Abstract
Global fitting algorithms have been shown to improve effectively the accuracy and precision of the analysis of fluorescence lifetime imaging microscopy data. Global analysis performs better than unconstrained data fitting when prior information exists, such as the spatial invariance of the lifetimes of individual fluorescent species. The highly coupled nature of global analysis often results in a significantly slower convergence of the data fitting algorithm as compared with unconstrained analysis. Convergence speed can be greatly accelerated by providing appropriate initial guesses. Realizing that the image morphology often correlates with fluorophore distribution, a global fitting algorithm has been developed to assign initial guesses throughout an image based on a segmentation analysis. This algorithm was tested on both simulated data sets and time-domain lifetime measurements. We have successfully measured fluorophore distribution in fibroblasts stained with Hoechst and calcein. This method further allows second harmonic generation from collagen and elastin autofluorescence to be differentiated in fluorescence lifetime imaging microscopy images of ex vivo human skin. On our experimental measurement, this algorithm increased convergence speed by over two orders of magnitude and achieved significantly better fits.
INTRODUCTION
Biological fluorescent probes are usually characterized by their wavelength of absorption and emission, which depend on the energy spacing between the ground state and the first electronic excited state. Another photophysical property of those fluorophores is the residence time in the excited state called fluorescence lifetime (τ). For common fluorophores used in microscopy, τ is on the order of a few nanoseconds. However, this lifetime can be strongly influenced by the fluorophore's microenvironment. Fluorescence lifetime imaging microscopy (FLIM) has been used in many different biological systems to discriminate between different histological structures (Laiho et al., 2004), to probe intracellular concentrations of cations (Despa et al., 2000; Agronskaia et al., 2003) or pH (Carlsson et al., 2000; Lin et al., 2003) or to map protein interactions using FRET (Verveer et al., 2000b; Becker et al., 2001; Krishnan et al., 2003).
The measurement of fluorescence lifetime is becoming widespread in microscopy due to the commercial availability of gated CCD cameras (Dowling et al., 1998), and single-photon counting cards (Becker et al., 2001; Bascskai et al., 2003). However, the analysis of microscopic data obtained by these instruments is not straightforward, especially if one wants to associate these lifetime measurements with specific photophysical processes such as fluorescence resonance energy transfer. Fluorescence decay, F(t), can often be satisfactorily modeled as a sum of first order kinetic processes and is mathematically represented as a sum of exponentials:
 |
(1) |
where τi is the lifetime of component i and ai its intensity contribution to the fluorescence decay. A major problem in the analysis of time domain lifetime measurements is that this sum of exponential is convoluted with the instrument response G(t). Thus the total intensity measured I(t) is:
 |
(2) |
The fitting of a sum of exponentials convoluted to an instrument response to experimental data is traditionally done by iterative convolution (IC). A guess of the coefficients ai and τi, is used to calculate F(t), which is convoluted with the known instrument response G(t) to obtain Imodel. Comparison between the model and the data, allows the coefficients to be refined in the next iteration. This process requires the successive calculation of a large number of convolutions, which is a time-consuming process. This situation is particularly severe in FLIM where the analysis of each image requires fitting of over one million decay curves.
Since the invention of fluorescent lifetime measurements, many researchers have addressed this problem and have developed different approaches to circumvent the tedious deconvolution process by using Fourier transforms, Laplace transforms, or integrated differential equations. The relative efficiency of these different techniques has been reviewed (McKinnon et al., 1977; O'Connor et al., 1979; Good et al., 1984). Unfortunately, the typical experimental conditions of FLIM (i.e., low fluorophore concentration and short pixel residence time) lead to images with small signal to noise ratio (SNR) and a limited number of time points for each measured decay, which makes those alternative techniques not suitable to our problem.
The most important characteristic of FLIM is the fact that on the order of a million decay curves must be analyzed for a single image. Although it is possible to analyze the decay curve at each pixel independently, the poor SNR of the typical FLIM data results in significant uncertainty in quantifying the parameters of the underlying photophysical model. Previous studies have shown that fitting accuracy can be significantly improved by using a priori information. Global analysis makes use of prior information, such as the spatial invariance of the lifetime of each fluorescent specie in the image, to significantly reduce the degree of freedom in the fitting algorithm resulting in a better measure of the relevant parameters. In systems with two exponential decays, it has been shown for frequency domain imaging that a global fitting, assuming spatially invariant lifetimes, can accurately model parameters whereas pixel-by-pixel analysis can only extract a single average lifetime (Verveer et al., 2000a; Verveer and Bastiaens, 2003).
Although global analysis is a powerful approach, the coupled nature of the analysis routine presents a major challenge. Specifically, global analysis requires simultaneous minimization of over one million decay curves with at least the same number of free parameters. The χ-square surface of this large scale minimization problem is plagued by the presence of many local minima. These local minima significantly hinder the algorithm to locate the true minimum and greatly increase the convergence time.
In this article, we will present a fast global fitting algorithm using iterative convolution to extract the two lifetime components from simulated and measured FLIM images. We further realize that fluorophore distribution and their biochemical environment are in general correlated with the morphology of cells and tissues. We develop a global analysis procedure where the initial guesses for the global fitting parameters are informed by the image morphology. This global fitting technique is compared to other fitting algorithms to show that it delivers the best analysis of the data in terms of fastest convergence and minimal χ-square value.
