Abstract
Using a solvent-referenced energy calculation, a 16-residue peptide with alanine side chains folded into predominantly alpha-helical conformations during constant temperature (274 K) simulations. From different initial conformations, helical conformations were reached and the multiple energy minima did not become a serious problem. Under the same conditions, the simulation did not indiscriminately fold a sequence such as polyglycine into stable helices. Interesting observations from the simulations relate to the folding mechanism. The electrostatic interactions between the successive amides favored extended conformations (or beta strands) and caused energy barriers to helix folding. beta-bends were observed as intermediates during helix nucleation. The helix propagation toward the C-terminus seemed faster than that toward the N-terminus. In helical conformations, hydrogen bond oscillation between the i,i+ 4 and the i,i+3 patterns was observed. The i,i+3 hydrogen bonds occurred more frequently during helix propagation and deformation near both ends of the helical segment.
Full text
PDF
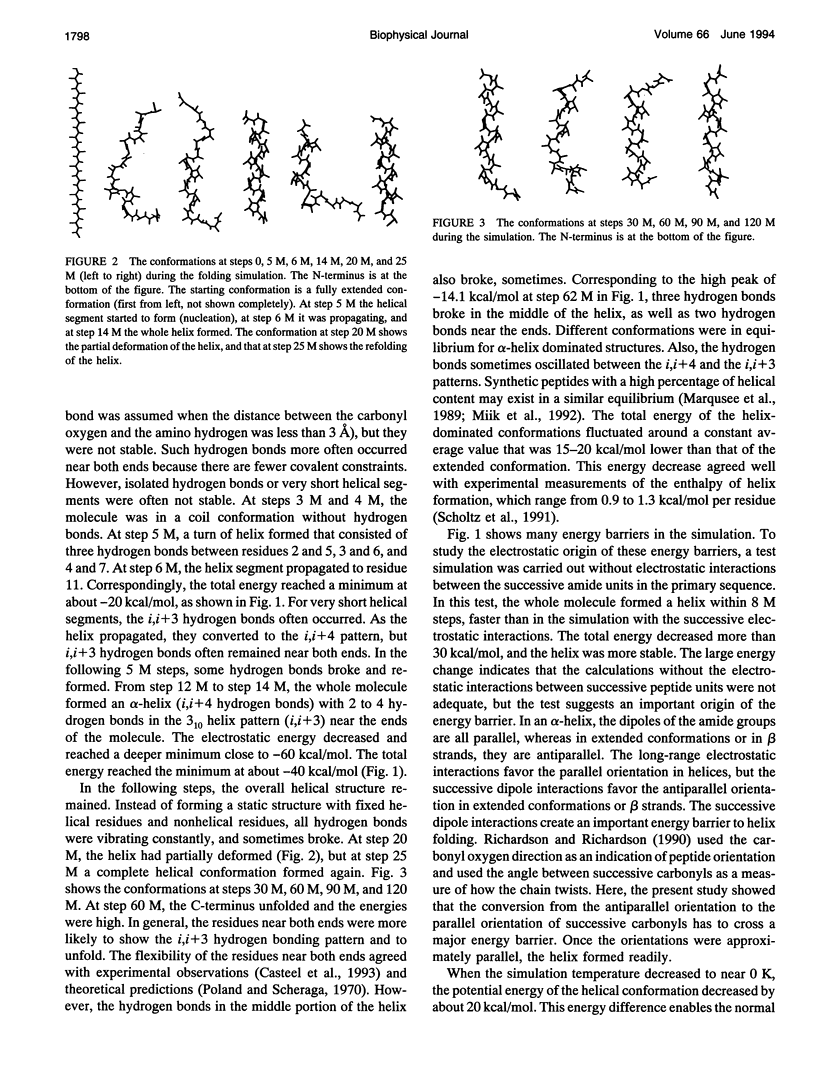
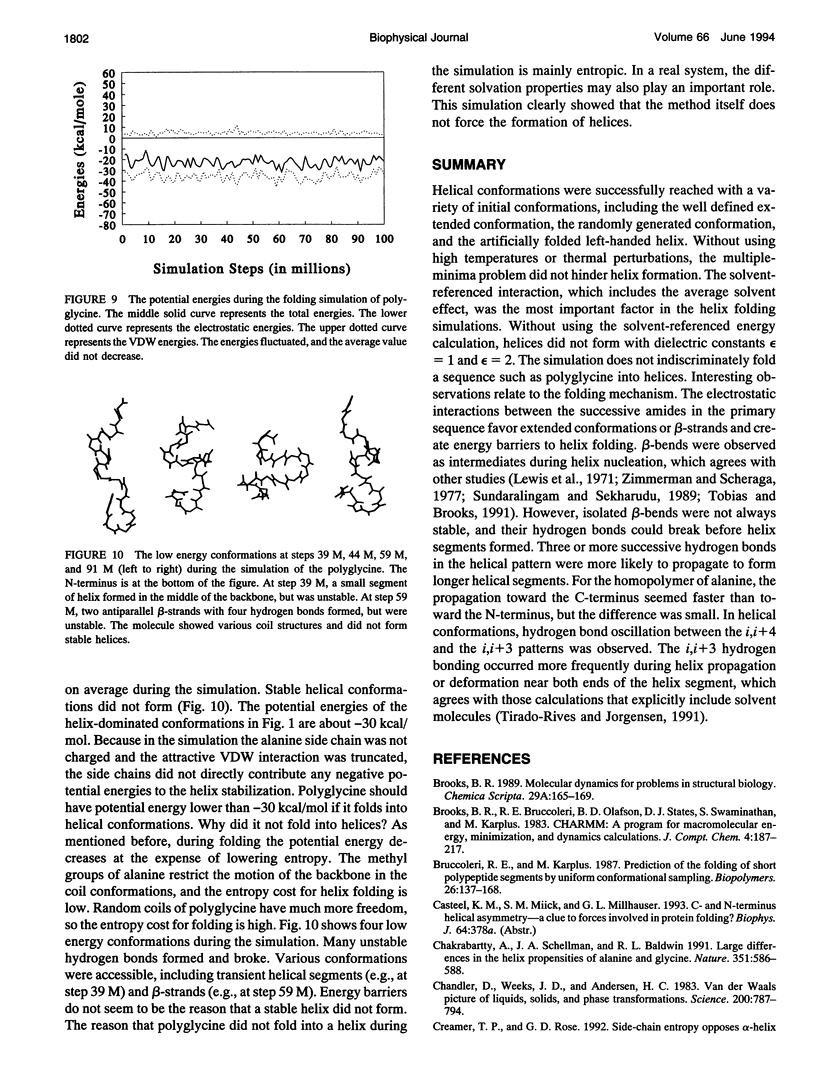
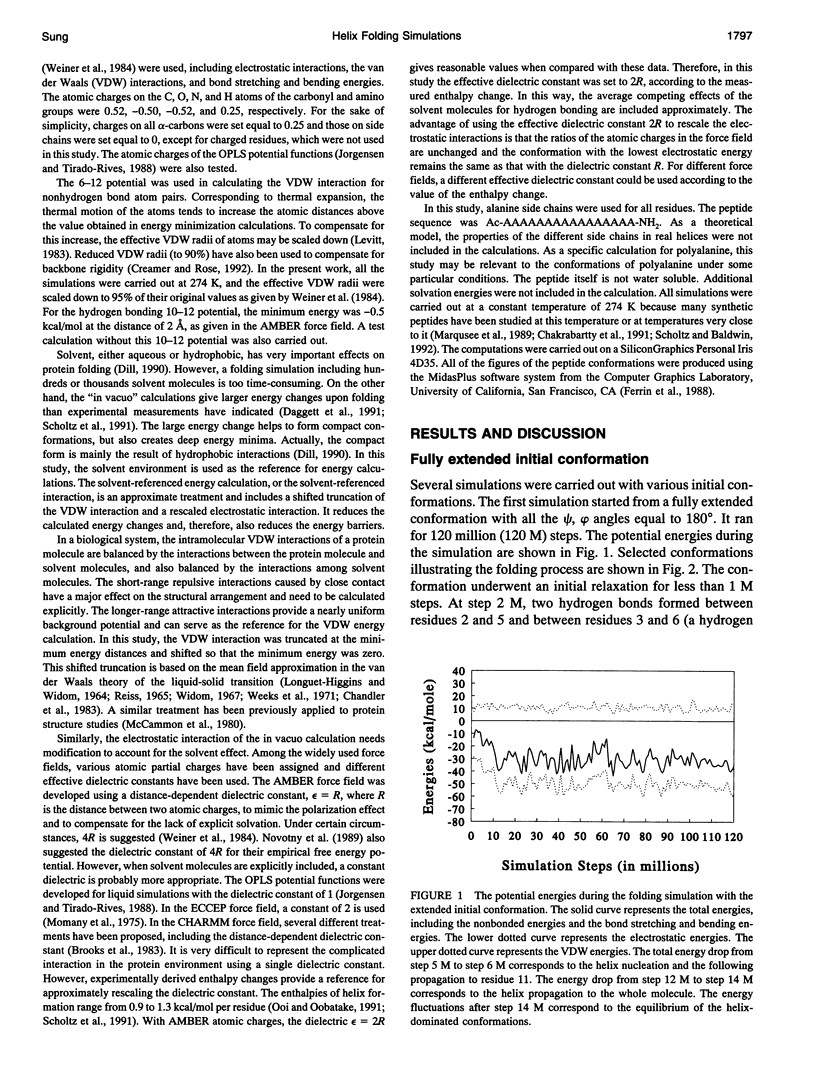

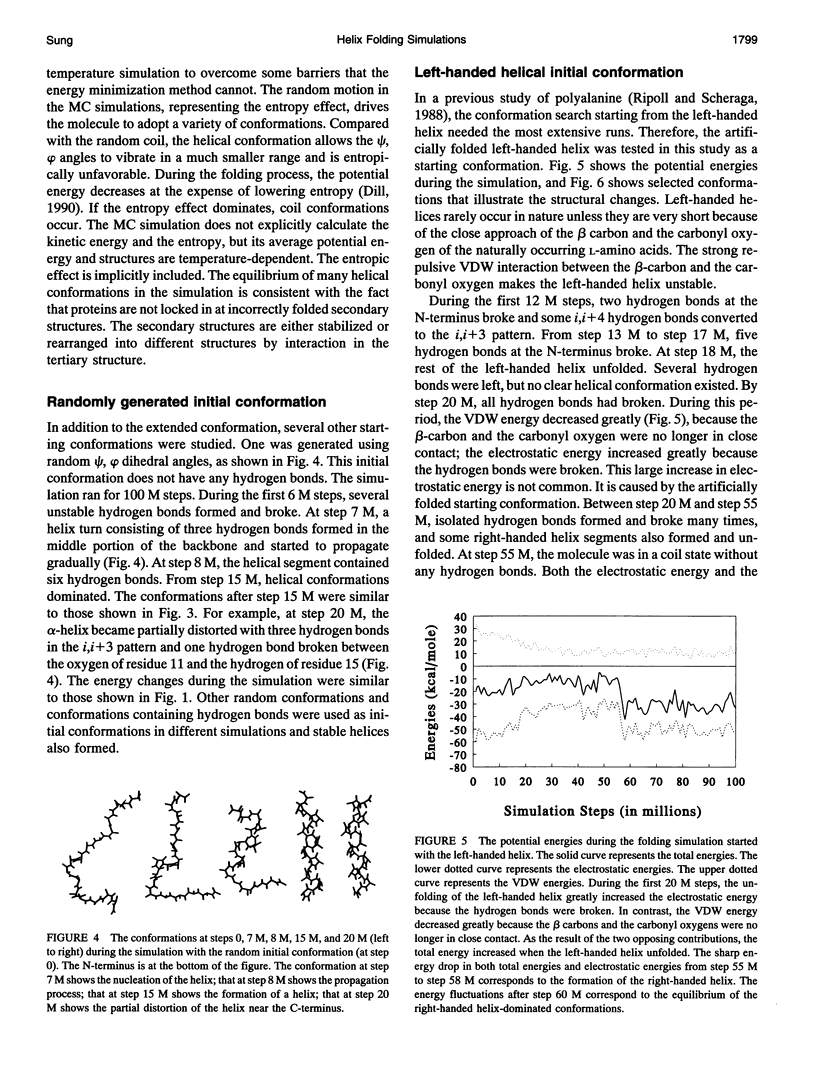
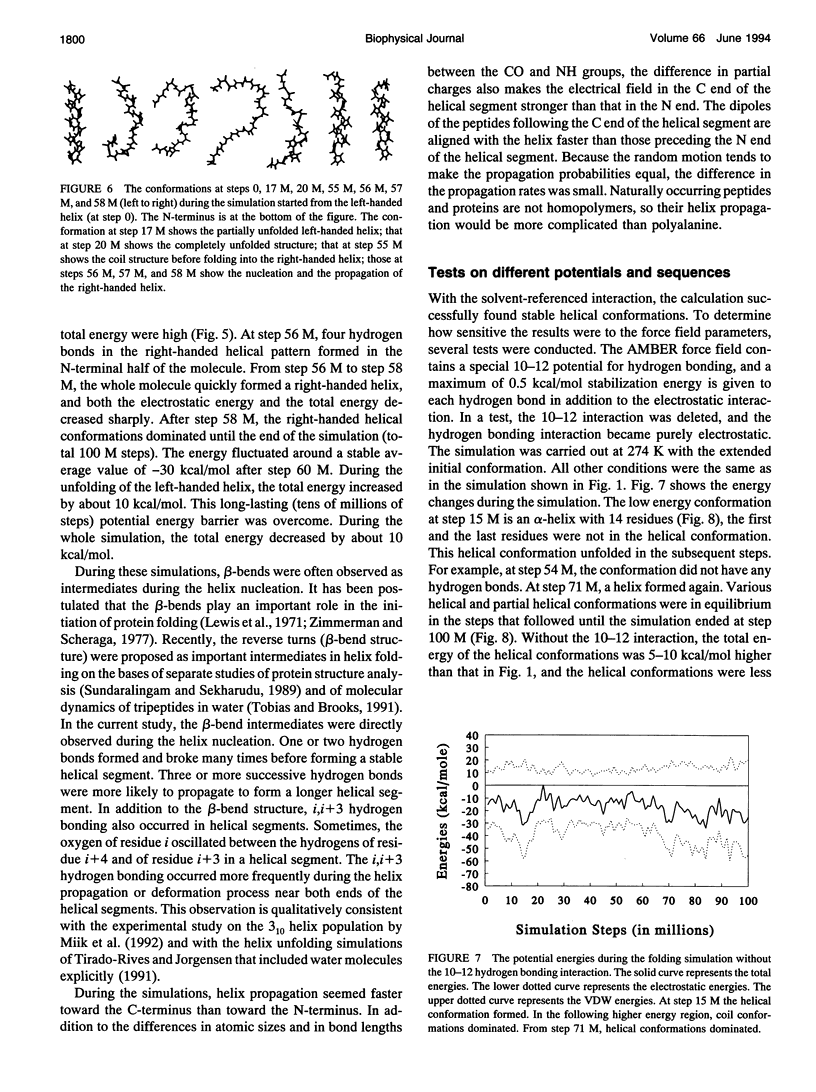



Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bruccoleri R. E., Karplus M. Prediction of the folding of short polypeptide segments by uniform conformational sampling. Biopolymers. 1987 Jan;26(1):137–168. doi: 10.1002/bip.360260114. [DOI] [PubMed] [Google Scholar]
- Chakrabartty A., Schellman J. A., Baldwin R. L. Large differences in the helix propensities of alanine and glycine. Nature. 1991 Jun 13;351(6327):586–588. doi: 10.1038/351586a0. [DOI] [PubMed] [Google Scholar]
- Chandler D., Weeks J. D., Andersen H. C. Van der waals picture of liquids, solids, and phase transformations. Science. 1983 May 20;220(4599):787–794. doi: 10.1126/science.220.4599.787. [DOI] [PubMed] [Google Scholar]
- Creamer T. P., Rose G. D. Side-chain entropy opposes alpha-helix formation but rationalizes experimentally determined helix-forming propensities. Proc Natl Acad Sci U S A. 1992 Jul 1;89(13):5937–5941. doi: 10.1073/pnas.89.13.5937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Daggett V., Kollman P. A., Kuntz I. D. A molecular dynamics simulation of polyalanine: an analysis of equilibrium motions and helix-coil transitions. Biopolymers. 1991 Aug;31(9):1115–1134. doi: 10.1002/bip.360310911. [DOI] [PubMed] [Google Scholar]
