Abstract
Carbapenem resistance mediated by acquired carbapenemase genes has been increasingly reported, particularly for clinical isolates of Pseudomonas aeruginosa and Acinetobacter spp. Of 1,234 nonduplicate isolates of carbapenem-resistant Pseudomonas spp. and Acinetobacter spp. isolated at a tertiary-care hospital in Seoul, Korea, 211 (17%) were positive for metallo-β-lactamase (MBL). Of these, 204 (96%) had either the blaIMP-1 or blaVIM-2 allele. In addition, seven Acinetobacter baumannii isolates were found to have a novel MBL gene, which was designated blaSIM-1. The SIM-1 protein has a pI of 7.2, is a new member of subclass B1, and exhibits 64 to 69% identity with the IMP-type MBLs, which are its closest relatives. All SIM-1-producing isolates exhibited relatively low imipenem and meropenem MICs (8 to 16 μg/ml) and had a multidrug resistance phenotype. Expression of the cloned blaSIM-1 gene in Escherichia coli revealed that the encoded enzyme is capable of hydrolyzing a broad array of β-lactams, including penicillins, narrow- to expanded-spectrum cephalosporins, and carbapenems. The blaSIM-1 gene was carried on a gene cassette inserted into a class 1 integron, which included three additional cassettes (arr-3, catB3, and aadA1). The strains were isolated from sputum and urine specimens from patients with pneumonia and urinary tract infections, respectively. All patients had various underlying diseases. Pulsed-field gel electrophoresis of SmaI-digested genomic DNAs showed that the strains belonged to two different clonal lineages, indicating that horizontal transfer of this gene had occurred and suggesting the possibility of further spread of resistance in the future.
Gram-negative bacilli have a propensity to develop and acquire resistance to multiple antimicrobials. A significant increase in the prevalence of multidrug-resistant gram-negative bacilli, even among isolates recovered at admission, was reported (20). Carbapenems have been the most successful β-lactam antibiotics in evading bacterial resistance (14). However, carbapenem resistance, mediated by acquired carbapenemase genes, has been increasingly reported, particularly for clinical isolates of Pseudomonas aeruginosa and Acinetobacter spp. (7). Although some enzymes of molecular classes A and D can hydrolyze carbapenems, metallo-β-lactamases (MBLs) are the most prevalent acquired carbapenemases (5, 14, 18, 24, 27, 28).
Of acquired MBLs, the IMP- and VIM-type enzymes are the most common and exhibit a worldwide distribution (7, 8, 19, 24, 26). Recently, however, two additional types, SPM and GIM, have been reported for P. aeruginosa isolates from Brazil and Germany, respectively (2, 22). In Korea, a prevalence of both IMP-1- and VIM-2-type MBLs has been reported (10, 12). Here we report the detection of a new acquired MBL in clinical isolates of Acinetobacter baumannii from Korea.
MATERIALS AND METHODS
Bacterial strains.
A total of 1,234 nonduplicate imipenem-resistant clinical isolates of Pseudomonas spp. and Acinetobacter spp. isolated in 2003-2004 at a tertiary-care hospital in Seoul, Korea, were investigated in this study. The isolates were identified by conventional methods (21) or by using the ATB 32GN system (bioMerieux, Marcy l'Etoile, France). The phenotypic identification of four randomly selected VIM-2 MBL-producing Pseudomonas putida isolates was confirmed by 16S rRNA sequencing as described previously (15). The Escherichia coli strain XL-1 Blue (Stratagene Inc., La Jolla, Calif.) was used as the host for expression of the cloned blaSIM-1 gene.
Susceptibility testing.
Imipenem susceptibility was tested by a disk diffusion method (17). MICs for the SIM-1-producing A. baumannii isolates were determined by an agar dilution method (16) and interpreted according to the guidelines of the Clinical Laboratory Standards Institute (3). MICs for E. coli XL-1(pBC-SIM1) were determined by a broth microdilution method (16).
β-Lactamase assays.
The Hodge test (9), using MacConkey agar instead of Mueller-Hinton agar, which allows a better performance (unpublished result), was used for screening of carbapenemase production. The imipenem and EDTA-sodium mercaptoacetic acid (SMA) double-disk synergy test (11) and Etest MBL strips (AB BIODISK, Solna, Sweden) were used for screening of MBL production. A spectrophotometric assay was used to measure the MBL activity in crude cell extracts, as described previously (8). Analytical isoelectric focusing of β-lactamase was performed using the nitrocefin chromogenic substrate (8).
