Abstract
Continued improvements of adenoviral vectors require the investigation of novel genome configurations. Since adenovirus can be generated directly by transfecting packaging cell lines with viral genomes isolated from plasmid DNA, it is possible to separate genome construction from virus production. In this way failure to generate a virus is not associated with an inability to generate the desired genome. We have developed a novel lambda-based system that allows rapid modification of the viral genome by double homologous recombination in Escherichia coli. The recombination reaction and newly generated genome may reside in a recombination-deficient bacterial host for enhanced plasmid stability. Furthermore, the process is independent of any restriction endonucleases. The strategy relies on four main steps: (i) homologous recombination between an adenovirus cosmid and a donor plasmid (the donor plasmid carries the desired modification[s] and flanking regions of homology to direct its recombination into the viral genome); (ii) in vivo packaging of the recombinant adenoviral cosmids during a productive lambda infection; (iii) transducing a recombination-deficient E. coli lambda lysogen with the generated lysate (the lysogen inhibits the helper phage used to package the recombinant andenoviral cosmid from productively infecting and destroying the host bacteria); (iv) effectively selecting for the desired double-recombinant cosmid. Approximately 10,000 double-recombinant cosmids are recovered per reaction with essentially all of them being the correct double-recombinant molecule. This system was used to generate quickly and efficiently adenoviral genomes deficient in the E1/E3 and E1/E3/E4 regions. The basis of this technology allows any region of the viral genome to be readily modified for investigation of novel configurations.
The ability to modify adenovirus is essential in order to explore its full biological potential as a therapeutic agent in gene therapy. Several modified vectors have been developed, including those that delete portions or a set of the E1, E2A, E2B, E3, E4, and L5 regions (1, 3, 7, 16, 17, 20, 27, 29, 34) which are propagated on cell lines that complement for the functions deleted from the virus (1, 3, 5, 16, 17, 25, 35, 36). Multiply deficient virus vectors with the capacity to carry 10 to 11 kb of foreign DNA have been built. Utilizing these vector configurations for biological and clinical uses has proceeded in a rapid pace in the past few years. However, to sculpt the 36-kb genome of the frequently used adenovirus 2 (Ad2) or Ad5 virus into the desired configuration can be an extremely challenging and tedious task.
Initial vector construction technology primarily relied on in vitro ligation (31, 32) or homologous recombination occurring in a complementing cell line between an adenoviral arm and a shuttle plasmid (23, 28; for a review, see Berkner [2]). These processes are time-consuming, requiring the generation of viral arms and the plaquing of the recombinants away from potential contaminants introduced with the viral arm. By using a plasmid-derived packaging-deficient viral arm, the potential contamination could be effectively eliminated (4). However, in all of these instances if no recombinants arise the researcher does not know if the failure is technical in nature or due to biological impediments. By the creation of a clonal DNA stock of the desired viral genome prior to transducing the complementing cell line, the need for plaque purification can essentially be eliminated and failure to generate the virus is not due to poor ligation or recombination efficiencies.
Various methods have been developed to create viral genomes in noncomplementing cells. The feasibility was initially demonstrated in Escherichia coli and demonstrated that at least one free viral inverted terminal repeat (ITR) is required to produce an infectious genome (3, 19). This demonstration has been expanded to include, by using homologous recombination, yeast (24) and bacteria (9, 10, 21) and in vitro cosmid cloning (15, 26). All of these methods have some limitations. In vitro cosmid cloning only allows insertion of sequences into a predetermined site(s). While homologous recombination can theoretically target any region of the molecule for modification, most require linearization of at least one of the two DNA molecules. The need to avoid particular restriction sites reduces the flexibility of these systems. Although these limitations were circumvented by Crouzet et al. by using two uncut plasmids, the method is limited by the use of a specialized replicon (IncP derived) that function in a polA E. coli host. In all of the homologous recombination methods, the resulting product resides in recombination competent cells. If the product contains multiple regions of homology, the use of these systems will be severely challenged.
To overcome these constraints, a new technology was developed. It is based on homologous recombination in bacteria with preferential packaging of cosmid vectors in vivo by a lambda helper phage and selection for desired products in lysogenic bacteria. The method is quick, efficient (∼100% of the clones are correct), and independent of restriction enzyme digestion. Unlike other described systems the entire process can be carried out in recombination-deficient bacteria. This enhances the opportunity to generate complex vectors and maintain their genomic integrity.
MATERIALS AND METHODS
BN dual expression cassette.
The PstI/SalI fragment-containing EM-7 Sc-ble from pCMV/Zeo (Invitrogen, Inc.) was cloned into pBluescript SK(+) II (Stratagene, Inc.) restricted with PstI/SalI. The amino-terminal 57 amino acids of LacZ were cloned immediately 3′ of the EM-7 promoter inframe with Sc-ble. This expression cassette was subsequently cloned into pSCV-1 on a BamHI/XhoI fragment into the same sites. This placed the EM-7 Sc-Ble expression cassette next to the rat NP-1 defensin that is under Tac promoter control in pSCV-1 (8, 11) to create the BN dual expression cassette that is flanked by SalI restriction sites.
Plasmid description.
All of the cosmid vectors contain the Ad5 genome cloned into pACYC177 (New England BioLabs) between the DrdI and DraI sites to retain the p15 origin of replication and kanamycin (KAN) resistance gene. The adenovirus left ITR resides nearest to the DraI site. The left and right ITRs are flanked with PacI restriction sites. The lambda phage cos packaging site is 32 bp from the viral left ITR. The LacIQ expression cassette is oriented to direct transcription toward the cos site. Cosmids pACE1(f)10 and pACE1(Z)10 are isogenic with an expression cassette replacing E1 sequences 356 to 3327 under the control of the cytomegalovirus (CMV) promoter and SV40 polyadenylation signal expressing green fluorescent protein (GFP) and LacZ, respectively, and a deletion in the E3 region spanning bp 28593 to 30470. The expression cassettes direct expression toward the left viral ITR. pACE1(f)11D is isogenic to pACE1(f)10 with replacement of E4 sequences 32825 to 35565 with the β-glucuronidase gene flanked by the bovine growth hormone (BGH) and simian virus 40 (SV40) polyadenylation sequences at its 5′ and 3′ termini, respectively. pACE1(f)ΔE3/4(BN) is isogenic to pACE1(f)10, with the BN expression cassette replacing adenovirus sequences from 27087 to 35564.