- Daggett V., Levitt M. Molecular dynamics simulations of helix denaturation. J Mol Biol. 1992 Feb 20;223(4):1121–1138. doi: 10.1016/0022-2836(92)90264-k. [DOI] [PubMed] [Google Scholar]
- Dill K. A. Dominant forces in protein folding. Biochemistry. 1990 Aug 7;29(31):7133–7155. doi: 10.1021/bi00483a001. [DOI] [PubMed] [Google Scholar]
- Kirkpatrick S., Gelatt C. D., Jr, Vecchi M. P. Optimization by simulated annealing. Science. 1983 May 13;220(4598):671–680. doi: 10.1126/science.220.4598.671. [DOI] [PubMed] [Google Scholar]
- Levitt M. Molecular dynamics of native protein. I. Computer simulation of trajectories. J Mol Biol. 1983 Aug 15;168(3):595–617. doi: 10.1016/s0022-2836(83)80304-0. [DOI] [PubMed] [Google Scholar]
- Lewis P. N., Momany F. A., Scheraga H. A. Folding of polypeptide chains in proteins: a proposed mechanism for folding. Proc Natl Acad Sci U S A. 1971 Sep;68(9):2293–2297. doi: 10.1073/pnas.68.9.2293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Z., Scheraga H. A. Monte Carlo-minimization approach to the multiple-minima problem in protein folding. Proc Natl Acad Sci U S A. 1987 Oct;84(19):6611–6615. doi: 10.1073/pnas.84.19.6611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marqusee S., Robbins V. H., Baldwin R. L. Unusually stable helix formation in short alanine-based peptides. Proc Natl Acad Sci U S A. 1989 Jul;86(14):5286–5290. doi: 