Abstract
The results of normal-mode analyses are in accord with the proposal that a low-frequency motion of the HhaI methyltransferase enzyme is responsible for base flipping in bound DNA. The vectors of the low-frequency normal mode of residues Ser-85 and Ile-86 point directly to the phosphate and ribose moieties of the DNA backbone near the target base in position to rotate the dihedral angles and flip the base out of the DNA duplex. The vector of residue Gln-237 on the major groove is in the proper orientation to assist base separation. Our results favor the major groove pathway and the protein active process in base flipping.
Mammalian DNA (cytosine-5)-methyltransferases methylate certain CpG sequences that are believed to modulate gene expression and cell differentiation (1). Bacterial DNA (cytosine-5)-methyl transferases are a component of restriction-modification systems and serve as important tools for the manipulation of DNA structure and the analysis of protein–nucleic acid interactions. HhaI methyltransferase (M.HhaI) is one of the many S-adenosyl-l-methionine (AdoMet)-dependent DNA-modifying enzymes found in bacteria, plants, and animals (2). It catalyzes the AdoMet-dependent methylation of cytosine residues in specific sequences of DNA to give DNA-(5-methyl)cytosine and AdoHcy (S-adenosyl-l-homocysteine) (3). The 327-aa protein recognizes the sequence 5′-GCGC-3′ (4) in double-stranded DNA and methylates the first cytosine residue to yield 5′-GmeCGC-3′ (5).
Base Flipped
The structure of M.HhaI has been characterized extensively by x-ray crystallography in complexes with various forms of its DNA substrate (6–9) and without the substrate (10). The enzyme is folded into three parts: a large domain, a small domain, and a hinge region, which correspond to the three sides surrounding a cleft large enough to bind DNA in a binary form (without the DNA substrate) (10). The large domain (amino acids 1–193 and 304–327) is a mixed alpha/beta structure consisting of the N-terminal two-thirds of the protein followed by a crossover connection to an alpha helix from the C-terminal. The small domain contains seven strands (amino acids 194–275). The hinge region (amino acids 276–303) connects the two domains by forming the bottom of the cleft.
In a 1994 report of a ternary crystal structure of M.HhaI, its DNA substrate, and the reaction product S-adenosyl-l-homocysteine (SAM) (6), it was detailed for the first time that the target cytosine in the DNA substrate had swung completely out of the helix and into the active-site pocket of the enzyme. The base had flipped out with a conformational shift of 180°. The DNA binding induces a large-scale movement of the catalytic loop (residues 80–89) closure.
Catalysis is initiated by the attack of Cys-81 to the cytosine C6, followed by a slower transfer of the methyl group from AdoMet to the C5 position (6, 11). Beta elimination regenerates the active enzyme, and product release dominates kcat. Kinetic isotope studies and molecular dynamics (MD) simulations have shown that AdoMet-dependent methyltransferase uses active-site compressions to assist catalysis (12–15).
The Origins of Base Flipping
NMR dynamics studies of the base-flipping motions in the binary M.HhaI–DNA and the ternary M.HhaI–DNA-cofactor systems in solutions show that addition of the cofactor analog S-adenosyl-l-homocysteine greatly enhances the trapping of the target cytosine in the catalytic pocket detected by a clear signal that is attributed to the flipped state of the target base (16). The enhancement indicates that flipping of the cytosine base out of the DNA helix does not depend on binding of the target base in the catalytic pocket and suggests an active role of the enzyme in the flipping of the base (16). It is clear from the crystal structures (17) of M.HhaI and substrates containing mismatches [G:A, G:U, or G:AP (AP, abasic)] at the target base that the enzyme does not require a flipped-out cytosine residue as part of its initial recognition mechanism. Particularly interesting is that when M.HhaI binds to an abasic site, it flips the deoxyribose ring and its flanking phosphates into the same conformation that is adopted during base flipping on its normal substrates. It is believed that the push is not on the base, but rather on the sugar-phosphate backbone. Thus, rotation of the DNA backbone may be the key to base flipping, with the base merely carried along with it.
