Abstract
CHARGE syndrome is a well-established multiple-malformation syndrome with distinctive consensus diagnostic criteria. Characteristic associated anomalies include ocular coloboma, choanal atresia, cranial nerve defects, distinctive external and inner ear abnormalities, hearing loss, cardiovascular malformations, urogenital anomalies, and growth retardation. Recently, mutations of the chromodomain helicase DNA-binding protein gene CHD7 were reported to be a major cause of CHARGE syndrome. We sequenced the CHD7 gene in 110 individuals who had received the clinical diagnosis of CHARGE syndrome, and we detected mutations in 64 (58%). Mutations were distributed throughout the coding exons and conserved splice sites of CHD7. Of the 64 mutations, 47 (73%) predicted premature truncation of the protein. These included nonsense and frameshift mutations, which most likely lead to haploinsufficiency. Phenotypically, the mutation-positive group was more likely to exhibit cardiovascular malformations (54 of 59 in the mutation-positive group vs. 30 of 42 in the mutation-negative group; P=.014), coloboma of the eye (55 of 62 in the mutation-positive group vs. 30 of 43 in the mutation-negative group; P=.022), and facial asymmetry, often caused by seventh cranial nerve abnormalities (36 of 56 in the mutation-positive group vs. 13 of 39 in the mutation-negative group; P=.004). Mouse embryo whole-mount and section in situ hybridization showed the expression of Chd7 in the outflow tract of the heart, optic vesicle, facio-acoustic preganglion complex, brain, olfactory pit, and mandibular component of the first branchial arch. Microarray gene-expression analysis showed a signature pattern of gene-expression differences that distinguished the individuals with CHARGE syndrome with CHD7 mutation from the controls. We conclude that cardiovascular malformations, coloboma, and facial asymmetry are common findings in CHARGE syndrome caused by CHD7 mutation.
CHARGE syndrome (MIM 214800) is a distinctive syndrome with a complex constellation of multiple congenital anomalies. The incidence of CHARGE syndrome may be as high as 1 in 8,500 births (Issekutz et al. 2005). It is characterized by variable occurrence of coloboma, choanal atresia or cleft lip and/or palate, cranial nerve dysfunction, characteristic external ear malformations with distinct temporal bone anomalies, cardiovascular malformations, and hypogonadotropic hypogonadism with genitourinary anomalies (Hall 1979; Hittner et al. 1979; Pagon et al. 1981; Blake et al. 1998). Inner ear defects, including Mondini malformation and/or semicircular canal hypoplasia/aplasia are common and often cause hearing loss and vestibular abnormalities in the affected individuals (Amiel et al. 2001). Many types of cardiovascular malformations have been reported, but conotruncal and aortic arch malformations appear to be more common than the others (Blake et al. 1998). Growth delay, distinctive facial features, DiGeorge sequence, and tracheoesophageal fistula are some of the additional clinical features of this condition. In the 25 years since its initial characterization, the diagnostic criteria for the original CHARGE acronym were revised and standardized (Blake et al. 1998), with a further proposal to update them (Verloes 2005). What was considered a prototypic “association” is now viewed as a syndrome in many well-defined cases (Lubinsky 1994), for which the name “Hall-Hittner syndrome” may also apply (Graham 2001). Vissers et al. (2004) reported mutations in the CHD7 gene, which encodes chromodomain helicase DNA-binding protein, in ∼60% of individuals with CHARGE syndrome. Chromatin remodeling is a recognized mechanism of gene-expression regulation, and the CHD7 gene is likely to play a significant role in embryonic development and cell-cycle regulation (Woodage et al. 1997). The prevalence of CHD7 mutations in a large cohort of individuals with CHARGE syndrome and the possible correlation of specific mutations with clinical characteristics have not been formally addressed. We report the systematic molecular and phenotypic evaluation of 110 individuals with clinical CHARGE syndrome. We identified mutations in the CHD7 gene in 58% (64/110) of the affected individuals and analyzed the phenotypic data of the study cohort, using multivariate analysis conditional on the mutation status. Our results indicate significant phenotypic differences between CHD7 mutation carriers and noncarriers and are consistent with locus heterogeneity as the likely explanation for the group of individuals with CHARGE syndrome whose sequence analysis of CHD7 was normal. Nucleotide sequence data reported herein are available in the DDBJ/EMBL/GenBank databases; for details, see the Web Resources section.
