Abstract
Drug resistance in malarial parasites has become a major obstacle in the control of the disease. Strategies are urgently needed to control the development of resistance and to possibly reverse existing resistance. One key element required to reverse malaria drug resistance is for the parasites to “pay” a biological “cost” or suffer a loss of fitness when acquiring resistance to antimalarial drugs. Such a situation would be a disadvantage to the resistant parasites in the absence of drug pressure. We compared here the relative fitness of atovaquone-resistant Plasmodium falciparum K1 clones with single and double base mutations in their cytochrome b genes to their parent clones during erythrocytic stages in the absence of drug pressure. We found that the double amino acid mutation (M133I and G280D) is associated with a 5 to 9% loss of fitness and that the single amino acid change of M133I did not result in any detectable loss of fitness. Molecular modeling of the interaction of P. falciparum cytochrome b with ubiquinone led to the prediction that a loss of fitness of the malaria parasites would result from the G280D mutation due to its close proximity to the putative ubiquinone-binding site. This appears to have resulted in a weakening of the cytochrome b-ubiquinone complex, thereby causing the electron transport chain to become less efficient. Our results suggest that the prevalence of resistant parasites may decrease after the drug usage is discontinued.
During the past 3 decades, Plasmodium falciparum has developed resistance to almost every commonly available antimalarial drug, including chloroquine, pyrimethamine, cycloguanil, and sulfadoxine. The rapid spread of these resistant parasites has dramatically affected the successful treatment of patients; some drugs are now almost useless in many parts of the world.
To slow the development of resistance and prolong the effectiveness of antimalarial drugs, it is generally recommended that combinations of synergetic antimalarials should be used (37). It has also been suggested that the usage of a particular drug should be stopped when a certain level of treatment failure is observed (World Health Organization Expert Committee on Malaria, 1998 [http://mosquito.who.int/docs/ecr20_6.htm.]). Removal of the selection pressure exerted by the drug may enable the population of susceptible parasites to reestablish itself and gradually replace resistant parasites. However, for this strategy to be effective, three essential elements are required: (i) a proportion of parasites that are still susceptible to the drug (that is, they do not carry the mutation); (ii) drug-resistant parasites that are less fit compared to the susceptible wild-type parasites in the absence of drug pressure; and (iii) transmission levels that are high enough to result in some multiple infections.
Resistance to antimalarial drugs can result from mutations to a single target locus in the parasites (e.g., pyrimethamine resistance [9]) or from alterations to multiple loci (e.g., chloroquine resistance [12, 24]), or it can result from upregulation or duplication of genes associated with the influx or efflux of the antimalarial drugs (14). In many organisms, mutations that confer resistance to drugs clearly provide the organism with a survival advantage under these drug pressures. However, genetic mutations resulting in drug resistance are often associated with a loss of fitness to the organism due to alterations to functionally or structurally important regions of the target molecules (6, 11, 35, 38). This loss of fitness causes resistant organisms to become disadvantaged relative to the wild-type parasites when the drug pressure is withdrawn.
In the case of malaria, the relative fitness of the resistant parasites in the absence of drug pressure is unclear. Rosario et al. (26) reported that in mice infected with a mixture of chloroquine-resistant and -susceptible Plasmodium chabaudi, a higher proportion of chloroquine-resistant P. chabaudi survived to recrudesce between days 26 and 36 (i.e., 26 and 36 generations), suggesting a selection advantage to chloroquine-resistant parasites. However, in the same study, experiments on pyrimethamine-resistant parasites did not show a clear advantage for resistant over sensitive parasites. Shinondo et al. (31) reported that pyrimethanime-resistant and -sensitive Plasmodium berghei grew similarly during blood stages but that the resistant parasites proceeded through sporogonic development more slowly than the susceptible parasites. In P. falciparum, studies were performed to demonstrate that chloroquine-resistant parasites have transmission advantages over sensitive parasites, as indicated by higher parasitemia levels in asymptomatic carriers (36) and higher gametocyte prevalence and density (16, 25). However, none of these studies measure the relative fitness of resistant parasites to the sensitive parasites because of the continuous usage of chloroquine in the communities.
In this study, we compared the relative fitness of atovaquone-resistant K1 clones, with double (M133I, G280D) and single (M133I) base mutations in the cytochrome b gene, to their parent clones without these mutations. We found that the double amino acid mutation (M133I, G280D) resulted in approximately a 5 to 9% loss in fitness compared to the wild type. However, the single-amino-acid change M133I did not result in any detectable loss in fitness. We propose that the observed loss of fitness results from the reduced binding of ubiquinone to cytochrome b caused by the G280D mutation occurring close to a putative atovaquone-binding and ubiquinone-binding site.
MATERIALS AND METHODS
Parasites. (i) Wild-type parasites.
P. falciparum K1-P, K1-B, K1-C, K1-D, and K1-E are wild-type sibling clones of K1 originating from five separate parasite-containing wells produced by a limiting dilution procedure (19). These parasites are considered clonal since only 7% of the wells contained parasites after the limiting dilution procedure.
(ii) Mutant parasites.
P. falciparum ATV-M1, ATV-M4-1, and ATV-M4-2 are mutant clones selected by exposing K1-P to atovaquone (19). ATV-M1 is a clone of resistant parasites that contains a single-base mutation in the parasite cytochrome b gene resulting in a M133I change at the amino acid level. ATV-M4-1 and ATV-M4-2 are two sibling clones of resistant parasites that each contains two single-base mutations in the cytochrome b gene, resulting in amino acid changes at M133I and G280D.
