Abstract
Cationic lipid-DNA (CL-DNA) complexes are abundantly used in nonviral gene therapy clinical applications. Surface functionality is the next step in developing these complexes as competent, target-specific gene carriers. Poly(ethylene glycol) (PEG) is the natural choice to serve as a protective coat or act as a tether for a specific ligand on the surface of these complexes due to its biocompatibility and ability to convey stealth-like properties. Understanding the effect of PEG on the internal structure and surface properties of CL-DNA complexes is essential in developing vectors with more complex derivatives of PEG, such as Arg-Gly-Asp (RGD)-based peptide-PEG-lipids. We report on x-ray diffraction studies to probe the internal structure of CL-DNA complexes consisting of a ternary mixture of cationic lipids, neutral lipids, and PEG-lipids. The PEG-coated complexes are found to exhibit a structure consistent with the lamellar phase. In addition, three distinct DNA interchain interaction regimes were found to exist, due to a), repulsive long-range electrostatic forces; b), short-range repulsive hydration forces; and c), novel polymer-induced depletion attraction forces in two dimensions. Optical microscopy and reporter gene assays further demonstrate the incorporation of the PEG-lipids into the lamellar CL-DNA complexes under biologically relevant conditions, revealing surface modification. Both techniques show that PEG-lipids with a polymer chain of molecular weight 400 do not provide adequate shielding of the PEGylated CL-DNA complexes, whereas PEG-lipids with a polymer chain of molecular weight 2000 confer stealth-like properties. This surface functionalization is a crucial initial step in the development of competent vectors for in vivo systemic gene delivery and suggests that a second type of surface functionality can be added specifically for targeting by the incorporation of peptide-PEG-lipids.
INTRODUCTION
The structure of cationic lipid-DNA (CL-DNA) complexes has been the object of intense experimental (Koltover et al., 1998; Lasic et al., 1997; Rädler et al., 1997) and theoretical interest (Bruinsma and Mashl, 1998; Harries et al., 1998). Cationic lipids have been shown to be advantageous carriers of DNA in gene delivery (Chesnoy and Huang, 2000; Felgner, 1997; Felgner and Rhodes, 1991; Lin et al., 2003; Mahato and Kim, 2002), although their transfection efficiency, a measure of the successful transfer of a foreign gene to a host cell followed by expression, still remains low. Knowledge of the correlation between the physical attributes of these complexes and their functional activity is crucial in understanding and improving them as synthetic carriers of genes (Ewert et al., 2002; Lin et al., 2000, 2003; Mahato and Kim, 2002; Safinya, 2001). In particular, incorporation of PEG-lipids is an essential aspect in making CL-DNA complexes a viable option as an in vivo gene delivery system (Allen, 1994; Woodle, 1995). The presence of a polymeric shell makes the complexes less susceptible to attack by the immune system, specifically charge-mediated opsonization (Bradley et al., 1998; Lasic and Martin, 1995a), and more persistent in blood circulation due to steric stabilization conferring stealth-like properties (Lasic and Martin, 1995a; Lasic and Papahadjopoulos, 1995b). However, the introduction of a polymeric shell screens the electrostatic interaction between the cationic complex and the negative sulfated proteoglycans on the cell surface (Mislick and Baldeschwieler, 1996). Utilizing ligand-receptor mediated attraction, such as an RGD (GRGDSP) sequence (Suh et al., 2002; Woodle et al., 2001), which is expected to bind with high affinity to the cell's integrins (Midoux and Pichon, 2002), requires a polymeric tether of the specific ligand to the complex. As for stealth properties, PEG again is the natural choice for such a tether because of its hydrophilicity and flexibility, variability in chain length, and the abundance of protocols for its chemical derivatization (Harris and Zalipsky, 1997; Lasic and Martin, 1995a; Zalipsky, 1995).
The mixing of cationic liposomes and DNA results in their spontaneous self-assembly into 0.2-μm size particles of CL-DNA complexes. The driving force for this is the large increase in entropy resulting from the release of tightly bound counterions from the cationic lipid membranes and the anionic DNA rods. In the lamellar CL-DNA complexes, the DNA monolayers, sandwiched between lipid bilayers, are ordered with a well-defined separation distance between adjacent DNA rods (Koltover et al., 2000; Rädler et al., 1997). We show that by tuning different parameters of the lipid membrane, in particular with the introduction of PEG-lipids, we can adjust this DNA distance between 25 Å and 60 Å.
The structure, morphology, and function of PEG-CL-DNA complexes were characterized using x-ray diffraction, optical microscopy, and reporter gene assays, respectively. X-ray diffraction probed the inner structure of the complex, in particular the incorporation of the polymer into the inside of the complex, whereas optical microscopy and reporter gene assays allowed us to investigate the surface functionalization achieved by PEGylation of the complexes.