Material and Methods
Clinical Evaluation
The study sample included 110 individuals with CHARGE syndrome who either had four major criteria for the diagnosis of CHARGE syndrome (coloboma, choanal atresia, characteristic ear abnormalities, and cranial nerve dysfunction), had three major and three minor criteria (genital hypoplasia, developmental delay, cardiovascular malformation, growth deficiency, orofacial cleft, tracheoesopageal fistula, and characteristic face), or had one or two major criteria with several minor criteria (Blake et al. 1998) (fig. 1). According to Blake et al. (1998), individuals with all four major criteria or with three major and three minor criteria definitely have CHARGE syndrome, and individuals with one or two major criteria and several minor criteria possibly have the syndrome. The children with CHARGE syndrome and their parents were enrolled through the CHARGE Syndrome Foundation meetings in 1999 and 2001 and through the Baylor College of Medicine Cardiovascular Genetics Web site. Informed consent was obtained from participant families. Clinical histories were obtained for all probands, as well as for any other possibly affected family members. The medical records of the individuals with CHARGE syndrome were analyzed. The probands were ascertained by one of the dysmorphologists (S.R.L., J.W.B., C.A.B., J.M.G., or S.L.H.D.). Several individuals with CHARGE syndrome were also enrolled through evaluation of medical records. Photographs were reviewed when the individual with CHARGE syndrome was not available for clinical assessment. The research protocol was reviewed and approved by the Baylor College of Medicine Institutional Review Board.
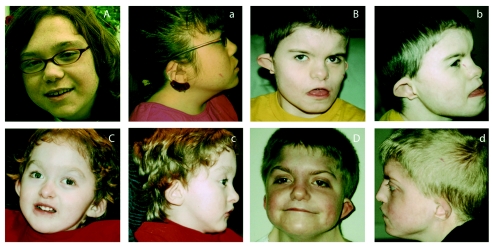
Figure 1.
Individuals with CHARGE syndrome who have CHD7 mutations. A and a, A 17-year-old girl with retinal coloboma, characteristic ear malformation, hearing loss, partial atrioventricular canal defect, growth retardation, and delayed puberty who has a frameshift mutation in exon 37. B and b, An 11-year-old boy with colobomas of the iris and the macula, choanal stenosis, characteristic ear malformation, bilateral Mondini malformation, profound hearing loss, submucous cleft palate, bilateral facial palsy, undescended testes, and pulmonary atresia with ventricular septal defect who has a de novo frameshift mutation in exon 18. C and c, A 3-year-old boy with bilateral iris coloboma, choanal stenosis, ear malformation, hearing loss, facial palsy, and abnormal cochlea but without cardiovascular malformation who has a nonsense mutation in exon 13. D and d, A 12-year-old boy with coloboma involving the optic nerve, choanal atresia, characteristic ear malformation, profound hearing loss, abnormal inner ear, patent ductus arteriosus, and genital abnormalities who has a frameshift mutation in exon 2. Note that the individuals in panels B and D illustrate the characteristic facial appearance of CHARGE syndrome.
DNA Sequencing
Primers were designed using the program Primer 3, on the basis of the annotated genomic sequence across each exon, including regions of at least 40–50 bp of flanking intervening sequence (table A1). Larger exons were subdivided to allow for optimal product lengths. All PCR reactions were performed in a 25-μl reaction volume containing 40 ng of genomic DNA, 10 pmol of each primer, and 12.5 μl of Qiagen Multiplex PCR Master Mix. Conditions for PCR were 15 min of initial activation step at 95°C, followed by 40 cycles of denaturation at 94°C for 30 s and annealing at 60°C for 90 s, and a final extension at 72°C for 60 s. An annealing temperature of 55°C was used for PCR amplification of exons 30 and 38b. Sequencing of the CHD7 coding region (exons 2–38) for mutations was performed using Big Dye Terminator Cycle Sequencing Reactions on an ABI 3700 (PE Applied Biosystems) by use of the manufacturer’s standard protocol and reagents. Sequence electropherograms were compared with gene sequences from GenBank accession number NM_017780. Sequence analysis of both the sense and antisense strands of the PCR-amplified fragments was performed with the aid of Sequencher 4.5 software.
FISH
FISH was performed on DNA from 23 individuals with CHARGE syndrome whose sequence analysis of CHD7 was normal, by use of two clones, RP11-33I11 (GenBank accession number AC113143) and RP11-174G1 (GenBank accession number AC022182 ), encompassing the CHD7 gene. The BAC clones were obtained from the Children’s Hospital Oakland Research Institute BACPAC Resources Center. FISH analysis was performed as described elsewhere (Lalani et al. 2003). After hybridization and washing, biotinylated probes were detected by avidin-fluorescein, and digoxigenin-labeled probes were detected using anti-digoxigenin antibody coupled with rhodamine. The chromosome preparations were counterstained with 4′,6-diamidino-2-phenylindole and were analyzed with a Zeiss Axioskop fluorescence microscope.
Mouse Embryo In Situ Hybridization
Whole-mount mouse embryo in situ hybridization was performed at 10.5 d post coitum (dpc), with modification of the protocol described elsewhere (Wilkinson 1992) and with the use of blocking reagent (Boehringer Mannheim). Probes were labeled using a DIG RNA Labeling kit (Boehringer Mannheim). Chd7 riboprobes were prepared from a cDNA fragment (1587–2034) of KIAA1416 (GenBank accession number BC034239). The probe spanning 447 bp in the 3′ region of the gene was generated using PCR. The primers used for PCR were forward primer 5′-GGAAACGACAATGAAGACGAGAAC-3′ and reverse primer 5′-CGTTTCCGGTGTTTTCTATCTCAT-3′. In situ hybridization on sagittal sections of 10.5-dpc embryos was performed using the same probe.