In vitro cultivation of P. falciparum.
Parasites were maintained in vitro under conditions modified from those described by Trager and Jensen (34).
Growth competition experiments.
The wild-type and mutant clones of parasites were synchronized by using 5% sorbitol for the ring stage (20) and 63% Percoll for the schizont stage (28). The number of infected erythrocytes in each culture was determined by multiplying the number of erythrocytes by the percentage of parasitemia. Two growth competition experiments were carried out as follows. In experiment 1, K1-P and K1-E were each mixed with either ATV-M4-1 or ATV-M4-2 at a ratio of ca. 1:1 of wild-type to mutant parasites. The wild-type and mutant parasites were also cultured in separate flasks as controls. All cultures were maintained in duplicate for 108 days, after which they were cryopreserved (8). In an extension of experiment 1, cryopreserved wild-type and mutant control parasites from experiment 1 were thawed, remixed at a 1:1 ratio, and cultured for a further 60 days. In experiment 2, K1-P, -B, -C, -D, and -E were each mixed with ATV-M1 at a ratio of ca. 1:1. Again, the wild-type and mutant parasites were cultured in separate flasks as controls. All cultures were maintained in duplicate for 105 days.
DNA extraction.
A fraction of the parasite cultures were collected every 20 days and parasite genomic DNA extracted by using a method previously described (7). The extracted DNA were resuspended in 100 μl of TE (10 mM Tris pH 8.0 and 1 mM EDTA, pH 8.0) and stored at −20°C. The DNA was diluted 100-fold for use in the competitive PCR.
Quantitative PCR.
A competitive PCR was performed to quantify the ratio of wild-type to mutant parasites. A pair of primers, cytb1 (5′-CTC TAT TAA TTT AGT TAA AGC ACA C-3′) and cytb4 (5′-ATT TTT AAT GCT GTA TCA TAC CCT-3′), were designed to amplify a fragment at the 5′ end of cytochrome b flanking the M133I mutation. A forward primer cytb-s (5′-TTG CAC CCC AAT AAC TC-3′) was designed to anneal to the wild-type cytochrome b gene with its 3′ end matching ATG (M133), and a reverse primer cytb-m1 (5′-TAC CAT GGG GTC AAA TA-3′) was designed to anneal to the mutant type cytochrome b with its 3′ base matching ATA (I133). PCR was performed with 2 mM Mg2Cl, 0.2 mM concentrations of each deoxynucleoside triphosphate (Promega, Madison, Wis.), 75 ng of each of the four primers, 1.25 U of AmpliTaqGold (PE Applied Biosystems), and 1 μl of diluted DNA. The reaction mixture was initially heated at 93°C for 10 min and then underwent a touchdown cycling program. The cycling conditions were as follows: 20 cycles at 93°C for 50 s, an initial 65°C decreasing by 1°C per cycle for 50 s, and 70°C for 50 s, followed by 20 cycles of 93°C for 50 s, 45°C for 50 s, and 70°C for 50 s.
Quantitation.
PCR products were separated on a 2% agarose gel and visualized on a UV transilluminator. Gel images were captured by using a Kodak Digital Science EDAS 120 system, and net intensities for wild-type and mutant cytochrome b bands were measured by using Kodak ID analysis software. A series of controls were made by mixing genomic DNA from wild-type parasites with that of mutant parasites at 1:1, 3:1, 10:1, 30:1, and 100:1 ratios. The controls were amplified in parallel with the samples. After each PCR, a standard curve was generated from the controls and the ratio of wild-type to mutant parasites in a sample was determined based on the standard curve.
DNA sequencing.
Full-length cytochrome b was amplified with the primers cytb1 and cytb2 (19) and then sequenced from both orientations.
DNA fingerprinting.
Parasite genomic DNA collected from each sample at the beginning and end of the growth competition experiments was digested with restriction enzyme AccI, electrophoresed on a 1% agarose gel, and transferred to Hybond-N+ membranes. A digoxigenin (DIG)-labeled 7H8/6-specific probe (21) was generated by PCR with the M13 forward and reverse primers and DIG-labeled dUTPs (Roche Diagnostics, Pty., Ltd., Castle Hill, Australia). The probe was hybridized to the membrane overnight at 65°C. The membranes were washed three times with 2× SSC (1× SSC is 0.15 M NaCl plus 0.015 M sodium citrate)-0.1% sodium dodecyl sulfate at 65°C. An anti-DIG antibody alkaline phosphatase conjugate (Roche) was bound to the probe and detected by using CDP-Star chemiluminescence substrate (Roche) exposed to Kodak Biomax Light-1 film (Amersham Pharmacia Biotech, Pty., Ltd., Castle Hill, Australia).
Chromosomal mapping.
Parasite chromosomal DNA was prepared in 0.8% low-melting-point agarose blocks (18) and separated by pulsed-field gel electrophoresis (29) in a 0.8% agarose gel by using the CHEF-DR III system (Bio-Rad Laboratories, Hercules, Calif.). Electrophoresis was performed for 24 h at 3 V/cm, with a switch time of 270 to 500 s, and then 25 h at 2.4 V/cm and 500 to 900 s, followed by 20 h at 1.8 V/cm and 1,200 s, and then finally 18 h at 1.8V/cm and 1,600 s (M. Tchavtchitch, Queensland Institute of Medical Research [unpublished data]). The electrophoresis buffer was maintained at a constant 14°C and changed every 24 h. The separated chromosomes were visualized by staining with ethidium bromide. The gel was transferred to a Hybond-N+ membrane and hybridized with a DIG-labeled 7H8/6 probe.
