Abstract
The upstream regulatory region of the Drosophila melanogaster hsp26 gene includes two DNase I-hypersensitive sites (DH sites) that encompass the critical heat shock elements. This chromatin structure is required for heat shock-inducible expression and depends on two (CT)n•(GA)n elements bound by GAGA factor. To determine whether GAGA factor alone is sufficient to drive formation of the DH sites, we have created flies with an hsp26/lacZ transgene wherein the entire DNA segment known to interact with the TFIID complex has been replaced by a random sequence. The replacement results in a loss of heat shock-inducible hsp26 expression and drastically diminishes nuclease accessibility in the chromatin of the regulatory region. Chromatin immunoprecipitation experiments show that the decrease in TFIID binding does not reduce GAGA factor binding. In contrast, the loss of GAGA factor binding resulting from (CT)n mutations decreases TFIID binding. These data suggest that both GAGA factor and TFIID are necessary for formation of the appropriate chromatin structure at the hsp26 promoter and predict a regulatory mechanism in which GAGA factor binding precedes and contributes to the recruitment of TFIID.
The positioning of nucleosomes is of critical importance in regulating gene expression in eukaryotes. Packaging promoter DNA in a nucleosome prevents TATA binding protein (TBP) from binding to the TATA element, blocking association of the full TFIID complex and RNA polymerase II (for reviews, see references 1, 3, and 37). It has generally been observed that the TATA box and critical regulatory elements of an active or inducible gene lie in DNase I-hypersensitive sites (DH sites), discontinuities in the nucleosome array (reviewed in reference 12). Creation of appropriate DH sites appears to be essential for gene activity in many cases. Multiple mechanisms that are involved in counteracting nucleosomal repression have been identified. In particular, a number of remodeling and histone modification activities that facilitate the binding of DNA-specific factors with a concomitant change in the nucleosome array have been identified (for a review, see reference 1).
In vivo studies have shown that GAGA factor plays a critical role in establishing the nucleosome-free DH sites observed at the promoter and upstream regulatory elements of the hsp26 and hsp70 heat shock genes in Drosophila melanogaster prior to activation (reviewed in references 17 and 30). It appears that the chromatin remodeling complex NURF can have a key role in this process (42, 43). The NURF complex can alter nucleosome structure in an ATP-dependent manner to allow GAGA factor binding in an in vitro chromatin assembly assay, resulting in nuclease sensitivity at the hsp70 promoter (42, 43). However, our understanding of the steps required to obtain the in vivo nucleosome-free DH site, and the potential for activation, remains incomplete.
The Drosophila hsp26 gene, as well as the hsp70 gene, is characterized not only by preset DH sites in the promoter region but also by the presence of a paused RNA polymerase II, having stalled after synthesis of ∼25 bases of RNA (reviewed in references 29 and 30). Heat shock induces the formation of trimers of the heat shock factor (HSF); binding of the HSF to its target sequences, located in the DH sites, releases RNA polymerase II to proceed with elongation. An illustration of the hsp26 promoter in the inactive but inducible state is shown in Fig. 1A. GAGA factor, the TFIID complex, and RNA polymerase II are all present in the DH sites, while the regions upstream and downstream are packaged in a specific nucleosome array. Folding of the DNA around the nucleosome positioned between the two DH sites appears likely to bring the distal regulatory site into proximity with the proximal regulatory site. The above considerations suggest a possible cooperation of GAGA factor with TFIID and other members of the preinitiation complex in creation of the DH sites and positioning of nucleosomes in the upstream region of the hsp26 gene.
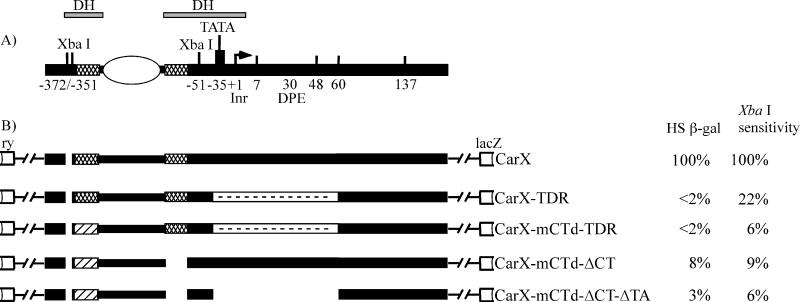
FIG. 1.
CarX-based constructs of the hsp26 gene promoter used in this work. (A) Current view of the organization and chromatin structure of the endogenous hsp26 gene promoter. The white oval represents a positioned nucleosome separating the two DH sites, and the cross-hatched boxes represent regions containing (CT)n repeats (the target sequences for GAGA factor) and the major heat shock elements (the target sequences for the gene-specific activator, heat shock factor). The TATA box (binding site for TBP) and Initiator Element (Inr) and Downstream Promoter Element (DPE) are also shown. Numbers below the line mark the borders (in base pairs) of various changes made in the transgenic constructs depicted in panel B. +1 is the transcription start site (indicated by a bent arrow). (B) Structures of various CarX-based constructs used in this work. The hatched boxes depict mutation in the distal GAGA factor binding site, the gaps show deletions of the corresponding sequences, and the white boxes with dashes depict replacement with random sequence. The white boxes depict ry, the rosy gene, a visible marker used in all transgenic constructs. The constructs in panel B are based on the CarX construct (32). The heat shock-inducible β-galactosidase activity (HS β-gal) of each construct, normalized to the corresponding control CarX transgene (set at 100%), is shown to the right of the constructs. XbaI cleavage of chromatin (from larval nuclei of each line, no heat shock) at the proximal DH site, normalized to the CarX control transgene (set at 100%), is shown in the XbaI sensitivity column. These results are the averages of measurements in three to seven independent lines for each transgene (the variation was ±5%).
In this study, we ask whether the presence of GAGA factor in the 5′ regulatory region of the hsp26 gene is sufficient to direct formation of the DH sites or whether this transition also requires the presence of an adjacent TFIID complex. Do these components work independently, recruit each other, or act synergistically in nucleosome displacement? For this investigation we have created a new set of transgenic constructs with replacement of the TFIID binding site and have utilized some transgenic constructs made previously (20, 32) with various changes in the hsp26 promoter region. We have analyzed the impact of mutations in GAGA factor and TFIID binding sites on generation of the proximal DH site and on the binding of these proteins to the altered regulatory region. The results show that replacement of the TFIID binding site by a random sequence abolishes nuclease accessibility to an extent similar to that caused by the loss of GAGA factor binding sites (32). These data indicate that both GAGA factor and the TFIID complex are necessary for the formation of an appropriate chromatin structure (DH sites) at the hsp26 promoter. Chromatin immunoprecipitation experiments show that GAGA factor can bind its target sequences when TFIID binding is reduced. In contrast, binding of the TFIID complex to its target sequence is decreased when GAGA factor binding sites are mutated. These results suggest a model in which GAGA factor binds to its target independently and prior to binding of the TFIID complex; to complete formation of the characteristic chromatin structure of the uninduced but activatable hsp26 promoter, binding of the TFIID complex at its normal site is required.
MATERIALS AND METHODS
Plasmid construction.
Experimental manipulations of recombinant DNA were performed as described by Sambrook et al. (39). The structures of all hsp26 transgenes used in this work are depicted schematically in Fig. 1B and 6B. Transformation constructs CarX (32) and cP-351 (20) were used as the starting plasmids. Both are based on transformation vector Car20T (52) and carry a genomic fragment of the rosy gene as a visible transformation marker. The cP-351 transformation plasmid contains the hsp26 gene promoter sequences starting from a distal XbaI site at position −351, fused in frame with the lacZ gene at position +632. This plasmid and its derivative, cP-ΔCT (same as cPΔCT·GA), with a deletion of the proximal GAGA factor binding site, were described by Glaser et al. (20). Plasmids with downstream deletions cP-ΔD1 (deletion of positions +7 to + 48) and cP-ΔD2 (deletion of positions +48 to +137) were made by standard techniques from cP-351, and cP-ΔCT-ΔD1 and cP-ΔCT-ΔD2 were made by standard techniques from cP-ΔCT.