Microarray Analysis of Gene Expression
RNA was isolated from the lymphoblastoid cell lines of seven mutation-positive, three mutation-negative, and five control samples by use of the Qiagen RNeasy Mini kit. The quality and purity of total RNA was assayed in a 2% agarose gel, and the recovery was calculated after absorbance was measured with a spectrophotometer at 260 nm. Samples were labeled using the standard Affymetrix T7 oligo(dT) primer protocol and reagents supplied in the Affymetrix One-Cycle Target Labeling and Hybridization kit (catalog number 900493). Total RNA was reverse transcribed to produce double-stranded cDNA. The cDNA product was used as a template for the in vitro transcription reaction, producing biotin-labeled cRNA. The labeled cRNA was quantified by spectrophotometry (NanoDrop ND-1000). Fifteen μg of the labeled cRNA was fragmented and rechecked for concentration. A hybridization cocktail containing Affymetrix spike-in controls and fragmented labeled cRNA was loaded onto GeneChip U133 2.0 Plus arrays. The array was hybridized overnight, for 16 h at 45°C with rotation at 60 rpm. Arrays were then washed and stained on Affymetrix Fluidics Station 400 equipment with a strepavidin, R-phycoerythrin conjugate stain. Signal amplification was done using biotinylated anti-streptavidin. The stained array was scanned on the Affymetrix GeneChip Scanner 3000. The images were analyzed, and quality-control metrics were recorded using Affymetrix GCOS software, version 1.1.2. By use of reference standards, data were normalized from different experiments and were compared with multiple experiments on a quantitative level. The expression profiles of both groups were compared and further evaluated with control samples.
Affymetrix Data Analysis
Microarray data were analyzed with ChipST2C 1.14 (see ChipST2C Web site). Normalization of the probe-level hybridization data was performed using the Robust Multiarray Average measure (Affy, R-package) (Irizarry et al. 2003a, 2003b). Pairwise T tests were used to compare expression values from individuals with CHARGE syndrome and CHD7 mutations (n=7) with those from matched controls (n=5) by use of ChipST2C. With a threshold of P<.001, 116 transcripts were identified. The analysis was repeated for these transcripts but with the addition of the individuals with CHARGE syndrome without detectable CHD7 mutation (n=3), by use of an F test (analysis of variance) and within-gene permutation to evaluate the empirical P values (B=1,000 iterations). Storey q values were calculated to estimate the positive false-discovery rate (Storey and Tibshirani 2003). This resulted in 85 transcripts, with q<0.01 for all transcripts (positive false-discovery rate 1%). The results are displayed after hierarchical cluster analysis for genes (fig. 2).
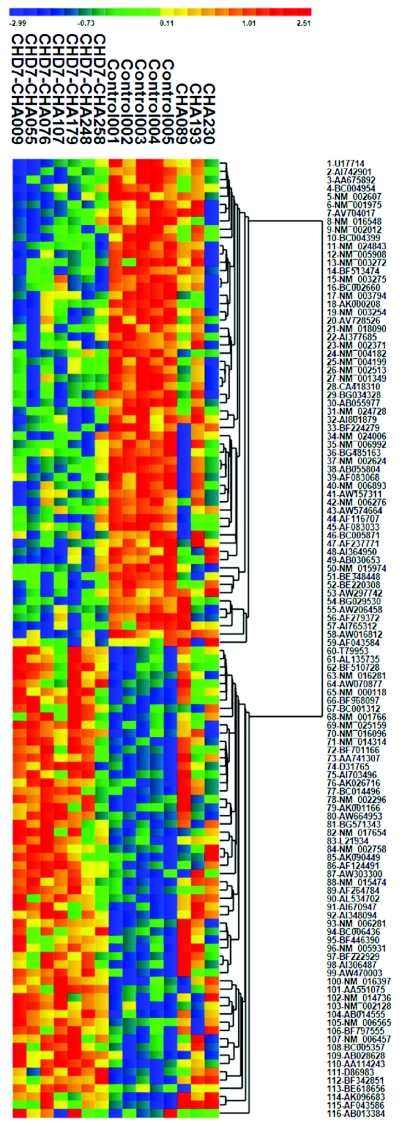
Figure 2.
Microarray gene-expression profile and the resulting cluster analysis. Patterns of gene expression were assessed on Affymetrix U133Plus2.0 arrays (see “Material and Methods” section for details). After identification of a subset of transcripts with apparently significant differences in expression between the individuals with CHARGE syndrome with CHD7 mutations and the controls, the combined data were used to produce a hierarchical cluster analysis. Expression values shown were standardized using the gene-specific mean and SD of log2 expression. Note that the profile of CHA230 (who had no detectable CHD7 mutation) is highly similar to those of individuals with CHD7 mutations.
