Abstract
GerE from Bacillus subtilis is the smallest member of the LuxR-FixJ family of transcription activators. Its 74-amino-acid sequence is similar over its entire length to the DNA binding domain of this protein family, including a putative helix-turn-helix (HTH) motif. In this report, we sought to define regions of GerE involved in promoter activation. We examined the effects of single alanine substitutions at 19 positions that were predicted by the crystal structure of GerE to be located on its surface. A single substitution of alanine for the phenylalanine at position 6 of GerE (F6A) resulted in decreased transcription in vivo and in vitro from the GerE-dependent cotC promoter. However, the F6A substitution had little effect on transcription from the GerE-dependent cotX promoter. In contrast, a single alanine substitution for the leucine at position 67 (L67A) reduced transcription from the cotX promoter, but not from the cotC promoter. The results of DNase I protection assays and in vitro transcription reactions lead us to suggest that the F6A and L67A substitutions define two regions of GerE, activation region 1 (AR1) and AR2, that are required for activation of the cotC and cotX promoters, respectively. A comparison of our results with those from studies of MalT and BvgA indicated that other members of the LuxR-FixJ family may use more than one surface to interact with RNA polymerase during promoter activation.
Proteins that have a structure similar to LuxR and FixJ comprise a large subfamily of transcriptional activator proteins, including NarL, UhpA, and MalT (23). Members of the LuxR-FixJ family contain a carboxy-terminal domain that includes an amino acid sequence similar to the helix-turn-helix (HTH) motif found in other DNA-binding proteins (22, 23) (Fig. 1). The crystal structure of NarL (2) identified four specific α-helices in the C-terminal region of the protein that are characteristic of DNA-binding proteins (14). These include a region that shares significant homology with the HTH motif in the region of RNA polymerase sigma factors that interacts with the DNA at the −35 regions of their cognate promoters (18).
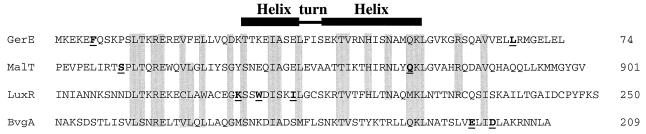
FIG. 1.
Alignment of amino acid sequences from GerE (8), MalT (5), LuxR (10,13), and BvgA (1). The numbers at the right indicate the positions of the rightmost amino acid shown in each sequence. The areas shaded gray represent regions of similarity between the proteins. The underlined boldface lettering indicates the amino acids that have been shown to be important for transcription activation by GerE, MalT, LuxR, and BvgA. The horizontal bars at the top represent the location of the HTH motif in the proposed DNA binding region (6).
GerE, the smallest member of the LuxR-FixJ family, is homologous over its entire sequence to the carboxy terminus of the LuxR-FixJ family of transcriptional activators (Fig. 1). This 74-amino-acid protein is required for activation or repression of several ςK-dependent promoters (e.g., cotC, cotX, and cotD) during the late stages of sporulation in Bacillus subtilis (7, 8). The structure of GerE has been solved at a 2.05-Å resolution by X-ray crystallography (11), and a consensus GerE binding site, RWWTRGGYNNYY (where R = G/A, W = A/T, Y = C/T, and N = any nucleotide), has been identified by DNase I footprint analysis of several GerE-dependent promoters (6, 16, 25, 28, 29).
Transcriptional activators in bacteria can be classified by the position of their binding sites on promoters. Class I activators bind upstream from the −35 region of promoters, and several have been shown to function by interacting with the carboxy-terminal domain of the α-subunit (α-CTD) of the RNA polymerase (17). A classic example of a class I activator is catabolite gene activator protein (CAP) from Escherichia coli (for a recent review, see reference 4). Other class I activators include FNR (19) and φ29 protein p4 (21). Class II activators bind near the −35 region of the promoter DNA and usually interact with the N-terminal domain of the α subunit of RNA polymerase or with the ς subunit (4). Examples of class II activators that interact with ς factors include λ cI (20) and Spo0A from B. subtilis (3). In addition to its function as a class I activator, CAP acts as a class II activator that can associate with the N-terminal domain of the RNA polymerase α subunit on some promoters.
DNase I footprint results show that GerE, like CAP, can bind to various locations on promoters. These results suggest that GerE may act as both a class I and class II transcriptional activator, depending on the promoter to which it is binding. For example, GerE binds to the cotC promoter at three locations, centered at −134.5, −68.5, and −40.5, respectively (29; unpublished data), suggesting that GerE may make multiple contacts with the RNA polymerase (e.g., the α-CTD and ςK). In addition, GerE binds to a region of DNA surrounding the −35 region on the cotX promoter (28). On this promoter, GerE may act as a class II activator, associating with the α subunit or ςK of the RNA polymerase. Evidence that GerE interacts with ςK at the cotX promoter comes from the studies of Wade et al. (27), who found that an amino acid substitution in ςK prevents GerE-dependent activation of the cotX promoter, but has no effect on another GerE-dependent promoter, cotD.
It is not known what region of GerE interacts with ςK at the cotX promoter or whether more than one surface of GerE interacts with RNA polymerase to activate class I and class II type promoters. In order to identify regions of GerE that are required for promoter activation, we examined the effects of single alanine substitutions in 19 positions on the surface of GerE. Our results define two regions of GerE, activation region 1 (AR1) and AR2, that are required for transcription activation of the cotC and cotX promoters, respectively. A comparison of our results with studies from other LuxR-FixJ family members is also discussed.
MATERIALS AND METHODS
Plasmids and strains.
The plasmids and strains used in this study are listed in Tables 1 and 2, respectively. pOR356 contains gerE flanked by approximately 100 bp upstream of the transcription start site and 226 bp downstream of the translation stop codon. Oligonucleotides used for PCR amplifications are listed in Table 3. In order to create a gerE deletion strain, the 5′ flanking DNA of gerE was generated from JH642 chromosomal DNA by PCR with oligonucleotides GerE-5′For (HindIII end) and GerE-5′Rev (XbaI end) (Table 3). This fragment of DNA was cloned into HindIII-XbaI-digested pUC19 by using the Rapid DNA ligation kit (Roche Diagnostics Corp., Indianapolis, Ind.), producing pDC991. The kanamycin gene from pKΔ102 was inserted into pDC991 at the unique BamHI site, thus producing pDC992. The 3′ flanking DNA of gerE was generated from JH642 chromosomal DNA by PCR with oligonucleotides GerE-3′For (KpnI end) and GerE-3′Rev (EcoRI end) (Table 3) and cloned into KpnI-EcoRI-digested pDC992, producing pDC993.
TABLE 1.
