Abstract
Vitamin B12 (cobalamin) is an essential cofactor of two enzymes, methionine synthase and methylmalonyl-CoA mutase. The conversion of the vitamin to its coenzymes requires a series of biochemical modifications for which several genetic diseases are known, comprising eight complementation groups (cblA through cblH). The objective of this study was to clone the gene responsible for the cblA complementation group thought to represent a mitochondrial cobalamin reductase. Examination of bacterial operons containing genes in close proximity to the gene for methylmalonyl-CoA mutase and searching for orthologous sequences in the human genome yielded potential candidates. A candidate gene was evaluated for deleterious mutations in cblA patient cell lines, which revealed a 4-bp deletion in three cell lines, as well as an 8-bp insertion and point mutations causing a stop codon and an amino acid substitution. These data confirm that the identified gene, MMAA, corresponds to the cblA complementation group. It is located on chromosome 4q31.1–2 and encodes a predicted protein of 418 aa. A Northern blot revealed RNA species of 1.4, 2.6, and 5.5 kb predominating in liver and skeletal muscle. The deduced amino acid sequence reveals a domain structure, which belongs to the AAA ATPase superfamily that encompasses a wide variety of proteins including ATP-binding cassette transporter accessory proteins that bind ATP and GTP. We speculate that we have identified a component of a transporter or an accessory protein that is involved in the translocation of vitamin B12 into mitochondria.
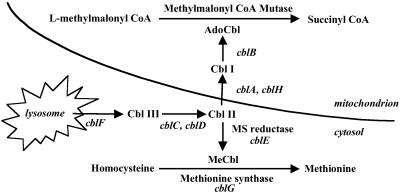
Vitamin B12 (cobalamin, Cbl) is one of the most complex vitamins found in nature (1). It acts as a coenzyme in variety of molecular rearrangements to generate oxidizable substrates. In mammals, Cbl is involved in two enzyme systems: methionine synthase (MS), a key enzyme in one carbon metabolism, and methylmalonyl-CoA mutase (MCM), an enzyme in the pathway of branched chain amino acid catabolism (Fig. 1). The conversion of the vitamin to its coenzyme forms requires a series of processing steps in cells. Following entry of Cbl into the lysosome as a complex with transcobalamin II, it is released by proteolytic digestion and transported into the cytosol across the lysosomal membrane. The free vitamin is reduced from cob(III)alamin to cob(II)alamin and cob(I)alamin, upon which it can be methylated to form MeCbl or adenosylated to AdoCbl. The reductive methylation of Cbl occurs on the MS protein and is catalyzed by MS reductase. The reactions leading to the synthesis of AdoCbl are less well understood. MCM is a mitochondrial enzyme, and the final steps of AdoCbl synthesis occur in the mitochondrion. The current view is that the first reductive step, cob(III)alamin to cob(II)alamin, occurs in the cytosol. Subsequently, Cbl, probably as cob(II)alamin, is transported into the mitochondrion, where the next reduction step, to cob(I)alamin, occurs followed by addition of the adenosyl group to form AdoCbl (2).
Fig 1.
Intracellular processing of Cbl (modified from ref. 2). Steps are identified according to complementation group names. Their gene (uppercase italics) and predicted functions are: cblA (MMAA), this study; cblB, cob(I)alamin adenosyltransferase; cblC/D, possibly cob(III)alamin reductase; cblE (MTRR), methionine synthase reductase; cblF, lysosomal Cbl efflux; cblG (MTR), methionine synthase; and cblH, possibly a component of Cbl transport or cob(II)alamin reductase.
Blocks in any of the Cbl transport or modification steps usually cause severe disease in infancy or early childhood, although they may also present later in life (2). Depending on the nature of the metabolic block, affected patients may present with homocystinuria or methylmalonic aciduria or both. The combined defects are due to deficits in early steps of Cbl processing, whereas deficient activity of only MS or MCM indicates a block at a later step of Cbl metabolism or a primary defect of the apoenzyme. Eight complementation groups, cblA to cblH, have been identified that account for these disorders (Fig. 1). Two of the corresponding genes have been cloned: MTRR encoding MS reductase (cblE group; ref. 3) and MTR encoding MS (cblG group; refs. 4–6). Of the remaining complementation groups, three, cblA, cblB, and cblH, are uniquely involved in AdoCbl synthesis, whereas the remainder, cblC, cblD, and cblF, affect both MeCbl and AdoCbl synthesis (2).
Our aim was to identify the gene responsible for the cblA complementation group (7), a disorder that classically is associated with vitamin B12-responsive methylmalonic aciduria (8). The biochemical features of this disorder have been described (9–13). Fibroblast cultures from cblA patients typically have decreased propionate incorporation into cellular protein and reduced synthesis of AdoCbl, whereas MeCbl synthesis remains normal (13). In contrast, extracts from the same fibroblast cultures are able to synthesize AdoCbl in the presence of chemical reducing agents, such as DTT (and ATP) (10–13). It has been proposed that cblA corresponds to a defect of a mitochondrial, NADPH-dependent aquacobalamin reductase (13, 14), or that it may have a block in mitochondrial binding or transport of Cbl (9, 10, 12).