Phenotypic Data Analysis
Association of CHD7 mutations with clinical phenotypes was computed using SPSS 12.0 (SPSS). The analysis was performed for each clinical variable, and the analysis was compared between mutation-positive and mutation-negative individuals. Types of mutations—including nonsense, frameshift, missense, and splice-site mutations—and the position of the mutations within the protein were analyzed and compared with the phenotypic data of the individuals with CHARGE syndrome. The Fisher’s exact test with 1 df was used for all comparisons. The significance threshold was set at P<.05. All P values were two sided.
Results
Mutation Analysis
Among 110 individuals with clinical CHARGE syndrome in the study group, there were 105 sporadic cases; four familial cases and one pair of MZ twins were also included in the study. The individuals with CHARGE syndrome comprised 61 (55%) females and 49 (45%) males. Apparently pathogenic mutations in CHD7 were found in 64 (58%) of the individuals with CHARGE syndrome, distributed throughout the gene (fig. 3 and table 1). Truncating mutations were seen in 47 (73%) of patients, including 28 stop-codon mutations and 19 frameshift mutations. Seven (11%) were missense mutations, and 9 (14%) were intron-exon boundary mutations (table 2). Either one or both parental samples were unavailable for 17 (26%) of the mutation-positive patients. All truncating mutations for which parental samples were available were found to be de novo. Of the seven missense mutations, parental samples were unavailable for one, five were found to be de novo, and one was inherited from an affected mother. Of the nine intron-exon boundary mutations, seven were found to be de novo; parental samples were unavailable for the other two. Three familial cases in this study group were found to have mutations in the CHD7 gene (fig. 4). The missense mutation R2319S, seen in CHA166, was also identified in the mother, who was mildly affected. In another familial case, the E1271X mutation was identified in both MZ twins. The nonsense mutation W2332X was identified in a mildly affected girl whose brother died of complications of CHARGE syndrome in infancy. Although most mutations were unique, R1339X in exon 17, involving the HELICc domain, was present in both CHA151 and CHA184; R1810X in exon 26 was observed in both CHA16 and CHA85; W2332X in exon 33 was present in both CHA76 and CHA176; and R2653X in exon 36, involving the BRK domain, was present in three individuals: CHA131, CHA255, and CHA304. One mutation was detected in the chromodomain, six in the SNF2 domain, three in the HELICc domain, and three in the BRK domain.
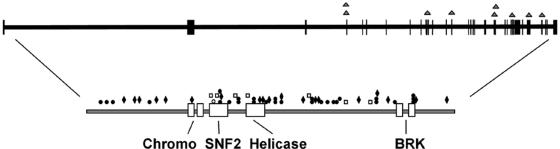
Figure 3.
Genomic and protein map of CHD7, indicating the spectrum of mutations in CHARGE syndrome. On the protein map, blackened circles signify nonsense mutations, blackened diamonds signify frameshift mutations, unblackened squares signify missense mutations, and the unblackened circle specifies a mutation with an in-frame deletion of 3 aa. On the genomic sequence, triangles represent splice-site mutations.
Table 1.
CHD7 Mutations Observed in 64 Individuals with CHARGE Syndrome
| MutationNumber | Exon | Individuala | Mutation | Originb | Domain |
| 1 | 2 | 95 | Q112X | NA | |
| 2 | 2 | 330 | W145X | De novo | |
| 3 | 2 | 3 | Q180X | De novo | |
| 4 | 2 | 315 | T262fsX304 | De novo | |
| 5 | 2 | 125 | V424fsX466 | NA | |
| 6 | 2 | 107 | P366fsX377 | De novo | |
| 7 | 2 | 134 | R494X | De novo | |
| 8 | 2 | 284 | H556fsX564 | De novo | |
| 9 | 3 | 28 | G623fsX626 | De novo | |
| 10 | 4 | 308 | c.2097−1G→A | De novo | |
| 11 | 4 | 297 | c.2238+1G→A | De novo | |
| 12 | 8 | 34 | Q839fsX843 | De novo | Chromo |
| 13 | 11 | 22 | c.2836−15C→G | NA | |
| 14 | 12 | 116 | L1007C | NA | SNF2 |
| 15 | 12 | 71 | T1027_P1029del | De novo | SNF2 |
| 16 | 12 | 233 | W1031G | De novo | SNF2 |
| 17 | 13 | 66 | R1069X | De novo | SNF2 |
| 18 | 13 | 281 | R1069X | De novo | SNF2 |
| 19 | 13 | 119 | V1070fsX1071 | NA | SNF2 |
| 20 | 14 | 307 | E1152X | NA | |