METHODS
Synthetic images
To test the different fitting algorithms, two types of synthetic data sets were generated. These images were constructed to resemble real time correlated single-photon counting (TCSPC) data but with well-defined image characteristics. In the first type of synthetic images, the decay curve in each pixel is a double exponential of 2 and 4 ns decay convoluted with a Gaussian instrument response of 0.25 ns with Poisson noise added to the theoretical intensity to simulate the photon counting process. The first image shown in Fig. 1 corresponds to a 64 × 64 pixels image where the intensity ratio of one exponential decay to the other varies linearly from one pixel to the other across the image. The total intensity in each pixel is allowed to vary; images with an average total photon count of 103 or 104 photons per pixel are examined. The second type of synthetic image represents a simulated cell image (Fig. 3) where different regions of the image have a given ratio (with 5% Gaussian noise) between the two exponential decays at 2 and 4 ns. All regions except the background have the same initial intensity.
FIGURE 1.
Comparison of the ability of lifetime invariant fit, integrated differential equation and iterative convolution global fits to retrieve the two lifetimes and their ratio hidden in the simulated FLIM image. The lower left image shows five typical decay curves extracted from this image.
FIGURE 3.
Schematic of the segmentation technique. (A) Intensity image of the simulated cell showing the six different intensity regions selected. (B) Summed lifetime information from the six regions selected. (C) Map of the ratio coefficient after the initial fitting procedure. (D) One pixel decay and its fits from the five high intensity regions in the image. (E) Map of the ration coefficients after the global fit. (F) True map of the ratio coefficients.
FLIM images
The FLIM images were obtained in custom-built two-photon microscope as previously published but modified for lifetime imaging (So et al., 2001). This two-photon microscope is based on a modified inverted microscope (Axiovert 110, Zeiss, Göttingen, Germany) using the pulse train of a femtosecond laser (Mira, Coherent, Santa Clara, CA) to excite a subfemtoliter volume of the sample at the focal point of a 40× objective (Fluar, Zeiss). The emitted light is collected by the same objective, spectrally filtered to select the detection wavelength range and measured by a photon counting photomultiplier tube (R7400P, Hamamatsu, Bridgewater, NJ). The signal from the photomultiplier tube is sent to a TCSPC card (SPC-730, Becker-Hickl, Berlin, Germany) housed in the central computer that also controls the scanning of the sample. The TCSPC module generates a histogram of the time of arrival of the photons for each pixel.
Ex vivo human skin was examined in this study. The specimen was stored at −3°C before use. A 1 cm × 1 cm specimen was sandwiched between a cover slip and a damp sponge to maintain its moisture. The dermis was surgically exposed and was placed closest to the cover slip. The femtosecond laser was tuned at 780 nm and the emitted light was selected by a short-pass Schott BG-39 filter. The instrument response was extracted from second harmonic generation (SHG) signal emitted by the sample.
Fibroblast cells (NIH 3T3) were cultured in fetal calf serum, 105 cells were plated in a chambered cover-glass system (4.2 cm2 surface area, Lab-Tek, Nagle Nunc International, Naperville, IL) and incubated overnight before the staining procedure. The culture medium was replaced by 1 ml phosphate-buffered saline (PBS) buffer and 5 μl calcein AM (C3100, Molecular Probes, Eugene, OR) 50 μM in DMSO for ten minutes. The solution was rinsed and replaced by 1 ml PBS with 2 μl of Hoechst 33342 (H-3570, Molecular Probes) for ten more minutes. The staining solution was rinsed away and replaced by PBS buffer. The cells were imaged in the two-photon microscope with the laser tuned at 780 nm. Contribution of scattered light and autofluorescence was minimized by filtering the emission with a 650 nm short pass filter and a BG-39 filter (Chroma, Brattleboro, VT). The instrument response was obtained by convoluting a Gaussian with a 1.35 ns single exponential decay to fit the fluorescence decay of POPOP in methanol. The full width at half maximum (FWHM) is 0.26 ns.
Global fitting
The global fitting model describes the intensity Imodel in each pixel i for the whole image as a function of one set of coefficient ci that needs to be optimized:
 |
(3) |
This is a two-exponential decay model assuming two independent fluorophore species with two lifetimes corresponding to the coefficients c1 and c2. The initial intensity, c2i+1, and the intensity ratio of the two lifetime components, c2i+2, at each pixel, i, are allowed to vary spatially. Note that c2i+2 is set to vary only between 0 and 1 to prevent any negative contributions from the two exponentials. For a 256 × 256 pixel image with ∼50 time points per pixels, the global fitting requires the simultaneous optimization of 6.5 104 coefficients with 3.3 106 experimental data points. Using a least-square estimate χ2 as merit function, the minimization algorithm will optimize the value of all the ci coefficients.
 |
(4) |
The global fitting is realized on a desktop computer (Dell workstation PWS650 with two Intel Xeo 2.4 Ghz processors and 1 GB of RAM, Austin, TX) in MATLAB (The MathWorks, Natick, MA) using the optimization function fmincon based on a reflective Newton method (MathWorks). To reach convergence with this large data set, this function requires the calculation of the least-square (χ2), its gradient (∂χ2/∂ci) and the Hessian (∂2χ2/∂ci∂cj) at every step in the iteration. The convergence criterion corresponds to χ2 changes of <10−13. Before the fitting, an intensity discrimination is performed to remove all background pixels that don't have enough counts to contain valuable information.