Shuttle vectors pE1-IX(BN) and pAdCLxGLP contain parts of adenovirus sequences 1 to 5788 cloned into a pGEM plasmid (Promega, Inc.). pE1-IX(BN) contains the “BN” dual expression cassette flanked by Ad5 sequences 1 to 355 and 4122 to 5788. pAdCLxGLP contains a CMV promoter and SV40 polyadenylation signal to regulate the GFP variant gene GLP (LifeTechnologies, Inc.), all of which was flanked by Ad5 sequences 1 to 355 and 3328 to 5788. The expression cassette directs transcription toward the left viral ITR. pGBS11D is a pGEM-based vector with Ad5 sequences 21562 to 35935. The adenovirus sequences were modified with an XbaI deletion in E3 (sequences 28593 to 30470) and replacement of E4 sequences 32825 to 35565 with the β-glucuronidase gene flanked by the BGH and SV40 polyadenylation sequences at its 5′ and 3′ termini, respectively.
Bacterial DH5(DE3) lysogen strain.
E. coli BL21(DE3) is a lambda lysogenic strain (F− ompT hsdS gal dcm) that carries integration-deficient immunity 21 phage containing the gene for T7 RNA polymerase under the control of LacUV5 promoter (33). BL21(DE3) was lysogenized with wild-type lambda to generate a double lysogen with dual immunity. A fresh overnight culture of this double lysogen was used to release the spontaneously induced phage, which was then plated onto a lawn of E. coli lysogenized with lambda to purify DE3 phage with immunity 21. Well-isolated plaques of DE3 phage were then used to prepare high-titer liquid lysates on E. coli LE392. Strain DH5α was lysogenized with bacteriophage DE3 by using lambda as the helper phage. DH5(DE3) lysogens were tested for sensitivity to lambda clear and immunity to phage DE3 and also for IPTG (isopropyl-β-d-thiogalactopyranoside)-induced expression of T7 RNA polymerase.
Phage-directed recombination.
E. coli LE392 or DH5α (18, 30) was cotransformed with 100 ng of the cosmid and plasmid vectors before being plated onto Luria-Bertani (LB) agar plates (8 g of Bacto Agar, 10 g of Bacto Tryptone, 5 g of Bacto Yeast, and 5 g of NaCl in 1 liter of water) containing 50 μg of KAN and 100 μg of ampicillin (AMP)/ml and incubated overnight at 30°C. A 2-ml overnight culture in LB medium (10 g of Bacto Tryptone, 5 g of Bacto Yeast, and 5 g of NaCl in 1 liter of water) containing 50 μg of KAN/ml was started from a single colony. The bacteria were isolated by centrifugation and resuspended in 1 ml of 10 mM MgSO4. Helper phage DE3 (33) was plaque purified in top agar (LB medium with 10 mM MgSO4, 0.6% Bacto Agar) at ∼100 plaques/60-mm agarose plate according to standard protocols (30). Four fresh plaques of DE3 were added to 100 μl of the bacterial suspension and allowed to incubate for 30 to 60 min at room temperature before 2 ml of LB medium containing 10 mM MgSO4 was added. The infection was incubated at 37°C at 250 rpm for up to 5 h or until cell lysis was observed before the addition of 200 μl of CHCl3. The lysate was removed from the cell debris and organic phase after being clarified by centrifugation. The lysate is stable at 4°C. A 2-ml overnight LB culture of DH5(DE3) was pelleted by centrifugation before being resuspended in 1 ml of 10 mM MgSO4. Then, 100 μl of the cell suspension and viral lysate were incubated at room temperature for 30 min before 2 ml of LB medium was added. The culture was incubated for 60 min at 37 oC. Transductants were plated on LB agar plates, and selection was applied. To select for the cosmid backbone but against the BN cassette (OUT reaction) 50 μg of KAN/ml and 1 mM IPTG plates were used. To select for the presence of the BN cassette (IN reaction) cells are plated on LB plates with 50 μg of zeocin (ZEO)/ml.
Adenovirus conversion.
Cosmid DNA was restricted with PacI and extracted with phenol and then with 25:25:1 phenol-CHCl3-isoamyl alcohol before being precipitated with ethanol, dried, and resuspended in pyrogen-free water. Next, 5 μg of DNA was brought to a final volume of 250 μl in 100 mM CaCl2. A total of 250 μl of 2× HBS (250 mM NaCl; 50 mM HEPES, pH 7.4; 1.5 mM NaHPO4) was quickly added, mixed, and incubated at room temperature for 1 to 2 min before being added to a 60-mm plate that contained 106 293 or 293-Orf6 cells (6, 17) that had been fed with Dulbecco minimum Eagle’s medium (DMEM) containing 5% fetal calf serum 2 h before. At 4 h posttranfection, the cells were washed once with 1 ml of 1 mM EGTA in 1× HBS and twice with 2 ml of DMEM-2% fetal calf serum and then fed 5 ml of DMEM-5% fetal calf serum. After 5 days the cells were freeze-thawed three times. Then 1 ml of lysate was used to infect a 10-cm plate that was 70% confluent with 293 or 293-Orf6 cells for 1 h before they were fed with 5 ml of DMEM containing 5% fetal calf serum. The virus was passaged as just described until viral cytopathic infection was observed.
RESULTS
Cosmid structure.