| 21 | 15 | 206 | Q1214R | De novo | |
| 22 | 15 | 158 | c.3778+1G→A | NA | |
| 23 | 16 | 172 | E1271X | NA | |
| 24 | 16 | 182 | E1271X | NA | |
| 25 | 16 | 215 | L1294P | De novo | HELICc |
| 26 | 17 | 151 | R1339X | De novo | HELICc |
| 27 | 17 | 184 | R1339X | De novo | HELICc |
| 28 | 18 | 236 | V1409fsX1420 | De novo | |
| 29 | 18 | 248 | M1442fsX1443 | De novo | |
| 30 | 19 | 221 | R1465X | NA | |
| 31 | 19 | 55 | R1493fsX1501 | De novo | |
| 32 | 20 | 146 | W1534X | De novo | |
| 33 | 20 | 171 | L1545X | De novo | |
| 34 | 21 | 9 | R1571fsX1572 | De novo | |
| 35 | 21 | 25 | S1577X | De novo | |
| 36 | 21 | 113 | c.4850+1G→A | De novo | |
| 37 | 22 | 58 | c.5050+3A→T | De novo | |
| 38 | 24 | 49 | E1751fsX1752 | NA | |
| 39 | 26 | 16 | R1810X | De novo | |
| 40 | 26 | 85 | R1810X | De novo | |
| 41 | 26 | 98 | L1815P | De novo | |
| 42 | 26 | 154 | G1835fsX1843 | De novo | |
| 43 | 26 | 143 | c.5534+1G→T | De novo | |
| 44 | 29 | 300 | A1896fsX1898 | NA | |
| 45 | 29 | 42 | W1903X | De novo | |
| 46 | 29 | 290 | R1945X | NA | |
| 47 | 30 | 287 | C2017X | De novo | |
| 48 | 30 | 29 | R2024X | De novo | |
| 49 | 30 | 339 | c.6103+8C→T | De novo | |
| 50 | 31 | 258 | R2053X | De novo | |
| 51 | 31 | 292 | H2096R | De novo | |
| 52 | 31 | 92 | E2202fsX2208 | De novo | |
| 53 | 33 | 166 | R2319S | Inherited | |
| 54 | 33 | 76 | W2332X | De novo | |
| 55 | 33 | 176 | W2332X | De novo | |
| 56 | 33 | 179 | P2333fsX2334 | De novo | |
| 57 | 34 | 239 | E2450GfsX2468 | De novo | |
| 58 | 34 | 203 | K2510X | De novo | |
| 59 | 35 | 224 | c.7830+11A→G | De novo | |
| 60 | 36 | 131 | R2653X | NA | BRK |
| 61 | 36 | 255 | R2653X | NA | BRK |
| 62 | 36 | 304 | R2653X | De novo | BRK |
| 63 | 37 | 37 | Q2691fsX2709 | NA | |
| 64 | 38 | 77 | L2916fsX2940 | NA |
For individual IDs, add “CHA” before the number shown.
NA = not available.
Table 2.
Summary of Mutations Observed in the CHD7 Gene in Individuals with CHARGE Syndrome
| Type of Mutation | Frequency(n) | De novo | Inherited | ParentalSamplesUnavailable |
| Nonsense | 28 | 20 | 0 | 8 |
| Frameshift | 19 | 13 | 0 | 6 |
| In-frame deletion (3 aa) | 1 | 1 | 0 | 0 |
| Missense | 7 | 5 | 1 | 1 |
| Splice site | 9 | 7 | 0 | 2 |
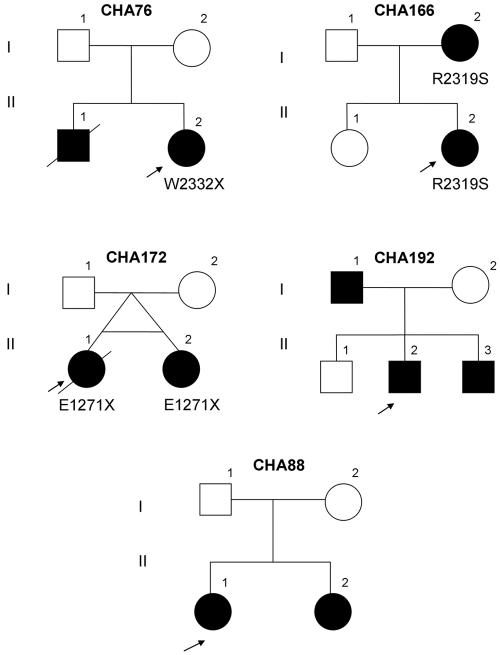
Figure 4.
Five pedigrees with CHARGE syndrome illustrating four familial cases and an affected pair of MZ twins. CHD7 mutations were identified in pedigrees CHA76, CHA166, and CHA172. Germline mosaicism is suggested in CHA76, whereas CHA166 indicates autosomal dominant transmission of the missense mutation in this mildly affected family. CHA172 shows a nonsense mutation in MZ twins. Mutations in CHD7 were not identified in families CHA192 and CHA88. Probands are indicated by arrows.
Of 64 individuals with the CHD7 mutation, there were 40 (63%) females and 24 (37%) males. No sex difference was noted when they were compared with the group without CHD7 mutations. Three of the clinical criteria showed statistically significant positive correlations with the CHD7 mutation: cardiovascular malformations (54 of 59 in the mutation-positive group vs. 30 of 42 in the mutation-negative group; P=.014), coloboma of the iris, retinal, or optic nerve (55 of 62 in the mutation-positive group vs. 30 of 43 in the mutation-negative group; P=.022), and facial asymmetry (36 of 56 in the mutation-positive group vs. 13 of 39 in the mutation-negative group; P=.004 ). There were no significant differences between the two groups with respect to choanal atresia, external or inner ear malformations, deafness, cleft lip and palate, tracheoesophageal fistula, esophageal atresia, and urogenital abnormalities (table 3). The major and minor clinical diagnostic criteria scores for each individual, as suggested by the criteria from Blake et al. (1998) and Verloes (2005), were also compared with mutation status. All 10 individuals with the combined three major signs of Verloes (coloboma, atresia of choanae, and hypoplastic semicircular canals) were found to have mutations in the CHD7 gene (P=.005).