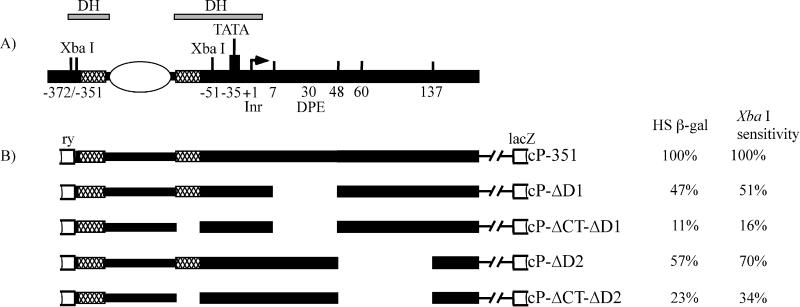
FIG. 6.
cP-351-based constructs of the hsp26 gene promoter used in this work. (A) Current view of the organization and chromatin structure of the endogenous hsp26 gene promoter. For explanation of symbols, see the legend to Fig. 1. The constructs in panel B are based on cP-351 (20). The heat shock-inducible β-galactosidase activity of each construct (HS β-gal), normalized to the corresponding control transgene (cP-351) (set at 100%), is shown to the right of the constructs. XbaI cleavage of the chromatin from each line, again normalized to the cP-351 control transgene, is shown in the XbaI sensitivity column.
CarX contains a larger fragment of the hsp26 gene (from positions −1917 to +632) with the same lacZ gene fusion as in cP-351. The XbaI-SacI fragment (from positions −52 to +490) of CarX was recloned in the pGEM-3Zf(+) plasmid; subsequent manipulations to change the hsp26 gene promoter region were performed using this plasmid. Final versions of the new transgenes were made by reinserting the modified fragment into transformation vector Car20T. CarX and CarX-mCTd-ΔCT were described by Lu et al. (32).
Creation and cloning of a random sequence replacement and point mutations in the hsp26 TATA box.
The random sequence introduced in transgenic constructs designated TDR (TATA-downstream replacement) starts from the first nucleotide of the TATA box (position −35) and extends to position +60 downstream from the transcription start site (total of 95 bp). The TDR sequence was computer generated and inspected for the absence of obvious motifs for known DNA binding proteins, including target sequences for GAGA factor. To facilitate cloning and mapping, restriction sites for NheI and ApaI were introduced at the ends of the sequence; the second base in the NheI recognition site replaces the first base in the TATA box. To form 95 bp of random sequence, two overlapping synthetic oligonucleotides (58 bases each) were hybridized and extended with Sequenase version 2.0 DNA polymerase (U.S. Biochemical Corp., Cleveland, Ohio). A pGEM-3Zf(+) plasmid containing a XbaI-SacI fragment of the hsp26 upstream region (positions −52 to + 490) was PCR amplified outward using a pair of primers to introduce NheI and ApaI sites in the desired positions. The random sequence fragment, trimmed by NheI and ApaI, was ligated with pGEM-3Zf(+) plasmid containing the modified XbaI-SacI fragment of the hsp26 upstream region (positions −52 to + 490) cut by NheI and ApaI. Thus, the original sequence from positions −39 to +64 at the hsp26 gene promoter,5′cgggTATAAAAGCAGCGTCGCTTGACGAACAGAGCACAGATCGAATTCAAAAATCGAGCAGTGAACAACTCAAAGCAACTTTGCGCAAAAGCAAAACTTcaaa3′ (sequences to be replaced in capital letters; the TATA box underlined; lowercase letters are unchanged sequences), was replaced by 5′cgggCTAGCAAAGCGAACCTAAAATACGGTCCGAATAGACAGGAGGTCCAACATGTTGAAAAAGCAACAGGGCCAACATACCGCATTACCATAAGGGCCcaaa3′ (new, random sequence in capital letters and italic type; NheI and ApaI sites in bold type). The modified XbaI-SacI fragment with the random sequence was reintegrated with the SalI fragments from CarX and CarX-mCTd (which has a mutation in the distal GAGA factor binding site) and then placed in the transformation vector Car20T, creating the constructs CarX-TDR and CarX-mCTd-TDR, respectively.
A modified XbaI-SacI fragment without the random sequence inserted was also reintroduced in the transgenic construct CarX-mCTd-ΔCT, creating the construct CarX-mCTd-ΔCT-ΔTA with a mutation in the distal GAGA factor binding site, deletion of the proximal GAGA factor binding site, and a 67-bp deletion in the region of TFIID binding. The final sequence of the above region is 5′cgggCTAGCa agcag/gcgca aaagcaaGGG CCcaaa3′ (nucleotides remaining from the wild-type hsp26 promoter are in lowercase letters,/indicates the junction of the original hsp26 fragments, and NheI and ApaI recognition sites are in bold type).
In addition, we have made a new mutation changing the two first bases in the TATA box (hsp26mTATA) for in vitro studies of TFIID binding. In this mutation, the wild-type sequence 5′tccagcgggtataaaagcagcgtcgc3′ (from positions −44 to −19) is replaced with the sequence 5′tccagcAGTACtaaaagcagcgtcgc3′, introducing a ScaI recognition site (TATA box remnants are underlined; new sequence is in capital letters, with the ScaI site in bold type).
In vitro binding of the TFIID complex.
To assess binding of the TFIID complex to the hsp26 gene variant promoters in vitro, we generated PCR fragments from the transformation plasmids described above containing the wild-type (hsp26X) or mutant versions of the TATA box region (hsp26TDR, hsp26mTATA, and hsp26ΔTA). In all cases the left primer was the same, but the right primer was chosen to yield fragments of different length for each sequence studied. These PCR fragments were end labeled, mixed, and incubated with TFIID from Drosophila nuclear extract immobilized on protein G-Sepharose beads with an antibody against the TAFII230 subunit of the TFIID complex, as previously described (51). Purified DNA from immunoprecipitated material was analyzed on a denaturing polyacrylamide gel. A phosphorimager and NIH Image software were used to quantitatively assess the relative amount of DNA in each of the bands. The fractions of bound DNA to input DNA in the control lanes (mock precipitation) were subtracted from the fractions of bound DNA to input DNA in the experimental lanes (beads treated with the anti-TFIID antibodies).
Fly strains and germ line transformation.
Transformed lines (designated CarX-TDR, CarX-mCTd-TDR, CarX-mCTd-ΔCT-ΔTA, cP-ΔD1, cP-ΔCT-ΔD1, cP-ΔD2, and cP-ΔCT-ΔD2) were obtained using P elements as described previously (25, 40). The presence of a single copy of the P element and the integrity of the transgenes were verified by Southern hybridization using both ry sequences and the lacZ gene as probes (data not shown). Either the transformed lines were made homozygous or the transgenes were maintained over an appropriate balancer chromosome. In some cases, to increase the number of lines with inserts in different locations, the original transgene was mobilized in genetic crosses using the stock w; PΔ2-3/TM6, Tb e (see description at http://flybase.bio.indiana.edu). Three to seven independent lines were obtained for all constructs described in this work.
Transgenic flies containing the intact hsp70 promoter (line WT D7) and the hsp70 promoter with a 15-bp deletion encompassing the TATA box (line ΔTATA C) were described by Wu et al. (51).
Analysis of inducible expression.
Third-instar larvae were heat shocked for 90 min at 37°C. β-Galactosidase activity was measured as described by Lu et al. (32) using the CPRG (chlorophenol red/β-d-galactopyranoside) assay.
Analysis of XbaI and DNase I cleavage in chromatin from larval nuclei.
Experiments were performed as previously described (46). Briefly, nuclei were isolated from 1 g of flash-frozen third-instar larvae from each strain studied and treated with increasing concentrations of XbaI or DNase I. DNA from treated nuclei was purified, cut with restriction enzymes appropriate for the indirect end-labeling procedure (EcoRV and SmaI), electrophoresed in 1% agarose, transferred to a nylon filter (Roche Diagnostics, Indianapolis, Ind.) and hybridized with a digoxigenin-labeled probe (Roche Diagnostics). The probe was a 1.1-kb fragment of the lacZ gene (downstream from the hsp26 promoter). This provides qualitative assessment (with DNase I) and quantitative assessment (with XbaI) of the accessibility to nucleases at the proximal DH site. X-ray films were scanned, and bands were quantitatively scored on a Fluoro S MultiImager documentation system with QuantityOne software (Bio-Rad).