Base-Flipping Pathways
Theoretical methods based on empirical force fields provide alternative ways to view and to study microscopically the dynamics of base flipping in time and in space. It was originally thought that flipping occurred via the minor groove because the M.HhaI DNA recognition domain approached the DNA from the major groove side, blocking this as a potential pathway (6). DNA-only MD simulations of abasic systems supported the minor groove pathway hypothesis (18). MD simulations were carried out to reveal the energetic and conformational feature of the base-flipping states via opening into the minor groove or major groove of the double helix (19, 20). Umbrella sampling methods simulating the opening of the central cytosine base indicated that the minor or major groove pathways had similar energetic barriers. However, the major groove route is generally assumed for steric reasons (21). Movement of the base from the base stacked position toward the major groove occurs smoothly, providing a structure in accord with the crystal structure of M.HhaI in terms of the backbone conformational change. A major groove pathway is suggested based on free energy calculations of flipping in DNA complexed to M.HhaI (22). When the target cytosine is replaced by an abasic south-constrained pseudosugar, the resulting crystal structure (23) shows the pseudosugar to be trapped on the DNA major groove side rotated by ≈90° about its flanking phosphodiester bonds. This angle corresponds to the mid-point along the flipping pathway. MD calculations suggest that the new crystal structure corresponds to an intermediate stage of the sugar and phosphodiester backbone moieties halfway along the flipping corridor (23). This finding is consistent with the major groove pathway.
Structural, kinetic, and computational data for M.HhaI are available in abundance. However, the amount of studies for the mechanism of the base flipping in terms of long-time protein dynamics is limited. For example, it is not known which specific residues are involved in the possible pushing of the base out of the DNA helix, and how the “push” rotates the backbone and positions the base out of the helix. Normal-mode analysis (NMA) is a method for studying long-time dynamics and elasticity of biological systems. Many studies have shown that the results from NMA can also provide a proper description of the functionally important motions of the protein (24–36).
In this study, we probe the low-frequency normal mode of the M.HhaI molecule to understand how this mode of protein motion affects the base-flipping process.
Methods
NMA was performed by using the elastic network model (37), applying the single-parameter Hookean potential (38) and protein models. Several studies have shown that a single parameter potential is sufficient to reproduce complex slow dynamics in good detail (31, 37, 38). Moreover, the elastic network model is an efficient way to study the characters and directions of low-frequency motions. A lot of information about the nature of the large conformational changes of biological systems can be obtained from a single normal mode. For example, a single low-frequency normal mode can be found whose direction compares well with the conformational change or gives a good description of the pattern of the atomic displacement as observed experimentally. The single-parameter Hookean potential (38) was also tested, using an all-atom model, on a periplasmic maltodextrin binding protein. The results indicated that the slowest modes closely map the open form going to the closed form (38).
In our elastic network model, protein residues were represented by their Cα atoms. To reduce space requirements, 27 residues from the C terminus were removed, leaving 300 residues. A Hessian matrix was constructed by using a force constant of 1 kJ·Å–2·mol–1 and a cutoff of 8 Å. The figures showing the normal-mode vectors were generated by using the program vmd (39).
Results and Discussion
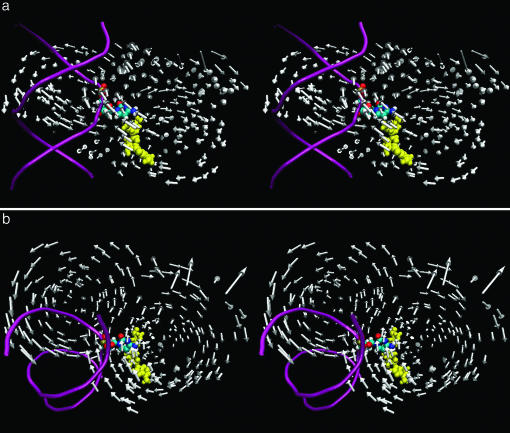
Fig. 1 shows an overall picture of the low-frequency mode of the enzyme interaction. Each arrow in gray represents the direction of motion of the Cα atom. The enzyme residues are omitted for easier viewing. The DNA backbone structure, the flipped out base, and the cofactor SAM are shown in pink, yellow, and atom color, respectively. In Fig. 1a, in front of SAM and the flipped out base of the large domain (amino acids 1–193) that appear on the right of the figure, there are a group of arrows pointing up toward the phosphodiester and sugar groups of the base-flipped nucleotide in the 5′ direction. Details of this region and these arrows are shown in Fig. 2. For the small domain (amino residues 194–275) at the back of the DNA backbone (in pink), there are a group of arrows pointing toward the orphan base guanine of the complementary strand from the major groove. Details of this region are shown in Fig. 3.