10.1073/pnas.86.14.5286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miick S. M., Martinez G. V., Fiori W. R., Todd A. P., Millhauser G. L. Short alanine-based peptides may form 3(10)-helices and not alpha-helices in aqueous solution. Nature. 1992 Oct 15;359(6396):653–655. doi: 10.1038/359653a0. [DOI] [PubMed] [Google Scholar]
- Novotny J., Bruccoleri R. E., Saul F. A. On the attribution of binding energy in antigen-antibody complexes McPC 603, D1.3, and HyHEL-5. Biochemistry. 1989 May 30;28(11):4735–4749. doi: 10.1021/bi00437a034. [DOI] [PubMed] [Google Scholar]
- Némethy G., Scheraga H. A. Protein folding. Q Rev Biophys. 1977 Aug;10(3):239–252. doi: 10.1017/s0033583500002936. [DOI] [PubMed] [Google Scholar]
- Ooi T., Oobatake M. Prediction of the thermodynamics of protein unfolding: the helix-coil transition of poly(L-alanine). Proc Natl Acad Sci U S A. 1991 Apr 1;88(7):2859–2863. doi: 10.1073/pnas.88.7.2859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ripoll D. R., Scheraga H. A. On the multiple-minima problem in the conformational analysis of polypeptides. II. An electrostatically driven Monte Carlo method--tests on poly(L-alanine). Biopolymers. 1988 Aug;27(8):1283–1303. doi: 10.1002/bip.360270808. [DOI] [PubMed] [Google Scholar]
