Abstract
Usher syndrome type I is an autosomal recessive disorder marked by hearing loss, vestibular areflexia, and retinitis pigmentosa. Six Usher I genetic subtypes at loci USH1A–USH1F have been reported. The MYO7A gene is responsible for USH1B, the most common subtype. In our analysis, 151 families with Usher I were screened by linkage and mutation analysis. MYO7A mutations were identified in 64 families with Usher I. Of the remaining 87 families, who were negative for MYO7A mutations, 54 were informative for linkage analysis and were screened with the remaining USH1 loci markers. Results of linkage and heterogeneity analyses showed no evidence of Usher types Ia or Ie. However, one maximum LOD score was observed lying within the USH1D region. Two lesser peak LOD scores were observed outside and between the putative regions for USH1D and USH1F, on chromosome 10. A HOMOG χ2(1) plot shows evidence of heterogeneity across the USH1D, USH1F, and intervening regions. These results provide conclusive evidence that the second-most-common subtype of Usher I is due to genes on chromosome 10, and they confirm the existence of one Usher I gene in the previously defined USH1D region, as well as providing evidence for a second, and possibly a third, gene in the 10p/q region.
Usher syndrome is defined as congenital neurosensory hearing loss with retinitis pigmentosa (RP). Its frequency is estimated to be 3.5/100,000 in Scandinavia (Hallgren 1959; Nuutila 1970; Grondahl 1987) and 4.4/100,000 in the United States (Boughman et al. 1983). Although relatively rare, Usher syndrome has been estimated to account for 50% of all individuals who are both deaf and blind and for ∼3%–6% of all children who are deaf (Vernon 1969).
Usher syndrome is both clinically and genetically heterogeneous. Clinical heterogeneity is demonstrated by the variation in the severity and progression of the hearing impairment, the age at onset of retinal degeneration, and the presence or absence of vestibular areflexia. There are three clinical types of Usher syndrome. Usher type I is the most severe subtype, characterized by congenital profound deafness, early-onset RP (usually diagnosed before puberty), and absent or severely diminished vestibular responses. Usher type II is marked by a congenital moderate-to-severe hearing impairment that is identified by a characteristic sloping audiogram, a later diagnosis of RP (during the 2d decade of life), and normal vestibular responses. Usher type III is characterized by a progressive hearing loss, with variable RP and progressive vestibular dysfunction (Kimberling and Moller 1995; Smith et al. 1995). Types I and II are the most common forms of Usher syndrome.
At least 10 loci have been identified for Usher syndrome: 6 for Usher type I (USH1A–USH1F), 3 for Usher type II (USH2A–USH2C) and 1 for Usher type III (Kimberling et al. 1990, 1992; Kaplan et al. 1992; Smith et al. 1992; Sankila et al. 1995; Wayne et al. 1996, 1997; Chaib et al. 1997; Hmani et al. 1999; Pieke-Dahl et al. 2000) (table 1). The most common form of Usher type I, Ib, is localized to 11q13.5 (locus USH1B [MIM 276903]). The other five loci are believed to be uncommon: the gene for Usher type Ia maps to 14q32 and has been observed in families with ancestry from the Poitou-Charentes region of France (Kaplan et al. 1992); type Ic maps to 11p15.1 and has been reported in the French Acadian population (Smith et al. 1992); type Ie has been mapped to 21q21 by homozygosity mapping in a Moroccan family (Chaib et al. 1997); and types Id and If both map to chromosome 10 and have been identified in single Pakistani (Id) and Hutterite (If) families, respectively (Wayne et al. 1996, 1997). Usher syndrome type II is known to have at least three loci, and there is evidence for an additional, as yet unlocalized, subtype (Pieke-Dahl et al. 2000). Usher type IIa is the most common of the milder forms of Usher syndrome and maps to 1q41 (Kimberling et al. 1990). Type IIb maps to chromosome 3p23-24.2 (Hmani et al. 1999), and Usher type IIc maps to 5q14.3-21.3 (Pieke-Dahl et al. 2000). There is only one Usher III locus, USH3, mapping to 3q21-25 (Sankila et al. 1995). Families reported to have Usher type III are primarily of Finnish origin; however, there are reports of one Italian family (Gasparini et al. 1998) and two Spanish families that showed linkage to the 3q region (Espinos et al. 1998).