Two other fitting algorithms using the same optimization function fmincon have also been implemented. The first one is a time invariant fit (Verveer et al., 2000a) where the two time constants τ1 and τ2 are extracted from a decay curve calculated by the sum of all pixels and are kept as constants to fit the whole image. The model fluorescence decay for each pixel i is then described by the following equation:
 |
(5) |
The second fitting technique is based on integrated differential equation (IDE) (Apanasovich and Novikov, 1990, 1996) also called phase plane method. It provides a linear relationship between integrated forms of Idata(t) and G(t):
 |
(6) |
where  ,
,
 |
and n is the number of lifetimes that one wants to extract from the image (two in this case). The cj and ρj which are the coefficients optimized during the fit are nonlinear functions of the lifetime τi and intensities ai (Apanasovich and Novikov, 1990). As the cj coefficients depend only on τi, it is fairly easy to extend this deconvolution method to global analysis (Apanasovich and Novikov, 1996).
RESULTS
Synthetic images
Fig. 1 shows the test image used to compare the efficiency of IDE and time invariant fitting methods to global fitting. The reference image shown on the left displays the true ratio coefficients c2i+2 and can be compared with the images obtained from the fit. When the average number of count is sufficiently high (104), all three methods are relatively comparable in terms of image and time constant retrieval. One thousand counts per pixel is often considered as the lower limit of counts needed to be able to extract two lifetimes in a fluorescence decay (Gratton et al., 2003). As can be seen in Fig. 1, there is a great variability in intensity between different pixels, which explain the large noise in the c2i+2 coefficients image. The IDE method clearly underperforms the two other techniques but is also the fastest to reach convergence. Time invariant and IC global fitting both give relatively similar images but the time constants are slightly more accurate with the latter. The comparison of the ratio coefficient c2i+2 obtained from the global fit or the time invariant fit with the true ratio hidden in the image is shown in Fig. 2. The effect of the intensity is clearly visible on the accuracy of the fit when comparing panel A and B of the image. A histogram of the deviation of the fitted coefficients from the expected value (cfit − cexpected) is shown for these two situations. The two different optimization methods provide almost identical deviations in the two situations.
FIGURE 2.
(A) and (B) Comparison of the ratio coefficients c2i+2 obtained from the iterative convolution fit (○) and from the time invariant fit (Δ) with the true values of these coefficients for an average photon count number of 103 (A) or 104 (B). (C and D) Histogram of the deviation of the coefficients from the true value for IC fit (solid line) and time invariant fit (dotted line) for an average photon count number of 103 (C) or 104 (D).
The main drawback of a global fitting strategy is the time needed to reach convergence. For this small image this process is almost ten times slower than the time invariant method. And this difference will be exacerbated when trying to fit larger images. However, this situation can be improved by providing an educated guess. This guess can be more or less elaborate and consist of only a few averaged parameters, such as the two lifetimes, or a complete set of coefficients given by a fast converging method.
Other types of method can make use of the image morphology. That is the case, for instance, of the division method, which is based on the fact that neighboring pixels in the image tend to have similar fluorescent decays. Starting with an averaged image made of only sixteen pixels and dividing them further and further until the final pixel size is reached, one seed each step of the optimization with averaged coefficients, which are very close to the optimized value if the region averaged is uniform.
The other method that is based on the image morphology is the segmentation technique. In that case one stipulates that the fluorescence decay rates are correlated with the morphological structures in the image. The distinct morphological structures can be identified by image segmentation based on pixel intensity, lifetime obtained based on fast, local pixel-based estimation, or even image texture. In Fig. 3, we choose to use intensity-based segmentation. By selecting pixels of similar intensity, one should select mostly pixels, which have similar decay curves. Using this technique on the image shown in Fig. 3 A, one can recognize six different intensity levels, which are selected manually. The algorithm then adds up all pixels with similar intensities and a global fit is performed on this very limited set of decay curves (shown in Panel B). The coefficients obtained are then used as initial guess for the final global fitting step. Panel C of this figure shows the map of the ratio of the two lifetimes obtained after the initial fit and can be compared with the true solution Panel F and the result of the global fit Panel E. Panel D displays one typical decay and its fit extracted from one pixel in each region of the image.
To compare the efficiency of the segmentation technique, an intensity-independent division of the image is realized using the sum of the pixels in each quadrant of the image to feed the result to the global optimization algorithm. This strategy will be referred to as quadrant average. Table 1 gives an overview of the different fitting strategies used in this article.
TABLE 1.
Fitting strategies
| Fit type | Deconvolution | Initial guess | Description |
|---|---|---|---|
| Global fit | Iterative convolution (IC) | Full guess | All coefficients are randomly chosen in a physically meaningful range. |
| Image average | Time constants obtained from a fit of the decay curve generated by the sum of all the pixels in the image. | ||
| Quadrant average | Time constants and average coefficients obtained from four decay curves obtained from the sum of all pixels in each quadrant of the image. | ||
| Division | Time constant and coefficients obtained by dividing the image in 16 squares. After convergence, each square is then divided in four until full resolution of the image is achieved. | ||
| Segmentation | Manual selection of different regions in the image to add all similar pixels and obtain a limited number of decay curves from which the time constants and coefficients are obtained. | ||
| IDE guess | Use the IDE deconvolution algorithm to obtain time constants and coefficients for all the pixels in the image | ||
| IDE | Division | Fast deconvolution algorithm based on the integration of the response and decay functions. | |
| Restrained Fit | IC | Image average | The two time constants obtained from a decay curve generated by the sum of all pixels are kept constant whereas the fit optimizes the intensity and ratio coefficients in each pixel. |
Characterization of every optimization schemes in terms of fit types, deconvolution algorithms, and initial guess generations.