We have identified the gene responsible for the cblA defect through examination of bacterial gene clusters by using the National Center for Biotechnology Information (NCBI) utility, Clusters of Orthologous Groups of proteins (www.ncbi.nlm.nih.gov/COG). It was detected as one of the potential genes in an operon containing MCM. Several mutations, including two frameshift mutations, were identified in cblA patients, confirming the authenticity of the gene identification. Based on its predicted domain structure, we hypothesize that we have identified a component of a transporter required for the translocation of Cbl into mitochondria. We propose the gene name, MMAA, to denote methylmalonic acidemia linked to the cblA complementation group.
Materials and Methods
Patient Cell Lines.
Fibroblast lines from six patients classified as members of the cblA complementation group were used in this study. Patient cell line WG1192 was reported to synthesize AdoCbl in vitro (11). Propionate uptake was 1.26 ± 0.10 nmol/mg protein/18 h without hydroxylcobalamin (OHCbl) added to medium (13) compared to control values of 10.8 ± 3.2 nmol/mg protein/18 h. After incubation of cells in 25 pg/ml [57Co]cyanocobalamin for 4 days, AdoCbl was at 5.0% of total Cbl as compared to normal levels of 15.29 ± 4.2%. WG1776 is from a white female who presented at 7 mo with acidosis, ketosis, and lethargy. [14C]propionate uptake was 2.3 nmol/mg protein/18 h without and 4.6 nmol/mg protein/18 h with OHCbl in the culture medium (normal with OHCbl added, 10.9 ± 3.5 nmol/mg protein/18 h). AdoCbl was at 6.6% after [57Co]cyanocobalamin incubation. WG1943 is from a 2-mo-old East Indian male with vomiting, lethargy, and Cbl-responsive methylmalonic acidemia. [14C]propionate uptake was 3.4 nmol/mg protein/18 h without and 9.0 nmol/mg protein/18 h with OHCbl in the culture medium. AdoCbl was at 6.8%. WG2014 is from a 3-mo-old white male who presented with fever, projectile vomiting, dehydration, generalized fine tremors, and Cbl-responsive methylmalonic acidemia. [14C]propionate uptake was 1.2 nmol/mg protein/18 h without and 7.1 nmol/mg protein/18 h with OHCbl in the culture medium. AdoCbl was at 7.5% (15). WG3080 is from a white female who presented at 7 days with lethargy, poor feeding, and seizures. The [14C]propionate uptake was 0.6 nmol/mg protein/18 h without and 2.4 nmol/mg protein/18 h with OHCbl in the culture medium. AdoCbl was at 3.5%. There is no known consanguinity associated with any of the patient cell lines.
DNA and RNA Extraction and PCR Amplification.
Genomic DNA was isolated as described (16) or by using Genomic-tip (Qiagen) as described in the manufacturer's protocol. PCR reactions were performed on genomic DNA with primers designed to amplify exons of the coding sequence and portions of the flanking introns (Table 1). Total cellular RNA was purified from a human liver hybridoma cell line (Huh-1) by using Trizol (Invitrogen). The total cDNA was amplified by using oligo-dT as described by the Omniscript protocol (Qiagen), followed by PCR amplification with RNA-specific primers (Table 1). The PCR products were purified with the QIAquick PCR purification kit (Qiagen) and cloned into pGEM (Promega) according to the supplier's instructions.
Table 1.
Splice junctions in the MMAA gene and primers used for exon/flanking sequence amplifications
| Exon |
Size, bp |
PCR, bp |
Sense
primer sequence |
Antisense primer
sequence |
Splice sites | Exon boundaries |
|
|---|---|---|---|---|---|---|---|
| 5′ | 3′ | ||||||
| 1 | ≥19 | ND | ND | ND | … GCGGgtg | ≥−85 to −68 | |
| 2 | 506 | 515 | tagggaggtcacaatcacattgagc | agactgacCTACTCGAAATGCTAGTG | tagGGAG… | … GTAGgtc | −67 to 439 |
| 3 | 123 | 397 | tctctttccaccgtagGATTGTCTG | gatcatcttcaacttctggcacctg | tagGATT… | … GGTGgta | 440–562 |
| 4 | 171 | 507 | ggaactggctgataattgacccgta | cctaatgtttacagttctcctctggc | tagGATC… | … GTTGgtg | 563–733 |
| 5 | 87 | 470 | ggttgactgtgtgaccatgagtatg | gctggagcgatccatgtatttcagt | cagGTGT… | … GCAGgta | 734–820 |
| 6 | 149 | 586 | ctgccactcaatcatgttgtgacttg | tcatgactacacactccgtctttgtc | tagGGTA… | … AAAGgta | 821–969 |
| 7 | 352 | 527 | taagaattaactggcaggtatcagc | ctaacttccttcaagttttcaagcatg | tagGTAA… | 970–1317 | |
| RNA | 1,359 | cttcgggcggggaggtcacaatcaca | gtaaaattattatacaggatgaa | ||||
Splice sites were determined based on a comparison of the cDNA and genomic sequences. The coding sequences are uppercase, whereas intron and noncoding sequences are lowercase. ND, not done.
