Abstract
More than 500 unrelated patients with neurofibromatosis type 1 (NF1) were screened for mutations in the NF1 gene. For each patient, the whole coding sequence and all splice sites were studied for aberrations, either by the protein truncation test (PTT), temperature-gradient gel electrophoresis (TGGE) of genomic PCR products, or, most often, by direct genomic sequencing (DGS) of all individual exons. A total of 301 sequence variants, including 278 bona fide pathogenic mutations, were identified. As many as 216 or 183 of the genuine mutations, comprising 179 or 161 different ones, can be considered novel when compared to the recent findings of Upadhyaya and Cooper, or to the NNFF mutation database. Mutation-detection efficiencies of the various screening methods were similar: 47.1% for PTT, 53.7% for TGGE, and 54.9% for DGS. Some 224 mutations (80.2%) yielded directly or indirectly premature termination codons. These mutations showed even distribution over the whole gene from exon 1 to exon 47. Of all sequence variants determined in our study, <20% represent C→T or G→A transitions within a CpG dinucleotide, and only six different mutations also occur in NF1 pseudogenes, with five being typical C→T transitions in a CpG. Thus, neither frequent deamination of 5-methylcytosines nor interchromosomal gene conversion may account for the high mutation rate of the NF1 gene. As opposed to the truncating mutations, the 28 (10.1%) missense or single-amino-acid-deletion mutations identified clustered in two distinct regions, the GAP-related domain (GRD) and an upstream gene segment comprising exons 11–17. The latter forms a so-called cysteine/serine-rich domain with three cysteine pairs suggestive of ATP binding, as well as three potential cAMP-dependent protein kinase (PKA) recognition sites obviously phosphorylated by PKA. Coincidence of mutated amino acids and those conserved between human and Drosophila strongly suggest significant functional relevance of this region, with major roles played by exons 12a and 15 and part of exon 16.
Introduction
Neurofibromatosis type 1 (NF1; also known as “von Recklinghausen neurofibromatosis” [MIM 162200]) is a common autosomal dominant disorder affecting ∼1 per 3,000–5,000 people. It is fully penetrant and exhibits a mutation rate some 10-fold higher than that reported for most other disease genes. As a consequence, a high number of sporadic cases (up to 50%) is observed (Upadhyaya and Cooper 1998). NF1 is clinically characterized by cutaneous neurofibromas, café-au-lait spots, iris hamartomas (Lisch nodules), and freckling of axillary and inguinal regions. These features are present in >90% of patients at puberty. Further manifestations, such as deeply situated and arbitrarily located neurofibromas, plexiform neurofibromas, optic glioma, macrocephaly, short stature, learning difficulties, scoliosis, and pseudarthrosis, as well as certain malignancies, occur less frequently among patients with NF1 (Riccardi 1992; Huson and Hughes 1994; Friedman and Birch 1997). NF1 is notable for its extreme phenotypic variability, both within and between families (Riccardi 1992). In addition to multiple allelism of the NF1 locus, the influence of modifying genes (Easton et al. 1993) and the impact of stochastic events (Riccardi et al. 1993) have been suggested as causes of this variability. So far, none of these hypotheses has been tested satisfactorily. A comprehensive analysis of the mutational spectrum might help to elucidate the role of different NF1 mutations.
The NF1 gene maps to chromosome 17q11.2 and is considered to be a tumor-suppressor gene, because loss-of-function mutations have been associated with the occurrence of benign and malignant tumors in neural-crest–derived tissues (Legius et al. 1993; Colman et al. 1995; Serra et al. 1997) as well as myeloid malignancies (Shannon et al. 1994). The NF1 gene was cloned in 1990 (White and O’Connell 1991). It spans over 350 kb of genomic DNA comprising ⩾60 exons. The open-reading frame codes for 2,867 amino acids, including three alternatively spliced exons. The most common transcript codes for a polypeptide of 2,818 amino acids called neurofibromin (Marchuk et al. 1991; Danglot et al. 1995; Li et al. 1995). A central region of about 360 amino acids shows significant homology to the mammalian GTPase-activating protein (GAP) (Xu et al. 1990). This region, the NF1 GAP-related domain (GRD), is encoded by the exons 20–27a and has been found to interact with p21ras (Martin et al. 1990). As a result of this interaction, GTP•Ras levels may be elevated in human NF1 peripheral nerve tumors, in which neurofibromin is usually found to be totally absent or, at least, dramatically reduced (Guha et al. 1996). Commensurate with its pivotal functional significance, several missense mutations were identified in the GRD (Li et al. 1992; Upadhyaya et al. 1997; Kim and Tamanoi 1998; Klose et al. 1998a).
In general, mutation analysis of the NF1 gene has turned out to be a major challenge, for several reasons. The large size of the gene under investigation and the diversity of the underlying pathological lesions (Korf 1998; Upadhyaya and Cooper 1998) make analysis problematic. In addition, the presence in the genome of several unprocessed NF1 pseudogenes severely hampers PCR approaches to the analysis of genomic DNA samples, because of coamplification of homologous sequences of the NF1-like loci at other chromosomes (Purandare et al. 1995; Régnier et al. 1997). Most of the NF1 mutations are point mutations or other small lesions leading to premature termination codons (PTCs) (Upadhyaya and Cooper 1998). Hence, the application of the protein truncation test (PTT) was thought to substantially accelerate the rate of identification of NF1 mutations. In an initial study, mutations were identified in 14 (67%) of 21 individuals (Heim et al. 1995). Before PTT was introduced, the best results had been obtained by using chemical mismatch cleavage analysis. In a study covering 70% of the coding sequence, seven mutations were found in 25 patients (28%) (Purandare et al. 1994). In a more recent study, by using a combined heteroduplex/SSCP approach, researchers identified only 13 disease-causing mutations in 67 unrelated NF1 patients (a 19% detection rate), even though as many as 45 exons were tested (Abernathy et al. 1997). Most other analyses focused on one or a few arbitrarily selected exons but sometimes were performed in very large cohorts of NF1 patients (e.g., Maynard et al. 1997).
Together with multiple reports of isolated mutations, most of these data have been submitted to the NF1 Genetic Analysis Consortium. The consortium was organized in the autumn of 1992 to facilitate mutation detection in the NF1 gene and to pool mutation data into a common database for better interpretation of the growing body of information. By November 1997 a total of 240 mutations, including 173 minor lesions (72%) had been reported to the consortium (Korf 1998). The mutational spectrum obtained from these data, however, suffers from a strong bias towards more thoroughly investigated regions of the NF1 gene. To overcome this drawback, we launched a mutation-screening project in Germany aimed at the complete analysis of the entire NF1 gene in a large cohort of unrelated NF1 patients. According to the preferences of the participating centers, PTT, temperature-gradient gel electrophoresis (TGGE) analysis (Riesner et al. 1989; Wartell et al. 1990), and direct genomic sequencing (DGS) of all individual exons were used for our analysis.
Materials and Methods
Patient Samples
The study was approved by the ethics committee of the Charité University Hospital. NF1 diagnostic criteria (Gutmann et al. 1997) were used to identify 521 unrelated patients with NF1, all of whom were of German or Turkish descent. Once informed consent was obtained, DNA from peripheral blood was obtained from each patient, applying one of several standard procedures. RNA was also obtained from some patients, either from fresh blood or from cell cultures (i.e., fibroblast-like cells or Epstein-Barr virus–transformed lymphocytes) by using the RNeasy total RNA purification kit (Qiagen). The SuperScript preamplification system (BRL Gibco) was used to prepare cDNA from the RNA samples. The cDNA reaction was performed in a 20-μl reaction volume containing 2–3 μg total RNA, 1.5 μg random hexamers (BRL Gibco), 0.5 μg single-stranded binding protein (Promega), 20 units RNAsin (Promega), 4 mM each dNTP, and 300 U Superscript II reverse transcriptase (RT) (BRL Gibco). cDNA was stored at -20°C.
Analysis of NF1 Gene Expression
Allele-specific expression level studies were performed by RT-PCR reactions by using the following primer pair to amplify the polymorphic exon 5 (Hoffmeyer and Assum 1994) from cDNA: 5′-TAGTCGCATTTCTACCAGGTTA-3′ (sense primer) and 5′-GAGAATGGCTTACTTGGATTAAA-3′ (antisense primer). Thirty PCR cycles of 1 min at 92°C, 1 min at 54°C, and 1 min at 72°C were used. After RsaI restriction, the fragments were analyzed by agarose gel electrophoresis and ethidium bromide staining. Subsequently, the band intensities were measured densitometrically by using a charge-coupled-device camera and the ImageQuant software package distributed with the Phosphoimager from Molecular Dynamics.
PTT
RT-PCR
Hot-start PCR reactions to amplify each of the five overlapping segments required that 2.5–3 μl cDNA be added to a lower-phase PCR mixture containing 1× PCR buffer (50 mM KCl, 10 mM Tris, pH 8.3, and 2 mM MgCl2), 0.8 mM each dNTP, and 10 pmol each primer (primer sequences were as described elsewhere (Heim et al. 1995) in a total volume of 40 μl. A wax gem (Perkin Elmer) was added to each tube. Reaction mixtures were heated to 80°C for 5 min, followed by cooling to room temperature. A 10-μl upper-phase mixture containing 1× PCR buffer, 0.25 μg single-stranded binding protein, and 2.5 units Taq polymerase (Pharmacia) was layered onto the solidified wax. Reactions were performed in a Cetus model 9600 thermal cycler (PE Biosystems), under the following conditions: 95°C for 1 min; 40 cycles of 95°C for 30 s, 61.5°C for 30 s, and 72°C for 90 s; and 72°C for 10 min. The PCR products were analyzed by agarose gel electrophoresis. Apparent splice defects, for example, exon skipping events, were verified by cDNA sequencing by using radioactive labeling and the Thermo Sequenase kit (Amersham Life Science). Subsequently, the presumably mutated exons or exon/intron boundaries were sequenced from the patients’ genomic DNA samples.
In vitro transcription/translation and analysis of the peptides
For each in vitro transcription/translation reaction, 4–9.5 μl PCR product (volume depends on the quality of the PCR product) and 0.5 μl 35S-methionine (specific activity: >1000 Ci/mmol; Amersham) were added to the commercially available TNTTM Coupled Reticulocyte Lysate System (Promega). Reactions were performed under conditions recommended by the manufacturer, with 1 exception: for all components, only a one-half volume was added to the PTT reaction (e.g., 12.5 μl TNT Rabbit Reticulocyte Lysate). After incubation of the complete reaction for 1.5 h at 30°C, 5 μl of the reaction was added to 5 μl loading buffer (10% glycerol, 5% 2-mercaptoethanol, 2% SDS, and 0.1% bromphenol blue), and was heated for 5 min, then placed on ice. Samples were subjected to electrophoresis in a 14% SDS-polyacrylamide gel for 1.5–2 h at 40 mA (Rainbow 14C methylated protein (Amersham) was used as a protein-weight marker). The gel was fixed in a solution of 40% methanol and 10% acetic acid and dried for 1.5 h at 80°C in a gel dryer. The synthesized polypeptides were visualized by means of a Phosphoimager (Molecular Dynamics) after the gel had been exposed overnight to a special film slide (Molecular Dynamics). On the basis of the determined fragment size, the positions of the mutations causing shortened polypeptides in the PTT were predicted and the presumably affected exons were sequenced from the patient DNA samples. Either the Thermo Sequenase kit (Amersham Life Science) and radioactive labeling were used for DNA sequencing or samples were analyzed on an ABI 377 PRISM DNA Sequencer by using the Big Dye terminator chemistry (see below).
TGGE
For TGGE analysis, the individual NF1 exons were usually amplified directly from genomic DNA. For several exons, however, preamplification of a larger genomic fragment was necessary (see table 1) to improve specificity—that is, either to avoid coamplification of NF1 pseudogenes or simply to relieve constraints of TGGE primer design. Final TGGE primers were chosen by using the programs MELT87 and SQHTX (Lerman and Silverstein 1987), which allow prediction of the impact of mutations on the melting behavior of the PCR products. Sequences containing two or more melting domains were split into several overlapping fragments. One or both oligonucleotides of each primer pair were 5′-modified with psoralen (table 2). Using two psoralen-modified primers in one reaction results in bipolar clamping of the fragments that may improve the sensitivity of mutation detection (Gille et al. 1998). For PCR, some 100 ng of genomic DNA (for the nested PCRs, 50 ng PCR product is required) were added to 25 μl standard Perkin Elmer PCR buffer containing 1.5 mM MgCl2, 200 μM each dNTP, 0.2 μM each primer, and 0.5 U Taq polymerase. All reactions were performed on a Perkin Elmer thermocycler (model T1), under the following conditions: 95°C for 3 min; 35 amplification cycles of 95°C for 30 s, specific annealing temperature Ta (seetable 2) for 30 s, and 72°C for 50 s (final extension for 10 min). A commercially available apparatus was used for TGGE (Diagen GmbH, Germany). Prior to electrophoresis, the PCR products were placed directly under a portable UV light source (365 nm) for 15 min to permit cross-linking of the psoralen moiety to the thymine residues of the opposite DNA strand. For exons 2–49, TGGE was done on an 8% polyacrylamide gel with 8 M urea in 3–N-morpholino–propanesulfonic acid (MOPS) buffer. For exon 1, an 8% polyacrylamide gel with 7 M urea and 35% formamide in MOPS buffer was used. The individual running conditions for each exon are given intable 2. Silver staining was used for visualization of bands. Samples that showed altered mobility patterns were subjected to DNA sequencing, as described below. Alternatively, radioactive labeling and the Thermo Sequenase kit (Amersham Life Science) were used for sequencing.
Table1.
Pre-Amplification PCR Primers and Conditions for Nested PCRs in the TGGE Approach
| Exon | Primer Sequencesa (5′→3′) | Fragment Size (bp) | Ta (°C)b |
| 1 | CAGACCCTCTCCTTGCCTCTT | 433 | 66 |
| GGATGGAGGGTCGGAGGCTG | |||
| 7 | TAATTTGCTATAATATTAGCTAC | 323 | 54 |
| ATAATACTTATGCTAGAAAATTC | |||
| 9 | AGAAATAATCTGCTTTTTTTTTTC | 1,341 | 59 |
| TACAATTTAACTCAGTGATACTC | |||
| 14 - 18 | CTCTGCCTAAGTTGTAGATAGTGAAG | 3,595 | 53→51 |
| TTAGGGAAATTTGAGAAGAGATGT | |||
| 23–1 | TTTGTATCATTCATTTTGTGTGTA | 282 | 56 |
| AAAAACAGCGGTTCTATGTGAAAAG | |||
| 23–2 | CTTAATGTCTGTATAAGAGTCTC | 268 | 56 |
| ACTTTAGATTAATAATGGTAATCTC | |||
| 23a | AGCCAGAAATAGTAGACATGATTGGGT | 446 | 56 |
| CTATTTTGTGCCAGAATTAGTAGA | |||
| 24 | TTGAACTCTTTGTTTTCATGTCTT | 266 | 58 |
| GGAATTTAAGATAGCTAGATTATC | |||
| 32–36 | TTGATTAGGCTGTTCCAATGAATA | 1,936 | 53→50 |
| TCATAAATATTTGGGAGAAGTGAGG |
Primers for the amplification of exons 1, 23-1, 23-2, 23a, and 24 are from Purandare et al. (1995).
Two different annealing temperatures (Ta) connected with an arrow indicate a touch-down PCR protocol with the first 8 cycles being performed at the higher temperature.
Table 2.