MATERIALS AND METHODS
Materials
CL-DNA complexes were prepared by mixing liposomes and the appropriate DNA (Koltover et al., 1998; Rädler et al., 1997). For x-ray samples, highly purified λ-phage DNA (48,502 bp), purchased from New England Biolabs (Beverly, MA), was used. For microscopy and transfection experiments, plasmid DNA containing the luciferase gene was used (pGL3-Control vector, Promega, Madison, WI). DOTAP (1,2-dioleoyl-3-trimethyl-ammonium-propane, molecular weight 698.55) and the neutral “helper” lipid DOPC (1,2-dioleoyl-sn-glycero-3-phosphatidylcholine, 786.13 mol wt) were purchased from Avanti Polar Lipids (Alabaster, AL). A novel set of PEG-lipids, some of which incorporate two charged amino groups at the end of the polymeric chain, was synthesized according to the procedure previously described (Schulze et al., 1999). Two different cationic PEG-lipids, PEG4002+-lipid (〈n〉 = 9, 1394 mol wt) and PEG20002+-lipid (〈n〉 = 45, 3010 mol wt) were investigated. These were compared with similar neutral PEG-lipids, PEG400-lipid (〈n〉 = 9, 1035 mol wt) and PEG2000-lipid (〈n〉 = 45, 2635 mol wt). The chemical structures of these molecules are shown in Fig. 1. Microscopy and transfection experiments were done with PEG2000-lipid (1,2-dioleoyl-sn-glycero-3-phosphoethanolamine-N-[poly(ethylene glycol)-2000]) (〈n〉 = 45, 2802 mol wt), purchased from Avanti Polar Lipids.
FIGURE 1.
Chemical structures and names of the PEG-lipids used in this study.
Liposomes
Lipid mixtures were prepared by dissolving the lipids in a chloroform:methanol mixture and then mixing the three components in the desired ratio, keeping the mol fraction of DOTAP constant for any given line of data. Microscopy samples required the inclusion of 0.2 wt % Texas Red DHPE fluorescent label purchased from Molecular Probes (Eugene, OR). The solvents were then evaporated, first under a stream of nitrogen and subsequently in a high vacuum to ensure removal of all solvents. The lipid mixtures were then hydrated with the appropriate amount of deionized water (final concentration of 25 mg/ml for x-ray samples; final concentration of 0.5 mg/ml for transfection and microscopy samples), incubated at 37°C for at least 6 h, sonicated for 8–10 min with a VibraCell from Sonics and Materials (Newtown, CT), and filtered through a 0.2-μm Whatman filter (Maidstone, UK). The obtained liposome solutions were then stored at 4°C.
X-ray diffraction
PEG-CL-DNA complexes were prepared by mixing 50–100 μg of λ-phage DNA at 5 mg/ml with liposome mixtures at 25 mg/ml in an Eppendorf centrifuge for ∼3 h. Samples were prepared at the isoelectric (charge neutral) point. After three days of storage at 4°C to allow the samples to reach equilibrium, the samples were transferred to 1.5-mm diameter quartz x-ray capillaries.
The high-resolution x-ray diffraction experiments were carried out at the Stanford Synchrotron Radiation Laboratory at an x-ray energy of 10 KeV using a double bounce Si (111) crystal. Powder diffraction measurements were taken with a PMT Bicron detector (Paris, France).
Optical microscopy
PEG-CL-DNA complexes were prepared by mixing 2 μg of pGL3 plasmid DNA at 0.1 mg/ml, fluorescently tagged with YOYO dye purchased from Molecular Probes, with Texas Red DHPE labeled PEG-CLs at 0.5 mg/ml. Samples were prepared in Dulbecco's modified Eagle's medium (DMEM; Gibco, Gaithersburg, MD) at a charge ratio of 2.8 to mimic the conditions of in vitro experiments. Differential interference contrast and fluorescence images were taken with a Nikon Diaphot 300 microscope (Nikon, Tokyo, Japan) and imaged with a QE Sensicam (Cooke, Auburn Hills, MI).
Cell transfection
Mouse fibroblast L-cells were cultured in DMEM supplemented with 1% (v/v) penicillin-streptomycin (Gibco BRL) and 5% (v/v) Fetal Bovine Serum (Gibco BRL) at 37°C in a 5% CO2 atmosphere, splitting the cells every 2–4 days to maintain monolayer coverage. The cells were transfected at a confluency of 60–80% in six well plates (35 mm diameter per well). Using liposome (0.5 mg/ml) and DNA (1 mg/ml) stock solutions, liposome and DNA stocks were further diluted in DMEM to a final volume of 0.5 ml and complexes, containing 2 μg of pGL3-DNA, were prepared at a cationic/anionic charge ratio of 2.8. The cells were incubated with complexes for 6 h, rinsed 3 times with PBS (Phosphate Buffered Saline, Gibco BRL), and incubated in supplemented DMEM for an additional 24 h (sufficient for a complete cell cycle) to allow expression of the luciferase gene. Luciferase gene expression was measured with the Luciferase Assay System from Promega and light output readings were performed on a Berthold AutoLumat luminometer (Wallac, Gaithersburg, MD). Transfection efficiency was normalized to milligrams of total cellular protein using the Bio-Rad Protein Assay Dye Reagent (Bio-Rad, Hercules, CA) and is expressed as relative light units per mg of total cellular protein.