The full-length coding sequence was amplified by RT-PCR from position −74 to 1285, using the “RNA” primers.
Heteroduplex Analysis and DNA Sequencing.
Initial examination of PCR products was performed by heteroduplex analysis, as described (17). Heteroduplexes were obtained by mixing, melting, and reannealing the PCR products of mutant and control DNA before electrophoresis. Positive cell lines also underwent the same procedure without mixing with control to determine whether the mutation was on one or both alleles. The observation of a heteroduplex, with implication of a multibase mutation, was taken as indication that the correct gene was identified. DNA sequencing was accomplished in 96-well plates in 10-μl reactions made up of 2 μl of purified PCR product and 1 μl of BigDye Terminator Cycle Sequencing Version 2.0 (Applied Biosystems) and analyzed with an ABI 3700 sequencer (Applied Biosystems).
Northern Blot.
A multiple human tissue Northern blot (CLONTECH) was used for determining the level and size of mRNA species. A probe was generated from a 508-bp cDNA segment amplifying regions 380–887 bp of the full-length cDNA, using the Huh-1 cell line as the source of RNA. Detection of β-actin mRNA was used as control. All procedures followed the protocol supplied by CLONTECH.
Results
Identification of the MMAA cDNA.
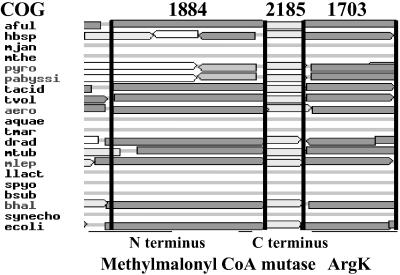
Initial efforts to identify sequences in model genomes that might represent an NAD(P)H and flavin-dependent aquacobalamin reductase were unsuccessful, owing to the excessive number of potential candidates. However, with the availability of the COG database, it became possible to narrow the search to genes in microbial organisms that were arranged in clusters with genes encoding Cbl-dependent enzymes. A search for MCM yielded COG2185, identified as corresponding to the Cbl-binding, carboxyl-terminal domain of MCM. It is present as a member of a cluster in 16 of the 43 complete genomes represented in the database (Fig. 2). In 10 of these, COG1703 was found immediately adjacent to COG2185. COG1703 is characterized as “putative periplasmic protein kinase ArgK and related GTPases of G3E family.” COG1703 was found in the genomes of Archaea (five genera), Bacteria (five genera), and Eukaryota (one genus), as listed in the COG database. One member of this COG, namely ArgK, has been described as a coupled ATPase of the lysine-arginine-ornithine (LAO) transport system in E. coli (18). Given the proximity of COG1703 to MCM and its frequency in microbial genomes, we pursued it as a candidate for involvement in the AdoCbl pathway.
Fig 2.
Cluster of orthologous genes examining the MCM N terminus (COG1884) and C terminus (COG2185) and the genes identified as comprising COG1703. Bacteria discussed in this study include Archaeoglobus fulgidus (aful) and Escherichia coli K12 (ecol); others are listed at www.ncbi.nlm.nih.gov/COG.
The A. fulgidus sequence (GenBank accession no. AAB89957) from COG1703 was used as the template for a blast (tblastn) search of the human Expressed Sequence Tags (dbest) database and Unigene clusters at NCBI. This search yielded 10 ESTs listed in Unigene cluster Hs.126216 and 18 others in Hs.21017. Alignment of the ESTs permitted construction of a composite 1,404-bp-long cDNA that contains a putative start codon with in-frame upstream stop codon at the 5′ end and polyA tail at the 3′ end (Figs. 3 and 4).
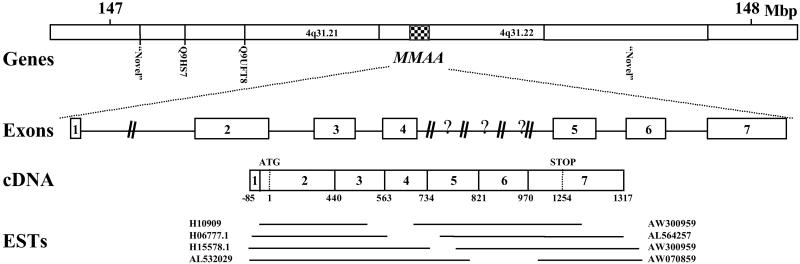
Fig 3.