TGGE-PCR Primers and Conditions
| Exona | Primer Sequences (5′→3′)b | Fragment Size (bp) | Ta (°C)c | TGGE Conditionsd (gradient, running time × voltage) |
| 1 (nested) | TATTCTCTCCGTCGCCCCCC | 137 | 59 | 30–60°C, 2 h × 300 V |
| p-AACCCCCACTCCGCTCCC | ||||
| 2 | TAAGGATAAGCTGTTAACGTG | 221 | 55 | 25–55°C, 3 h × 300 V |
| p-AATTCCCCAAAACACAGTA | ||||
| 3 | p-AATGGTAGCAGAAAGTGAAAC | 175 | 58 | 35–60°C, 1 h 30 min × 300 V |
| AATTCACAAAGCCTGCCTAC | ||||
| 4a | TTGTTCTGTGTGTGTGTTTG | 316 | 56 | 32–53°C, 2 h 30 min × 300 V |
| p-A-ACTTCAGTAGTCCCATGTGG | (35–70°C, 5 h × 300 V)e | |||
| 4b | CTGTCCCCTAATACTTAATT | 209 | 52 | 29–52°C, 3 h 15 min × 300 V |
| p-A-ATACTAGTTTTTGACCCAGT | ||||
| 4c | p-AA-GCTCTGAGTTGTATTTGTGT | 190 | 51 | 27–53°C, 3 h 15 min × 300 V |
| ACAACAGCAAATTTTACATC | ||||
| 5 | GAAGGAAGTTAGAAGTTTGTG | 213 | 54 | 30–60°C, 1 h 30 min × 300 V |
| p-AA-CACAAGTAGGCATTTAAAAGA | ||||
| 6 | p-A-AATGCCAGGGATTTTGT | 247 | 47 | 35–60°C, 1 h 30 min × 300 V |
| p-AA-TAGATATAATGGAAATAATTTTG | ||||
| 7 (nested) | GAAACTTCATATATTATCTTA | 254 | 52 | 30–60°C, 2 h 30 min × 300 V |
| p-AA-CTTAAAGTTTGTAGTAGA | ||||
| 8 | p-AA-TTGTGTGGGTAATGTGTTGA | 248 | 52 | 30–60°C, 2 h 15 min × 300 V |
| AAATATAGTTAGATAAAAACCAATG | ||||
| 9 (nested) | p-AATCTGCTTTTTTTTTTCTTT | 136 | 51 | 30–60°C, 1 h 30 min × 300 V |
| p-AA-TTTAGTAATTTAGCAATAC | ||||
| AATCTGCTTTTTTTTTTCTTT | 136 | 51 | 30–60°C, 2 h × 300 V | |
| p-AA-TTTAGTAATTTAGCAATAC | ||||
| 9br | p-AATTGTGAAATATTTTTGTCTAC | 102 | 52 | 32–62°C, 55 min × 300 V |
| CTTGGAACCCAGAAAGGAA | ||||
| 10a | p-AATTTTGTACTTTTTCTTCC | 217 | 53 | 30–60°C, 2 h 10 min × 300 V |
| AATAGAAAGGAGGTGAGA | ||||
| 10b | p-A-ATTATCCTGAGTCTTACGTC | 229 | 51 | 30–70°C, 50 min × 300 V |
| p-AA-TAACTTAGTGTGATAATTT | ||||
| 10c | ATTTTTTTTAATTGAAGTTTC | 186 | 50 | 30–60°C, 2 h 30 min × 300 V |
| p-AA-TTCCTTCAAGAACATGGAA | ||||
| 11 | AAGTACTCCAGTGTTATGTTTAC | 205 | 52 | 20–45°C, 3 h × 300 V |
| p-A-ACAAAATAAAATTTAAAAGTTGAA | ||||
| 12a | CTAAGCTTCTCTAAACTTGTATTCA | 297 | 62 | 25–55°C, 3 h × 300 V |
| p-AATCTCTCACCATTACCATTCC | ||||
| 12b | GTGTTTATTCCTCTTGGTTGTC | 274 | 59 | 25–55°C, 3 h 30 min × 300 V |
| p-AA-ATTCAGAAAACAAACAGAGCAC | ||||
| 13 | CCCAAGTTGCAAATATATGTC | 277 | 60 | 35–65°C, 4 h 30 min × 300 V |
| p-A-ATTGCTGACAGAGGCAAACTC | ||||
| CTCTTGCCCAACTATAACAC | 89 | 58 | 35–65°C, 2 h 30 min × 300 V | |
| p-A-AGTAAAAAAAACCACTATTCAC | ||||
| 14 (nested) | p-AA-CCTACTCCTTTTGGGTGGAGCTTA | 191 | 55 | 31–61°C, 2 h 40 min × 400 V |
| GGGGTGTTTCTGTTGCTA | ||||
| 15 (nested) | p-AATATACATCAAGTTTGAAACTT | 208 | 55 | 28–60°C, 2 h 10 min × 400 V |
| AGTTAACAGACAAAAGTCAACTT | ||||
| 16 (nested) | p-AATGTTGGATAAAGCATAATTT | 330 | 55 | 31–61°C, 2 h 40 min × 400 V |
| p-AA-CGTTTCCCTCTGAAGACA | ||||
| p-AAGGGTTCTATGATTTCATGAT | 275 | 55 | 31–61°C, 2 h 40 min × 400 V | |
| p-AA-CACTCTATTCATAGAGAAAGGTG | ||||
| 17 (nested) | p-AA-TCATCTCTCTAGGGGGTCT | 240 | 55 | 31–61°C, 2 h 40 min × 400 V |
| ATGTTTCTGCTACATTTCAGTA | ||||
| GATGGCAAATCATTAATGTAT | 204 | 55 | 28–60°C, 2 h 10 min × 400 V | |
| p-AATGTTTCTGCTACATTTCAG | ||||
| 18 (nested) | p-AA-TGATAATTTTTTTATTGTTTCTATG | 230 | 55 | 31–61°C, 2 h 40 min × 400 V |
| TGTACTTCATTGGACATATTAAGATTTACA | ||||
| p-AA-TATTTAGAATGCCTTCTCTTTTG | 222 | 55 | 28–60°C, 2 h 10 min × 400 V | |
| TACAAGACCCTACATTGCTC | ||||
| 19a | GTGTGTGTGTGGCTTCAAAA | 192 | 60 | 30–55°C, 3 h 30 min × 300 V |
| p-AACTCTCACAGTAAAACCCACTAA | ||||
| 19b | TTTTTTTTCAGAGATTTGGACCA | 174 | 58 | 30–55°C, 3 h 3 min × 300 V |
| p-AA-GCTTTATTTGCTTTTTGCTTTA | ||||
| 20 | p-A-ATACGGCCTTCACTATGTAAAGGT | 276 | 58 | 30–60°C, 3 h × 300 V |
| GCCAGTTCTCTAGGTTTTGT | ||||
| 21 | p-AA-GCATGTAAGAGAAGCAAAAATTA | 328 | 55 | 35–60°C, 3 h × 300 V |
| CTATGTGCCAGGCACTTTTC | ||||
| 22 | TCTTTAGCTTCCTACCTAAGAA | 262 | 59 | 30–55°C, 4 h × 300 V |
| p-A-ACACACATACACACAAAATGAA | ||||
| 23-1 (nested) | p-AAGGTAAAATATATGGAGCAG | 191 | 63 | 25–55°C, 3 h × 300 V |
| AAAACAGCGGTTCTATGTG | ||||
| 23-2 (nested) | p-AAGTGTTAGGATTTTATTTTTATTTTTT | 93 | 59 | 35–65°C, 1 h 30 min × 300 V |
| TTTTCAGTCATCTGAAGGAGG | ||||
| GTTAGAACCATCAGAGAGC | 159 | 59 | 33–63°C, 3 h 30 min × 300 V | |
| p-AA-GTAATCTCTAACTGTAAGCAT | ||||
| 23a (nested) | GCTGTATGTAGTCGGTGCT | 78 | 58 | 33–63°C, 1 h 30 min × 300 V |
| p-AA-TTTATTCAGTAGGGAGTGGCA | ||||
| TGCTCATCTCTGTTCTGTA | 113 | 55→53 | 33–63°C, 1 h 15 min × 300 V | |
| p-AACAGCTAATAAAAAGTTCTCC | ||||
| 24 (nested) | AACTCCTTGTTTTTAGGTGG | 171 | 56 | 37–67°C, 2 h 30 min × 400 V |
| p-AA-GACATTAACTTCAAGCCC | ||||
| ACCTTTGAACTCTTTGTTTTCAT | 230 | 52 | 37–67°C, 2 h 30 min × 400 V | |
| p-AA-GACATTAACTTCAAGCCC | ||||
| p-AA-TTCAAACCTTATACTCAATTC | 238 | 48 | 30–60°C, 2 h 10 min × 400 V | |
| AAGGGGAATTTAAGATAGCTAG | ||||
| 25 | p-AACCCTGTTTTATTGTGTAG | 138 | 62 | 30–60°C, 5 h 20 min × 300 V |
| GTAAGTGGCAAGAAAATTAC | ||||
| 26 | TGAAAATTCTAATGACTTTGC | 212 | 64 | 30–60°C, 2 h 40 min × 300 V |
| p-A-AGTGTTCACTATCCCCATGAC | ||||
| 27a | TGTGTAGTGCTAAATGTG | 263 | 54 | 30–60°C, 2 h 30 min × 300 V |
| p-AA-GCAAACTCTCCTTCTCAAC | ||||
| 27b | p-AA-CTACTCAGTAGACAACAT | 195 | 55 | 25–55°C, 3 h × 300 V |
| GTTAAGAGACCCAAAACATAG | ||||
| 28 | TCTTTGTCTTTTTTGTCATTTTCC | 509 | 60 | SSCPd |
| AGTCAAGAAAAGCAATGAATCGT | ||||
| 29 | TCACCCCGTCACCACCACTTT | 411 | 60 | SSCPd |
| GCAACAACCCCAAATCAAACTGA | ||||
| 30 | p-AA-GGAACTATAAGGAAAAATACG | 181 | 57 | 25–55°C, 3 h 30 min × 300 V |
| ACTAATAGAGACAATAAAGAGGG | ||||
| GCCAGTTACTAGAGACAT | 182 | 50 | 25–55°C, 3 h 30 min × 300 V | |
| p-AATAATTGTTAGTATTAAAGAAAA | ||||
| 31 | p-A-ATTGACCATCACATGCTAATAG | 260 | 62 | 30–55°C, 2 h 30 min × 300 V |
| CTAGATAAATATTTGAGCAAACTC | ||||
| 32 (nested) | p-AA-TGATTAGGCTGTTCCAATGAATA | 156 | 48 | 30–64°C, 2 h 30 min × 400 V |
| p-AA-TGATCCCAAGCCACCTGTT | ||||
| p-AA-CTTGATGTTGTACTAGACAGTT | 156 | 48 | 30–64°C, 2 h 30 min × 400 V | |
| p-AA-TACAGAAGGCAAAAGAAAAGTGAT | ||||
| 33 (nested) | p-AATTAAACTGAACTTTTTTGTGCTA | 204 | 48 | 30–64°C, 2 h 30 min × 400 V |
| p-AA-GAAGGACAGCATCAGCATGTA | ||||
| p-A-ATGTGGGATGATATTGCTATTTTA | 202 | 48 | 30–64°C, 2 h 30 min × 400 V | |
| p-AATGAAGCTGTGAACAAGTACA | ||||
| p-A-ATGGACTGGTCATTAATATCATT | 114 | 48 | 30–64°C, 2 h 30 min × 400 V | |
| p-AA-GAAGTAAAATGGAGAAAGGAACT | ||||
| 34 (nested) | p-A-ATGGAACTTTAGAAATTAAAAAGTA | 187 | 48 | 30–64°C, 2 h 30 min × 400 V |
| p-AA-CCCGGTAACTGGAACGGAA | ||||
| p-AA-CAAAGTCAAGTCAGCTGCTGT | 179 | 48 | 30–64°C, 2 h 30 min × 400 V | |
| p-AAGATTTTAACTTAGTTTCTTATCA | ||||
| 35 (nested) | p-A-ATCATAATAAACATTATTTAAACAGTT | 168 | 48 | 30–60°C, 2 h 30 min × 400 V |
| p-AA-TTTACAACTTATATTTAATTTAGGA | ||||
| 36 (nested) | p-AAGGTTTTTATAAGTTCTGTGGAT | 189 | 48 | 30–60°C, 2 h 30 min × 400 V |
| p-AA-CATAAATATTTGGGAGAAGTGAGG | ||||
| 37 | TCCGAGATTCAGTTTAGGAGT | 203 | 51 | 25–55°C, 3 h × 300 V |
| p-AATGCACTCATTTTCTATACAGTA | ||||
| 38 | p-AA-TTGAAAGAGACTATGTCATGAT | 235 | 62 | 30–60°C, 2 h × 300 V |
| p-AA-GAGTAATCTAGGAACCTCAAG | ||||
| 39 | CCTTTAAAGAAAGCTACTG | 215 | 48 | 30–60°C, 2 h × 300 V |
| p-A-ATAAAAATATTCTAAATAAGGC | ||||
| 40 | p-A-AGATGCTTGTTCAAAAAATTA | 241 | 52 | 30–60°C, 4 h × 300 V |
| TATATATAGATGTAGCAAGAT | ||||
| AAGATGCTTGTTCAAAAAATTA | 241 | 52 | 30–60°C, 4 h × 300 V | |
| p-AA-TATATATAGATGTAGCAAGAT | ||||
| 41 | GAGACTGTAAGAAGTTCATC | 268 | 50 | 25–49°C, 2 h 30 min × 300 V |
| p-AATTGAAACTTGATTATGATTA | ||||
| 42 | p-AA-TAATTGATTTTTCTCTATTG | 230 | 52 | 30–60°C, 4 h 30 min × 300 V |
| AATAATAAAAAAAATCTACATAAT | ||||
| 43 | p-A-ATGTCCAAACATTTTCTTTTT | 251 | 53 | 30–60°C, 3 h 30 min × 300 V |
| CCACCTTATTTTCAATGATC | ||||
| 44 | p-AACATTGAAATAGTTAGGTGAA | 253 | 52 | 35–65°C, 2 h 45 min × 300 V |
| TAGACTGGAATAAAAATTTG | ||||
| 45 | p-AA-CCACAAAGTAAAAATGTTGT | 179 | 49 | 30–58°C, 2 h 15 min × 300 V |
| AAGGTGAATTAAAATCAAAA | ||||
| 46 | p-A-ATTTTCATTTAATTTTCCTCT | 246 | 48 | 30–55°C, 1 h 50 min × 300 V |
| ATGTTAGCAAGTTCATCAAC | ||||
| 47 | p-A-ATTTAATTTCTGTTACAATT | 143 | 44 | 26–52°C, 3 h × 300 V |
| CAAAAGTTAGAGAAAAATAT | ||||
| 48 | p-AACTAAAATAATTTCCTATTTTCC | 294 | 58 | 35–55°C, 3 h 45 min × 300 V |
| p-AA-GTCTTATATTGTTGCTCAAAGT | ||||
| 48a | p-AATCTTATGAACATCACTTACTT | 134 | 54 | 35–55°C, 2 h × 300 V |
| CGTGCAAAGATGATGAAAA | ||||
| 49 | CCTGGAAGGAAAAGAAGA | 216 | 56 | 35–60°C, 4 h × 300 V |
| p-AA-GATTTTAAAAAAGAAAGCAA |
For TGGE, exons were amplified directly from genomic DNA or, as indicated, from a preamplified PCR product (see table 1) using nested TGGE-PCR primers.
Psoralen modification is indicated by “p-”; note that psoralen is followed by two adenosine residues irrespective of the target sequence.
Ta is the annealing temperature for the primers during PCR.
The exons 28 and 29 were investigated by single-strand conformation polymorphism (SSCP) analysis in lieu of TGGE (see Peters et al. [1999b]).
A second TGGE analysis of exon 4a was performed after digesting the PCR product with Tru 9 I.
DGS
For DGS-PCR, samples were cycled with a first activating step for HotStarTaq (Qiagen) at 95°C for 3 min. Primer sequences, fragment sizes, and annealing temperatures are given in table 3. Each cycle consisted of a denaturation step at 95°C for 90 s, annealing for 40 s (48°C to 65°C, see table 3) and an extension step for 1 min at 72°C. 35 cycles were performed in a 20-μl reaction volume on a 9700 thermal cycler (PE Applied Biosystems). PCR products were purified by using the PCR Product Purification Kit (Qiagen) according to the manufacturer’s protocol. Cycle sequencing was performed in a 5-μl volume at 60°C with 25 cycles. Sequencing products were purified on Sephadex G-50-80 columns (Sigma) and analyzed on an ABI 377 automated DNA sequencer (PE Applied Biosystems).
Table 3.
