Abstract
Micro RNAs comprise a large family of small, functional RNAs with important roles in the regulation of protein coding genes in animals and plants. Here we show that human and mouse miRNA22 precursor molecules are subject to posttranscriptional modification by A-to-I RNA editing in vivo. The observed editing events are predicted to have significant implications for the biogenesis and function of miRNA22 and might point toward a more general role for RNA editing in the regulation of miRNA gene expression.
Keywords: RNA editing, miRNA, inosine, pri-miRNA, adenosine deaminase, ADAR
RNA editing by adenosine (A) to inosine (I) modification generates RNA and protein diversity in higher eukaryotes, selectively altering coding and noncoding sequences in nuclear transcripts (for reviews, see Bass 2002; Maas et al. 2003). The enzymes responsible for A-to-I editing, the adenosine deaminases acting on RNA (ADARs), are ubiquitously expressed in mammals and specifically recognize partially double-stranded (ds) RNA structures where they modify individual adenosines depending on the local structure and sequence environment (Bass 2002). Long extended dsRNAs undergo massive editing (Morse et al. 2002), whereas RNA duplex structures with bulges and loops are subject to site-selective editing, as observed in several nuclear encoded pre-mRNAs (Bass 2002; Schmauss and Howe 2002). Because inosine acts as guanosine during translation, A-to-I conversion in coding sequences leads to amino acid changes and often entails changes in protein function. The best studied examples are neuronal glutamate receptor (Glu-R) genes where single amino acid changes alter gating behavior and kinetic properties of the ion channels (Seeburg et al. 1998; Seeburg and Hartner 2003) and a serotonin receptor subunit where editing regulates G-protein coupling efficiency (Burns et al. 1997). Furthermore, editing within noncoding regions of pre-mRNAs can lead to alternative splicing (Rueter et al. 1999) or might affect the translation, transport, or stability of RNAs (Morse et al. 2002). Theoretically, any RNA with the ability to form a dsRNA fold-back structure could be a target for editing.
Micro (mi) RNAs (Lai 2003; Bartel 2004) are generated as long pri-miRNA transcripts that are processed into ~60–70-nt pre-miRNAs in the nucleus by the ribonuclease III-like enzyme Drosha (Lee et al. 2003). After nuclear export the pre-miRNAs are further processed by Dicer into ~20–22-nt mature miRNAs (Bernstein et al. 2001). The stem–loop structures predicted for all miRNA precursors are reminiscent of the partially double-stranded fold-back structures of known editing substrates (Bass 2002), raising the possibility that miRNA precursor molecules might be targeted by ADARs in vivo. Because ADARs are predominantly nuclear enzymes, the most likely targets for miRNA editing are pri-miRNA and pre-miRNA precursor molecules prior to nuclear export.
To test this hypothesis, we analyzed brain-specific cDNAs from human and mouse pri-miRNA22 for evidence of A-to-I RNA editing. In contrast to many other known human miRNA genes, miRNA22 is widely expressed in mammalian tissues (Lagos-Quintana et al. 2001, 2002) and sequences that correspond to pri-miRNA22 transcripts can be found in expressed sequence databases (dbEST; Boguski et al. 1993). Interestingly, human and mouse EST sequences corresponding to miRNA22 precursors appear to be spliced from a longer, primary transcript with several exons and show evidence of alternative splicing (Fig. 1A), properties usually associated with RNA polymerase II transcripts (Goldstrohm et al. 2001). We specifically amplified pri-miRNA22 molecules corresponding to two alternative splice forms (see Fig. 1A) and sequence analyzed a 197-nt region including the complete miRNA22 precursor stem–loop of 85 nt, which encompasses 17 adenosines.
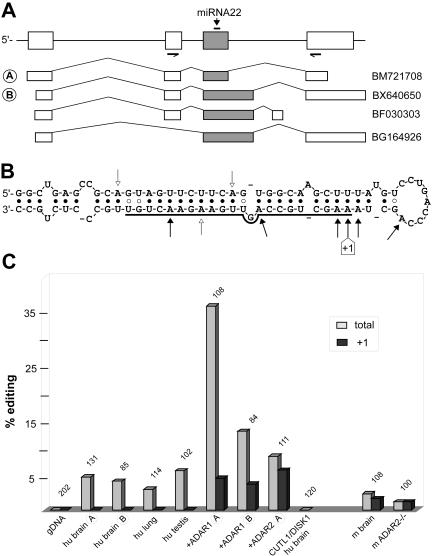
FIGURE 1.