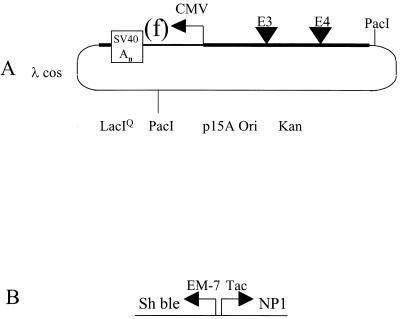
The structure of a representative cosmid vector used in these studies is shown in Fig. 1A. These replicons carry the entire adenovirus genome with modifications, the low-copy-number p15A origin of replication for enhanced plasmid stability, the LacIQ expression cassette, and the lambda cos packaging signal and confer KAN resistance. These cosmids carry the pACE prefix in their names. In the example described here, the transgene expression cassette resides in the E1 region and is regulated by the CMV promoter and SV40 polyadenylation signal. When lacZ or GFP genes are expressed in this cassette their names are abbreviated (as “Z” and “f,” respectively). Cosmids in which the viral genome carry an E1 and E3 deletion contain the designation “10.” The designation “11D” was applied if the viral genome is further modified so that the E4 region is replaced with a transcriptional inert spacer element. In this example, the inert spacer is the β-glucuronidase gene in which the BGH polyadenylation signal resides 5′ of the coding sequence (D. E. Brough et al., unpublished data).
FIG. 1.
Cosmid and BN dual selection cassette structure. (A) pACE1(f)11D. The Ad5 genome was cloned into pACYC177, which was modified to contain LacIQ and the phage lambda packaging site (λcos). The Ad5 genome was modified. E1 sequences 356 to 3327 were replaced with a CMV promoter, SV40 early polyadenylation signal (SV40 An), and GFP gene (f). E3 and E4 deletions are represented as triangles. The E3 deletion spans bp 28593 to 30470. E4 sequences 32825 to 35565 were replaced with the β-glucuronidase gene, oriented right to left with respect to the map, flanked by the BGH and SV40 polyadenylation sequences at its 5′ and 3′ termini, respectively. PacI restriction sites flank the viral genome. (B) Positive and negative dual selection cassette BN. The Sh bleomycin gene under control of the synthetic constitutive EM-7 promoter confers resistance to zeocin. EM-7 is comprised of a constitutive and T7 polymerase responsive promoter. Transcribed in the opposite orientation under the hybrid Trp/Lac (Tac) promoter is a gene that encodes the 32-amino-acid peptide with antimicrobial activity from the rat defensin NP1 gene.
Strategy to alter the adenovirus genome.
A two-step IN-OUT homologous recombination strategy was developed to allow for rapid modification of the viral genome. The IN step is used to mark a region of the viral genome into which modifications are to be targeted. For example, the E1 region can be marked so that any expression cassette flanked with homologous sequences can be directed to this part of the virus. In this way a stock of the marked vector can be made and used to generate many different expression vectors, all having the same viral backbone. The OUT step is the replacement of the marked region with the modification of interest.
The two-step replacement strategy utilizes a positive selection schema for the IN reaction and a negative selection strategy for the OUT reaction. To accommodate these divergent functions, the BN dual selection cassette (Fig. 1B) was created. Positive selection is provided by constitutive expression of the Sh bleomycin gene (12) from the EM-7 promoter (Invitrogen), resulting in ZEO drug resistance. Negative selection is tightly regulated by IPTG. In the presence of IPTG the rat defensin NP-1 and the phage T7 RNA polymerase are expressed. The rat defensin NP-1 sequence is limited to the 32-amino-acid peptide with antimicrobial activity (22). The T7 polymerase, located in the bacterial genome, activates transcription of the Sh bleomycin gene from the T7 component in the EM-7 promoter. Overexpression of transgenes by T7 polymerase is known to inhibit bacterial growth (33). Activation of this “death cassette” by IPTG results in only 1 in 104 to 1 in 105 bacteria that are capable of growing. This effectively eliminates all background bacteria that retain the BN cassette during the OUT reaction.
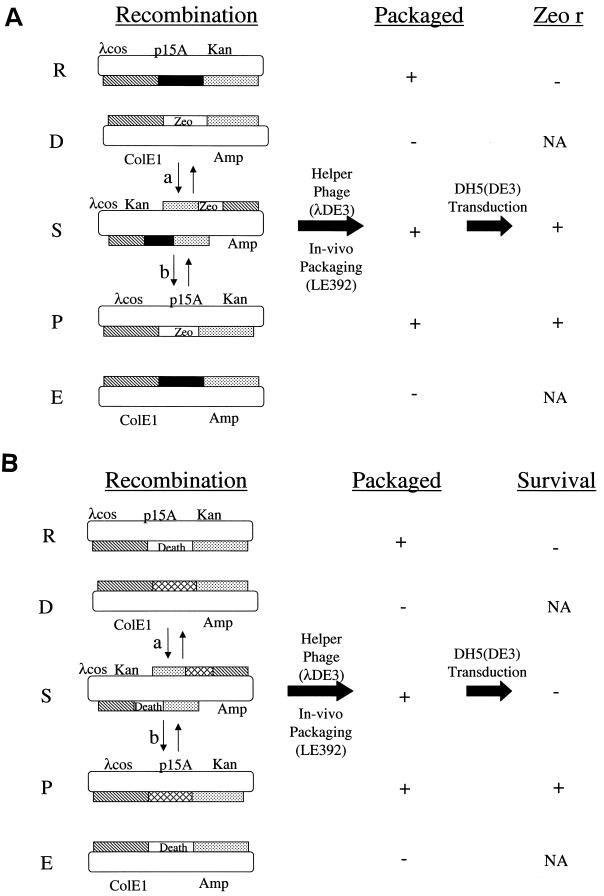
The outline of the IN strategy used to modify the viral genome is depicted in Fig. 2A. The IN step uses the gain of ZEO resistance to mark a particular region of the viral genome with the death cassette. To initiate the process, a pair of replicons with two regions of homology is introduced into a bacterium. The desired product arises by replacement of a region (solid box) in the “recipient” cosmid (R) with the BN cassette (ZEO) from the “donor” plasmid (D) to generate recombinant cosmid (P) by double homologous recombination. Two additional products are generated: the single recombinant cosmid (S) and the second double-recombinant plasmid (E). The recipient cosmid and donor plasmids retain different antibiotic resistance genes, KAN and AMP, respectively, along with compatible origins of replication. This ensures stable maintenance of the two replicons in the same recombination competent bacterium under appropriate positive selective pressure. The culture is infected with a lambda helper phage, which results in the recipient (R), product (P), and single (S) recombinant cosmid clones being packaged into phage heads. The plasmids (D and E) will not be packaged since they lack the phage packaging cos site. The desired recombinant (P), as well as the single recombinant (S), is recovered from the lysate after transduction of a bacterial lambda lysogen by selection for ZEO-resistant colonies. The bacterial lambda lysogen is immune to and therefore not destroyed by the helper phage. The double-recombinant P is readily distinguished from the single recombinant S by its drug-resistant phenotype. The single recombinant S confers KAN, AMP, and ZEO resistance, whereas the double-recombinant P is AMP sensitive.
