Abstract
The interaction of the αα, ββ, and αβ smooth muscle tropomyosin (Tm) isoforms with F-actin was systematically studied in the absence and in the presence of myosin subfragment 1 (S1) using multifrequency phase/modulation Förster resonance energy transfer (FRET). A Gaussian double distance distribution model was adopted to fit FRET data between a 5-(2-iodoacetyl-amino-ethyl-amino)naphthalene-1-sulfonic acid donor at either Cys-36 of the β-chain or Cys-190 of the α-chain and a 4-dimethylaminophenylazophenyl 4′-maleimide acceptor at Cys-374 of F-actin. Experimental data were obtained for singly and doubly labeled αβ Tm (donor only at α, only at β, or both) and for doubly labeled αα or ββ Tm. Data for singly labeled αβTm were combined in a global analysis with doubly labeled αβTm. In all doubly labeled isoforms, upon S1 binding, one donor-acceptor “apparent” distance increased slightly by 0.5–2 Å, whereas the other decreased by 6–9 Å. These changes are consistent with a uniform “rolling” motion of Tm over the F-actin surface. The analysis indicates that Tm occupies relatively well-defined positions, with some flexibility, in both the predominantly closed (−S1) and open (+S1) thin-filament states. The results for the αβTm heterodimer indicate that the local twofold symmetry of αα or ββ Tm is effectively broken in αβTm bound to F-actin, which implies a difference between the α- and β-chains in terms of their interaction with F-actin.
INTRODUCTION
Biochemical studies have shown that the F-actin skeletal tropomyosin-troponin (F-actin·Tm·Tn) thin filament equilibrates between three states: blocked, closed, and open (McKillop and Geeves, 1993). The equilibrium is shifted from mostly blocked to mostly closed by Ca2+, allowing myosin heads to bind in a weak complex with F-actin. The isomerization of myosin heads to a strong complex with F-actin shifts the system to the fully open or active state. Thus, myosin heads cooperatively “turn on” the system assisted by Ca2+ (Lehrer and Geeves, 1998; Geeves and Lehrer, 2002). Electron microscopy studies have shown that skeletal Tm takes on three positions on F-actin associated with the three biochemical states (Lehman et al., 2000). Recently, we have used Förster resonance energy transfer (FRET), in the frequency domain, to obtain data in solution that shows that Ca2+ causes movement of skeletal muscle Tm and that the degree of movement is in reasonable agreement with the structural data. The movement appears to be a “rolling” motion around the actin filament (Bacchiocchi and Lehrer, 2002).
Smooth muscle does not contain troponin and is primarily regulated by the phosphorylation of myosin (Kamm and Stull, 1985). With steady-state FRET, Graceffa obtained evidence for gizzard muscle Tm movement upon the binding of myosin heads to actin (Graceffa, 1999), which appeared to be sensitive to phosphorylation (Graceffa, 2000), suggesting that phosphorylation and Tm movement are both involved in smooth muscle regulation. Recent electron microscopy studies of different Tm isoforms in troponin-free thin filaments (Lehman et al., 2000) have shown that minor amino acid sequence differences among isoforms can modify the position of Tm on actin due to the small free energy difference between the blocked and closed states. This may explain why electron microscopy located smooth Tm in a blocking position near the outer domain of actin even though biochemical studies indicated that it is mainly in the closed state (Maytum et al., 2001).
Solution studies have revealed differences in interaction and stability among the various skeletal and smooth Tm isoforms (αβ, αα, and ββ). In skeletal Tm, the stability of the different isoforms is quite similar whereas in smooth Tm the most thermodynamically stable isoform is αβ (Lehrer and Stafford, 1991). In skeletal muscle the α/β ratio is >1:1 and the Tm isoforms present are mainly a mixture of αα and αβ with a minimal amount of ββ. In smooth muscle the α/β ratio is 1:1 and Tm is almost entirely formed by the αβ heterodimer (for a review, see e.g., Smillie, 1996, and references therein). The head-to-tail interaction for smooth Tm is stronger than in skeletal Tm and is associated with stronger binding to actin (Jancso and Graceffa, 1991) and a regulatory cooperative unit twice as large (Lehrer et al., 1997).
Despite the above studies, the differences in structural and interaction properties among the various Tm isoforms, in vitro, and the changes that take place when activated by myosin heads are still poorly understood. Therefore, we used FRET and analyses developed in our earlier study for a similar multisite system (Bacchiocchi and Lehrer, 2002).
We donor labeled the α- and β-isoforms of gizzard Tm at positions 190 and 36, respectively, and used frequency domain FRET to study the movement of the reconstituted dimers (αβ, αα, and ββ) on F-actin by myosin subfragment 1 (S1). Two different models were used to fit the data: we used first a distance distribution (DD) model in which the “apparent” donor-acceptor (D-A) distances (Bacchiocchi and Lehrer, 2002) between one or two donors on Tm and the array of acceptors on F-actin are fitted to one or two distance distributions, respectively; then we adopted an atomic coordinate-distance distribution (AC-DD) model, where the apparent distances of the DD model are interpreted in terms of actual positions and orientations of the labeled proteins, using published atomic coordinates of F-actin·Tm (Lorenz et al., 1993, 1995) and varying the position and orientation of Tm on F-actin. Our results showed a very good agreement between the DD and the AC-DD model, which provided a direct link between distance changes and structure of the protein complex. The use of different isoforms both singly and doubly labeled, allowed for the global treatment of many fluorescence measurements that could be interpreted consistently in terms of a simple model. The results indicate that Tm rolls over actin uniformly in the presence of saturating myosin heads. They also indicate that the native heterodimer, αβTm, binds specifically to F-actin.
The key element in our accurate time-resolved measurements and global analysis is the ability to recover one or two D-A apparent distances in the Tm·F-actin ± S1 complex. This ability has been assessed in our previous article (Bacchiocchi and Lehrer, 2002) via thorough analysis of simulated data. A preliminary report of this work has been presented (Bacchiocchi et al., 2002).
EXPERIMENTAL PROCEDURES
Protein preparation
Skeletal muscle actin was prepared and labeled at Cys-374 with the DABMI acceptor (Molecular Probes, Eugene, OR) and chicken gizzard smooth muscle Tm was prepared and labeled at Cys-36 of the β-chain or Cys-190 of the α-chain with the 1,5-IAEDANS donor (Molecular Probes) according to published procedures (Graceffa, 1999, 2000). The labeled and unlabeled Tm chains were assembled to form doubly or singly labeled heterodimers as described (Graceffa, 1999). In the native state smooth muscle Tm is a heterodimer. In all our experiments, actin was always taken from a fresh stock solution, of high concentration (≈150 μM) and used within a week. Typical labeling ratios were 0.80 for Tm and 0.85 for F-actin. A typical sample composition was: [Tm] = 1.0 μM; [F-actin] = 10.0 μM; [S1] = 10.0 μM in F-buffer (10 mM Hepes, 25 mM NaCl, 5 mM MgCl2, and 0.1 mM CaCl2, pH 7.5 at 25°C). In the chosen conditions (Graceffa, 1999, 2000), labeling did not impair normal Tm binding on actin and the fraction of Tm bound to F-actin was always larger than 0.95. We also used an excess ofF-actin to further limit the presence of donor-labeled Tm not bound toF-actin, to minimize the amount of donor fluorescence not quenched by the acceptor. In practice, a certain amount of unquenched donor fluorescence is always present in this kind of experiment and, even if we can take it into account in the analysis (see the Results section), it must be kept as low as possible because it does not provide any information about D-A distances, thereby reducing the quality of the FRET measurements. Using an excess of F-actin does not introduce other unwanted fluorescence signals because DABMI is not fluorescent and the small increase in light scattering of the solution (in particular when S1 is also present) is filtered out (see next section).
Fluorescence lifetime measurements
Frequency-domain fluorescence data were collected at 20°C with an ISS K2 cross-correlation, phase, and modulation fluorometer (ISS, Urbana, IL), using the 325-nm excitation of a Liconix 3220N He-Cd laser (Melles-Griot, Carlsbad, CA). Twenty-five frequencies were recorded between 3 and 200 MHz for each measurement. A 500-nm interference filter, 30-nm bandwidth was used to isolate the AEDANS emission and to reject excitation light. A 10-μM PPO solution in ethanol was used as reference (1.4 ns monoexponential decay; Lakowicz, 1999). A pair of identical 4-mm light-path cuvettes was used in all measurements to avoid inner filter effect and to minimize the targeting error (Lakowicz, 1999). The acquisition statistics of the instrument was set to a standard deviation δφ = 0.2° for the phase and δm = 0.005 for the modulation.