Fig. 1.
An overall picture of the low-frequency mode of the enzyme interaction. Each arrow in gray represents the direction of motion of the Cα atom. The enzyme residues are omitted for easier viewing. The DNA backbone structure, the flipped out base, and the cofactor SAM are shown in pink, yellow, and atom color, respectively. b is a top view of a. The arrows representing the large (lower right) and small (upper left) domains qualitatively form two circles. The group of arrows entering the minor groove (from the right of the helix) point directly toward the phosphodiester and sugar backbone around the flipped base. The arrows entering the major groove from the back (left) of the helix point directly to the complementary orphan guanine.
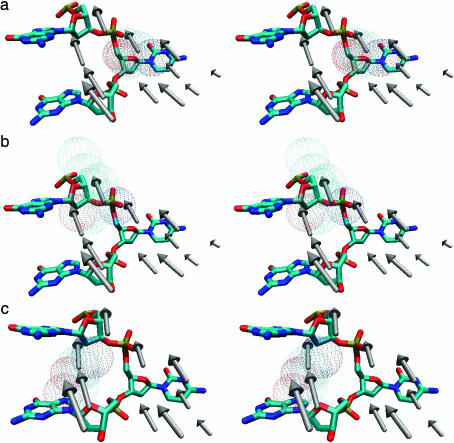
Fig. 2.
An enlarged local view of the vectors for the residues 79–89 (Phe-79, Pro-80, Cys-81, Gln-82, Ala-83, Phe-84, Ser-85, Ile-86, Ser-87, Gly-88, and Lys-89). The flipped out base and the adjacent bases of the same strand are shown explicitly. (a) Ser-85 is shown in a dotted surface fashion. The vector of this residue is located on the lower right within the top trio of vectors. (b) The vector of Ile-86, shown as a dotted surface, is located at the center and on top of the trio of vectors. (c) The Ser-87 vector, shown in a dotted surface fashion, is located on the lower left of the top trio of vectors. Ser-87 is situated on the minor groove side of the DNA between the adjacent bases. The Ser-87 vector also points to the sugar of the adjacent guanine on the 5′ side.
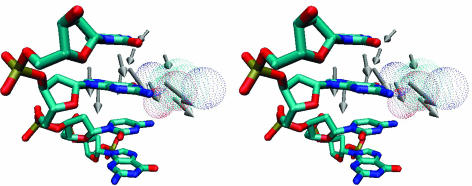
Fig. 3.
An enlarged local view of the vectors for the residues 230–240 (Gly-230, Ile-231, Val-232, Gly-233, Lys-234, Gly-235, Gly-236, Gln-237, Gly-238, Glu-239, and Arg-240). These vectors are located in the back of the major groove of the small domain and point toward the complementary orphan guanine base (Fig. 1 a and b). The orphan guanine and the adjacent bases of the same strand are shown explicitly. Residue Gln-237, shown in a dotted surface fashion, is situated on the major groove side in front of the group of vectors.
Fig. 1b is a top view of Fig. 1a. The arrows representing the large (lower right) and small (upper left) domains qualitatively form two circles moving toward the middle of the DNA duplex. A closer look at Fig. 1a shows that these circles merge at an angle. In Fig. 1b, the group of arrows entering the minor groove (from the right of the helix) point directly toward the phosphodiester and sugar backbone around the flipped base. The arrows entering the major groove from the back (left) of the helix point directly to the complementary orphan guanine.
