Abstract
We introduce an unbiased protocol for performing rotational moves in rigid-body dynamics simulations. This approach—based on the analytic solution for the rotational equations of motion for an orthogonal coordinate system at constant angular velocity—removes deficiencies that have been largely ignored in Brownian dynamics simulations, namely errors for finite rotations that result from applying the noncommuting rotational matrices in an arbitrary order. Our algorithm should thus replace standard approaches to rotate local coordinate frames in Langevin and Brownian dynamics simulations.
INTRODUCTION
Generalized numerical methods for Langevin or Brownian dynamics (BD) involving rigid bodies require operators for applying arbitrary finite rotational moves to three-dimensional objects. Typically, a local body-fixed coordinate frame, which defines a body's three-dimensional orientation, is associated with each body in a system (García de la Torre and Bloomfield, 1981; Allison, 1991; Fernandez and García de la Torre, 2002). Such methods are important and commonly used to study large biomolecular systems, such as long DNA (Wu et al., 1991; Chirico and Langowski, 1996; Jian et al., 1997; Klenin et al., 1998) or large nucleoprotein complexes (Beard and Schlick, 2001; Huang and Schlick, 2002). During every time-step, each body is translated and rotated according to a given discretization of the governing dynamical equations (Ermak and McCammon, 1978; Allison, 1991; Fernandez and García de la Torre, 2002). The translational step is straightforward; it involves updating the three coordinates of a body's center of rotation. The rotational step is more subtle, because operators for rotation about orthogonal axes do not commute and thus their order of application defines different protocols. The related difficulties have been ignored in the past (e.g., Allison, 1991; Fernandez and García de la Torre, 2002); that is, the noncommuting operators are usually applied in some particular (arbitrary) order, introducing errors and bias into the rotational move. Although most prior models with BD simulation protocols have been assessed relative to available experimental data and found to be satisfactory overall, the previously-used rotational operators are strictly appropriate only in the limit of small rotational angles, where these errors are negligible. Here we address this limitation by exploring the magnitudes of angles for which errors become apparent in a simple model and introduce an unbiased accurate operator for finite rotations. Given that our protocol has the same computational cost as the biased protocol, it should be used in general in BD algorithms to avoid any possible errors from this computational component.
ROTATIONAL MOVES
We define a right-handed local particle-fixed coordinate frame for a rigid body using three orthogonal unit vectors  which translate and rotate in space as a dynamics trajectory evolves (e.g., Fig. 1 of Beard and Schlick, 2001). In this article, we focus on how a given rotation is applied to a coordinate frame; we do not consider issues related to how the dynamical equations are discretized to determine the rotation and translation vectors for each time-step.
which translate and rotate in space as a dynamics trajectory evolves (e.g., Fig. 1 of Beard and Schlick, 2001). In this article, we focus on how a given rotation is applied to a coordinate frame; we do not consider issues related to how the dynamical equations are discretized to determine the rotation and translation vectors for each time-step.
FIGURE 1.
The unit sphere in the {x,y,z} fixed lab frame. The initial local frame  corresponds to the fixed lab frame axes. The rotation vector
corresponds to the fixed lab frame axes. The rotation vector  is oriented in the x-y plane;
is oriented in the x-y plane;  denotes
denotes  after being rotated about
after being rotated about 
A finite rotation is denoted by  where the components of
where the components of  are rotations about the three orthogonal local axes. The central problem is posed as follows. Given a rotation vector
are rotations about the three orthogonal local axes. The central problem is posed as follows. Given a rotation vector  how can this rotation be applied to a coordinate frame
how can this rotation be applied to a coordinate frame  ? This issue is key in simulating Brownian dynamics of rigid bodies (Allison, 1991; Fernandez and García de la Torre, 2002). In previous works, a three-dimensional rotation is applied to a coordinate frame by successively applying individual rotations about each local frame. In this approach, the successive rotation operators are given by
? This issue is key in simulating Brownian dynamics of rigid bodies (Allison, 1991; Fernandez and García de la Torre, 2002). In previous works, a three-dimensional rotation is applied to a coordinate frame by successively applying individual rotations about each local frame. In this approach, the successive rotation operators are given by
 |
(1) |
and the cumulative rotational operator R is defined by applying these rotations in a particular order. For example, applying rotations about the  then
then  axes, yields
axes, yields
 |
(2) |
However, this choice of R is not unique. The rotation operators do not commute: Ra Rb Rc ≠ Rc Rb Ra. Thus, the operator R introduces bias into the coordinate rotation. As we show below for a given random rotation vector chosen from an unbiased distribution, resulting rotations will be biased toward an axis which is determined by the choice of the order of operators applied in Eq. 2.