The instrument performance in terms of intensity of the excitation light and level of modulation was optimized on a daily basis to ensure the best quality of the collected data. This care was essential in particular when measuring the singly labeled samples where the amount of donor-labeled protein (and hence the fluorescence emission) is relatively low as described in the published procedures (Graceffa, 1999, 2000).
COMPUTATIONAL DETAILS
Gaussian distance distribution model
To obtain information about the flexibility of the donor-acceptor-labeled F-actin·Tm ± S1 complex, frequency-domain phase and modulation FRET data were analyzed in terms of a Gaussian distance distribution model. In this model the protein complex is assumed to be rigid on the energy transfer timescale (ns) but flexible on a longer timescale, resulting in a static distribution of D-A distances. This model has been presented and discussed in detail (Lakowicz et al., 1988) and subsequently extended (Lakowicz et al., 1991; Cheung et al., 1991) to take into account incomplete acceptor labeling.
The energy transfer (ET) system is formed by one or two donors on one or both of the two Tm chains and an array of acceptors in spectroscopically equivalent positions on F-actin. Because the ET rates are additive, this set of acceptors will behave as a single deactivation channel of the donor excitation energy or, in other words, as an apparent single acceptor. On the other hand, because the two donor-labeled Tm chains can be located at a different distance from the acceptors on the actin filament, we will need, in general, two apparent D-A distances to describe the ET in the system. Therefore, assuming complete acceptor labeling, the fluorescence intensity decay I(t) of the system can be modeled by the following double Gaussian distance distribution equation:
 |
(1) |
where αDi is the fractional amplitude of the i-th donor relaxation τDi and R0 is the critical transfer distance. The summations on i and j are extended to L exponential components of the donor decay and two different distance distributions, respectively. Fj(x) has the usual Gaussian form for the distribution of the D-A distance x:
 |
(2) |
where aj is the fractional contribution of the j-th distribution of mean apparent distance raj and width σj. The frequency response (phase shift φω and demodulation mω) can be calculated as the sine and cosine transforms of the decay law 1 (Lakowicz et al., 1984).
Data were fitted using the GAUDIS fitting function (CFS_LS global fitting program; Johnson, 2000), which implements the Gaussian distance distribution model of Eq. 1 and that can also fit the fraction of acceptor labeling fl (Lakowicz et al., 1991; Cheung et al., 1991). A detailed study on the capability of the GAUDIS model to resolve the required two distance distributions was presented in a previous article (Bacchiocchi and Lehrer, 2002).
Atomic coordinate-distance distribution model
In the DD model presented above, the experimental data are analyzed in terms of apparent distances raj. Unfortunately, the results cannot be directly translated into different structures of the protein complex and, particularly interesting in our study, into different positions and orientations of Tm on F-actin. To provide a link between the experimental data and the structure of the protein complex we developed a second model to interpret the apparent distances in terms of actual positions and orientations of the labeled proteins. This was done in two steps.
First, we consider that each apparent distance ra in the DD model can be related to the individual D-A distances ri via the equation (Bacchiocchi and Lehrer, 2002)
 |
(3) |
where the sum is extended over N acceptors located within a chosen cutoff radius (six actin subunits in our case, see below) from the donor to include all the significant energy transfers.
Second, in the rigid-body approximation, an appropriate atomic coordinate model of the protein complex can be used to provide an estimate of the positions of the donors and acceptors allowing for the calculation of the individual D-A distances ri. The positions of the proteins are then varied to yield the best fit to the experimental data. The resulting model can be viewed as a combination of an atomic coordinate and a distance distribution model where the individual D-A distances ri, calculated from the atomic coordinates, are included in a distance distribution model (Eqs. 1 and 2) via Eq. 3. We will indicate this model as the atomic coordinate-distance distribution (AC-DD) model.
The AC-DD model used the published coordinates of F-actin with ADP, Ca2+ and phalloidin (Lorenz et al., 1993), and the coordinates of Tm bound to F-actin, model-built as described (Lorenz et al., 1995). The structures of the D-A labels AEDANS and DABMI were modeled using the molecular modeling package Insight II (Accelrys, San Diego, CA). Mean label positions were chosen as follows. A stereochemically reasonable average position for AEDANS attached to the sulfur atom of the cysteine residues of the α- or β-chain of Tm was modeled with the program “O” (Jones et al., 1991). DABMI was linked to the sulfur atom of Cys-374 of F-actin and its average position, defined by the average orientation of the main molecular axis, was included as a variable parameter in the data analysis. Details about the chosen D-A positions are presented in the Discussion section. The D-A cutoff radius was chosen so as to take into account all the significant D-A interactions while keeping the computing time to a reasonable amount. A good compromise was to consider only the individual transfers with an efficiency larger than 1%, which corresponds, for R0 = 39.9 Å for the AEDANS-DABMI D-A pair (Tao et al., 1983) to a D-A distance cut-off radius of ≈86 Å and, in practice, it means to consider the acceptors located on six actin subunits closest to the donors (Fig. 1).
FIGURE 1.
Atomic coordinate model of the F-actin·Tm complex from Lorenz et al. (1993, 1995). The model is formed by six actin monomers (wire frame) and two αβTm molecules (α-chain, cyan ribbon; β-chain, blue ribbon) on opposing sides of the F-actin helix. The donor and acceptor moieties (space-fill) are AEDANS attached to βTm Cys-36 (D1) and DABMI attached to F-actin Cys-374 (A1–A6), respectively. Displayed using the program RasMol (Bernstein, 1999).
Experimental data were fitted using a search algorithm that inputs the labeled Tm and F-actin atomic coordinates and systematically varies the Tm position on F-actin (both proteins are considered as rigid bodies) by changing the azimuthal position (angle around the F-actin axis) and the axial orientation (angle around the Tm interchain axis). Angles are calculated with respect to the original (unchanged) Lorenz position. A positive angle represents a clockwise Tm rotation as viewed from the F-actin pointed end. The average position of the acceptors on F-actin was also optimized whereas the donors were kept fixed on Tm. The detailed procedure followed in the fit is described in the Results section. The analysis was performed using a software package that combines an implementation of a modified Gauss-Newton-Marquardt nonlinear least-squares global fitting (Arcioni et al., 1993) with a modified version of the FORTRAN code for the calculation of the frequency response (phase and modulation data) of the system (Bacchiocchi and Lehrer, 2002). Confidence intervals were estimated at a probability of 95% using a Monte-Carlo method (Straume and Johnson, 1992). The assumptions involved in the adoption of the Lorenz model, obtained for skeletal Tm, to analyze data for smooth Tm are presented in the Discussion section.
RESULTS
Donor-only fluorescence decays
The phase and modulation data of the donor-only (D-only) samples were fitted to one, two, and three exponentials (CFS_LS global fitting program, HETANL-1, 2, and 3 fitting functions (Johnson, 2000). The mono or double-exponential model was not appropriate to fit the data as clearly indicated by a high value of the  (>10) obtained for all the data sets and nonrandom residuals (data not shown). A three-exponential model gave, instead, values of the
(>10) obtained for all the data sets and nonrandom residuals (data not shown). A three-exponential model gave, instead, values of the  close to unity and random residuals for all the data sets (see Table 1 and Fig. 2, a and c). The decays are all very similar showing that the environment of the donor label is essentially the same and very similar to our previous study with skeletal muscle Tm. A change in the two shorter-lived components occurred when Tm formed a complex with F-actin and S1 (Tables 2 and 3). Because both these components have, in general, a relatively small fractional contribution, αD2 and αD3, to the total decay, the phase and modulation data for all the samples are very similar (Fig. 2 b). A larger contribution of the shortest component αD3 for the singly labeled samples in the presence of S1 (first two rows in Table 3) might be due to a very small amount of light scattering and to the low emission of these singly labeled samples (see also the Protein preparation section) and thus a fraction of scattered light, which is estimated to be ∼1–2% of the total intensity, cannot be avoided. This effect is anyway present to the same extent in both the D-only and in the D-A samples and therefore does not interfere with the ET analysis, which is always obtained by a global fit of the D-only and the corresponding D-A data set. The residuals of the fit always had small and random deviations (Fig. 2 c). The errors on the recovered lifetimes, in particular for the two shorter components, are sometimes relatively large as a consequence of a certain correlation between the parameters, typical of a three-exponential fit. Again, the complexity of the D-only decay does not limit the ability to recover meaningful distances from the analysis of the D-A samples, because the D-only decay is included in the fit (Lakowicz et al., 1988).