The accessibility of the proximal site to digestion by XbaI was calculated as the ratio of the measured intensity of signal in the proximal band to the sum of intensities of signal from the proximal, distal, and parental bands (32). Three to six lines with each transgenic construct were studied; the experiments were repeated at least three times with each line. Variation in the measurement of XbaI cleavage is approximately ±5%. To simplify the comparison of relative values of XbaI cleavage between different constructs, values for transgenes shown in Fig. 1 and 6 (XbaI sensitivity values) have been normalized to those of CarX and cP-351, respectively.
Immunoprecipitation of cross-linked chromatin (ChIP).
ChIP analyses of hsp26 transgenes were performed using 0- to 12-h-old embryos as described by Cavalli et al. (8) with minor modifications. Polyclonal rabbit anti-GAGA or anti-TBP antibodies (2 μl of each per 500 μl of chromatin) were used. The GAGA factor used to immunize rabbits was that expressed from the plasmid pAR-GAGA and purified as described previously (42), with the following modifications. Fractions from the heparin-Affigel column were diluted to a final concentration of 0.2 M HEMGN (HEPES-EDTA-magnesium-glycerol-Nonidet p-40). This material was loaded onto a Mono S column, washed with 3 column volumes of 0.15 M HEMGN, and eluted with a 0.15 to 1 M HEMGN gradient. Fractions containing GAGA factor were collected. Anti-GAGA factor serum was obtained by immunizing rabbits with the recombinant GAGA factor (service provided by the Pennsylvania State University Animal Facility). A polyclonal antiserum against Drosophila TBP was raised in rabbits as previously described (13). Antibodies were evaluated and shown to have the desired specificity by immunostaining of polytene chromosomes and by Western blotting (data not shown).
For each chromatin preparation, mock precipitations without antibodies were performed; the amount of DNA recovered in mock precipitations was usually 5- to 10-fold less than that from precipitation with antibodies. To visualize the products, we employed two different approaches, with comparable results. In both cases (see Fig. 4A), the immunoprecipitated DNA was first amplified using primers P1 and P2 (30 cycles [1 cycle consisting of 1 min at 95°C, 1 min at 55°C, and 1 min at 72°C], followed by 10 min at 72°C). To achieve quantitative PCR results following chromatin immunoprecipitation, a series of PCR amplifications was conducted using serially diluted input DNA, and all immunoprecipitated samples were assayed in a range where the PCR products showed a strict, linear correlation to input DNA. In the first method, the resulting PCR products were digested with restriction enzyme DraI, electrophoresed in a 15% polyacrylamide gel in 1× Tris-borate-EDTA (TBE) buffer for 3 to 4 h at 200 V, and electroblotted for 2 h in 0.5× TBE buffer at 40 V at 4°C to a nylon filter. The filter was hybridized with a 32P-labeled PCR-generated fragment (made with the same primers [P1 and P2]) from a cloned genomic fragment of the wild-type hsp26 gene (see Fig. 4A) (plasmid p88B13 [10]), and exposed using X-ray film and/or a Bio-Rad phosphorimager screen. QuantityOne software was used for quantitation of the data. The length of the transgene fragment is 21 bp shorter than the length of the endogenous gene fragment due to the loss of a 21-bp XbaI fragment during the original cloning. The lengths of the endogenous and transgenic DraI restriction fragments are 142 and 121 bp, respectively. This allows one to measure hybridization signals for the transgene and the endogenous gene in the same lane. Results were calculated as (Tab/Eab)/(Tm/Em), where T is the length of the transgene fragment, E is the length of the endogenous fragment, and the ab and m superscripts indicate that the samples were precipitated with antibodies or without antibodies (mock), respectively. This calculation assumes that if the transgene with a given mutation immunoprecipitates without any difference from the wild-type transgene, then the T/E ratio will be the same following immunoprecipitation as it is in the mock precipitation. Thus, Tab/Eab = Tm/Em where Eab is expected to be constant for all flies. Deviations from this equality should be attributable to the differences in protein binding to the transgene. PCR amplification of DNA recovered from three to seven independent preparations of cross-linked chromatin was repeated a total of 6 to 16 times for each case.
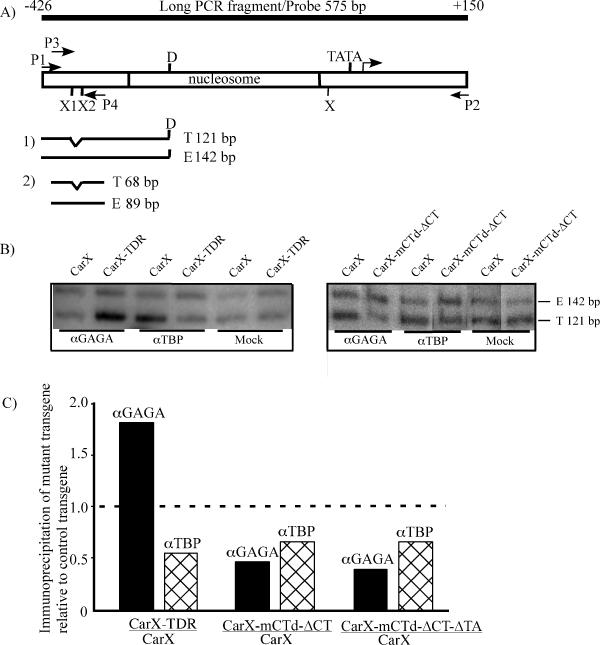
FIG. 4.
Binding of GAGA factor and TBP to different variants of the hsp26 promoter as shown by chromatin immunoprecipitation. (A) The general approach used to visualize and quantify the immunoprecipitated chromatin fragments is diagrammed. Immunoprecipitated DNA was amplified by PCR with primers P1 and P2 and digested with restriction enzyme DraI (D), followed by polyacrylamide gel electrophoresis and hybridization with the PCR-generated wild-type 32P-labeled probe shown in the top line (the anticipated fragments shown in panel 1). In some experiments, immunoprecipitated material was first amplified using the P1 and P2 primers, followed by a second round of amplification with primers P3 and P4 (the anticipated fragments are shown in panel 2). The lengths (in base pairs) of the anticipated fragments of the transgene (T) and endogenous gene (E) are indicated. The numbers above the top line are the left and right map positions of the probe. X, X1, and X2 are XbaI sites in the endogenous hsp26 promoter; all fall within the DH sites. (B) Representative examples of the chromatin immunoprecipitation data. (Left) Comparison between the control CarX line and the line with random replacement of sequences from positions −35 to +60, CarX-TDR. (Right) Comparison between the control line CarX and the line with two mutations in GAGA factor binding sites, CarX-mCTd-ΔCT. Note that the amount of DNA recovered in mock precipitations is 5- to 10-fold less than that from precipitations with antibodies; however, equal amounts of DNA have been used here to establish the ratio of transgene to endogenous gene in the precipitate. The positions of the 142-bp DraI fragment from the endogenous gene (E) and the 121-bp DraI fragment from the transgene (T) are shown to the right. αGAGA, anti-GAGA antibody; αTBP, anti-TBP antibody. (C) Comparison of immunoprecipitation of the hsp26 promoter chromatin fragments from the control (CarX) and mutant hsp26 transgenes. The data presented were obtained by the first approach described above. The results are shown as the value (normalized T/E) for the mutant transgene divided by the value (normalized T/E) for the control (wild-type) transgene (Tables 1 and 2). The broken line indicates the theoretically expected value (1.0) if the promoter mutations do not affect GAGA factor or TBP binding. See Materials and Methods for details.
In the second approach, the long PCR product (without restriction enzyme digestion) was subjected to a second round of PCR amplification using primers P3 and P4 (see Fig. 4A) in the presence of 32P-labeled dATP and dCTP. The conditions of PCR amplification were the same, except that linear amplification was observed with 20 cycles. The labeled products were similarly separated by electrophoresis in a polyacrylamide gel and recorded using X-ray film. The sizes of the endogenous and transgenic fragments were in this case 89 and 68 bp, respectively. Results were calculated as described above.