Table 1.
Summary of the Clinical and Genetic Subtypes of the Usher Syndromes
| Usher Typea and Location | Gene | Screening Markersb |
| I | ||
| USH1A-14q32 | D14S78, D14S250, D14S292, D14S260 | |
| USH1B-11q13.5 | MYO7A | D11S527, D11S4186, D11S906, D11S911 |
| USH1C-11p15.1 | USH1C | D11S921, D11S902, D11S4160, D11S899 |
| USH1D-10q21-22 | D10S529, D10S195, D10S202, D10S573 | |
| USH1E-21q21 | D21S1905, D21S1914, D21S269, D21S1913 | |
| USH1F-10 | D10S193, D10S1791, D10S220, D10S1790 | |
| II | ||
| USH2A-1q41c | USH2A | D1S237, AFM143, AFM144, D1S490 |
| USH2B-3p | ||
| USH-5qc | D5S617, D5S484, D5S505, D5S485 | |
| III | ||
| USH3-3q21-25 | D3S1555, D3S1308, D3S1279, D3S1280 |
Phenotypes are described in the text.
Markers typed for linkage analysis in the present study.
Families were tested for linkage for Usher II and III markers. The results were negative and are not reported here.
For the 10 known Usher loci, 3 genes have been identified. MYO7A is responsible for Usher Ib; USH2A is a novel gene, responsible for Usher IIa, that codes for a protein now called “usherin”; and the third gene, USH1C, which codes for a PDZ (PSD95, Dlg, ZO-1) domain–containing protein named “harmonin,” has been recently identified (Verpy et al. 2000; Bitner-Glindzicz et al. 2000). Mutations in MYO7A have been found to account for ∼60% of all Usher I cases (W.J.K., unpublished data). Although a wide spectrum of mutations in the MYO7A gene have been described in patients with Usher Ib, the gene has also been shown to harbor mutations producing nonsyndromic deafness, including DFNA11 and DFNB2, as well as atypical Usher syndrome with progressive hearing loss (Levy et al. 1997; Liu et al. 1997a, 1997b, 1998).
Here we report the results of linkage and mutation analysis of 169 families with Usher that represent diverse backgrounds from the United States, Europe, Colombia, and South Africa. Our results show that one or more genes on chromosome 10 are responsible for a majority of the families with Usher I that is not linked to MYO7A (MYO7A−).
We studied 169 independent families, including 242 affected individuals, with Usher syndrome. Of these families, 151 had Usher I, 12 had Usher II, and 6 had Usher III (table 2). The results for Usher types II and III will be reported elsewhere. Sixteen families with Usher I were consanguineous. The families were collected in the United States (86 families), The Netherlands (37 families), Sweden (31 families), South Africa (5 families), Spain (4 families), Colombia (3 families), and England (3 families); however, the ethnic composition is primarily of European extraction. The five South African and three Colombian families were of European ancestry. The sample group also included two French Acadian families who showed linkage to the USH1C region.
Table 2.
Distribution of MYO7A Mutations in 169 Families with Usher Syndrome[Note]
| UsherPhenotype(s) | Mutation | Linkage Informative? | No. |
| I | MYO7A+ | Yes | 46 |
| I | MYO7A+ | No | 18 |
| I | MYO7A− | Yes | 54 |
| I | MYO7A− | No | 33 |
| II and III | MYO7A− | No | 18 |
| Total | 169 |
Note.— Families are divided by phenotype, the presence or absence of at least one pathologic MYO7A mutation, and informativeness. MYO7A+ represents 64 families with Usher I with at least one identifiable MYO7A mutation; only 46 of these families are informative for linkage. The remaining families revealed no mutation (MYO7A−). The analysis reported here has focused on the 54 MYO7A− families with Usher I that were informative for linkage analysis.