DNA analysis and manipulation methodology.
PCR detection of the blaIMP-1 and blaVIM-2 genes was carried out using the IMP1 and VIM2 primers (Table 1) as described previously (29). SIM primers, which were designed after an analysis of the sequence of the novel MBL gene, were used to detect the presence of the allele in the remaining six non-IMP-, non-VIM-type MBL-producing isolates. PCR was performed with 1 μl of heat-extracted DNA template, 20 pmol of each primer, and PreMix (Bioneer, Cheongwon, Korea) containing 1 U of Taq DNA polymerase in a total volume of 20 μl. A Mastercycler instrument (Eppendorf, Hamburg, Germany) was used with the following reaction conditions: 94°C for 5 min, 25 cycles of 94°C for 30 s, 56°C for 30 s, and 72°C for 45 s, and finally, 72°C for 7 min.
TABLE 1.
Primers used for sequencing of the blaSIM-1 gene and for detection of MBL genes
| Target | Primer | Use | Sequence (5′ to 3′) | Positiona | Reference |
|---|---|---|---|---|---|
| 5′-CS | INT1-F | Detection and sequencing | GGCATCCAAGCAGCAAG | 1256-1272 | 13 |
| 3′-CS | INT2-R | Detection and sequencing | AAGCAGACTTGACCTGA | 4336-4620 | |
| intI1 | INT7-R | Sequencing | GTTCTTCTACGGCAAGGTGC | 956-937 | 12 |
| arr-3 | ARR3-F | Sequencing | GGTGACTTGCTAACCACAG | 91-109 | 25 |
| ARR3-R | Sequencing | ACAGTGACATAGCAAGTTCAG | 211-191 | ||
| aadA1 | AADA1-F | Sequencing | TGATTTGCTGGTTACGGTGAC | 144-164 | 12 |
| AADA1-R | Sequencing | CACTACATTTCGCTCATCG | 561-543 | ||
| blaIMP-1 | IMP1-F | Detection | CATGGTTTGGTGGTTCTTGT | 601-620 | 29 |
| IMP1-R | Detection | ATAATTTGGCGGACTTTGGC | 1048-1029 | ||
| blaVIM-2 | VIM2-F | Detection | ATGTTCAAACTTTTGAGTAAG | 1-21 | 19 |
| VIM2-R | Detection | CTACTCAACGACTGAGCG | 801-784 | ||
| blaSIM-1 | SIM1-F | Detection and sequencing | TACAAGGGATTCGGCATCG | 127-145 | This study |
| SIM1-R | Detection and sequencing | TAATGGCCTGTTCCCATGTG | 697-678 | ||
| blaSIM-1 | SIM1-Exp/f | Cloning and expression | GGTCTAGAAGGAGAGTTAAAAATGAGAACTTTATTGATTT | 1-19 | This study |
| SIM1-Exp/r | Cloning and expression | CCGGATCCTTAATTAATGAGCGGCGGTTTTG | 719-741 | ||
| 16S rRNA | 8F | Sequencing | AGAGTTTGATCCTGGCTCAG | 8-27 | 15 |
| 1541R | Sequencing | AAGGAGGTGATCCAGCCGCA | 1541-1522 |
Position numbers correspond to the nucleotides of the coding sequences, except for the integron, for which position number 1 is the first nucleotide of the 5′ conserved segment (AY046276). For the SIM1-Exp primers, the regions annealing to the blaSIM-1 coding sequence are underlined.
The amplicons generated with primers INT1-F and INT2-R (13) were used to analyze the sequence of the new MBL gene and other gene cassettes. The reactions were carried out in a total volume of 50 μl, with 5 μl of heat-extracted template DNA, 20 pmol of each primer, and 3 U of LA Taq (Takara, Shiga, Japan). The cycling conditions were as follows: 94°C for 12 min and then 35 cycles of 94°C for 1 min, 56°C for 1 min, and 72°C for 5 min. The extension time was increased by 5 s for each cycle. Nucleotide sequences were determined for both strands of DNA by the dideoxy chain-termination method using an ABI 3700 autosequencer (Perkin-Elmer, Foster City, Calif.). Three independently generated amplicons were used to determine the nucleotide sequences. Sequence analysis and comparisons were performed using programs available at the NCBI (http://www.ncbi.nlm.nih.gov/) and ExPASy (http://www.expasy.org/) servers.