To analyze the interaction of GAGA factor with the wild-type hsp70 promoter compared to the hsp70 promoter with a deletion in the TATA box, chromatin immunoprecipitation experiments were performed using isolated, intact salivary glands of third-instar larvae as a source of material. Salivary glands (15 pairs per sample) were dissected from WT D7 or ΔTATA C transgenic larvae in Ringer solution (130 mM NaCl, 5 mM KCl, 1.5 mM CaCl2) and cross-linked with 1% formaldehyde for 10 min at room temperature. To stop the cross-linking reaction, 1 M glycine was added to a final concentration of 125 mM and the glands were incubated at room temperature for 5 min. The glands were then washed twice with 1 ml of Tris-buffered saline (140 mM NaCl, 20 mM Tris-Cl [pH 7.5]) and suspended in 200 μl of sodium dodecyl sulfate lysis buffer (1% sodium dodecyl sulfate, 10 mM EDTA, 50 mM Tris-Cl [pH 8]). The suspension was then sonicated in four 15-s pulses at output level 1.6 with a Branson Sonifier 450 instrument (VWR Scientific). Subsequent treatment with antibody plus protein A beads and recovery of immunoprecipitated DNA followed the ChIP protocol from Upstate Biotechnology, except that whole Sepharose beads were subjected to proteinase K digestion to release the DNA from the immunoprecipitate. Multiplex PCRs were performed with the purified DNA as template using two pairs of primers to amplify the fragments of the hsp70 promoter and of the white gene.
RESULTS
TFIID does not bind the random sequence in vitro.
The main goal of this study is to assess the possible role of the TFIID complex in formation of the organized chromatin structure observed at the promoter of the hsp26 gene prior to activation by heat shock. To address this question, we have generated transgenic constructs (CarX-TDR and CarX-mCTd-TDR) with the region from positions −35 to +60 replaced with a random sequence. We anticipated that none of the TFIID subunits would bind to this sequence. The TFIID complex is bound at the promoters of the hsp26 and hsp70 genes prior to heat shock, occupying primarily sequences from positions −35 to around +30 (13, 19, 38). DNase I footprints of some preparations of the TFIID complex extend to position +47 (41). In vitro binding experiments confirm that TFIID does not bind to a PCR-generated fragment of the construct with the random replacement sequence (hsp26TDR) or to PCR fragments generated with a deletion of this region at the hsp26 gene promoter (hsp26ΔTA) (Fig. 2). To assess the sensitivity of this in vitro approach, we analyzed binding of TFIID to a PCR fragment with a point mutation (changing the TATA box from TATAAAA to ACTAAAA). TFIID binding is significantly reduced but not abolished in this case (hsp26mTA; 29% binding in comparison with the wild-type construct). This agrees very well with previous data for different point mutations in the TATA box (33). Thus, the constructs prepared can serve the desired purpose of reducing the association of TFIID with the promoter DNA.
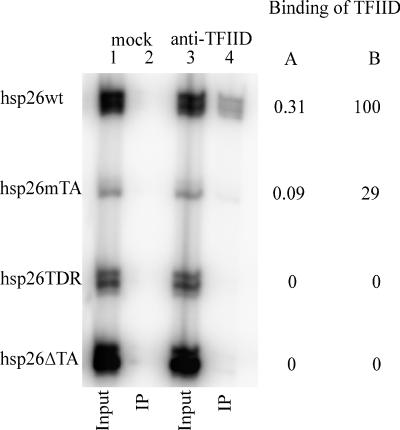
FIG. 2.
In vitro binding of the TFIID complex to hsp26 promoter DNA fragments. A mixture of 32P-labeled PCR-generated fragments containing the wild-type hsp26 (hsp26wt) promoter and promoter mutants (hsp26mTA, hsp26TDR, and hsp26ΔTA) was incubated with TFIID that had been immobilized on protein G-Sepharose with an antibody against TAFII230. As a negative control (mock precipitation), an equivalent mixture of 32P-labeled DNA fragments was incubated with protein G-Sepharose that had been incubated with Drosophila nuclear extract in the absence of the TAFII230 monoclonal antibody. Bound (IP) and input DNA were analyzed on a sequencing gel. To the right of the gel, the fraction of each fragment recovered in the bound fraction calculated as described in Materials and Methods is shown in column A, and binding as a percentage of binding for the hsp26wt fragment is shown in column B.
The hsp26 promoter is not heat shock inducible when the TFIID binding region is replaced by the random sequence.
The heat shock inducibility of various mutant constructs was measured by analysis of β-galactosidase activity following heat shock; several independent transgenic lines were assayed for each of the constructs. When all TFIID-interacting sequences were replaced by the random sequence (constructs CarX-TDR and CarX-mCTd-TDR), gene activity was found to be less than 2% of the control wild-type transgene (Fig. 1, HS β-gal column). Thus, in accordance with previous data for a point mutation in the TATA box, which results in 3% activity (33), loss of the TATA box and downstream sequences is very detrimental for heat shock inducibility of the hsp26 gene. Elimination of the GAGA sequences (by mutation of the distal site and deletion of the proximal site) also reduces transcription to very low levels, as previously reported (32) (Fig. 1).
Generation of accessible DH sites is drastically curtailed in flies where the TFIID binding region is replaced by the random sequence.
The chromatin structure of the uninduced hsp26 promoter, as shown by maintenance of the proximal DH site, was assessed for the various transgenes by digestion of isolated larval nuclei with XbaI. XbaI sites are located within both the proximal and distal DH sites of the hsp26 promoter (Fig. 1A), and cleavage with excess restriction enzyme correlates very well with relative sensitivity to DNase I. The extent of XbaI cleavage was determined using the indirect end-labeling protocol (see Materials and Methods). Sample lumigraphs are shown in Fig. 3A, and the quantitative results for each of the constructs studied are given in Fig. 1 (XbaI sensitivity column), normalized to the corresponding control line, CarX. It is evident that replacement of the region from positions −35 to +60 with random sequence (CarX-TDR) results in a dramatic loss in accessibility at the proximal DH site (22% cleavage in comparison to control line CarX). The presence of an additional mutation in the distal GAGA factor binding site (CarX-mCTd-TDR) decreased the cleavage to 6%, indicating an additional effect. However, in the presence of mutation or deletion of the GAGA sites, no further loss in accessibility can be detected on loss of the TFIID binding site (compare CarX-mCTd-ΔCT-ΔTA to CarX-mCTd-ΔCT), suggesting that the GAGA factor plays an upstream role.
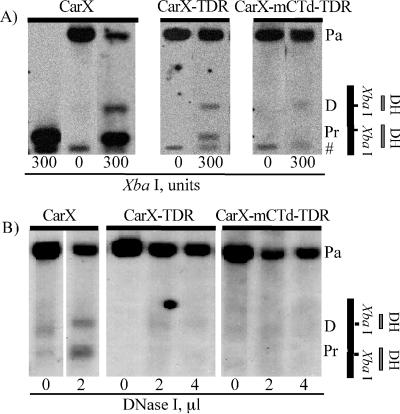
FIG. 3.
Cleavage by XbaI or DNase I in the proximal DH site is reduced in constructs CarX-TDR and CarX-mCTd-TDR. (A) Lumigraphs of DNA from nuclei digested with XbaI, analyzed by the indirect end-labeling procedure using the lacZ probe. Numbers below the lanes show the amount of enzyme used; results are shown for one representative transgenic line carrying the CarX-TDR construct and one carrying the CarX-mCTd-TDR construct. The positions of the parental band (which is created by EcoRV digestion of DNA not cleaved in nuclei by XbaI) (Pa), the distal DH site (D), the proximal DH site (Pr), and an unrelated band (#) are indicated to the right. The far left lane shows results from digestion of purified DNA, rather than chromatin. Quantitative results for all lines studied are presented in Fig. 1 in the XbaI sensitivity column. (B) Lumigraphs of the parallel experiment with limited digestion of nuclei with DNase I. In panels A and B, a map with the positions of the DH sites and XbaI sites of the hsp26 promoter is shown at the extreme right.