RESULTS AND DISCUSSION
Representative small angle x-ray scattering (SAXS) scans for the PEG-CL-DNA complexes are shown in Fig. 2. All complexes were prepared at the isoelectric point at a fixed MDOTAP = 0.33. To study the effect of PEG-lipids, we increased their molar fraction (MPEG-lipid) at fixed molar fraction of DOTAP (MDOTAP). Therefore, the contribution of DOTAP to the membrane charge density remains the same. The reference scan at the bottom of Fig. 2 shows the typical powder-average diffraction pattern without added PEG-lipid. The set of periodically spaced sharp peaks (q00n) originates from the lipid lamellar structure, with an interlayer spacing, d = 2π/q001 = 68.3 Å, that corresponds to the membrane thickness (δm) plus water gap (δw) between the lipid bilayer. The broader peak at qDNA, marked with an arrow, arises from the 2D smectic ordering of the DNA rods, separated by a distance dDNA = 2π/qDNA = 47.6 Å. The second order of the DNA-DNA correlation peak is also visible (DNA II). The dotted line in Fig. 2 corresponds to the position of the DNA peak for the sample without PEG-lipid.
FIGURE 2.
Synchrotron SAXS scans for isoelectric PEG-CL-DNA complexes at fixed MDOTAP = 0.33 as a function of increasing concentration of (A) PEG4002+-lipid; (B) PEG20002+-lipid; (C) PEG400-lipid; and (D) PEG2000-lipid. A reference scan of a CL-DNA complex without PEG-lipid is shown at the bottom (shaded plot). The dotted line marks the DNA correlation peak for the reference scan, and the arrows mark the DNA correlation peaks for the others. With the exception of PEG400-lipid, the addition of PEG-lipid leads to a decrease in the DNA spacing compared to the reference scan.
Fig. 2 A shows the effect of incorporating PEG4002+-lipid into CL-DNA complexes. SAXS scattering profiles reveal one phase of stable complexes with a well-ordered lamellar structure. The addition of PEG4002+-lipid at fixed MDOTAP leads to a shift of the DNA peak to higher values of q, corresponding to a tighter packing of the DNA rods. A similar trend is observed with the inclusion of PEG20002+-lipid as shown in Fig. 2 B, but we also detect the existence of two different phases at higher MPEG20002+-lipid values (∼6%). Figs. 2, C and D, show the effect of incorporating neutral PEG400-lipid and PEG2000-lipid. In the case of PEG400-lipid we see that the DNA-spacing remains constant, within error, compared to the reference sample without PEG-lipid, whereas PEG2000-lipid behaves similar to PEG20002+-lipid. We examine this behavior below.
Fig. 3 shows the variation of the distance between DNA chains with increasing molar fraction of PEG4002+-lipid at fixed MDOTAP. Different symbols correspond to different values of MDOTAP. The series at lower MDOTAP show a large decrease in dDNA as a function of increasing molar fraction of PEG4002+-lipid, indicating a significant compaction of the DNA rods. For higher MDOTAP almost no variation is observed (Fig. 3 A).
FIGURE 3.
Interaxial DNA spacing, dDNA, results for CL-DNA complexes incorporating PEG4002+-lipid at various MDOTAP values, obtained from SAXS data. (A) Decrease of the DNA interaxial distance, dDNA, with increasing molar fraction of PEG4002+-lipid at different fixed molar fractions of DOTAP. (B) The experimental dDNA values represented in A are plotted against the membrane charge density (symbols as in A). The solid circles represent the experimental dDNA for the reference system (DOTAP/DOPC without PEG-lipid) and increasing MDOTAP (from left to right, 0.27, 0.33, 0.38, 0.43, 0.53, 0.63, and 0.82). The continuous line represents the predicted dDNA, calculated as described in the text. The data for complexes with added PEG4002+-lipid follow this curve exactly, implying that PEG4002+-lipid behaves as a cationic colipid. The inset shows that no further condensation of DNA occurs upon addition of neutral PEG400-lipid at fixed MDOTAP = 0.33. This verifies the domination of electrostatic interactions.