Southern blot hybridization was performed on pulsed-field gel electrophoresis (PFGE)-separated DNA after its transfer to a nylon membrane (Bio-Rad, Hercules, Calif.), using a digoxigenin-labeled blaSIM-1 probe which was prepared according to the reagent manufacturer's instructions (Roche Diagnostics, Mannheim, Germany).
Plasmid pBC-SIM1 was constructed by cloning a PCR amplicon containing the blaSIM-1 gene which was generated with primers SIM1-Exp/f (adding an XbaI restriction site and a ribosome binding site upstream of the blaSIM-1 start codon) and SIM-1-Exp/r (adding a BamHI restriction site downstream of the blaSIM-1 stop codon) (Table 1). PCR was performed in a 50-μl volume, using 3.5 U of the Expand High-Fidelity PCR system (Roche Diagnostics) in the buffer supplied by the manufacturer, with 50 pmol of each primer and 10 ng of A. baumannii YMC 03/9/T104 genomic DNA, under the following cycling conditions: 95°C for 3 min, 30 cycles of 95°C for 1 min, 55°C for 1 min, and 72°C for 2 min, and finally, 72°C for 7 min. The XbaI- and BamHI-digested amplicon was ligated to the pBC-SK(+) plasmid vector (Stratagene), which had been digested with the same enzymes. The authenticity of the cloned blaSIM-1 gene carried by pBC-SIM1 was confirmed by sequencing.
Gene transfer experiments.
Imipenem resistance transfer was tested by the agar mating method (1), using rifampin-resistant P. aeruginosa PAO 4089 and azide-resistant Escherichia coli J53 as recipients and ceftazidime (4 μg/ml) plus rifampin (200 μg/ml) or azide (100 μg/ml) for the selection of transconjugants. Ceftazidime was used for the selection of transconjugants instead of a carbapenem because the SIM-1-producing isolates were inhibited by a relatively low concentration of imipenem.
Genotyping.
PFGE of SmaI-restricted genomic DNAs of the SIM-1-producing A. baumannii isolates was performed using the CHEF-DR II system (Bio-Rad) according to the manufacturer's protocols. The banding patterns were analyzed with UVIband/Map software (UVItech Ltd., Cambridge, United Kingdom) to generate a dendrogram based on the unweighted-pair group method using arithmetic averages from the Dice coefficients.
Nucleotide sequence accession number.
The nucleotide sequence data reported in this paper are listed in the GenBank database under accession number AY887066.
RESULTS AND DISCUSSION
MBL production in carbapenem-resistant Pseudomonas spp. and Acinetobacter spp.
Of 1,234 nonduplicate imipenem-resistant clinical strains isolated in 2003-2004 at a tertiary-care hospital, 211 (17%) were positive for carbapenemase and MBL production by the Hodge test and the imipenem and EDTA-SMA double-disk synergy test, respectively (Table 2). PCR analysis of the MBL producers revealed the presence of a blaVIM-2 allele in 157 (74%) isolates and of a blaIMP-1 allele in 47 (22%) isolates, but 7 (4%) A. baumannii isolates yielded negative results for both genes. MBL activities were detectable in crude extracts of these isolates by a spectrophotometric assay (data not shown). With these isolates, the imipenem and EDTA-SMA double-disk synergy test was clearly positive only when the two disks were placed 15 mm (edge-to-edge) apart from each other, due to a relatively large imipenem inhibition zone, while the Etest for MBL was not informative due to the relatively low imipenem MICs for these isolates (8 to 16 μg/ml) (Table 3).
TABLE 2.
MBL-producing Pseudomonas spp. and Acinetobacter spp. detected at a tertiary-care hospital in 2003-2004
| Species (no. of isolates) | No. of imipenem- resistant isolates | No. (%) of isolates positive for:
|
|||
|---|---|---|---|---|---|
| MBL screeninga | blaIMP-1 | blaVIM-2 | blaSIM-1 | ||
| P. aeruginosa (5,970) | 644 | 34 (5) | 2 (6) | 32 (94) | 0 |
| P. putida/P. fluorescens (211) | 45b | 42 (93) | 0 | 42 (100) | 0 |
| Acinetobacter spp. (3,294) | 545 | 135 (25) | 45 (33) | 83 (61) | 7 (6) |
| Total (9,475) | 1,234 | 211 (17) | 47 (22) | 157 (74) | 7 (4) |
By the Hodge test and the imipenem and EDTA-SMA double-disk synergy test.