Degree of loss of fitness.
The loss of fitness of the mutant parasites relative to the wild type was determined by assuming that the ratio of mutant to wild-type parasites is defined as follows: mutant/wild-type ratio = 1:(starting ratio/[1 − loss]n), where “loss” is the theoretical loss of fitness of the mutant (as a proportion), and n is the number of generations since the start of the experiment.
Least-squares linear regression of the form ln(experimental ratio) = c + mn was used to estimate the loss of fitness, where m = −ln(1 − loss) and c = ln(starting ratio).
Molecular modeling.
A homology model of P. falciparum CYT b (19) was used to investigate the possible effect of mutations in cytochrome b to the ubiquinone binding site in the same molecule. The ubiquinone molecule was docked to the CYT b model based on predictions by Bartoschek et al. (3) and Crofts et al. (10) and by using the program GOLD (17). The resulting complex and the M133I and M133I-G280D mutated complexes were energy minimized by using the program Discover.
Molecular simulations.
The models of the wild-type, M133I, and M133I-G280D mutant cytochrome b P. falciparum models were simulated in vacuo at 300K by using the GROMACS 3.0 package (4, 22) and the GROMOS 43a1 force field (30) in a 0.5-ns interval with an integration step size of 1 fs. Chemical bonds were restricted only at the start by using the SETTLE algorithm (27) and were represented by a harmonic potential throughout the simulation. Position restraints were imposed on all atoms except hydrogens with a force constant of 1 MJ/mol · nm2. Temperature coupling was carried out by using the Berendsen temperature-coupling scheme (5). Short- and long-range electrostatics were simulated by using a Coulomb type interaction with a distance to neighbors of 1 nm, a van der Waals cutoff of 1.4 nm, an electrostatic cutoff of 1.4 nm, and a dielectric constant of 1.
RESULTS
Quantitative PCR and standard curve.
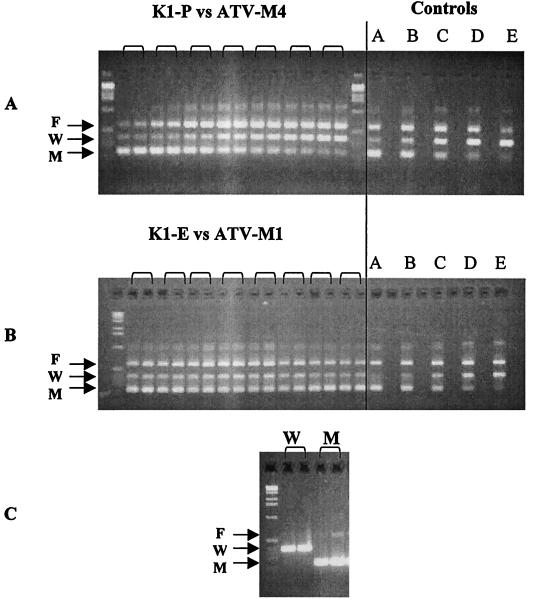
The quantitative PCR generated three specific bands: band F, amplified with primers 1 and 4; band W, amplified with primers 1 and S; and band M, amplified with primers 4 and M1 (Fig. 1). The ratio of band intensity between bands W and M was measured for the control samples in which wild-type and mutant parasite DNAs were mixed at 1:1, 3:1, 10:1, 30:1, and 100:1. A log linear regression curve was calculated and used as the standard curve.
FIG. 1.
Gel electrophoresis of PCR product showing the change of the wild-type/mutant ratio over time in duplicate in K1-P and ATV-M4-1 mixtures (A), K1-E and ATV-M1 mixtures (B), and wild type or mutant parasite alone (C). In the control panels, lanes A, B, C, D, and E represents the wild-type parasite DNA mixed with mutant parasite DNA at 1:1, 3:1, 10:1, 30:1, and 100:1, respectively. Bands: F, full-length product; W, wild type; M, mutant.
ATV-M4 has a slower growth rate than the wild-type parasites.
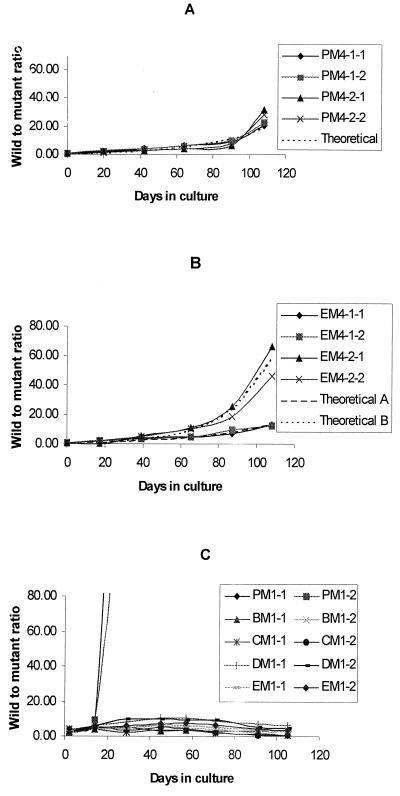
Two sibling clones of ATV-M4 were each mixed with two sibling clones of K1 (K1-P and K1-E) and cultured continuously for 108 days. The starting ratios of wild-type to mutant parasites were between 0.40 and 0.90, and these ratios increased steadily to beween 18 and 66 by day 108 (Fig. 2A and B; Table 1). Figure 1 shows an example of the change in the ratio of wild-type to mutant type occurring in the K1-P and ATV-M4-1 mixture as measured by quantitative PCR. During the culture period, the wild-type band (W) increased in intensity, while the mutant band decreased in intensity. No change was seen in the wild-type and mutant parasites cultured alone.