Structural organization of the human MMAA gene. The top schematic (“Genes”) illustrates the flanking genes and map positions surrounding MMAA according to contigview in Ensembl (www.ensembl.org). Note that NCBI maps MMAA to the 147.36–147.38 megabase pairs (Mbp) location of chromosome 4. Both Ensembl and NCBI localize MMAA to 4q31.21–22. The next line (“Exons”) illustrates the location of exons (numbered boxes) and introns (lines) along the MMAA gene located in Contig NT_016606.11. Our independently confirmed sequences from genomic DNA were deposited in GenBank under the accession nos. AF524841–AF524846. Introns are drawn to scale, except where “?” is used to denote a region of unknown sequence and length and “//” is used to denote a longer region than illustrated. Next the cDNA (“cDNA”) is illustrated, with splice and start/stop codons indicated, showing that it was constructed from the numerous ESTs (“ESTs”) belonging to Unigene clusters Hs.126261 (10 ESTs) and Hs.21017 (18 ESTs) and verified by RT-PCR sequencing. Only some ESTs are illustrated here.
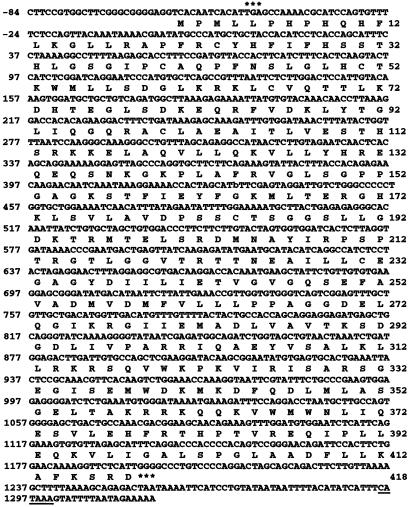
Fig 4.
Nucleotide and deduced amino acid sequence of human MMAA. The nucleotide and amino acid residues are numbered on the left and right margins, respectively. Two in-frame stop codons are indicated by *** immediately upstream and downstream of the coding sequence. A candidate polyA signal starting at 1,296 is underlined.
The 5′ and 3′ ends of the composite human cDNA were extended further by using a computer-generated version of EST walking. Sequential searches were made by using 5′ or 3′ most sequences from successively identified ESTs until no further extensions were identified (Fig. 3). At the 5′ end, the human EST, AL532029 from Hs.21017, was used to blast the EST database. It identified H10909 as having an additional nine nucleotides at the 5′ end. H10909, in turn, was used as the subject for blast in the next round. Three more ESTs were found, H06777, Z44188, and H15578, with 14, 15, and 15 nucleotides of additional sequence, respectively. No further 5′ additions were detected. At the 3′ end, seven of the 10 ESTs from Hs.21017 contained polyA tails, all starting at the same location. No alternate 3′ polyA sites were identified.
The cDNA sequence was verified experimentally by RT-PCR and sequencing of mRNA from the transformed human liver cell line Huh-1. It contains an ORF 1,254 bp in length, predicting a polypeptide of 418 aa with a calculated molecular weight of 46.5 kDa (Fig. 4). During the course of this work, the deduced protein sequence (XP 094094) was placed in GenBank as a sequence from contig NT_006177 and agrees with our construction. Based on all ESTs examined in the database, no splice variants appear to be present. Examination of the database of single nucleotide polymorphisms (dbSNP) revealed a SNP at c.1089.C → G that would result in an amino acid change, L363V. This change was not observed in any of the patient or seventy control cell lines that were sequenced experimentally, suggesting that it is a rare mutation.
Orthologous Proteins.
Sequences corresponding to the candidate MMAA cDNA were also identified in the mouse EST database. This allowed us to verify reading frame throughout, and to assure the likelihood that the sequence is full length. The mouse Unigene cluster Mm.26510 contains 54 ESTs. These were assembled into a composite cDNA sequence that verified the corresponding GenBank entry (BC021954). The mouse cDNA is 76.4% identical to the human cDNA. The corresponding proteins are 78.8% identical, with similarity based on conserved amino acid properties of 84.5% (Fig. 5).
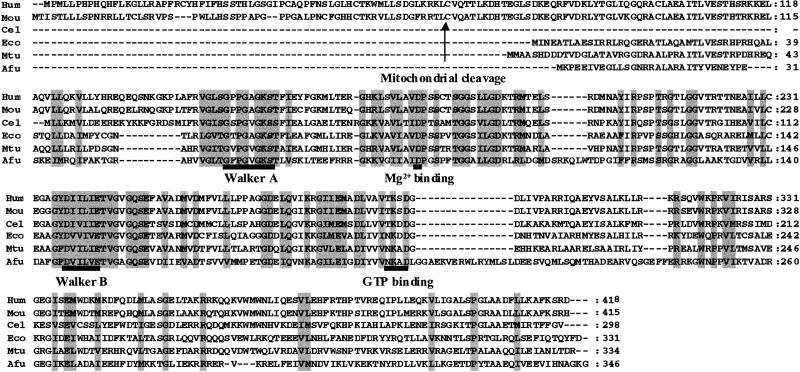
Fig 5.