Fig. 3 B collects all the experimental dDNA data from Fig. 3 A and plots it as a function of the membrane charge density, σM, of the lipid bilayer. This parameter was calculated taking into account the contribution of charge from both DOTAP and the cationic PEG-lipid, σM = e(1NDOTAP + 2NPEG-lipid)/(ADOTAPNDOTAP + ADOPCNDOPC + APEG-lipidNPEG-lipid). Assuming that the headgroup size is the same for all lipids (Alipid = 72 Å (Gruner et al., 1988; Tristram-Nagle et al., 1998)), this equation simplifies to σM = e(MDOTAP + 2MPEG-lipid)/Alipid. The solid circles, corresponding to dDNA for the reference system complexes (no PEG-lipids), show a perfect agreement with predicted dDNA values, represented by the continuous line. These predicted values were obtained for isoelectric complexes, taking into account only electrostatic interactions and geometric contributions. Complex formation, driven by counterion release, dictates that the DNA chains should occupy all the available membrane area while matching the charges of the cationic lipids. As reported earlier (Koltover et al., 1999; Rädler et al., 1997), by defining a unit cell in the lamellar complex (assuming an equidistant smectic array of DNA rods) and equating the charge density contributions of the lipid bilayer and the DNA rods, we obtain the average spacing between DNA chains for an isoelectric complex (dDNAiso):
 |
where ρDNA = 1.7 g/cm3 and ρlipid = 1.07 g/cm3 are the densities of lipid and DNA respectively; ADNA is the cross section area of a DNA molecule, ρiso = 2.2 is the weight ratio of DOTAP and DNA for stoichiometrically charge neutral complexes, and ΦDOPC is the weight fraction of DOPC.
The experimental values for dDNA collected in Fig. 3 B reveal the existence of two distinct DNA interaction regimes for CL-DNA complexes with short cationic PEG-lipids (PEG4002+-lipid). For 0.27 < MDOTAP < 0.43 and 0 < MPEG4002+-lipid < 9%, the results show very good agreement with the dDNA predicted values. This defines the electrostatic regime, where dDNA depends purely on the membrane charge density (σM) for a complex at the isoelectric point. For MDOTAP > 0.53 a strong repulsive hydration barrier (Salditt et al., 1998) between DNA rods becomes relevant, making it impossible to further condense the DNA by adding PEG4002+-lipid.
These results indicate that PEG4002+-lipid is acting as a cationic colipid, leading to condensation of DNA through an increase in the membrane charge density. As mentioned earlier, the inset in Fig. 3 B, comparing the addition of cationic PEG4002+-lipid with neutral PEG400-lipid, shows that condensation only occurs when cationic PEG-lipid is used. This suggests that the PEG chain length of PEG4002+-lipid (〈n〉 = 9) is too short to show any polymeric behavior, even in a 2D system. Instead, repulsive electrostatics and hydration forces dominate the system as found in the reference system without PEG-lipids (Salditt et al., 1998). We can further conclude that PEG4002+-lipid completely incorporates into the CL-DNA complexes since it forms one phase even at high amounts of PEG4002+-lipid (MPEG4002+-lipid = 30%). Finally, we conclude that the condensation of DNA by different lipid systems forming lamellar complexes follows a universal charge-condensation curve in the electrostatic regime (dDNA > 31 Å) (Lin et al., 2003).
We now examine the influence of a PEG-lipid with a longer polymer chain. Fig. 4 A shows the variation of the distance between DNA chains in single-phase PEG-CL-DNA complexes upon adding PEG20002+-lipid for various series of fixed MDOTAP. These results seem similar to those obtained with PEG4002+-lipid, but when plotted as a function of membrane charge density, significant differences become apparent.
FIGURE 4.
Interaxial DNA spacing, dDNA, results for CL-DNA complexes incorporating PEG20002+-lipid at various MDOTAP values, obtained from SAXS data. (A) Decrease of the DNA interaxial distance dDNA with increasing molar fraction of PEG20002+-lipid at different molar fractions of DOTAP. (B) Experimental dDNA data from A are collected and plotted against the membrane charge density (symbols as in A). The continuous line represents the predicted dDNA, calculated as described in the text. The data for complexes with added PEG20002+-lipid show a DNA spacing smaller than the predicted value, implying enhanced DNA condensation. The inset shows similar behavior for the inclusion of neutral PEG2000-lipid and PEG20002+-lipid with increasing concentration of PEG-lipid at fixed MDOTAP = 0.33. This verifies the domination of polymer-induced depletion attraction interactions.
For complexes incorporating the long-chain cationic PEG-lipid (PEG20002+-lipid) at 0.27 < MDOTAP < 0.43 and 0 < MPEG20002+-lipid < 4.5%, we find the existence of a depletion attraction interaction regime (Fig. 4 B). The distances between DNA rods, as measured from x-ray scattering, deviate from the universal charge-condensation curve to lower values as MPEG20002+-lipid is increased. This indicates the existence of an additional attractive force that decreases the DNA-DNA spacing further than the membrane charge density predicts. The enhanced condensation can be explained by taking into account the existence of a depletion attraction interaction. This attraction is due to the presence of a polymer chain in the confined space between the lipid bilayers where the DNA chains reside. For complexes with MDOTAP > 0.53, we reproduce the results found in the PEG4002+-lipid system. This demonstrates that the repulsive hydration forces dominate the interactions between the DNA rods at short range, independent of the PEG-lipid chain length.
Unlike for the short-chain PEG, the inset in Fig. 4 B shows that for PEG2000-lipid, the cationic and the neutral lipid both yield the same result. The DNA spacing, plotted for MDOTAP = 0.33, decreases with increasing MPEG20002+-lipid and MPEG2000-lipid. These results show that the polymer chain-DNA interaction dominates in this system, as opposed to electrostatics as seen in the case of PEG4002+-lipid and PEG400-lipid. The data also confirms that the depletion attraction force is indeed responsible for the additional DNA condensation with increasing amounts of MPEG20002+-lipid, even at remarkably low concentrations of PEG-lipid (MPEG20002+-lipid < 1%).