Phenotypic differentiation of P. fluorescens and P. putida was performed for all imipenem-resistant isolates. The phenotypic identification of P. putida was confirmed by sequencing 16S rRNA from four randomly selected isolates.
TABLE 3.
Clinical features of patients and characteristics of A. baumannii isolates with blaSIM-1 cassette-carrying class 1 integrons
| Patient no. | Isolatea | Age (yrs)/sex | Underlying disease | Hospital ward | Days since admissionb | Previous antimicrobial therapy (days)c | Specimen/ diagnosisc | MIC (μg/ml)c
|
PFGE pattern | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SAM | PIP | CTX | CAZ | FEP | ATM | IPM | MEM | GEN | TOB | AMK | CIP | |||||||||
| 1 | 03/9/T104 | 9/F | Cerebral palsy | Intensive care unit | 30 | MEM (12), ATM (9), FEP (5) | Sputum/ pneumonia | 16 | 64 | >256 | >256 | 64 | 128 | 8 | 16 | 16 | 16 | 128 | 0.5 | A1 |
| 2 | 04/5/U2266 | 66/F | Hypothyroidism, osteoporosis, hypertension | Outpatient | NA | ROX (10) | Urine/UTI | 16 | 64 | >256 | >256 | 64 | 128 | 8 | 16 | 16 | 16 | 128 | 0.5 | A1 |
| 3 | 04/8/U1575 | 64/M | Herniated lumbar disc | Internal medicine | 9 | FMX (7) | Urine/UTI | 16 | 32 | >256 | >256 | 64 | 128 | 16 | 16 | 16 | 16 | 128 | 0.5 | A2 |
| 4 | 04/11/U1817 | 45/M | Intracranial hemorrhage | Internal medicine | 11 | ISP (8), CTX (7), CFM (4) | Urine/UTI | 16 | 32 | >256 | >256 | 64 | 128 | 8 | 16 | 16 | 16 | 128 | 0.5 | A2 |
| 5 | 04/12/U575 | 13/M | Chronic myelogenous leukemia | Pediatrics | 4 | None | Urine/UTI | 16 | 128 | >256 | >256 | 128 | 128 | 16 | 16 | 16 | 16 | 128 | 0.5 | B1 |
| 6 | 04/12/U837 | 5/M | Acute lymphocytic leukemia | Pediatrics | 3 | AMC (4) | Urine/UTI | 16 | 256 | >256 | >256 | 64 | 128 | 8 | 16 | 16 | 16 | 128 | 0.5 | B1 |
| 7 | 04/12/U1398 | 3/M | Acute myelogenous leukemia | Pediatrics | 3 | SXT (3), AMC (2), ISP (2) | Urine/UTI | 16 | 128 | >256 | >256 | 128 | 128 | 8 | 16 | 16 | 16 | 64 | 0.5 | B2 |
The first two numbers, separated by a slash, indicate the year and month of isolation, respectively.
Days between admission and the isolation of SIM-1-producing A. baumannii.
Abbreviations: SAM, ampicillin-sulbactam; PIP, piperacillin; CTX, cefotaxime; CAZ, ceftazidime; FEP, cefepime; ATM, aztreonam; IPM, imipenem; MEM, meropenem; GEN, gentamicin; TOB, tobramycin; AMK, amikacin; CIP, ciprofloxacin; ROX, roxithromycin; FMX, flomoxef; ISP, isepamycin; CFM, cefixime; AMC, amoxicillin-clavulanate; SXT, trimethoprim-sulfamethoxazole; NA, not applicable; UTI, urinary tract infection.
Identification of a new MBL determinant.
The new MBL determinant and the integron content of the first non-IMP- and non-VIM-type MBL-producing isolate (YMC 03/9/T104) were investigated by sequencing, using PCR products obtained with the INT1-F and INT2-R primers (13). This analysis revealed the presence of two class 1 integrons, with one containing two gene cassettes (arr-3 and aacA4) and the other containing four gene cassettes (Fig. 1). The first cassette of the latter integron carried a new MBL gene, which we designated blaSIM-1 (for Seoul imipenemase).
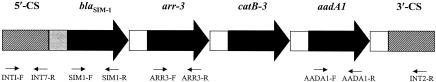
FIG. 1.