Table 3.
Distribution of CHARGE Syndrome Features in CHD7 Mutation-Positive and Mutation-Negative Individuals
|
No. of Individuals withFeature/No. Evaluated (%) |
|||
| Syndrome Feature | WithCHD7 Mutation | WithoutCHD7 Mutation | Pa |
| Cardiovascular malformation | 54/59 (92%) | 30/42 (71%) | .014 |
| Coloboma | 55/62 (89%) | 30/43 (70%) | .022 |
| Facial asymmetry | 36/56 (64%) | 13/39 (33%) | .004 |
| Choanal atresia | 34/57 (60%) | 23/39 (59%) | 1.000 |
| Hearing loss | 54/59 (92%) | 36/38 (95%) | .701 |
| Inner ear abnormality | 21/22 (95%) | 9/10 (90%) | .490 |
| External ear malformation | 59/62 (95%) | 39/42 (93%) | .683 |
| Urogenital abnormality | 29/53 (55%) | 26/39 (67%) | .287 |
| Cleft lip/palate | 18/60 (30%) | 9/41 (22%) | .493 |
| Tracheoesophageal fistula | 10/55 (18%) | 3/40 (8%) | .226 |
All P values are two sided. Significant P values are in bold italics.
The phenotype-genotype analysis of the CHD7 mutation-positive group revealed no clear correlation between the severity of the clinical findings and the type of mutation. Individuals with a milder phenotype were found to have mutations involving both the middle third and the C-terminal part of the protein. Ten SNPs with minor-allele frequency >0.01 in the white subjects (n=86) were detected within the CHD7 gene (table 4). There were no significant differences in allele frequencies between mutation carriers and noncarriers.
Table 4.
Common CHD7 Polymorphisms
|
Frequency |
|||
| Exon | Polymorphism | Allele 1 | Allele 2 |
| 2 | c.1665+34G→A | .793 | .207 |
| 4 | c.2238+39G→A | .802 | .198 |
| 6 | c.2442+38A→T | .976 | .024 |
| 6 | c.2442+93A→G | .810 | .190 |
| 9 | c.2614−45G→A | .779 | .221 |
| 10 | c.2698−54A→G | .540 | .460 |
| 15 | c.3523−39C→G | .989 | .011 |
| 19 | c.4533+45G→A | .731 | .269 |
| 31 | c.6276G→A | .971 | .029 |
| 34 | c.7356A→G | .951 | .049 |
Mouse Embryo In Situ Hybridization
Previous research has shown that CHD7 expression is widespread in human tissues, but the method used did not allow estimation of relative expression levels or correlation of expression pattern with the defects in organogenesis observed in individuals with CHARGE. Therefore, we examined the expression of the mouse homolog, Chd7, at a stage in which the relevant anlage might be visualized. Mouse embryo whole-mount and section in situ hybridization at 10.5 dpc showed expression of Chd7 in the cardiac outflow tract, truncus arteriosus, facio-acoustic (VII–VIII) preganglion complex, hindbrain, forebrain, mandibular component of the first branchial arch, otic vesicle, optic stalk/optic vesicle, and olfactory pit (fig. 5) (Kaufman 1992). Although Chd7 is widely expressed, these tissues in the early embryo exhibit differentially increased expression. The tissues with high levels of expression appear to correspond to the mature structures that are most severely affected in CHARGE syndrome.
Figure 5.
Mouse embryo whole-mount and section in situ hybridization showing the expression of Chd7. A, Whole-mount in situ hybridization of the embryo at 10.5 dpc, demonstrating expression of Chd7 in the craniofacial region. B, Expression of Chd7 observed in cardiac outflow tract (OFT). LA = left atrium; RA = right atrium. C, Sagittal section of embryo at 10.5 dpc, showing expression in the facio-acoustic (VII–VIII) preganglion complex (FA), hindbrain (HB), forebrain (FB), mandibular component of the first branchial arch (BA), and otic vesicle (OV). D and F, Sagittal section of embryo with expression in optic stalk/optic vesicle (OPV) and olfactory pit (OP). E and G, Magnified images of panel C, indicating expression of Chd7 in facio-acoustic preganglion complex and otic vesicle, with hematoxylin and eosin stain. H, Sagittal section of the embryo, demonstrating expression in truncus arteriosus (TA) at 10.5 dpc.