DGS-PCR Primers and Conditions
| Exon | Primer Sequencesa (5′→3′) | Fragment Size (bp) | Ta(°C)b |
| 1 | CACAGACCCTCTCCTTGCCTCTTC | 233 | 66 |
| TACCTCCCCTCACCTACTCTGTCC | |||
| 2 | TCTGTGGTTGATGCAGTTTTCC | 371 | 62 |
| TATATCCAAAGTCCACAGAAAATC | |||
| 3 | TGGTAGCAGAAAGTGAAACTA | 224 | 56 |
| ATAGGACTGTCTCTGGTCCATC | |||
| 4a | GTTTGAAAATTTTCATAATAGAAAATG | 419 | 60 |
| GGAGGTCAAAGCTGCTGTGAG | |||
| 4b | GAACTCCTGGCCTCAAGTGGTC | 349 | 62 |
| ATGTCATTATAAAATCCAGTTTGGTG | |||
| 4c | GCAAAAGTAATACGTAAATGGAAAG | 904 | 62 |
| TGATGTACCCAAGCAACAAAGAC | |||
| 5 | (amplification as for 4c) | 904 | 62 |
| [TGACTTGAGTGATAGTTTCACAT] | |||
| 6 | CATGTTTATCTTTTAAAAATGTTGCC | C333 | 62 |
| TTGAGTAAAATAAACTGCTTCACAT | |||
| 7 | ATTTGCTATAATATTAGCTACATCTGG | 387 | 62 |
| AAAGCAAGTCCTATGAACTTATCAAC | |||
| 8 | GGATTTTACTGCCATTTGTGTG | 237 | 60 |
| TAACAGCATCAGTAAATATAGTTAGATA | |||
| 9 | CTTAGTGTTTTTTTTTTAQAACTTTCTA | 224 | 54 |
| ATTTTTTTAGTAATTTAGCAATACC | |||
| 9br | GCTTAAAATTTGTATACAATAAAC | 193 | 62 |
| ATGCATGAAGCACCACTCCAGG | |||
| 10a | GATAAAACTTAAAAQCTACAGTGATAAACAG | 270 | 62 |
| GCAATAGAAAGGAGGTGAGATTC | |||
| 10b | CTTTAAAGTGATAGCTATTACCTGAGTC | 288 | 60 |
| CTAAAAAGTATCCTCCAGGTCTTG | |||
| 10c | AAACTTGGTACCCTTTAGCAGTC | 348 | 60 |
| GTATAGACATAAACATACCATTTC | |||
| 11 | GTACTCCAGTGTTATGTTTACC | 268 | 64 |
| AAAACCTTTGGAAGTGTAAGTTTTAC | |||
| 12a | TGTATTCATTATGGGAGAATGCC | 267 | 64 |
| ATTACCATTCCAAATATTCTTCCA | |||
| 12b | AAGTTGGGGCATAGAGATTGAGAG | 363 | 66 |
| GATGAAATTTACCAAATTTCATTCAG | |||
| 13 | CACAGTTTATTGCATTGTTAG | 376 | 60 |
| GCCATGTGCTTTGAGGCAG | |||
| 14 | TCCTACTCCTTTTGGGTGGAG | 511 | 54 |
| TTAACAGATAAAAGTCAACTTTACAG | |||
| 15 | (amplification as for 14) | 511 | 54 |
| [TGTGATCAGGAATAGCTTTTG] | |||
| 16 | TGGATAAAGCATAATTTGTCAAGT | 628 | 60 |
| GGAAATTTTTAAAAACTGTGAGTACC | |||
| TCAGTGAACGTAAGGGTTCTATG | |||
| 17 | GGTGCACTTACTCTGTGTGTTTAG | 808 | 64 |
| GGGAAATTTGAGAAGAGATGTAGAG | |||
| 18 | (amplification as for 17) | 808 | 64 |
| [GCCATTCTTTACTGCACACAAAC] | |||
| 19a | GCCATTCTTTACTGCACACAAAC | 985 | 60 |
| AAACAAAAGTTTGACATCTCAAAAG | |||
| [TCATGTCACTTAGGTTATCTGG] | |||
| 19b | ATTTAAGGGGAAGTGAAAGAAC | 242 | 56 |
| TGGTGGGGGGCTTTATTTGC | |||
| 20 | CTATATCAGGTAAAATCATGTCCAAC | 758 | 62 |
| GATTTGCTATGTGCCAGGGAC | |||
| 21 | (amplification as for 20) | 758 | 62 |
| [GGTCTCATGCACTCCATAGGTG] | |||
| 22 | GCTACTCTTTAGCTTCCTACCTAAG | 518 | 60 |
| AAAAACAGCGGTTCTATGTGAAAAG | |||
| [CCTTAAAAGAAGACAATCAGCC] | |||
| 23-1 | (amplification as for 22) | 518 | 60 |
| [TTTGTATCATTCATTTTGTGTGTA] | |||
| 23-2 | GGCTTAATGTCTGTATAAGAGTCTC | 270 | 60 |
| ACTTTAGATTAATAATGGTAATCTC | |||
| 23a | AGCCAGAAATAGTATACATGATTGGGT | 448 | 60 |
| CTATTTTCTGCCAGAATTAGTAGA | |||
| 24 | AATAAGACAAGCTATGTCTTGACCTAG | 1023 | 60 |
| GAGTTTTTATGCAAAGTTTGACC | |||
| 25 | (amplification as for 24) | 1023 | 60 |
| [TTAAGTACTAGCAGAAATTATATCAATGAG] | |||
| 26 | GCTTTGTCTAATGTCAAGTCAC | 339 | 62 |
| TTAAACGGAGAGTGTTCACTATC | |||
| 27a | ATGGTCCTGAGGTCTTTTTG | 360 | 66 |
| CTAACAAGTGGCCTGGTGGC | |||
| 27b | TTGCTTTTAAAATATTTTTTCATTTTAG | 329 | 56 |
| ACCTCCTGTTAAGTCAACTGGG | |||
| 28 | TTTAAAAAATGAATCCAGACTTTGAAG | 696 | 62 |
| GCCTTACGTGACATTTTATACACCAC | |||
| [TGCAAAGCCATATGAAATTGTAGTG] | |||
| 29 | GTTGGTTTCTGGAGCCTTTTAG | 489 | 60 |
| AAATGGTCTCATTTTAAAAGCAAC | |||
| 30 | TTAAGGGGTATTTTGGTTTTACTG | 530 | 62 |
| AGGATACCACAGATATAAAATCAGAAG | |||
| 31 | TTTTTTCCCCGAATTCTTTATG | 429 | 56 |
| GTGAGTGTCTACATGCTTTCTGAAG | |||
| 32 | TGACAGGCCTGTAAATAAAATCTAG | 885 | 58 |
| TTTTGGTAATATTTCATGTCATTACTG | |||
| 33 | (amplification as for 32) | 885 | 58 |
| [TTTTGGTAATATTTCATGTCATTACTG] | |||
| 34 | GTTTGATTTAGGGAACATGAT | 794 | 62 |
| CTGCAATTAAAAGATCCACAGAAC | |||
| 35 | (amplification as for 34) | 794 | 62 |
| [CTGTGTTATTGGTAACAGGTCAC] | |||
| 36 | GCTGGACCAGTGGACAGAAC | 392 | 62 |
| TCATTGACCTCAAATTTAAACGTC | |||
| 37 | ATAGCATGAGAAATCATTCTAG | 439 | 62 |
| CAAGCGCTTGAGAACATACTATC | |||
| 38 | GTTCTCAGTCCAGCTAACAGTGTC | 530 | 60 |
| ATGTAAAAAATATGCATTCAAGTTTAC | |||
| 39 | TGCTTGACTGTCTTGCACCAG | 511 | 66 |
| CAAGTGATCCTCCTGCCTCAG | |||
| 40 | ATTCACATATGCATGTTTTACCTTC | 556 | 64 |
| GATAACAGAGAGCCACTGTAGTGTC | |||
| 41 | TGCTTATTAAATCTCTCTGTATATTTC | 436 | 64 |
| TTTCACTTACTCTTCCTAGGCCATC | |||
| 42 | CCTAGGGATACACCAAGAGTTTG | 428 | 62 |
| ACATGGAAAATTTTGATAATCCTG | |||
| 43 | CTCAGTGAAAGCTTAAACACTTTATG | 1144 | 60 |
| AAAGCTAAAATCACTATCACAATATCTC | |||
| 44 | (amplification as for 43) | 1144 | 60 |
| [CGAATAGTAATTCTCTATGATGTTTATG] | |||
| 45 | (amplification as for 43) | 1144 | 60 |
| [CCCTCAAATTTTTATTCCAGTC] | |||
| 46 | ATTGTAGAAAATTTGGAAAATGAAG | 856 | 58 |
| ATGAGAAACTTTTTATAAAAGTAACATATG | |||
| 47 | (amplification as for 46) | 856 | 58 |
| [TTTAGTTGCTTTGACACTCATTC] | |||
| 48 | AAGGAAGAAAAATAGTAAATTAAGTCC | 421 | 60 |
| AAGGAGCAAAATTTGCTATAAAC | |||
| 48a | ATTCAATAATTAAAACCAGATTCC | 327 | 64 |
| GGTGGCTTTACAAGTTCCTAAAG | |||
| 49 | TTCCTAGAATGTGTCCCCGTTG | 931 | 62 |
| ATATATGTGGTCGCACTTATTTTCCTG | |||
| [CCATGTTGTAATGCTGCACTTC] |
Primers used for sequencing but not for amplification are shown in square brackets.
Ta is the annealing temperature for the primers during PCR.
Results
Synopsis of Mutation Screening
We analyzed 521 unrelated patients with NF1 by means of PTT, TGGE, or DGS for mutations in the whole coding sequence and the splice sites of the NF1 gene. Twenty patients were included in both the PTT and the TGGE screening program. In total, we identified 301 sequence changes (table 4), 278 of which were considered pathogenic. The detection rates for the different methods were 40/85 (47.1%) for PTT, 65/121 (53.7%) for TGGE, and 184/335 (54.9%) for DGS. The mutations of 11 patients were identified independently by PTT as well as by TGGE. Among the 278 mutations identified, we observed 84 (30.2%) nonsense mutations and 140 (50.4%) frameshift mutations, comprising 78 deletions and 32 insertions of one or a few base pairs, as well as 30 out-of-frame exon skipping events. Thus, as many as 224 (80.6%) mutations caused, directly or indirectly, a premature termination codon (PTC). Another 25 (9.0%) of the mutations caused in-frame exon skipping. Furthermore, 28 (10.1%) missense mutations and one initiation codon mutation were also observed. Most mutations were found only once in any one patient. The total number of different mutations identified was 216. Among the recurrent mutations, the three most common were R1513X in exon 27a (7 patients), 499delTGTT in exon 4b (6 patients), and 6789delTTAC in exon 37 (five patients). Furthermore, we found that 216 of our mutations, of which 179 were different, were not listed in the most recent review in the field (Upadhyaya and Cooper 1998). Overall, the total number of different minor lesion mutations of the NF1 gene described to date well exceeds 276, because 97 other mutations of this type have been published before (Upadhyaya and Cooper 1998) and some more unpublished mutations have been reported to the International NF1 Genetic Analysis Consortium.
Table 4.
Summary of NF1 Mutations and Sequence Variants Detected with PTT, TGGE, and DGS
| Patienta | Location | Sequence Changeb | CpGc(CpNpG) | RNA Leveld | Protein Leveld,c | Mutation Type | Reference; NF1 Mutation Database No. |
| 2318 | Exon 1 | T2C | M1T; initiation at M68, M102 or M108 (?) | (Missense); faulty initiation | This report | ||
| 34 | Exon 1 | G26A | No | W9X | Nonsense | This report | |
| 650 | Exon 1 | G55T | E19X | Nonsense | This report | ||
| 1852 | Exon 2 | C147G | Y49X | Nonsense | This report | ||
| 348 | Exon 2 | C168T | Yes | S56S | Silent | This report | |
| 2732 | Exon 2 | C168T | Yes | S56S | Silent | This report | |
| 2938 | Intron 2 | 204+1G→T | Skip of exon 2 (?) | In frame; −48 aa | Splice site | This report | |
| 710 | Intron 2 | 204+2T→G | Skip of exon 2 (?) | In frame; −48 aa | Splice site | This report | |
| 1972 | Intron 2 | 205−1G→A | No | Skip of exon 3 (?) | In frame; −28 aa | Splice site | This report |
| 119 | Exon 3 | 220delG | aa 74; PTC 84 | 1-bp deletion | This report | ||
| 214 | Exon 3 | 227insA | aa 76; PTC 106 | 1-bp insertion | This report | ||
| 2170 | Exon 4a | 426delATTTT | aa 142−144; PTC 153 | 5-bp deletion | This report | ||
| 2253 | Exon 4b | T482A | L161X | Nonsense | This report | ||
| 420 | Exon 4b | 496delGT | aa 166; PTC 172 | 2-bp deletion | Toliat et al. (1999); 844 | ||
| 461 | Exon 4b | 499delTGTT | aa 167-168; PTC 176 | 4-bp deletion | This report | ||
| 1062 | Exon 4b | 499delTGTT | aa 167-168; PTC 176 | 4-bp deletion | This report | ||
| 110 | Exon 4b | 499delTGTT | aa 167-168; PTC 176 | 4-bp deletion | Toliat et al. (1999); 843 | ||
| U-36/375 | Exon 4b | 499delTGTT | aa 167-168; PTC 176 | 4-bp deletion | Toliat et al. (1999); 848 | ||
| E13795 | Exon 4b | 499delTGTT | aa 167-168; PTC 176 | 4-bp deletion | Toliat et al. (1999) | ||
| E13332 | Exon 4b | 499delTGTT | aa 167-168; PTC 176 | 4-bp deletion | Toliat et al. (1999) | ||
| 696 | Exon 4b | 527delA | aa 176; PTC 177 | 1-bp deletion | This report | ||
| 1899 | Exon 4b | T528A | D176E | (Missense); polymorphism | This report | ||
| 2406 | Exon 4b | T528A | D176E | (Missense); polymorphism | This report | ||
| 190 | Exon 4b | T528A | D176E | (Missense); polymorphism | Toliat et al. (1999); 545 | ||
| 702 | Exon 4b | T528A | D176E | (Missense); polymorphism | Toliat et al. (1999); 849 | ||
| 422 | Exon 4b | T539A | L180X | Nonsense | Toliat et al. (1999); 845 | ||
| NF58a/124 | Exon 4b | 540insA | Unequal expression | aa 180; PTC 200 | 1-bp insertion | Däschner et al. (1997); 97-003 | |
| 1559 | Exon 4b | C574T | Yes | R192X | Nonsense | This report | |
| 380 | Exon 4b | C574T | Yes | Unequal expression | R192X | Nonsense | Toliat et al. (1999); 847 |
| 1511 | Intron 4b | 586+1delG | Skip of exon 4b (?) | Out of frame; PTC 164 | 1-bp deletion; splice site | This report | |
| 3399 | Intron 4b | 586+1G→A | No | Skip of exon 4b (?) | Out of frame; PTC 164 | Splice site | This report |
| U-76 | Intron 4b | 586+1G→A | No | Skip of exon 4b; unequal expression | Out of frame; PTC 164 | Splice site | This report |
| 2395 | Exon 4c | T647C | L216P | Missense | This report | ||
| U-61 | Intron 4c | 655−2A→T | Skip of exon 5 | Out of frame; PTC 255 | Splice site | This report | |
| 314 | Intron 4c | 655−1G→A | No | Skip of exon 5 | Out of frame; PTC 255 | Splice site | Horn et al. (1996); 550 |
| 2048 | Exon 5 | 703delTA | aa 235; PTC 240 | 2-bp deletion | This report | ||
| 434 | Exon 5 | C715T | (Yes) | Q239X | Nonsense | Horn et al. (1996); 549 | |
| 3358 | Exon 6 | 754delT | aa 252; PTC 280 | 1-bp deletion | This report | ||
| 2880 | Exon 6 | G801A | No | W267X | Nonsense | This report | |
| 1862 | Exon 6 | 838delATAA | aa 280-281; PTC 293 | 4-bp deletion | This report | ||
| NF84a | Exon 6 | 887delA | Unequal expression | aa 296; PTC 316 | 1-bp deletion | This report | |
| 1008 | Intron 6 | 888+1G→A | No | Skip of exon 6 (?) | Out of frame; PTC 261 | Splice site | This report |
| 3080 | Intron 6 | 888+1G→A | No | Skip of exon 6 (?) | Out of frame; PTC 261 | Splice site | This report |
| 620 | Intron 6 | 889−2A→G | Skip of exon 7 | In frame; −58 aa | Splice site | Klose et al. (1999) | |
| U-15 | Exon 7 | C910T* | Yes | Skip of exon 7; unequal expression | R304X; in frame; −58 aa | (Nonsense); splice error | Hoffmeyer et al. (1998), 0156 |
| 2180 | Exon 7 | C910T* | Yes | Skip of exon 7 (?) | R304X; in frame; −58 aa | (Nonsense); splice error | This report |
| U-5 | Exon 7 | 918delT | No exon skipping | aa 306; PTC 316 | 1-bp deletion | Hoffmeyer et al. (1998), 97-012 | |
| 1815 | Exon 7 | 918delT | aa 306; PTC 316 | 1-bp deletion | This report | ||
| 1163 | Exon 7 | 955delAG | aa 319; PTC 328 | 2-bp deletion | This report | ||
| 3109 | Exon 7 | 1019delCT | aa 340; PTC 351 | 2-bp deletion | This report | ||
| 3293 | Exon 7 | 1019delCT | aa 340; PTC 351 | 2-bp deletion | This report | ||
| 737 | Exon 7 | 1019delCT | aa 340; PTC 351 | 2-bp deletion | This report | ||
| 750 | Exon 7 | A1060T | K354X | Nonsense | This report | ||
| 448 | Exon 7 | G1062A | No | Skip of exon 7; unequal expression | K354K; in frame; −58 aa | (Silent); splice site | This report |
| 2719 | Intron 7 | 1062+67T→C | ? | ? | Splice error ? | This report | |
| 2239 | Intron 7 | 1063−13G→A | No | ? | ? | Splice error ? | This report |
| 2837 | Exon 8 | T1070C | L357P | Missense | This report | ||
| 256 | Exon 8 | 1111insT | aa 371; PTC 378 | 1-bp insertion | This report | ||
| NF172 | Intron 8 | 1185+1G→A | No | Skip of exon 8; unequal expression | In frame;−41 aa | Splice site | Hoffmeyer et al. (1995); report w/o no. |
| 59 | Intron 8 | 1185+1G→T | Skip of exon 8 | In frame;−41 aa | Splice site | Horn et al. (1996); 551 | |
| 889 | Intron 8 | 1185+3insTAAA | Skip of exon 8 (?) | In frame;−41 aa | 4-bp insertion; splice site | This report | |
| 763 | Exon 9 | T1224G | Y408X | Nonsense | This report | ||
| U-51 | Exon 9 | C1246T | Yes | R416X | Nonsense | This report | |