Plasmids used in this study
| Plasmid | Contents | Source |
|---|---|---|
| pOR356 | HindIII-EcoRI fragment of pER19 in pLD30 (see Materials and Methods) | R. Losick |
| pGerE-E5A | pOR356 with the E5A substitution | This study |
| pGerE-F6A | pOR356 with the F6A substitution | This study |
| pGerE-Q7A | pOR356 with the Q7A substitution | This study |
| pGerE-S8A | pOR356 with the S8A substitution | This study |
| pGerE-K9A | pOR356 with the K9A substitution | This study |
| pGerE-P10A | pOR356 with the P10A substitution | This study |
| pGerE-S11A | pOR356 with the S11A substitution | This study |
| pGerE-K14A | pOR356 with the K14A substitution | This study |
| pGerE-Q25A | pOR356 with the Q25A substitution | This study |
| pGerE-K27A | pOR356 with the K27A substitution | This study |
| pGerE-K30A | pOR356 with the K30A substitution | This study |
| pGerE-F37A | pOR356 with the F37A substitution | This study |
| pGerE-K41A | pOR356 with the K41A substitution | This study |
| pGerE-R44A | pOR356 with the R44A substitution | This study |
| pGerE-H46A | pOR356 with the H46A substitution | This study |
| pGerE-Q52A | pOR356 with the Q52A substitution | This study |
| pGerE-L67A | pOR356 with the L67A substitution | This study |
| pGerE-E73A | pOR356 with the E73A substitution | This study |
| pGerE-L74A | pOR356 with the L74A substitution | This study |
| pGerE-Ex | gerE coding sequence in pET26b (Novagen) | J. Brannigan |
| pGerE-ExF6A | pGerE-Ex with the F6A substitution | This study |
| pGerE-ExL67A | pGerE-Ex with the L67A substitution | This study |
| pGerE-ExL74A | pGerE-Ex with the L74A substitution | This study |
| pUC19 | New England Biolabs | |
| pKΔ102 | Source of kanamycin resistance cassette | A. Henriques |
| pDC993 | pUC19 containing the 5′ and 3′ regions of gerE flanking the kanamycin cassette of pKΔ102 | This study |
| pKW25 | PcotC in pCR2.1-TOPO (Invitrogen) | 27 |
| pJZ6 | cotX and cotY in pUC18 | 28 |
TABLE 2.
Bacterial strains used in this study
| Strain | Genotype | Source |
|---|---|---|
| B. subtilis | ||
| JH642 | trpC2 pheA1 | J. Hoch |
| AH131 | trpC2 metC3 amyE::erm | A. Henriques |
| EUDC9901 | JH642 gerE::kan | This study |
| EUDC9902 | EUDC9901 SPβcotC-lacZ | This study |
| EUDC990201 | EUDC9902 amyE::WT gerE | This study |
| EUDC990202 | EUDC9902 amyE::E5A-gerE | This study |
| EUDC990203 | EUDC9902 amyE::F6A-gerE | This study |
| EUDC990204 | EUDC9902 amyE::Q7A-gerE | This study |
| EUDC990205 | EUDC9902 amyE::S8A-gerE | This study |
| EUDC990206 | EUDC9902 amyE::K9A-gerE | This study |
| EUDC990207 | EUDC9902 amyE::P10A-gerE | This study |
| EUDC990208 | EUDC9902 amyE::S11A-gerE | This study |
| EUDC990209 | EUDC9902 amyE::K14A-gerE | This study |
| EUDC990210 | EUDC9902 amyE::Q25A-gerE | This study |
| EUDC990211 | EUDC9902 amyE::K27A-gerE | This study |
| EUDC990212 | EUDC9902 amyE::K30A-gerE | This study |
| EUDC990213 | EUDC9902 amyE::F37A-gerE | This study |
| EUDC990214 | EUDC9902 amyE::K41A-gerE | This study |
| EUDC990215 | EUDC9902 amyE::R44A-gerE | This study |
| EUDC990216 | EUDC9902 amyE::H46A-gerE | This study |
| EUDC990217 | EUDC9902 amyE::Q52A-gerE | This study |
| EUDC990218 | EUDC9902 amyE::L67A-gerE | This study |
| EUDC990219 | EUDC9902 amyE::E73A-gerE | This study |
| EUDC990220 | EUDC9902 amyE::L74A-gerE | This study |
| EUDC9903 | EUDC9901 SPβcotX-lacZ | This study |
| EUDC990301 | EUDC9903 amyE::WT-gerE | This study |
| EUDC990302 | EUDC9903 amyE::E5A-gerE | This study |
| EUDC990303 | EUDC9903 amyE::F6A-gerE | This study |
| EUDC990304 | EUDC9903 amyE::Q7A-gerE | This study |
| EUDC990305 | EUDC9903 amyE::S8A-gerE | This study |
| EUDC990306 | EUDC9903 amyE::K9A-gerE | This study |
| EUDC990307 | EUDC9903 amyE::P10A-gerE | This study |
| EUDC990308 | EUDC9903 amyE::S11A-gerE | This study |
| EUDC990309 | EUDC9903 amyE::K14A-gerE | This study |
| EUDC990310 | EUDC9903 amyE::Q25A-gerE | This study |
| EUDC990311 | EUDC9903 amyE::K27A-gerE | This study |
| EUDC990312 | EUDC9903 amyE::K30A-gerE | This study |
| EUDC990313 | EUDC9903 amyE::F37A-gerE | This study |
| EUDC990314 | EUDC9903 amyE::K41A-gerE | This study |
| EUDC990315 | EUDC9903 amyE::R44A-gerE | This study |
| EUDC990316 | EUDC9903 amyE::H46A-gerE | This study |
| EUDC990317 | EUDC9903 amyE::Q52A-gerE | This study |
| EUDC990318 | EUDC9903 amyE::L67A-gerE | This study |
| EUDC990319 | EUDC9903 amyE::E73-gerE | This study |
| EUDC990320 | EUDC9903 amyE::L74A-gerE | This study |
| EUDC9904 | EUDC9901 SPβgerE-lacZ | This study |
| EUDC990401 | EUDC9904 amyE::WT-gerE | This study |
| EUDC990402 | EUDC9904 amyE::E5A-gerE | This study |
| EUDC990403 | EUDC9904 amyE::F6A-gerE | This study |
| EUDC990404 | EUDC9904 amyE::Q7A-gerE | This study |
| EUDC990405 | EUDC9904 amyE::S8A-gerE | This study |
| EUDC990406 | EUDC9904 amyE::K9A-gerE | This study |
| EUDC990407 | EUDC9904 amyE::P10A-gerE | This study |
| EUDC990408 | EUDC9904 amyE::S11A-gerE | This study |
| EUDC990409 | EUDC9904 amyE::K14A-gerE | This study |
| EUDC990410 | EUDC9904 amyE::Q25A-gerE | This study |
| EUDC990411 | EUDC9904 amyE::K27A-gerE | This study |
| EUDC990412 | EUDC9904 amyE::K30A-gerE | This study |
| EUDC990413 | EUDC9904 amyE::F37A-gerE | This study |
| EUDC990414 | EUDC9904 amyE::K41A-gerE | This study |
| EUDC990415 | EUDC9904 amyE::R44A-gerE | This study |
| EUDC990416 | EUDC9904 amyE::H46A-gerE | This study |
| EUDC990417 | EUDC9904 amyE::Q52A-gerE | This study |
| EUDC990418 | EUDC9904 amyE::L67A-gerE | This study |
| EUDC990419 | EUDC9904 amyE::E73A-gerE | This study |
| EUDC990420 | EUDC9904 amyE::L74A-gerE | This study |
| EUDC9905 | EUDC9901 SPβsigK-lacZ | This study |
| EUDC990501 | EUDC9905 amyE::WT-gerE | This study |
| EUDC990502 | EUDC9905 amyE::E5A-gerE | This study |
| EUDC990503 | EUDC9905 amyE::F6A-gerE | This study |
| EUDC990504 | EUDC9905 amyE::Q7A-gerE | This study |
| EUDC990505 | EUDC9905 amyE::S8A-gerE | This study |
| EUDC990506 | EUDC9905 amyE::K9A-gerE | This study |
| EUDC990507 | EUDC9905 amyE::P10A-gerE | This study |
| EUDC990508 | EUDC9905 amyE::S11A-gerE | This study |
| EUDC990509 | EUDC9905 amyE::K14A-gerE | This study |
| EUDC990510 | EUDC9905 amyE::Q25A-gerE | This study |
| EUDC990511 | EUDC9905 amyE::K27A-gerE | This study |
| EUDC990512 | EUDC9905 amyE::K30A-gerE | This study |
| EUDC990513 | EUDC9905 amyE::F37A-gerE | This study |
| EUDC990514 | EUDC9905 amyE::K41A-gerE | This study |
| EUDC990515 | EUDC9905 amyE::R44A-gerE | This study |
| EUDC990516 | EUDC9905 amyE::H46A-gerE | This study |
| EUDC990517 | EUDC9905 amyE::Q52A-gerE | This study |
| EUDC990518 | EUDC9905 amyE::L67A-gerE | This study |
| EUDC990519 | EUDC9905 amyE::E73A-gerE | This study |
| EUDC990520 | EUDC9905 amyE::L74A-gerE | This study |
| E. coli | ||
| DH5α | Gibco BRL | |
| BL21(DE3) (pLysS) | Stratagene | |
| EUKW9820 | BL21(DE3)(pLysS)(pGerE-Ex) | 27 |
| EUDC0010 | BL21(DE3)(pLysS)(pGerE-ExF6A) | This study |
| EUDC0011 | BL21(DE3)(pLysS)(pGerE-ExL67A) | This study |
TABLE 3.