Alignment of amino acid sequences similar to that of human (Hum) MMAA, including mouse (Mou), Caenorhabditis elegans (Cel), E. coli (Eco), Mycobacterium tuberculosis (Mtu), and A. fulgidus (Afu). Sequence motifs defining the G3E family of proteins are indicated, including the GxxGxGK[S/T] Walker A, the Mg2+-binding aspartate residue, the DHbHbHbHbE Walker B motif with Hb denoting hydrophobic residues, and an [N/T]KxD GTP-binding motif, as described by Leipe et al. (27). The arrow denotes the predicted mitochondrial leader sequence cleavage site. Amino acids that are similar in all species are shaded with a gray background.
In addition to the mouse sequence and the sequences identified in the COG database, a survey of the NCBI database for additional orthologous protein sequences by using blast analysis (19) yielded a predicted protein in C. elegans (NP 495466) and M. tuberculosis (NP 335996) and an LAO/AO transport system kinase (ArgK) in E. coli (P27254). These sequences were also aligned with the proposed human-coding sequence (Fig. 5).
Structure of the MMAA Gene and Chromosomal Location.
The MMAA gene was localized to chromosome 4q31.1–2 based on the linkage of contig NT_006177.5 to locus LOC166785 (Fig. 3) and the mapping of sequence-tagged sites (STS) including WI-14208, A005L23, and STSG2679. To define the structure of the human gene, the cDNA sequence was compared to the whole human genome by using blastn. Seven hits were obtained with 100% continuous alignment in each segment, indicating that the gene contains seven exons (Fig. 3). Based on comparison of the assembled EST and genomic sequence, only 19 bp were found to constitute the first exon. No recognizable CAP site suggesting its potential length was identified. However, because an in-frame stop codon precedes the first in-frame ATG in exon 2, the first exon was concluded to be untranslated and was not included in mutation analysis (below). All exon–intron boundaries conserve the AG-GT rule and maintain the ORF as predicted by the cDNA sequence. The exons range in size from 89 bp (excluding exon 1) to 586 bp. Four introns range in size from 1,003 bp to 3,502 bp, and intron 3 is estimated to be ≈21,600 bp. In total, the human MMAA gene was found to encompass ≈17.1 kb and is flanked by an Ensembl (www.ensembl.org)-predicted gene (Q9UFT8) having homology to the mothers against decapentaplegic 1 (MAD1) gene on the 5′-side ≈100 kb away and by a novel gene (“novel”) with a zinc finger domain ≈160 kb away on the 3′-side of the human gene map (using the contig viewer utility at Ensembl).
Comparison of the human and mouse genes showed preservation of all seven exon–intron splice junctions. Interestingly, the mouse and human chromosomes show a large amount of synteny around the MMAA gene including the MAD1 homolog and zinc finger protein. This gene pattern is in agreement with the human–mouse homology map (www.ncbi.nlm.nih.gov) that links human 4q31.3–4q35 with chromosome 8 of mouse.
Expression in Tissues.
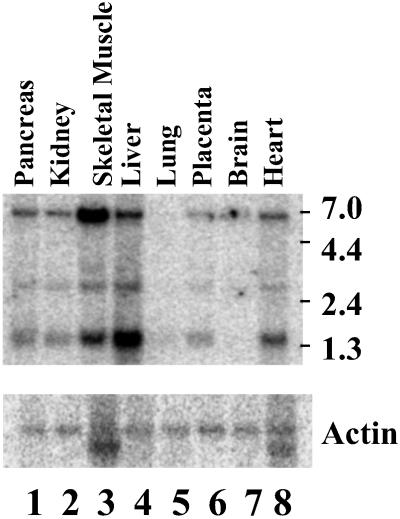
The 508-bp probe hybridized to three RNA species of ≈1.4, 2.6, and 5.5 kb on the Northern blot, with highest levels of expression in liver and skeletal muscle. Skeletal muscle expressed the largest transcript predominantly, whereas in liver the smallest one predominated (Fig. 6). The 1.4-kb species is essentially identical to the predicted length of the composite cDNA.
Fig 6.
Northern blot analysis of MMAA mRNA. A Northern blot of poly(A)+ RNA from the indicated human tissues. The membranes were hybridized with a 32P-labeled 508-bp probe from MMAA mRNA (A) or a β-actin probe (B). Molecular size marker positions are indicated on the right.
Mutations in the cblA Complementation Group.