FIG. 2.
Genome alteration strategy. The recombination column represents the bacteria LE392 cotransformed with both a recipient (R) cosmid (p15A; origin of replication, Kan; r, resistance gene; λcos, lambda packaging sequences), a donor (D) plasmid (ColE1; origin of replication; Amp, resistance gene), and predicted recombinants (S, P, and E). Regions of homology between the two replicons R and D are represented by the hatched and stippled rectangles. The single- and double-recombination pathways are indicated by the lowercase letters “a” and “b,” respectively. The resultant products of these reactions are the single-recombinant (S) and double-recombinant product (P) cosmids and episomal plasmid (E). Helper phage (λDE3) is used to package the cosmid vectors in vivo to generate a lysate. Under the packaged column, “+” and “−” indicate whether the replicons is or is not packaged, respectively. Not indicated is the presence of the packaged helper phage in the lysate. Lysogenic bacteria DH5(DE3) are transduced with the lysate and seeded on agar plates. (A) IN strategy. The BN cassette is indicated by Zeo. Cosmids that do (+) or do not (−) support colony formation in the presence of ZEO are indicated in column marked “Zeo r.” (B) OUT strategy. The BN cassette is indicated by death. Cosmids that do (+) or do not (−) support colony formation in the presence of IPTG and KAN are indicated in the column marked “Survival.” NA, not applicable since the nonpackaged plasmid cannot transduce the lysogen.
The marked double-recombinant cosmid is recovered and used in a second recombination reaction, the OUT step depicted in Fig. 2B. Once the viral genome has been marked with the BN cassette (labeled “death”), this region is readily modified by many different donor plasmids (D). The OUT step takes advantage of the ability to induce the BN death cassette's antigrowth properties to eliminate all transductants that retain it such as the nonrecombinant (R) and single-recombinant (S) cosmids. Nontransduced bacteria perish under KAN selection that is also applied during this antigrowth selection strategy. Only bacteria that harbor the double-recombinant cosmid (P) that has lost the death cassette will survive these selection conditions. The result is a final clone that is seamless, containing no unwanted ancillary sequences.
Positive selection strategy to modify adenovirus E1 region.
To demonstrate the feasibility of this technology, the BN dual selection cassette carrying the ZEO resistance marker was transferred into the E1 region of the viral genome. The BN cassette was flanked by Ad5 sequences 1 to 355 and 4122 to 5788 in plasmid pE1-IX(BN). This plasmid was cotransformed with pACE1(Z)10 into the bacterium LE392 to create the double-recombinant pACE1(BN)10 (Fig. 3A). pE1-IX(BN) is a plasmid with a ColE1 origin of replication, which also confers AMP resistance. Bacteriophage lambda lysates were prepared and used to infect lambda lysogenic bacteria. ZEO-resistant transductants were selected for and obtained at a frequency of 1 × 10−5 to 2 × 10−5 PFU of the phage lysate. The transductants were found to be KAN resistant, suggesting that they comprised a population of single and possibly doubly recombinant molecules.
FIG. 3.
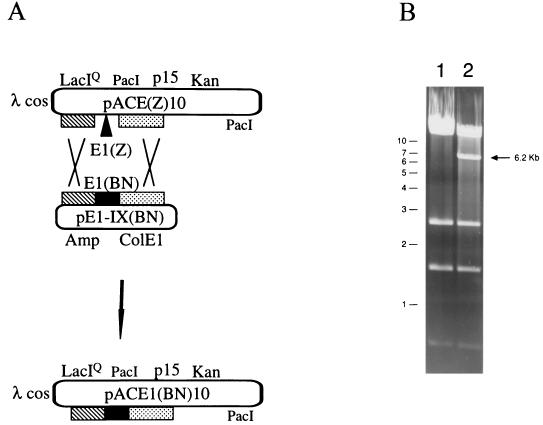
pACE1(BN)10 Cosmid construction: IN selection. (A) Schematic of double-recombination reaction in the E1 region. Regions of homology between the replicons are represented by the hatched and stippled rectangles. The recipient cosmid is pACE1(Z)10, the donor plasmid is pAdE1-IX(BN), and the desired cosmid product is pACE1(BN)10. The cosmid backbone and the Ad5 E3(10) deletion are described in the legend to Fig. 2B. E1(Z), E1 sequences 356 to 3327 were replaced with LacZ under the transcriptional regulation of a CMV promoter and SV40 early polyadenylation signal. E1(BN), the dual selection cassette (Fig. 2) flanked by Ad5 sequences 1 to 355 and 4122 to 5788. The donor plasmid comprises an origin of replication (ColE1) and resistance marker (Amp). (B) Xho restriction analysis of pACE1(Z)10 (lane 1) and pACE1(BN)10 (lane 2) is shown. A 6.2-kb band is unique to pACE1(BN)10. Molecular size markers are shown to the left in kilobases.