Oligonucleotides used for PCR, sequencing, and mutagenesis
| Name | Sequence (5′→3′) |
|---|---|
| T7 promoter | TAATACGACTCACTATAGGG |
| GerE-Seq1 | GCTCTCACCTTAGCAAG |
| GerE-964F | GCCCTTCTTACATATGATATCTCG |
| GerE-5′For | GGAGATGAAAAAAGATGAAG |
| GerE-5′Rev | GCCAGTCTAGAGTAACCCTCCTTGCTAG |
| GerE-3′For | GCCGGGTACCGCTGTTTGTTTGGAACATTTCG |
| GerE-3′Rev | GCCGGAATTCGATGTATCCTGACAACACGTCTG |
| E5A-For | GAAGGAGAAAGCATTTCAATCG |
| E5A-Rev | CGATTGAAATGCTTTCTCCTTC |
| F6A-For | GAAGGAGAAAGAAGCTCAATCGAAGCCG |
| F6A-Rev | CGGCTTCGATTGAGCTTCTTTCTCCTTC |
| Q7A-For | GAGAAAGAATTTGCATCGAAGCCGTCG |
| Q7A-Rev | CGACGGCTTCGATGCAAATTCTTTCTC |
| S8A-For | GAAAGAATTTCAAGCGAAGCCGTCG |
| S8A-Rev | CGACGGCTTCGCTTGAAATTCTTTC |
| K9A-For | GAATTTCAATCGGCGCCGTCGCTAACG |
| K9A-Rev | CGTTAGCGACGGCGCCGATTGAAATTC |
| P10A-For | GAATTTCAATCGAAGGCGTCGCTAACG |
| P10A-Rev | CGTTAGCGACGCCTTCGATTGAAATTC |
| S11A-For | CAATCGAAGCCGGCGCTAACGAAAAG |
| S11A-Rev | CTTTTCGTTAGCGCCGGCTTCGATTG |
| K14A-For | GCCGTCGCTAACGGCAAGAGAAAGAGAAG |
| K14A-Rev | CTTCTCTTTCTCTTGCCGTTAGCGACGGC |
| Q25A-For | CGAATTGCTCGTTGCAGATAAGACAAC |
| Q25A-Rev | GTTGTCTTATCTGCAACGAGCAATTCG |
| K27A-For | GCTCGTTCAAGATGCGACAACAAAGGAG |
| K27A-Rev | CTCCTTTGTTGTCGCATCTTGAACGAGC |
| K30A-For | CAAGATAAGACAACAGCAGAGATTGCAAGCGAGC |
| K30A-Rev | GCTCGCTTGCAATCTCTGCTGTTGTCTTATCTTG |
| F37A-For | GATTGCAAGCGAGCTAGCTATCAGTGAGAAAACCG |
| F37A-Rev | CGGTTTTCTCACTGATAGCTAGCTCGCTTGCAATC |
| K41A-For | CTATTTATCAGTGAGGCAACCGTTCGAAAACC |
| K41A-Rev | GGTTTCGAACGGTTGCCTCACTGATAAATAG |
| R44A-For | GTGAGAAAACCGTTGCAAACCATATTTCAAATG |
| R44A-Rev | CATTTGAAATATGGTTTGCAACGGTTTTCTCAC |
| H46A-For | GAAAACCGTTCGAAACGCTATTTCAAATGCCATG |
| H46A-Rev | CATGGCATTTGAAATAGCGTTTCGAACGGTTTTC |
| Q52A-For | CAAATGCCATGGCGAAATTGGGTGTG |
| Q52A-Rev | CACACCCAATTTCGCCATGGCATTTG |
| L67A-For | GTTGTTGAGCTTGCTAGAATGGGTGAG |
| L67A-Rev | CTCACCCATTCTAGCAAGCTCAACAAC |
| E73A-For | GGGTGAGCTAGCGCTTTAATCCTTG |
| E73A-Rev | CAAGGATTAAAGCGCTAGCTCACCC |
| L74A-For | GGGTGAGCTAGAGGCTTAATCCTTGCCGG |
| L74A-Rev | CCGGCAAGGATTAAGCCTCTAGCTCACCC |
| AmyE-Back | ACCTGCATTTGCCAGCACAACGCC |
| CotC139-For | TGGGCCGATGAAAAATCGGCTC |
| CotC-Rev | TGTCGTAATCACAGTCATAATC |
| CotX-BcII | GAGCAGCTCATTGAACTGATCA |
| CotX-Hind | CGTGTTTTTCTTAAGCTTTCTC |
pDC993 was linearized by restriction endonuclease digestion with ScaI and transformed into JH642 as previously described (15). Chromosomal DNA was isolated from kanamycin (10 μg/ml)-resistant colonies, as described in reference 15, and subjected to PCR to determine if the gene replacement of kanamycin for gerE occurred. The following primer combinations were used: GerE-Seq2 and GerE-3′Rev (Table 3) to verify the absence of gerE and GerE-5′For and GerE-3′Rev (Table 3) to show the increased size of the kanamycin replacement (1 kb) compared to that of the wild-type gerE (224 bp). All resulting colonies gave identical PCR fragment profiles, and one was chosen as EUDC9901 (Table 2).
Transduction of EUDC9901.
In order to measure the effect that the gerE mutations had on gerE-dependent and gerE-independent transcription, EUDC9901 was transduced with an SPβ lysate containing either cotC-lacZ, cotX-lacZ, gerE-lacZ, or sigK-lacZ as previously described (15). Chloramphenicol (5 μg/ml)- and kanamycin-resistant colonies were then plated on Difco sporulation media (DSM) agar that contained the appropriate antibiotics and X-Gal (5-bromo-4-chloro-3-indolyl-β-d-galactopyranoside [20 μg/ml]) to determine the level of transcription from the promoter-lacZ fusions in the absence of gerE. The resulting strains (EUDC9902, EUDC9903, EUDC9904, and EUDC9905) are listed in Table 2.
Protein modeling.
The coordinates for the crystal structure of GerE (11) (RCSB protein data bank accession code 1FSE) were provided by J. Brannigan (University of York). RasMol version 2.6 (command language and program by Roger Sayle, Glaxo Wellcome, United Kingdom) was used to create the model of the GerE-GerE dimer and to determine surface-exposed amino acids of GerE.
Alanine scanning mutagenesis.