| NF56 | Exon 9 | C1246T | Yes | Unequal expression | R416X | Nonsense | This report |
| 1994 | Exon 9 | C1246T | Yes | R416X | Nonsense | This report | |
| 2413 | Exon 9 | C1246T | Yes | R416X | Nonsense | This report | |
| 1023 | Exon 9 | 1255delA | aa 419; PTC 472 | 1-bp deletion | This report | ||
| 248 | Exon 10a | G1275A | No | W425X | Nonsense | This report | |
| U-65 | Exon 10a | C1318T | Yes | R440X | Nonsense | This report | |
| 1138 | Exon 10a | C1318T | Yes | R440X | Nonsense | This report | |
| 2349 | Exon 10a | C1318T | Yes | R440X | Nonsense | This report | |
| 1571 | Exon 10a | 1338delA | aa 446; PTC 472 | 1-bp deletion | This report | ||
| E11034 | Exon 10a | C1381T | Yes | R461X | Nonsense | This report | |
| U-57 | Exon 10b | 1398insT | aa 466; PTC 469 | 1-bp insertion | This report | ||
| 33 | Exon 10b | 1436insA | aa 479; PTC 490 | 1-bp insertion | This report | ||
| 2788 | Exon 10b | A1466G | Skip of the 3′ end of Exon 10b (?) | Y489C; out of frame; PTC 489 | (Missense); splice error | This report | |
| 250 | Exon 10b | A1472G | Y491C | Missense | This report | ||
| 936 | Exon 10b | A1472G | Y491C | Missense | This report | ||
| 1917 | Exon 10b | 1484delCC | aa 495; PTC 509 | 2-bp deletion | This report | ||
| 2859 | Exon 10b | 1519insT | aa 507; PTC 509 | 1-bp insertion | This report | ||
| 1288 | Intron 10b | 1527+1delG | Skip of exon 10b (?) | In frame; −45 aa | 1-bp deletion; splice site | This report | |
| 22 | Exon 10c | 1541delAG | aa 514; PTC 556 | 2-bp deletion | Robinson et al. (1996); 544 | ||
| 190 | Exon 10c | 1541delAG | No exon skipping | aa 514; PTC 556 | 2-bp deletion | Robinson et al. (1996); 545 | |
| 535 | Exon 10c | 1541delAG | aa 514; PTC 556 | 2-bp deletion | This report | ||
| 1169 | Exon 10c | 1546delC | aa 516; PTC 525 | 1-bp deletion | This report | ||
| 2243 | Intron 10c | 1641+1G→T | Skip of exon 10c (?) | In frame; −38 aa | Splice site | This report | |
| 3918 | Intron 10c | 1642−8A→G | Exon 11 enlarged by 7 bp (?) | Out of frame; PTC 559 (?) | Splice error | This report | |
| 1003 | Exon 11 | T1646C | L549P | Missense | This report | ||
| 279 | Exon 11 | G1721C | Skip of exon 11 (?) | S574T; out of frame; PTC 560 | (Missense); splice site | This report | |
| 945 | Exon 11 | G1721A | No | Skip of exon 11 (?) | S574N; out of frame; PTC 560 | (Missense); splice site | This report |
| 1070 | Intron 11 | 1721+3A→G | Skip of exon 11 (?) | Out of frame; PTC 560 | Splice site | This report | |
| 1377 | Intron 11 | 1721+3A→G | Skip of exon 11 (?) | Out of frame; PTC 560 | Splice site | This report | |
| 3855 | Exon 12a | T1742C | 1581T | Missense | This report | ||
| 1584 | Exon 12a | A1748G | K583R | Missense | This report | ||
| 1700 | Exon 12a | A1748G | K583R | Missense | This report | ||
| 3853 | Exon 12a | 1756delACTA | aa 586-587; PTC 603 | 4-bp deletion | This report | ||
| 2869 | Exon 12a | 1817insT | aa 606; PTC 609 | 1-bp insertion | This report | ||
| 753 | Intron 12a | 1845+1delGTAAG | Skip of exon 12a (?) | Out of frame; PTC 589 | 5-bp insertion; splice site | This report | |
| 2273 | Exon 12b | 1935delG | aa 645; PTC 687 | 1-bp deletion | This report | ||
| 2760 | Exon 12b | C1994T | No | S665F | Missense | This report | |
| NF92 | Exon 12b | 1998insCCTCT | aa 666; PTC 689 | 5-bp insertion | Böddrich et al. (1995); 546 | ||
| 2889 | Exon 13 | 2027delC | aa 676; PTC 687 | 1-bp deletion | This report | ||
| 3259 | Exon 13 | 2027insC | aa 676; PTC 699 | 1-bp insertion | This report | ||
| NF176 | Exon 13 | 2033insC | Unequal expression | aa 678; PTC 699 | 1-bp insertion | 97-014 | |
| 212 | Exon 13 | C2041T* | Yes | R681X | Nonsense | This report | |
| 666 | Exon 13 | C2041T* | Yes | R681X | Nonsense | This report | |
| 2052 | Exon 13 | C2041T* | Yes | R681X | Nonsense | This report | |
| 3806 | Exon 13 | C2041T* | Yes | R681X | Nonsense | This report | |
| 2296 | Exon 13 | C2076G | Y692X | Nonsense | This report | ||
| 1982 | Exon 13 | T2084C | L695P | Missense | This report | ||
| 4132 | Exon 13 | 2190delCCTCT | aa 730-732; PTC 734 | 5-bp deletion | This report | ||
| 520 | Intron 13 | 2252−31A→T | Polymorphism | This report | |||
| E11225 | Intron 13 | 2252−31A→T | No exon skipping | Polymorphism | This report | ||
| E13339 | Intron 13 | 2252−31A→T | Polymorphism | This report | |||
| 168 | Exon 14 | 2272delAG | aa 758; PTC 766 | 2-bp deletion | 552 | ||
| 173 | Exon 14 | T2288C | L763P | Missense | 553 | ||
| 66 | Exon 15 | G2330C | W777S | Missense | This report | ||
| 640 | Exon 15 | C2339A | T780K | Missense | 554 | ||
| 1137 | Exon 15 | A2342C | H781P | Missense | This report | ||
| U-55 | Exon 15 | C2356T | No | Q786X | Nonsense | This report | |
| 1624 | Exon 16 | 2427insGTCTT/2430delG | aa 810; PTC 815 | Insertion/deletion | This report | ||
| U-62 | Exon 16 | C2446T | Yes | No exon skipping; equal expression | R816X | Nonsense | This report |
| 1551 | Exon 16 | C2446T | Yes | R816X | Nonsense | This report | |
| 2746 | Exon 16 | C2446T | Yes | R816X | Nonsense | This report | |
| 942 | Exon 16 | T2540C | L847P | Missense | This report | ||
| NF213 | Exon 16 | 2590insTATA | Unequal expression | aa 864; PTC 865 | 4-bp insertion | Report w/o no. | |
| 2332 | Exon 16 | 2666delC | aa 889; PTC 901 | 1-bp deletion | This report | ||
| 633 | Exon 16 | 2674delA | aa 892; PTC 901 | 1-bp deletion | This report | ||
| 626 | Exon 16 | C2842T | No | Q948X | Nonsense | This report | |
| 1031 | Exon 16 | 2844delA | aa 948; PTC 953 | 1-bp deletion | This report | ||
| 3212 | Exon 16 | 2845insT | aa 949; PTC 955 | 1-bp insertion | This report | ||
| 2803 | Exon 16 | 2850insTT | aa 950; PTC 954 | 2-bp insertion | This report | ||
| 1742 | Exon 17 | 2970delAAT | aa 990-991; 991delM | 3-bp deletion | This report | ||
| E13563 | Exon 17 | 2970delAAT | aa 990-991; 991delM | 3-bp deletion | This report | ||
| NF183 | Exon 17 | 2972insT | Unequal expression | aa 991; PTC 1020 | 1-bp insertion | 97-013 | |
| U-88 | Intron 17 | 2991−2A→G | Skip of exon 18; equal expression | In frame; −41 aa | Splice site | This report | |
| 646 | Intron 17 | 2991−1G→A | No | Skip of exon 18; equal expression | In frame; −41 aa | Splice error | This report |
| U-27/507 | Intron 17 | 2991−1G→C | Skip of exon 18; equal expression | In frame; −41 aa | Splice site | This report | |
| 3365 | Exon 18 | T2994A | Y998X | Nonsense | This report | ||
| 858 | Exon 18 | 3060delA | aa 1020; PTC 1021 | 1-bp deletion | This report | ||
| 223 | Exon 19a | 3178delG | aa 1060; PTC 1061 | 1-bp deletion | This report | ||
| NF23 | Exon 19a | 3193insA | aa 1065; PTC 1087 | 1-bp insertion | Klose et al. (1998c); 440 | ||
| 1572 | Exon 20 | 3394insAG | aa 1132; PTC 1142 | 2-bp insertion | This report | ||
| 3400 | Exon 20 | 3456delACTC | aa 1152-1153; PTC 1156 | 4-bp deletion | This report | ||
| 255 | Exon 20 | 3456delACTC | aa 1152-1153; PTC 1156 | 4-bp deletion | This report | ||
| 460 | Exon 20 | 3456delACTC | aa 1152-1153; PTC 1156 | 4-bp deletion | This report | ||
| 3809 | Exon 20 | A3467G* | N1156S | Missense | This report | ||
| 520 | Intron 20 | 3496+2T→C | Skip of exon 20 | Out of frame; PTC 1133 | Splice site | Klose et al. (1998c); 444 | |
| 1747 | Exon 21 | 3525delAA | aa 1175-1176; PTC 1193 | 2-bp deletion | This report | ||
| 2861 | Exon 21 | 3525delAA | aa 1175-1176; PTC 1193 | 2-bp deletion | This report | ||
| 3340 | Exon 21 | G3628T | E1210X | Nonsense | This report | ||
| 878 | Exon 21 | 3643delATG | 1215delM | 3-bp deletion | This report | ||
| 2340 | Exon 21 | G3707A | No | W1236X | Nonsense | This report | |
| 742 | Exon 22 | C3721T* | Yes | No exon skipping | R1241X | Nonsense | This report |
| 2266 | Exon 22 | 3737delTGTT | aa 1246-1247; PTC 1264 | 4-bp deletion | This report | ||
| 1528 | Exon 22 | G3749C | R1250P | Missense | This report | ||
| 76 | Exon 22 | G3773A | (Yes) | W1258X | Nonsense | This report | |
| 51 | Exon 22 | 3822delCT | aa 1274-1275; PTC 1282 | 2-bp deletion | This report | ||
| 1186 | Exon 22 | C3826T* | Yes | R1276X | Nonsense | This report | |
| 157 | Exon 22 | C3826T* | Yes | R1276X | Nonsense | Klose et al. (1998b); 441 | |
| U-63 | Exon 22 | C3826T* | Yes | R1276X | Nonsense | This report | |
| 1899 | Exon 22 | G3827A | Yes | R1276Q | Missense | This report | |
| 1939 | Exon 22 | G3827A | Yes | R1276Q | Missense | This report | |
| 364 | Exon 22 | G3827C | No exon skipping | R1276P | Missense | Klose et al. (1998a); 442 | |
| 288 | Exon 22 | C3831T | No | G1277G | Silent | This report | |
| 2235 | Exon 23-1 | 3909dclT | aa 1303; PTC 1308 | 1-bp deletion | This report | ||
| 578 | Exon 23-1 | 3911dclT | aa 1304; PTC 1308 | 1-bp deletion | This report | ||
| 258 | Exon 23-1 | C3916T | Yes | “CpG, RNA editing site” | R1306X | Nonsense | This report |
| 2601 | Exon 23-1 | C3916T | Yes | “CpG, RNA editing site” | R1306X | Nonsense | This report |
| 71 | Exon 23-1 | C3916T | Yes | “CpG, RNA editing site” | R1306X | Nonsense | This report |
| 3979 | Exon 23-1 | C3916T | Yes | “CpG, RNA editing site” | R1306X | Nonsense | This report |
| 82 | Intron 23-1 | 3975−2A→G | Skip of exon 23-2; equal expression | Out of frame; PTC 1339 | Splice site | This report | |
| 796 | Exon 23-2 | C4006T | (Yes) | Q1336X | Nonsense | This report | |
| 471 | Exon 23-2 | 4016delT | aa 1339; PTC 1342 | 1-bp deletion | This report | ||
| 206 | Exon 23-2 | C4084T | Yes | No exon skipping | R1362X | Nonsense | This report |
| 415 | Exon 23-2 | C4084T | Yes | R1362X | Nonsense | This report | |
| E13562 | Intron 23-2 | 4110+1G→C | Skip of exon 23-2; unequal expression | Out of frame; PTC 1339 | Splice site | This report | |
| U-19 | Exon 24 | G4243T | E1415X | Nonsense | This report | ||
| 1765 | Exon 24 | 4247ins74-bp from intron 25 | aa 1416;PTC 1422 | 74-bp insertion | This report | ||
| 895 | Exon 25 | T4274C | L1425P | Missense | This report | ||
| 219 | Exon 25 | T4274C | L1425P | Missense | Peters et al. (1999a) | ||
| 278 | Exon 25 | 4311delAGAA | aa 1437-1438; PTC 1446 | 4-bp deletion | This report | ||
| 238 | Intron 25 | 4368−46G→C | rare variant | This report | |||
| 333 | Intron 25 | 4368−1G→T | Skip of exon 26 | In frame; −49 aa | Splice site | This report | |
| 3745 | Exon 26 | 4374insT | aa 1459; PTC 1460 | 1-bp insertion | This report | ||
| 1517 | Exon 26 | 4431delC | aa 1477; PTC 1478 | 1-bp deletion | This report | ||
| U-77 | Exon 26 | G4473A | No | W1491X | Nonsense | This report | |
| 31 | Exon 26 | 4486delA | aa 1496; PTC 1552 | 1-bp deletion | This report | ||
| U-66 | Exon 26 | 4497insG | aa 1499; PTC 1508 | 1-bp insertion | This report | ||
| 831 | Intron 26 | 4514+1G→A | No | Skip of exon 26 (?) | In frame; −49 aa | Splice site | This report |
| 2928 | Intron 26 | 4515−2A→G | Skip of exon 27a (?) | In frame; −49 aa | Splice site | This report | |
| U-39 | Exon 27a | C4537T* | Yes | R1513X | Nonsense | This report | |
| U-40 | Exon 27a | C4537T* | Yes | R1513X | Nonsense | This report | |
| 401 | Exon 27a | C4537T* | Yes | R1513X | Nonsense | This report | |
| 740 | Exon 27a | C4537T* | Yes | R1513X | Nonsense | This report | |
| 2488 | Exon 27a | C4537T* | Yes | R1513X | Nonsense | This report | |
| 2713 | Exon 27a | C4537T* | Yes | R1513X | Nonsense | This report | |
| 2789 | Exon 27a | C4537T* | Yes | R1513X | Nonsense | This report | |
| U-87a | Exon 27a | G4614A | No | W1538X | Nonsense | This report | |
| 966 | Exon 27a | 4649insG | aa 1550; PTC 1554 | 1-bp insertion | This report | ||
| U-29 | Exon 27b | 4703delC | aa 1558; PTC 1569 | 1-bp deletion | This report | ||
| 1784 | Exon 27b | C4719G | Y1573X | Nonsense | This report | ||
| 2646 | Exon 27b | A4750G | 11584V | Missense | This report | ||
| 486 | Exon 28 | T4839G | No exon skipping; unequal expression | Y1613X | Nonsense | Peters et al. (1999b); 97-001 | |
| 3696 | Exon 28 | 4936insT | aa 1646; PTC 1660 | 1-bp insertion | This report | ||
| 56 | Exon 28 | 5050delAGGCTTG | aa 1684-1686; PTC 1686 | 7-bp deletion | This report | ||
| 102 | Exon 28 | 5055insT | aa 1686; PTC 1696 | 1-bp insertion | Peters et al. (1999b); 450 | ||
| 345 | Exon 28 | A5106G | Q1702Q | Silent | Peters et al. (1999b) | ||
| 734 | Exon 28 | 5152delG | aa 1718; PTC 1725 | 1-bp deletion | This report | ||
| B-212 | Exon 28 | 5168delTC | aa 1723; PTC 1734 | 2-bp deletion | Peters et al. (1999b); 842 | ||
| 342 | Exon 28 | G5172A | No | K1724K | Silent | Peters et al. (1999b) | |
| 186 | Exon 28 | G5172A | No | K1724K | Silent | Peters et al. (1999b) | |
| B-250 | Exon 28 | 5205delAGTAA | Skip of exon 28 (?) | Out of frame; PTC 1599 | 5-bp deletion; splice site | Peters et al. (1999b); 97-002 | |
| 1218 | Exon 29 | C5242T | Yes | R1748X | Nonsense | This report | |
| 2886 | Exon 29 | C5242T | Yes | R1748X | Nonsense | This report | |
| U-34/40 | Exon 29 | C5242T | Yes | No exon skipping | R1748X | Nonsense | Peters et al. (1999b); 97-011 |
| 68 | Exon 29 | 5248delAAA | 1750delK | 3-bp deletion | This report | ||
| 1079 | Exon 29 | T5286G | Y1762X | Nonsense | This report | ||
| 388 | Exon 29 | C5329T | (Yes) | Q1777X | Nonsense | This report | |