FIG. 2.
Change of wild-type/mutant ratio. (A) K1-P and ATV-M4 mixtures (PM4-1-1/2 and PM4-2-1/2); (B) K1-E and ATV-M4 mixtures (EM4-1-1/2 and EM4-2-1/2); (C) K1-P, -B, -C, -D, -E and ATV-M1 mixtures (PM1-1/2, BM1-1/2, CM1-1/2, DM1-1/2, and EM1-1/2). Theoretical curves were produced by fitting a regression model to the grouped data for the four K1-P and ATV-M4 mixtures (R2 = 0.929, P < 0.001) (A) and the two groups of K1-E and ATV-M4 mixtures (theoretical A, R2 = 0.918, P < 0.001 for EM4-1-1 and EM4-1-2 data; theoretical B, R2 = 0.969, P < 0.001 for EM4-2-1 and EM4-2-2) (B).
TABLE 1.
Total change in the wild type (K1-P and K1-E) to mutant parasite (ATV-M4-1 and ATV-M4-2) ratios and estimated difference in growth rate during the initial 108-day period
| Wild-type/mutant mixture | End/start ratio | Predicted % difference in growth rate (95% CI)a |
|---|---|---|
| K1-P/M4-1-1 | 22.64 | 4.99 (3.5-6.5) |
| K1-P/M4-1-2 | 26.94 | 5.27 (3.8-6.7) |
| K1-P/M4-2-1 | 39.61 | 5.55 (3.1-8.0) |
| K1-P/M4-2-2 | 28.53 | 5.53 (3.7-7.3) |
| K1-E/M4-1-1 | 26.32 | 4.97 (2.4-7.4) |
| K1-E/M4-1-2 | 17.97 | 4.97 (3.6-6.3) |
| K1-E/M4-2-1 | 165.58 | 8.67 (6.3-11.0) |
| K1-E/M4-2-2 | 86.83 | 7.35 (5.7-8.9) |
CI, confidence interval.
DNA sequencing confirmed the changes observed by PCR. At the beginning of the experiment, sequencing of the DNA from the mixed cultures clearly showed mixed nucleotide signals at the bases where the mutations were located; at the end of the experiment only the wild-type nucleotides were seen in these positions (data not shown).
The relatively slow growth rate in ATV-M4 is stable.
Parasites that were frozen after 108 days of culture were thawed, remixed in a 1:1 ratio, and cultivated in vitro for an additional 60 days. The same trend observed in the first 108 days of culture was seen in the change of the wild-type/mutant ratio during this 60-day period (data not shown).
The difference in growth rates was predicted to be 5 to 9%.
For the first 108-day experiment, the growth rate of the ATV-M4 clone was predicted to be between 4.97 to 8.67% slower than that of the wild-type K1-P and K1-E (Table 1), averaging at 5.97%. There is agreement in the predicted ratio of growth rates between the replicates and across replicates and also in the experimental ratio (Fig. 2).
DNA fingerprinting with a stable probe 7H8/6 showed that all of the clones were identical in banding patterns at both the start and the end of the experiment (data not shown). These results indicate that there was no cross-contamination of the cultures from other sources and that no major genomic rearrangements occurred in these parasite clones during culturing.
ATV-M1 has the same growth rate as the wild-type parent parasites.
ATV-M1 was mixed with five sibling wild-type K1 clones separately and cultured for 105 days in vitro. As shown in Fig. 2C, the starting ratio of wild-type to ATV-M1 parasites varied from 2.59 to 4.28. The ratios of K1-B, -C, -D, and -E to ATV-M1 fluctuated during the time of culture but did not change substantially from the start to the end of the experiment (Fig. 2C; Table 2), i.e., ATV-M1 grew at approximately the same rate as K1-B, -C, -D, and -E (1.7 to −0.1% faster; Table 2). The ratio of the growth rate of K1-P to ATV-M1 changed dramatically during the culture. By day 29 the K1-P to ATV-M1 ratio was higher than 100:1, and the mutant type was not detectable from day 45 onward (Fig. 2C). The estimated difference in the growth rate of K1-P compared to ATV-M1 was between 29.1 and 36.4% higher (Fig. 2C, Table 2). To investigate whether significant genomic or chromosomal rearrangement occurred in this K1-P clone during the culture period, DNA fingerprinting and chromosomal mapping were performed. No genetic or chromosomal rearrangement was detected in this clone of the parasite during the period of culturing.
TABLE 2.