The Quick Change site-directed mutagenesis kit (Stratagene, La Jolla, Calif.) was used to create mutations in gerE that resulted in single alanine substitutions. Briefly, pOR356 (used for recombination at the amyE locus of the B. subtilis chromosome) or pGerE-Ex (used for protein overexpression [described below]) was subjected to oligonucleotide-directed mutagenesis with the For-Rev combination of primers (Table 3), and the resulting plasmids are listed in Table 1. Each plasmid was subjected to DNA sequencing (Emory University DNA Sequencing Facility) by using GerE-964F (for the pOR356 derivatives) or the T7 promoter primer (for the pGerE-Ex derivatives) (Table 3) to ensure the presence of the desired mutation.
Each mutant derivative of pOR356, as well as pOR356 (for the wild-type control), was linearized with ScaI and transformed into competent AH131 (Table 2) for insertion into the amyE locus. Chromosomal DNA was prepared from spectinomycin (50 μg/ml)-resistant, erythromycin (1 μg/ml)-sensitive clones and subjected to PCR with GerE-964F and amyE-Back (Table 3) to indicate that the recombination event occurred at the correct location on the chromosome. The resulting PCR fragments were then sequenced with GerE-Seq1 (Table 3) to determine if the desired mutation was present.
The same chromosomal DNA preparations were then used to transform competent EUDC9902, EUDC9903, EUDC9904, and EUDC9905 (Table 2) in order to transfer the amyE::gerE mutations. Antibiotic-resistant clones (kanamycin, chloramphenicol, and spectinomycin) were then patched onto Luria-Bertani (LB) agar containing the appropriate antibiotics plus starch (1%) to determine the replacement of amyE. In addition, the clones were patched onto DSM agar plates containing the respective antibiotics plus X-Gal to determine the relative amount of gerE-dependent and gerE-independent transcription of the promoter-lacZ fusions as compared to no replacement at amyE. Chromosomal DNA was prepared from respective clones and subjected to PCR with GerE-964F and amyE-Back in order to confirm recombination at the amyE locus. These PCR fragments were then sequenced with GerE-Seq1 to ensure that the desired mutation was present.
β-Galactosidase activity.
EUDC9902, EUDC9903, EUDC9904, and EUDC9905 with either wild-type GerE or individual GerE alanine substitutions at the amyE locus were grown in duplicate in 25 ml of DSM with the appropriate antibiotics to the initiation of sporulation (t0). Every hour from t3 to t11, two 300-μl aliquots of each culture were taken; one was taken to measure the optical density at 600 nm, and the other was stored at −80°C until all time points were complete. β-Galactosidase activity was then measured from the frozen samples as previously described by Henriques et al. (15) and reported in Miller units.
Overexpression and partial purification of GerE.
BL21(DE3)(pLysS) cells containing pGerE-Ex, pGerE-ExF6A, or pGerE-ExL67A were grown at 37°C to an optical density at 600 nm of approximately 0.8 in LB medium containing chloramphenicol (25 μg/ml) and kanamycin (30 μg/ml). Expression of GerE was then induced with isopropyl-β-d-thiogalactopyranoside (IPTG) to a final concentration of 1 mM at 37°C. After 30 min, rifampin was added to a final concentration of 200 μg/ml, and the cells were incubated for another 2.5 h at 37°C. The cells were harvested and subjected to purification over a 5-ml HighTrap heparin column as described by Wade et al. (27) with an Acta Explorer 900 and version 3 of the Unicorn software (Amersham Pharmacia Biotech). The concentration of GerE present in each of the preparations was determined by the Bio-Rad protein assay (Bio-Rad Laboratories, Hercules, Calif.) as per the manufacturer’s instructions. The purity of the GerE was determined by Coomassie blue staining (Bio-Rad Laboratories) after electrophoresis into an 18% polyacrylamide gel containing sodium dodecyl sulfate; we routinely achieved >95% purity. The active fraction of protein in each preparation was determined by a DNase I footprint analysis of radiolabeled cotC promoter (described below) with a constant amount of protein and various concentrations of cold cotC promoter DNA.
Preparation of end-labeled DNA and DNase I footprinting.
The oligonucleotides cotC139-FOR and cotX-BclI (50 pmol; Table 3) were end labeled with [γ-32P]ATP by using T4 polynucleotide kinase (Promega, Madison, Wis.) as per the manufacturer’s instructions. Unincorporated 32P was removed by a G-25 Microspin column (Amersham Pharmacia Biotech, Piscataway, N.J.). The labeled and purified oligonucleotides were then subjected to PCR with 36 pmol of cotC-REV primer (with cotC139-FOR) or cotX-Hind (with cotX-BclI) (Table 3) and Herculase polymerase (Stratgene). The final PCR products (cotC, 268 bp; cotX, 236 bp) were then cleaned with a G-50 Microspin column (Amersham Pharmacia Biotech) to remove any unincorporated oligonucleotides, and the specific activity of the DNA probe was measured with a Beckman LS-6500 scintillation counter. We routinely recovered greater than 90% of the probe with a typical specific activity of approximately 5 × 106 cpm/μg.
The end-labeled DNA probe was subjected to DNase I footprinting reactions with wild-type, F6A-, or L67A-substituted GerE as previously described by Zheng et al. (29). Briefly, DNA fragments labeled at one end were incubated in separate reaction mixtures without protein or with various concentrations of purified GerE in a 42-μl reaction mixture containing 10 mM HEPES (pH 7.5), 50 mM NaCl, 1 mM EDTA (pH 8), 1 mM dithiothreitol, and 10% glycerol. Poly(dI-dC) was added to a final concentration of 1.2 μg/ml, and the reaction mixtures were incubated at 37°C for 10 min. DNase I (3 μl of 0.0004 mg/ml) was then added to the reaction mixtures, and after 1 min at 37°C, the digests were terminated by addition of 50 μl of STOP buffer (100 mM Tris-HCl [pH 8], 50 mM EDTA, 200 μg of yeast tRNA per ml) and incubation for 2 min at 65°C. The DNA in each reaction mixture was then subjected to ethanol precipitation followed by electrophoresis in a 7 M urea-polyacrylamide gel containing 6% acrylamide.
In vitro transcription.
Plasmids pKW25, digested with XbaI (PcotC), and pJZ6, digested with BglII (PcotX), were used as templates for in vitro transcription reactions. The reactions were performed as previously described by Wade et al. (27) with purified ςK-associated RNA polymerase and wild-type or F6A- or L67A-substituted GerE. Specific transcription from the cotC and cotX promoters produced nucleotide transcripts of 182 and 183 bp, respectively.
RESULTS
Substitutions in GerE that affect specific promoters in vivo.
We used the model of the GerE structure produced by X-ray crystallography (11) to identify surface-exposed residues. By using the Rasmol program (see Materials and Methods), we observed that approximately half of the amino acids of GerE were located on the surface of the protein, and we chose 12 amino acids that resulted in an even distribution over the entire surface. We also chose seven amino acids at the amino-terminal end of the protein, because the crystal structure suggested that these amino acids form an “arm” that is positioned away from the core structure of the protein. We replaced each of these amino acids with an alanine residue. Mutant alleles of gerE were introduced in the bacterial chromosome at the amyE locus by transformation of B. subtilis strains that contained a null allele of gerE (gerE::kan, EUDC9901) and either a cotC-lacZ (EUDC9902), cotX-lacZ (EUDC9903), gerE-lacZ (EUDC9904), or sigK-lacZ promoter fusion (EUDC9905) at the SPβ locus (Table 2). Strains transformed with wild-type GerE served as positive controls, whereas untransformed strains served as negative controls.