Further analysis confirmed the presence of the double-recombinant molecule pACE1(BN)10. Transductants with this molecule would be phenotypically KAN and ZEO resistant but AMP sensitive. About 2% of the ZEO-resistant transductants were KAN resistant and AMP sensitive. Restriction analysis confirmed the cosmids to be pACE1(BN)10. Figure 3B shows the XhoI restriction pattern of plasmid DNA preparations from a phenotypically correct, independent isolate (lane 2) and its parental control pACE1(Z)10 (lane 1). The cosmid showed the loss of a 10,503-bp fragment from its parental control and the appearance of a 6,232-bp fragment. Also present is a doublet of 14,502 and 11,203 bp that is common to both cosmids. A total of four independent isolates were tested, and all were shown to be correct. Further confirmation of pACE1(BN)10 was done by PacI restriction analysis (data not shown). This demonstrated the ability to direct sequence-specific recombination into the adenovirus genome and to recover double-recombinant molecules in lambda phage heads.
E1 region modification.
To demonstrate the technical feasibility and the ease with which transgenes were moved into the E1 region of the adenoviral genome, the above-described E1 region vector pACE1(BN)10 was chosen as the recipient and pAdCLxGLP was chosen as the donor replicon (Fig. 4A). Plasmid pAdCLxGLP confers ampicillin resistance on a ColE1 replicon. A CMV promoter and SV40 polyadenylation signal regulate GFP expression, all of which was flanked by Ad5 sequences 1 to 355 and 3328 to 5788. E. coli LE392, cotransformed by these two replicons, was used as the host to prepare bacteriophage lambda lysates, which were then used to transduce lambda lysogens. Recovery of the putative double recombinant, pACE1(f)10, as measured by KAN- and IPTG-resistant transductants was obtained at a frequency of 2 × 10−6 to 3 × 10−6 PFU in the phage lysate. All 100 transductants tested were scored as AMP and ZEO sensitive when assayed on selective LB agar plates, a finding consistent with the identification of double-recombinant molecules. There were ca. 400 and 1,700 base pairs of homology flanking the GFP expression cassette in these experiments.
FIG. 4.
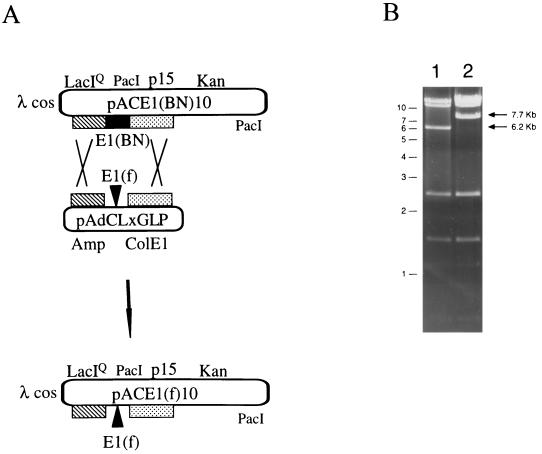
E1 replacement construction: OUT selection. (A) Schematic of double-recombination reaction in E1 region. Regions of homology between the replicons are represented by the hatched and stippled rectangles. The recipient cosmid is pACE1(BN)10, the donor plasmid is pAdCLxGLP, and the desired cosmid product is pACE1(f)10. The cosmid backbone, Ad5 E1(f), and E3(10) modifications are described in the legend to Fig. 2B. E1(BN), the dual selection cassette (Fig. 2A) replaced Ad5 sequences 356 to 4122. The donor plasmid comprises an origin of replication (ColE1) and resistance marker (Amp). (B) Xho restriction analysis of pACE1(BN)10 (lane 1) and pACE1(f)10 (lane 2) are shown. A 6.2-kb band is unique to pACE1(BN)10, while a 7.7-kb fragment is unique to pACE1(Z)10. Molecular size markers are shown to the left in kilobases.
DNA restriction analysis with XhoI of 18 putative pACE1(f)10 confirmed them all to be correct. A representative is shown in Fig. 4B (lane 2). The positive transductant showed the loss of the 6,232-bp fragment from the parental pACE1(BN)10 (lane 1) and the appearance of a 7,701-bp fragment with pACE1(f)10. Further confirmation of this double-crossover homologous recombination event was done by PacI restriction analysis (data not shown). Approximately 12,000 to 16,000 positive clones were calculated to have been generated since a 2 ml portion of lysates was prepared and one-tenth of the 10−2 dilution was plated to generate six to eight pACE1(f)10 transductants. Similar levels of double recombinants were obtained with the recombination-deficient bacteria DH5α.
E3/E4 region modification.
A similar IN-OUT selection strategy was used to replace sequences in the adenovirus E3 and E4 regions. The BN dual selection cassette was moved into pACE1(f)10 replacing adenovirus sequences from 27087 to 35564, creating pACE1(f)ΔE3/4(BN). The deletion encompassed coding sequences for protein VIII through all of the E4 region. pACE1(f)ΔE3/4(BN) was only 32,566 bp in size and was too small to be efficiently packaged in a phage head. Therefore, an alternative construction method (Brough et al., unpublished) was used in which a linear fragment containing BN and a circular cosmid were transfected into E. coli BJ5183 (recBC sbcBC) (18) and ZEO-resistant clones were selected. The transductants were also KAN resistant and confirmed by HindIII restriction analysis to be pACE1(f)ΔE3/4(BN) (data not shown).