close to unity and random residuals for all the data sets (see Table 1 and Fig. 2, a and c). The decays are all very similar showing that the environment of the donor label is essentially the same and very similar to our previous study with skeletal muscle Tm. A change in the two shorter-lived components occurred when Tm formed a complex with F-actin and S1 (Tables 2 and 3). Because both these components have, in general, a relatively small fractional contribution, αD2 and αD3, to the total decay, the phase and modulation data for all the samples are very similar (Fig. 2 b). A larger contribution of the shortest component αD3 for the singly labeled samples in the presence of S1 (first two rows in Table 3) might be due to a very small amount of light scattering and to the low emission of these singly labeled samples (see also the Protein preparation section) and thus a fraction of scattered light, which is estimated to be ∼1–2% of the total intensity, cannot be avoided. This effect is anyway present to the same extent in both the D-only and in the D-A samples and therefore does not interfere with the ET analysis, which is always obtained by a global fit of the D-only and the corresponding D-A data set. The residuals of the fit always had small and random deviations (Fig. 2 c). The errors on the recovered lifetimes, in particular for the two shorter components, are sometimes relatively large as a consequence of a certain correlation between the parameters, typical of a three-exponential fit. Again, the complexity of the D-only decay does not limit the ability to recover meaningful distances from the analysis of the D-A samples, because the D-only decay is included in the fit (Lakowicz et al., 1988).
TABLE 1.
Triple-exponential decay parameters for the AEDANS-labeled Tm chains, α* and β*, in the singly and doubly labeled Tm dimer
| Tm sample | τD1 (ns) | αD1 | τD2 (ns) | αD2 | τD3 (ns) | αD3 |  |
|---|---|---|---|---|---|---|---|
| α*β | 14.3 ± 0.7 | 0.46 ± 0.17 | 9.4 ± 0.6 | 0.36 ± 0.04 | 1.1 ± 0.1 | 0.19 ± 0.01 | 1.16 |
| αβ* | 14.3 ± 0.6 | 0.59 ± 0.16 | 8.6 ± 1.8 | 0.22 ± 0.12 | 1.2 ± 0.1 | 0.19 ± 0.01 | 1.06 |
| α*β* | 14.3 ± 0.6 | 0.60 ± 0.20 | 8.5 ± 1.9 | 0.21 ± 0.10 | 1.22 ± 0.04 | 0.19 ± 0.01 | 1.07 |
| α*α* | 13.7 ± 1.0 | 0.60 ± 0.20 | 8 ± 3 | 0.25 ± 0.16 | 1.2 ± 0.2 | 0.15 ± 0.02 | 1.11 |
| β*β* | 14.3 ± 0.6 | 0.51 ± 0.12 | 10.2 ± 1.4 | 0.34 ± 0.06 | 1.7 ± 0.3 | 0.15 ± 0.02 | 1.11 |
FIGURE 2.
Frequency-domain phase and modulation data and fits for AEDANS-labeled Tm alone and in complex with F-actin and S1. The phase angle increases and the modulation decreases with increasing frequency. (a) Tm alone (all combinations of α- and β-chains, singly and doubly labeled are shown). (b) Doubly labeled Tm heterodimer (α*β* Tm) + F-actin in the absence (□) and in the presence (+) of S1. The Tm-alone fit (thicker line) is shown for comparison. (c) Phase (δφ) and modulation (δm) residuals for a typical fit. The parameters recovered from the fits are reported in Tables 1–3.
TABLE 2.
Triple-exponential decay parameters for the AEDANS-labeled Tm chains, α* and β*, in the singly and doubly labeled Tm dimer complexed with F-actin
| Tm sample | τD1 (ns) | αD1 | τD2 (ns) | αD2 | τD3 (ns) | αD3 |  |
|---|---|---|---|---|---|---|---|
| α*β | 13.3 ± 0.4 | 0.68 ± 0.08 | 5.0 ± 2.2 | 0.14 ± 0.06 | 0.7 ± 0.2 | 0.18 ± 0.02 | 1.14 |
| αβ* | 13.9 ± 0.2 | 0.61 ± 0.02 | 4.6 ± 0.8 | 0.16 ± 0.02 | 0.6 ± 0.1 | 0.23 ± 0.01 | 1.07 |
| α*β* | 14.8 ± 1.2 | 0.54 ± 0.26 | 9 ± 3 | 0.3 ± 0.2 | 1.5 ± 0.2 | 0.16 ± 0.02 | 1.13 |
| α*α* | 13.4 ± 0.5 | 0.64 ± 0.06 | 6.2 ± 2.6 | 0.15 ± 0.06 | 0.6 ± 0.1 | 0.21 ± 0.01 | 1.01 |
| β*β* | 13.6 ± 0.2 | 0.72 ± 0.02 | 4.3 ± 1.0 | 0.15 ± 0.02 | 0.6 ± 0.3 | 0.14 ± 0.01 | 1.04 |
TABLE 3.
Triple-exponential decay parameters for the AEDANS-labeled Tm chains, α* and β*, in the singly and doubly labeled Tm dimer complexed with F-actin + S1
| Tm sample | τD1 (ns) | αD1 | τD2 (ns) | αD2 | τD3 (ns) | αD3 |  |
|---|---|---|---|---|---|---|---|
| α*β | 13.9 ± 0.2 | 0.48 ± 0.02 | 3.9 ± 0.8 | 0.17 ± 0.01 | 0.5 ± 0.1 | 0.35 ± 0.01 | 1.09 |
| αβ* | 14.5 ± 0.3 | 0.51 ± 0.03 | 4.5 ± 1.0 | 0.13 ± 0.02 | 0.4 ± 0.1 | 0.36 ± 0.03 | 1.26 |
| α*β* | 14.4 ± 0.6 | 0.70 ± 0.09 | 7 ± 3 | 0.15 ± 0.08 | 1.3 ± 0.3 | 0.14 ± 0.02 | 1.23 |
| α*α* | 13.8 ± 0.3 | 0.66 ± 0.03 | 4.7 ± 1.8 | 0.12 ± 0.02 | 0.6 ± 0.2 | 0.22 ± 0.01 | 1.17 |
| β*β* | 14.6 ± 0.5 | 0.69 ± 0.08 | 7.8 ± 2.6 | 0.16 ± 0.08 | 1.6 ± 0.2 | 0.15 ± 0.01 | 1.02 |
Distance distribution model analysis
Phase and modulation data for the DABMI acceptor-labeled F-actin in complex with the different Tm isoforms, donor labeled with AEDANS on one or both of the two chains, in the absence and in the presence of S1, were analyzed using the DD model described above. A first series of fits was done by a global analysis of each pair of D-only and D-A data sets. The lifetime decay was fixed at the value obtained for the D-only sample and all of the remaining parameters were kept variable: the mean D-A apparent distances ra1, ra2; the distribution widths σ1, σ2; and the fraction, fl, of acceptor labeling, which in practice corresponds to the fraction of F-actin-bound Tm, fb, when fl > 0.8, as we have previously discussed for this particular multiacceptor system (Bacchiocchi and Lehrer, 2002).