All families were assigned to a specific subtype of Usher syndrome on the basis of the pattern and severity of hearing loss, the presence or absence of vestibular areflexia, and the presence of RP. The clinical diagnosis was determined from the patient’s medical history and results of audiometric, ophthalmologic, and vestibular examinations. The human-subjects committee at each location approved this study, and written, informed consent was obtained from all participants.
Blood samples were collected, and genomic DNA was extracted using a Puregene kit (Gentra Systems). Fluorescent (6-FAM, TET, or HEX) oligonucleotide primers were used to amplify polymorphic markers surrounding the various Usher loci (for the specific markers that were typed, see table 1). Markers were amplified according to the following protocol: 95°C for 5 min, 30 cycles (94°C for 1 min, 60°C for 1 min, and 72°C for 1 min) and 60°C for 30 min. PCR products were then pooled, denatured, and separated on 4.25% polyacrylamide gels in an ABI 377 automated sequencer (PE Biosystems), and fragment analysis was performed using GENESCAN 2.1 software. Genotyping data were collected and were analyzed using GENOTYPER 2.0.
All families with Usher I in the present report had been subjects of a previous report on a mutation screen of 14 exons (Weston et al. 1996). However, the mutation analysis in the present report involved screening 46 of the 48 coding exons for all 169 families in our sample. A summary of the mutations observed is available on the GeneClinics home page, and a report on these mutations will be published elsewhere.
Linkage analysis of DNA markers was performed using the LINKAGE programs (Lathrop et al. 1985). Two-point and multipoint analyses were computed for all 54 families with MYO7A− Usher syndrome. Rolling multipoint analysis across the entire span of chromosome 10 markers generated LOD scores by comparison of overlapping sets of four contiguous marker loci with the test locus (five-point analysis). The pattern of inheritance of Usher syndrome was assumed to be recessive with full penetrance for the mutant homozygote. Gene frequency of the mutant allele was set at .001. The number of alleles was taken from information in the Whitehead Institute for Biomedical Research/MIT Center for Genome Research home page and the The Genome Database home page. The frequencies of the marker alleles were assumed to be equal. Heterogeneity analysis was done using the rolling multipoint LOD scores, generated as outlined above, with HOMOG, version 2.4.
Of the 169 families initially studied, 151 had Usher type I. Of the families with Usher I, 64 had at least one detectable MYO7A mutation. Of the families with MYO7A− Usher I, 54 were informative for linkage and were analyzed using the LINKAGE and HOMOG computer programs. These 54 families were screened with flanking markers, as summarized in table 1, for all known USH1 loci. Both two-point and rolling multipoint analyses were performed on all families, using all USH1 loci. Except for chromosome 10 markers, none of the LOD scores were significantly positive—that is, there was no evidence for linkage outside chromosome 10. All sets of scores generated from the rolling multipoint analysis were used for a multiple HOMOG analysis. Table 3 summarizes the results of the HOMOG analyses for each Usher locus. The χ2 values of 0.81 and 0.11 are clearly not significant, providing no evidence for the presence of either USH1A or USH1E in this sample. However, the χ2(1) values of 3.98, 4.15, and 36.30 suggest the presence of a mixture of USH1B, USH1C, USH1D, or USH1D/USH1F within the 54 families.
Table 3.
Summary of Heterogeneity Analysis of 54 Families with Usher Type I[Note]
| Usher Subtype | Marker Set | α | Heterogeneity χ2 |
| IA | 14q | .05 | .11 |
| IB | 11q | .15 | 3.98a |
| IC | 11p | .15 | 4.15a |
| ID/IF | 10 | .75 | 36.30a |
| IE | 21q | .05 | .81 |
Note.— Heterogeneity analysis was performed with the HOMOG program using markers spanning each of the USHI loci.
Significant, providing evidence for presence of subtypes.
Two French Acadian families were included in the MYO7A screening to test the possibility of a digenic effect involving the MYO7A gene; however, no mutations were detected. The sensitivity of the HOMOG program is demonstrated by the detection of linkage heterogeneity as a result of the inclusion of these two families. In addition, we identified three families that each had LOD scores >2.0 for 11q markers. We believe that these families have MYO7A mutations that were not detectable by our testing strategy. When these five families were removed from the analysis, the significant χ2(1) HOMOG values for USH1B and USH1C disappeared.