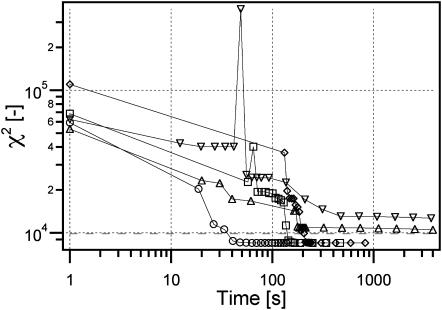
As can be seen in Fig. 4, the evolution of the least-square estimate is greatly influenced by the initial guess of the global fit. This figure clearly demonstrates the advantage of having a preoptimization strategy. Starting from a completely random set of coefficients, the evolution of the algorithm is extremely slow and after more than one hour, the χ2 is still far from the optimum solution depicted by the horizontal dashed line. Comparatively, effective educated guesses, such as the segmentation and the quadrant average methods, lead to convergence in five minutes, which is also the time needed for the time invariant method to produce its result.
FIGURE 4.
Evolution of the least-square estimate χ2 during the global fit (including the prefitting time), using different technique to generate the initial guess: full guess (∇), image average (⋄), division (Δ), segmentation (○), and quadrant average (□). The dashed line is the optimum χ2 calculated using the true coefficients used to generate the image.
The time constants retrieved by the global fit with the segmentation (4.09 and 2.15 ns) or the time invariant fit (3.96 and 1.92 ns) are close to the expected lifetime of 4 and 2 ns. Fig. 5 compares the retrieval of the coefficients determining the ratio between the two lifetimes for these two techniques. Panels A and B represent a plot of the expected ratio coefficients versus the fitted ones. The deviations from the expected value (cfit − cexpected) for both fits were well fitted with Gaussian curves shown in panel C. The mean error in estimating the ratio parameter can be seen from the centers of the distribution. The centers of both fitting methods are skewed from zero. The global fit has a tendency to provide larger ratio with a mean deviation at 0.1 from the expected value and the uncertainty on the ratio value given by the half of the FWHM is ± 0.1. The time invariant fit has a mean deviation at −0.5 from the expected ratio but the FWHM is 0.3. Deviations from the other global IC fits using different optimization strategies that reach convergence are identical to the one obtained for the segmentation.
FIGURE 5.
Comparison of the ratio coefficients c2i+2 obtained from the iterative convolution fit (A) and from the time invariant fit (B). (C) Gaussian fit of the histogram of the difference between the true coefficients and the ones obtained the iterative convolution (solid line) and time invariant (dotted line) optimizations.
TCSPC data
The first experimental data set used to test the efficiency of the segmentation-based global fit is a FLIM image from the dermis of the skin. The dermis is constituted from a collagen matrix and with elastin fibers. It is well-known that the collagen generates a strong SHG signal (Georgiou et al., 2000), whereas elastin is one of the many endogenous fluorophores found in the skin (Zeng et al., 1995; König and Riemann, 2003). Fig. 6 B shows the lifetime of low and high autofluorescence regions in the dermis. One clearly sees the difference between the longer duration of the fluorescence emission from elastin compared to the instantaneous response from SHG due to collagen. Due to this large difference in decay time, a map of the lifetime generated using the rapid lifetime determination method (Ballew and Demas, 1989) shows more contrast than a simple intensity image, thus a lifetime-based segmentation, instead of intensity-based segmentation, was used to generate the initial guess for the fit.
FIGURE 6.
(A) Map of the ratio of SHG versus autofluorescence obtained from the global fit. (B) Sample of decay pixels and their fits from high (○) and low (□) fluorescence regions. (C) Evolution of the least-square estimate χ2 during the optimization using different strategies: full guess (∇), image average (⋄), division (Δ), segmentation (○), and quadrant average (□).
Panel A of Fig. 6 shows the result from the global fit and is a map of the ratio of SHG versus autofluorescence intensity and allows to highlight the distribution of the elastin fibers in the image. Panel B of this figure shows the typical instantaneous response of the collagen matrix and the emission from the elastin fiber pixels with a long decay constant due to the fluorescence of this compound. The two time constants extracted from the image are 45 ps, which is much shorter than the instrument response of 350 ps, and 1.84 ns, which matches well with the 2.3 ns expected from elastin (Maarek et al., 2000).
Similarly to what has been done with the synthetic image of the cell, a comparison of the efficiency of the different fitting strategies is shown in Fig. 6 C. Except for the random initial guess and the image average, the other fitting techniques seem relatively equivalent and yield very comparable time constants and images. The convergence of the fit is obtained in 12 min for the segmentation and quadrant average and 15 min for the division technique. The full guess and the image average do not lead to full convergence in <3h, but the time constants are close to the expected values, as seen on Table 1. This is the opposite of the IDE guess and time invariant techniques which both converge quickly but fail to recover the nanosecond time constant for the fluorescence contribution to the image.