| 528 | Exon 29 | T5339A | L1780X | Nonsense | This report | ||
| 1698 | Exon 29 | T5339A | L1780X | Nonsense | This report | ||
| 3443 | Exon 29 | C5353T | (Yes) | Q1785X | Nonsense | This report | |
| 1038 | Exon 29 | 5399delT | aa 1800; PTC 1841 | 1-bp deletion | This report | ||
| 63 | Exon 29 | C5458T | No | Q1820X | Nonsense | Peters et al. (1999b); 97-010 | |
| 3344 | Exon 29 | 5484delT | aa 1828; PTC 1841 | 1-bp deletion | This report | ||
| U-25/702 | Exon 29 | G5546A | Yes | Skip of exon 29 | R1849Q; Out of frame; PTC 1740 | (Missense); splice site | This report |
| 1990 | Exon 29 | G5546A | Yes | Skip of exon 29 (?) | R1849Q; Out of frame; PTC 1740 | (Missense); splice site | This report |
| 2191 | Intron 29 | 5546+1G→A | (Yes) | Skip of exon 29 (?) | Out of frame; 1740 | Splice site | This report |
| NF113 | Intron 29 | 5546+2T→G | Skip of exon 29; unequal expression | Out of frame; PTC 1740 | Splice site | report w/o no. | |
| 1164 | Exon 30 | 5584delAC | aa 1862; PTC 1863 | 2-bp deletion | This report | ||
| 527 | Exon 30 | 5592delTTTAA | aa 1864-1866; PTC 1889 | 5-bp deletion | Harder et al. ((1999)); 439 | ||
| 224 | Exon 31 | C5839T | Yes | R1947X | Nonsense | Klose et al. (1999); 437 | |
| 1197 | Exon 31 | C5839T | Yes | R1947X | Nonsense | This report | |
| 2672 | Exon 31 | C5839T | Yes | R1947X | Nonsense | This report | |
| 394 | Exon 31 | 5847delAG | aa 1949-1950; PTC 1954 | 2-bp deletion | This report | ||
| 2097 | Intron 31 | 5943+1G→A | No | Skip of exon 31 (?) | Out of frame; PTC 1922 | Splice site | This report |
| 472 | Intron 31 | 5944−5A→G | Skip of exon 32 (?) | In frame; −47 aa | Splice site | This report | |
| 1215 | Intron 31 | 5944−2A→G | Skip of exon 32 (?) | In frame; −47 aa | Splice site | This report | |
| 2369 | Exon 33 | 6220delG | aa 2074; PTC 2089 | 1-bp deletion | This report | ||
| 2774 | Exon 34 | 6468delC | aa 2156; PTC 2178 | 1-bp deletion | This report | ||
| 608 | Exon 34 | 6470delT | aa 2157; PTC 2178 | 1-bp deletion | This report | ||
| 2912 | Exon 34 | 6471delC | aa 2157; PTC 2178 | 1-bp deletion | This report | ||
| 16 | Exon 34 | T6566A | L2189X | Nonsense | This report | ||
| U-64 | Intron 34 | 6579+2T→G | Skip of exon 34; unequal expression | Out of frame; PTC 2148 | Splice site | This report | |
| 1126 | Intron 34 | 6579+45T→A | ? | ? | Splice error ? | This report | |
| 514 | Intron 34 | 6579+87G→A | No | ? | ? | Splice error ? | This report |
| 2086 | Exon 35 | 6604delT | aa 2202; PTC 2211 | 1-bp deletion | This report | ||
| 234 | Exon 35 | G6628T | E2210X | Nonsense | This report | ||
| 3568 | Intron 35 | 6641+2delT | Skip of exon 35 (?) | Out of frame; PTC 2199 | 1-bp deletion; splice site | This report | |
| 2406 | Intron 35 | 6642-1G→T | Skip of exon 36 (?) | Out of frame; PTC 2220 | Splice site | This report | |
| 1832 | Exon 36 | C6709T | Yes | R2237X | Nonsense | This report | |
| U-67/572 | Exon 36 | C6709T | Yes | R2237X | Nonsense | This report | |
| 3634 | Exon 36 | C6709T | Yes | R2237X | Nonsense | This report | |
| 952 | Intron 36 | 6756+1G→A | No | Skip of exon 36 (?) | Out of frame; PTC 2220 | Splice site | This report |
| NF33 | Exon 37 | 6789delTTAC | aa 2263-2264; PTC 2268 | 4-bp deletion | Hoffmeyer et al. (1998) | ||
| U116 | Exon 37 | 6789delTTAC | No exon skipping | aa 2263-2264; PTC 2268 | 4-bp deletion | Robinson et al (1995); 543 | |
| 342 | Exon 37 | 6789delTTAC | No exon skipping; equal expression | aa 2263-2264; PTC 2268 | 4-bp deletion | Böddrich et al. (1997); 547 | |
| 1706 | Exon 37 | 6789delTTAC | aa 2263-2264; PTC 2268 | 4-bp deletion | This report | ||
| 3467 | Exon 37 | 6789delTTAC | aa 2263-2264; PTC 2268 | 4-bp deletion | This report | ||
| 407 | Exon 37 | 6790insTT | No exon skipping; equal expression | aa 2264; PTC 2270 | 2-bp insertion | Böddrich et al. (1997); 548 | |
| 1354 | Exon 37 | 6791insA | aa 2264; PTC 2285 | 1-bp insertion | This report | ||
| 25 | Exon 37 | C6792A | Skip of exon 37; equal expression | Y2264X; in frame; −34 aa | (Nonsense); splice error | Robinson et al. (1995); 541 | |
| 1 | Exon 37 | C6792A | Skip of exon 37; equal expression | Y2264X; in frame; −34 aa | (Nonsense); splice error | Robinson et al. (1995); 542 | |
| E11225 | Exon 37 | C6792A | Skip of exon 37; equal expression | Y2264X; in frame; −34 aa | (Nonsense); splice error | This report | |
| 1828 | Exon 37 | C6792G | Skip of exon 37 (?) | Y2264X; in frame; −34 aa | (Nonsense); splice error | This report | |
| 2472 | Exon 37 | 6792insA | aa 2265; PTC 2285 | 1-bp insertion | This report | ||
| U-43/547 | Exon 37 | 6797delGT | No exon skipping | aa 2266; PTC 2284 | 2-bp deletion | This report | |
| 1920 | Exon 37 | T6839G | L2280X | Nonsense | This report | ||
| 3110 | Exon 39 | 7080insA | aa 2361; PTC 2364 | 1-bp insertion | This report | ||
| 3578 | Exon 39 | 7095delT | aa 2365; PTC 2374 | 1-bp deletion | This report | ||
| E15470 | Exon 40 | 7208delGA | aa 2403; PTC 2405 | 2-bp deletion | This report | ||
| 260 | Exon 40 | C7237T | (Yes) | Q2413X | Nonsense | This report | |
| 61 | Exon 40 | G7258C | Skip of exon 40 (?) | A2420P; in frame; −44 aa | (Missense); splice site | This report | |
| B-214 | Intron 40 | 7259−17C→T | No | ? | ? | Splice error? | 841 |
| B-212 | Intron 40 | 7259−14C→T | Yes | rare variant | 842 | ||
| 400 | Exon 41 | 7268delCA | aa 2423; PTC 2425 | 2-bp deletion | This report | ||
| 3541 | Exon 41 | 7285delC | aa 2429; PTC 2434 | 1-bp deletion | This report | ||
| 1777 | Exon 41 | C7285T | Yes | R2429X | Nonsense | This report | |
| E14377 | Exon 41 | C7285T | Yes | R2429X | Nonsense | This report | |
| E14071 | Exon 41 | 7367delCC | aa 2456; PTC 2460 | 2-bp deletion | This report | ||
| 395 | Intron 41 | 7395−1G→A | (Yes) | Skip of exon 42 (?) | Out of frame; PTC 2471 | Splice site | 850 |
| U-20/9723 | Exon 42 | CT7424AG | S2475X | Nonsense | This report | ||
| 1000 | Exon 42 | C7457T | No | T24861 | Missense | This report | |
| 934 | Exon 42 | C7486T | Yes | R2496X | Nonsense | This report | |
| 2717 | Exon 42 | C7486T | Yes | R2496X | Nonsense | This report | |
| U-46 | Exon 42 | 7528ins1 bp | Duplication of bases 7515-7528 | aa 2510; PTC 2531 | 14-bp insertion | This report | |
| 698 | Exon 42 | 7544insGA | aa 2515; PTC 2527 | 2-bp insertion | This report | ||
| 2714 | Exon 43 | 7569delT | aa 2523; PTC 2526 | 1-bp deletion | This report | ||
| 1675 | Exon 43 | 7633insC | aa 2545; PTC 2555 | 1-bp insertion | This report | ||
| U-87b | Exon 44 | C7699T | No | Q2567X | Nonsense | This report | |
| 2253 | Exon 44 | A7701G | Q2567Q | Silent | This report | ||
| 90 | Exon 44 | C7702T | (Yes) | Q2568X | Nonsense | This report | |
| 2973 | Exon 44 | C7702T | (Yes) | Q2568X | Nonsense | This report | |
| 2569 | Exon 45 | C7846T | Yes | R2616X | Nonsense | This report | |
| 383 | Intron 45 | 7907+1G→A | No | Skip of exon 45 | Out of frame; PTC 2604 | Splice site | 555 |
| 735 | Intron 45 | 7908-2A→G | Skip of exon 46 (?) | Out of frame; PTC 2640 | Splice site | This report | |
| 1480 | Exon 46 | 7926insT | aa 2643; PTC 2643 | 1-bp insertion | This report | ||
| 2499 | Exon 46 | 8024delC | aa 2675; PTC 2717 | 1-bp deletion | This report | ||
| 578 | Intron 46 | 8050+20A→G | No exon skipping | Polymorphism | This report | ||
| 586 | Intron 46 | 5080+20A→G | Polymorphism | This report | |||
| U-82/310 | Exon 47 | 8092insTT | aa 2698; PTC 2718 | 2-bp insertion | This report |
The same patient may have different ID numbers because some patients were included in both the PTT and the TGGE screening panels. Some patients appear twice in the list because two separate sequence changes were identified (see table 5).
Numbering system based on NF1 cDNA (Genbank M82814), 1 = first base of methionine (ATG) at start of cDNA. The asterisk indicates congruence to NF1 pseudogene sequences: C910T to D26141/AC004527 (#21); C2041T to U35696 (#14); A3467G, C3721T, C3826T, and C4537T to M84131 (#15).
All G→A and C→T transitions were checked to see whether they occurred in a CpG or a CpNpG motif. In the latter case, “yes” was put in parentheses.
A question mark in parentheses means “predicted but not tested.” A question mark without parentheses means “highly speculative” and that the underlying mutations were not considered pathogenic.
In the case of frameshift mutations, the affected amino acid (aa) positions are given, as well as the position of the premature termination codon (PTC) in the predicted aberrant polypeptide. In the case of in-frame deletions caused by exon skipping, the number of amino acids (aa) expected to be lost in the predicted aberrant polypeptide is given.
Distribution of NF1 Mutations
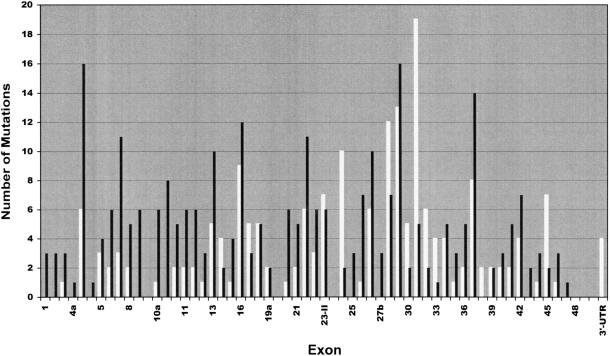
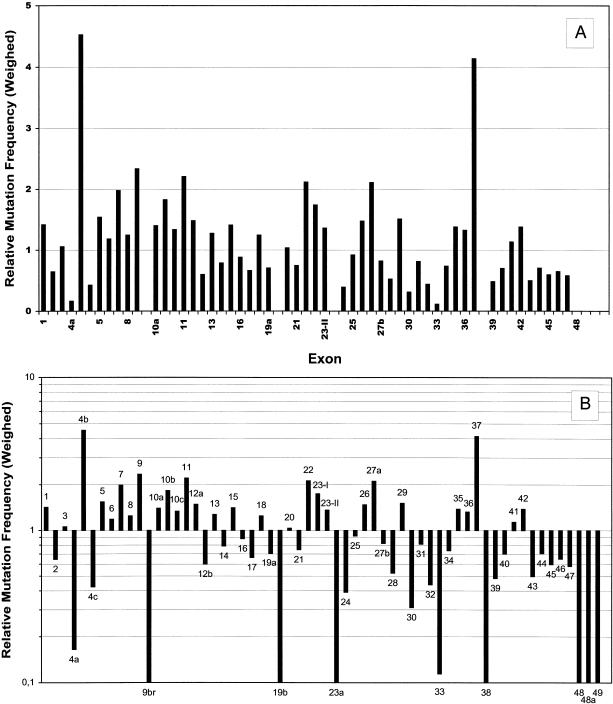
To get an idea of their distribution over the NF1 gene, we plotted the 278 pathogenic mutations exon by exon into a diagram (fig. 1). Intronic mutations were allocated to the nearest exon. Direct comparison with the 173 minor lesions that had been reported to the International NF1 Genetic Analysis Consortium by November 1997 (Korf 1998) (white columns in fig. 1) underscores the strong bias towards central parts of the NF1 gene inherent in these data. In particular, exons 28–36—the 9 exons that were described first (Cawthon et al. 1990)—and the GRD region were analyzed in hundreds of NF1 patients by the consortium members, whereas exons of other regions, mainly at the 5′ end of the gene, were not investigated at all. In our study, all parts of the gene were analyzed with equal intensity. Thus, we suppose our data to be an approximate representation of the actual distribution of mutations over the NF1 gene, at least in our study population, which is mainly of German origin, but includes also some patients of Turkish extraction. As indicated by figure 1, it appears likely that all regions of the NF1 gene are subject to mutation to a similar extent. However, if data are weighed for exon size (fig. 2a), exons 4b and 37 stand out as sites of a remarkably high mutation density. The reason for this may be found in some structural elements prone to mutation, a short tandem repeat structure in exon 4b (Toliat et al. 1999) and a quasisymmetric element in exon 37 (Robinson et al. 1995; Böddrich et al. 1997). If presented in a logarithmic mode (fig. 2b), the data reveal a slight tendency to fewer mutations towards the 3′ end of the gene that may be caused by the distribution of the missense mutations, as discussed later. Furthermore, several exons are easily recognized in which no mutation was found (exons 9br, 19b, 23a, 38, and 48–49) or that are extremely underrepresented (exons 4a and 33).
Figure 1.
Histogram of number of mutations, exon by exon. Black columns represent the 278 pathogenic mutations of table 4. White columns represent the mutations reported to the NF1 Genetic Analysis Consortium as of November 1997 (Korf 1998).
Figure 2.
Weighed distribution of mutations over the NF1 gene. For each exon, the number of pathogenic mutations was divided by the number of base pairs (bp). Ten bp were added to each exon, to allow for splice-site mutations. Values shown are ratios between the exon-specific mutation densities and the average mutation density for the whole gene (278/9,204 bp). A, linear presentation. B, logarithmic presentation.
Cryptic Splice Mutations
We identified a total of 55 splicing-error mutations (19.8% of all the mutations we found). In 22 cases, an exon skip was revealed in the patients’ mRNA. Several mutations were found to affect the splicing process, although they do not pertain to the group of typical splice-site mutations. At first glance, many of these were interpreted as faults, but analysis of mRNA as part of the PTT made us aware of the problem, and we initiated a routine checking of mRNA in all ambiguous cases (if mRNA was available). As reported elsewhere (Hoffmeyer et al. 1998), the two nonsense mutations (R304X and Y2264X) result in a skipping of exons 7 and 37, respectively. The mutation Y2264X may be caused by either of two transversions, C6792A or C6792G, both of which result in skipping of exon 37 (Messiaen et al. 1997). Five other nonsense mutations, as well as six small deletions or insertions, were also tested thoroughly, but did not show any influence on the splicing process (see table 4). However, in addition to the nonsense mutations, we identified a silent mutation (K354K) and several missense mutations (Y489C, S574T, S574N, R1849Q, and A2420P) that also cause exon skipping, presumably or definitely (see table 4). Most of these mutations affect the last base of the exon that is part of the splice donor and is usually a guanine (Krawczak et al. 1992). Messiaen et al. (1998) reported that mutation Y489C generates a new splice donor converting the 3′ end of exon 10b into an intronic sequence that is subsequently removed during splicing as part of intron 10b. The mutation Y491C might act in the same manner; however, no mRNA was available to test this possibility.