The singly donor-labeled samples were well fitted to a single distance distribution giving small errors on all recovered parameters. A double distance distribution was required to fit the doubly labeled samples. The fits were all good but the recovered parameters exhibited relatively large correlations and uncertainties (results not shown). A more reliable fit was obtained by assuming the same value of σ for both distances and repeating the optimization for a different value of σ until the best possible fit was achieved. Typical fits and corresponding residuals are shown in Fig. 3, the recovered parameters are reported in Table 4. The goodness of the fits was assessed by the  values and by the analysis of the residuals (see Fig. 3 b). All confidence intervals were calculated with the support-plane method (Johnson, 2000) at a probability of 95%. The fraction of F-actin-bound Tm fb was always larger than 95% both in the absence and in the presence of S1, in agreement with the binding assays. The recovered width of the distance distribution σ (penultimate column) is larger than 2.5 Å, the distribution expected for a fraction fl of acceptor labeling = 0.85 in a rigid system (Bacchiocchi and Lehrer, 2002). Therefore we must assume a certain degree of flexibility in the F-actin·Tm ± S1 complex, which can be due to flexibility of the proteins, to a distribution of positions of Tm on F-actin and to a slow component of the segmental motions of the labels resulting in a static (on the FRET timescale) distribution of label positions. This last effect, which was observed also in our previous work on the F-actin·Tm·Tn complex (Bacchiocchi and Lehrer, 2002), seems indeed to be relevant, because the width of the distance distribution, σ, appears to be only slightly affected by the presence of S1, which may be expected to reduce considerably the Tm mobility. Instead, the distribution width appears to be more related to the label position on Tm, with a smaller σ value for the label attached to the α-chain at position 190 (6–7 Å) and a slightly largerσ-value for the label attached to the β-chain at position 36 (8–9 Å). These values are also in good agreement with those obtained in our previous work (Bacchiocchi and Lehrer, 2002) for AEDANS on skeletal Tm, again suggesting that the distribution width is mainly a label-related parameter in its local environment, which is indeed similar for residues 190 and 36 being both in the same “a” position of the pseudoheptapeptide repeat, “abcdefg,” located in the Tm hydrophobic ridge.
values and by the analysis of the residuals (see Fig. 3 b). All confidence intervals were calculated with the support-plane method (Johnson, 2000) at a probability of 95%. The fraction of F-actin-bound Tm fb was always larger than 95% both in the absence and in the presence of S1, in agreement with the binding assays. The recovered width of the distance distribution σ (penultimate column) is larger than 2.5 Å, the distribution expected for a fraction fl of acceptor labeling = 0.85 in a rigid system (Bacchiocchi and Lehrer, 2002). Therefore we must assume a certain degree of flexibility in the F-actin·Tm ± S1 complex, which can be due to flexibility of the proteins, to a distribution of positions of Tm on F-actin and to a slow component of the segmental motions of the labels resulting in a static (on the FRET timescale) distribution of label positions. This last effect, which was observed also in our previous work on the F-actin·Tm·Tn complex (Bacchiocchi and Lehrer, 2002), seems indeed to be relevant, because the width of the distance distribution, σ, appears to be only slightly affected by the presence of S1, which may be expected to reduce considerably the Tm mobility. Instead, the distribution width appears to be more related to the label position on Tm, with a smaller σ value for the label attached to the α-chain at position 190 (6–7 Å) and a slightly largerσ-value for the label attached to the β-chain at position 36 (8–9 Å). These values are also in good agreement with those obtained in our previous work (Bacchiocchi and Lehrer, 2002) for AEDANS on skeletal Tm, again suggesting that the distribution width is mainly a label-related parameter in its local environment, which is indeed similar for residues 190 and 36 being both in the same “a” position of the pseudoheptapeptide repeat, “abcdefg,” located in the Tm hydrophobic ridge.
FIGURE 3.
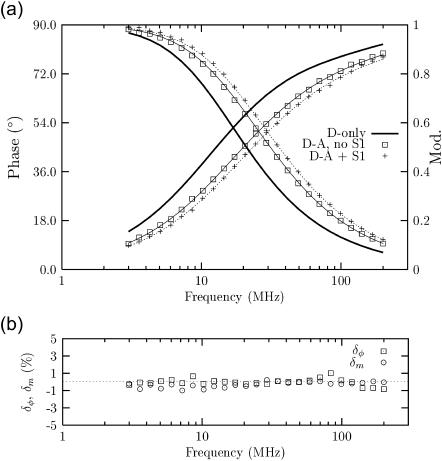
(a) Frequency-domain phase and modulation data in the presence of energy transfer and fits for α*β* AEDANS donor-labeled Tm in complex with DABMI acceptor-labeled F-actin in the absence (□) and in the presence (+) of S1. The fit of the corresponding donor-only F-actin·Tm complex (D-only, thicker line) is shown for comparison. (b) Phase (δφ) and modulation (δm) residuals for a typical fit. The parameters recovered from the fits are reported in Tables 4 and 5.
TABLE 4.
Single and double Gaussian distance distribution analysis of FRET in the F-actin·Tm ± S1 complex for the different Tm isoforms
| Tm isoform | fb | ra1 (Å) | ra2 (Å) | σ (Å) |  |
|
|---|---|---|---|---|---|---|
| −S1 | α*β | 0.99 ± 0.01 | 40.4 ± 0.1 | 6.9 ± 0.1 | 1.1 | |
| −S1 | αβ* | 0.99 ± 0.01 | – | 47.3 ± 0.1 | 9.0 ± 0.4 | 1.2 |
| −S1 | α*β* | 0.99 ± 0.01 | 40.4 ± 0.6 | 48.4 ± 0.7 | 6.8 ± 0.6 | 1.2 |
| −S1 | α*α* | 0.99 ± 0.01 | 40.4 ± 0.3 | 40.0 ± 0.2 | 6.0 ± 0.8 | 1.1 |
| −S1 | β*β* | 0.99 ± 0.01 | 40.0 ± 0.4 | 47.1 ± 0.3 | 8.6 ± 0.9 | 1.5 |
| +S1 | α*β | 0.99 ± 0.01 | 41.1 ± 0.1 | – | 7.2 ± 0.4 | 1.5 |
| +S1 | αβ* | 0.99 ± 0.01 | – | 38.6 ± 0.1 | 8.0 ± 0.1 | 1.3 |
| +S1 | α*β* | 0.99 ± 0.01 | 41.9 ± 0.8 | 38.8 ± 0.9 | 6.0 ± 0.7 | 1.3 |
| +S1 | α*α* | 0.99 ± 0.01 | 40.9 ± 0.3 | 34.7 ± 0.2 | 6.2 ± 0.8 | 1.1 |
| +S1 | β*β* | 0.99 ± 0.01 | 41.8 ± 0.5 | 38.6 ± 0.4 | 8.1 ± 0.9 | 1.1 |
Recovered parameters are fb, fraction of F-actin-bound Tm; ra1 and ra2, donor-acceptor apparent distances; and σ, width of the distance distribution.
The apparent distances recovered for the two singly labeled αβTm heterodimers appeared to be consistent with those obtained for the doubly labeled heterodimer, both in the absence and in the presence of S1 (Table 4, row number 1, 2, and 3 (−S1) and row number 6, 7, and 8 (+S1)). Therefore, a larger global analysis, comprehensive of all the data of the αβTm heterodimer was performed, to verify this consistency. The results, presented in Table 5, confirmed the hypothesis and were also in reasonable agreement with the previous steady-state results (Graceffa, 1999, 2000). The quality of each global fit was comparable with that of the independent analysis, thus indicating that the distances recovered from the singly labeled samples are indeed the same obtained in the doubly labeled ones. Due to the globalization of many independent data sets, the correlation among the parameters was significantly reduced and it was possible to fit all the parameters independently. By joining together singly and doubly labeled samples, it was also possible to assign each apparent distance to the corresponding Tm chain (see Table 5).
TABLE 5.
Double Gaussian distance distribution analysis: global fit of the singly and doubly labeled αβTm heterodimer samples in complex with F-actin ± S1
| ra1 (Å) α-chain | σ1 (Å) | ra2 (Å) β-chain | σ2 (Å) |  |
|
|---|---|---|---|---|---|
| −S1 | 39.7 ± 0.3 | 6.8 ± 0.4 | 46.5 ± 0.2 | 7.2 ± 0.6 | 1.4 |
| Graceffa (1999) | 38 | – | 43 | – | – |
| Graceffa (2000) | – | – | 45 | – | – |
| – | – | – | – | – | |
| +S1 | 41.1 ± 0.3 | 6.4 ± 0.3 | 39.0 ± 0.2 | 7.2 ± 0.5 | 1.5 |
| Graceffa (1999) | 43 | – | 37 | – | – |
Recovered parameters are ra1 and ra2, donor-acceptor apparent distances; and σ1 and σ2, widths of the distance distributions. The distances obtained in previous steady-state studies are shown for comparison (Graceffa, 1999, 2000).
Atomic coordinate-distance distribution (AC-DD) model analysis
Phase and modulation data for the DABMI acceptor-labeled F-actin in complex with the αβTm heterodimer, donor-labeled with AEDANS on one or both of the two chains, in the absence and in the presence of S1, were also analyzed using the AC-DD model, described above, according to the following procedure.
First, a rough estimate for the average position of the DABMI acceptor moiety attached to F-actin Cys-374 was obtained by a preliminary global analysis of the data sets for the singly and doubly labeled αβTm in the absence of S1. The fit was performed by keeping Tm fixed at the original (unchanged) Lorenz position (Lorenz et al., 1995) to model the −S1 state. The distribution widths σ1, σ2, and the fraction of F-actin-bound Tm, fb were fixed at the values recovered by the DD model and the average position of the acceptor was then optimized to obtain best fit to the data.