Structure of the variable region of the blaSIM-1-containing class 1 integron from isolate YMC 03/9/T104. Large arrows indicate the resistance genes carried by the gene cassettes and their transcriptional direction. The gray box indicates the attI site, open boxes indicate the attC sites (59-base elements) of the gene cassettes, and hatched boxes indicate the partially sequenced 5′ and 3′ conserved sequences. The locations of primers (described in Table 1) used to map the integron and determine its sequence are also shown (small arrows).
The SIM-1 enzyme is a new subclass B1 MBL (4) whose closest relatives are the IMP-type enzymes (69% identity with IMP-12 and 64% identity with IMP-9), while the other MBLs are more divergent (Fig. 2 and 3). The theoretical molecular weight and pI of the mature SIM-1 protein are 25,439 and 7.28, respectively, considering the presence of an 18-amino-acid leader peptide (http://www.cbs.dtu.dk/services/SignalP/). The attC (59-base element) recombination site of the blaSIM-1 cassette was 88 bp long and not related to those of known blaIMP cassettes but was very similar (90% identity) to that of a cassette (ypar13) of the Pseudomonas alcaligenes In55044 superintegron (23), suggesting that the source of the blaSIM-1 cassette could be a similar element. It was interesting that the other cassettes carried arr-3, a rifampin ADP ribosyltransferase gene, and catB3, a chloramphenicol acetyltransferase gene, which are rarely found in MBL-encoding integrons (6).
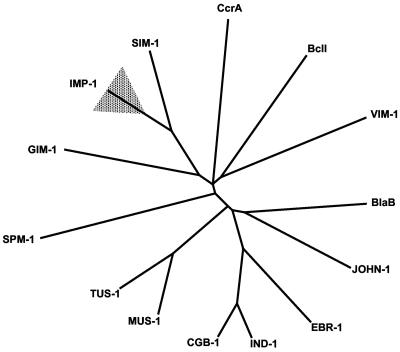
FIG. 2.
Unrooted tree showing the relationships between SIM-1 and other subclass B1 MBLs. The shaded triangle indicates the sequence variability observed within the IMP lineage. Accession numbers for the sequences are as follows: IMP-1, AAB30289; CcrA, P25910; BcII, P04190; VIM-1, CAB46686; BlaB, CAA65601; JOHN-1, AAK38324; EBR-1, AAN32638; IND-1, AAD20273; CGB-1, AAL55263; MUS-1, AAN63647; TUS-1, AAN63648; SPM-1, CAD37801; and GIM-1, CAF05908.
FIG. 3.
Amino acid sequence comparison between SIM-1 and representatives of the other four types of acquired MBLs. Residues conserved in all proteins are shown in bold. The residues known to be involved in metal binding are indicated by asterisks. Numbering is according to the updated BBL scheme (4). Accession numbers are the same as those in the legend to Fig. 2.
Using the SIM primers, the blaSIM-1 gene was also detected in the remaining six isolates. The structures of the blaSIM-1-containing class 1 integrons were apparently identical in these isolates, according to the results of PCR mapping using the INT1-F and INT-2R primers in combination with the SIM1-R and SIM1-F primers, respectively, and of partial sequencing of the amplicons (data not shown).
Repeated attempts failed to transfer ceftazidime resistance from each of the seven isolates to either P. aeruginosa or E. coli recipient cells by conjugation. In a Southern blot analysis of SmaI-restricted and PFGE-separated genomic DNAs, a blaSIM-1 probe recognized a band of approximately 300 kb in all seven isolates (data not shown). This suggested that the blaSIM-1 gene was located on the chromosome. Further studies are required to ascertain the chromosomal or plasmid location of the blaSIM-1 gene.
Functional characterization of SIM-1.
The functional properties of SIM-1 were investigated by expression of the enzyme in E. coli XL-1 Blue and testing of the susceptibility of this strain to several β-lactams. The E. coli XL-1(pBC-SIM1) strain produced an EDTA-inhibitable carbapenemase (Table 4), and analytical isoelectric focusing of a crude extract revealed the presence of a β-lactamase band with a pI of 7.2 (consistent with the theoretical value calculated for the mature SIM-1 protein [see above]). Compared to the host strain, XL-1(pBC-SIM1) exhibited decreased susceptibility to a broad array of β-lactams, including penicillins, narrow- to expanded-spectrum cephalosporins, and carbapenems (Table 4), indicating that SIM-1 has a broad overall substrate specificity. Only the MICs of aztreonam and piperacillin were unchanged, suggesting that SIM-1 is not active against these compounds.
TABLE 4.