Fig. 2 shows the vectors for residues 79–89 (Phe-79, Pro-80, Cys-81, Gln-82, Ala-83, Phe-84, Ser-85, Ile-86, Ser-87, Gly-88, and Lys-89). These vectors are located in front of the large domain and point up toward the phosphodiester group of the flipped out base (Fig. 1 a and b). Fig. 2 provides an enlarged local view of the picture. The flipped out base and the adjacent bases of the same strand are shown explicitly to display their relative positions. In Fig. 2a, residue Ser-85 is shown in a dotted surface fashion. The vector of this residue is located on the lower right within the top trio of vectors. Obviously, Ser-85 has direct influence on the phosphodiester and sugar group of the flipped out nucleotide. In Fig. 2b, the vector of residue Ile-86, shown as a dotted surface, is located at the center and on top of the trio of vectors. Ile-86 has direct influence on the phosphodiester group of the flipped out nucleotide. In Fig. 2c, the Ser-87 vector, shown in a dotted surface fashion, is located on the lower left of the top trio of vectors. Because Ser-87 is situated on the minor groove side of the DNA between the adjacent bases, the natural way for the base to flip out must be from the major groove. Possible functions of this residue are to push the cytosine base from the minor groove side or to block the minor groove pathway so as to lead the base to flip toward the major groove pathway. Also, the Ser-87 vector points to the sugar of the adjacent guanine on the 5′ side. Another possible role of this vector is to push the 5′ guanine to elongate the backbone and to allow more space for the flipped base.
The remaining vectors in Fig. 2 located behind the trio are pushed by the rest of the large domain residues (shown in Fig. 1 a and b), so they will push the trio of vectors on the top of the closing circle. This group of vectors has an upwards direction toward the phosphodiester group, changing the dihedral conformation and deforming the space between the bases of the adjacent nucleosides. The new configuration may also have elongated the nearby backbone, leaving additional space for the base to flip out.
According to previous crystal structure findings (figure 5 of ref. 18), the changes in backbone dihedral angles correlate with the base flipping out. The local phosphodiester dihedrals were obtained from the flipped region of nine DNA-methyltransferase crystal structures. For the backbone of the flipped base, there are significant differences in the γ and ζ dihedrals. Beyond the flipped base, significant changes also occur in the adjacent ε dihedral on the 5′ side and in the β dihedral on the 3′ side. Perturbations on backbone dihedral angles were also observed by NMR studies (40, 41). Our study shows that the vectors of the low-frequency normal mode of residues Ser-85 and Ile-86 point directly to the phosphate and sugar groups of the DNA backbone near the target base (Fig. 2) in position to rotate the dihedral angles and flip the base out of the DNA duplex.
Fig. 3 shows the vectors for the residues 230–240 (Gly-230, Ile-231, Val-232, Gly-233, Lys-234, Gly-235, Gly-236, Gln-237, Gly-238, Glu-239, and Arg-240). These vectors are located in the back of the major groove of the small domain and point toward the complementary orphan guanine base (Fig. 1 a and b). Fig. 3 provides an enlarged local view of the region. The orphan guanine and the adjacent bases of the same strand are shown explicitly to display their relative positions. In Fig. 3, Gln-237 (shown in a dotted surface fashion) is situated on the major groove side in front of the group of vectors. The motion of these vectors is in the proper orientation to assist base separation.
Residues Ser-87 and Gln-237 enter the helix (from the minor and major grooves, respectively) and form a hydrogen bond with each other. Gln-237 also forms three hydrogen bonds with the orphan guanine, which originally complements the target cytosine. Our study shows that Ser-87 is in position to push the cytosine base from the minor groove side or to block the minor groove to lead the base flipping toward the major groove passage. The Gln-237 vector points directly to the orphan guanine, providing the “push” to perturb the base-pairing and allowing the side chain to then penetrate into the DNA duplex. However, mutation of Gln 237 to any of the other 19 amino acids (42) shows little change in the enzyme function but greatly reduces the stability of the protein–DNA complex. From our model, this implies that the network of the low-frequency vectors might be unchanged in the mutation such that the “push” is still provided, but the hydrogen bonds to the orphan guanine by the mutated residue are altered such that the stability of the protein–DNA complex is affected. Protein sequences may have been designed through evolutionary process to allow the protein to follow the direction of a single mode, because of the shape it happened to have, as a starting point for its large amplitude functional motions.