An estimate of the position of Tm in the presence of S1 was then obtained by keeping the acceptor fixed at the previously estimated position with σ1, σ2, and fb still fixed. Due to Tm flexibility, it is possible that the displacement of Tm in the presence of S1 is different for the region around Cys-36 with respect to the region around Cys-190. Therefore, we did two independent fits of the data sets in the presence of S1: a global fit of only the data sets corresponding to Tm labeled at position 36 and another global fit of only the data sets corresponding to Tm labeled at position 190. Best-fit Tm azimuthal positions were determined with a search centered at the original Lorenz position, searching over a range of ±30° in step of 1° and optimizing the Tm position at each step. Best-fit parameters (see Table 6) indicated a similar displacement, within the experimental error, for both labeled positions. The previous global analysis was then repeated using the data sets for the singly and doubly labeled αβTm in the presence of S1 and assuming the same displacement at the two labeled positions. Best fit was obtained for a Tm azimuthal position of 23 ± 3° and an axial orientation of 19 ± 12°.
TABLE 6.
Best-fit azimuthal and axial positions of Tm obtained from the atomic coordinate-distance distribution analysis of the donor-acceptor fluorescence decays
| Parameters | Cys-36 | Cys-190 |
|---|---|---|
| Tm azimuthal position | 22 ± 3° | 24 ± 1° |
| Tm axial orientation | 22 ± 36° | 17 ± 24° |
Finally, a global analysis was performed including all data sets for the singly and doubly labeled αβTm heterodimer in the absence and in the presence of S1. This was done in two steps. Initially, the average position of the DABMI moiety was again kept fixed, to further optimize the Tm azimuthal position and axial orientation by varying also σ1, σ2, and fb, which were previously kept fixed. The best fit values obtained in this way were then used as the initial guess for the final optimization where all the parameters were variable. Final best fit was obtained for a Tm azimuthal movement of 22.6°, an axial rotation of 16.5°, and a fraction of F-actin-bound Tm, fb of 0.99. The recovered parameters are listed in Table 7; the azimuthal and axial movement of the αβTm heterodimer is shown in Figs. 4 and 5. The corresponding D-A apparent distances and distribution widths are presented in Table 8. The best fit for the average position of the DABMI moiety attached to F-actin Cys-374 is displayed in Fig. 6 (left) where a comparison with the position of a TMR moiety, linked to G-actin Cys-374, determined by x-ray crystallography (Otterbein et al., 2001) is also shown (right). We notice that the position of the DABMI moiety is very different from the TMR, in agreement with the observation that the latter inhibits G-actin polymerization whereas the former does not.
TABLE 7.
Best-fit azimuthal and axial positions of Tm and fraction of bound Tm obtained from the atomic coordinate-distance distribution analysis of the donor-acceptor fluorescence decays
| Parameters | −S1 | +S1 |
|---|---|---|
| Tm azimuthal position | 0 ± 1.5° | 22.6 ± 1.6° |
| Tm axial orientation | 0 ± 1.1° | 16.5 ± 1.3° |
| Fraction of bound Tm | 0.99 ± 0.01 | 0.99 ± 0.01 |
Global fit of the data for the singly and doubly labeled αβTm heterodimer samples in complex with F-actin ± S1. Angles are relative to the original Lorenz model (Lorenz et al., 1995).
FIGURE 4.
S1-induced azimuthal and axial movement of the αβTm heterodimer corresponding to the atomic coordinate-distance distribution analysis reported in Table 7. The longitudinal view of the thin filament along theF-actin axis (C) shows two adjacent actin monomers (green and red wire frame); the ±S1 fitted positions of the Tm region around the donor at Cys-190 (Glu-187–Leu-193, blue and cyan ribbons; Cys-190, space-fill) and at Cys-36 (Glu-33–Leu-39, blue and cyan ribbons; Cys-36, space-fill); and the fitted average position of the DABMI moiety (space-fill; see text for details). The fitted Tm movement is a combination of an azimuthal (around the F-actin axis) and an axial (around its own interchain axis) rotation. Displayed using the program RasMol (Bernstein, 1999).
FIGURE 5.
Comparison between the position of the region around the donor at Cys-190 of the α-chain and the position of the region around the donor at Cys-36 of the β-chain in the αβTm heterodimer in presence of S1, corresponding to the atomic coordinate-distance distribution analysis reported in Table 7 (+S1). The longitudinal view of the thin filament along the F-actin axis (C) is similar to Fig. 4 and shows two adjacent actin monomers (green and red wire frame); the +S1 fitted positions of the Tm region around the donor at Cys-190 (Glu-187–Leu-193, blue and cyan space-fill; Cys-190 and AEDANS, space-fill) and at Cys-36 (Glu-33–Leu-39, blue and cyan space-fill; Cys-36 and AEDANS, space-fill); and the fitted average position of the DABMI moiety (space-fill; see text for details). Displayed using the program RasMol (Bernstein, 1999).
TABLE 8.
Donor-acceptor apparent distances, ra1, ra2, and widths of the distance distributions, σ1, σ2, corresponding to the atomic coordinate-distance distribution analysis reported in Table 7
| ra1 (Å) α-chain | σ1 (Å) | ra2 (Å) β-chain | σ2 (Å) | |
|---|---|---|---|---|
| −S1 | 40.6 | 7.4 ± 0.6 | 47.2 | 7.4 ± 0.6 |
| +S1 | 40.7 | 7.4 ± 0.6 | 39.0 | 7.4 ± 0.6 |
FIGURE 6.
Comparison between best-fit average position of the DABMI moiety attached to F-actin Cys-374 (left) and the position of a TMR moiety linked to Cys-374 of G-actin (same backbone orientation as F-actin is shown), determined by x-ray crystallography (Otterbein et al., 2001) (right). The view, along the F-actin axis, shows one actin monomer (red wire frame), a ribbon rendering of the F-actin C-terminal region (from Gly-366 to Cys-374, left, or to Arg-372, right) and the attached moieties (DABMI, left, and TMR, right, space-fill). Displayed using the program RasMol (Bernstein, 1999).
DISCUSSION
Orientation factor κ2
In the calculation of R0, for the D-A pair considered in this study, the orientation factor κ2 was 2/3 (Förster, 1948; Van Der Meer et al., 1991) the value averaged over a fast-reorienting and random distribution of label orientations. Discussions on the validity of this approximation, on the basis of substantial rapid reorientation of both donor and acceptor on the fluorescence timescale, have been presented: for Tm donor-labeled with AEDANS (Tao et al., 1983), for F-actin acceptor-labeled with DABMI at Cys-374 (Tao, 1978). The good agreement between the modeled and the measured interchain distances on Tm, presented in a previous work (Bacchiocchi and Lehrer, 2002), is a further confirmation of the correctness of this approximation. The fact that more than one acceptor is involved also contributes to making 2/3 an appropriate value for κ2 (Censullo et al., 1992).
Relating movement to atomic coordinates
The D-A distances in the AC-DD model were calculated between the centers of the aromatic systems of the AEDANS and DABMI labels attached to the F-actin·Tm protein complex. These coordinates were obtained by linking a model of the AEDANS and DABMI labels (Insight II molecular modeling package, Accelrys, San Diego, CA) to their respective labeling sites on the Lorenz atomic coordinate model of the protein complex (Lorenz et al., 1993, 1995).