An E1, E3, and E4 modified genome was readily constructed. LE392 cotransfected with pACE1(f)ΔE3/4(BN) and pGBS11D was used to generate a lysate with the helper phage. pGBS11D carried a ColE1 origin of replication, AMP resistance gene, and Ad5 sequences 21562 to 35935. The adenovirus sequences were modified with an XbaI deletion in E3 (sequences 28593 to 30470) and the replacement of E4 sequences 32825 to 35565 with the β-glucuronidase gene flanked by the BGH and SV40 polyadenylation sequences at its 5′ and 3′ termini, respectively. The lysate was used to infect a lambda lysogen. KAN- and IPTG-resistant transductants were scored as the double-recombinant pACE1(f)11D and found at a frequency of 1 × 10−6 to 10 × 10−6 PFU in the phage lysate in two independent experiments. Again, all 50 KAN- and IPTG-resistant transductants tested from each experiment were sensitive to both AMP and ZEO, a finding consistent with the occurrence of a double-crossover homologous recombination event. In these experiments, there were ca. 400 and 5,500 bp of homology to the late and early sides flanking the region of interest, respectively.
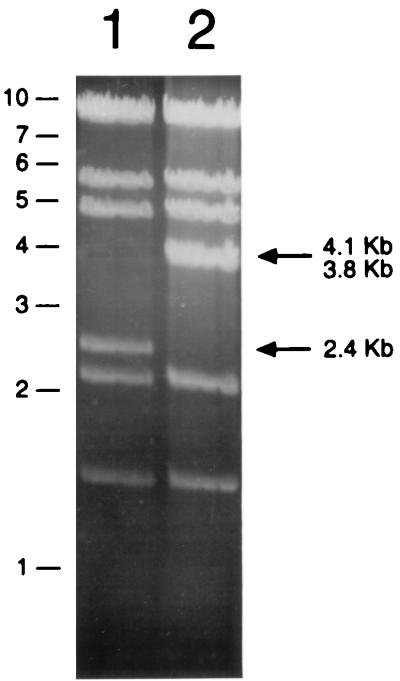
DNA restriction analysis confirmed the generation of pACE1(f)11D. Figure 5 shows the HindIII restriction pattern of a representative of cosmid DNAs prepared from nine independent transductants (lane 2), along with the parental control pACE1(f)ΔE3/4(BN) (lane 1). All nine positive transductants of pACE1(f)11D showed the loss of a 2,408-bp fragment from the parental control and the appearance of the two new predicted fragments of 4,075 and 3,787 bp. The total number of predicted double recombinants was again in the 5,000 to 10,000 range, similar to that in the E1 region experiments. Likewise, similar levels of double recombinants were obtained with the recombination-deficient bacteria DH5α.
FIG. 5.
E4 replacement analysis. HindIII restriction analysis of the product cosmid pACE1(f)11D (lane 2, described in the legend to Fig. 2B) and recipient cosmid pACE1(f)ΔE3/4(BN) (lane 1) is shown. pACE1(f)11D showed the loss of a 2.4-kb fragment from the parental control (lane 2) and the appearance of the two new predicted fragments of 4.1 and 3.8 kb.
An equal number of double-recombinant transductants was found with the E1 and E3/E4 replacement vectors. In both cases, ca. 400 bp of homology was available to one side, while the other contained either 1,600 or 5,500 bp of homology. This suggests that 1,600 bp of homology is saturating in these recombination reactions. Similar results were found when the recombination reactions were carried out in the recombination-deficient DH5α strain. In this case, the recombinase was provided by the phage Red/Gam gene products in this RecA strain.
Recombinant adenoviral plasmid constructs are infectious in permissive mammalian cells.
PacI restriction sites flank the viral genomes of both pACE1(f)10 and an isogenic vector derived from pACE1(f)11D in which the E1 region contains a luciferase expression cassette. The plasmids were restricted with PacI and transfected into human 293 (E1 complementing) and 293-Orf6 (E1 and E4 complementing) cells, respectively (6, 17). At 5 days posttransfection the cells were lysed by freeze-thawing and passaged onto their respective complementing cell lines. By the second and third passages, full viral cytopathic infection was observed. The identity of the viral genomes was confirmed by PCR analysis (data not shown).
DISCUSSION
A novel phage-dependent technology was developed that provides efficient modification of the adenovirus genome by homologous recombination in E. coli. Unlike other recombination systems the entire process of this technology can be carried out in recombination-deficient bacteria with efficiencies similar to those obtained when recombination-competent bacteria are used. The recombinase is putatively provided by lambda's Red/Gam gene products and is only present during the helper phage portion of the process. It is therefore estimated that the recombinase is present for less than an hour, the time required for the lytic infection. This provides the opportunity to work with unstable genetic elements and reduces the risk of generating mutations in the final vector genome.
The use of a dual selection cassette allows targeting of any region of the molecule for recombination while eliminating the need for restriction enzymes. The method relies on the ability of bacteria to carry out homologous recombination between a plasmid and cosmid replicon, packaging of the cosmid during a lambda infection, and transducing lysogenic bacteria under conditions to identify the desired double-recombinant cosmid (Fig. 1). The E1, E3, and E4 regions were modified with an efficiency approaching 100% of selected clones being correct and yielding ca. 10,000 positive clones. According to this protocol, E1/E3- and E1/E3/E4-deficient adenovirus vectors were produced. The E2, L2, and L5 regions have also been modified with similar efficiencies (data not shown).
To recover specifically and efficiently the desired double-recombinant adenoviral plasmid, several important features were incorporated into this system. The two replicons used to generate the recombinant cosmid contained compatible origins of replication and confer different drug resistances to enhance stable cotransduction within a single bacterium. The large adenovirus-containing recipient replicon contains the low-copy-number origin of replication, p15, and confers KAN resistance, while the donor plasmid carries a ColE1 origin of replication and AMP resistance. Since the KAN-resistant replicon contains the bacteriophage lambda cos packaging signal, only the parental and recombinant cosmids are packaged into phage heads during the lambda infection stage. A negative selection cassette, inducible with IPTG, resides in the locus to be replaced by homologous recombination. Selection for transductants on KAN plates in the presence of IPTG yields only double-recombinant cosmids, which contain the modification of interest. The use of lambda lysogenic strains of E. coli that are immune to the helper phage prevents lytic growth of the helper phage and subsequent destruction of the transduced cells.