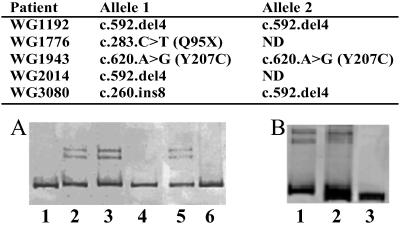
Fibroblast lines from patients from the cblA complementation group were analyzed for mutations by PCR-heteroduplex analysis and DNA sequencing. Three cell lines, WG1192, WG2014, and WG3080, gave heteroduplexes on PAGE of PCR products obtained by amplification of exon 4 and flanking sequences (Fig. 7). The latter two cell lines showed heteroduplexes in the presence or absence of mixing with control DNA. These results suggest that WG1192 is homoallelic, and that WG2014 and WG3080 are heteroallelic for a multibase change within the amplified sequence. DNA sequencing revealed the mutation to be a 4-bp deletion, c.592.delACTG, which would disrupt reading frame and introduce a downstream stop codon in the transcribed sequence. WG3080 produced a second heteroduplex on PCR of exon 2 (Fig. 7). DNA sequencing revealed this change to be an 8-bp insertion, c.260.insATAAACTT. It would also result in a severely truncated protein. Given the presence of two different frameshift mutations on each allele of WG3080 and the 4-bp deletion on both alleles of WG1192, both cell lines would be null for a functional protein.
Fig 7.
Mutations reported here and heteroduplex tests. (A) Heteroduplex test for c.592.delACTG: lane 1, control cells; lane 2, WG2014 mixed with control cells in a 1:1 ratio; lane 3, WG2014 alone; lane 4, WG1192 alone; lane 5, WG1192 mixed with control; lane 6, WG1943 mixed with control cells in a 1:1 ratio. (B) Heteroduplex for c.259.delATAAACTT: lane 1, WG3080 alone; lane 2, WG3080 and control cells; lane 3, control cells alone.
Two point mutations were also detected. Cell line WG1776 contains a premature stop codon, c.283.C>T (Q94X), on one allele, which would also disrupt protein expression. The second allele is unidentified. WG1943 contains a mutation, c.620.G>A, resulting in the amino acid substitution, Y207C, which was not identified in 70 control cell lines sequenced. This mutation is present on both alleles and suggests inactivation or instability of the resulting protein; although expression of the mutant protein would be required to clarify the effect of this substitution.
Discussion
We have combined the analysis of prokaryotic gene arrangements with the identification of mutations in affected patients to determine the identity of the disease-causing gene in patients from the cblA complementation group. The key finding confirming the identity of the MMAA gene was the detection of heteroduplexes harboring mutations in the candidate gene for five cblA cell lines. DNA sequencing revealed a 4-bp deletion and an 8-bp insertion in patient cell lines that would be expected to produce frameshifts and truncated proteins. Significantly, the 4-bp deletion was shown to be homoallelic in one cell line, whereas another was heteroallelic for both the deletion and insertion. These results demonstrate that in at least two patients, the mutations on both alleles are of such severity that the expected outcome is the complete absence of protein function. Based on the nature of mutations identified in these patients, we conclude that the MMAA gene is responsible for the cblA complementation group. Given the similarity of the cblH to cblA complementation groups, we also tested a cblH cell line for mutations, but found none, indicating that cblH likely corresponds to a different gene.
It was anticipated that the MMAA gene would code for a Cbl reductase or perhaps a mitochondrial transporter of vitamin B12. Early studies pointed to a mitochondrial defect because the reduction in AdoCbl synthesis was localized to mitochondria, and the activity was restored if the assay was conducted in cell extracts rather than whole cells (10). These findings are compatible with a block in mitochondrial transport, but no such transporter was identified (20, 21). Instead, Watanabe and coworkers (22, 23) identified an NADPH-dependent aquacobalamin reductase in rat liver and Euglena mitochondria, which was deficient in fibroblasts from one cblA cell line (14). They took this observation as evidence that the MMAA gene might code for a reductase. Our results are consistent with mitochondrial targeting of the MMAA protein. It has a predicted leader sequence and signal cleavage site (P = 0.998; www.inra.fr/predotar; Fig. 5), but it is unlikely to be an aquacobalamin reductase because it does not contain domains suggestive of flavin or NAD(P)H-binding sites.
In considering a possible role for the MMAA gene, we first examined the published work on ArgK, the E. coli protein that is 61% similar to MMAA. The argK gene was initially identified as the site of mutations resulting in a deficiency in uptake of lysine (L), ornithine (O), and arginine (A) (18). The affected gene, argK, was expressed and shown to code for a protein that has intrinsic ATPase activity and could phosphorylate the LAO and AO periplasmic binding proteins in vitro (18). However, a revertant of the original mutant had restored ATPase activity and LAO uptake but remained defective in the protein kinase activity. It was concluded that ArgK is an essential, common accessory to the LAO and AO transport systems through its ATPase activity, but that its putative kinase activity was not involved (18). The LAO/AO and arginine periplasmic binding proteins have been shown to be associated with two independent inner membrane transport systems, HisP and ArtP. Both are ATPases with ATP-binding cassette signatures (24, 25). This challenges the notion that ArgK, which lacks an ATP-binding cassette signature, is an LAO/AO transport-associated ATPase.