The depletion attraction phenomenon (Vroege and Lekkerkerker, 1992) in our system is analogous to the polymer-induced condensation of DNA solutions (Vasilevskaya et al., 1995). In our experiments, we find that when PEG2000-lipid is incorporated within the CL-DNA complexes, it induces a depletion attraction force between the DNA rods adsorbed on the cationic membranes because the DNA has a depletion region around it, which excludes the PEG2000 part of the PEG-lipid molecule (Fig. 5 A). This depletion attraction force causes a phase separation between the polymer and the DNA rods within the layers inside the complex as shown schematically in Fig. 5 B. The complex has DNA-rich domains, shown in green, and polymer-rich domains, shown in blue (Fig. 5 C). Our predicted values of dDNA, from the universal charge-condensation curve, dictate a water gap between DNA rods of 5–35 Å to accommodate the added polymer. As PEG2000 has an Rg of 35 Å (Devanand and Selser, 1991; Warriner et al., 1996), we should expect PEG2000-lipid to be excluded from regions between DNA rods (Israelachvili, 1992). Thus, the closer DNA spacing observed in our data is a result of the osmotic stress exerted on the DNA domains due to the PEG2000 chains. This is the 2D analog of what happens in 3D solutions of DNA containing the polymer PEG where depletion attraction induces bundling of DNA with the polymer PEG being excluded from the bundle region. Again, the polymer PEG is known to exert an osmotic stress on the bundle phase, which increases as the concentration of polymer is increased, and leads to the DNA interaxial spacing decreasing.
FIGURE 5.
Schematic of a lamellar CL-DNA complex containing long-chain PEG-lipids. (A) The presence of the polymer is forcing the DNA rods closer together than predicted by electrostatics (depletion attraction force). (B) Cross section of PEG-CL-DNA complex with DNA rich domains (green) and polymer rich domains (blue) between lipid bilayers (gray). (C) Enlarged view of PEG-CL-DNA complex showing internal phase separation as well as the outer shell of polymer chains.
The novel 2D depletion attraction is only seen with the higher molecular weight polymer, PEG2000-lipid, as the polymer chain of the PEG400-lipids is too short to induce phase separation. Indeed, we find that CL-DNA complexes containing PEG4002+-lipid follow the predictions of the universal charge-condensation curve, which implies that the PEG400-chains are mixed with the DNA chains.
To investigate the incorporation and behavior of PEG-lipids on the surface of CL-DNA complexes, we used optical microscopy as well as established transfection protocols. All of these experiments were conducted in DMEM, a biologically relevant cell culturing medium containing ∼150 mM monovalent salt, at a charge ratio of +2.8 (leading to positively charged complexes).
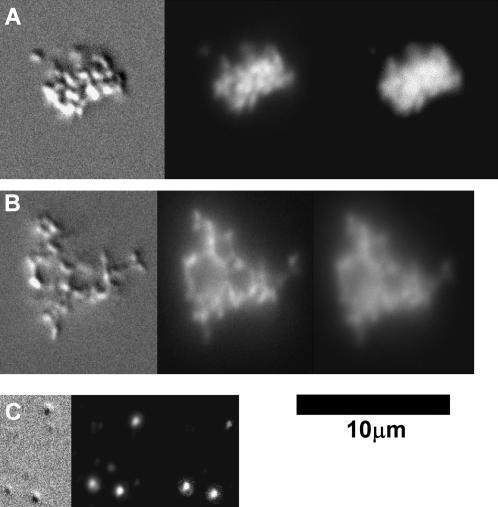
Optical microscopy at MDOTAP = 0.82 confirms the existence of complexes through colocalization of DNA and lipid and further demonstrates the different behavior of the complexes depending on polymer chain length. Fig. 6 A shows complexes without PEG-lipid. Some degree of aggregation of the complexes is observed, due to the presence of DMEM. Fig. 6 B shows a complex at MPEG400-lipid = 10%. Again the aggregation of the complexes is clearly evident. Only for complexes prepared using the longer polymer lipid at MPEG2000-lipid = 10% do we see an effect of the PEG-lipid. No aggregation of complex particles is observed due to steric repulsion conferred by a shell of PEG2000-lipid polymer chains even in the presence of salt. This shows that the PEG-lipid is coating the surfaces of the complexes, whereas x-ray diffraction results clearly demonstrate that the PEG-lipid is located internally, as well. As mentioned earlier, this polymeric stabilization is important for developing a viable in vivo gene delivery system (Allen, 1994; Harvie et al., 2000; Pitard et al., 2001; Wheeler et al., 1999).
FIGURE 6.