Our study shows that in low-frequency normal mode, the vectors exert forces directly on the DNA from minor and major groove sides and on the backbone near the target cytosine on the 5′ end, supporting the conclusion that our results favor the active process and major groove pathway. If backbone rotation is the key to base flipping, then amino acids in the enzyme that interact with the DNA backbone (Ser-85, Ile-86, and Ser-87 in this study) will be the prime candidates for site-specific mutagenesis to probe their functions. The method applied in this study can also be used to test the network of vectors upon mutation.
An earlier MD study (14), using CHARMM (43, 44), derived a dynamic cross-correlation map (figure 2A of ref. 14) based on the high-resolution cr ystal structure of the enzyme·cofactor·dsDNA structure M.HhaI·AdoMet·d(CCATGCGCTGAC)2 (pdb6mht.ent) solved at 2.05 Å (9). The cross-correlation map (45–49) was calculated from residue–residue-based correlated motions. Fig. 4 shows the preliminary qualitative assignments of the anticorrelated motions mapped onto the 3D stereo DNA methyltransferase M.HhaI complex molecular structure. Detailed cross-correlations are discussed in the legend of Fig. 4. The correlated motions, analyzed in a nanosecond time frame (14), can be summarized qualitatively in three directions (x, y, and z). These motions perturb the active site at higher frequencies (MD time frame) in a molecular breathing fashion. Two amino acid residues are located on opposite sides of the active site that move toward each other (anticorrelated motion) and push the reactants together. The low-frequency motions seen by normal-mode analysis (NMA) close the catalytic loop in the large domain and at the same time flip the base out of the duplex into the catalytic pocket. It is interesting to note the correlated character of the two circles of normal-mode vector networks revealed in Fig. 1.
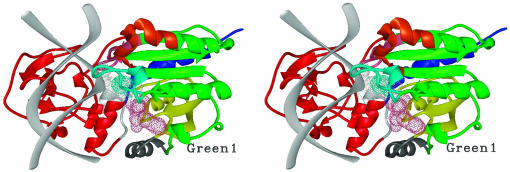
Fig. 4.
Anticorrelated motions mapped onto the 3D stereo DNA methyltransferase M.HhaI in a complex [d(CCATGCGCTGAC)2 and AdoMet]. The DNA backbone is shown as a light gray ribbon. The flipped out base and AdoMet are shown by light gray and pink dotted surfaces, respectively. The Cys-81 side chain is shown as a dotted cyan surface. Different anticorrelated regions are color coded for easy viewing. Residues 186–279 and 291–304 (red) of the small domain are anticorrelated with residues 1–40 (yellow), 55–79, 98–120, 135–150, and 165–185 (green) and 305–327 (blue) in large catalytic domain. Residues 280–290 (light gray) of the small domain are anticorrelated with residues 80–97 (cyan) in the large catalytic domain. Residues 80–97 are also anticorrelated, within the large catalytic domain, with residues 1–40 (yellow) and 305–327 (blue). The following anticorrelated motions are also within the large catalytic domain: residues 121–134 (orange) are anticorrelated with residues 1–40 (yellow) and residues 41–54 (gray). The pink region, residues 151–164, is anticorrelated with residues 41–54 (gray) and residues 55–79 (labeled green 1).
Acknowledgments
We thank Jing Yuan for assistance. This work was supported by National Institutes of Health Grant 5R37DK09171.
Author contributions: T.C.B. designed research; J.L. performed research; J.L. analyzed data; and J.L. and T.C.B. wrote the paper.
Abbreviations: AdoMet, S-adenosyl-l-methionine; HhaI, HhaI methyltransferase; MD, molecular dynamics.