Total change in the wild-type (K1-P, -B, -C, -D, and -E)/mutant parasite (ATV-M1) ratios and estimated differences in growth rate during a 105-day period
| Wild-type/mutant mixture | End/start ratio | Ratio (%) of growth rate (95% CI)a |
|---|---|---|
| K1-P/M1-1 | >110.08 | 36.4∗ |
| K1-P/M1-2 | >72.99 | 29.1∗ |
| K1-B/M1-1 | 1.30 | −0.1 (−0.6 to 0.4) |
| K1-B/M1-2 | 1.02 | −0.3 (−1.0 to 0.4) |
| K1-C/M1-1 | 0.19 | −1.2 (−1.8 to −0.6) |
| K1-C/M1-2 | 0.19 | −1.7 (−2.9 to −0.6) |
| K1-D/M1-1 | 1.05 | −0.1 (−0.8 to 0.6) |
| K1-D/M1-2 | 1.51 | 0.1 (−0.9 to 1.1) |
| K1-E/M1-1 | 0.73 | −0.3 (−0.9 to 0.4) |
| K1-E/M1-2 | 0.78 | −0.2 (−0.9 to 0.5) |
CI, confidence interval. ∗, Confidence intervals were not calculated since there are only three datum points.
The slower growth rate is associated with a change in the ubiquinone-binding site.
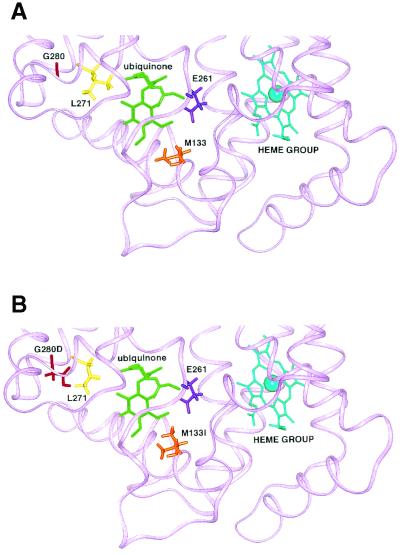
Molecular modeling revealed that the G280D change in ATV-M4 caused an alteration of the orientation of a putative ubiquinone-binding residue, L271(Fig. 3). After energy minimization of the ubiquinone-cytochrome b complex containing the G280D mutation, steric interference and an altered electrostatic environment was observed in the putative ubiquinone-binding site (Fig. 3). In contrast, the M133I mutation was relatively conserved both chemically and structurally and caused little variation when minimized (Fig. 3).
FIG. 3.
(A) Ribbon representation of wild-type cytochrome b bound to ubiquinone. Amino acid side chains, ubiquinone, and heme are represented in stick form, and the Fe center of heme is represented in CPK form. (B) Ribbon representation of G280D-M133I cytochrome b bound to ubiquinone after energy minimization. The G280D mutation affects L271 and hence the ubiquinone-binding site.
Molecular simulations of the M133I, M133I-G280D, and wild-type cytochrome b-modeled proteins without ubiquinone resulted in disruption to the structure when G280D was introduced. These results support the notion that the G280D mutation is responsible for the loss binding of ubiquinone in the putative ubiquinone-binding site (Fig. 3).
DISCUSSION
Drug resistance in malarial parasites often results from mutations in a gene that then expresses a protein with a lower binding capacity for a particular drug or the duplication or upregulation of a gene that leads to an increased expression of a particular protein. In each case, changes in a single locus or in multiple loci may be involved in resistance. In this study, we studied the loss of fitness associated with mutations that occurred in a single locus, the cytochrome b locus of the parasite genome.
The atovaquone-resistant mutants ATV-M1 and ATV-M4 were selected by exposing the parent parasite clone to atovaquone and then cloning the resistant parasites. The selection and cloning of the resistant parasites was carried out over ca. 110 days, i.e., the mutant parasite clones used in the fitness experiment were 55 generation lineages from the parent parasite clone. Thus, genomic differences between the mutant and parent parasites were limited. The resistant mutations occurred in the cytochrome b gene of the mitochondria genome in the mutant parasites. When the mutant parasites (ATV-M4) were mixed with parent parasites at ca. 1:1 ratio and then cultured in the same flask and under the same conditions, the proportion of the ATV-M4 parasites (two amino acid changes) decreased steadily over time. Similar changes were observed in replicates of each parent-mutant mixture. Therefore, this change in proportion did not result from clonal variation but was due to a relatively slower growth rate of the mutant parasite ATV-M4 compared to the wild-type K1-P and K1-E. The results demonstrate that there is a loss of fitness associated with the ATV-M4 mutant parasites.
In contrast, for ATV-M1 that has a single amino acid change, the proportion of wild-type to mutant parasites did not change over time in four of five mixtures, suggesting that there was no loss of fitness associated with ATV-M1 since the mutant parasites had the same growth rates as four of the five wild-type parasites. In the fifth mixture, in which the parent clone K1-P was mixed with ATV-M1 and a rapid takeover of K1-P was observed, it is likely that uncharacterized changes occurred in this K1-P genome that allowed the parasite to grow faster than ATV-M1, since a similar change was not observed in the four other mixtures with ATV-M1. This change may have occurred in the period between the thawing of this K1-P and the start of this particular experiment since K1-P was the only parasite clone that had been substantially cultured (∼100 days) prior to the start of experiment, and a different vial of K1-P used in the ATV-M4 mixtures did not exhibit the same behavior. Chromosome mapping and DNA fingerprinting of the parasites at the start and the end of culture failed to detect any rearrangement in the genome and no extra mutations were found in the cytochrome b gene of the parasite, suggesting that the change or changes in this particular K1-P are likely to be mutations that have occurred in a locus outside of cytochrome b.
The loss of fitness in ATV-M4 was calculated to be between 5 and 9% (averaging 5.97%). This estimate demonstrates the biological disadvantage sustained by the mutant parasites at the erythrocytic stages. Whether the mutations also alter parasite transmission abilities, such as the production and viability of gametocytes, the fertilization process, and development in mosquitoes, requires further investigation.