Gene-Expression Analysis
Because CHD7 is broadly expressed and because its protein motifs suggest that it may play a role in global regulation of chromatin structure, and hence gene expression, we wished to determine whether a characteristic pattern of expression abnormality is associated with mutation. Microarray analysis of gene-expression patterns by use of lymphoblastoid cell lines from seven CHD7 mutation-positive individuals showed significant gene expression differences when compared with those from five control individuals and three affected individuals with no detected mutation of CHD7 (fig. 2). The gene-expression signature created by the 116 transcripts with the best statistical support shows that the cases and controls cluster into two distinct groups. Interestingly, individual CHA230 appears to cluster with the case patients bearing a CHD7 mutation, although we have not been able to find an alteration of the coding sequence in that individual.
Discussion
CHARGE syndrome is a recognizable multiple-malformation syndrome, the etiology of which was unknown until recently (Vissers et al. 2004). Autosomal dominant inheritance in rare families (Mitchell et al. 1985), concordance in MZ twins (Blake et al. 1989), advanced paternal age (Tellier et al. 1996), and variable chromosomal abnormalities in rare individuals with CHARGE syndrome (Clementi et al. 1991; Lee et al. 1995; North et al. 1995; Devriendt et al. 1998; De Krijger et al. 1999; Lev et al. 2000) supported a genetic basis for the condition. Genomewide deletion screening by use of comparative genomic hybridization (CGH) (Sanlaville et al. 2002), STR markers (Lalani et al. 2003), and SNPs (Lalani et al. 2005) in affected families did not identify a common microdeletion in this syndrome. Vissers et al. (2004) identified mutations in the gene for a chromodomain helicase DNA-binding protein, CHD7, in 10 of 17 individuals with CHARGE syndrome, after BAC CGH analysis of two unrelated individuals with CHARGE syndrome, one with an apparently balanced translocation involving chromosomes 6 and 8. These two subjects had overlapping microdeletions in chromosome 8q12, both of which spanned the CHD7 locus. Subsequent mutation analysis demonstrated that de novo point mutations in this gene are associated with CHARGE syndrome. Chromatin remodeling is an important mechanism for regulation of gene expression. Like the other CHD genes, CHD7 is likely to play a significant role in development and cell-cycle regulation. The gene encodes a protein with chromatin remodeling, an SNF2-like ATPase/helicase domain, and DNA-binding motifs. CHD7 is 188 kb in length and is composed of 38 exons, of which 37 are coding. In our study, pathogenic mutations were identified throughout the gene, although many mutations were identified in exons that encode the middle third of the protein. Most of the mutations are truncating mutations that are likely to result in haploinsufficiency. Most of the affected individuals have de novo mutations in the CHD7 gene. However, rare familial cases with recurrence were observed. In at least one family in our study cohort, autosomal dominant inheritance from the mother was documented. In CHA166, the R2319S mutation seen in the proband was also found in the affected mother who had iris coloboma, facial palsy, hearing loss, and delayed puberty. Neither the mother nor the child had a cardiovascular malformation. CHA76, a mildly affected girl, was found to have truncating mutation W2332X, not present in either of her parents. Interestingly, this child had a sibling who died at age 6 mo due to complications related to the diagnosis of CHARGE syndrome. The phenotype of the deceased infant included coloboma, choanal atresia, hearing loss, pulmonary atresia with aortic stenosis and ventricular septal defect, kidney abnormality, duodenal atresia, and swallowing difficulties that necessitated G-tube and Nissen fundoplication. A tissue sample was not available for analysis from this sibling. Nonpaternity was excluded for this study subject. These findings suggest germline mosaicism in this family.
Proband CHA151 was found to have a de novo heterozygous mutation, R1339X, in the CHD7 gene. His features included coloboma, choanal atresia, deafness, ear malformation, undescended testes, and partial atrioventricular canal. His sister was born with coarctation of the aorta, and their mother had unilateral deafness and mild facial palsy. The parents and sister were negative for the CHD7 R1339X mutation detected in the proband. We conclude that, in this apparently familial case, the clinical features in other family members are unrelated to the R1339X mutation seen in the proband.
This study group also included a pair of female MZ twins. Both had the nonsense mutation E1271X in exon 16. They were born prematurely at 31 wk gestation. Twin A died at age 4 mo due to sepsis, and a complete autopsy was performed. The phenotype included bilateral optic-nerve coloboma, complete atrioventricular canal defect, mild-to-moderate hypoplasia of the ascending aorta, anomalous aortic arch branches including common brachiocephalic trunk, unilateral choanal atresia, olfactory bulb hypoplasia, growth retardation, unilateral cleft lip and palate, bilateral simplified ears, and renal glomerular cysts. Twin B survived and had bilateral coloboma, choanal stenosis, external ear malformation, profound bilateral hearing loss, bilateral cleft lip and palate, ventricular septal defect and patent ductus arteriosus, growth retardation, and normal renal sonogram (table 5). The causes of phenotypic variation between MZ twins remain poorly understood. However, the variation of phenotype in these twins could be explained by differential epigenetic regulation, such as variable methylation, resulting in altered gene expression and cellular phenotypes.
Table 5.