References
- 1.Doerfler, W. (1983) Annu. Rev. Biochem. 52, 93–124. [DOI] [PubMed] [Google Scholar]
- 2.Schubert, H. L., Blumenthal, R. M. & Cheng, X. (2003) Trends Biochem. Sci. 28, 329–335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Wu, J. C. & Santi, D. V. (1987) J. Biol. Chem. 262, 4778–4786. [PubMed] [Google Scholar]
- 4.Roberts, R. J., Myers, P. A., Morrison, A. & Murray, K. (1976) J. Mol. Biol. 103, 199–208. [DOI] [PubMed] [Google Scholar]
- 5.Mann, M. B. & Smith, H. O. (1979) in Proceedings of the Conference on Transmethylation, eds. Usdin, E., Borchardt, R. T. & Creveling, C. R. (Elsevier, New York), pp. 483–492.
- 6.Klimasauskas, S., Kumar, S., Roberts, R. J. & Cheng, X. (1994) Cell 76, 357–769. [DOI] [PubMed] [Google Scholar]
- 7.O'Gara, M., Klimasauskas, S., Roberts, R. J. & Cheng, X. (1996) J. Mol. Biol. 261, 634–645. [DOI] [PubMed] [Google Scholar]
- 8.O'Gara, M., Roberts, R. J. & Cheng, X. (1996) J. Mol. Biol. 263, 597–606. [DOI] [PubMed] [Google Scholar]
- 9.Kumar, S., Horton, J. R., Jones, G. D., Walker, R. T., Roberts, R. J. & Cheng, X. (1997) Nucleic Acids Res. 25, 2773–2783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cheng, X., Kumar, S., Posfai, J., Pflugrath, J. W. & Roberts, R. J. (1993) Cell 74, 299–307. [DOI] [PubMed] [Google Scholar]
- 11.Svedruzic, Z. & Reich, N. O. (2004) Biochemistry 43, 11460–11473. [DOI] [PubMed] [Google Scholar]
- 12.Lau, E. Y. & Bruice, T. C. (1998) J. Am. Chem. Soc. 120, 12387–12394. [Google Scholar]
- 13.Lau, E. Y. & Bruice, T. C. (1999) J. Mol. Biol. 293, 9–18. [DOI] [PubMed] [Google Scholar]
- 14.Estabrook, R. A., Luo, J., Purdy, M. M., Sharma, V., Weakliem, P., Bruice, T. C. & Reich, N. O. (2005) Proc. Natl. Acad. Sci. USA 102, 994–999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Takusagawa, F., Fulioka, M., Spies, A. & Schowen, R. L. (1998) in Comprehensive Biological Catalysis, ed. Sinnot, M. (Academic, San Diego), pp. 1–30.
- 16.Klimasauskas, S., Szyperski, T., Serva, S. & Wuthrick, K. (1998) EMBO J. 17, 317–324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.O'Gara, M., Horton, J. R., Roberts, R. J. & Cheng, X. (1998) Nat. Struct. Biol. 5, 872–877. [DOI] [PubMed] [Google Scholar]
- 18.Wang, P. Y., Brank, A. S., Banavali, N. K., Nicklaus, M. C., Marquez, V. E., Christman, J. K. & MacKerell, A. D., Jr. (2000) J. Am. Chem. Soc. 122, 12422–12434. [Google Scholar]
- 19.Várnai, P. & Lavery, R. (2002) J. Am. Chem. Soc. 124, 7272–7273. [DOI] [PubMed] [Google Scholar]
- 20.Banavali, N. K. & MacKerell, A. D., Jr. (2002) J. Mol. Biol. 319, 141–160. [DOI] [PubMed] [Google Scholar]
- 21.Ramstein, J. & Lavery, R. (1988) Proc. Natl. Acad. Sci. USA 85, 7231–7235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Huang, N., Banavali, N. K. & MacKerell, A. D., Jr. (2003) Proc. Natl. Acad. Sci. USA 100, 68–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Horton, J. R., Ratner, G., Banavali, N. K., Huang, N., Choi, Y., Maier, M. A., Marquez, V. E., MacKerell, A. D., Jr., & Cheng, X. (2004) Nucleic Acids Res. 32, 3877–3886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Go, N., Noguti, T. & Nishikawa, T. (1983) Proc. Natl. Acad. Sci. USA 80, 3696–3700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Brooks, B. & Karplus, M. (1983) Proc. Natl. Acad. Sci. USA 80, 6571–6575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Levitt, M., Sander, C. & Stern, P. S. (1985) J. Mol. Biol. 181, 423–447. [DOI] [PubMed] [Google Scholar]