Therefore, with the possibility that ArgK might be involved with vitamin B12 transport in microbes, we reviewed the components of the vitamin B12 transport system in E. coli (26). Vitamin B12 is transported across the inner membrane by two identical membrane-spanning components, BtuC, and an associated ATPase, BtuD. A comparison of the sequence of BtuD with that of ArgK shows them to be distinct and related only in the Walker A motif, commonly found in ATP-binding proteins. An extensive phylogenetic analysis of nucleotide-binding proteins included ArgK as a member, proposing that it belongs to one of four subfamilies comprising the G3E family of P-loop GTPases (27). The G3E branch of the phylogenic tree is defined by the specific sequence signatures associated with nucleotide binding, including Walker A and Walker B motifs, a Mg2+-binding aspartate residue and a sequence specific to GTP defined by an [N/T]KxD motif (Fig. 5). In addition to ArgK, the G3E family comprises three additional subfamilies represented by: UreG, an accessory urease subunit involved in the assembly of the metallocentre of nickel metalloenzymes; HypB, a hydrogenase expression protein that is also involved in metal binding and metalloenzyme assembly; and, interestingly, CobW, a protein in the biosynthetic pathway of vitamin B12 whose role remains undefined. It is significant that all of these proteins are involved in the assembly of metalloenzymes which, conceptually, reinforces the idea of a similar role for ArgK in Cbl metabolism. Nonetheless, although the proximity of argK to the MCM gene in bacterial gene clusters has been recognized previously, efforts to identify a role for the ArgK protein in this area of metabolism have been unsuccessful (28, 29).
In sum, the studies on ArgK do not readily point to a function for the protein and offer only limited insight into predicted function of the MMAA protein. Nevertheless, our knowledge of the biochemical abnormalities of cblA patients strongly suggests that MMAA and, by extension, ArgK have specific involvement in vitamin B12 utilization. An interesting aspect of previous biochemical studies included the finding that vitamin B12 uptake in isolated rat liver mitochondria depended on mitochondrial swelling mediated by calcium or phosphate addition (11, 30). This appears to occur through opening of the mitochondrial permeability transition pore (MPTP) that is permeable to molecules of <1,500 Da, a value bordering on the molecular mass of vitamin B12 (1,350 Da) (31). Acute opening of the MPTP causes mitochondrial swelling and apoptosis and can be initiated by the addition of calcium or phosphate salts (32). We speculate that vitamin B12 uptake might occur via MPTP pores, opened within a physiological range, which might be stimulated by the ATP or GTPase activity of the associated MMAA protein. A search for transmembrane domains was equivocal for human MMAA, and negative for MMAA homologs in other species. If MMAA is an accessory protein, it would likely be on the matrix side of the MPTP. As a mechanism, this conjecture would obviate the need for a unique transport protein, for which there seems to be little evidence, but would offer the possibility of a requirement for calcium/phosphate stimulation and ATP dependence as originally detected (11). Further, if the system is inherently leaky so that there is a low level of transport in the absence of activation, then this might provide an explanation for the vitamin B12 responsiveness in these patients (8). In conclusion, we propose a role for MMAA in the transport of vitamin B12 into the mitochondrion, either as a component of a transport system or as a soluble accessory protein required for translocation across a nonspecific membrane pore, possibly the MPTP.
Acknowledgments
We acknowledge the contributions of Pierre Lepage (Genome Quebec Innovation Centre, Montreal) for aiding in sequencing, Nora Matiaszuk and Gail Dunbar (Royal Victoria Hospital, Montreal) for growing cell cultures, and David Watkins (Royal Victoria Hospital) for helpful discussions. These studies were supported by grants from the Canadian Institute for Health Research, the National Institutes of Health–National Heart and Lung Institute, and the March of Dimes. Scholarship support to C.M.D. was provided by the Canadian Institute for Health Research and the Alberta Heritage Foundation for Medical Research.
Abbreviations
MCM, methylmalonyl-CoA mutase
Cbl, cobalamin
AdoCbl, adenosylcobalamin
OHCbl, hydroxylcobalamin
MS, methionine synthase
LAO, lysine-arginine-ornithine
MPTP, mitochondrial permeability transition pore
MMAA, methylmalonic acidemia linked to the cblA complementation group
References
- 1.Raux E., Schubert, H. L. & Warren, M. J. (2000) Cell. Mol. Life Sci. 57 1880-1893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Rosenblatt D. S. & Fenton, W. A. (2001) in The Metabolic and Molecular Bases of Inherited Disease, eds. Scriver, C. R., Beaudet, A. L., Valle, D. & Sly, W. S. (McGraw–Hill, New York), pp. 3897–3933.