MICs of β-lactams for E. coli XL-1 (pBC-SIM1), expressing the SIM-1 enzyme, and for the E. coli host containing only the cloning vector pBC-SK
| Antibiotic | MIC (μg/ml)
|
|
|---|---|---|
| XL-1(pBC-SIM1) | XL-1(pBC-SK) | |
| Ampicillin | 64 | 2 |
| Ticarcillin | >128 | 4 |
| Piperacillin | 1 | 1 |
| Cephalothin | >128 | 4 |
| Cefoxitin | >128 | 2 |
| Cefuroxime | >128 | 4 |
| Cefotaxime | 32 | ≤0.06 |
| Ceftriaxone | 32 | ≤0.06 |
| Ceftazidime | 32 | 0.12 |
| Cefepime | 2 | ≤0.06 |
| Imipenem | 1 | 0.12 |
| Meropenem | 2 | ≤0.06 |
| Panipenem | 2 | 0.12 |
| Aztreonam | 0.25 | 0.25 |
| Carbapenemase sp act (nmol imipenem/min/mg protein) | 2,800 ± 400a | <1 |
The carbapenemase activity was reduced by 80% after incubation of the extract with 5 mM EDTA for 20 min at 25°C.
Clonal relatedness of SIM-1-producing isolates and clinical features of patients.
PFGE of SmaI-restricted genomic DNAs showed that the SIM-1-producing isolates belonged to two quite different clusters (Fig. 4). The first SIM-1-producing A. baumannii isolate was from a sputum specimen from a patient with pneumonia, while the remaining six were from urines from patients with urinary tract infections. All patients had underlying diseases, and all but one were inpatients. The single outpatient, however, had a long history of ambulatory treatment due to several comorbidities. The six patients had been in the hospital for 3 to 30 days before the isolation of a SIM-1-producing isolate (Table 3). Only the first patient had received meropenem therapy for 12 days before the isolation of the SIM-1-producing isolate. The other patients had received other antimicrobial agents, except for one pediatric patient, who did not receive any antimicrobial agents during the 4 days of hospitalization preceding the isolation of the SIM-1-producing isolate. This suggests that the acquisition of SIM-1-producing A. baumannii was not strictly related to the use of carbapenems.
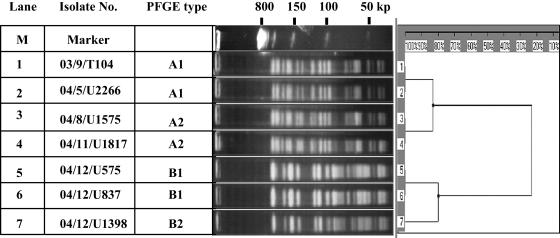
FIG. 4.
PFGE patterns of SmaI-digested genomic DNAs from SIM-1-producing isolates of A. baumannii. The patterns of the first four isolates are quite different from those of the last three isolates.
The first four SIM-1-producing isolates were detected sporadically, from different settings, during a period of 14 months (September 2003 to November 2004), while the last three isolates were detected over a period of 11 days from patients in the same ward, suggesting the occurrence of a small outbreak. Of the two clonal lineages of SIM-1-producing isolates, one was represented by the sporadic isolates, and the other was represented by those involved in the outbreak in the pediatric ward (Table 3; Fig. 4).
All isolates were susceptible to ciprofloxacin but resistant or intermediate to other antimicrobial agents, including expanded-spectrum cephalosporins, ampicillin-sulbactam, and all aminoglycosides tested (Table 3). Levofloxacin was used in one patient, and isepamycin was used in the other patients. All patients eventually recovered from the infection.
In conclusion, a novel acquired MBL gene, blaSIM-1, was detected in seven clinical isolates of A. baumannii from Korea, where VIM-2- and IMP-1-type MBL-producing P. aeruginosa and Acinetobacter spp. have been prevalent. SIM-1 is encoded by a class 1 integron-borne gene cassette and is more closely related to IMP-type enzymes than to other MBLs. The presence of apparently the same integron in isolates belonging to different clonal lineages indicates that horizontal transfer of this gene has occurred and suggests that further spread of the resistance may occur in the future.
Acknowledgments
This work was supported in part by the Brain Korea 21 Project for Medical Sciences, Yonsei University, in 2004.
We thank Myung Sook Kim for collecting the isolates and Yonghee Suh and Chasoon Lee for screening the MBL-producing isolates.
REFERENCES
- 1.Bergquist, P. L. 1988. Incompatibility, p. 37-78. In K. G. Hardy (ed.), Plasmids. IRL Press, Oxford, United Kingdom.