The adoption of the Lorenz model, obtained for skeletal Tm, to mimic smooth Tm can be questioned. For example, recent studies have shown that the azimuthal position of smooth Tm, in the troponin-free thin filament, is different from skeletal (Lehman et al., 2000). On the other hand, given the large similarities in sequence between skeletal and smooth Tm (Smillie, 1996 and references therein) and the constraints imposed by the coiled-coil structure and by the interaction with F-actin that determine the tertiary structure, the length of a Tm molecule and its periodicity on the thin filament are the same in striated and smooth muscle Tm (Matsumura and Lin, 1982). The relative positions of the donor labels with respect to the protein itself are also likely to be quite similar in skeletal and smooth Tm. In fact, the donor labels are attached to the same type of position in the pseudoheptapeptide repeat of either skeletal or smooth Tm. The most reasonable assumption for a label attached to this position is to model it by minimizing clashes within the two chains. Therefore, a stereochemically reasonable average position for AEDANS attached to the sulfur atom of the single cysteine residue of the α- or β-chain of Tm was modeled with the program “O” (Jones et al., 1991) minimizing the interaction of the label with the neighboring side chains. In the resulting position, AEDANS is essentially perpendicular to the local interchain Tm axis, exposing the hydrophilic  group and the hydrophobic aromatic group. By a detailed analysis of the AEDANS structure, it could be argued that a folded conformation of the label linker arm, with the aromatic group less exposed, might also be possible. The presence of more than one conformation of AEDANS is indeed consistent with the observed three-exponential lifetime decay of the D-only samples and it would explain, at least in part, the distance distribution obtained for the D-A samples. Due to the above considerations, we believe that the Lorenz model can still be used in a search algorithm, which will optimize its position on F-actin, to mimic the positions of the donor labels attached to smooth Tm in the Tm·F-actin complex.
group and the hydrophobic aromatic group. By a detailed analysis of the AEDANS structure, it could be argued that a folded conformation of the label linker arm, with the aromatic group less exposed, might also be possible. The presence of more than one conformation of AEDANS is indeed consistent with the observed three-exponential lifetime decay of the D-only samples and it would explain, at least in part, the distance distribution obtained for the D-A samples. Due to the above considerations, we believe that the Lorenz model can still be used in a search algorithm, which will optimize its position on F-actin, to mimic the positions of the donor labels attached to smooth Tm in the Tm·F-actin complex.
A different approach was used to estimate the position of the DABMI label on F-actin. DABMI is attached on Cys-374 at the C-terminus of F-actin, a region known to be quite flexible and not well resolved in the F-actin structure (Lorenz et al., 1995). Therefore, the average position of the DABMI label was included as an extra variable parameter in the data analysis performed with the AC-DD model. To avoid the correlation between the positions of the donors and acceptors, in the preliminary fit of the DABMI average position, Tm was fixed at the original Lorenz position (Lorenz et al., 1995) to model the −S1 state (see also the Results section). This relatively arbitrary choice represents, nevertheless, a reasonable initial guess for the position of Tm. In fact, the radial distance of Tm from F-actin is quite well determined (Lehman et al., 2000 and references therein) and is essentially the same for skeletal and smooth Tm, therefore, it has been kept fixed to the Lorenz model value in the analysis. The azimuthal position of Tm in the Lorenz model is shifted ∼10–20° toward the F-actin inner domain with respect to the real position of smooth Tm (Lehman et al., 2000) whereas the axial and longitudinal (z-stagger) offset of the model with respect to the real position of smooth Tm is completely unknown. Due to the continuous array of acceptors along F-actin, the FRET data are almost independent from Tm z-stagger (data not shown), which means that the relevant parameters in the analysis of the localization of Tm are only its azimuthal position and axial orientation. In the next step of the fit, the acceptor label was, in turn, kept fixed to the preliminary position to obtain a first estimate of the +S1 Tm position. Finally, a global fit of the −S1 and +S1 data sets was performed, using the preliminary estimated positions as initial guess, to further optimize the Tm positions in the absence and in the presence of S1 and the average position of DABMI, in the assumption that this position is independent of [S1]. In fact, a net movement of the acceptor label would result in a change of the same sign in both recovered apparent distances, whereas, as we observed, in agreement with previous results (see Table 5; Graceffa, 1999, 2000), one apparent distance increased and the other decreased.
Any azimuthal and axial offset of the modeled −S1 Tm position with respect to the real position of smooth Tm will affect the average position of the DABMI acceptor label. If this position is the same in the absence and in the presence of S1, the offset of the Lorenz model will also be the same in the absence and in the presence of S1. This means that the difference between these two positions, i.e., the Tm movement, can still be considered essentially unaffected by any offset. Therefore, although we cannot determine the absolute position of smooth Tm to high precision, we can determine relative positions and movement.
In Fig. 6 the best-fit average position of the DABMI label attached to F-actin is compared to the position of a TMR label attached to G-actin (Otterbein et al., 2001). The evident difference of the positions is in agreement with the observation that the DABMI-labeled G-actin retains its ability to polymerize to F-actin, whereas the TMR-labeled G-actin does not.
Localization of Tm on F-actin
The main conclusions of this work have been obtained by comparing the distances and the distance distributions recovered via the DD model global analysis, listed in Tables 4 and 5.
Localization of singly and doubly labeled αβTm in the absence of S1
The apparent D-A distance of the singly labeled αβTm is 40.4 Å when the donor label is on the α-chain (Table 4, first row) and 47.3 Å when the donor label is on the β-chain (Table 4, second row). The apparent D-A distances of the doubly labeled αβTm are 40.4 Å and 48.4 Å (Table 4, third row), in quite good agreement with the distances found separately for the two singly labeled αβTm. The agreement was confirmed by the quality of a global analysis of all the above data sets (see Table 5, first row). This result clearly indicates that the localization of αβTm on F-actin, in the absence of S1, is independent of the labeling.
Localization of singly and doubly labeled αβTm in presence of S1
The same correspondence among the distances can be found also in the presence of S1. The apparent D-A distance of the singly labeled αβTm is 41.1 Å when the donor label is on the α-chain (Table 4, sixth row) and 38.6 Å when the donor label is on the β-chain (Table 4, seventh row), similar to the distances found for the doubly labeled αβTm, which are 41.9 Å and 38.8 Å, respectively (Table 4, eighth row), indicating, even in this case, that the localization of αβTm on F-actin is independent of the labeling.
Localization of αβTm in comparison to ααTm and ββTm in the absence and in the presence of S1
The first of the two distances of the doubly labeled ααTm, in the absence of S1, is 40.4 Å (Table 4, fourth row) and the second of the two distances of the doubly labeled ββTm is 47.1 Å (Table 4, fifth row). These distances are in quite good agreement with the distances found for the doubly labeled αβTm, in the absence of S1 (40.4 Å and 48.4 Å, Table 4, third row). In other words, the distance of one of the two α-chains in the ααTm is very similar to the distance of the α-chain in the αβTm and the same is true for the β-chain, suggesting that one of the two chains in the homodimers has the same localization of the corresponding chain in the heterodimer. Even if this might be just a coincidence, we believe that it is an indication that the homodimers have a localization similar to the heterodimer on F-actin, in the absence of S1. Indeed, this seems to be true also in the presence of S1. In this case, we notice that the distance of one of the two α- or β-chains in the doubly labeled homodimer (namely, 40.9 Å for ααTm and 38.6 Å for ββTm, Table 4, ninth and tenth rows), is quite similar to the distance of the corresponding chain in the heterodimer (41.9 Å and 38.8 Å, Table 4, eighth row), again suggesting that, also in the presence of S1, the homodimers have a localization similar to the heterodimer on F-actin.
From a careful inspection of the results presented in Table 4, it is possible to recognize a general trend: in all isoforms, upon S1 binding, the shorter D-A apparent distance showed a small increase of 0.5–2 Å, whereas the longer distance showed a larger decrease of 6–9 Å. This observation suggests that: i), the movement is likely to be a “rolling” motion of Tm on the surface of F-actin, where the Tm chain closer to the F-actin surface undergoes a smaller net displacement than the Tm chain farther from F-actin. In fact, the AC-DD model analysis showed that when Tm rotates around the thin filament axis and also around its own interchain axis (combination of an azimuthal and axial movement, rolling motion), the apparent distance of the donor on the Tm chain closer to actin exhibits a smaller and positive change and the apparent distance of the donor on the Tm chain farther from actin exhibits a larger and negative change; ii), because this combination of distance changes is the same for all isoforms, it is reasonable to conclude that all isoforms are displaced in a similar way by the full binding of myosin heads to F-actin; and iii), the combination of distance changes, and hence the displacement, is the same for the two Tm regions around the two positions 36 and 190 and because these two positions are at about one- and five-sevenths of the Tm length, respectively, this is also an indication that the Tm movement on F-actin is uniform when fully decorated with S1.
Tm mobility
The mobility of Tm can be, at least in part, monitored by the recovered width of the distance distribution, σ. The binding of myosin heads to the thin filament is expected to reduce considerably this mobility and hence the corresponding distribution widths. Instead, the widths appear to be only slightly affected by the presence of S1. In fact, all distributions are very similar, with a variation of σ within ±2 Å, both in the absence and in the presence of S1 (Table 4). This result indicates, therefore, that Tm occupies relatively well-defined positions in both the predominantly closed (−S1) and open (+S1) thin filament states.