A-to-I RNA editing of miRNA22. (A) The exon/intron structure of the human miRNA22 gene as deduced from expressed sequences with GenBank accession numbers. Exons are shown as boxes, introns as lines. The location of the mature miRNA22 sequence and the positions of the oligonucleotide primers used for editing analysis of two different splice variants A and B are indicated. (B) Predicted RNA secondary structure of pre-miRNA22 with the mature miRNA sequence underlined. Edited positions in human and/or mouse tissues (filled arrowheads) and additional sites found only in ADAR1 overexpressing cell lines (open arrowheads) are indicated. (C) Editing levels of miRNA22 precursor molecules in various human tissues, human cell lines, wild-type mouse brain, and brains of ADAR2 knock-out mice. A PCR product derived from the human miRNA22 genomic region (gDNA), as well as the reverse transcribed and amplified coding sequences from human CUTL1 (CCAAT displacement proteinlike) and DISK1 (disrupted in Schizophrenia 1) genes, were analyzed as negative controls. The number of individually sequenced clones is indicated above each column. In addition to the total extent of editing (percent of molecules edited at one or more of the 17 adenosines within the pre-miRNA), the results for the +1 position of miRNA22 are shown because they point toward a pronounced site selectivity of ADAR2 versus ADAR1. For RNA editing analysis, total RNA was isolated from human brain (Harvard Brain Tissue Collection) using Trizol Reagent (Invitrogen) and reverse transcribed with Superscript Reverse Transcriptase II (Invitrogen) and N6 random oligonucleotides as described (Maas et al. 2001). Genomic DNA was isolated from the same specimen. Additional sources for cDNA preparation were mouse brain total RNA prepared from SV129 mice, human lung and testis RNAs (Clontech), and total RNA extracted from stably transfected human cell lines (Maas et al. 2001). cDNA from ADAR2 knock-out mice (Higuchi et al. 2000) were a kind gift from Miyoko Higuchi and Peter H. Seeburg (Heidelberg). Primers miRNA22D (5′-TGAGGAGCCTGTTCCTCTCACG-3′) and miRNA22U (5′-CGCACTATGGTGCCACATCTCG-3′) for cDNA and primers gmiRNA22D (5′-GGAATTCCTTAGGAGCCTGTTCCTCTCACG-3′) and gmiRNA22U (5′-CCAGGAATCTAGAGTCTGGGC-3′) for genomic DNA were used for miRNA22-specific PCRs. Amplicons were subcloned into pBSII vector plasmid (Stratagene) and recombinant clones were isolated by blue–white selection. At least 100 individual clones were sequenced (C-tracked) from each subcloned PCR fragment. Splice form A was analyzed for all samples and splice variant B where detectable. The theoretical background of A/G discrepancies resulting from errors during reverse transcription and PCR was calculated as P = a · c + b · c · d = 3.4 · 10−5 · 17 + 0.8 · 10−5 · 17 · 24 = 0.4 · 10−2, with a being the error rate of Superscript reverse transcriptase (3.4 · 10−5; Potter et al. 2003), b the error rate of Taq DNA polymerase (0.8 · 10−5; Cline et al. 1996), c the number of adenosines analyzed, and d the number of template doublings during PCR (estimated for 30 cycles of amplification). We would therefore expect 0.4 base changes affecting adenosine in 100 sequences, corresponding to 1 base change due to mutation in 250 sequences. Because changes of A to C and A to T are also possible, this error rate is likely an overestimation because the analysis only detects A-to-G changes. The results from analysis of genomic DNA isolated from human brain is in line with this value, as within 200 sequences clones we did not detect a single A/G change. Analysis of genomic DNA from HEK293 cells yielded the same result (data not shown), as well as RT-PCR and sequencing of coding regions from human CUTL1 (BC025422) and DISC1 (AK025293) RNAs.
In brains from human and mouse, as well as in human lung and testis, low-level editing is detectable at several positions, including nucleotides within the mature miRNA sequence (Fig. 1B). The occurrence of miRNA22 editing in two mammalian species further suggests a functional relevance for editing. Within our sample the positions of editing events, however, are different for human tissues (nucleotide positions −5, −1,+9,+18) and mouse tissues (−41,+1,+2).