- 3.Leclerc D., Odievre, M., Wu, Q., Wilson, A., Huizenga, J. J., Rozen, R., Scherer, S. W. & Gravel, R. A. (1999) Gene 240 75-88. [DOI] [PubMed] [Google Scholar]
- 4.Leclerc D., Campeau, E., Goyette, P., Adjalla, C. E., Christensen, B., Ross, M., Eydoux, P., Rosenblatt, D. S., Rozen, R. & Gravel, R. A. (1996) Hum. Mol. Genet. 5 1867-1874. [DOI] [PubMed] [Google Scholar]
- 5.Li Y. N., Gulati, S., Baker, P. J., Brody, L. C., Banerjee, R. & Kruger, W. D. (1996) Hum. Mol. Genet. 5 1851-1858. [DOI] [PubMed] [Google Scholar]
- 6.Chen L. H., Liu, M. L., Hwang, H. Y., Chen, L. S., Korenberg, J. & Shane, B. (1997) J. Biol. Chem. 272 3628-3634. [PubMed] [Google Scholar]
- 7.Gravel R. A., Mahoney, M. J., Ruddle, F. H. & Rosenberg, L. E. (1975) Proc. Natl. Acad. Sci. USA 72 3181-3185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Matsui S. M., Mahoney, M. J. & Rosenberg, L. E. (1983) N. Engl. J. Med. 308 857-861. [DOI] [PubMed] [Google Scholar]
- 9.Mahoney M. J. & Rosenberg, L. E. (1971) J. Lab. Clin. Med. 78 302-308. [PubMed] [Google Scholar]
- 10.Mahoney M. J., Hart, A. C., Steen, V. D. & Rosenberg, L. E. (1975) Proc. Natl. Acad. Sci. USA 72 2799-2803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Fenton W. A. & Rosenberg, L. E. (1978) Annu. Rev. Genet. 12 223-248. [DOI] [PubMed] [Google Scholar]
- 12.Cooper B. A., Rosenblatt, D. S. & Watkins, D. (1990) Am. J. Hematol. 34 115-120. [DOI] [PubMed] [Google Scholar]
- 13.Watkins D., Matiaszuk, N. & Rosenblatt, D. S. (2000) J. Med. Genet. 37 510-513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Watanabe F., Saido, H., Yamaji, R., Miyatake, K., Isegawa, Y., Ito, A., Yubisui, T., Rosenblatt, D. S. & Nakano, Y. (1996) J. Nutr. 126 2947-2951. [DOI] [PubMed] [Google Scholar]
- 15.Gordon B. A. & Carson, R. A. (1976) Can. Med. Assoc. J. 115 233-236. [PMC free article] [PubMed] [Google Scholar]
- 16.Wilson A., Leclerc, D., Rosenblatt, D. S. & Gravel, R. A. (1999) Hum. Mol. Genet. 8 2009-2016. [DOI] [PubMed] [Google Scholar]
- 17.Leclerc D., Wilson, A., Dumas, R., Gafuik, C., Song, D., Watkins, D., Heng, H. H., Rommens, J. M., Scherer, S. W., Rosenblatt, D. S., et al. (1998) Proc. Natl. Acad. Sci. USA 95 3059-3064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Celis R. T., Leadlay, P. F., Roy, I. & Hansen, A. (1998) J. Bacteriol. 180 4828-4833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Altschul S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. (1990) J. Mol. Biol. 215 403-410. [DOI] [PubMed] [Google Scholar]
- 20.Mellman I. S., Youngdahl-Turner, P., Willard, H. F. & Rosenberg, L. E. (1977) Proc. Natl. Acad. Sci. USA 74 916-920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kolhouse J. F. & Allen, R. H. (1977) Proc. Natl. Acad. Sci. USA 74 921-925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Watanabe F. & Nakano, Y. (1997) Methods Enzymol. 281 295-305. [DOI] [PubMed] [Google Scholar]
- 23.Watanabe F., Nakano, Y. & Kitaoka, S. (1987) Biochem. J. 247 679-685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wissenbach U., Six, S., Bongaerts, J., Ternes, D., Steinwachs, S. & Unden, G. (1995) Mol. Microbiol. 17 675-686. [DOI] [PubMed] [Google Scholar]
- 25.Higgins C. F. & Ames, G. F. (1981) Proc. Natl. Acad. Sci. USA 78 6038-6042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Locher K. P., Lee, A. T. & Rees, D. C. (2002) Science 296 1091-1098. [DOI] [PubMed] [Google Scholar]
- 27.Leipe D. D., Wolf, Y. I., Koonin, E. V. & Aravind, L. (2002) J. Mol. Biol. 317 41-72. [DOI] [PubMed] [Google Scholar]
- 28.Haller T., Buckel, T., Retey, J. & Gerlt, J. A. (2000) Biochemistry 39 4622-4629. [DOI] [PubMed] [Google Scholar]
- 29.Bobik T. A. & Rasche, M. E. (2001) J. Biol. Chem. 276 37194-37198. [DOI] [PubMed] [Google Scholar]
- 30.Fenton W. A., Ambani, L. M. & Rosenberg, L. E. (1976) J. Biol. Chem. 251 6616-6623. [PubMed] [Google Scholar]
- 31.Chernyak B. V. & Bernardi, P. (1996) Eur. J. Biochem. 238 623-630. [DOI] [PubMed] [Google Scholar]
- 32.Letuve S., Druilhe, A., Grandsaigne, M., Aubier, M. & Pretolani, M. (2002) Am. J. Respir. Cell Mol. Biol. 26 565-571. [DOI] [PubMed] [Google Scholar]