BIAS-FREE ROTATIONAL APPROACH
A bias-free approach to performing these rotational steps can be formulated from the dynamics equations for rotation of an orthogonal coordinate system at a constant angular velocity (see Eq. 4-62 of Goldstein et al., 2002):
 |
(3) |
where {ωa, ωb, ωc} are the three components of angular velocity. For constant ωa, ωb, ωc, Eq. 3 is a constant-coefficient linear system with solution (which was obtained using the program MAPLE, Waterloo Maple, Waterloo, Ontario, Canada):
 |
(4) |
where {Ωa, Ωb, Ωc} = {ωaΔt, ωbΔt, ωcΔt}, and  .
.
Using the notation of Fernandez and García de la Torre (2002), Eq. 4 can be expressed compactly as
 |
(5) |
where the triad vectors  denote the initial local frame (at time to), and
denote the initial local frame (at time to), and  correspond to the local axes following rotation (at time t). The matrix
correspond to the local axes following rotation (at time t). The matrix  is a transformation from the local frame at time t to the fixed lab (l) frame. Thus Eq. 5 can be written
is a transformation from the local frame at time t to the fixed lab (l) frame. Thus Eq. 5 can be written  where the matrix U, defined by Eq. 5, replaces the operator R inEq. 2.
where the matrix U, defined by Eq. 5, replaces the operator R inEq. 2.
The matrix operator U in Eq. 5 is an exact representation for the rotation of the vectors  by an angle Ω through the axis of rotation
by an angle Ω through the axis of rotation  where
where  is the unit vector defined by
is the unit vector defined by  Through geometric arguments, Goldstein et al. (2002) have shown that this rotation is expressed as
Through geometric arguments, Goldstein et al. (2002) have shown that this rotation is expressed as
 |
(6) |
where  and
and  represent a vector before and after rotation, respectively. Eq. 6—derived by rotating the frame to make the z-axis point along the axis of rotation, applying the rotation around this axis, and then rotating back—is equivalent to our matrix operation of Eq. 5.
represent a vector before and after rotation, respectively. Eq. 6—derived by rotating the frame to make the z-axis point along the axis of rotation, applying the rotation around this axis, and then rotating back—is equivalent to our matrix operation of Eq. 5.
EXAMPLE OF ROTATIONAL ERROR
To compare the performance of the above rotational operators, we consider the geometry illustrated in Fig. 1. Without loss of generality, we assume that the initial local frame  coincides with the lab fixed coordinate system, indicated by the x-, y-, and z-axes. We consider rotations where
coincides with the lab fixed coordinate system, indicated by the x-, y-, and z-axes. We consider rotations where  is directed in the x-y plane. In particular,
is directed in the x-y plane. In particular,  where θ is the polar angle of
where θ is the polar angle of  (see Fig. 1) and the superscript T denotes the vector transpose. The local frame following rotation is denoted by
(see Fig. 1) and the superscript T denotes the vector transpose. The local frame following rotation is denoted by  with rotated
with rotated  component,
component,  The endpoint of
The endpoint of  directed out of the origin, lies on a circle located in the plane at z = cos Ω. The transformation given by Eq. 5 produces the expected result, whereas the approximations
directed out of the origin, lies on a circle located in the plane at z = cos Ω. The transformation given by Eq. 5 produces the expected result, whereas the approximations  and the other variations are not exact, as we next show.
and the other variations are not exact, as we next show.
Specifically, we perform rotations on  at 18° intervals in θ for Ω of 10°, 30°, and 60°. In Fig. 2, we plot the resulting endpoints of
at 18° intervals in θ for Ω of 10°, 30°, and 60°. In Fig. 2, we plot the resulting endpoints of  for the six variations on the combinations of the noncommuting operators in R. Thex- and y-components of the rotated
for the six variations on the combinations of the noncommuting operators in R. Thex- and y-components of the rotated  are plotted separately in Fig. 2; the three concentric circles in each plot correspond to the three different magnitudes of Ω. The solid circles correspond to the exact operator of Eq. 5; the unfilled circles correspond to the R operators as indicated in each plot (a–f). The behavior of the R operator is clearly dependent on the definition used, with the magnitude of the error increasing with the size of the rotation, Ω.
are plotted separately in Fig. 2; the three concentric circles in each plot correspond to the three different magnitudes of Ω. The solid circles correspond to the exact operator of Eq. 5; the unfilled circles correspond to the R operators as indicated in each plot (a–f). The behavior of the R operator is clearly dependent on the definition used, with the magnitude of the error increasing with the size of the rotation, Ω.
FIGURE 2.