To establish a direct link between RNA editing enzyme activity and base changes in pri-miRNA sequences, the analysis was extended to human cell lines that ectopically overproduce ADAR1 or ADAR2 (Maas et al. 2001). In cells with the highest ADAR1 intracellular activity, 35% of endogenous pri-miRNA22 transcripts are edited at one or more positions (nt −30, −1,+1,+2,+15,+18) within the precursor sequence (see Fig. 1B,C). Interestingly, ADAR2 hyperactivity yields a moderately increased rate of editing (~10%; nucleotide positions +1, +2), but with high site selectivity for the +1 position (7%). Taken together, these results show that overexpression of both ADAR enzymes leads to an increase in editing levels at sites that are also subject to editing in mammalian tissues; however, ADAR1 overexpression results in a pattern of modification that includes bases that do not get modified under physiological conditions. In other cases of nuclear RNA editing, ADAR1 and 2 also show overlapping but different site specificities and often additional adenosines are modified in conjunction with ADAR hyperactivity (Bass 2002). In brains from ADAR2 knock-out mice (Higuchi et al. 2000) pri-miRNA22 editing is still detectable at position +1, indicating that the endogenous ADAR1 editing enzymes can target this position (Fig. 1C).
The current data set does not clearly point to the physiological role of miRNA22 editing because of the low levels of editing observed in adult human and mouse tissues and because miRNA22 function is unknown. The editing events might simply represent “background” activity of the editing machinery targeting this miRNA stem–loop without a significant impact on its function. However, considering the transient and regulatory role of characterized miRNAs (Bartel 2004) and the regulatory function of RNA editing, a specific time point or place may exist where editing of pre-miRNA22 is transiently or locally induced to much higher levels. Recently several candidate target genes for miRNA22 have been identified in computational studies (Lewis et al. 2003) and their functional characterization might lead to insights into the role of miRNA22 as well as the biological consequences of its post-transcriptional modification by ADARs.
The editing of miRNA precursors by ADARs could have major implications for miRNA analysis, biogenesis, and function. When cloning mature miRNAs, editing could result in misidentification, and miRNA expression analysis by Northern hybridization could yield incorrect results due to edited RNA/probe mismatches. It is well known that point mutations within the miRNA precursor can strongly influence processing activity by dicer (Zeng and Cullen 2003), and the −1 and +1 positions, both subject to editing in pri-miRNA22, are positioned right around the cleavage site. Therefore, A-to-I modification of miRNA precursors could modulate processing kinetics and outcome, or even prevent further maturation altogether. Importantly, editing at both positions +1 and +2 is predicted to influence miRNA strand choice (Khvorova et al. 2003; Schwarz et al. 2003) because they would weaken the 5′ end of the pre-miRNA22 dicer product.
Finally, a single A-to-I difference within a 20–22mer could redirect a miRNA to a different target sequence. Alternatively, editing of miRNAs might fine tune miRNA function by modulating target sequence interactions. Hopefully, the discovery of miRNA precursor editing will encourage additional studies regarding other posttranscriptional mechanisms for regulation of miRNA function, such as the alternative splicing of precursor molecules, which we also observe for miRNA22. Like the recently proposed regulation of miRNAs by other miRNAs (Lai et al. 2004), A-to-I RNA editing might provide another layer of regulatory control within the complex network of RNA-mediated gene function.
Acknowledgments
We thank Alekos Athanasiadis for helpful discussions and Miyoko Higuchi and Peter H. Seeburg for brain cDNA from ADAR2 knock-out mice. This work was supported in part by the 2020 Bioscience and Biotechnology Initiative at Lehigh University.
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 USC section 1734 solely to indicate this fact.
Article and publication are at http://www.rnajournal.org/cgi/doi/10.1261/rna.7350304.