Transformants were patched onto DSM agar plates containing X-Gal to determine the relative amount of gerE-dependent and gerE-independent transcription of the promoter-lacZ fusions compared to that with no replacement at amyE (Table 4). The results indicated that gerE-lacZ expression was unaffected by each of the alanine substitutions, which was expected, since GerE does not play a role in the transcription of its own gene, gerE. In addition, the strains containing the sigK-lacZ fusion produced white colonies, which indicated that all of the alanine-substituted forms of GerE could still repress transcription from the sigK promoter (16). Two alanine substitutions in GerE (R44A and H46A) appeared to reduce both cotC-lacZ and cotX-lacZ expression. Another substitution (F6A) appeared to reduce cotC-lacZ expression, but seemed to have no effect on cotX-lacZ expression. Expression of cotX-lacZ, but not of cotC-lacZ, appeared to be reduced by the substitutions L67A and L74A.
TABLE 4.
Effects of alanine substitutions in GerE on promoter activity in vivo
| GerEa | β-Galactosidase activity (Miller units)b | |||
|---|---|---|---|---|
| cotC-lacZ | cotX-lacZ | gerE-lacZ | sigK-lacZ | |
| Wild type | 220 (+) | 219 (+) | 108 (+) | − |
| gerE::kanc | <10 (−) | <10 (−) | 168 (+) | + |
| E5A | 236 (+) | 154 (+) | 60 (+) | − |
| F6A | <10 (−) | 125 (+) | 121 (+) | − |
| Q7A | 209 (+) | 290 (+) | 102 (+) | − |
| S8A | 320 (+) | 234 (+) | 113 (+) | − |
| K9A | 276 (+) | 214 (+) | 131 (+) | − |
| P10A | 292 (+) | 410 (+) | 113 (+) | − |
| S11A | 325 (+) | 252 (+) | 60 (+) | − |
| K14A | 253 (+) | 184 (+) | 67 (+) | − |
| Q25A | 291 (+) | 72 (+/−) | 81 (+) | − |
| K27A | 281 (+) | 66 (+/−) | 100 (+) | − |
| K30A | 359 (+) | 261 (+) | 155 (+) | − |
| F37A | 99 (+/−) | 85 (+/−) | 196 (+) | − |
| K41A | 226 (+) | 118 (+) | 174 (+) | − |
| R44A | <10 (−) | 88 (+/−) | 172 (+) | − |
| H46A | <10 (−) | <10 (−) | 132 (+) | − |
| Q52A | 234 (+) | 123 (+) | 101 (+) | − |
| L67A | 106 (+/−) | 35 (−) | 126 (+) | − |
| E73A | 178 (+) | 92 (−) | 100 (+) | − |
| L74A | 160 (+) | <10 (−) | 91 (+) | − |
gerE allele at the amyE locus.
β-Galactosidase activity at t10, cotC-lacZ; t9, cotX-lacZ; t8, gerE-lacZ. Colony type: +, blue colony; −, white colony; +/−, mixture of blue and white. The values shown are the average of two independent experiments. The variation between experiments was less than 5% in all cases.
gerE deletion mutant without a gerE allele at amyE.
To obtain quantitative measures of the effects of the mutations in GerE, the strains were cultured in liquid DSM, and the β-galactosidase accumulation in the cultures was monitored throughout sporulation. These results correlated well with the agar plate analysis (Table 4). The colonies that were blue on agar plates also produced high levels of β-galactosidase activity, whereas the colonies that were white expressed very low levels of β-galactosidase activity. The F37A, R44A, and H46A substitutions in GerE resulted in reduced expression from both GerE-dependent promoters. The F6A substitution in GerE abolished the expression of cotC-lacZ, whereas cotX-lacZ expression was reduced by only twofold (Fig. 2A and B, respectively). This indicated that F6 may be important specifically for activation of PcotC. In addition, the Q25A, K27A, L67A, E73A, and L74A substitutions in GerE resulted in lower levels of cotX-lacZ expression (Table 4 and Fig. 2D), whereas these substitutions had little if any effect on transcription from cotC-lacZ (Fig. 2C). These amino acids may therefore be important for specific activation from PcotX.
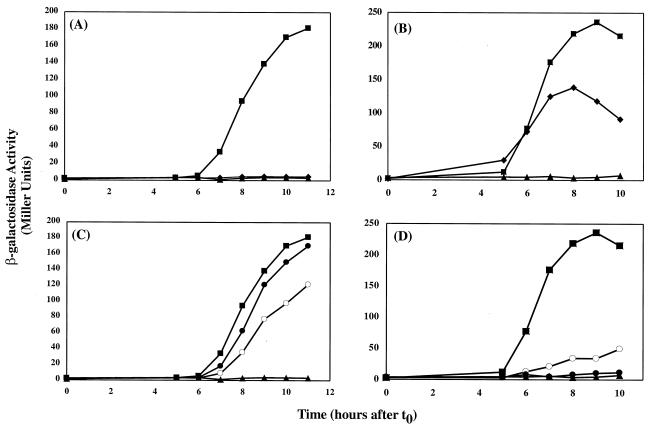
FIG. 2.
Effects of alanine substitutions in GerE on transcription from the cotC and cotX promoters. B. subtilis strains were grown in DSM liquid medium. Samples were taken at the onset of sporulation (t0) and at 1-h intervals thereafter (t3 to t11) and assayed for β-galactosidase accumulation. Two independent cultures were assayed for each of the strains mentioned above, and the averages are shown. (A and C) Measurement of cotC-lacZ expression in B. subtilis strains EUDC9902 (triangles; gerE::kan), EUDC990201 (squares; wild-type GerE), EUDC990203 (diamonds; F6A-GerE), EUDC990218 (open circles; L67A-GerE), and EUDC990220 (solid circles; L74A-GerE). (B and D) Measurement of cotX-lacZ expression in B. subtilis strains EUDC9903 (triangles; gerE::kan), EUDC990301 (squares; wild-type GerE), EUDC990303 (diamonds; F6A-GerE), EUDC990318 (open circles; L67A-GerE), and EUDC990320 (solid circles; L74A-GerE).
F6A-substituted GerE binds to PcotC but fails to activate cotC transcription in vitro.
Two models might explain how a mutant form of GerE would activate one GerE-dependent promoter, but fail to activate another. In one model, the mutant form of GerE binds to one promoter, but not the other. In the alternative model, the mutant form of GerE binds to both promoters, but fails to activate one promoter. Since F6A-substituted GerE failed to stimulate cotC-lacZ expression in vivo, we purified F6A-substituted GerE, examined its binding to cotC in DNase I footprinting and electrophoretic mobility shift assay (EMSA) experiments in vitro, and tested whether this form of GerE stimulated transcription in vitro. Approximately 700 pM wild-type (lane d) or F6A-substituted (lane g) GerE was needed to protect the primary GerE binding site on the cotC promoter from DNase I cleavage (Fig. 3). Similar results were observed in the EMSA experiment (data not shown). This indicated that the F6A substitution had no large effect on the protein’s ability to bind to the cotC promoter. We also found that the F6A-substituted GerE bound with apparent affinities equal to those of wild-type GerE to the cotX and sigK promoters in DNase I footprinting and EMSA experiments (data not shown).
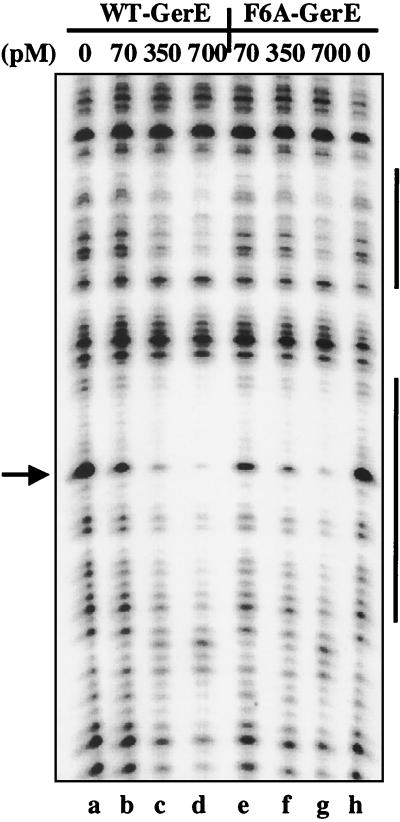
FIG. 3.