The highly significant χ2(1) value of 36.30 for chromosome 10 markers prompted us to further examine the USH1D and USH1F regions. The USH1D and USH1F loci have been mapped to chromosomal locations 10q21.22 and 10pq (fig. 1). The broad critical regions and close proximity of these two loci to each other necessitated the selection of 12 chromosome 10 markers spanning both critical regions, as well as the intervening space, covering a distance of ∼50 cM. These markers were used in two-point and rolling multipoint analyses. Table 4 shows the results of the two-point analysis for the individual markers, for USHID/USH1F. The highest LOD score, 5.27, was obtained for marker D10S529 at θ=.10.
Figure 1 .
Map of chromosome 10. The critical regions of USH1D and USH1F are outlined, and the markers used in the linkage analysis are listed, with their position (in cM) from the p-arm telomere.
Table 4.
Results of Two-Point Analysis, on Chromosome 10, for 54 Families with Usher Type I
|
Two-Point LOD Score at θ = |
||||||
| Marker | 0 | .05 | .1 | .2 | .3 | .4 |
| D10S193 | −∞ | −8.648 | −3.316 | .019 | .527 | .264 |
| D10S1791 | −∞ | −3.962 | −.213 | 1.487 | 1.130 | .411 |
| D10S220 | −∞ | −4.106 | −.375 | 1.378 | .999 | .278 |
| D10S1784 | −∞ | .147 | .120 | .072 | .033 | .009 |
| D10S1790 | −∞ | −3.245 | .099 | 1.545 | 1.027 | .240 |
| D10S1756 | −∞ | −.652 | .680 | 1.115 | .723 | .240 |
| D10S1743 | −∞ | 3.097 | 4.210 | 3.517 | 1.918 | .560 |
| D10S1665 | −∞ | .669 | 2.219 | 2.106 | 1.136 | .259 |
| D10S529 | −∞ | 3.970 | 5.268a | 4.297 | 2.339 | .629 |
| D10S195 | −∞ | 1.859 | 4.219 | 4.175 | 2.494 | .802 |
| D10S202 | −∞ | 1.050 | 3.142 | 3.198 | 1.916 | .621 |
| D10S573 | −∞ | −3.035 | .261 | 1.658 | 1.138 | .338 |
Highest two-point LOD score.
Multipoint analysis of the non-USH1B families, with the exclusion of the two families of French Acadian ancestry and of three families with 11q linkage, resulted in one significant maximum LOD score (Zmax), 3.77, lying on the qter side, just beyond the USH1D critical region. In addition, a LOD of 1.03 was observed between markers D10S1743 and D10S1665, which is between the two previously defined critical regions of USH1D and USH1F. In addition, a Zmax of 2.89 also lies within the USH1D region. Interestingly, the analysis also provided a LOD score of 1.60 just pter of the putative region of USH1F (fig. 2). On the surface, these results do not support the hypothesis of an Usher gene in the USH1F region. However, the impact that heterogeneity due to linked genes has on LOD scores is not well understood; and it is conceivable that a mixture of families that are heterogeneous because of linked genes could yield a LOD-score distribution with maxima that are shifted from the true location of the genes actually responsible. We had expected to see bimodality in the heterogeneity χ2(1) but found only a single maximum corresponding to the presence of an USH1D locus (fig. 3). It may be that Usher If is not frequent and that there are too few families in our sample for the second maximum to be detected. An alternative explanation is that USH1F is incorrectly mapped and that it actually lies between markers D10S1743 and D10S1665 or that there is another Usher I gene lying between USH1D and USH1F.
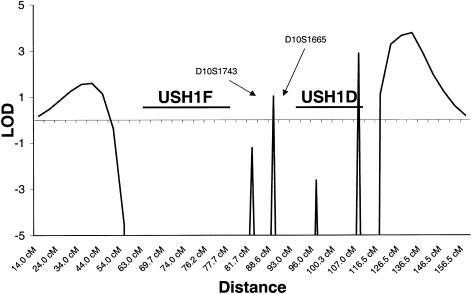
Figure 2 .