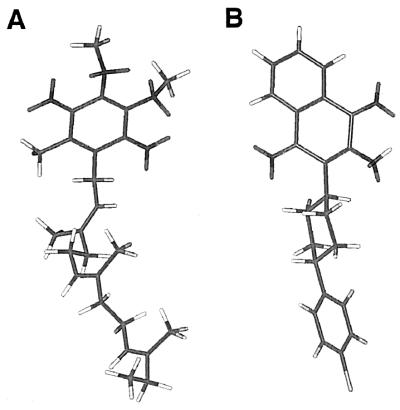
Atovaquone is widely used in combination with proguanil under the trade name of Malarone. Atovaquone acts against malaria parasites by inhibiting mitochondrial electron transport (15) and collapsing mitochondrial membrane potential (32). Due to its structural similarity to ubiquinone, it is postulated that the action is achieved by binding to cytochrome b as a ubiquinone analogue (Fig. 4). Therefore, mutations at the atovaquone-binding site may affect the parasite's ability to bind ubiquinone. Molecular modeling predicted that the loss of fitness in ATV-M4 had resulted from the G280D mutation interfering with the binding of parasite cytochrome b to ubiquinone. The G280D mutation was predicted to occur in close vicinity to the putative ubiquinone-binding site, causing a loss of the hydrophobic interaction between L271 and ubiquinone. This change was not fatal to the parasites and explains the reduced efficiency of the parasites electron transport chain mechanism. In contrast, the M133I change was relatively conserved and, although M133 is putatively involved directly in binding to ubiquinone, the change had little effect on the binding. In the presence of atovaquone, the loss of fitness was outweighed by the parasite's ability to resist the drug pressure. However, when drug pressure was withdrawn, the resistant parasites were disadvantaged due to an impaired electron transport chain. The resistance profile of the mutants provides additional support for this differential effect caused by these mutations. M133I, which is predicted to have a moderate effect on the atovaquone binding, conferred a 25-fold resistance to atovaquone, while the combination of M133I and G280D, predicted to significantly affect the drug binding site, conferred an 800-fold reduction in susceptibility to atovaquone (19).
FIG. 4.
Chemical structures of ubiquinone (truncated tail) (A) and atovaquone (B).
This is the first study showing a loss of fitness associated with drug-resistant P. falciparum. The outcome has significant implications in the formulation of drug policies in geographic areas where malaria is endemic, especially in countries such as Africa, where malaria transmission is high. Our results suggest that it is important to monitor the prevalence of resistance and that the usage of a drug should be stopped before the prevalence of resistance reaches a high level. These may allow the prevalence of resistance to drugs to be reduced over a number of years. In the case of single locus resistance, the speed of this decrease in resistance depends first of all on the prevalence of resistance at the time drug usage is stopped. For instance, if drug usage is ceased when resistance prevalence is only 50%, the reversal will occur in a shorter time than if the prevalence has reached 90%. Second, the speed of the decrease in the prevalence of drug resistance depends on the frequency of mixed infections in the human host since the decreased fitness will only be manifested in competition with wild-type parasites. Generally, the frequency of mixed infections will increase as the transmission intensity increases. Thus, the reversal of drug resistance will be more difficult in populations in low-transmission areas where infections are predominantly clonal (1, 2, 13). Finally, the degree of loss of fitness associated with the development of resistance will determine the speed of the decline. This degree of loss may vary between various drugs. It should be noted that, although the loss was stable in vitro culture system for a period of ca. 170 days, it is possible that the mutant parasites may develop compensatory mutations after a number of generations. These compensatory mutations might decrease the degree of loss or even totally cancel the deficit they carried.
Although our study only examined loss of fitness associated with atovaquone-resistant parasites, our results highlight the possibility of decreasing the prevalence of resistance to other drugs. In fact, decreases in the prevalence of resistance to chloroquine have been observed in several countries where drug usage has officially ceased for a number of years. A study conducted in Hainan, China, showed that the prevalence of chloroquine-resistant P. falciparum on Hainan island decreased from 84.2% in 1981 to 40% in 1991 after the official cessation of chloroquine usage on the island in 1979 (23). A recent study in Vietnam also demonstrated a similar trend after chloroquine was replaced with artemisinin derivatives (33). Loss of fitness in resistant parasites could be one of the reasons for these declines. This presents a possibility that an antimalarial drug may be reintroduced after the drug usage has ceased for a number of years and the prevalence of resistant parasites has decreased. To avoid an immediate reselection of mutant parasites, the drug should be reintroduced in combination with another antimalarial drug.
Acknowledgments
The project is funded by a NIH grant (AI47500-2), and the work was performed at the Australian Army Malaria Institute.
We thank David van der Spoel for advice with the GROMACS package and the High Performance Computing group from the Queensland University of Technology for supplying hardware.