DNase I footprint of wild-type (WT) and F6A-substituted GerE on PcotC. The cotC promoter, end labeled on the nontranscribed strand, was incubated in separate reactions with 0 (lanes a and h), 70 (lanes b and e), 350 (lanes c and f), or 700 (lanes d and g) pM wild-type (lanes b to d) or F6A-substituted (lanes e to g) GerE and subjected to DNase I digestion. The top vertical bar on the right represents the GerE binding site centered at −40.5 relative to the transcription start site, and the bottom vertical bar on the right represents the GerE binding site centered at −68.5 relative to the transcription start site on the cotC promoter. The arrow indicates a band that is protected by GerE from DNase I cleavage.
We next assayed the ability of F6A-substituted GerE to stimulate ςK-dependent transcription of cotC or cotX in an in vitro reaction by runoff transcription experiments. Transcription reactions that included wild-type GerE produced a strong cotC transcript with 350 or 700 pM GerE (Fig. 4, top panel, lanes c and d), whereas reactions that included 700 pM F6A-substituted GerE did not stimulate ςK-RNA polymerase-dependent transcription of cotC (top panel, lane g), even though this was a concentration of F6A-substituted GerE sufficient to bind PcotC in the DNase I protection assay. No difference was observed between the abilities of wild-type and F6A-substituted GerE to stimulate transcription of cotX (Fig. 4, bottom panel, lanes d and g). Taken together, these data suggested that when bound to the cotC promoter, the F6A-substituted form of GerE failed to stimulate ςK-dependent transcription of cotC.
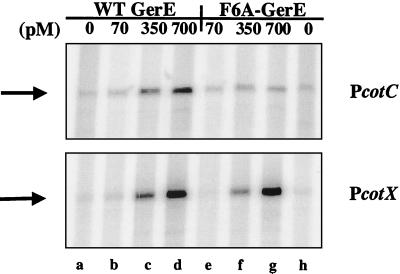
FIG. 4.
In vitro transcription of PcotC and PcotX with wild-type (WT) and F6A-substituted GerE. Shown are portions of scanned images of radiolabeled transcripts that were subjected to electrophoresis on a 6% (wt/vol) polyacrylamide gel containing 7 M urea. DNA templates (1 μg of each template, indicated on the right) were incubated with ςK-associated RNA-P alone (lanes a and h) or with 70 (lanes b and e), 350 (lanes c and f), or 700 (lanes d and g) pM wild-type (lanes b to d) or F6A-substituted (lanes e to g) GerE. The positions of the runoff transcripts are indicated on the left by arrows.
L67A-substituted GerE binds but fails to activate cotX transcription in vitro.
Since L67A-substituted GerE failed to stimulate cotX-lacZ expression in vivo, we purified the L67A-substituted form of GerE and examined its binding to the cotX promoter region. Approximately 350 pM wild-type or L67A-substituted GerE completely protected the GerE binding sites on the cotX promoter in the DNase I footprint experiment (Fig. 5, lanes c and f). Wild-type and L67A-substituted GerE were also equally active in binding to the cotX promoter DNA in gel mobility shift assays (data not shown). Therefore, we tested whether the L67A-substituted GerE would stimulate ςK-dependent transcription of cotC or cotX in an in vitro reaction with runoff transcription experiments. Transcription reaction mixtures that included 350 or 700 pM wild-type or L67A-substituted GerE stimulated cotC transcription (Fig. 6, top panel, lanes c and d and f and g). Moreover, reactions that included 350 or 700 pM wild-type GerE also stimulated cotX transcription (Fig. 6, bottom panel, lanes c and d). However, 700 pM L67A-substituted GerE, which was sufficient for binding to cotX in the DNase I protection assays, did not stimulate ςK-RNA polymerase-dependent transcription of cotX (Fig. 6, bottom panel, lane g). Taken together, these data suggested that when bound to the cotX promoter, L67A-substituted GerE is unable to stimulate ςK-dependent transcription of cotX.
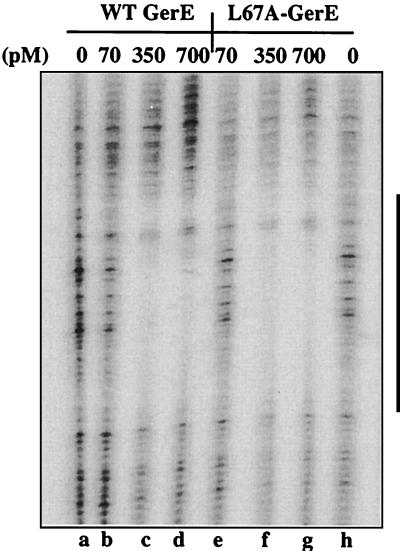
FIG. 5.
DNase I footprint of wild-type (WT) and L67A-substitued GerE on PcotX. The cotX promoter, end labeled on the nontranscribed strand, was incubated in separate reactions with 0 (lane g), 70 (lanes a and d), 350 (lanes b and e), or 700 (lanes c and f) pM wild-type (lanes a to c) or L67A-substituted (lanes d to f) GerE and subjected to DNase I digestion. The vertical bar on the right represents the GerE binding site centered at −42.5 relative to the transcription start site on the cotX promoter.
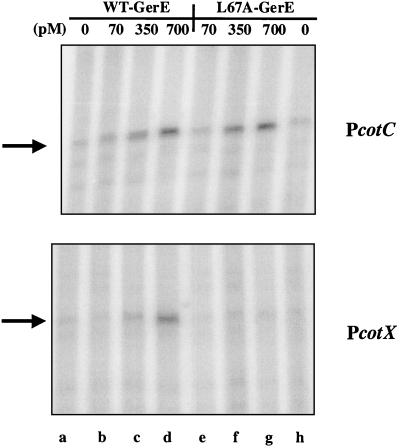
FIG. 6.
In vitro transcription of PcotC and PcotX with wild-type (WT) and L67A-substituted GerE. Portions of scanned images of radiolabeled transcripts that were subjected to electrophoresis on a 6% polyacrylamide gel are shown. DNA templates (1 μg of each template, indicated at the right) were incubated with ςK-associated RNA-P alone (lanes a and h) or with 70 (lanes b and e), 350 (lanes c and f), or 700 (lanes d and g) pM wild-type (lanes b to d) or L67A-substituted (lanes e to g) GerE. The positions of the runoff transcripts are indicated on the left by arrows.
DISCUSSION
We examined the effects of single alanine substitutions in GerE on expression of GerE-dependent promoters. Three of these mutations, F37A, R44A, and H46A, reduced transcription from both the cotC and cotX promoters. These substitutions may have reduced the levels of GerE, reduced its ability to bind DNA, or altered the interaction with the RNA polymerase. The R44A and H46A substitutions lie with the HTH motif that is thought to directly contact DNA upon binding; therefore, these mutations probably affected binding to the cotC and cotX promoters. Using purified R44A- and H46A-substituted GerE, we observed a 10- to 20-fold decrease in the ability of these mutant forms of GerE to bind to DNA (data not shown). The sigK promoter is repressed by GerE, and the R44A-, H46A-, and F37A-substituted forms of GerE appeared to repress sigK-lacZ expression when assayed on plates. These results suggest that these proteins retain some specific binding affinity for DNA. However, GerE binds with higher affinity to the site on the sigK promoter than to the cotC or cotX promoters (6); therefore, these substituted forms of GerE may bind more weakly than wild-type GerE to all sites, but the affinity for sigK remains high enough to cause repression.