Map of multipoint LOD scores for chromosome 10 markers and the families with MYO7A− Usher I: results of chromosome 10 multipoint analysis for all families, with the exclusion of the three 11q-linked and two French Acadian families. A LOD score of 1.03 lies between markers D10S1743 and D10S1665 between the two defined regions of USH1D and USH1F, and a LOD score of 2.89 lies within the USH1D region. Additional LOD scores, of 1.60 and 3.77, lie outside the regions of USH1F and USH1D, respectively.
Figure 3 .
Plot of chromosome 10 HOMOG analysis. Rolling χ2 values generated from the chromosome 10 multipoint analyses (data from fig. 2) are plotted against position (in cM). A maximum χ2 of 40.44 lies in USH1D and a second, lesser χ2 value of 28.71 lies just outside the USH1F region.
These data provide an initial estimate of the proportion of the various Usher type I subtypes. Our original sample of 169 families with Usher syndrome was subdivided on the basis of the presence or absence of a MYO7A mutation. After the exclusion of the 18 families with Usher II and Usher III and of the 51 families with non–linkage-informative Usher I, the remaining sample consisted of 100 families with linkage-informative Usher I (table 2). Forty-six families, representing 46% of the group with linkage-informative Usher I, had a pathologic MYO7A mutation. The remaining 54 families with Usher I did not have an identified mutation. These families represented 54% of our sample. From our analysis of families with MYO7A− Usher I, the proportion of individuals with Usher type Ic in our sample is estimated at 2%. The estimate for Usher Ic is obviously biased, because the two families were purposefully included in the analyses. In addition, the three families with Usher 1b linked to 11q constitute ∼3% of the sample. Thus, in our sample, 49% of families with Usher I had a predicted or confirmed MYO7A mutation (i.e., Usher Ib). Of the remaining families, 33 lacking the MYO7A link to the USH1D and USH1F regions constitute 61% of the MYO7A− sample, or 33% of all families with linkage-informative Usher I. The remaining individuals with Usher I are unconfirmed for linkage to any of the six Usher I loci, and they constitute 16% of the sample.
The findings presented here have important implications for characterizing and defining the heterogeneity of Usher syndrome type I. These results support the genetic heterogeneity of Usher syndrome and also indicate for the first time that USH1D/USH1F is the second-most-common Usher I subtype.
The genetic heterogeneity of Usher syndrome type I is complex and raises the question of clinical differentiation. Our clinical findings have not yet suggested any obvious clinical variance between these subgroups. However, deeper and more-extensive examinations of the audiologic, vestibular, and retinal phenotypes may reveal differences that, because of heterogeneity, had not previously been noticed. Consequently, our next efforts will focus on the use of both clinical and genetic factors to differentiate, characterize, and further define the heterogeneity of the Usher I subtypes.
Acknowledgments
This work was supported in part by National Institute on Deafness and other Communicative Disorders (NIDCD) grants R01 DC00677-07 (to W.J.K.), P01 DC01813 (to W.J.K.), P60 DC00982 (to L.M.A.) and by a Foundation Fighting Blindness grant (to W.J.K.). The project was also supported by the Van Overbeek Foundation, De Drie Lichten, De Gelderse Blinden Verneniging (support to C.W.R.J.C. and M.W.), NIDCD grant R01 DC02842 (to R.J.H.S.), support by the Retinal Preservation Foundation of South Africa and a Foundation for Fighting Blindness in South Africa grant (to J.G.), National Eye Institute grant EY05627 (to S.G.J.), and by Research to Prevent Blindness and the Eye and Ear Foundation of Pittsburgh grants (to M.B.G.).