REFERENCES
- Bartel, D.P. 2004. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 116: 281–297. [DOI] [PubMed] [Google Scholar]
- Bass, B.L. 2002. RNA editing by adenosine deaminases that act on RNA. Annu. Rev. Biochem. 71: 817–846. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernstein, E., Caudy, A.A., Hammond, S.M., and Hannon, G.J. 2001. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409: 363–366. [DOI] [PubMed] [Google Scholar]
- Boguski, M.S., Lowe, T.M., and Tolstoshev, C.M. 1993. dbEST—Database for “expressed sequence tags.” Nat. Genet. 4: 332–333. [DOI] [PubMed] [Google Scholar]
- Burns, C.M., Chu, H., Rueter, S.M., Hutchinson, L.K., Canton, H., Sanders-Bush, E., and Emeson, R.B. 1997. Regulation of serotonin-2C receptor G-protein coupling by RNA editing [see comments]. Nature 387: 303–308. [DOI] [PubMed] [Google Scholar]
- Cline, J., Braman, J.C., and Hogrefe, H.H. 1996. PCR fidelity of pfu DNA polymerase and other thermostable DNA polymerases. Nucleic Acids Res. 24: 3546–3551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldstrohm, A.C., Greenleaf, A.L., and Garcia-Blanco, M.A. 2001. Co-transcriptional splicing of pre-messenger RNAs: Considerations for the mechanism of alternative splicing. Gene 277: 31–47. [DOI] [PubMed] [Google Scholar]
- Higuchi, M., Maas, S., Single, F.N., Hartner, J., Rozov, A., Burnashev, N., Feldmeyer, D., Sprengel, R., and Seeburg, P.H. 2000. Point mutation in an AMPA receptor gene rescues lethality in mice deficient in the RNA-editing enzyme ADAR2. Nature 406: 78–81. [DOI] [PubMed] [Google Scholar]
- Khvorova, A., Reynolds, A., and Jayasena, S.D. 2003. Functional siRNAs and miRNAs exhibit strand bias. Cell 115: 209–216. [DOI] [PubMed] [Google Scholar]
- Lagos-Quintana, M., Rauhut, R., Lendeckel, W., and Tuschl, T. 2001. Identification of novel genes coding for small expressed RNAs. Science 294: 853–858. [DOI] [PubMed] [Google Scholar]
- Lagos-Quintana, M., Rauhut, R., Yalcin, A., Meyer, J., Lendeckel, W., and Tuschl, T. 2002. Identification of tissue-specific microRNAs from mouse. Curr. Biol. 12: 735–739. [DOI] [PubMed] [Google Scholar]
- Lai, E.C. 2003. microRNAs: Runts of the genome assert themselves. Curr. Biol. 13: R925–936. [DOI] [PubMed] [Google Scholar]
- Lai, E.C., Wiel, C., and Rubin, G.M. 2004. Complementary miRNA pairs suggest a regulatory role for miRNA:miRNA duplexes. RNA 10: 171–175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee, Y., Ahn, C., Han, J., Choi, H., Kim, J., Yim, J., Lee, J., Provost, P., Radmark, O., Kim, S., et al. 2003. The nuclear RNase III Drosha initiates microRNA processing. Nature 425: 415–419. [DOI] [PubMed] [Google Scholar]
- Lewis, B.P., Shih, I.H., Jones-Rhoades, M.W., Bartel, D.P., and Burge, C.B. 2003. Prediction of mammalian microRNA targets. Cell 115: 787–798. [DOI] [PubMed] [Google Scholar]
- Maas, S., Patt, S., Schrey, M., and Rich, A. 2001. Underediting of glutamate receptor GluR-B mRNA in malignant gliomas. Proc. Natl. Acad. Sci. 98: 14687–14692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maas, S., Rich, A., and Nishikura, K. 2003. A-to-I RNA editing: Recent news and residual mysteries. J. Biol. Chem. 278: 1391–1394. [DOI] [PubMed] [Google Scholar]
- Morse, D.P., Aruscavage, P.J., and Bass, B.L. 2002. RNA hairpins in noncoding regions of human brain and Caenorhabditis elegans mRNA are edited by adenosine deaminases that act on RNA. Proc. Natl. Acad. Sci. 99: 7906–7911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Potter, J., Zheng, W., and Lee, J. 2003. Thermal stability and cDNA synthesis capability of SuperScript reverse transcriptase. Focus (In-vitrogen) 25: 19–14. [Google Scholar]
- Rueter, S.M., Dawson, T.R., and Emeson, R.B. 1999. Regulation of alternative splicing by RNA editing. Nature 399: 75–80. [DOI] [PubMed] [Google Scholar]
- Schmauss, C. and Howe, J.R. 2002. RNA editing of neurotransmitter receptors in the mammalian brain. Sci. STKE 2002: PE26. [DOI] [PubMed] [Google Scholar]
- Schwarz, D.S., Hutvagner, G., Du, T., Xu, Z., Aronin, N., and Zamore, P.D. 2003. Asymmetry in the assembly of the RNAi enzyme complex. Cell 115: 199–208. [DOI] [PubMed] [Google Scholar]
- Seeburg, P.H. and Hartner, J. 2003. Regulation of ion channel/neuro-transmitter receptor function by RNA editing. Curr. Opin. Neuro-biol. 13: 279–283. [DOI] [PubMed] [Google Scholar]
- Seeburg, P.H., Higuchi, M., and Sprengel, R. 1998. RNA editing of brain glutamate receptor channels: Mechanism and physiology. Brain Res. Brain Res. Rev. 26: 217–229. [DOI] [PubMed] [Google Scholar]
- Zeng, Y. and Cullen, B.R. 2003. Sequence requirements for micro RNA processing and function in human cells. RNA 9: 112–123. [DOI] [PMC free article] [PubMed] [Google Scholar]