- 27.Gibrat, J. F. & Go, N. (1990) Proteins 8, 258–279. [DOI] [PubMed] [Google Scholar]
- 28.Marques, O. & Sanejouand, Y. H. (1995) Proteins 23, 557–560. [DOI] [PubMed] [Google Scholar]
- 29.Wako, H., Tachikawa, M. & Ogawa, A. (1996) Proteins 26, 72–80. [DOI] [PubMed] [Google Scholar]
- 30.Thomas, A., Field, M. J. & Perahia, D. (1996) J. Mol. Biol. 257, 1070–1087. [DOI] [PubMed] [Google Scholar]
- 31.Hinsen, K. (1998) Proteins 33, 417–429. [DOI] [PubMed] [Google Scholar]
- 32.Kitao, A. & Go, N. (1999) Curr. Opin. Struct. Biol. 9, 164–169. [DOI] [PubMed] [Google Scholar]
- 33.Miller, D. W. & Agard, D. A. (1999) J. Mol. Biol. 286, 267–278. [DOI] [PubMed] [Google Scholar]
- 34.Kikuchi, H., Wako, H., Yura, K., Go, M. & Mimuro, M. (2000) Biophys. J. 79, 1587–1600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Berendsen, H. J. C. & Hayward, S. (2000) Curr. Opin. Struct. Biol. 10, 165–169. [DOI] [PubMed] [Google Scholar]
- 36.Chacon, P., Tama, F. & Wriggers, W. (2003) J. Mol. Biol. 326, 485–492. [DOI] [PubMed] [Google Scholar]
- 37.Tama, F. & Sanejouand, Y.-H. (2001) Protein Eng. 14, 1–6. [DOI] [PubMed] [Google Scholar]
- 38.Tirion, M. M. (1996) Phys. Rev. Lett. 77, 1905–1908. [DOI] [PubMed] [Google Scholar]
- 39.Humphrey, W., Dalke, A. & Schulten, K. (1996) J. Mol. Graphics 14, 33–38. [DOI] [PubMed] [Google Scholar]
- 40.Meints, G. A. & Drobny, G. P. (2001) Biochemistry 40, 12436–12443. [DOI] [PubMed] [Google Scholar]
- 41.Marcourt, L., Cordier, C., Couesnon, T. & Dodin, G. (1999) Eur. J. Biochem. 265, 1032–1042. [DOI] [PubMed] [Google Scholar]
- 42.Mi, S., Alonso, D. & Roberts, R. J. (1995) Nucleic Acids Res. 23, 620–627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Brooks, B. R., Bruccoleri, R. E., Olafson, B. D., States, D. J., Swaminathan, S. & Karplus, M. (1983) J. Comput. Chem. 4, 187–217. [Google Scholar]
- 44.MacKerell, A. D., Bashford, D., Bellott, M., Dunbrack, R. L., Evanseck, J. D., Field, M. J., Fischer, S., Gao, J., Guo, H., Ha, S., et al. (1998) J. Phys. Chem. B 102, 3586–3616. [DOI] [PubMed] [Google Scholar]
- 45.Ichiye, T. & Karplus, M. (1991) Proteins Struct. Funct. Genet. 11, 205–217. [DOI] [PubMed] [Google Scholar]
- 46.McCammon, A. J. & Harvey, S. C. (1986) Dynamics of Proteins and Nucleic Acids (Cambridge Univ. Press, Cambridge, U.K.).
- 47.Swaminathan, S., Harte, W. E., Jr., & Beveridge, D. L. (1991) J. Am. Chem. Soc. 113, 2717–2721. [Google Scholar]
- 48.Radkiewicz, J. L. & Brooks, C. L., III (2000) J. Am. Chem. Soc. 122, 225–231. [Google Scholar]
- 49.Luo, J. & Bruice, T. C. (2002) Proc. Natl. Acad. Sci. USA 99, 16597–16600. [DOI] [PMC free article] [PubMed] [Google Scholar]