The meaning of apparent distance
As mentioned above, the apparent distances, ra, are related to the individual D-A distances, ri, by Eq. 3. To understand which actin subunit principally contributes to each apparent distance (i.e., it bears the closest acceptor to each donor), we have used the AC-DD model to show the individual D-A distances from each donor on the doubly labeled αβTm, taken as an example, to each of the six closest acceptors on the F-actin subunits, in the absence and presence of S1 (Table 9). As can be seen, in the absence of S1, the shortest D-A distance is on different actin subunits for each donor (rA3 = 43.2 Å for D1 and rA4 = 51.1 Å for D2, boldface). S1 causes one donor to transfer more efficiently to a different acceptor (rA4 = 44.3 Å for D1, boldface), whereas the other donor transfers more efficiently to the same acceptor (rA4 = 39.8 Å for D2, boldface). We also notice that the observed apparent distance is always smaller than the corresponding shortest individual D-A distance, indicating that ET is occurring to multiple acceptors.
TABLE 9.
Apparent D-A distances, ra (Å) and individual D-A distances, rA1–rA6 (Å), from each donor, D1 (α-chain) and D2 (β-chain), on the doubly labeled αβTm, to each of the six closest acceptors, A1–A6, on the F-actin subunits, calculated according to the AC-DD model (see text for details), in the absence and presence of S1
| ra | rA1 | rA2 | rA3 | rA4 | rA5 | rA6 | ||
|---|---|---|---|---|---|---|---|---|
| −S1 | D1 | 40.6 | 65.8 | 67.4 | 43.2 | 54.4 | 74.7 | 86.1 |
| D2 | 47.2 | 79.6 | 71.9 | 62.6 | 51.1 | 86.3 | 79.9 | |
| +S1 | D1 | 40.7 | 71.6 | 59.4 | 52.5 | 44.3 | 79.6 | 81.6 |
| D2 | 39.0 | 87.2 | 63.0 | 71.8 | 39.8 | 91.7 | 75.9 |
For each donor, the apparent and the shortest D-A distance is shown in boldface.
F-actin-binding specificity of αβTm
We observe that, due to the twofold local symmetry of Tm, a rotation of the smooth αβTm heterodimer by 180° around its local interchain axis, will swap the α- with the β-chain and vice versa. Given the large similarities between the two chains (Smillie, 1996 and references therein), it might seem reasonable to assume that the difference in interaction energy between the two pseudosymmetric axial orientations of αβTm with F-actin is small and that both interactions are present in the reconstituted protein complex in solution. If this is the case, we should observe two apparent distance distributions even when only one chain is labeled, because in the two possible pseudosymmetric interactions of Tm with F-actin, the donor will be in two different places. On the other hand, the observation of only one distance would suggest that only one of the two possible interactions is present.
The results from the DD-model analysis of the three singly and doubly labeled αβTm heterodimer samples, in complex with F-actin ± S1 (Table 4, row number 1, 2, and 3 (−S1) and row number 6, 7, and 8 (+S1)), discussed in the previous section, showed that only one apparent distance distribution was needed to fit the FRET data for the singly labeled heterodimers but that two apparent distances were necessary to obtain a good fit for the doubly labeled heterodimer. These two apparent distances, in particular in the absence of S1, are different enough to be well resolved by our analysis. Moreover, these apparent distances were very similar to those found separately for the Tm heterodimer labeled only on one chain or the other. A global fit of the data sets obtained for singly and doubly labeled Tm heterodimers confirmed that indeed the distances were the same (see Table 5). The result of the global fit strongly suggests that the αβTm heterodimer exhibits only one apparent distance when labeled only on one chain. Therefore, for the Tm heterodimer the two chains are nonequivalent in terms of their interaction with F-actin. In other words, the twofold local symmetry of the αα- or ββTm homodimers is effectively broken in the αβTm heterodimer so that a specific interaction with the surface of F-actin is present which, in turn, implies a specific Tm-Tm interaction in the overlap region between two subsequent Tm molecules along the thin filament. Because αβTm is the native dimer, the specificity of interaction with F-actin may have functional importance.
Relationship to F-actin·Tm regulatory states
Previous biochemical studies have indicated that the skeletal F-actin·Tm thin filament equilibrates between two states, closed and open (or myosin-induced), which correspond to off-and-on ATPase activity states, respectively. In the presence of troponin, removal of Ca2+ mainly produces the blocked state (McKillop and Geeves, 1993; Lehrer, 1994; Lehrer and Geeves, 1998). Tm from gizzard muscle inhibits acto-S1 ATPase to a similar extent as rabbit skeletal muscle Tm at low [S1] (Lehrer and Morris, 1984; Williams et al., 1984) and similar equilibrium constants for cooperative S1 binding, but with a cooperative unit size twice as large, are obtained (Maytum et al., 2001), Thus, the F-actin·Tm skeletal and smooth systems are predominantly in the closed biochemical state in the absence of myosin heads, in apparent disagreement with electron microscopy reconstruction studies that indicated that smooth muscle Tm is located near the outer domain of F-actin, a position associated with the blocked state. This probably reflects the small free energy difference between the two states. However, the observation in this work, that S1 shifts smooth Tm to a position further toward the inner domain of F-actin is in agreement with electron microscopy reconstruction data of effects of myosin on Tm position in skeletal thin filaments (Craig and Lehman, 2001). A similar shift would be expected to occur for smooth Tm in view of S1 effects on F-actin·Tm·S1 ATPase. Our previous work showed that Ca2+ shifts skeletal Tm in F-actin·Tm·Tn ∼17° toward the inner domain of F-actin (Bacchiocchi and Lehrer, 2002). Our current work shows that S1 shifts smooth Tm in F-actin·Tm ∼23° toward the inner domain. Although we do not as yet have FRET data for the S1-induced movement of skeletal Tm from the closed state, it can be expected that the movement would be similar. Recent electron microscopy studies on skeletal thin filaments indicated that Tm moves differentially from the blocked to the closed state with the C-terminal half mostly moving to an unblocked site whereas the N-terminal half does not move much (Narita et al., 2002). However, recent FRET studies with a donor at position 245 in the C-terminal one-third of a Cys mutant of skeletal Tm, showed no movement of Tm on binding Ca2+ to reconstituted thin filaments (Bacchiocchi et al., 2003). Clearly, further studies need to be done to understand the differences between the structural and the solution studies.
Acknowledgments
We thank Dr. R. Dominguez for help in the modeling of the label positions on Tm using the program “O” and for the structure match of the Lorenz model for F-actin (Lorenz et al., 1995) and the recent structure of TMR-actin (Otterbein et al., 2001).
This work was supported by National Institutes of Health (AR41637, HL22461, and HL66219).
Corrado Bacchiocchi's present address is Dipartimento di Chimica Fisica e Inorganica, Università, Viale Risorgimento, 4, 40136 Bologna, Italy.
Abbreviations used: FRET, Förster resonance energy transfer; ET, energy transfer; D-only, donor-only; D-A, donor-acceptor; Tm, tropomyosin; S1, single-headed fragment of skeletal muscle myosin (myosin subfragment 1); 1,5-IAEDANS, 5-(2-iodoacetyl-amino-ethyl-amino)naphthalene-1-sulfonic acid; AEDANS, 1,5-IAEDANS after reaction with a sulfhydryl group; TMR, tetramethylrhodamine; DABMI, 4-dimethyl amino phenyl azophenyl 4′-maleimide; PPO, 2, 5-diphenyloxazole.
References
- Arcioni, A., R. Tarroni, and C. Zannoni. 1993. Global target analysis of fluorescence depolarization in model membranes using exponential splines. J. Chem. Soc. Faraday Trans. 89:2815–2822. [Google Scholar]
- Bacchiocchi, C., P. Graceffa, and S. S. Lehrer. 2002. A global FRET study of the myosin-induced movement of αα, ββ and αβ smooth muscle tropomyosin. Biophys. J. 82:241a. (Abstr.) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bacchiocchi, C., and S. S. Lehrer. 2002. Ca2+-induced movement of tropomyosin in skeletal muscle thin filaments observed by multi-site FRET. Biophys. J. 82:1524–1536. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bacchiocchi, C., M. Miki, T. Wakabayashi, and S. S. Lehrer. 2003. The C-terminal region of Tm does not exhibit Ca2+- induced movement in reconstituted thin filaments. Biophys. J. 84:214a. (Abstr.) [Google Scholar]
- Bernstein, H. J. 1999. RasMol 2.7.1. Molecular Graphics Visualization Tool (based on RasMol 2.6 by R. Sayle). http://www.bernstein-plus-sons.com/software/rasmol/.