The substitutions that affected activity of one GerE-dependent promoter more than another are the ones most likely to define regions directly involved in promoter activation, because activity on one promoter indicates that the mutant protein folds properly, accumulates, and has no general defect in DNA binding. The F6A substitution reduced cotC promoter activity, but had little effect on cotX promoter activity in vivo and in vitro. The F6A-substituted form of GerE, like wild-type GerE, bound to the cotC promoter in vitro, but failed to activate transcription from the cotC promoter in vitro with purified ςK RNA polymerase. F6 is located on the N-terminal arm of GerE, which is extended from the body of the GerE dimer (Fig. 7). The side chain of F6 probably interacts with RNA polymerase at the cotC promoter to stimulate transcription. The F6A substitution defines a region of GerE required for activation of the cotC promoter, but not binding to this promoter. We refer to this region as AR1. The F6A substitution did not have a large effect on cotX transcription; therefore, a different region of GerE may be required for cotX transcription.
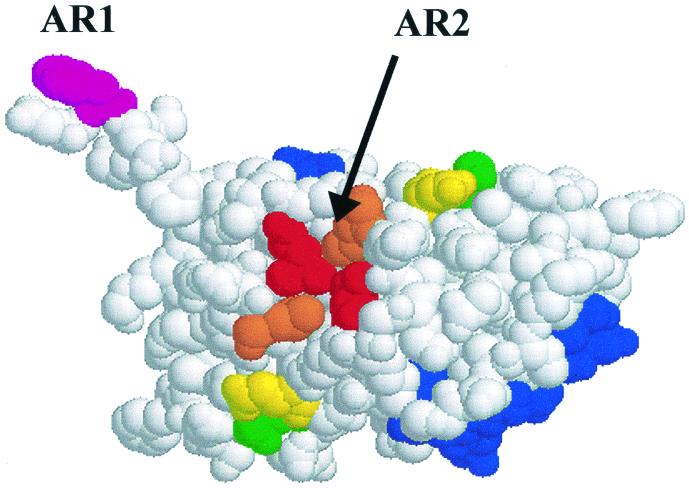
FIG. 7.
Activation regions on the GerE dimer. The amino acids colored in blue (positions 40 to 53 in Fig. 1) form the DNA binding helix in each subunit of the GerE dimer. AR1 is defined by F6 (shaded pink). The amino acids in AR2 are shaded as follows: L67, red; L74; orange; Q25, yellow; K27, green.
The second region of GerE that is required for promoter activation, AR2 is defined by the L67A substitution (Fig. 7). The L67A-substituted form of GerE bound to the cotX promoter, but when bound, it failed to stimulate transcription in vivo and in vitro. This substitution had little or no effect on cotC transcription. L67 is located at the top of GerE dimer on the opposite face of the structure from the DNA binding helix (Fig. 7), where it forms part of a hydrophobic pocket. This hydrophobic pocket may be the site at which GerE interacts with ςK RNA polymerase at the cotX promoter. Other amino acid substitutions that we examined may also lie within AR2. The L74A substitution also specifically affected cotX, but not cotC transcription in vivo (Table 4) and in vitro (not shown). Since cotC promoter activity was stimulated by the L74A form of GerE, it is likely that its DNA binding properties are unaltered. DNase I footprinting studies showed that the L74A form of GerE binds the cotX promoter, albeit with about twofold less affinity than wild-type GerE (data not shown). We note that L74 and L67 interact at the dimer interface to form the hydrophobic pocket that may be the site of interaction with ςK RNA polymerase. Therefore, the L74A substitution may have the additional effect of destabilizing dimer formation, causing its apparent lower affinity for promoter DNA.
The effects of two additional substitutions, Q25A and K27A, indicate that they may also define part of AR2. These substituted forms of GerE had no effect on cotC transcription, but reduced cotX transcription at least twofold (Table 4). Q25 and K27 are located on the same face of GerE as L67 and L74 (Fig. 7), but do not form part of the dimer interface. This suggests that substitutions at Q25 and K27 are unlikely to affect dimer formation and may also directly participate in RNA polymerase contacts made by AR2.
GerE binds at different positions relative the start point of transcription on different promoters, and different surfaces of GerE are required for activation of different promoters. AR1, defined by substitution F6A, is required for activation of the cotC promoter, whereas, AR2, which may include L67, L74, Q25, and K27, is required for cotX transcription. AR1 and AR2 probably interact with ςK RNA polymerase, but which subunits of RNA polymerase are the targets for this interaction is not known. A single amino acid substitution in ςK (H225Y) prevented activation of cotX while having no effect on cotD (27); the effect that this substitution had on cotC transcription was not tested. These results suggested that ςK might be the target of GerE interaction at the cotX promoter. If there were only a single contact between GerE and ςK RNA polymerase at the cotX promoter, we would speculate that AR2 of GerE interacts with ςK. However, whether GerE makes one or more contacts with ςK RNA polymerase at the cotX promoter remains to be shown. Regardless of its target on RNA polymerase, it is remarkable that the small GerE molecule uses different surfaces to stimulate RNA polymerase at different promoters.
Our results may provide insights into the function of other LuxR-FixJ family members. AR1 in GerE and a region of MalT that is required for promoter activation are located in similar positions (Fig. 1). The effects of an amino acid substitution at S834 in MalT indicate that this amino acid plays a role in promoter activation (9). An alignment of the amino acid sequences shows that S834 of MalT corresponds to P10 in GerE (Fig. 1). Single alanine substitutions at P10 and at the adjacent position, S11, in GerE had small effects, slightly increasing expression from the cotX and cotC promoters, respectively. We have not tested whether the alanine substitutions at positions 10 and 11 in GerE have greater effects on other GerE-dependent promoters. The structure of MalT is unknown; however, the similarity of the amino acid sequences of GerE and MalT suggests that the structure of the carboxy-terminal domain of MalT is similar to that of GerE. Therefore, we suggest that MalT and GerE possess activation regions located in similar positions, near the N terminus of their DNA binding domains. The glutamine at position 876 (Q876) in MalT also appears to be involved in promoter activation. Therefore, MalT, like GerE, appears to use two regions on its surface for promoter activation.
Another member of the LuxR-FixJ family may use two regions on its surface for promoter activation. Substitutions at glutamate 198 and aspartate 201 in BvgA, another LuxR-FixJ family member, reduce activation of one BvgA-dependent promoter, ptx, but have no effect on another BvgA-dependent promoter, fha (24). The effects of these mutations were not tested in vitro; however, if the substitutions at positions 198 and 201 of BvgA define an activation site required for ptx promoter activity, a second region of BvgA must be required for fha promoter activation.
Only a single putative activation region has been described in LuxR. Egland and Greenberg identified three single substitutions of alanine for amino acids in LuxR (K198, W201, and I206) (Fig. 1) that may be involved in activation of transcription of the lux operon (12). However, the effects of mutations in LuxR have been assayed only on a single LuxR-dependent promoter (12, 26). Therefore, whether a second region of LuxR would interact with the RNA polymerase when the protein is bound to a different promoter is unknown.
In conclusion, GerE and possibly other LuxR-FixJ family members may use more than one surface to interact with RNA polymerase during promoter activation. Additional studies may reveal whether the members of the LuxR-FixJ family use a small set of homologous surfaces or whether these proteins use a large variety of surfaces to activate promoters.