Fig. 7 displays the results obtained with the cells stained with calcein and Hoechst. Calcein is typically used as a live cell label and is distributed throughout the cell. Hoechst is a nucleic label and is localized in the nucleus. In panel A of this figure, the intensity distribution of the fluorescence is shown. The nucleic region of the cell is approximately twice as fluorescent as the rest of the cytoplasm due to the additive emission from the two dyes in this region. Very low photon count is measured in the background of the image, which is discarded for the global fitting optimization. Panel B shows typical decays recorded in the cytoplasmic and nucleic regions of the cell. The solid lines represent the result of the fit for these decay curves. In the inset, these fitting curves are normalized to allow for a better comparison of the lifetime and demonstrate the small difference between the emission decays arising from these two different regions of the cells. The two time constants extracted from the image using the global fit based on an intensity segmentation method are 3.67 ns and 2.37 ns. These two time constants are in good agreement with the values found in the literature for the lifetime of these dyes in water: 3.85 for calcein (Periasamy et al., 1991) and 2.3 for Hoechst (Sailer et al., 1997). The distribution of the ratio of the shortest lifetimes throughout the image is shown in the panel C of Fig. 7. A histogram of the distribution of the ratio coefficient c2i+2 in the nucleic region of the image (see panel D) is well fitted by a Gaussian centered at 0.6 with 0.28 FWHM. If one excludes the large peak at zero, the histogram of the ratio in the cytoplasmic domain of the image can be represented by a Gaussian distribution centered at 0.3 with a FWHM of 0.5.
FIGURE 7.
(A) Intensity map of the fluorescence emitted by cells stained with calcein AM and Hoechst 33342. (B) Two typical lifetime measured from both the cytoplasmic (□) and nucleic (○) regions of the cells. The solid lines represent the fit of these decays. The inset shows these same fits normalized. (C) Map of the proportion of 2.4 ns decay versus 3.7 ns decay in each pixel of the image. (D) Histogram of the lifetime ratio for the cytoplasmic (solid line) and nucleic (dotted line) regions.
Fig. 8 compares the speed of convergence for the different optimization methods already tested above. In this image, where the two lifetimes are very close, the best method to reach the optimum is clearly the segmentation technique. The only other method to reach convergence is the quadrant average, but the time constants do not correspond to the lifetimes of the dyes. On the opposite, the division and image average strategies provide meaningful time constants but do not reach convergence in <2 h. The time invariant delivers meaningless information because it is not able to differentiate between the two different decays.
FIGURE 8.
Evolution of the least-square estimate χ2 during the optimization using different strategies: full guess (∇), image average (·), division (Δ), segmentation (○), and quadrant average (□).
DISCUSSION
The performance of the time invariant fit and the global fitting are similar when applied to simulated images. However, with both real TCSPC images, the time invariant fit fails to provide the correct time constants and validates the use of the global fitting. The simultaneous optimization of the lifetime and the intensity in each pixel is a very powerful technique to extract the lifetime information especially from very noisy data. There are mainly two disadvantages to this technique. The first one is that it requires the simultaneous fitting of very large data sets. Fortunately, the large-scale algorithms, which are implemented in many commercial software such as MATLAB, prove to be very capable of handling the many parameters to optimize.
The second problem is the time needed to reach convergence that can really be prohibitive. Unfortunately, alternative deconvolution methods are not applicable due to the high noise of the data. The results obtained with IDE optimization are acceptable with high photon counts per pixels, but become very unreliable at lower counts found in real TCSPC data. Thus the only appropriate deconvolution method that can be used is the iterative convolution. As mentioned above, for each iterative step, not only the Imodel has to be calculated, but also the gradient and the Hessian matrices. These three matrices all require the convolution of the theoretical exponential decay or its derivatives with the instrument response, which leads to a tremendous number of convolutions to be calculated at each step. Due to the limited number of time points per pixel acquired in a FLIM image, this convolution process is not a too large burden.
Enhancement of the iterative step calculation was realized by profiling of the code, but it is not sufficient to obtain a fast converging algorithm. Only a hundred iterations require close to an hour of calculation. Thus, the total number of IC calculated on whole image has to be decreased and replaced by less time-consuming iteration with fewer pixels.
The results summarized in Table 2 clearly demonstrate the efficiency of the segmentation technique for all the different images tested. In comparing the different strategies used to reach an optimal guess for the optimization, one can understand why this technique is so efficient. Using a single decay curve to extract τ1 and τ2 leads to too much ambiguity in the time constants as can be seen with the image average or time invariant techniques and using a few number of decays allows this uncertainty to be resolved and to obtain the correct time constants quickly. The two optimization schemes (i.e., the segmentation and the quadrant average) that make use of this proved to be the most efficient ones. The former has some advantages over the latter. First, the selection of the different regions based on intensity or lifetime allows an initial set of decay curves to be obtained. These curves are representative of the various regions of the image and thus of the different types of decays observed. As each decay curve is considered equal in this first optimization step, it allows the weight of certain regions of the image to increase even if they are representative of only a small number of pixels. This is clearly not the case for the quadrant average, were the difference in decay curves is based only on asymmetries in the image but the four regions represent the same number of pixels. The demonstration of this is given by the fit of the doubly stained cells, where the difference in lifetime is minimal and proper choice of the set of decay curve helps to reach a meaningful optimum quickly. The second advantage is that the intensity coefficients obtained by the segmentation guess can be used to generate an educated guess, which closely resembles the final image. Since the segmentation and the quadrant average require almost the same time to reach convergence, this feature does not seem to lead to a major advantage in the synthetic cell image analysis and the dermis TCSPC fitting. The explanation may be that the optimization of the individual pixel parameter is done quickly once the correct time constants are obtained because they are independent of each other as calculated in the Hessian matrix.
TABLE 2.