Electronic-Database Information
The accession number and URLs for data in this article are as follows:
- GeneClinics, http://www.geneclinics.com
- Genome Database, The, http://www.gdb.org
- Online Mendelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/Omim (for MYO7A [MIM 276903])
- Whitehead Institute for Biomedical Research/MIT Center for Genome Research, http://www.genome.wi.mit.edu
References
- Bitner-Glindzicz M, Lindley KJ, Rutland P, Blaydon D, Smith VV, Milla PJ, Hussain K, Furth-Lavi J, Cosgrove KE, Shepherd RM, Barnes PD, O'Brien RE, Farndon PA, Sowden J, Liu XZ, Scanlan MJ, Malcolm S, Dunne MJ, Aynsley-Green A, Glaser B (2000) A recessive contiguous gene deletion causing infantile hyperinsulinism, enteropathy and deafness identifies the Usher type 1C gene. J Med Genet 37:623–627 [DOI] [PubMed] [Google Scholar]
- Boughman JA, Vernon M, Shaver KA (1983) Usher syndrome: definition and estimate of prevalence from two high-risk populations. J Chronic Dis 36:595–603 [DOI] [PubMed]
- Chaib H, Kaplan J, Gerber S, Vincent C, Ayadi H, Slim R, Munnich A, Weissenbach J, Petit C (1997) A newly identified locus for Usher syndrome type I, USH1E, maps to chromosome 21q21. Hum Mol Genet 6:27–31 [DOI] [PubMed] [Google Scholar]
- Espinos C, Najera C, Millan JM, Ayuso C, Baiget M, Perez-Garrigues H, Rodrigo O, Vilela C, Beneyto M (1998) Linkage analysis in USH1 families from Spain. J Med Genet 35:391–398 [DOI] [PMC free article] [PubMed]
- Gasparini P, De Fazio A, Croce AI, Stanziale P, Zelante L (1998) Usher syndrome type III (USH3) linked to chromosome 3q in an Italian family. J Med Genet 35:666–667 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grondahl J (1987) Estimation of prognosis and prevalence of retinitis pigmentosa and Usher syndrome in Norway. Clin Genet 31:255–264 [DOI] [PubMed]
- Hallgren B (1959) Retinitis pigmentosa combined with congenital deafness, with vestibulo-cerebellar ataxia and neural abnormality in a proportion of cases. Acta Psychiatr Scand Suppl 138:1–101 [PubMed] [Google Scholar]
- Hmani M, Ghorbel A, Boulila-Elgaied A, Ben Zina Z, Kammoun W, Drira M, Chaabouni M, Petit C, Ayadi (1999) A novel locus for Usher syndrome type II, USH2B, maps to chromosome 3 at p23-24.2. Eur J Hum Genet 7:363–367 [DOI] [PubMed] [Google Scholar]
- Kaplan J, Gerber S, Bonneau D, Rozet JM, Delrieu O, Briard ML, Dollfus H, Ghazi I, Dufier JL, Frezal J (1992) A gene for Usher syndrome type I (USH1A) maps to chromosome 14q. Genomics 14:979–987 [DOI] [PubMed] [Google Scholar]
- Kimberling WJ, Moller C (1995) Clinical and molecular genetics of Usher syndrome. J Am Acad Audiol 6:63–72 [PubMed] [Google Scholar]
- Kimberling WJ, Moller CG, Davenport S, Priluck IA, Beighton PH, Greenberg J, Reardon W, Weston MD, Kenyon J (1992) Linkage of Usher syndrome type I gene (USH1B) to the long arm of chromosome 11. Genomics 14:988–994 [DOI] [PubMed] [Google Scholar]
- Kimberling WJ, Weston MD, Moller C, Davenport SL, Shugart YY, Priluck IA, Martini A, Milani M, Smith RJ (1990) Localization of Usher syndrome type II to chromosome 1q. Genomics 7:245–249 [DOI] [PubMed] [Google Scholar]
- Lathrop GM, Lalouel JM, Julier C, Ott J (1985) Multilocus linkage analysis in humans: detection of linkage and estimation of recombination. Am J Hum Genet 37:482–498 [PMC free article] [PubMed] [Google Scholar]