(A–C) Microscopy images of positive (charge ratio = 2:8) DOTAP/DOPC CL-DNA complexes at MDOTAP = 0.33 with plasmid DNA in the presence of the cell culture medium DMEM, taken in differential interference contrast (left), lipid fluorescence (center), and DNA fluorescence modes. The images show complexes prepared (A) without PEG-lipid, (B) with 10% PEG400-lipid, and (C) with 10% PEG2000-lipid. The aggregation of particles that is seen when no PEG-lipid or 10% PEG400-lipid are added is eliminated by the addition of 10% PEG2000-lipid, indicating steric stabilization by the long-chain polymer.
Fig. 7 shows experiments using PEG-CL-DNA complexes as vectors for gene delivery. The transfection efficiency (TE) was measured in relative light units using a standard luciferase assay system. Nonspecific electrostatic attraction between the positively charged CL-DNA complexes (the CL-DNA charge ratio in our transfection experiments is +2.8) and the negatively charged cell membrane is important for transfection because it leads to attachment of the complexes to the cell. The relationship between TE and lipid composition of the two-component DOTAP/DOPC lipid system is well established (Lin et al., 2003), showing low and high TE in regions of low σM and high σM, respectively. We investigated the effect of added PEG-lipid in both of these regimes.
FIGURE 7.
Transfection efficiency in fibroblast L-cells as a function of increasing molar fraction of PEG-lipid (MPEG-lipid) added to a DOTAP/DOPC liposome mixture. (A) When initial TE is low, in the low σM regime at MDOTAP = 0.30, addition of PEG400-lipid (open squares) does not effect TE whereas adding the PEG4002+-lipid (open circles) increases TE due to an increase in the overall σM of the DOTAP/DOPC/PEG-lipid system. The PEG2000-lipids (solid squares) and PEG20002+-lipids (solid circles) both decrease TE, due to the presence of the long PEG chains on the CL-DNA surface. (B) In the high σM regime at MDOTAP = 0.80, TE is already at a maximum and addition of PEG4002+-lipid does not aid transfection. Adding PEG20002+-lipid or PEG2000-lipid leads to a decrease in TE by nearly two orders of magnitude with only 6% added PEG-lipid. These findings show that the PEG2000-lipid and PEG20002+-lipid are coating the CL-DNA particles. As a reference, typical TE for naked DNA is on the order of 300,000 RLU/mg protein.
In the low TE regime, i.e., at a fixed MDOTAP = 0.30 (Fig. 7 A), the efficiency behavior varies for the two PEG chain lengths. Addition of neutral PEG400-lipid does not alter TE, whereas the addition of cationic PEG4002+-lipid increases efficiency by acting as a cationic colipid and increasing σM, as evident from the x-ray data (Fig. 3, inset). By contrast, a decrease in the already low efficiency occurs with the addition of long-chain PEG-lipids (PEG2000-lipid), regardless of the presence of cationic charges on the PEG-lipid. At 2% added PEG2000-lipid, TE is less affected by the cationic lipid, but at 6% PEG-lipid, the effects of the neutral and charged PEG2000-lipid are similar. A fraction of the PEG2000-lipid is most likely coating the outside of the CL-DNA complexes, thus shielding cell-complex interaction and subsequent transfection. Our data indicates that PEG coverage of the CL-DNA complexes occurs with 6% PEG2000-lipid. Data points were not taken beyond 10% in anticipation of phase separation between CL-DNA complexes and the PEG2000-lipids. In direct contrast, the CL-DNA complexes can accommodate up to 20% PEG400-lipid without phase separation, but there is no indication that shielding occurs with these shorter amphiphiles.
The long-chain PEG-lipids show a similar behavior in the high TE regime at MDOTAP = 0.80 (Fig. 7 B). Transfection efficiency is high without added PEG-lipid but decreases nearly two orders of magnitude upon addition of 6% PEG2000-lipid, either cationic or neutral. The addition of cationic PEG4002+-lipid in this case does not lead to higher efficiency; in fact, both cationic and neutral PEG400-lipid behave similarly, only negligibly affecting TE. In this high initial σM regime, TE already is at a maximum and further addition of cationic colipid does not aid transfection (Lin et al., 2003).
These results are promising and point to future experiments. The transfection data, in combination with optical microscopy, proves that PEG2000 is coating the CL-DNA complexes. A cell-specific targeting component, i.e., a peptide sequence such as RGD, could be tethered to the PEG2000 molecules to promote specific attachment to cells while retaining sterically stabilized PEG2000-CL-DNA complexes.
CONCLUSIONS
The inclusion of a biocompatible polymer shell is essential in developing a synthetic gene delivery system that can successfully evade the immune system for systemic in vivo applications. Furthermore, it is important to have the ability to introduce polymer chains of variable length to act as tethers for target-specific peptide sequences, increasing the complexes' functionality. In such a case, the cationic lipid acts as the DNA condensing agent whereas the functionalized PEG-lipids are responsible for the cell-complex interaction. Thus, understanding the effects of incorporating PEG-lipids into CL-DNA complexes is crucial for the next step in gene delivery by CL-DNA complexes—adding target specificity with derivatives of PEG-lipids. Investigating a model system for such PEG-CL-DNA complexes, we have shown that a critical value of polymer chain length exists above which stealth properties and steric stabilization becomes evident.