Summary of optimizations results
| Synthetic image | Dermis | Calcein/Hoechst cells | |
|---|---|---|---|
| Expected lifetimes | 4.00 2.00 ns | 2.30 0.0 ns | 3.85 2.35 ns |
| Segmentation | 5 2.15 | 2 0.063 | 3.67 2.37 |
| 40 (5 min) | 16 (12 min) | 217 (23 min) | |
| Full guess | 3.49 3.23 | 1.78 0.043 | 3.02 2.89 |
| 500 (1 h)* | 300 (3 h)* | 1000 (1 h 42)* | |
| Image average | 5 2.17 | 1.39 0.065 | 3.56 2.37 |
| 79 (11 min) | 300 (4 h)* | 1000 (1 h 45)* | |
| Quadrant average | 4.10 2.15 | 2 0.052 | 15.1 2.83 |
| 41 (5 min) | 18 (12 min) | 451 (47 min) | |
| Division | 5 1.85 | 1.84 0.027 | 4 2.40 |
| 500 (1 h)* | 27 (15 min.) | 1000 (1 h 51)* | |
| Time invariant | 3.96 1.92 | 0.22 0.17 | 3.07 3.07 |
| 44 (5 min) | 44 (8 min) | 25 (3 min) | |
| IDE guess | 3.83 2.23 ns | 7.3e12 0.34 | |
| 500 (1 h 12)* | 36 (30 min) |
Summary of the time constants obtained for the optimizations using different strategies to generate the initial guess. The number of iterations to reach convergence is given below with the average time needed for the total optimization.
Fits were stopped before reaching full convergence.
Analysis of dermis data by lifetime measurement delivers a way to separate efficiently the contribution of the autofluorescence of the tissue and the SHG from the collagen. The corroboration of the lifetime data and spectrally resolved measurements proves unambiguously the presence of elastin fibers within the collagen matrix in the dermis (Laiho et al., 2004). We see that global fitting of this image is done easily with all kind of different optimization path due to the large difference in the two lifetimes contained in the image. The equivalence of the different optimization approaches is also a result from the relatively uniform nature of this image and does not readily segmented into few discrete regions.
The advantage of the segmentation technique becomes clearly evident when the two lifetimes are much closer and the image can be segmented into few discrete regions as is the case with the image of the cell stained with calcein and Hoechst. Interestingly, the optimized guess generated by the fit on the segmentated parts yields a higher least-square estimate than all the other educated guesses generated by different techniques, and also higher than some completely random guesses of coefficients. This indicates that the optimization hypersurface is relatively flat, which is understandable due to the closeness of the two time constants. Trying to minimize χ2 from any point on the hypersurface does not always lead to the global minimum and this emphasizes the need for a good starting point for the optimization as can be given by the segmentation method.
The time constants obtained from the optimization match well with the values found in the literature for calcein and Hoechst. In the nucleic region, the contribution of the two dyes to the fluorescence signal estimated from cells stained with a single marker is in good agreement with the ratio coefficients of 60% retrieved by the global fit. The value of the FWHM of 0.28 is comparable to the 0.2 obtained for the synthetic image at similar intensity levels. However, the ratio coefficients obtained for the cytoplasmic regions of the cells are surprisingly high. One would expect to measure only fluorescence from the calcein in this part of the cell and thus obtain ratio close to zero for the contribution from the fast lifetime. However, when looking at cells stained only with Hoechst, one can see a small emission coming from the cytoplasmic region. This signal, which varies strongly throughout the cytoplasm, could account for a ratio from zero up to 30% at most, when comparing with the intensity of cells stained only with calcein. As it has already been observed in Fig. 5, at low values of the ratio coefficients and comparable photon count levels, the global fit has a tendency to provide larger coefficients than expected.
One has to concede that the large noise in the TCSPC traces cannot allow for a perfect recovery of these two contributions especially in regions of lower photon count rate like the cytoplasm. Multiple exponential fits on noisy data have a tendency to overfit the data (i.e., a double exponential would produce a lower χ2 even for a single exponential decay). This might become an even bigger problem if one wants to apply this algorithm to retrieve a larger number of time constants.
The quantification of the ratio of the two lifetimes in a FLIM is linked to a subject that has attracted a lot of attention, which is the possibility to use fluorescence lifetime imaging to obtain quantitative FRET maps of the distributions of protein interactions inside a cell (Verveer et al., 2000b; Krishnan et al., 2003). In this kind of study, one expects to measure two different lifetimes produced by interacting and noninteracting proteins. The noninteracting labeled proteins exhibit a natural lifetime of the dye, whereas the interacting proteins will exhibit a shorter lifetime due to the quenching of the emission by energy transfer. Assuming that there is a given geometry of interaction between the proteins, a constant FRET lifetime should be measured. The ratio of these two lifetimes throughout the cells gives the ratio of interactions between the studied proteins. Thus the global fitting algorithm developed here will be ideal to analyze this kind of data. The results obtained with dual cell labeling clearly show the validity of the technique but also stresses the need for high FRET efficiencies (i.e., large separation in lifetime) and high photon counts per pixels to decrease the uncertainty in the measurement.
CONCLUSION
Global fitting algorithms have been shown to work for time domain lifetime microscope images. Due to the large number of coefficients that need to be optimized simultaneously, generating an educated guess as starting point for global optimization can reduce tremendously the time needed for convergence. Using simulated data as well as experimental TCSPC images, it has been shown that segmentation of the image based either on intensity or lifetime provides the best way to reach the optimum in a timely fashion for all our data sets.