- Levy G, Levi-Acobas F, Blanchard S, Gerber S, Larget-Piet D, Chenal V, Liu XZ, Newton V, Steel KP, Brown SD, Munnich A, Kaplan J, Petit C, Weil D (1997) Myosin VIIA gene: heterogeneity of the mutations responsible for Usher syndrome type IB. Hum Mol Genet 6:111–116 [DOI] [PubMed] [Google Scholar]
- Liu XZ, Hope C, Walsh J, Newton V, Ke XM, Liang CY, Xu LR, Zhou JM, Trump D, Steel KP, Bundey S, Brown SD (1998) Mutations in the myosin VIIA gene cause a wide phenotypic spectrum, including atypical Usher syndrome. Am J Hum Genet 63:909–912 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu XZ, Walsh J, Mburu P, Kendrick-Jones J, Cope MJ, Steel KP, Brown SD (1997a) Mutations in the myosin VIIA gene cause non-syndromic recessive deafness. Nat Genet 16:188–190 [DOI] [PubMed] [Google Scholar]
- Liu XZ, Walsh J, Tamagawa Y, Kitamura K, Nishizawa M, Steel KP, Brown SDM (1997b) Autosomal dominant non-syndromic deafness caused by mutation in the myosin VIIA gene. Nat Genet 17:268–269 [DOI] [PubMed] [Google Scholar]
- Nuutila A (1970) Dystrophia retinae pigmentosa—dysacusis syndrome (DRD): a study of the Usher, or Hallgren, syndrome. J Genet Hum 18:57–88 [PubMed]
- Pieke-Dahl S, Moller CG, Kelley PM, Astuto LM, Cremers CW, Gorin MB, Kimberling WJ (2000) Genetic heterogeneity of Usher syndrome type II: localization to chromosome 5q. J Med Genet 37:256–262 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sankila EM, Pakarinen L, Kaariainen H, Aittomaki K, Karjalainen S, Sistonen P, de la Chapelle A (1995) Assignment of an Usher syndrome type III (USH3) gene to chromosome 3q. Hum Mol Genet 4:93–98 [DOI] [PubMed] [Google Scholar]
- Smith RJ, Berlin CI, Hejtmancik JF, Keats BJ, Kimberling WJ, Lewis RA, Moller CG, Pelias MZ, Tranebjaerg L (1994) Clinical diagnosis of the Usher syndromes: Usher Syndrome Consortium. Am J Med Genet 50:32–38 [DOI] [PubMed]
- Smith RJ, Lee EC, Kimberling WJ, Daiger SP, Pelias MZ, Keats BJ, Jay M, Bird A, Reardon W, Guest M (1992) Localization of two genes for Usher syndrome type I to chromosome 11. Genomics 14:995–1002 [DOI] [PubMed] [Google Scholar]
- Vernon, M (1969) Usher's syndrome: deafness and progressive blindness: clinical cases, prevention, theory and literature survey. J Chronic Dis 22:133–151 [DOI] [PubMed] [Google Scholar]
- Verpy E, Leibovici M, Zwaenepoel I, Liu XZ, Gal A, Salem N, Mansour A, Blanchard S, Kobayashi I, Keats BJ, Slim R, Petit C (2000) A defect in harmonin, a PDZ domain-containing protein expressed in the inner ear sensory hair cells, underlies Usher syndrome type 1C. Nat Genet 26:51–55 [DOI] [PubMed] [Google Scholar]
- Wayne S, Der Kaloustian V, Schloss M, Polomeno R, Scott DA, Hejtmancik JF, Sheffield VC, Smith RJ (1996) Localization of the Usher syndrome type ID gene (Ush1D) to chromosome 10. Hum Mol Genet 5:1689–1692 [DOI] [PubMed] [Google Scholar]
- Wayne S, Lowry RB, McLeod DR, Knaus R, Farr C, Smith RJH (1997) Localization of the Usher syndrome type IF to chromosome 10. Am J Hum Genet Suppl 61:A300 [Google Scholar]
- Weston MD, Kelley PM, Overbeck LD, Wagenaar M, Orten DJ, Hasson T, Chen ZY, Corey D, Mooseker M, Sumegi J, Cremers C, Moller C, Jacobson SG, Gorin MB, Kimberling WJ (1996) Myosin VIIa screening in 189 Usher syndrome type 1 patients. Am J Hum Genet 59:1074–1083 [PMC free article] [PubMed] [Google Scholar]