, F. J., M. E. Jones, B. Hofmann, B. Hansen, S. Scheuring, M. Luckefahr, A. Fluit, J. Verhoef, U. Hadding, H. P. Heinz, and K. Kohrer. 1998. Characterization of grlA, grlB, gyrA, and gyrB mutations in 116 unrelated isolates of Staphylococcus aureus and effects of mutations on ciprofloxacin MIC. Antimicrob. Agents Chemother. 42:1249-1252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Wiedemann, B., and H. Grimm. 1996. Susceptibility to antibiotics, p. 900-1168. In V. Lorian (ed.), Antibiotics in laboratory medicine. Williams and Wilkins, Baltimore, Md.