Images of the dermis, where collagen and elastin contribute to the emitted signal due to SHG and fluorescence respectively were analyzed with the global fitting algorithm. Differentiation between the two components based on lifetime is easy due to the large difference in emission decay time and yields a map of the elastin fibers embedded in the collagen matrix. Most of the guess generating strategies converged with the segmentation method reaching convergence the fastest.
Staining of the cell by calcein and Hoechst resulted in an image containing two close lifetimes that were harder to dissociate. Again, the segmentation technique proved to be the most adequate because it generated an educated guess that was very close to the final optimum and thus required only a limited time to converge. The repartition of the two dyes in the final image corresponds with the expected one and the time constants retrieved are in good agreement with literature values. This proves that this algorithm is able to discriminate between different structures in biological specimen based on their emission lifetime, and that it will be applicable to FRET-FLIM measurements.
Acknowledgments
S. Pelet thanks the Swiss National Science Foundation.
The authors acknowledge the support of the National Institutes for Health grant NIHPOIHL64858.
APPENDIX
The FLIM images data can be represented by a two dimensional matrix where each row contains the temporal evolution of the fluorescence at a given pixel. To obtain the least-square value, one has to calculate a model matrix from a vector c of fitting coefficients and along a time axis t. The fluorescence decay is modeled by a double exponential decay:
 |
(A1) |
where NP is the number of pixels and NT the number of time points. The Model matrix is then equal to the convolution (*) between the Decay matrix and a Gaussian impulse response represented by a vector G:
 |
(A2) |
The least-square value estimates the difference between the model and the data:
 |
(A3) |
This value is the merit function used by the optimization algorithm implemented in MATLAB. It will determine if the optimization reaches the minimum. To improve convergence, the algorithm requires the calculation of the first and second derivatives as function of the coefficients, the gradient, and Hessian, respectively:
 |
(A4) |
 |
(A5) |
Note that for the coefficients c(2i + 1) or c(2i + 2), the derivative of the Decay(i,j) relative to the coefficient c(k), where 2i + 1 (resp. 2i + 2) ≠ k, is equal to zero. Thus for all the coefficients except c(1) and c(2), the sum over all i can be omitted and one need to sum only over all j when 2i + 1 (resp. 2i + 2) = k. This also implies that the Hessian matrix is mostly sparse. With only the two first column and rows which contains the cross-derivatives as function of c(1) and c(2) which are full as well as the diagonal and the off-diagonal elements where k = 2i + 1 and l = 2i + 2 corresponding to the cross-derivative for a given pixel decay.
It is clear that estimating the gradient and Hessian along with the χ2 at each step of the iteration for a large number of pixels is a time-consuming process in part due to the large number of convolutions to be calculated. Obtaining educated values for the coefficients using a reduce set of pixel representative of different regions of the image allows to reduce greatly the time needed to reach convergence.
In the first part of the program, the computer removes all pixels in the image that contain intensity lower than a certain threshold value. The remaining pixels are then summed in different groups to generate a reduced number of pixels for the first run of the optimization loop. In the case of the image average all the pixels are added to generate a single decay curve and in the quadrant average, the pixels in each quadrant of the image are sum to form four decays.
The intensity-based segmentation algorithm works in a similar fashion. It requires the user to select a few numbers of subregions that are representative of the different intensities found in the image. The program averages the intensity in each subregion and goes through the image to group all pixels with intensity values close to each subregion together. The decays of these grouped pixels are added together to generate a few decay curves used as the initial guess for the optimization.
An identical procedure is followed for the lifetime segmentation. The difference relies in the criteria used for the segmentation. In that case, a lifetime image is generated using the rapid lifetime method (Ballew and Demas, 1989), which produces an average lifetime for each pixel:
 |
(A6) |
where Tmax is the maximum of the decay curve close to time zero and Δt is a time interval that can be adjusted depending on the lifetime range measured. All pixels are grouped to subregions as before and the decay curves for each subregion are used as initial guesses for the optimization.
The MATLAB code for the global fitting algorithm with different optimization schemes will be made available through our web site: http://www.mit.edu/∼solab/.
References
- Agronskaia, A. V., L. Tertoolen, and H. C. Gerristen. 2003. High frame rate fluorescent lifetime imaging. J. Phys. D: Appl. Phys. 36:1655–1662. [Google Scholar]
- Apanasovich, V. V., and E. G. Novikov. 1990. Deconvolution method for fluorescence decays. Opt. Commun. 78:279–282. [Google Scholar]
- Apanasovich, V. V., and E. G. Novikov. 1996. The method of fluorescence decays simultaneous analysis. Rev. Sci. Instrum. 67:48–54. [Google Scholar]
- Ballew, R. M., and J. N. Demas. 1989. An error analysis of the rapid lifetime determination method for the evaluation of single exponential decays. Anal., Chem. 61:30–33. [DOI] [PubMed] [Google Scholar]
- Bascskai, B. J., J. Skoch, G. A. Hickey, R. Allen, and B. T. Hyman. 2003. Fluorescence resonance energy transfer determinations using multiphoton fluorescence lifetime imaging microscopy to characterize amyloid-beta plaques. J. Biomed. Opt. 8:368–375. [DOI] [PubMed] [Google Scholar]
- Becker, W., K. Benndorf, A. Bergmann, C. Biskup, K. Koenig, U. Tirplapur, and T. Zimmer. 2001. FRET Measurements by TCSPC Laser Scanning Microscopy. European Conference on Biomedical Optics, ECBO, Munich, Germany.