Targeting the region of the genome for modification was simplified with the BN dual selection cassette. The dual selection cassette is composed of a positive cassette (ZEO resistance) and negative cassette that are juxtaposed to each other (Fig. 2A). By flanking this cassette with predetermined sequences, a positive selection scheme can be used to position the negative selection cassette precisely within the adenovirus genome.
Negative selection with this system is provided by two components of the dual selection cassette, both of which are IPTG regulated. One component was the 32-amino-acid peptide with antimicrobial activity from the rat defensin gene NP1 under the control of the Tac promoter (8). The antimicrobial activity has been attributed to the ability of defensins to form pores in bacterial membranes (22). Additional negative selection is provided by activation of the phage T7 promoter contained within the EM-7 constitutive promoter that is part of the ZEO resistance positive selection cassette. In the presence of T7 polymerase, the growth of the bacteria is inhibited due to its action on the phage T7 promoter contained within the positive selection cassette. T7 polymerase is expressed from the lysogen in this system after IPTG induction. Growth inhibition has also been attributed to overexpression of genes under T7 polymerase control (33).
The recipient BN cosmid vectors are designed to take advantage of phage genetics and are below optimum packaging size. Lambda is capable of packaging DNA in the size range of 78% (38 kb) to 105% (51 kb) of that of wild-type lambda DNA (13, 14). Therefore, by making the recipient cosmid either too small or too large the nonrecombinant background is reduced. Illegitimate recombinants and those arising from internal deletions may also be selected against.
The system's packaging of recombinants into phage heads also provides additional benefits. The packaged cosmids are no longer in a recombination-competent environment but in an essentially inert one. This eliminates the possibility of additional nondesired recombination events occurring. Therefore, the packaged cosmids can be stored for extended periods of time in this stable state and assayed when desired. The transduction and propagation of the cosmids into a recA host also provided greater confidence that no additional recombination events occur.
The ability to rapidly modify the adenovirus genome greatly enhances the prospects of investigating many potentially useful vector configurations. The present vectors and methodology are designed to quickly and efficiently modify any region of the adenovirus genome independent of restriction enzymes. These cosmid vectors also provide the opportunity to change the virus completely in vitro by using the classical ligation and packaging reactions. The combination of creating the viral genome of choice coupled with in vitro cosmid cloning provides the ability to make custom-designed complex adenovirus libraries, the full potential of which has yet to be explored.
Acknowledgments
We thank Joe Bruder, Jason Gall, Thomas Wickham, and Rick King for critical reading of the manuscript.
REFERENCES
- 1.Amalfitano, A., M. A. Hauser, H. Hu, D. Serra, C. R. Begy, and J. S. Chamberlain. 1998. Production and characterization of improved adenovirus vectors with the E1, E2b, and E3 genes deleted. J. Virol. 72:926-933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Berkner, K. L. 1988. Development of adenovirus vectors for the expression of heterologous genes. BioTechniques 6:616-629. [PubMed] [Google Scholar]
- 3.Berkner, K. L., and P. A. Sharp. 1983. Generation of adenovirus by transfection of plasmids. Nucleic Acids Res. 11:6003-6020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bett, A. J., W. Haddara, L. Prevec, and F. L. Graham. 1994. An efficient and flexible system for construction of adenovirus vectors with insertions or deletions in early regions 1 and 3. Proc. Natl. Acad. Sci. USA 91:8802-8806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Brough, D. E., V. Cleghon, and D. F. Klessig. 1992. Construction, characterization, and utilization of cell lines which inducibly express the adenovirus DNA-binding protein. Virology 190:624-634. [DOI] [PubMed] [Google Scholar]
- 6.Brough, D. E., A. Lizonova, C. Hsu, V. A. Kulesa, and I. Kovesdi. 1996. A gene transfer vector-cell line system for complete functional complementation of adenovirus early regions E1 and E4. J. Virol. 70:6497-6501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Challberg, S. S., and G. Ketner. 1981. Deletion mutants of adenovirus 2: isolation and initial characterization of virus carrying mutations near the right end of the viral genome. Virology 114:196-209. [DOI] [PubMed] [Google Scholar]
- 8.Chan, R. Y., R. G. Palfree, L. F. Congote, and S. Solomon. 1994. Development of a novel type of cloning vector for suicide selection of recombinants. DNA Cell Biol. 13:311-319. [DOI] [PubMed] [Google Scholar]
- 9.Chartier, C., E. Degryse, M. Gantzer, A. Dieterle, A. Pavirani, and M. Mehtali. 1996. Efficient generation of recombinant adenovirus vectors by homologous recombination in Escherichia coli. J. Virol. 70:4805-4810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Crouzet, J., L. Naudin, C. Orsini, E. Vigne, L. Ferrero, A. Le Roux, P. Benoit, M. Latta, C. Torrent, D. Branellec, P. Denefle, J. F. Mayaux, M. Perricaudet, and P. Yeh. 1997. Recombinational construction in Escherichia coli of infectious adenoviral genomes. Proc. Natl. Acad. Sci. USA 94:1414-1419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.de Boer, H. A., L. J. Comstock, and M. Vasser. 1983. The tac promoter: a functional hybrid derived from the trp and lac promoters. Proc. Natl. Acad. Sci. USA 80:21-25. [DOI] [PMC free article] [PubMed]