- 2.Castanheira, M., M. A. Toleman, R. N. Jones, F. J. Schmidt, and T. R. Walsh. 2004. Molecular characterization of a β-lactamase gene, blaGIM-1, encoding a new subclass of metallo-β-lactamase. Antimicrob. Agents Chemother. 48:4654-4661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Clinical and Laboratory Standards Institute. 2005. Performance standards for antimicrobial disk susceptibility testing. Fifteenth informational supplement. M100-S15. Clinical and Laboratory Standards Institute, Wayne, Pa.
- 4.Garau, G., I. García-Sáez, C. Bebrone, C. Anne, P. Mercuri, M. Galleni, J.-M. Frère, and O. Dideberg. 2004. Update of the standard numbering scheme for class B β-lactamases. Antimicrob. Agents Chemother. 48:2347-2349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Héritier, C., A. Dubouix, L. Poirel, N. Marty, and P. Nordmann. 2005. A nosocomial outbreak of Acinetobacter baumannii isolates expressing the carbapenem-hydrolysing oxacillinase OXA-58. J. Antimicrob. Chemother. 55:115-118. [DOI] [PubMed] [Google Scholar]
- 6.Houang, E. T. S., Y.-W. Chu, W.-S. Lo, K.-Y. Chu, and A. F. B. Cheng. 2003. Epidemiology of rifampin ADP-ribosyltransferase (arr-2) and metallo-β-lactamase (blaIMP-4) gene cassettes in class 1 integrons in Acinetobacter strains isolated from blood cultures in 1997 to 2000. Antimicrob. Agents Chemother. 47:1382-1390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Jacoby, G. A., and L. S. Munoz-Price. 2005. The new β-lactamases. N. Engl. J. Med. 352:380-391. [DOI] [PubMed] [Google Scholar]
- 8.Lauretti, L., M. L. Riccio, A. Mazzariol, G. Cornaglia, G. Amicosante, R. Fontana, and G. M. Rossolini. 1999. Cloning and characterization of blaVIM, a new integron-borne metallo-β-lactamase gene from a Pseudomonas aeruginosa clinical isolate. Antimicrob. Agents Chemother. 43:1583-1590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Lee, K., Y. Chong, H. B. Shin, Y. A. Kim, D. Yong, and J. H. Yum. 2001. Modified Hodge test and EDTA-disk synergy tests to screen metallo-β-lactamase-producing strains of Pseudomonas and Acinetobacter species. Clin. Microbiol. Infect. 7:88-91. [DOI] [PubMed] [Google Scholar]
- 10.Lee, K., W. G. Lee, Y. Uh, G. Y. Ha, J. Cho, Y. Chong, and Korean Nationwide Surveillance of Antimicrobial Resistance Group. 2003. VIM- and IMP-type metallo-β-lactamase-producing Pseudomonas spp. and Acinetobacter spp. in Korean hospitals. Emerg. Infect. Dis. 9:868-871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Lee, K., Y. S. Lim, D. Yong, J. H. Yum, and Y. Chong. 2003. Evaluation of the Hodge test and the imipenem-EDTA double disk synergy test for differentiating metallo-β-lactamase-producing isolates of Pseudomonas spp. and Acinetobacter spp. J. Clin. Microbiol. 41:4623-4629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lee, K., J. B. Lim, J. H. Yum, D. Yong, Y. Chong, J. M. Kim, and D. M. Livermore. 2002. blaVIM-2 cassette-containing novel integrons in metallo-β-lactamase-producing Pseudomonas aeruginosa and Pseudomonas putida isolates disseminated in a Korean hospital. Antimicrob. Agents Chemother. 46:1053-1058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Levesque, C., and P. H. Roy. 1993. PCR analysis of integrons, p. 590-594. In D. H. Persing, T. F. Smith, F. C. Tenover, and T. J. White (ed.), Diagnostic molecular microbiology: principles and applications. American Society for Microbiology, Washington, D.C.
- 14.Livermore, D. M. 2002. The impact of carbapenemases on antimicrobial development and therapy. Curr. Opin. Investig. Drugs 3:218-224. [PubMed] [Google Scholar]
- 15.Loeffler, F. E., Q. Sun, J. Li, and J. M. Tiedje. 2000. 16S rRNA gene-based detection of tetrachloroethene-dechlorinating Desulfuromonas and Dehalococcoides species. Appl. Environ. Microbiol. 66:1369-1374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.National Committee for Clinical Laboratory Standards. 2003. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically. Approved standard, 6th ed. M7-A6. National Committee for Clinical Laboratory Standards, Wayne, Pa.