Two Sequence Changes
All 60 exons were analyzed in all patients; therefore, multiple sequence changes should have been identified if present. In addition to the known biallelic polymorphisms in exons 5 (702A/G) and 13 (2034G/A) and those in introns 28 (5205+23C/T) and 41 (7395-29A/G), we identified three novel biallelic polymorphisms: 1528-29delT, 1641+39C/T, and 3315-130G/C (located in introns 10b, 10c, and 19b, respectively) with allele frequencies higher than 30% for both alleles. The frequent polymorphisms were not taken into account when looking for additional sequence changes in the NF1 gene, yet two rare sequence changes were found in 10 of the patients (table 5). Determining which change is pathogenic is straightforward if a truncating mutation is observed among them; however, the occurrence of two pathogenic mutations is possible. We tested the mutations 2252-31A→T and 8050+20A→G for their influence on the splicing process, but no exon skipping was observed. In the case of patient 1899, a choice between two missense mutations, D176E and R1276Q, was necessary; the position of the latter gives a helpful hint. Arg1276 is the arginine finger of the GRD—the most essential catalytic element for RasGAP activity. Its mutation into proline was shown to completely disable GAP activity without impairing the secondary and tertiary protein structure (Klose et al. 1998a). Mutation R1276Q compromises GAP-stimulated GTP hydrolysis some 500-fold (Ahmadian et al. 1997). Compared with the 8,000-fold reduction by mutation R1276P (Klose et al. 1998a), this is still a remarkable residual activity. However, it may be sufficient to cause NF1, because other missense mutations of the GRD with moderate reduction of GAP activity have already been reported (Kim and Tamanoi 1998). It is tempting to speculate about a modifying effect of D176E on the NF1 phenotype. This might occur in cis as well as trans to another missense mutation or in trans to any other pathogenic mutation. In the instance of patient 1899, it is not known whether D176E and R1276Q are situated on the same allele or on different alleles. In the other three cases, both sequence changes cosegregate; that is, D176E is in cis to a truncating mutation, with little or no chance of exerting an effect on the phenotype because of the absence of the mutant gene product normally observed with this type of mutation (Hoffmeyer et al. 1995). Recently, we analyzed the unaffected father of patient 702, who is a sporadic case himself. Because the father is also a carrier of D176E, we were able to determine that this amino acid exchange alone is not sufficient to cause NF1.
Table 5.
Patients with Two Sequence Changes in the NF1 Gene
| Patient | First Sequence Change, Nonpathogenic | Second Sequence Change, Pathogenic |
| 1899 | T528A = D176E (exon 4b) | G3827A = R1276Q (exon 22) |
| 2406 | T528A = D176E (exon 4b) | 6642-1G→T (intron 35) |
| 190 | T528A = D176E (exon 4b) | 1541delAG (exon 10c) |
| 702 = U-25 | T528A = D176E (exon 4b) | G5546A = R1849Q (skip of exon 29) |
| 520 | 2252-31A→T (intron 13) | 3496+2T→C (intron 20) |
| E11225 | 2252-31A→T (intron 13) | C6792A = Y2264X (skip of exon 37) |
| 342 | G5172A = K1724K (exon 28) | 6789del4bp (exon 37) |
| B-212 | 7259-14C→T (intron 40) | 5168delTC (exon 28) |
| 2253 | A7701G = Q2567Q (exon 44) | T482A = L161X (exon 4b) |
| 578 | 8050+20A→G (intron 46) | 3911delT (exon 23-1) |
CpG Mutations
CpG dinucleotides show a high mutation rate in the human genome caused by spontaneous deamination of 5-methylcytosine; ∼25% of all single base-pair substitutions involve this dinucleotide (Cooper and Krawczak 1993). In the coding region of the NF1 gene, there are 118 CpGs, and a C→T transition on either the coding or noncoding strand of 91 of these would result in a disease-related mutation (Krkljus et al. 1997). Methylation of CpGs is a prerequisite for this type of mutation. Interestingly, methylated CpGs have been reported to occur within and around the NF1 gene; for example, 92% of CpGs in exons 28, 29, and 31 were found to be methylated in sperm DNA (Rodenhiser et al. 1993; Andrews et al. 1996). This prompted us to check our data for typical CpG mutations. Among the 301 sequence changes, we identified 57 (18.9%) C→T or G→A transitions within a CpG dinucleotide (table 4). The 301 sequence changes include 181 single base-pair substitutions, and only 31.5% of these represent a typical CpG mutation. Twenty-one different CpGs from the coding region are involved; that is, 17.8% of the 118 CpGs present in the coding region. They include 1 silent, 2 missense, and 18 nonsense mutations. One of these CpG mutations, R1513X, is the most frequently observed recurrent mutation among all 278 mutations. In addition to CpGs, we identified 9 CpNpG motifs that were subject to a C→T or G→A transition in 10 patients. Although DNA methylation at these sites has also been reported (Clark et al. 1995), it is evident from our data that they do not play a significant role in NF1 mutagenesis.
Relations to Pseudogene Variants
Genetic variation within pseudogenes may serve as a template for pathological lesions in the original gene by means of gene conversion even if located on different chromosomes (Eikenboom et al. 1994). In view of the large number of NF1 pseudogenes, it has been hypothesized that they are reservoirs of preformed mutations and may account for the high mutation rate of the NF1 gene (Cummings et al. 1993). Therefore, we analyzed all NF1-like sequences available from the public databases for any sequence deviation from the authentic NF1 gene. We located 31 entries (table 6) and identified 235 discrepancies (of which 196 were different) between the sequences of the pseudogenes and the NF1 gene (not shown). Only 6 (3.1%) of the 196 pseudogene variations were also found among the NF1 mutations in table 4 (indicated by an asterisk); however, 4 of the 6 were recurrent mutations (including R1513X, which was found in seven patients). Unfortunately, all but 1 of the 18 mutations with a pseudogene equivalent happened to occur at highly mutable CpG sites (see above). This definitely weakens further the evidence of NF1 gene conversion. Moreover, for gene conversion to explain the high mutation rate, mutations should predominantly affect the central part of the gene, which is not the case.
Table 6.
NF1 Pseudogene Sequences in the Public Databases
| Accession Number | Exons | Chromosome | Reference |
| AC004527 | 7–9 | 21 | R. B. Weiss et al.,a unpublished data |
| D26141 | 7–11 | 21 | H. Suzuki, unpublished data |
| U35688 | 8 | 18 | Purandare et al. (1995) |
| U35689 | 9 | 18 | Purandare et al. (1995) |
| U35696 | 13 | 18 | Purandare et al. (1995) |
| YO7853 | 13–15 | 17 | Régnier et al. (1997) |
| YO7854 | 13–15 | 14 | Régnier et al. (1997) |
| YO7855 | 13–15 | 14 | Régnier et al. (1997) |
| YO7856 | 13–15 | 15 | Régnier et al. (1997) |
| YO7857 | 13–15 | 15 | Régnier et al. (1997) |
| YO7858 | 13–15 | 2 | Régnier et al. (1997) |
| YO7859 | 13–15 | 22 | Régnier et al. (1997) |
| AF011743 | 14 | 15q11.2 | Kehrer-Sawatzki et al. (1997) |
| AF011744 | 15 | 15q11.2 | Kehrer-Sawatzki et al. (1997) |
| AF011746 | 15 | 15q11.2 | Kehrer-Sawatzki et al. (1997) |
| U35684 | 15 | 14 | Purandare et al. (1995) |
| U35685 | 15 | 15 | Purandare et al. (1995) |
| U35686 | 16 | 12 | Purandare et al. (1995) |
| U35687 | 16 | 14 | Purandare et al. (1995) |
| U35690 | 18 | 14 | Purandare et al. (1995) |
| U35691 | 18 | 15 | Purandare et al. (1995) |
| U35692 | 18 | 22 | Purandare et al. (1995) |
| U35693 | 19b | 15 | Purandare et al. (1995) |
| M84131 | 20–22 | 15 | Legius et al. (1992) |
| AF011748 | 21 | 15q11.2 | Kehrer-Sawatzki et al. (1997) |
| U35694 | 23-1 | 15 | Purandare et al. (1995) |
| U35695 | 24 | 15 | Purandare et al. (1995) |
| AF011745 | 24 | 15q11.2 | Kehrer-Sawatzki et al. (1997) |
| AF011747 | 24 | 15q11.2 | Kehrer-Sawatzki et al. (1997) |
| AF011749 | 24 | 15q11.2 | Kehrer-Sawatzki et al. (1997) |
| X72619 | 24 | 15q24-qter | Gasparini et al. (1993) |
| M84131 | 25–27b | 15 | Legius et al. (1992) |
Unpublished data from R. B. Weiss, D. M. Dunn, A. Aoyagi, L. Banks, B. Duval, C. Hamil, C. Holmes, M. Mahmoud, R. Rose, R. Stokes, M. D. Stump, P. Yu, L. Zhou, Y. Gitin, J. Nelson, and A. von Niederhausen.
Missense Mutations
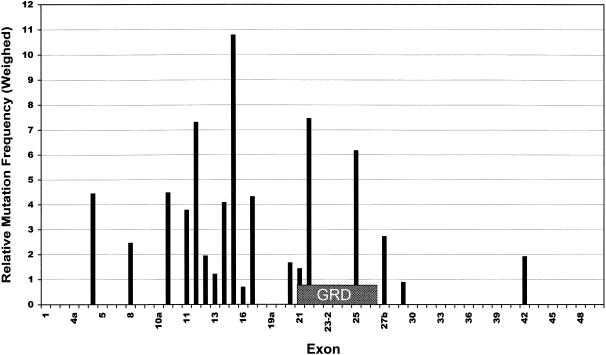
Formally, we identified a total of 39 missense mutations or deletions of single amino acids. However, as discussed earlier, D176E has to be considered a polymorphism, and others were shown to be splice mutations. Furthermore, substitution of threonine for the first methionine will cause a false start in translation, with one of the following methionines acting as the initiation codon (Neote et al. 1990; Patten et al. 1990). Hence, strictly speaking, only 28 are genuine missense mutations causing changes or losses of a particular amino acid in the polypeptide chain. The distribution of these mutations is shown in figure 3. It is evident that the mutations cluster in two regions: the GRD, and upstream between exons 10b and 17. If we assume the mutation Y491C to be a splice mutation like Y489C (see above), this region would shrink even further, to an interval from exon 11 to exon 17. Interestingly, a total of 11 substitutions into proline (39%) were found among the 28 missense mutations. Proline is known to destabilize helices and beta sheets.
Figure 3.
Weighed distribution of missense mutations over the NF1 gene. For each exon, the number of genuine missense or single-amino-acid-deletion mutations was divided by the number of base pairs (bp). Values shown are ratios between the exon-specific mutation densities and the average mutation density for the whole gene (28/8457 bp). Location of the GRD is indicated by the box.
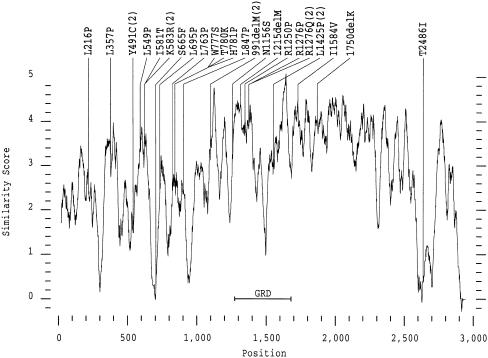
Except for patient 1899, who carries two missense mutations, R1276Q and D176E, all of the other 28 genuine missense mutations turned out to be the only sequence change present in the entire NF1 gene. This suggests that these mutations cause the disease, yet further evidence is needed to verify this hypothesis. To date, functional and structural assessment of particular amino acids has been obtained only for the GRD. For other regions, information about evolutionary conservation may help to evaluate the significance of an amino acid exchange. Therefore, we generated an amino acid sequence alignment (program PILEUP of the GCG-Wisconsin package, version 9; gap weight 12 and gap length weight 4) between the sequences of the human and the Drosophila NF1 proteins that are 60% identical over their entire length (The et al. 1997). As many as 15 of the 23 amino acid positions affected by missense mutation in our patients with NF1 turned out to be invariant, whereas, at two further positions, the amino acids were replaced by similar ones in the Drosophila protein (Ile1584 by Val and Arg1250 by Lys). At the other six positions nonconservative substitutions were found or the alignment generated a gap in the Drosophila protein (Y491: M; K583: Q; S665:−; H781: F; M1215: I; T2486:−). Moreover, we performed a similarity plot between the human and Drosophila sequences. Most of the mutations are situated on top of the similarity peaks or, at least, in regions of higher similarity, which may represent helices or beta sheets (fig. 4). Exceptions are Y491C—which might, in fact, be a splice mutation—S665F, and T2486I. These findings raise the possibility that the latter two mutations may actually be nonpathogenic.
Figure 4.
Missense mutations in the similarity plot of human versus Drosophila NF1 protein. The program PLOTSIMILARITY of the GCG-Wisconsin package, version 9 was used to calculate the amino-acid-sequence similarity profile of the human and Drosophila neurofibromins (accession numbers AAA59925 and AAB58976, respectively). A sliding window of 40 amino acids was chosen. Note that there are 2,963 instead of 2,839 amino acid positions in the alignment (program PILEUP; gap weight = 12 and gap-length weight = 4). For the first 20 and the last 20 positions, a value could not be calculated, because of the chosen window size. Mutations are located primarily in regions of higher similarity. Usually, helices and beta sheets have a higher local sequence similarity than coils.
Discussion
The systematic analysis of germline mutations in NF1 patients has been a major challenge for many laboratories during the last couple of years. Here, we present for the first time a comprehensive screen of minor lesions of the NF1 gene in a large cohort of >500 unrelated patients that is free of any assessment bias for special exons or regions. We launched this project hoping that the data would allow us to answer some long-standing questions about the NF1 gene and its proneness to mutate. Some of the questions were answered, but new questions were raised as well.
The Mutational Spectrum
A preponderance of truncating mutations is one of the most prominent features of NF1 mutations. Some 80% of all small lesions result in a PTC; that is, about one half of all NF1 mutations may be direct- or indirect-stop mutations. Distribution of this type of mutation is very even over the coding sequence. We identified the first one in exon 1 and the last in exon 47. At first glance, no difference in phenotype was observed; however, a detailed analysis of a possible relation between clinical features and position of the PTC is in progress and will be presented elsewhere (S. Tinschert, A. Buske, B. Müller-Myhsok, I. Naumann, C. Mischung, R. Fahsold, S. Hoffmeyer, D. Kaufmann, H. Peters, P. Nürnberg, unpublished data). Unequal expression of both NF1 alleles is a common observation in NF1 and has been related to the high number of truncating mutations (Hoffmeyer et al. 1995). This might explain the absence of a position effect of PTCs; however, we also observed equal expression in some cases, including patients U-62, 82, 342, and 407 (table 4). Of all mutations identified in this study, ∼20% were splice defects, with nearly one-half of the latter causing in-frame exon skipping. Hence, analysis of mRNA, in combination with a PTC screening as performed in the PTT, is a good choice for mutation detection in NF1. In our hands, the detection rate was 47.1%, which is lower than the 67% detection rate originally reported (Heim et al. 1995) but in line with the genomic-screening methods applied in this study, taking into account the fact that missense mutations cannot be detected by this approach. According to our data, missense mutations represent ∼10% of all small NF1 lesions.
The identification of a recurrent NF1 mutation has often been heralded as the discovery of a mutation hotspot. The most widely known example is the mutation R1947X in exon 31 (Valero et al. 1994; Lázaro et al. 1995). At least 10 unrelated patients with NF1, among 563 tested, were reported to carry this mutation (recurrence rate ∼2%) (Upadhyaya and Cooper 1998). In our screening of 521 unrelated patients with NF1, however, only 3 presented with this mutation. The most frequent mutation in our study was R1513X in exon 27a found in seven patients. However, when looking for the highest density of mutations, exons 4b and 37 outnumbered all other exons with a value 4- to 5-fold higher than average. This suggests these exons include some minor hotspots. Nevertheless, with the large number of novel mutations presented in this paper, it becomes obvious that the worst-case scenario is operating: hundreds of private mutations for most of the families affected by NF1. Interestingly, all three alternatively spliced exons—9br, 23a, and 48a—seem to be irrelevant to mutation analysis. Likewise, two normally spliced exons—19b and 38—also failed to show a mutation in our study population, and other exons—such as 4a and 33—harbored only a single mutation. The reason for this underrepresentation in the compulsory exons is not clear and may be purely accidental. For exons 33 and 38, additional mutations have been reported to the consortium (Korf 1998).