Acknowledgments
We gratefully acknowledge Diana Caracino and Hamish McGinnis for assistance with isolation and characterization of several of the strains used in this study. We gratefully acknowledge J. A. Brannigan and other members of the Structural Biology Laboratory at University of York for helpful discussions.
This work was supported by grant MCB-9727722 to C.P.M. from the National Science Foundation. D.L.C. was supported in part by IRACDA grant 5K12-GM-00680 to the Emory University School of Medicine.
REFERENCES
- 1.Arico, B., J. F. Miller, C. Roy, S. Stibitz, D. Monack, S. Falkow, R. Gross, and R. Rappuoli. 1989. Sequences required for expression of Bordetella pertussis virulence factors share homology with prokaryotic signal transduction proteins. Proc. Natl. Acad. Sci. USA 86:6671–6675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Baikalov, I., I. Schroder, M. Kaczor-Grzeskowiak, K. Grzeskowiak, R. P. Gunsalus, and R. E. Dickerson. 1996. Structure of the Escherichia coli response regulator NarL. Biochemistry 35:11053–11061. [DOI] [PubMed] [Google Scholar]
- 3.Buckner, C. M., G. Schyns, and C. P. Moran, Jr. 1998. A region in the Bacillus subtilis transcription factor Spo0A that is important for spoIIG promoter activation. J. Bacteriol. 180:3578–3583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Busby, S., and R. H. Ebright. 1999. Transcription activation by catabolite activator protein (CAP). J. Mol. Biol. 293:199–213. [DOI] [PubMed] [Google Scholar]
- 5.Cole, S. T., and O. Raibaud. 1986. The nucleotide sequence of the malT gene encoding the positive regulator of the Escherichia coli maltose regulon. Gene 42:201–208. [DOI] [PubMed] [Google Scholar]
- 6.Crater, D. L., and C. P. Moran, Jr. 2001. Identification of a DNA binding region in GerE from Bacillus subtilis. J. Bacteriol. 183:4183–4189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Cutting, S., and J. Mandelstam. 1986. The nucleotide sequence and the transcription during sporulation of the gerE gene of Bacillus subtilis. J. Gen. Microbiol. 132:3013–3024. [DOI] [PubMed] [Google Scholar]
- 8.Cutting, S., S. Panzer, and R. Losick. 1989. Regulatory studies on the promoter for a gene governing synthesis and assembly of the spore coat in Bacillus subtilis. J. Mol. Biol. 207:393–404. [DOI] [PubMed] [Google Scholar]
- 9.Danot, O., D. Vidal-Ingigliardi, and O. Raibaud. 1996. Two amino acid residues from the DNA-binding domain of MalT play a crucial role in transcriptional activation. J. Mol. Biol. 262:1–11. [DOI] [PubMed] [Google Scholar]
- 10.Devine, J. H., C. Countryman, and T. O. Baldwin. 1988. Nucleotide sequence of the luxR and luxI genes and the structure of the primary regulatory region of the lux regulon in Vibrio fischeri ATCC 7744. Biochemistry 27:837–842. [Google Scholar]
- 11.Ducros, V. M.-A., R. J. Lewis, C. S. Verma, E. J. Dodson, G. Leonard, J. P. Turkenburg, G. N. Murshudov, A. J. Wilkinson, and J. A. Brannigan. 2001. Crystal structure of GerE, the ultimate transcriptional regulator of spore formation in Bacillus subtilis. J. Mol. Biol. 306:759–771. [DOI] [PubMed] [Google Scholar]
- 12.Egland, K. A., and E. P. Greenberg. 2001. Quorum sensing in Vibrio fischeri: analysis of the LuxR DNA binding region by alanine-scanning mutagensis. J. Bacteriol. 183:382–386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Engebrecht, J., and M. Silverman. 1987. Nucleotide sequence of the regulatory locus controlling expression of bacterial genes for bioluminescence. Nucleic Acids Res. 15:10455–10467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Harrison, S. C. 1991. A structural taxonomy of DNA-binding domains. Nature 353:715–719. [DOI] [PubMed] [Google Scholar]
- 15.Henriques, A. O., B. W. Beall, K. Roland, and C. P. Moran, Jr. 1995. Characterization of cotJ, a ςE-controlled operon affecting the polypeptide composition of the coat of Bacillus subtilis spores. J. Bacteriol. 177:3394–3406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ichikawa, H., R. Halberg, and L. Kroos. 1999. Negative regulation by the Bacillus subtilis GerE protein. J. Biol. Chem. 274:8322–8327. [DOI] [PubMed] [Google Scholar]
- 17.Ishihama, A. 1993. Protein-protein communication within the transcription apparatus. J. Bacteriol. 175:2483–2489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kahn, D., and G. Ditta. 1991. Modular structure of FixJ: homology of the transcriptional activator domain with the −35 binding domain of sigma factors. Mol. Microbiol. 5:987–997. [DOI] [PubMed] [Google Scholar]
- 19.Li, B., H. Wing, D. Lee, H. C. Wu, and S. Busby. 1998. Transcription activation by Escherichia coli FNR protein: similarities to, and differences from, the CRP paradigm. Nucleic Acids Res. 26:2075–2081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Li, M., H. Moyle, and M. M. Susskind. 1994. Target of the transcriptional activation function of phage lambda cI protein. Science 263:75–77. [DOI] [PubMed] [Google Scholar]
- 21.Monsalve, M., M. Mencia, M. Salas, and F. Rojo. 1996. Protein p4 represses phage phi 29 A2c promoter by interacting with the alpha subunit of Bacillus subtilis RNA polymerase. Proc. Natl. Acad. Sci. USA 93:8913–8918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Pao, G. M., and J. Saier. 1995. Response regulators of bacterial transduction systems: selective domain shuffling during evolution. J. Mol. Evol. 40:136–154. [DOI] [PubMed] [Google Scholar]
- 23.Pao, G. M., R. Tam, L. S. Lipschiz, and J. Saier. 1994. Response regulators: structure, function and evolution. Res. Microbiol. 145:356–362. [DOI] [PubMed] [Google Scholar]
- 24.Stibitz, S. 1994. Mutations in the bvgA gene of Bordetella pertussis that differentially affect regulation of virulence determinants. J. Bacteriol. 176:5615–5621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Takamatsu, H., Y. Chikahiro, T. Kodama, H. Koide, S. Kozuka, K. Tochikubo, and K. Watabe. 1998. A spore coat protein, CotS, of Bacillus subtilis is synthesized under the regulation of ςK and GerE during development and is located in the inner coat layer of spores. J. Bacteriol. 180:2968–2974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Trott, A. E., and A. M. Stevens. 2001. Amino acid residues in LuxR critical for its mechanism of transcriptional activation during quorum sensing in Vibrio fischeri. J. Bacteriol. 183:387–392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Wade, K. H., G. Schyns, J. A. Opdyke, and C. P. Moran, Jr. 1999. A region of ςK involved in promoter activation by GerE in Bacillus subtilis. J. Bacteriol. 181:4365–4373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Zhang, J., H. Ichikawa, R. Halberg, L. Kroos, and A. I. Aronson. 1994. Regulation of the transcription of a cluster of Bacillus subtilis spore coat genes. J. Mol. Biol. 240:405–415. [DOI] [PubMed] [Google Scholar]
- 29.Zheng, L., R. Halberg, S. Roels, H. Ichikawa, L. Kroos, and R. Losick. 1992. Sporulation regulatory protein GerE from Bacillus subtilis binds to and can activate or repress transcription from promoters for mother-cell-specific genes. J. Mol. Biol. 226:1037–1050. [DOI] [PubMed] [Google Scholar]