Similar results were obtained on partial digestion by DNase I (Fig. 3B); no significant signal could be detected in the position of the expected band in samples from transgenic flies with the constructs having a random replacement of sequences from positions −35 to +60. Thus, the loss of the TFIID binding site has a dramatic effect on the formation of proper chromatin structure in the core promoter region of the hsp26 gene. Coupled with earlier results from the analysis of GAGA factor sites, this suggests that both GAGA factor and TFIID are critical for creation of fully functional DH sites. However, it is unclear whether GAGA factor fails to bind its target sequences in the absence of TBP, whether their binding is mutually dependent, or whether the block to DH site formation lies downstream of successful binding of either component.
Chromatin immunoprecipitation experiments indicate that GAGA factor can bind its target sequences in the absence of TFIID and facilitates binding of TFIID at the hsp26 promoter.
We have performed immunoprecipitation experiments using chromatin from the transgenic lines described with different alterations in the hsp26 promoter region. Chromatin from 0- to 12-h-old embryos was cross-linked with formaldehyde, fragmented by sonication, purified in a CsCl gradient, and precipitated with antibodies to GAGA factor or to TBP. Immunoprecipitated chromatin was isolated by binding to protein A-Sepharose. The following genotypes were examined: a line with the control CarX construct, a line with the random replacement of the sequence from positions −35 to +60 (CarX-TDR), a line with a mutation in the distal GAGA factor binding site and deletion of the proximal GAGA factor binding site (CarX-mCTd-ΔCT), and a similar line with the additional deletion in the TFIID binding region (CarX-mCTd-ΔCT-ΔTA) (Fig. 1B and Materials and Methods). In each case, immunoprecipitation of the transgene was compared to immunoprecipitation of the endogenous hsp26 gene in the same sample. The results are presented in Tables 1 and 2, as the T/E ratio (transgene/endogenous gene) normalized to the same ratio from mock (no-antibody) control experiments, (Tab/Eab)/(Tm/Em), and in Fig. 4C as the ratio of this value in flies with the mutant transgene to the same value in flies with the control transgene.
TABLE 1.
Quantitative analysis of immunoprecipitation of cross-linked chromatin of CarX and CarX-TDR transgenes
| Antibody | Methoda | Signal intensity ratiob
|
|
|---|---|---|---|
| CarX (n) | CarX-TDR (n) | ||
| Anti-GAGA factor antibodies | 1 | 1.24 ± 0.33 (16) | 2.24 ± 0.55 (16) |
| 2 | 0.84 ± 0.14 (5) | 1.26 ± 0.17 (5) | |
| Anti-TBP antibodies | 1 | 1.99 ± 0.27 (16) | 1.12 ± 0.14 (16) |
| 2 | 1.27 ± 0.07 (5) | 0.78 ± 0.12 (5) | |
Method 1, DraI digestion of the long PCR product; method 2, a second round of PCR amplification (see the legend to Fig. 4 and Materials and Methods).
Mean ± standard error of the mean of the ratio of intensity of signal with the fragment of transgene to the fragment of endogenous gene in preparations treated by antibodies and normalized to the same ratio from the mock experiment (see Materials and Methods). The number of experiments for each transgene (n) is shown in parentheses.
The analysis shows that a random replacement of the sequence from positions −35 to +60 in CarX-TDR decreases TBP binding approximately twofold in comparison with the control CarX transgene, as shown in the example (Fig. 4B). At the same time, GAGA factor binding is preserved in the CarX-TDR hsp26 gene promoter. This indicates that GAGA factor does not depend on TBP (or the TFIID complex) in binding to its target sequences. In fact, GAGA factor binding is apparently increased at this transgene. Mutations in both distal and proximal GAGA factor sites (transgene CarX-mCTd-ΔCT) decrease GAGA factor binding approximately twofold (Fig. 4B). The binding of TBP is also significantly reduced on this transgene, indicating that GAGA factor is required for efficient TBP binding. The change observed in TBP binding, compared with the wild-type control transgene, is probably the limit of what can be detected with this method; flies with a transgene carrying two mutations in GAGA factor binding sites and a deletion in the TFIID binding region (CarX-mCTd-ΔCT-ΔTA), while showing a significant loss of GAGA factor binding, show a similar loss in TBP in comparison with the control CarX transgene. The results for this transgene are very similar to the results for the transgene with mutations only in GAGA factor binding sites (Fig. 4C).
Deletion of the TATA box in the hsp70 gene promoter does not affect binding of GAGA factor.
To demonstrate that GAGA factor binding in the absence of TBP is not a feature specific to the hsp26 promoter, we have performed chromatin cross-linking immunoprecipitation experiments with lines carrying a deletion of the TATA box at the hsp70 promoter using antibodies to GAGA factor. In vitro binding studies have shown that this mutation reduces the binding of TFIID by at least 20-fold (13). DNase I hypersensitivity was also decreased in transgenic flies with such a construct, although the association of GAGA factor was still evident in genomic footprinting analysis (47). Chromatin immunoprecipitation performed on isolated, intact salivary glands (Fig. 5) shows that deletion of the TATA box does not significantly impair the ability of GAGA factor to bind to its target sequences. We conclude that both heat shock genes behave similarly in this respect.
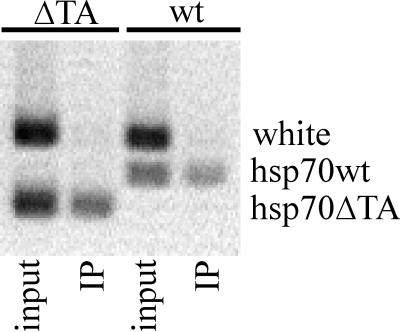
FIG. 5.
Binding of GAGA factor to wild-type (wt) and mutated hsp70 (ΔTA) promoters as shown by chromatin immunoprecipitation experiments. Quantitative PCR was performed with primers for the wild-type hsp70 promoter (hsp70wt) or the hsp70 promoter with a deletion of the TATA box (hsp70ΔTA), as well as with primers for the white gene promoter (control for specificity), followed by gel electrophoresis. GAGA factor is clearly able to interact with both the wild-type and mutant transgenes. Lanes: input, total DNA from flies; IP, DNA recovered after immunoprecipitation with anti-GAGA antibody. See Materials and Methods for details.
Other components of TFIID and/or the preinitiation complex may cooperate with GAGA factor in the formation of DH sites at the hsp26 promoter.
Binding of TBP to DNA facilitates multiple DNA contacts by other subunits of the TFIID complex. The hsp26 gene has all of the described sequences utilized in the initiation of transcription: a TATA box (from positions −35 to −30), Initiator Element (Inr; from positions−2 to +4) and Downstream Promoter Element (DPE; from positions +27 to +32) (27). The Drosophila Initiator sequence interacts with the TAFII150-TAFII230 dimer (9, 51), and the DPE interacts with the TAFII60-TAFII40 heterotetramer (6). To elucidate further whether these and/or other members of the TFIID complex take part in formation of the DH sites, we have analyzed heat shock inducibility and nuclease accessibility using XbaI digestion of transgenic lines carrying constructs with deletions downstream from the TATA box. cP-ΔD1 has a deletion from positions +7 to +48 that encompasses the DPE, and cP-ΔD2 has a deletion from positions +48 to +137. We anticipated that only the cP-ΔD1 deletion would affect accessibility and expression, as no reports in the literature indicate components of the preinitiation complex that contact DNA downstream from position +48. The loss of heat shock inducibility was significant, albeit less severe, in the transgenic flies in which the TATA box and Inr were maintained, but downstream portions of the TFIID-interacting sequences, containing the DPE were deleted (cP-ΔD1) (47% activity) (Fig. 6). [Note that the DPE consensus sequence is (A/G/T)(C/G)(A/T)(C/T)(A/C/G)(C/T); the hsp26 DPE is ACACCT, a complete match (27).] The smaller effect of a mutation in the downstream TFIID-interacting sequences is in accord with previous observations using the hsp70 promoter (51). Interestingly, deletion of sequences downstream of the known TFIID contact points, the ΔD2 mutation, also shows a significant decrease in heat shock inducibility (57% activity), indicating that DNA sequences well downstream from the DPE may be important for proper activation of hsp26.