REFERENCES
- 1.Ariey,F., W. Chalvet, D. Hommel, C. Peneau, A. Hulin, O. Mercereau-Puijalon, J. B. Duchemin, J. L. Sarthou, J. M. Reynes, and T. Fandeur. 1999. Plasmodium falciparum parasites in French Guiana: limited genetic diversity and high selfing rate. Am. J. Trop. Med. Hyg. 61:978-985. [DOI] [PubMed] [Google Scholar]
- 2.Arnot, D. 1998. Unstable malaria in Sudan: the influence of the dry season: clone multiplicity of Plasmodium falciparum infections in individuals exposed to variable levels of disease transmission. Trans. R. Soc. Trop. Med. Hyg. 92:580-585. [DOI] [PubMed] [Google Scholar]
- 3.Bartoschek, S., M. Johansson, B. H. Geierstanger, J. G. Okun, C. R. Lancaster, E. Humpfer, L. Yu, C. A. Yu, C. Griesinger, and U. Brandt. 2001. Three molecules of ubiquinone bind specifically to mitochondrial cytochrome bc1 complex. J. Biol. Chem. 276:35231-35234. [DOI] [PubMed] [Google Scholar]
- 4.Berendsen, H. J. C., D. van der Spoel, and R. van Drunen. 1995. GROMACS: a message-passing parallel molecular dynamics implementation. Comp. Physiol. Commun. 91:43-56. [Google Scholar]
- 5.Berendsen, H. J. C., J. P. M. Postma, A. DiNola, and J. R. Haak. 1984. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81:3684-3690. [Google Scholar]
- 6.Billington, O. J., T. D. McHugh, and S. H. Gillespie. 1999. Physiological cost of rifampin resistance induced in vitro in Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 43:1866-1869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Cheng, Q., G. Lawrence, C. Reed, A. Stowers, L. Ranford-Cartwright, A. Creasey, and A. Saul. 1997. Measurement of Plasmodium falciparum growth rate in vivo: a test of malaria vaccines. Am. J. Trop. Med. Hyg. 57:495-500. [DOI] [PubMed] [Google Scholar]
- 8.Chlichtherle, M., M. Walhlgren, H. Perlmann, and A. Scherf (ed.). 2000. Methods in malaria research, 3rd ed. American Type Culture Collection, Manassas, Va.
- 9.Cowman, A. F., M. J. Morry, B. A. Biggs, G. A. Cross, and S. J. Foote. 1988. Amino acid changes linked to pyrimethamine resistance in the dihydrofolate reductase-thymidylate synthase gene of Plasmodium falciparum. Proc. Nat. Acad. Sci. USA 85:9109-9113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Crofts, A. R., S. Hong, Z. Zhang, and E. A. Berry. 1999. Physicochemical aspects of the movement of the Rieske iron sulfur protein during quinol oxidation by the bc(1) complex from mitochondria and photosynthetic bacteria. Biochemistry 38:15827-15839. [DOI] [PubMed] [Google Scholar]
- 11.Devereux, H. L., V. C. Emery, M. A. Johnson, and C. Loveday. 2001. Replicative fitness in vivo of HIV-1 variants with multiple drug resistance-associated mutations. J. Med. Virol. 65:218-224. [DOI] [PubMed] [Google Scholar]
- 12.Djimde, A., O. K. Doumbo, J. F. Cortese, K. Kayentao, S. Doumbo, Y. Diourte, A. Dicko, X. Z. Su, T. Nomura, D. A. Fidock, T. E. Wellems, C. V. Plowe, and D. Coulibaly. 2001. A molecular marker for chloroquine-resistant falciparum malaria. N. Engl. J. Med. 344:257-263. [DOI] [PubMed] [Google Scholar]
- 13.Figtree, M., C. J. Pasay, R. Slade, Q. Cheng, N. Cloonan, J. Walker, and A. Saul. 2000. Plasmodium vivax synonymous substitution frequencies, evolution and population structure deduced from diversity in AMA 1 and MSP 1. Mol. Biochem. Parasitol. 108:53-66. [DOI] [PubMed] [Google Scholar]
- 14.Foote, S. J., J. K. Thompson, A. F. Cowman, and D. J. Kemp. 1989. Amplification of the multidrug resistance gene in some chloroquine-resistant isolates of P. falciparum. Cell 57:921-930. [DOI] [PubMed] [Google Scholar]
- 15.Fry, M., and M. Pudney. 1992. Site of action of the antimalarial hydroxynaphthoquinone, 2-[trans-4-(4′-chlorophenyl)cyclohexyl]-3-hydroxy-1,4-naphthoquinone (566c80). Biochem. Pharmacol. 40:914-919. [DOI] [PubMed] [Google Scholar]
- 16.Handunnetti, S. M., D. M. Gunewardena, P. P. S. L. Pathirana, S. Weerasinghe, and K. N. Mendis. 1996. Features of recrudescent choloroquine-resistant Plasmodium falciparum infections confer a survival advantage on parasites and have implications for disease control. Trans. R. Soc. Trop. Med. Hyg. 90:563-567. [DOI] [PubMed] [Google Scholar]
- 17.Jones, G., P. Willett, R. C. Glen, A. R. Leach, and R. Taylor. 1997. Development and validation of a genetic algorithm for flexible docking. J. Mol. Biol. 267:727-748. [DOI] [PubMed] [Google Scholar]
- 18.Kemp, D. J., L. M. Corcoran, R. L. Coppel, H. D. Stahl, A. E. Bianco, G. V. Brown, and R. F. Anders. 1985. Size variation in chromosomes from independent cultured isolates of Plasmodium falciparum. Nature 315:347-350. [DOI] [PubMed] [Google Scholar]
- 19.Korsinczky, M., N. Chen, B. Kotecka, A. Saul, K. Rieckmann, and Q. Cheng. 2000. Mutations in Plasmodium falciparum cytochrome b that are associated with atovaquone resistance are located in a putative drug-binding site. Antimicrob. Agents Chemother. 44:2100-2108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Lambros, C., and J. P. Vanderberg. 1979. Synchronization of Plasmodium falciparum erythrocytic stages in culture. J. Parasitol. 65:418-420. [PubMed] [Google Scholar]
- 21.Limpaiboon, T., M. W. Shirley, D. J. Kemp, and A. Saul. 1991. 7H8/6, a multicopy DNA probe for distinguishing isolates of Plasmodium falciparum. Mol. Biochem. Parasitol. 47:197-206. [DOI] [PubMed] [Google Scholar]
- 22.Lindahl, E., B. Hess, and D. van der Spoel. GROMACS 3.0: a package for molecular simulation and trajectory analysis. J. Mol. Mod., in press.