- Scholtz J. M., Baldwin R. L. The mechanism of alpha-helix formation by peptides. Annu Rev Biophys Biomol Struct. 1992;21:95–118. doi: 10.1146/annurev.bb.21.060192.000523. [DOI] [PubMed] [Google Scholar]
- Scholtz J. M., Marqusee S., Baldwin R. L., York E. J., Stewart J. M., Santoro M., Bolen D. W. Calorimetric determination of the enthalpy change for the alpha-helix to coil transition of an alanine peptide in water. Proc Natl Acad Sci U S A. 1991 Apr 1;88(7):2854–2858. doi: 10.1073/pnas.88.7.2854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sikorski A., Skolnick J. Monte Carlo simulation of equilibrium globular protein folding: alpha-helical bundles with long loops. Proc Natl Acad Sci U S A. 1989 Apr;86(8):2668–2672. doi: 10.1073/pnas.86.8.2668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skolnick J., Kolinski A., Yaris R. Dynamic Monte Carlo study of the folding of a six-stranded Greek key globular protein. Proc Natl Acad Sci U S A. 1989 Feb;86(4):1229–1233. doi: 10.1073/pnas.86.4.1229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skolnick J., Kolinski A., Yaris R. Monte Carlo simulations of the folding of beta-barrel globular proteins. Proc Natl Acad Sci U S A. 1988 Jul;85(14):5057–5061. doi: 10.1073/pnas.85.14.5057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sundaralingam M., Sekharudu Y. C. Water-inserted alpha-helical segments implicate reverse turns as folding intermediates. Science. 1989 Jun 16;244(4910):1333–1337. doi: 10.1126/science.2734612. [DOI] [PubMed] [Google Scholar]