Positions of the x- and y-components of  after rotating
after rotating  by
by  The initial local frame
The initial local frame  corresponds to the lab fixed frame (see Fig. 1). Rotations are performed for 20 values of θ between 0° and 360° and Ω of 10°, 30°, and 60°. Solid circles correspond to the exact move; unfilled circles correspond to rotations by each of the six R operators, as indicated in a–f. The R operators depend on the definition used, with the bias and magnitude of error dependent on that choice of R.
corresponds to the lab fixed frame (see Fig. 1). Rotations are performed for 20 values of θ between 0° and 360° and Ω of 10°, 30°, and 60°. Solid circles correspond to the exact move; unfilled circles correspond to rotations by each of the six R operators, as indicated in a–f. The R operators depend on the definition used, with the bias and magnitude of error dependent on that choice of R.
To quantify the error associated with R, we introduce a normalized error measure: 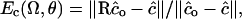 which measures the magnitude in the error of the R-rotated
which measures the magnitude in the error of the R-rotated  vector relative to magnitude of the exact move
vector relative to magnitude of the exact move  Contours of Ec(Ω, θ) for R = Rc Rb Ra are plotted in Fig. 3. As expected, the relative error increases with Ω. The error goes to zero as the rotational angle goes to zero. For other definitions of R, results are similar. This rotational operator is straightforward to implement in the standard Ermak and McCammon algorithm (Ermak and McCammon, 1978) and in higher-order BD updates, where repeated rotation applications may be necessary per time-step.
Contours of Ec(Ω, θ) for R = Rc Rb Ra are plotted in Fig. 3. As expected, the relative error increases with Ω. The error goes to zero as the rotational angle goes to zero. For other definitions of R, results are similar. This rotational operator is straightforward to implement in the standard Ermak and McCammon algorithm (Ermak and McCammon, 1978) and in higher-order BD updates, where repeated rotation applications may be necessary per time-step.
FIGURE 3.
Contours of  for the operator R = Rc Rb Ra in the Ω-θ plane.
for the operator R = Rc Rb Ra in the Ω-θ plane.
CONCLUSION
The commonly used rotation protocol for rigid-body dynamics introduces bias and error into the rotational move and hence the overall simulation. Here, we show that the bias and error are finite, yet diminish for small rotational angles. Our alternative rotational protocol (Eq. 5), applied to chromatin folding (Beard and Schlick, 2001), does not introduce additional numerical complexity and removes such biases and errors. It should thus be used to rotate local coordinate frames in rigid body dynamics.
Acknowledgments
The authors are grateful to Hong Qian for valuable discussions. They also thank Marshall Fixman and Bruce Robinson for comments on an alternative geometric approach.
References
- Allison, S. A. 1991. A Brownian dynamics algorithm for arbitrary rigid bodies. Applications to polarized dynamic light scattering. Macromolecules. 24:530–536. [Google Scholar]
- Beard, D. A., and T. Schlick. 2001. Computational modeling predicts the structure and dynamics of chromatin fiber. Structure. 9:105–114. [DOI] [PubMed] [Google Scholar]
- Chirico, G., and J. Langowski. 1996. Brownian dynamics simulations of supercoiled DNA with bent sequences. Biophys. J. 71:955–971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fernandez, M. X., and J. García de la Torre. 2002. Brownian dynamics of rigid particles of arbitrary shape in external fields. Biophys. J. 83:3039–3048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- García de la Torre, J., and V. A. Bloomfield. 1981. Hydrodynamic properties of complex, rigid, biological macromolecules. Theory and applications. Q. Rev. Biophys. 14:81–139. [DOI] [PubMed] [Google Scholar]
- Ermak, D. L., and J. A. McCammon. 1978. Brownian dynamics with hydrodynamic interactions. J. Chem. Phys. 69:1352–1360. [Google Scholar]
- Goldstein, H., C. Poole, and J. Safko. 2002. Classical Mechanics, 3rd Ed. Addison-Wesley, San Francisco, CA. 134–183.
- Huang, J., and T. Schlick. 2002. Macroscopic modeling and simulations of supercoiled DNA with bound proteins. J. Chem. Phys. 117:8573–8586. [Google Scholar]
- Jian, H., A. Vologodskii, and T. Schlick. 1997. A combined wormlike-chain and bead model for dynamic simulations of long linear DNA. J. Comp. Phys. 136:168–179. [Google Scholar]
- Klenin, K., H. Merlitz, and J. Langowski. 1998. A Brownian dynamics program for the simulation of linear and circular DNA and other wormlike chain polyelectrolytes. Biophys. J. 74:780–788. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu, P. G., B. S. Fujimoto, L. Song, and J. M. Schurr. 1991. Effect of ethidium on the torsion constants of linear and supercoiled DNAs. Biophys. Chem. 41:217–236. [DOI] [PubMed] [Google Scholar]