In general, the mutational spectrum of the NF1 gene seems to be very similar to that of other tumor-suppressor and mutator genes, which all seem to be characterized by a large proportion of truncating mutations—for example, nearly 100% for APC (Suzuki et al. 1998) and TSC1 (Jones et al. 1999), >80% for RB1 (Lohmann et al. 1996; 1997), ∼80% for BRCA1 (Miki et al. 1994) and ATM (Sandoval et al. 1999), and ∼60% for TSC2 (Jones et al. 1999), and all six NBS1 gene mutations identified to date (Varon et al. 1998). Similarly, 5%–20% missense mutations, as well as whole-gene deletions, have been found in some of these genes (RB1, ATM, TSC2). In addition, the problem of unsuccessful mutation detection in many of the patients is common to all these genes. Little more than half of all the NF1 alleles investigated in this study yielded a detectable mutation. It is true that the PTT protocol used here might have failed to detect missense as well as splicing-error mutations—especially those that result in in-frame skips of small exons— but, in most cases, all exons and adjacent intron regions of the NF1 gene were sequenced. DGS, like TGGE, is commonly thought to provide a sensitivity close to 100% in the analyzed regions, and, although some problems with heterozygote detection in automated sequencing are known to impair the identification rate of point mutations, this alone is unlikely to account for missing >40% of the NF1 mutations. Some of the unidentified mutations are definitely large deletions comprising the whole NF1 gene or a major part of it. When analyzing 15 TGGE-negative cases by fluorescence in situ hybridization, we identified three NF1 gene deletions (G. Thiel, unpublished data). Hence, we estimate the overall proportion of gross genomic deletions to be ∼10%. Among the remaining 35% of undetected mutations, multiexon deletions may represent the largest group (15% of all mutations, according to the consortium data [Korf 1998]). Furthermore, large duplications or inversions would also have escaped our PCR-based mutation scanning (Naylor et al. 1993). Incomplete assessment of intronic and regulatory sequences probably accounts for other unidentified cases. However, NF1 promoter mutations seem to be rare and the sequence variants identified so far are unlikely to cause the disease phenotype (Osborn and Upadhyaya 1998). For the RB1 gene, silencing of the promoter by allele-specific hypermethylation of a CpG island in the 5′ region was shown in 10% of retinoblastomas (Sakai et al. 1991; Greger et al. 1994). Interestingly, a hypomethylated region near the transcription start site within the NF1 CpG island could act as a target for silencing of the NF1 gene by local hypermethylation (Rodenhiser et al. 1998). Thus, epigenetic and genetic mechanisms may cause inactivation of NF1 transcription. Recent studies have shown that somatic mosaicism may be a frequent finding among sporadic cases of NF1 (Colman et al. 1996; Ainsworth et al. 1997; Tonsgard et al. 1997; Wu et al. 1997; Rasmussen et al. 1998). Provided the mutation was present in the tissues analyzed, the methods used for exon screening in this study are unlikely to have had sufficient sensitivity to detect low-level mosaicism. Unfortunately, we cannot exclude the possibility that patients who did not have NF1 were included in the screening, because some of the patients were diagnosed by others. Finally, the existence of genetic heterogeneity in NF1 is still an open discussion.
High Mutation Rate
The mutation rate for NF1 is one of the highest known for human disorders, estimated to be between 1×10−4 and 3.1×10−5 (Sergeyev 1975; Huson et al. 1989; Clementi et al. 1990). Thus, the NF1 gene seems to mutate 10 times more often than other disease genes. Several reasons have been proposed to explain the high mutability of the NF1 gene, including its relatively large size of ∼350 kb, gene-conversion events via pseudogenes, and mutational hotspots. Acting on the results of our study, we can definitely exclude the many NF1 pseudogenes as a major reason. Furthermore, pronounced hotspots of mutation have not yet been identified in the NF1 gene. It is true that some of the recurrent mutations, such as R1513X and R1947X, affect CpG dinucleotides, but the overall proportion of C→T or G→A transitions within CpGs is as low as 19% of all mutations and 32% of all the single-base-pair substitutions. Many other genes show similar or even higher percentages of CpG mutations—for example, 46% of factor VIII point mutations in hemophilia A (Antonarakis et al. 1995), 45%–50% of TP53 point mutations in colorectal cancer (Greenblatt et al. 1994), and 20%–25% of all RB1 mutations (Cowell et al. 1994). Likewise, overlap between pseudogene sequence variants and NF1 germline mutations as identified in this study is negligible. Therefore, we assume the large size of the NF1 gene to be the major reason for the high mutation rate, which is in fact not considerably higher than that of other large genes. In Duchenne muscular dystrophy, which is caused by mutations in an even larger gene (79 exons in 2.5 Mb), a similar high mutation rate is found (Vogel and Motulsky 1997). Interestingly, current estimates in Marfan syndrome place its prevalence at 1 per 3,000–5,000 people (Pyeritz 1996). This suggests that the FBN1 gene (65 exons in 110 kb), which is very similar in size to the NF1 gene, has a mutation rate nearly as high as in NF1.
Second Functional Domain
Our characterization of significant numbers of missense mutations and some small in-frame deletions is likely to be helpful in the investigation of putative functions of the NF1 gene product. The RasGAP activity of the central GAP-related domain and the structure of the GRD from neurofibromin have already been well characterized (Kim and Tamanoi 1998; Scheffzek et al. 1998), in particular, the effects of mutations of the arginine finger (Arg 1276) were studied in detail (Ahmadian et al. 1997; Klose et al. 1998a). Clustering of missense mutations in the GRD, as observed in this and other studies (fig. 5), is what we could expect from its functional relevance. However, there is another segment in the NF1 gene, upstream of the GRD, that harbors numerous missense mutations as well (see fig. 5). We assume this region defines a second functional domain of the neurofibromin molecule. Interestingly, this region coincides with the cysteine/serine-rich domain (CSRD) defined by Izawa et al. (1996). The CSRD comprises amino acid residues 543–909, in which three cysteine pairs (residues 622/632, 673/680, and 714/721) may be comparable to those that Maru et al. (1991) suggested form the ATP-binding domain of BCR protein. Furthermore, this sequence harbors three potential cAMP-dependent protein kinase A (PKA) recognition sites (Marchuk et al. 1991). These are, indeed, subject to phosphorylation by PKA (Izawa et al. 1996), and one is mutated in two of our NF1 families (mutation K583R in exon 12a). A second is situated in a small region (residues 815–834) of sequence similarity with two other proteins (MAP-2 and tau) that also associate with microtubules. It has been speculated that phosphorylation of this region might regulate the association of neurofibromin and microtubules (Gregory et al. 1993). The functional significance of the phosphorylation of neurofibromin by PKA is still unclear, but in view of the compelling evidence for the involvement of neurofibromin in cAMP-mediated signaling from Drosophila (Guo et al. 1997; The et al. 1997), it may explain the observed link between the second messenger cAMP and the Ras signaling pathway (Wu et al. 1993).
Figure 5.
Synopsis of all known NF1 missense mutations and deletions of 1 or 2 amino acids. Mutations on the left and right sides are data from this study and from the literature, respectively. Most published mutations have been reviewed in Upadhyaya and Cooper (1998). Some have been published very recently: R1204G (Krkljus et al. 1997); C39Y(1), Y489C, L847P, and 991delM(1) (Messiaen et al. 1998); and L2317P (Wu et al. 1999). Others have been submitted to the consortium but not published at this time. They are reported here with the consent of the consortium members who identified these mutations: 2366delNF(2) and R2616Q (L. Messiaen, personal communication); R1849G and L1932P (M. Upadhyaya, personal communication); 1658delIY and P2046R (D. Vidaud, personal communication). Mutations not representing genuine missense mutations including polymorphisms, splice mutations, or loss of the initiation codon are printed in italics. Number in parentheses after the mutation symbol indicates recurrent identifications. GRD region is shown in gray, whereas the exons 11–17, supposed to code for a new functional domain, are shown in black with a central light effect. To facilitate orientation, the numbers of some of the larger exons are given.
In summary, we have attributed functional relevance to a new region of the neurofibromin molecule on the basis of the missense mutations identified in a large cohort of NF1 patients. This protein segment was found upstream of the GRD and may be identical with the CSRD described elsewhere (Izawa et al. 1996). PKA phosphorylation sites within the domain suggest this region might be involved in cAMP-mediated signaling. However, further studies will be required to substantiate our hypothesis. To this end, the functional significance of the relevant missense mutations is currently being tested in the Drosophila model.
Acknowledgments
We thank the patients and their families for their willing cooperation, and we gratefully acknowledge the assistance of Peter N. Robinson, Annett Böddrich, Denise Horn, Fikret Erdogan, Mohammad R. Toliat, Anja Harder-Klose, Detlef Hess, Christiane Bielefeld, Andrea Lüder, Carmen Macsuga, Beatrice Heyne, and Kerstin Kamprath in the TGGE approach. We also thank Frances Hannan for his helpful comments on the manuscript. This work was supported in part by the Deutsche Krebshilfe.
Electronic-Database Information
Accession numbers and URLs for data in this article are as follows:
- GenBank, http://www.ncbi.nlm.nih.gov/Genbank/index.html (for human and Drosophila neurofibromin protein sequences [accession numbers AAA59925 and AAB58976, respectively] and human NF1 pseudogene sequences [accession numbers are listed in ])
- International NF1 Genetic Analysis Consortium, http://www.nf.org/nf1gene/nf1gene.home.html (for unpublished NF1 mutations)
- Messiaen L, Callens T, Mortier G, van Roy N, Speleman F, de Pape A (1998) Identification of mutations in the NF1 gene, including 3 different nonsense mutations and 1 missense mutation disrupting normal RNA splicing. (Abstract) http://nf.org/md1aspe1.htm
- Online Mendelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/Omim/ (for NF1 [162200])
References
- Abernathy CR, Rasmussen SA, Stalker HJ, Zori R, Driscoll DJ, Williams CA, Kousseff BG, et al (1997) NF1 mutation analysis using a combined heteroduplex/SSCP approach. Hum Mutat 9:548–554 [DOI] [PubMed]
- Ahmadian MR, Stege P, Scheffzek K, Wittinghofer A (1997) Confirmation of the arginine-finger hypothesis for the GAP-stimulated GTP-hydrolysis reaction of Ras. Nat Struct Biol 4:686–689 [DOI] [PubMed]
- Ainsworth PJ, Chakraborty PK, Weksberg R (1997) Example of somatic mosaicism in a series of de novo neurofibromatosis type 1 cases due to a maternally derived deletion. Hum Mutat 9:452–457 [DOI] [PubMed]
- Andrews JD, Mancini DN, Singh SM, Rodenhiser DI (1996) Site and sequence specific DNA methylation in the neurofibromatosis (NF1) gene includes C5839T: the site of the recurrent substitution mutation in exon 31. Hum Mol Genet 5:503–507 [DOI] [PubMed]
- Antonarakis SE, Kazazian HH, Tuddenham EG (1995) Molecular etiology of factor VIII deficiency in hemophilia A. Hum Mutat 5:1–22 [DOI] [PubMed]
- Böddrich A, Griesser J, Horn D, Kaufmann D, Krone W, Nürnberg P (1995) Reduced neurofibromin content but normal GAP activity in a patient with neurofibromatosis type 1 caused by a five base pair duplication in exon 12b of the NF1 gene. Biochem Biophys Res Commun 214:895–904 [DOI] [PubMed]
- Böddrich A, Robinson PN, Schülke M, Buske A, Tinschert S, Nürnberg P (1997) New evidence for a mutation hotspot in exon 37 of the NF1 gene. Hum Mutat 9:374–377 [DOI] [PubMed]
- Cawthon RM, Weiss R, Xu G, Viskochil D, Culver M, Stevens J, Robertson M, et al (1990) A major segment of the neurofibromatosis type 1 gene: cDNA sequence, genomic structure, and point mutations. Cell 62:193–201 [DOI] [PubMed]
- Clark SJ, Harrison J, Frommer M (1995) CpNpG methylation in mammalian cells. Nat Genet 10:20–27 [DOI] [PubMed]
- Clementi M, Barbujani G, Turolla L, Tenconi R (1990) Neurofibromatosis-1: a maximum likelihood estimation of mutation rate. Hum Genet 84:116–118 [DOI] [PubMed]
- Colman SD, Rasmussen SA, Ho VT, Abernathy CR, Wallace MR (1996) Somatic mosaicism in a patient with neurofibromatosis type 1. Am J Hum Genet 58:484–490 [PMC free article] [PubMed]
- Colman SD, Williams CA, Wallace MR (1995) Benign neurofibromas in type 1 neurofibromatosis (NF1) show somatic deletions on NF1 gene. Nat Genet 11:90–92 [DOI] [PubMed]
- Cooper DN, Krawczak M (1993) Human gene mutation. BIOS Scientific Publishers, Oxford [Google Scholar]
- Cowell JK, Smith T, Bia B (1994) Frequent constitutional C to T mutations in CGA-arginine codons in the RB1 gene produce premature stop codons in patients with bilateral (hereditary) retinoblastoma. Eur J Hum Genet 2:281–290 [DOI] [PubMed]
- Cummings LM, Glatefeller A, Marchuk DA (1993) NF1 related loci on chromosomes 2, 12, 14, 15, 20, 21, and 22: a potential role for gene conversion in the high spontaneous mutation rate of NF1? Am J Hum Genet Suppl 53:A672 [Google Scholar]
- Danglot G, Régnier V, Fauvet D, Vassal G, Kujas M, Bernheim A (1995) Neurofibromatosis 1 (NF1) mRNAs expressed in the central nervous system are differentially spliced in the 5′ part of the gene. Hum Mol Genet 4:915–920 [DOI] [PubMed]
- Däschner K, Assum G, Eisenbarth I, Krone W, Hoffmeyer S, Wortmann S, Heymer B, et al (1997) Clonal origin of tumor cells in a plexiform neurofibroma with LOH in NF1 intron 38 and in dermal neurofibromas without LOH of the NF1 gene. Biochem Biophys Res Commun 234:346–350 [DOI] [PubMed]
- Easton DF, Ponder MA, Huson SM, Ponder BA (1993) An analysis of variation in expression of neurofibromatosis (NF) type 1 (NF1): evidence for modifying genes. Am J Hum Genet 53:305–313 [PMC free article] [PubMed]
- Eikenboom JC, Vink T, Briet E, Sixma JJ, Reitsma PH (1994) Multiple substitutions in the von Willebrand factor gene that mimic the pseudogene sequence. Proc Natl Acad Sci USA 91:2221–2224 [DOI] [PMC free article] [PubMed]
- Friedman JM, Birch PH (1997) Type 1 neurofibromatosis: a descriptive analysis of the disorder in 1,728 patients. Am J Med Genet 70:138–143 [DOI] [PubMed]
- Gasparini P, Grifa A, Origone P, Coviello D, Antonacci R, Rocchi M (1993) Detection of a neurofibromatosis type I (NF1) homologous sequence by PCR: implications for the diagnosis and screening of genetic diseases. Mol Cell Probes 7:415–418 [DOI] [PubMed]
- Gille C, Gille A, Booms P, Robinson PN, Nürnberg P (1998) Bipolar clamping improves the sensitivity of mutation detection by temperature gradient gel electrophoresis. Electrophoresis 19:1347–1350 [DOI] [PubMed]
- Greenblatt MS, Bennett WP, Hollstein M, Harris CC (1994) Mutations in the p53 tumor suppressor gene: clues to cancer etiology and molecular pathogenesis. Cancer Res 54:4855–4878 [PubMed]
- Greger V, Debus N, Lohmann D, Hopping W, Passarge E, Horsthemke B (1994) Frequency and parental origin of hypermethylated RB1 alleles in retinoblastoma. Hum Genet 94:491–496 [DOI] [PubMed]