- Censullo, R., J. C. Martin, and H. C. Cheung. 1992. The use of the isotropic orientation factor in FRET studies of the actin filament. J. Fluorescence. 2:141–155. [DOI] [PubMed] [Google Scholar]
- Cheung, H. C., C.-K. Wang, I. Gryczynski, W. Wiczk, G. Laczko, M. L. Johnson, and J. R. Lakowicz. 1991. Distance distributions and anisotropy decays of troponin C and its complex with troponin I. Biochemistry. 30:5238–5247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Craig, R., and W. Lehman. 2001. Crossbridge and tropomyosin positions observed in native, interacting thick and thin filaments. J. Mol. Biol. 311:1027–1036. [DOI] [PubMed] [Google Scholar]
- Förster, T. 1948. Intermolecular energy transfer migration and fluorescence. Ann. Phys. 2:55–75. [Google Scholar]
- Geeves, M. A., and S. S. Lehrer. 2002. Cooperativity in the Ca2+-regulation of skeletal muscle contraction. In Molecular Control Mechanisms in Striated Muscle Contraction. R. J. Solaro and R. L. Moss, editors. Kluwer, Dordrecht, The Netherlands. 247–269.
- Graceffa, P. 1999. Movement of smooth muscle tropomyosin by myosin heads. Biochemistry. 38:11984–11992. [DOI] [PubMed] [Google Scholar]
- Graceffa, P. 2000. Phosphorylation of smooth muscle myosin heads regulates the head-induced movement of tropomyosin. J. Biol. Chem. 275:17143–17148. [DOI] [PubMed] [Google Scholar]
- Jancso, A., and P. Graceffa. 1991. Smooth-muscle tropomyosin coiled-coil dimers: subunit composition, assembly, and end-to-end interaction. J. Biol. Chem. 266:5891–5897. [PubMed] [Google Scholar]
- Johnson, M. L. 2000. CFS_LS Global, Non-Linear, Least-Squares Fitting Program. Center for Fluorescence Spectroscopy, Baltimore, MD. http://cfs.umbi.umd.edu/cfs/software/.
- Jones, T. A., J.-Y. Zou, S. W. Cowan, and M. Kjeldgaard. 1991. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A. 47:110–119. [DOI] [PubMed] [Google Scholar]
- Kamm, K. E., and J. T. Stull. 1985. The function of myosin and myosin light chain kinase phosphorylation in smooth muscle. Annu. Rev. Pharmacol. Toxicol. 25:593–620. [DOI] [PubMed] [Google Scholar]
- Lakowicz, J. R. 1999. Principles of Fluorescence Spectroscopy, 2nd ed. Kluwer Academic/Plenum, New York, NY.
- Lakowicz, J. R., I. Gryczynski, and H. C. Cheung. 1988. Distance distributions in proteins recovered by using frequency-domain fluorometry. Applications to troponin I and its complex with troponin C. Biochemistry. 27:9149–9160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lakowicz, J. R., I. Gryczynski, W. Wiczk, J. Kusba, and M. L. Johnson. 1991. Correction for incomplete labeling in the measurement of distance distributions by frequency-domain fluorometry. Anal. Biochem. 195:243–254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lakowicz, J. R., G. Laczko, H. Cherek, E. Gratton, and M. Limkeman. 1984. Analysis of fluorescence decay kinetics from variable-frequency phase shift and modulation data. Biophys. J. 46:463–477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lehman, W., V. Hatch, V. Korman, M. Rosol, L. Thomas, R. Maytum, M. A. Geeves, J. E. Van Eyk, L. S. Tobacman, and R. Craig. 2000. Tropomyosin and actin isoforms modulate the localization of tropomyosin strands on actin filaments. J. Mol. Biol. 302:593–606. [DOI] [PubMed] [Google Scholar]
- Lehrer, S. S. 1994. The regulatory switch of the muscle thin filament: Ca2+ or myosin heads? J. Muscle Res. Cell Motil. 15:232–236. [DOI] [PubMed] [Google Scholar]
- Lehrer, S. S., and M. A. Geeves. 1998. The muscle thin filament as a classical cooperative/allosteric regulatory system. J. Mol. Biol. 277:1081–1089. [DOI] [PubMed] [Google Scholar]
- Lehrer, S. S., N. L. Golitsina, and M. A. Geeves. 1997. Actin-tropomyosin activation of myosin subfragment 1 ATPase and thin filament cooperativity. The role of tropomyosin flexibility and end-to-end interactions. Biochemistry. 36:13449–13454. [DOI] [PubMed] [Google Scholar]
- Lehrer, S. S., and E. P. Morris. 1984. Comparison of the effects of smooth and skeletal tropomyosin on skeletal actomyosin subfragment 1 ATPase. J. Biol. Chem. 259:2070–2072. [PubMed] [Google Scholar]
- Lehrer, S. S., and W. F. Stafford. 1991. Preferential assembly of the tropomyosin heterodimer: equilibrium studies. Biochemistry. 30:5682–5688. [DOI] [PubMed] [Google Scholar]
- Lorenz, M., K. J. V. Poole, D. Popp, G. Rosembaum, and K. C. Holmes. 1995. An atomic model of the unregulated thin filament obtained by x-ray fiber diffraction on oriented actin-tropomyosin gels. J. Mol. Biol. 246:108–119. [DOI] [PubMed] [Google Scholar]
- Lorenz, M., D. Popp, and K. C. Holmes. 1993. Refinement of the F-actin model against x-ray fiber diffraction data by the use of a directed mutation algorithm. J. Mol. Biol. 234:826–836. [DOI] [PubMed] [Google Scholar]
- Matsumura, F., and J. J. Lin. 1982. Visualization of monoclonal antibody binding to tropomyosin on native smooth muscle thin filaments by electron microscopy. J. Mol. Biol. 157:163–171. [DOI] [PubMed] [Google Scholar]
- Maytum, R., M. Konrad, S. S. Lehrer, and M. A. Geeves. 2001. Regulatory properties of tropomyosin. Effects of length, isoform and N-terminal sequence. Biochemistry. 40:7334–7341. [DOI] [PubMed] [Google Scholar]
- McKillop, D. F. A., and M. A. Geeves. 1993. Regulation of the interaction between actin and myosin subfragment 1: evidence for three states of the thin filament. Biophys. J. 65:693–701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Narita, A., T. Yasunaga, T. Ishikawa, K. Mayanagi, and T. Wakabayashi. 2002. Ca2+-induced switching of troponin and tropomyosin on actin filaments as revealed by electron cryo-microscopy. J. Mol. Biol. 308:241–261. [DOI] [PubMed] [Google Scholar]
- Otterbein, L. R., P. Graceffa, and R. Dominguez. 2001. The crystal structure of uncomplexed actin in the ADP state. Science. 293:708–711. [DOI] [PubMed] [Google Scholar]
- Smillie, L. B. 1996. Tropomyosin. In Biochemistry of Smooth Muscle Contraction. M. Barany, editor. Academic Press, New York, NY. 63–75.
- Straume, M., and M. L. Johnson. 1992. Monte Carlo method for determining complete confidence probability distributions of estimated model parameters. In Numerical Computer Methods, Vol. 210 of Methods in Enzymology. L. Brand and M. L. Johnson, editors. Academic Press, San Diego, CA. 117–129. [DOI] [PubMed]
- Tao, T. 1978. Nanosecond fluorescence depolarization studies on actin labeled with 1, 5-IAEDANS and dansyl chloride. Evidence for label flexibility. FEBS Lett. 93:146–150. [DOI] [PubMed] [Google Scholar]
- Tao, T., M. Lamkin, and S. S. Lehrer. 1983. Excitation energy transfer studies of the proximity between tropomyosin and actin in reconstituted skeletal muscle thin filaments. Biochemistry. 22:3059–3066. [DOI] [PubMed] [Google Scholar]
- Van Der Meer, B. W., G. Coker III, and S.-Y. S. Chen. 1991. Resonance Energy Transfer, Theory and Data. Wiley-VCH, New York, NY.
- Williams, D. L., L. Greene, and E. Eisenberg. 1984. Comparison of the effects of smooth and skeletal tropomyosin on skeletal actomyosin subfragment 1 ATPase. Biochemistry. 23:4150–4155. [DOI] [PubMed] [Google Scholar]