XbaI cleavage is decreased to 51% (Fig. 6) using nuclei from larvae with the ΔD1 deletion, approximately to the same extent as in flies with point mutations in the TATA box (64%) (33). In transgenic lines with the combination of this deletion and a deletion in the proximal GAGA factor binding site, accessibility to nuclease was further decreased (51% versus 16% for cP-ΔD1 versus cP-ΔCT-ΔD1 [Fig. 6]). As is the case for CarX-TDR, this suggests cooperation between GAGA factor and members of the TFIID complex, impacted by subunits in addition to TBP, in the formation of proper chromatin structure at this promoter. The distal ΔD2 mutation has less impact than the more proximal ΔD1 mutation (XbaI cleavage is 70% [Fig. 6]). The combination of deletion ΔD2 and a deletion in the proximal GAGA factor binding site also leads to a further decrease in accessibility (70% versus 34% for cP-ΔD2 versus cP-ΔCT-ΔD2 [Fig. 6]). Considering that variation in the measurement of XbaI cleavage is approximately ±5%, this change in accessibility appears significant. The data for the ΔD2 mutation suggest that cooperativity between GAGA factor and unknown protein(s), interacting well downstream from the DPE, may be important for both chromatin structure formation and proper function of the gene, as the loss in accessibility correlates with a decrease in heat shock inducibility for this mutant (Fig. 6).
DISCUSSION
We have shown previously that mutations in the GAGA factor binding sites at the hsp26 promoter lead to a significant decrease in accessibility at the DH sites, with a concomitant failure in heat shock inducibility of the gene (32). Failure to establish the appropriate chromatin structure may indicate an absence of critical components, may limit the access of HSF after heat shock induction to its target sequences, and/or may block a downstream step in the transcription activation process. Changing the sequence of the TATA box from TATAAAA to CCCAAAA abolished heat shock inducibility but caused only partial loss in accessibility at the DH sites (33). In the latter case, however, TFIID was still capable, albeit less effectively, of binding to its target sequence in vitro. To clarify further the mechanics of chromatin structure organization, we have fully replaced the region from positions −35 to + 60 (encompassing the sequences which, according to existing data, interact with TBP and other subunits of the TFIID complex) with a random sequence. In vitro binding experiments confirmed that the latter sequence does not bind TFIID (tested with antibodies to TAFII230). Accordingly, this modified hsp26 promoter shows no heat shock inducibility in vivo. The sensitivity of the DH sites of this transgene to XbaI and DNase I cleavage has fallen approximately fivefold from that of the wild-type transgene; this is diminished even further in the presence of a mutation in the distal GAGA factor binding site. This strongly confirms the view that both proteins (complexes) participate and collaborate in the formation of the specialized chromatin structure of this promoter. Analysis of a downstream deletion from positions +7 to +48 has shown that this deletion has a similar impact on the chromatin structure as a point mutation in the TATA box (30 to 50% decrease in XbaI cleavage). This suggests that other subunits of the TFIID complex may also affect the chromatin structure at the hsp26 promoter. Surprisingly, analysis of a downstream deletion from positions +47 to +137 has shown that this deletion also has some impact on XbaI cleavage and heat shock inducibility, indicating the existence of additional interactions important for gene activity and the organization of the specific chromatin structure well downstream from known regulatory sequences.
These data alone do not identify the initial step in the construction of chromatin at the hsp26 promoter. Can the proteins bind to their target sequences independently from each other, or does one recruit another to build the corresponding chromatin structure? To address this question, we have performed immunoprecipitation experiments with formaldehyde cross-linked embryo chromatin. The results clearly show that the amount of GAGA factor is not decreased in flies with the random replacement of sequences from positions −35 to +60 at the hsp26 promoter (construct CarX-TDR [Table 1 and Fig. 4C]). GAGA factor can apparently bind its target sequences irrespective of depletion of TFIID. In contrast, TBP binding was diminished approximately twofold when GAGA factor binding was decreased (construct CarX-mCTd-ΔCT [Table 2]). We may conclude that GAGA factor binds to (CT)n repeats independently of TBP binding and facilitates binding of the latter directly or indirectly. Recruitment of TBP by GAGA factor is likely to be indirect, as Mason and Lis (34) did not observe any direct interaction of GAGA factor with TBP in pull-down experiments. Some other protein (or protein complex) could serve a bridging function. Alternatively, the presence of GAGA factor might trigger a chromatin modification that facilitates TFIID binding.
TABLE 2.
Quantitative analysis of immunoprecipitation of cross-linked chromatin of CarX, CarX-mCTd-ΔCT, and CarX-mCTd-ΔCT-ΔTA transgenes
| Antibody | Methoda | Signal intensity ratiob
|
||
|---|---|---|---|---|
| CarX (n) | CarX-mCTd-ΔCT (n) | CarX-mCTd-ΔCT-ΔTA (n) | ||
| Anti-GAGA factor antibodies | 1 | 1.21 ± 0.14 (9) | 0.57 ± 0.12 (6) | 0.49 ± 0.06 (9) |
| Anti-TBP antibodies | 1 | 1.36 ± 0.10 (9) | 0.91 ± 0.19 (8) | 0.90 ± 0.23 (8) |
It is interesting that GAGA factor binding appears to increase when TBP binding is decreased (using the TDR construct; Fig. 4C). This result may simply reflect an increase in accessibility of antigenic determinants in GAGA factor molecules in the absence of the bulky TFIID complex. Alternatively, the result suggests that in the wild-type promoter the binding of GAGA factor is limited by the TBP-based complex. In addition to the well-characterized larger (GA)n sites, GAGA factor can bind in vitro to GAG triplets (49). GAGA factor forms oligomeric complexes through its N-terminal and C-terminal domains (BTB/POZ and Q domains, respectively) (2, 4, 14, 26, 50). An increase of GAGA factor concentration in vitro has led to a significant expansion of its DNase I footprints on several natural promoters, possibly through formation of multimeric complexes and additional interactions with scattered GAG triplets (26). Optimization of the level of GAGA factor binding at a given promoter may be functionally important. The expansion of GAGA factor binding along the promoter might displace nucleosomes, given the ability of GAGA factor to bend DNA (26). Alternatively, the enrichment of GAGA factor in the region above a certain level might change the amount and/or species of proteins interacting with GAGA factor through its BTB/POZ and Q domains. However, the increased presence of GAGA factor in the CarX-TDR lines is insufficient to generate a normal DH site in the absence of TFIID binding.
GAGA factor is encoded by the Trl gene (16), a member of the trithorax group, and acts in several systems as a transcriptional activator (for reviews, see references 21 and 48; for recent experiments, see references 14, 36, 37, and 44). However, GAGA factor has also been shown to interact with dSAP18, a member of the dSin3 complex having histone deacetylase activity (15). GAGA factor has been found in complexes with Polycomb group proteins, involved in gene silencing (23), and has been shown to bind to two Polycomb-responsive elements in homeotic loci (7, 35). The involvement of GAGA factor in regulatory regions for both activation and repression of genes suggests a general role, perhaps involving nucleosome displacement at the regulatory site, with the regulatory outcome depending on the context and other participating proteins (for a review, see reference 18).
The new data allow more informed speculation on the order of events in creation of the chromatin structure at the hsp26 promoter (Fig. 7). It appears that nucleosomes impede GAGA factor interaction with its target sequences as in vitro (42, 43) as well as in vivo (11). Thus, GAGA factor binding to its target sequences must be facilitated by one or another chromatin remodeling complex, likely NURF (42, 43). Recently, Xiao and coworkers (53) have shown in pull-down experiments that the largest subunit of NURF, NURF301, specifically and stoichiometrically interacts with GAGA factor as well as with nucleosomes. The interaction of a chromatin remodeling complex with a nucleosome may destabilize it and increase the probability of GAGA factor binding to its target sequence. The binding of GAGA factor may stimulate DNA bending (26), leading to sliding or displacement of nucleosomes (42, 53). This alone appears to create a partially opened chromatin structure with a modestly developed DH site (20% XbaI accessibility observed for CarX-TDR). In the wild-type gene, this might allow the binding of TBP to the TATA box, further enhancing DNA bending and nucleosome sliding (31; for a review, see reference 37). Other subunits of the TFIID complex could then contact DNA sequences downstream from the TATA box (in the case of the hsp26 or hsp70 genes, including Inr and DPE). Histone acetyltransferase activity of the TAFII230 subunit of TFIID (3, 28) might further facilitate opening up the chromatin structure. Histone modification is widely observed to play a key role in determining the activation state (24). The sum of the interactions must, in the end, stabilize at least three nucleosomes, manifested as the two DH sites with a strictly positioned nucleosome between them, observed in vivo.