- 23.Liu, D.-Q, R.-J. Liu, D.-X. Ren, D.-Q. Gao, C.-Y. Zhang, C.-P. Qiu, X.-Z. Cai, C.-F. Ling, A.-H. Song, and X. Tang. 1995. Changes in the resistance of Plasmodium falciparum to chloroquine in Hainan, China. Bull. W. H. O. 73:483-486. [PMC free article] [PubMed]
- 24.Reed, M. B., K.J Saliba, S. R. Caruana, K. Kirk, and A. F. Cowman. 2000. Pgh1 modulates sensitivity and resistance to multiple antimalarials in Plasmodium falciparum. Nature 403:906-909. [DOI] [PubMed] [Google Scholar]
- 25.Robert, V., J.-F. Molez and J.-F. Trape. 1996. Gametocytes, chloroquine pressure, and the relative parasite survival advantage of resistant strains of falciparum malaria in West Africa. Am. J. Trop. Med. Hyg. 55:3550-3551. [DOI] [PubMed] [Google Scholar]
- 26.Rosario, V. E., R. Hall, D. Walliker, and G. H. Beale. 1978. Persistence of drug-resistant malaria parasites. Lancet i:185-187. [DOI] [PubMed]
- 27.Ryckaert, J., G. Ciccotti, and H. J. C. Berendsen. 1977. Numerical integration of the Cartesian equations of motion of a system with constraints; molecular dynamics of n-alkanes. J. Comp. Physiol. 23:327-341. [Google Scholar]
- 28.Saul, A., P. Myler, T. Elliott, and C. Kidson. 1982. Purification of mature schizonts of Plasmodium falciparum on colloidal silica gradients. Bull. W. H. O. 60:755-759. [PMC free article] [PubMed] [Google Scholar]
- 29.Schwartz, D. C., and C. R. Cantor. 1984. Separation of yeast chromosome-sized DNAs by pulsed field gradient gel electrophoresis. Cell 37:67-75. [DOI] [PubMed] [Google Scholar]
- 30.Scott, W. R. P., H. P. Hulnenberger, I. G. Tironi, A. E. Mark, S. R. Billeter, J. Fennen, A. E. Torda, T. Huber, P Kru1ger, and F. van Gunsteren. 1999. The GROMOS biomolecular simulation program package. J. Physiol. Chem. 103:3596-3607. [Google Scholar]
- 31.Shinondo, C. J., H. N. Lanners, R. C. Lowrie, Jr., and M. F. Wiser. 1994. Effect of pyrimethamine resistance on sporogony in a Plasmodium berghei/Anopheles stephensi model. Exp. Parasitol. 78:194-202. [DOI] [PubMed] [Google Scholar]
- 32.Srivastava, I. K., H. Rottenberg, and A. B. Vaidya. 1997. Atovaquone, a broad-spectrum antiparasitic drug, collapses mitochondria membrane potential in a malaria parasite. J. Biol. Chem. 272:3961-3966. [DOI] [PubMed] [Google Scholar]
- 33.Thanh, N. V., A. F. Cowman, D. Hipgrave, T. B. Kim, B. Q. Phuc, L. D. Cong, and B. A. Biggs. 2001. Assessment of susceptibility of Plasmodium falciparum to chloroquine, quinine, mefloquine, sulfadoxine-pyrimethamine and artemisinin in southern Vietnam. Trans. R. Soc. Trop. Med. Hyg. 95:513-517. [DOI] [PubMed] [Google Scholar]
- 34.Trager, W., and J. B. Jensen. 1976. Human malaria parasites in continuous culture. Science 193:673-675. [DOI] [PubMed] [Google Scholar]
- 35.Wang, J., S. Lu, R. Chen, and L. Wang. 1998. Relative fitness of three organophosphate-resistant strains of Culex pipiens pallens (Diptera: Culicidae). J. Med. Entomol. 35:716-719. [DOI] [PubMed] [Google Scholar]
- 36.Wernsdorfer, W. H., B. Landgraf, G. Wiedermann, and H. Kollaritsch. 1995. Chloroquine resistance of Plasmodium falciparum: a biological advantage? Trans. R. Soc. Trop. Med. Hyg. 89:90-91. [DOI] [PubMed] [Google Scholar]
- 37.White, N. J. 1988. Preventing antimalarial drug resistance through combinations. Drug Resist. Update 1:3-9. [DOI] [PubMed] [Google Scholar]
- 38.Zengel, J. M., R. Young, R. R. Dennis, and M. Nomura. 1977. Role of ribosomal protein S12 in peptide chain elongation: analysis of pleiotropic, streptomycine-resistant mutants of Escherichia coli. J. Bacteriol. 129:1320-1329. [DOI] [PMC free article] [PubMed] [Google Scholar]