- Tirado-Rives J., Jorgensen W. L. Molecular dynamics simulations of the unfolding of an alpha-helical analogue of ribonuclease A S-peptide in water. Biochemistry. 1991 Apr 23;30(16):3864–3871. doi: 10.1021/bi00230a009. [DOI] [PubMed] [Google Scholar]
- Tobias D. J., Brooks C. L., 3rd Thermodynamics and mechanism of alpha helix initiation in alanine and valine peptides. Biochemistry. 1991 Jun 18;30(24):6059–6070. doi: 10.1021/bi00238a033. [DOI] [PubMed] [Google Scholar]
- Widom B. Intermolecular Forces and the Nature of the Liquid State: Liquids reflect in their bulk properties the attractions and repulsions of their constituent molecules. Science. 1967 Jul 28;157(3787):375–382. doi: 10.1126/science.157.3787.375. [DOI] [PubMed] [Google Scholar]
- Wilson S. R., Cui W. L. Applications of simulated annealing to peptides. Biopolymers. 1990 Jan;29(1):225–235. doi: 10.1002/bip.360290127. [DOI] [PubMed] [Google Scholar]
- Zimmerman S. S., Scheraga H. A. Local interactions in bends of proteins. Proc Natl Acad Sci U S A. 1977 Oct;74(10):4126–4129. doi: 10.1073/pnas.74.10.4126. [DOI] [PMC free article] [PubMed] [Google Scholar]