Furthermore, for the first time we have shown the existence of three distinct interaction regimes within the lamellar structure of the CL-DNA complexes, where repulsive long-range electrostatic forces, short-range repulsive hydration forces, and polymer-induced attractive depletion forces dominate the interaction between DNA molecules. By tuning certain parameters, such as the polymer length and the membrane charge density, we can access any of these regimes.
Acknowledgments
This work was supported by National Institutes of Health grant GM-59288, and National Science Foundation grants DMR-0203755 and CTS 0103516. The Stanford Synchrotron Radiation Laboratory is supported by the U.S. Department of Energy. The Materials Research Laboratory at University of California Santa Barbara is supported by National Science Foundation grant DMR-0080034.
Ana Martin-Herranz's present address is Unilever R&D, Manuel de Falla 7, 28036 Madrid, Spain.
References
- Allen, T. M. 1994. Long-circulating (sterically stabilized) liposomes for targeted drug-delivery. Trends Pharmacol. Sci. 15:215–220. [DOI] [PubMed] [Google Scholar]
- Bradley, A. J., D. V. Devine, S. M. Ansell, J. Janzen, and D. E. Brooks. 1998. Inhibition of liposome-induced complement activation by incorporated poly(ethylene glycol) lipids. Arch. Biochem. Biophys. 357:185–194. [DOI] [PubMed] [Google Scholar]
- Bruinsma, R., and J. Mashl. 1998. Long-range electrostatic interaction in DNA cationic lipid complexes. Europhys. Lett. 41:165–170. [Google Scholar]
- Chesnoy, S., and L. Huang. 2000. Structure and function of lipid-DNA complexes for gene delivery. Annu. Rev. Biophys. Biomol. Struct. 29:27–47. [DOI] [PubMed] [Google Scholar]
- Devanand, K., and J. C. Selser. 1991. Asymptotic behavior and long-range interactions in aqueous solutions of poly(ethylene oxide). Macromolecules. 24:5943–5947. [Google Scholar]
- Ewert, K., A. Ahmad, H. M. Evans, H. W. Schmidt, and C. R. Safinya. 2002. Efficient synthesis and cell-transfection properties of a new multivalent cationic lipid for nonviral gene delivery. J. Med. Chem. 45:5023–5029. [DOI] [PubMed] [Google Scholar]
- Felgner, P. L. 1997. Nonviral strategies for gene therapy. Sci. Am. 276:102–106. [DOI] [PubMed] [Google Scholar]
- Felgner, P. L., and G. Rhodes. 1991. Gene therapeutics. Nature. 349:351–352. [DOI] [PubMed] [Google Scholar]
- Gruner, S. M., M. W. Tate, G. L. Kirk, P. T. C. So, D. C. Turner, D. T. Keane, C. P. S. Tilcock, and P. R. Cullis. 1988. X-ray diffraction study of the polymorphic behavior of N-methylated dioleoylphosphatidylethanolamine. Biochemistry. 27:2853–2866. [DOI] [PubMed] [Google Scholar]
- Harries, D., S. May, W. M. Gelbart, and A. Ben-Shaul. 1998. Structure, stability, and thermodynamics of lamellar DNA-lipid complexes. Biophys. J. 75:159–173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harris, J. M., and S. Zalipsky, editors. 1997. Poly(ethylene glycol) Chemistry and Biological Applications. American Chemical Society, Washington, DC.
- Harvie, P., F. M. P. Wong, and M. B. Bally. 2000. Use of poly(ethylene glycol)-lipid conjugates to regulate the surface attributes and transfection activity of lipid-DNA particles. J. Pharm. Sci. 89:652–663. [DOI] [PubMed] [Google Scholar]
- Israelachvili, J. 1992. Intermolecular and Surface Forces. Academic Press, London.
- Koltover, I., T. Salditt, J. O. Rädler, and C. R. Safinya. 1998. An inverted hexagonal phase of cationic liposome-DNA complexes related to DNA release and delivery. Science. 281:78–81. [DOI] [PubMed] [Google Scholar]
- Koltover, I., T. Salditt, and C. R. Safinya. 1999. Phase Diagram, Stability, and Overcharging of Lamellar Cationic Lipid-DNA Self-Assembled Complexes. Biophys. J. 77:915–924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koltover, I., K. Wagner, and C. R. Safinya. 2000. DNA condensation in two dimensions. Proc. Natl. Acad. Sci. USA. 97:14046–14051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lasic, D. D., and F. J. Martin, editors. 1995a. Stealth Liposomes. CRC Press, Boca Raton, FL.