- 17.National Committee for Clinical Laboratory Standards. 2003. Performance standards for antimicrobial disk susceptibility testing, 8th ed. M2-A8. National Committee for Clinical Laboratory Standards, Wayne, Pa.
- 18.Nordmann, P., and L. Poirel. 2002. Emerging carbapenemases in gram-negative aerobes. Clin. Microbiol. Infect. 8:321-331. [DOI] [PubMed] [Google Scholar]
- 19.Poirel, L., T. Naas, D. Nicolas, L. Collet, S. Bellais, J.-D. Cavallo, and P. Nordmann. 2000. Characterization of VIM-2, a carbapenem-hydrolyzing metallo-β-lactamase and its plasmid- and integron-borne gene from a Pseudomonas aeruginosa clinical isolate in France. Antimicrob. Agents Chemother. 44:891-897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Pop-Vicas, A. E., and E. M. C. D'Agata. 2005. The rising influx of multidrug-resistant gram-negative bacilli into a tertiary care hospital. Clin. Infect. Dis. 40:1792-1798. [DOI] [PubMed] [Google Scholar]
- 21.Schreckenberger, P. C., M. I. Daneshvar, R. S. Weyant, and D. G. Hollis. 2003. Acinetobacter, Chryseobacterium, Moraxella, and other nonfermentative gram-negative rods, p. 749-779. In P. R. Murray, E. J. Baron, J. H. Jorgensen, M. A. Pfaller, and R. H. Yolken (ed.), Manual of clinical microbiology, 8th ed. American Society for Microbiology, Washington, D.C.
- 22.Toleman, M. A., A. M. Simm, T. A. Murphy, A. C. Gales, D. J. Biedenbach, R. N. Jones, and T. R. Walsh. 2002. Molecular characterization of SPM-1, a novel metallo-β-lactamase isolated in Latin America: report from the SENTRY antimicrobial programme. J. Antimicrob. Chemother. 50:673-679. [DOI] [PubMed] [Google Scholar]
- 23.Vaisvila, R., R. D. Morgan, J. Posfai, and E. A. Raleigh. 2001. Discovery and distribution of super-integrons among pseudomonads. Mol. Microbiol. 42:587-601. [DOI] [PubMed] [Google Scholar]
- 24.Walsh, T. R., M. A. Toleman, L. Poirel, and P. Nordmann. 2005. Metallo-β-lactamases: the quiet before the storm? Clin. Microbiol. Rev. 18:306-325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Wang, M., J. H. Tran, G. A. Jacoby, Y. Zhang, F. Wang, and D. C. Hooper. 2003. Plasmid-mediated quinolone resistance in clinical isolates of Escherichia coli from Shanghai, China. Antimicrob. Agents Chemother. 47:2242-2248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Watanabe, M., S. Iyobe, and S. Mitsuhashi. 1991. Transferable imipenem resistance in Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 35:147-151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Woodford, N., P. M. Tierno, Jr., Y. Katherine, L. Tysall, M.-F. I. Palepou, E. Ward, R. E. Painter, D. F. Suber, D. Shungu, L. L. Silver, K. Inglima, J. Kornblum, and D. M. Livermore. 2004. Outbreak of Klebsiella pneumoniae producing a new carbapenem-hydrolyzing class A β-lactamase, KPC-3, in a New York medical center. Antimicrob. Agents Chemother. 48:4793-4799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Yigit, H., A. M. Queenan, G. J. Anderson, A. Domenech-Sanchez, J. W. Biddle, C. D. Steward, S. Alberti, K. Bush, and F. C. Tenover. 2001. Novel carbapenem-hydrolyzing β-lactamase, KPC-1, from a carbapenem-resistant strain of Klebsiella pneumoniae. Antimicrob. Agents Chemother. 45:1151-1161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Yum, J. H., K. Yi, H. Lee, D. Yong, K. Lee, J. M. Kim, G. M. Rossolini, and Y. Chong. 2002. Molecular characterization of metallo-β-lactamase-producing Acinetobacter baumannii and Acinetobacter genomospecies 3 from Korea: identification of two new integrons carrying the blaVIM-2 gene cassettes. J. Antimicrob. Chemother. 49:837-840. [DOI] [PubMed] [Google Scholar]