- 12.Drocourt, D., T. Calmels, J. P. Reynes, M. Baron, and G. Tiraby. 1990. Cassettes of the Streptoalloteichus hindustanus ble gene for transformation of lower and higher eukaryotes to phleomycin resistance. Nucleic Acids Res. 18:4009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Feiss, M., R. A. Fisher, M. A. Crayton, and C. Egner. 1977. Packaging of the bacteriophage lambda chromosome: effect of chromosome length. Virology 77:281-293. [DOI] [PubMed] [Google Scholar]
- 14.Feiss, M., and D. A. Siegele. 1979. Packaging of the bacteriophage lambda chromosome: dependence of cos cleavage on chromosome length. Virology 92:190-200. [DOI] [PubMed] [Google Scholar]
- 15.Fu, S., and A. B. Deisseroth. 1997. Use of the cosmid adenoviral vector cloning system for the in vitro construction of recombinant adenoviral vectors. Hum. Gene Ther. 8:1321-1330. [DOI] [PubMed] [Google Scholar]
- 16.Gorziglia, M. I., C. Lapcevich, S. Roy, Q. Kang, M. Kadan, V. Wu, P. Pechan, and M. Kaleko. 1999. Generation of an adenovirus vector lacking E1, e2a, E3, and all of E4 except open reading frame 3. J. Virol. 73:6048-6055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Graham, F. L., J. Smiley, W. C. Russell, and R. Nairn. 1977. Characteristics of a human cell line transformed by DNA from human adenovirus type 5. J. Gen. Virol. 36:59-74. [DOI] [PubMed] [Google Scholar]
- 18.Hanahan, D. 1983. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. 166:557-580. [DOI] [PubMed] [Google Scholar]
- 19.Hanahan, D., and Y. Gluzman. 1984. Rescue of functional replication origins from embedded configurations in a plasmid carrying the adenovirus genome. Mol. Cell. Biol. 4:302-309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Harrison, T., F. Graham, and J. Williams. 1977. Host-range mutants of adenovirus type 5 defective for growth in HeLa cells. Virology 77:319-329. [DOI] [PubMed] [Google Scholar]
- 21.He, T. C., S. Zhou, L. T. da Costa, J. Yu, K. W. Kinzler, and B. Vogelstein. 1998. A simplified system for generating recombinant adenoviruses. Proc. Natl. Acad. Sci. USA 95:2509-2514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hill, C. P., J. Yee, M. E. Selsted, and D. Eisenberg. 1991. Crystal structure of defensin HNP-3, an amphiphilic dimer: mechanisms of membrane permeabilization. Science 251:1481-1485. [DOI] [PubMed] [Google Scholar]
- 23.Ho, Y. S., R. Galos, and J. Williams. 1982. Isolation of type 5 adenovirus mutants with a cold-sensitive host range phenotype: genetic evidence of an adenovirus transformation maintenance function. Virology 122:109-124. [DOI] [PubMed] [Google Scholar]
- 24.Ketner, G., F. Spencer, S. Tugendreich, C. Connelly, and P. Hieter. 1994. Efficient manipulation of the human adenovirus genome as an infectious yeast artificial chromosome clone. Proc. Natl. Acad. Sci. USA 91:6186-6190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Klessig, D. F., D. E. Brough, and V. Cleghon. 1984. Introduction, stable integration, and controlled expression of a chimeric adenovirus gene whose product is toxic to the recipient human cell. Mol. Cell. Biol. 4:1354-1362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kojima, H., N. Ohishi, and K. Yagi. 1998. Generation of recombinant adenovirus vector with infectious adenoviral genome released from cosmid-based vector by simple procedure allowing complex manipulation. Biochem. Biophys. Res. Commun. 246:868-872. [DOI] [PubMed] [Google Scholar]
- 27.Legrand, V., D. Spehner, Y. Schlesinger, N. Settelen, A. Pavirani, and M. Mehtali. 1999. Fiberless recombinant adenoviruses: virus maturation and infectivity in the absence of fiber. J. Virol. 73:907-919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Miyake, S., M. Makimura, Y. Kanegae, S. Harada, Y. Sato, K. Takamori, C. Tokuda, and I. Saito. 1996. Efficient generation of recombinant adenoviruses using adenovirus DNA-terminal protein complex and a cosmid bearing the full-length virus genome. Proc. Natl. Acad. Sci. USA 93:1320-1324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Rice, S. A., and D. F. Klessig. 1985. Isolation and analysis of adenovirus type 5 mutants containing deletions in the gene encoding the DNA-binding protein. J. Virol. 56:767-778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Silhavy, T. J., M. L. Berman, L. W. Enquist, and Cold Spring Harbor Laboratory. 1984. Experiments with gene fusions. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
- 31.Stow, N. D. 1981. Cloning of a DNA fragment from the left-hand terminus of the adenovirus type 2 genome and its use in site-directed mutagenesis. J. Virol. 37:171-180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Stow, N. D. 1982. The infectivity of adenovirus genomes lacking DNA sequences from their left-hand termini. Nucleic Acids Res. 10:5105-5119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Studier, F. W., and B. A. Moffatt. 1986. Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J. Mol. Biol. 189:113-130. [DOI] [PubMed] [Google Scholar]
- 34.Von Seggern, D. J., C. Y. Chiu, S. K. Fleck, P. L. Stewart, and G. R. Nemerow. 1999. A helper-independent adenovirus vector with E1, E3, and fiber deleted: structure and infectivity of fiberless particles. J. Virol. 73:1601-1608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Von Seggern, D. J., J. Kehler, R. I. Endo, and G. R. Nemerow. 1998. Complementation of a fibre mutant adenovirus by packaging cell lines stably expressing the adenovirus type 5 fibre protein. J. Gen. Virol. 79:1461-1468. [DOI] [PubMed] [Google Scholar]
- 36.Weinberg, D. H., and G. Ketner. 1983. A cell line that supports the growth of a defective early region 4 deletion mutant of human adenovirus type 2. Proc. Natl. Acad. Sci. USA 80:5383-5386. [DOI] [PMC free article] [PubMed] [Google Scholar]