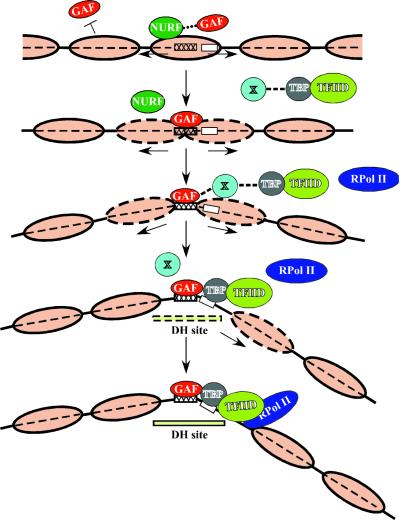
FIG. 7.
Proposed model of events leading to formation of proper chromatin structure at the hsp26 promoter. Nucleosomes are represented by large light brown ovals, with a dashed outline indicating their increased instability or movement. The GAGA factor (GAF) (red), chromatin remodeling complex (43) (NURF) (green), TATA binding protein (TBP) (gray), TFIID complex (yellow-green), RNA polymerase II (RPol II) (blue), unknown protein or protein complex (X) (light cyan), GAGA factor binding site (cross-hatched box), TATA box (white box), and DH site (light yellow box) (the broken outline indicating intermediate sensitivity to nucleases) are shown. See text for details. For simplicity, only one GAGA factor site is indicated. While X is suggested here to serve a bridging function, it may also indicate a process, such as histone modification, that facilitates TFIID binding.
Our data indicate that the TFIID complex plays an important role in formation of chromatin structure in cooperation with other proteins. The binding of TBP to a TATA box is not an obligatory step to attract other subunits of the TFIID complex to a promoter region. More than half of all core promoters have no TATA box (27); nevertheless, TBP is still important for transcription of many core promoters. TBP may bind to sequences other than the TATA box or may bind as part of a protein complex. Many TATA-less promoters are dependent on other proteins which appear to substitute for TBP (for example, TRF1 and TRF2) (3, 45). However, it appears that in all such cases the formation of a full or partial TFIID complex is still necessary. Several different TFIID complexes, which vary in their set of subunits, and some other complexes, performing coactivator functions and containing some of the TFIID subunits, are found in diverse experimental systems (for reviews, see references 3, 5, and 22). Many of these coactivator complexes contain histone acetyltransferase activities. The TFIID complex (or its subunits) appears likely to participate in various processes of chromatin modification and may be attracted to promoters not only by TBP binding but by other interactions.
In sum, our study indicates that both GAGA factor and the TFIID complex (or its subunits) play a significant role in the formation of the specific chromatin structure at the hsp26 promoter. The results obtained using the altered hsp26 regulatory regions we have constructed suggest a sequential process initiated by GAGA factor binding. It will be interesting to explore the modifications of histones that occur at the different steps in the reorganization of chromatin on the hsp26 promoter defined by these mutations.
Acknowledgments
We thank the members of the Elgin lab for helpful discussions.
This work was supported in part by the Public Health Service NIH grants GM 31532 to S.C.R.E. and GM7477 to D.S.G.
REFERENCES
- 1.Aalfs, J. D., and R. E. Kingston. 2000. What does “chromatin remodeling” mean? Trends Biochem. Sci. 25:548-555. [DOI] [PubMed] [Google Scholar]
- 2.Agianian, B., K. Leonard, E. Bonte, H. Van der Zandt, P. B. Becker, and P. A. Tucker. 1999. The glutamine-rich domain of the Drosophila GAGA factor is necessary for amyloid fibre formation in vitro, but not for chromatin remodelling. J. Mol. Biol. 285:527-544. [DOI] [PubMed] [Google Scholar]
- 3.Albright, S. R., and R. Tjian. 2000. TAFs revisited: more data reveal new twists and confirm old ideas. Gene 242:1-13. [DOI] [PubMed] [Google Scholar]
- 4.Benyajati, C., L. Mueller, N. Xu, M. Pappano, J. Gao, M. Mosammaparast, D. Conklin, H. Granok, C. Craig, and S. Elgin. 1997. Multiple isoforms of GAGA factor, a critical component of chromatin structure. Nucleic Acids Res. 25:3345-3353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Brown, C. E., T. Lechner, L. Howe, and J. L. Workman. 2000. The many HATs of transcription coactivators. Trends Biochem. Sci. 25:15-19. [DOI] [PubMed] [Google Scholar]
- 6.Burke, T. W., and J. T. Kadonaga. 1997. The downstream core promoter element, DPE, is conserved from Drosophila to humans and is recognized by TAFII60 of Drosophila. Genes Dev. 11:3020-3031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Busturia, A., A. Lloyd, F. Bejarano, M. Zavortink, H. Xin, and S. Sakonju. 2001. The MCP silencer of the Drosophila Abd-B gene requires both Pleiohomeotic and GAGA factor for the maintenance of repression. Development 128:2163-2173. [DOI] [PubMed] [Google Scholar]
- 8.Cavalli, G., V. Orlando, and R. Paro. 1999. Mapping DNA target sites of chromatin-associated proteins by formaldehyde cross-linking in Drosophila embryos, p. 21-37. In W. A. Bickmore (ed.), Chromosome structural analysis: a practical approach. Oxford University Press, Oxford, England.
- 9.Chalkley, G. E., and C. P. Verrijzer. 1999. DNA binding site selection by RNA polymerase II TAFs: a TAFII250-TAFII150 complex recognizes the Initiator. EMBO J. 18:4835-4845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Corces, V., R. Holmgren, R. Freund, R. Morimoto, and M. Meselson. 1980. Four heat shock proteins of Drosophila melanogaster coded within a 12-kilobase region in chromosome subdivision 67B. Proc. Natl. Acad. Sci. USA 77:5390-5393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cryderman, D. E., H. Tang, C. Bell, D. S. Gilmour, and L. L. Wallrath. 1999. Heterochromatic silencing of Drosophila heat shock genes acts at the level of promoter potentiation. Nucleic Acids Res. 27:3364-3370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Elgin, S. C. R. 1988. The formation and function of DNase I hypersensitive sites in the process of gene activation. J. Biol. Chem. 263:19259-19262. [PubMed] [Google Scholar]
- 13.Emanuel, P. A., and D. S. Gilmour. 1993. TFIID recognizes DNA sequences downstream of the TATA element in the hsp70 heat shock gene. Proc. Natl. Acad. Sci. USA 90:8449-8453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Espinás, M. L., E. Jimenez-Garcia, A. Vaquero, S. Canudas, J. Bernues, and F. Azorín. 1999. The N-terminal POZ domain of GAGA mediates the formation of oligomers that bind DNA with high affinity and specificity. J. Biol. Chem. 274:16461-16469. [DOI] [PubMed] [Google Scholar]
- 15.Espinás, M. L., S. Canudas, L. Fanti, S. Pimpinelli, J. Casanova, and F. Azorín. 2000. The GAGA factor of Drosophila interacts with SAP18, a Sin3-associated polypeptide. EMBO Rep. 1:253-259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Farkas, G., J. Gausz, M. Galloni, G. Reuter, H. Gyurkovics, and F. Karch. 1994. The Trithorax-like gene encodes the Drosophila GAGA factor. Nature 371:806-808. [DOI] [PubMed] [Google Scholar]
- 17.Farkas, G., B. A. Leibovitch, and S. C. R. Elgin. 2000. Chromatin organization and transcriptional control of gene expression in Drosophila. Gene 253:117-136. [DOI] [PubMed] [Google Scholar]
- 18.Francis, N. J., and R. E. Kingston. 2001. Mechanisms of transcriptional memory. Nat. Rev. Mol. Cell. Biol. 2:409-421. [DOI] [PubMed] [Google Scholar]