- Gregory PE, Gutmann DH, Mitchell A, Park S, Boguski M, Jacks T, Wood DL, et al (1993) Neurofibromatosis type 1 gene product (neurofibromin) associates with microtubules. Somat Cell Mol Genet 19:265–274 [DOI] [PubMed]
- Guha A, Lau N, Huvar I, Gutmann D, Provias J, Pawson T, Boss G (1996) Ras-GTP levels are elevated in human NF1 peripheral nerve tumors. Oncogene 12:507–513 [PubMed]
- Guo HF, The I, Hannan F, Bernards A, Zhong Y (1997) Requirement of Drosophila NF1 for activation of adenylyl cyclase by PACAP38-like neuropeptides. Science 276:795–798 [DOI] [PubMed]
- Gutmann DH, Aylsworth A, Carey JC, Korf B, Marks J, Pyeritz RE, Rubenstein A, et al (1997) The diagnostic evaluation and multidisciplinary management of neurofibromatosis 1 and neurofibromatosis 2. JAMA 278:51–57 [PubMed]
- Harder A, Macsuga C, Tinschert S, Nürnberg P, Peters H (1999) A novel 5-bp deletion in exon 30 of the neurofibromatosis type 1 (NF1) gene. Hum Mutat 13:259 [Google Scholar]
- Heim RA, Kam-Morgan LNW, Binnie CG, Corns DD, Cayouette MC, Farber RA, Aylsworth AS, et al (1995) Distribution of 13 truncating mutations in the neurofibromatosis 1 gene. Hum Mol Genet 4:975–981 [DOI] [PubMed]
- Hoffmeyer S, Assum G (1994) An RsaI polymorphism in the transcribed region of the neurofibromatosis (NF1)-gene. Hum Genet 93:481–482 [DOI] [PubMed]
- Hoffmeyer S, Assum G, Griesser J, Kaufmann D, Nürnberg P, Krone W (1995) On unequal allelic expression of the neurofibromin gene in neurofibromatosis type 1. Hum Mol Genet 4:1267–1272 [DOI] [PubMed]
- Hoffmeyer S, Nürnberg P, Ritter H, Fahsold R, Leistner W, Kaufmann D, Krone W (1998) Nearby stop codons in exons of the neurofibromatosis type 1 gene are disparate splice effectors. Am J Hum Genet 62:269–277 [DOI] [PMC free article] [PubMed]
- Horn D, Robinson PN, Böddrich A, Buske A, Tinschert S, Nürnberg P (1996) Three novel mutations of the NF1 gene detected by temperature gradient gel electrophoresis of exons 5 and 8. Electrophoresis 17:1559–1563 [DOI] [PubMed]
- Huson SM, Compston DA, Clark P, Harper PS (1989) A genetic study of von Recklinghausen neurofibromatosis in south east Wales. I. Prevalence, fitness, mutation rate, and effect of parental transmission on severity. J Med Genet 26:704–711 [DOI] [PMC free article] [PubMed]
- Huson SM, Hughes RAC (1994) The neurofibromatoses: a clinical and pathogenetic overview. Chapman & Hall, London [Google Scholar]
- Izawa I, Tamaki N, Saya H (1996) Phosphorylation of neurofibromatosis type 1 gene product (neurofibromin) by cAMP-dependent protein kinase. FEBS Lett 382:53–59 [DOI] [PubMed]
- Jones AC, Shyamsundar MM, Thomas MW, Maynard J, Idziaszczyk S, Tomkins S, Sampson JR, et al (1999) Comprehensive mutation analysis of TSC1 and TSC2—and phenotypic correlations in 150 families with tuberous sclerosis. Am J Hum Genet 64:1305–1315 [DOI] [PMC free article] [PubMed]
- Kehrer-Sawatzki H, Schwickardt T, Assum G, Rocchi M, Krone W (1997) A third neurofibromatosis type 1 (NF1) pseudogene at chromosome 15q11.2. Hum Genet 100:595–600 [DOI] [PubMed]
- Kim MR, Tamanoi F (1998) Neurofibromatosis 1 GTPase activating protein-related domain and its functional significance. In: Upadhyaya M, Cooper DN (eds) Neurofibromatosis type 1: from genotype to phenotype. BIOS Scientific Publishers, Oxford, pp 89–112 [Google Scholar]
- Klose A, Ahmadian MR, Schülke M, Scheffzek K, Hoffmeyer S, Gewies A, Schmitz F, et al (1998a) Selective disactivation of neurofibromin GAP activity in neurofibromatosis type 1 (NF1). Hum Mol Genet 7:1261–1268 [DOI] [PubMed]
- Klose A, Nürnberg P, Tinschert S, Peters H (1998b) Recurrence of the neurofibromatosis type 1 (NF1) mutation R1276X probably due to interchromosomal gene conversion. Hum Mutat 12:291 [Google Scholar]
- Klose A, Peters H, Hoffmeyer S, Buske A, Lüder A, Hess D, Lehmann R, et al (1999) Two independent mutations in a family with neurofibromatosis type 1 (NF1). Am J Med Genet 83:6–12 [PubMed]
- Klose A, Robinson PN, Gewies A, Kluwe L, Kaufmann D, Buske A, Tinschert S, et al (1998c) Two novel mutations in exons 19a and 20 and a BsaBI [correction of BsaI] polymorphism in a newly characterized intron of the neurofibromatosis type 1 gene. Hum Genet 102:367–371 [DOI] [PubMed]
- Korf BR (1998) The NF1 genetic analysis consortium. In: Upadhyaya M, Cooper DN (eds) Neurofibromatosis type 1: from genotype to phenotype. BIOS Scientific Publishers, Oxford, pp 57–63 [Google Scholar]
- Krawczak M, Reiss J, Cooper DN (1992) The mutational spectrum of single base-pair substitutions in mRNA splice junctions of human genes: causes and consequences. Hum Genet 90:41–54 [DOI] [PubMed]
- Krkljus S, Abernathy CR, Johnson JS, Williams CA, Driscoll DJ, Zori R, Stalker HJ, et al (1998) Analysis of CpG C-to-T mutations in neurofibromatosis type 1. Hum Mutat 11:411 [DOI] [PubMed]
- Lázaro C, Kruyer H, Gaona A, Estivill X (1995) Screening for the neurofibromatosis type 1 recurrent mutation R1947X. Med Genetik 7:272 (Abstract) [Google Scholar]
- Legius E, Marchuk DA, Collins FS, Glover TW (1993) Somatic deletion of the neurofibromatosis type 1 gene in a neurofibrosarcoma supports a tumour suppressor gene hypothesis. Nat Genet 3:122–126 [DOI] [PubMed]
- Legius E, Marchuk DA, Hall BK, Andersen LB, Wallace MR, Collins FS, Glover TW (1992) NF1-related locus on chromosome 15. Genomics 13:1316–1318 [DOI] [PubMed]
- Lerman LS, Silverstein K (1987) Computational simulation of DNA melting and its application to denaturing gradient gel electrophoresis. Methods Enzymol 155:482–501 [DOI] [PubMed]
- Li Y, Bollag G, Clark R, Stevens J, Conroy L, Fults D, Ward K, et al (1992) Somatic mutations in the neurofibromatosis 1 gene in human tumors. Cell 69:275–281 [DOI] [PubMed]
- Li Y, O’Connell P, Huntsman-Breidenbach H, Cawthon R, Stevens J, Xu G, Neil S, et al (1995) Genomic organization of the neurofibromatosis 1 gene (NF1). Genomics 25:9–18 [DOI] [PubMed]
- Lohmann DR, Brandt B, Höpping W, Passarge E, Horsthemke B (1996) The spectrum of RB1 germ-line mutations in hereditary retinoblastoma. Am J Hum Genet 58:940–949 [PMC free article] [PubMed]
- Lohmann DR, Gerick M, Brandt B, Oelschläger U, Lorenz B, Passarge E, Horsthemke B (1997) Constitutional RB1-gene mutations in patients with isolated unilateral retinoblastoma. Am J Hum Genet 61:282–294 [DOI] [PMC free article] [PubMed]
- Marchuk DA, Saulino AM, Tavakkol R, Swaroop M, Wallace MR, Andersen LB, Mitchell AL, et al (1991) cDNA cloning of the type 1 neurofibromatosis gene: complete sequence of the NF1 gene product. Genomics 11:931–940 [DOI] [PubMed]
- Martin GA, Viskochil D, Bollag G, McCabe PC, Crosier WJ, Haubruck H, Conroy L, et al (1990) The GAP-related domain of the neurofibromatosis type 1 gene product interacts with ras p21. Cell 63:843–849 [DOI] [PubMed]
- Maru Y, Witte ON (1991) The BCR gene encodes a novel serine/threonine kinase activity within a single exon. Cell 67:459–468 [DOI] [PubMed]
- Maynard J, Krawczak M, Upadhyaya M (1997) Characterization and significance of nine novel mutations in exon 16 of the neurofibromatosis type 1 (NF1) gene. Hum Genet 99:674–676 [DOI] [PubMed]
- Messiaen L, Callens T, De Paepe A, Craen M, Mortier G (1997) Characterisation of two different nonsense mutations, C6792A and C6792G, causing skipping of exon 37 in the NF1 gene. Hum Genet 101:75–80 [DOI] [PubMed]
- Miki Y, Swensen J, Shattuck-Eidens D, Futreal PA, Harshman K, Tavtigian S, Liu Q, et al (1994) A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 266:66–71 [DOI] [PubMed]
- Naylor J, Brinke A, Hassock S, Green PM, Giannelli F (1993) Characteristic mRNA abnormality found in half the patients with severe haemophilia A is due to large DNA inversions. Hum Mol Genet 2:1773–1778 [DOI] [PubMed]
- Neote K, Brown CA, Mahuran DJ, Gravel RA (1990) Translation initiation in the HEXB gene encoding the β-subunit of human β-hexosaminidase. J Biol Chem 265:20799–20806 [PubMed]
- Osborn M, Upadhyaya M (1998) Mutation analysis in the promoter and coding region of the NF1 gene. Am J Hum Genet Suppl 63:A378 [Google Scholar]
- Patten JL, Johns DR, Valle D, Eil C, Gruppuso PA, Steele G, Smallwood PM, et al (1990) Mutation in the gene encoding the stimulatory G protein of adenylate cyclase in Albright's hereditary osteodystrophy. N Engl J Med 322:1412–1419 [DOI] [PubMed]
- Peters H, Hess D, Fahsold R, Schülke M (1999a) A novel mutation L1425P in the GAP-region of the NF1 gene detected by temperature gradient gel electrophoresis (TGGE). Hum Mutat 13:337 [DOI] [PubMed]
- Peters H, Lüder A, Harder A, Schuelke M, Tinschert S (1999b) Mutation screening of neurofibromatosis type 1 (NF1) exons 28 and 29 with single strand conformation polymorphism (SSCP): five novel mutations, one recurrent transition and two polymorphisms in a panel of 118 unrelated NF1 patients. Hum Mutat 13:258 [DOI] [PubMed]
- Purandare SM, Huntsman-Breidenbach H, Li Y, Lin Zhu X, Sawada S, Neil SM, Brothman A, et al (1995) Identification of neurofibromatosis 1 (NF1) homologous loci by direct sequencing, fluorescence in situ hybridization, and PCR amplification of somatic cell hybrids. Genomics 30:476–485 [DOI] [PubMed]
- Purandare SM, Lanyon WG, Connor JM (1994) Characterisation of inherited and sporadic mutations in neurofibromatosis type-1. Hum Mol Genet 3:1109–1115 [DOI] [PubMed]
- Pyeritz RE (1996) Disorders of fibrillins and microfibrillogenesis: Marfan syndrome and other disorders of fibrillin. In: Rimoin DL, Connor JM, Pyeritz RE (eds) Emery and Rimoin's principles and practice of medical genetics, 3rd ed. Churchill Livingstone, New York, pp 1027–1066 [Google Scholar]
- Rasmussen SA, Colman SD, Ho VT, Abernathy CR, Arn PH, Weiss L, Schwartz C, et al (1998) Constitutional and mosaic large NF1 gene deletions in neurofibromatosis type 1. J Med Genet 35:468–471 [DOI] [PMC free article] [PubMed]
- Régnier V, Meddeb M, Lecointre G, Richard F, Duverger A, Nguyen VC, Dutrillaux B, et al (1997) Emergence and scattering of multiple neurofibromatosis (NF1)-related sequences during hominoid evolution suggests a process of pericentromeric interchromosomal transposition. Hum Mol Genet 6:9–16 [DOI] [PubMed]
- Riccardi VM (1992) Neurofibromatosis: phenotype, natural history, and pathogenesis, 2nd ed. John Hopkins University, Baltimore [Google Scholar]
- Riccardi VM (1993) Genotype, malleotype, phenotype, and randomness: lessons from neurofibromatosis-1 (NF1). Am J Hum Genet 53:301–304 [PMC free article] [PubMed]
- Riesner D, Steger G, Zimmat R, Owens RA, Wagenhöfer M, Hillen W, Vollbach S, et al (1989) Temperature-gradient gel electrophoresis of nucleic acids: analysis of conformational transitions, sequence variations, and protein-nucleic acid interactions. Electrophoresis 10:377–389 [DOI] [PubMed]
- Robinson PN, Böddrich A, Peters H, Tinschert S, Buske A, Kaufmann D, Nürnberg P (1995) Two recurrent nonsense mutations and a 4 bp deletion in a quasi-symmetric element in exon 37 of the NF1 gene. Hum Genet 96:95–98 [DOI] [PubMed]
- Robinson PN, Buske A, Neumann R, Tinschert S, Nürnberg P (1996) Recurrent 2-bp deletion in exon 10c of the NF1 gene in two cases of von Recklinghausen neurofibromatosis. Hum Mutat 7:85–88 [DOI] [PubMed]
- Rodenhiser DI, Coulter-Mackie MB, Singh SM (1993) Evidence of DNA methylation in the neurofibromatosis type 1 (NF1) gene region of 17q11.2. Hum Mol Genet 2:439–444 [DOI] [PubMed]
- Rodenhiser D, Mancini DN, Singh SM, Archer TK (1998) Site specific DNA methylation in the neurofibromatosis (NF1) promoter interferes with binding of CREB and SP1 transcription factors. Am J Hum Genet Suppl 63:A190 [DOI] [PubMed] [Google Scholar]
- Sakai T, Toguchida J, Ohtani N, Yandell DW, Rapaport JM, Dryja TP (1991) Allele-specific hypermethylation of the retinoblastoma tumor-suppressor gene. Am J Hum Genet 48:880–888 [PMC free article] [PubMed]
- Sandoval N, Platzer M, Rosenthal A, Dork T, Bendix R, Skawran B, Stuhrmann M, et al (1999) Characterization of ATM gene mutations in 66 ataxia telangiectasia families. Hum Mol Genet 8:69–79 [DOI] [PubMed]
- Scheffzek K, Ahmadian MR, Wiesmuller L, Kabsch W, Stege P, Schmitz F, Wittinghofer A (1998) Structural analysis of the GAP-related domain from neurofibromin and its implications. EMBO J 17:4313–4327 [DOI] [PMC free article] [PubMed]
- Sergeyev AS (1975) On the mutation rate of neurofibromatosis. Humangenetik 28:129–138 [DOI] [PubMed]
- Serra E, Puig S, Otero D, Gaona A, Kruyer H, Ars E, Estivill X, Lazaro C (1997) Confirmation of a double-hit model for the NF1 gene in benign neurofibromas. Am J Hum Genet 61:512–519 [DOI] [PMC free article] [PubMed]
- Shannon KM, O’Connell P, Martin GA, Paderanga D, Olson K, Dinndorf P, McCormick F (1994) Loss of normal NF1 allele from the bone marrow of children with type 1 neurofibromatosis and malignant myeloid disorders. N Engl J Med 330:597–601 [DOI] [PubMed]
- Suzuki T, Ishioka C, Kato S, Mitachi Y, Shimodaira H, Sakayori M, Shimada A, et al (1998) Detection of APC mutations by a yeast-based protein truncation test (YPTT). Genes Chromosomes Cancer 21:290–297 [PubMed]
- The I, Hannigan GE, Cowley GS, Reginald S, Zhong Y, Gusella JF, Hariharan IK, et al (1997) Rescue of a Drosophila Nf1 mutant phenotype by protein kinase A. Science 276:791–794 [DOI] [PubMed]
- Toliat MR, Erdogan F, Gewies A, Fahsold R, Buske A, Tinschert S, Nürnberg P (1999) Analysis of the NF1 gene by temperature gradient gel electrophoresis (TGGE) reveals a high incidence of mutations in exon 4b. Electrophoresis 21:541–544 [DOI] [PubMed] [Google Scholar]
- Tonsgard JH, Yelavarthi KK, Cushner S, Short MP, Lindgren V (1997) Do NF1 gene deletions result in a characteristic phenotype? Am J Med Genet 73:80–86 [DOI] [PubMed]
- Upadhyaya M, Cooper DN (1998) The mutational spectrum in neurofibromatosis 1 and its underlying mechanisms. In: Upadhyaya M, Cooper DN (eds) Neurofibromatosis type 1: from genotype to phenotype. BIOS Scientific Publishers, Oxford, pp 65–88 [Google Scholar]
- Upadhyaya M, Osborn MJ, Maynard J, Kim MR, Tamanoi F, Cooper DN (1997) Mutational and functional analysis of the neurofibromatosis type 1 (NF1) gene. Hum Genet 99:88–92 [DOI] [PubMed]
- Valero MC, Velasco E, Moreno F, Hernandez-Chico C (1994) Characterization of four mutations in the neurofibromatosis type 1 gene by denaturing gradient gel electrophoresis (DGGE). Hum Mol Genet 3:639–641 [DOI] [PubMed]
- Varon R, Vissinga C, Platzer M, Cerosaletti KM, Chrzanowska KH, Saar K, Beckmann G, et al (1998) Nibrin, a novel DNA double-strand break repair protein, is mutated in Nijmegen breakage syndrome. Cell 93:467–476 [DOI] [PubMed]
- Vogel F, Motulsky AG (1997) Human genetics: problems and approaches, 3rd ed. Springer-Verlag, Berlin [Google Scholar]
- Wartell RM, Hosseini SH, Moran CP (1990) Detecting base pair substitutions in DNA fragments by temperature-gradient gel electrophoresis. Nucleic Acids Res 18:2699–2705 [DOI] [PMC free article] [PubMed]
- White R, O’Connell P (1991) Identification and characterization of the gene for neurofibromatosis type 1. Curr Opin Genet Dev 1:15–19 [DOI] [PubMed]
- Wu B-L, Boles RG, Yaari H, Weremowicz S, Schneider GH, Korf BR (1997) Somatic mosaicism for deletion of the entire NF1 gene identified by FISH. Hum Genet 99:209–213 [DOI] [PubMed]
- Wu J, Dent P, Jelinek T, Wolfman A, Weber MJ, Sturgill TW (1993) Inhibition of the EGF-activated MAP kinase signaling pathway by adenosine 3′,5′-monophosphate. Science 262:1065–1069 [DOI] [PubMed]
- Wu R, López-Correa C, Rutkowski JL, Baumbach LL, Glover TW, Legius E (1999) Germline mutations in NF1 patients with malignancies. Genes Chromosomes Cancer 26:376–380 [PubMed]
- Xu G, O'Connell P, Viskochil D, Cawthon R, Robertson M, Culver M, Dunn D, et al (1990) The neurofibromatosis type 1 gene encodes a protein related to GAP. Cell 62:599–608 [DOI] [PubMed]