- Carlsson, K., A. Liljeborg, R. M. Andersson, and H. Brismar. 2000. Confocal pH imaging of microscopic specimens using fluorescence lifetimes and phase fluorometry: influence of parameter choice on system performance. J. Microsc. 199:106–114. [DOI] [PubMed] [Google Scholar]
- Despa, S., J. Vecer, P. Steel, and M. Ameloot. 2000. Fluorescence lifetime microscopy of the Na+ indicator sodium green in HeLa cells. Anal., Biochem. 281:159–175. [DOI] [PubMed] [Google Scholar]
- Dowling, K., M. J. Dayel, M. J. Lever, P. M. W. French, J. D. Hares, and A. K. L. Dymoke-Bradshaw. 1998. Fluorescence lifetime imaging with picosceond resolution for biomedical imaging. Opt. Lett. 23:810–812. [DOI] [PubMed] [Google Scholar]
- Georgiou, T., T. Theodossiou, V. Hovhannisyan, K. Politopoulos, G. S. Rapti, and D. Yova. 2000. Second and third optical harmonic generation in type I collagen by nanosecond laser irradiation, over broad spectral region. Opt. Commun. 176:253–260. [Google Scholar]
- Good, H. P., A. J. Kallir, and U. P. Wild. 1984. Comparison of fluorescence lifetime fitting techniques. J. Phys. Chem. 88:5435–5441. [Google Scholar]
- Gratton, E., S. Breusegem, Q. Ruan, and N. Barry. 2003. Fluorescence lifetime imaging for the two-photon microscope: time-domain and frequency-domain methods. J. Biomed. Opt. 8:381–390. [DOI] [PubMed] [Google Scholar]
- König, K., and I. Riemann. 2003. High-resolution multiphoton tomography of human skin with subcellular spatial resolution and picosecond time resolution. J. Biomed. Opt. 8:432–439. [DOI] [PubMed] [Google Scholar]
- Krishnan, R. V., A. Masuda, V. E. Centonze, and B. Herman. 2003. Quantitative imaging of protein-protein interactions by multiphoton fluorescence lifetime imaging microscopy using a streak camera. J. Biomed. Opt. 8:362–367. [DOI] [PubMed] [Google Scholar]
- Laiho, L. H., S. Pelet, T. M. Hancewicz, P. D. Kaplan, and P. T. C. So. 2004. Two-photon 3-D mapping of tissue endogenous fluorescence species based on fluorescence emission spectra. J. Biomed. Opt. Submitted. [DOI] [PubMed]
- Lin, H.-J., P. Herman, and J. R. Lacowicz. 2003. Fluorescence lifetime-resolved pH imaging of living cells. Cytometry A. 52A:77–89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maarek, J.-M. I., L. Marcu, W. J. Snyder, and W. S. Grundfest. 2000. Time-resolved fluorescence spectra of arterial fluorescent compounds: reconstruction with the Laguerre expansion technique. Photochem. Photobiol. 71:178–187. [DOI] [PubMed] [Google Scholar]
- McKinnon, A. E., A. G. Szabo, and D. R. Miller. 1977. The deconvolution of photoluminescence data. J. Phys. Chem. 81:1564–1570. [Google Scholar]
- O'Connor, D. V., W. R. Ware, and J. C. Andre. 1979. Deconvolution of fluorescence decay curves. A critical comparison of techniques. J. Phys. Chem. 83:1333–1343. [Google Scholar]
- Periasamy, N., M. Armijo, and A. S. Verkman. 1991. Picosecond rotation of small polar fluorophores in the cytosol of sea urchin eggs. Biochemistry. 30:11836–11841. [DOI] [PubMed] [Google Scholar]
- Sailer, B. L., A. J. Nastasi, J. G. Valdez, J. A. Steinkamp, and H. A. Crissman. 1997. Differential effects of deuterium oxide on the fluorescence lifetimes and intensities of dyes with different modes of binding to DNA. J. Histochem. Cytochem. 45:165–175. [DOI] [PubMed] [Google Scholar]
- So, P. T. C., K. H. Kim, C. Buehler, B. R. Masters, L. Hsu, and C. Y. Dong. 2001. Basic principles of multiphoton excitation microscopy. In Methods in Cellular Imaging. A. Periasamy, editor. Oxford University Press, New York. 147–161.
- Verveer, P. J., and P. I. H. Bastiaens. 2003. Evaluation of global analysis algorithms for single frequency fluorescence lifetime imaging microscopy data. J. Microsc. 209:1–7. [DOI] [PubMed] [Google Scholar]
- Verveer, P. J., A. Squire, and P. I. H. Bastiaens. 2000a. Global analysis of fluorescence lifetime imaging microscopy data. Biophys. J. 78:2127–2137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Verveer, P. J., F. S. Wouters, A. R. Reynolds, and P. I. H. Bastiaens. 2000b. Quantitative imaging of lateral ErbB1 receptor signal propagation in the plasma membrane. Science. 290:1567–1570. [DOI] [PubMed] [Google Scholar]
- Zeng, H., C. MacAulay, D. I. McLean, and B. Palcic. 1995. Spectroscopic and microscopic characteristics of human skin autofluorescence emission. Photochem. Photobiol. 61:639–645. [DOI] [PubMed] [Google Scholar]