- Lasic, D. D., and D. Papahadjopoulos. 1995b. Liposomes revisited. Science. 267:1275–1276. [DOI] [PubMed] [Google Scholar]
- Lasic, D. D., H. Strey, M. C. A. Stuart, R. Podgornik, and P. M. Frederik. 1997. The structure of DNA-liposome complexes. J. Am. Chem. Soc. 119:832–833. [Google Scholar]
- Lin, A. J., N. L. Slack, A. Ahmad, C. X. George, C. E. Samuel, and C. R. Safinya. 2003. Three-dimensional imaging of lipid gene-carriers: membrane charge density controls universal transfection behavior in lamellar cationic liposome-DNA complexes. Biophys. J. 84:3307–3316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin, A. J., N. L. Slack, A. Ahmad, I. Koltover, C. X. George, C. E. Samuel, and C. R. Safinya. 2000. Structure and structure-function studies of lipid/plasmid DNA complexes. J. Drug Target. 8:13–27. [DOI] [PubMed] [Google Scholar]
- Mahato, R. I., and S. W. Kim, editors. 2002. Pharmaceutical Perspectives of Nucleic Acid-Based Therapeutics. Taylor & Francis, London.
- Midoux, P., and C. Pichon. 2002. Peptide-based gene delivery systems. In Pharmaceutical Perspectives of Nucleic Acid-Based Therapeutics. S. W. Kim, editor. Taylor & Francis, London. 334–366.
- Mislick, K. A., and J. D. Baldeschwieler. 1996. Evidence for the role of proteoglycans in cation-mediated gene transfer. Proc. Natl. Acad. Sci. USA. 93:12349–12354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pitard, B., N. Oudrhiri, O. Lambert, E. Vivien, C. Masson, B. Wetzer, M. Hauchecorne, D. Scherman, J. L. Rigaud, J. P. Vigneron, J. M. Lehn, and P. Lehn. 2001. Sterically stabilized BGTC-based lipoplexes: structural features and gene transfection into the mouse airways in vivo. J. Gene Med. 3:478–487. [DOI] [PubMed] [Google Scholar]
- Rädler, J. O., I. Koltover, T. Salditt, and C. R. Safinya. 1997. Structure of DNA-cationic liposome complexes: DNA intercalation in multilamellar membranes in distinct interhelical packing regimes. Science. 275:810–814. [DOI] [PubMed] [Google Scholar]
- Safinya, C. R. 2001. Structures of lipid-DNA complexes: supramolecular assembly and gene delivery. Curr. Opin. Struct. Biol. 11:440–448. [DOI] [PubMed] [Google Scholar]
- Salditt, T., I. Koltover, O. Radler, and C. R. Safinya. 1998. Self-assembled DNA-cationic-lipid complexes: Two-dimensional smectic ordering, correlations, and interactions. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 58:889–904. [Google Scholar]
- Schulze, U., H. W. Schmidt, and C. R. Safinya. 1999. Synthesis of novel cationic poly(ethylene glycol) containing lipids. Bioconjug. Chem. 10:548–552. [DOI] [PubMed] [Google Scholar]
- Suh, W., S. O. Han, L. Yu, and S. W. Kim. 2002. An angiogenic, endothelial-cell-targeted polymeric gene carrier. Mol. Ther. 6:664–672. [PubMed] [Google Scholar]
- Tristram-Nagle, S., H. I. Petrache, and J. F. Nagle. 1998. Structure and interactions of fully hydrated dioleoylphosphatidylcholine bilayers. Biophys. J. 75:917–925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vasilevskaya, V. V., A. R. Khokhlov, Y. Matsuzawa, and K. Yoshikawa. 1995. Collapse of single DNA molecule in poly(ethylene glycol) solutions. J. Chem. Phys. 102:6595–6602. [Google Scholar]
- Vroege, G. J., and H. N. W. Lekkerkerker. 1992. Phase transitions in lyotropic colloidal and polymer liquid crystals. Rep. Prog. Phys. 55:1241–1309. [Google Scholar]
- Warriner, H. E., S. H. J. Idziak, N. L. Slack, P. Davidson, and C. R. Safinya. 1996. Lamellar biogels: fluid-membrane-based hydrogels containing polymer lipids. Science. 271:969–973. [DOI] [PubMed] [Google Scholar]
- Wheeler, J. J., L. Palmer, M. Ossanlou, I. MacLachlan, R. W. Graham, Y. P. Zhang, M. J. Hope, P. Scherrer, and P. R. Cullis. 1999. Stabilized plasmid-lipid particles: construction and characterization. Gene Ther. 6:271–281. [DOI] [PubMed] [Google Scholar]
- Woodle, M. C. 1995. Sterically stabilized liposome therapeutics. Adv. Drug Deliv. Rev. 16:249–265. [Google Scholar]
- Woodle, M. C., P. Scaria, S. Ganesh, K. Subramanian, R. Titmas, C. Cheng, J. Yang, Y. Pan, K. Weng, C. Gu, and S. Torkelson. 2001. Sterically stabilized polyplex: ligand-mediated activity. J. Control. Release. 74:309–311. [DOI] [PubMed] [Google Scholar]
- Zalipsky, S. 1995. Functionalized poly(ethylene glycol) for preparation of biologically relevant conjugates. Bioconjug. Chem. 6:150–165. [DOI] [PubMed] [Google Scholar]