- 19.Gilmour, D. S., T. J. Dietz, and S. C. R. Elgin. 1988. TATA box-dependent protein-DNA interactions are detected on heat shock and histone gene promoters in nuclear extracts derived from Drosophila melanogaster embryos. Mol. Cell. Biol. 8:3204-3214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Glaser, R. L., G. H. Thomas, E. Siegfried, S. C. R. Elgin, and J. T. Lis. 1990. Optimal heat-induced expression of the Drosophila hsp26 gene requires a promoter sequence containing (CT)n•(GA)n repeats. J. Mol. Biol. 211:751-761. [DOI] [PubMed] [Google Scholar]
- 21.Granok, H., B. A. Leibovitch, C. D. Shaffer, and S. C. Elgin. 1995. Chromatin: Ga-ga over GAGA factor. Curr. Biol. 5:238-241. [DOI] [PubMed] [Google Scholar]
- 22.Green, M. R. 2000. TBP-associated factors (TAFIIs): multiple, selective transcriptional mediators in common complexes. Trends Biochem. Sci. 25:59-63. [DOI] [PubMed] [Google Scholar]
- 23.Horard, B., C. Tatout, S. Poux, and V. Pirrotta. 2000. Structure of a Polycomb response element and in vitro binding of Polycomb group complexes containing GAGA factor. Mol. Cell. Biol. 20:3187-3197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Jenuwein, T., and C. D. Allis. 2001. Translating the histone code. Science 293:1074-1080. [DOI] [PubMed] [Google Scholar]
- 25.Karess, R. E., and G. M. Rubin. 1984. Analysis of P transposable element functions in Drosophila. Cell 38:135-146. [DOI] [PubMed] [Google Scholar]
- 26.Katsani, K. R., M. A. Hajibagheri, and C. P. Verrijzer. 1999. Co-operative DNA binding by GAGA transcription factor requires the conserved BTB/POZ domain and reorganizes promoter topology. EMBO J. 18:698-708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Kutach, A. K., and J. T. Kadonaga. 2000. The downstream promoter element DPE appears to be widely used as the TATA box in Drosophila core promoters. Mol. Cell. Biol. 20:4754-4764. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Lee, T. I., and R. A. Young. 2000. Transcription of eukaryotic protein-coding genes. Annu. Rev. Genet. 34:77-137. [DOI] [PubMed] [Google Scholar]
- 29.Lis, J., and C. Wu. 1995. Promoter potentiation and activation: chromatin structure and transcriptional induction of heat shock genes, p. 71-88. In S. C. R. Elgin (ed.),Chromatin structure and gene expression. IRL Press, Oxford, England.
- 30.Lis, J. T. 1998. Promoter-associated pausing in promoter architecture and post-initiation transcriptional regulation. Cold Spring Harbor Symp. Quant. Biol. 53:347-356. [DOI] [PubMed] [Google Scholar]
- 31.Lomvardas, S., and D. Thanos. 2001. Nucleosome sliding via TBP DNA binding in vivo. Cell 106:685-696. [DOI] [PubMed] [Google Scholar]
- 32.Lu, Q., L. L. Wallrath, H. Granok, and S. C. R. Elgin. 1993. (CT)n•(GA)n repeats and heat shock elements have distinct roles in chromatin structure and transcriptional activation of the Drosophila hsp26 gene. Mol. Cell. Biol. 13:2802-2814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lu, Q., L. L. Wallrath, P. E. Emanuel, S. C. R. Elgin, and D. S. Gilmour. 1994. Insensitivity of the preset hsp26 chromatin structure to a TATA box mutation in Drosophila. J. Biol. Chem. 269:15906-15911. [PubMed] [Google Scholar]
- 34.Mason, P. B., and J. T. Lis. 1997. Cooperative and competitive protein interactions at the hsp70 promoter. J. Biol. Chem. 272:33227-33233. [DOI] [PubMed] [Google Scholar]
- 35.Mishra, R. K., J. Mihaly, S. Barges, A. Spierer, F. Karch, K. Hagstrom, S. E. Schweinsberg, and P. Schedl. 2001. The iab-7 Polycomb response element maps to a nucleosome-free region of chromatin and requires both GAGA and Pleiohomeotic for silencing activity. Mol. Cell. Biol. 21:1311-1318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Pile, L. A., and I. L. Cartwright. 2000. GAGA factor-dependent transcription and establishment of DNase hypersensitivity are independent and unrelated events in vivo. J. Biol. Chem. 275:1398-1404. [DOI] [PubMed] [Google Scholar]
- 37.Pugh, B. F. 2000. Control of gene expression through regulation of the TATA-binding protein. Gene 255:1-14. [DOI] [PubMed] [Google Scholar]
- 38.Purnell, P. A., P. A. Emanuel, and D. S. Gilmour. 1994. TFIID sequence-recognition of the initiator and sequences further downstream in Drosophila class II genes. Genes Dev. 8:830-842. [DOI] [PubMed] [Google Scholar]
- 39.Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 40.Spradling, A. C. 1986. P element-mediated transformation, p. 175-197. In D. B. Roberts (ed.), Drosophila: a practical approach. IRL Press, Oxford, England.
- 41.Sypes, M. A., and D. S. Gilmour. 1994. Protein/DNA crosslinking of a TFIID complex reveals novel interactions downstream of the transcription start. Nucleic Acids Res. 22:807-814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Tsukiyama, T., P. B. Becker, and C. Wu. 1994. ATP-dependent nucleosome disruption at a heat-shock promoter mediated by binding of GAGA transcription factor. Nature 367:525-532. [DOI] [PubMed] [Google Scholar]
- 43.Tsukiyama, T., and C. Wu. 1995. Purification and properties of an ATP-dependent nucleosome remodeling factor. Cell 83:1011-1020. [DOI] [PubMed] [Google Scholar]
- 44.Vaquero, A., M. L. Espinás, F. Azorín, and J. Bernues. 2000. Functional mapping of the GAGA factor assigns its transcriptional activity to the C-terminal glutamine-rich domain. J. Biol. Chem. 275:19461-19468. [DOI] [PubMed] [Google Scholar]
- 45.Verrijzer, C. P. 2001. Transcription factor IID-not so basal after all. Science 293:2010-2011. [DOI] [PubMed] [Google Scholar]
- 46.Wallrath, L. L., M. J. Swede, and S. C. R. Elgin. 1998. Mapping chromatin structure in Drosophila, p. 59-77. In H. Gould (ed.), Chromatin: a practical approach. Oxford University Press, Oxford, England.
- 47.Weber, J. A., D. J. Taxman, Q. Lu, and D. S. Gilmour. 1997. Molecular architecture of the hsp70 promoter after deletion of the TATA box or the upstream regulation region. Mol. Cell. Biol. 17:3799-3808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Wilkins, R. C., and J. T. Lis. 1997. Dynamics of potentiation and activation: GAGA factor and its role in heat shock gene regulation. Nucleic Acids Res. 25:3963-3968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Wilkins, R. C., and J. T. Lis. 1998. GAGA factor binding to DNA via a single trinucleotide sequence element. Nucleic Acids Res. 26:2672-2678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Wilkins, R. C., and J. T. Lis. 1999. DNA distortion and multimerization: novel functions of the glutamine-rich domain of GAGA factor. J. Mol. Biol. 285:515-525. [DOI] [PubMed] [Google Scholar]
- 51.Wu, C.-H., L. Madabusi, H. Hishioka, P. Emanuel, M. Sypes, I. Archipova, and D. S. Gilmour. 2001. Analysis of core promoter sequences located downstream from the TATA element in the hsp70 promoter from Drosophila melanogaster. Mol. Cell. Biol. 21:1593-1602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Xiao, H., and J. T. Lis. 1988. Germline transformation used to define key features of heat-shock response elements. Science 239:1139-1142. [DOI] [PubMed] [Google Scholar]
- 53.Xiao, H., R. Sandaltzopoulos, H. M. Wang, A. Hamiche, R. Ranallo, K. M. Lee, D. Fu, and C. Wu. 2001. Dual functions of largest NURF subunit NURF301 in nucleosome sliding and transcription factor interactions. Mol. Cell 8:531-543. [DOI] [PubMed] [Google Scholar]