Clinical Features of CHARGE Syndrome in Familial Cases[Note]
| CHA76 |
CHA166 |
CHA172 |
CHA192 |
CHA88 |
|||||||
| Clinical Features | II1 | II2 | I2 | II2 | II1 | II2 | I1 | II2 | II3 | II1 | II2 |
| Coloboma | + | + | + | + | + | + | − | − | − | + | + |
| Choanal atresia | + | − | − | − | + | + | − | + | − | + | − |
| Cranial nerve anomalies | + | + | + | + | + | + | + | + | + | + | + |
| Hearing loss | + | + | + | + | U | + | + | + | + | + | + |
| Inner ear abnormality | U | U | U | U | U | U | U | + | + | U | U |
| External ear malformation | + | + | − | + | + | + | + | + | + | + | + |
| Cardiovascular malformation | + | + | − | − | + | + | − | − | − | + | − |
| Urogenital abnormality | + | − | + | − | + | − | − | + | − | + | − |
| Cleft lip/palate | − | − | − | − | + | + | + | − | + | − | − |
| Tracheoesophageal fistula | − | − | − | − | − | − | − | − | − | − | − |
| Growth retardation | − | + | − | − | + | + | + | + | − | + | + |
| Developmental delay | + | + | − | − | + | + | U | + | + | + | + |
Note.— U = unknown; + = presence; − = absence.
Cardiovascular malformations were more common in the CHD7 mutation-positive group than in the mutation-negative group. On the basis of its function in nucleosome remodeling and transcription regulation, this chromodomain helicase DNA-binding protein may affect the functions of multiple genes involved in the patterning of the cardiogenic developmental fields. Mouse in situ hybridization studies demonstrate expression of Chd7 in embryonic heart at 10.5 dpc. Assorted cardiovascular malformations are seen in the individuals with CHARGE syndrome with CHD7 mutations, including tetralogy of Fallot, septal defects, patent ductus arteriosus, coarctation of the aorta, and complete and partial atrioventricular canal. Patent ductus arteriosus is more common in the CHD7 mutation-positive group than in the mutation-negative group (31 of 54 vs. 7 of 38; P<.001). There was no clear difference between the two groups with respect to other cardiovascular malformations, including septal defects, left ventricular outflow-tract obstructive lesions, tetralogy of Fallot, and other outflow-tract anomalies. All five individuals with CHARGE syndrome with atrioventricular canal defects had CHD7 mutation; however, the number of individuals with CHARGE syndrome with this cardiovascular malformation was too small for meaningful comparison.
Postnatal growth retardation is known to be a distinct phenotype in CHARGE syndrome. Various studies indicate presence of growth retardation in ∼75% of individuals with CHARGE syndrome (Tellier et al. 1998). The etiology of short stature in these individuals is not understood in detail. Some children are known to have growth-hormone deficiency, although systematic studies of endocrine problems in these children are lacking. In our study, growth retardation was obvious in both CHD7 mutation carriers and noncarriers; however, the Z scores of stature (normalized for age and sex) were slightly lower for the mutation-positive group (mean Z=-2.5329) than for the mutation-negative group (mean Z=-1.7594). Detailed endocrine evaluation of these individuals with CHARGE syndrome may identify specific endocrine defects caused by aberration of CHD7 function.
Cranial nerve dysfunction, including facial and auditory/vestibular nerves, is a major characteristic of CHARGE syndrome. In our study, facial asymmetry was observed more often in the CHD7 mutation-positive group than in the mutation-negative group (36 of 56 vs. 13 of 39; P=.004) (fig. 1). Neuronal migration in vertebrate brain development is an important phenomenon, and the migration of embryonic facial motoneurones in mice and chicks has been studied extensively (Studer 2001). Despite detailed analyses, the cellular and molecular mechanisms that direct the migration of distinctive neuronal populations remain largely unknown. CHD7 may play a role in neuronal migration, either directly or through its interaction with various partners, thus causing the multiple cranial nerve dysfunction commonly observed in CHARGE syndrome.
We have detected CHD7 mutations in ∼60% of individuals with clinical CHARGE syndrome. Because CHARGE syndrome has considerable phenotypic heterogeneity, clinical criteria were developed to aid reliable diagnosis. Although individuals with four major criteria or with three major and three minor criteria were designated as having definite CHARGE syndrome, the clinical guidelines called for consideration of the diagnosis of CHARGE syndrome in children with either one or two major characteristics and multiple minor characteristics (Blake et al. 1998). In our study, we included these individuals given a possible diagnosis of CHARGE syndrome and found pathogenic mutations in three of them. Inner ear defects, including semicircular canal hypoplasia with or without Mondini malformation, is a distinct feature of CHARGE syndrome. In our study cohort, temporal bone CT scans were available for review for 32 individuals with CHARGE syndrome. Of these, 30 (94%) had abnormal findings, with hypoplastic or absent semicircular canals with or without Mondini malformation. There was no statistically significant difference in the carriers and noncarriers of CHD7 mutation when this clinical entity was separately analyzed (8 of 9 in the mutation-negative group vs. 22 of 23 in the mutation-positive group; P=.490). However, within this group of 32 individuals, for which temporal bone imaging scans were available, Verloes’s major signs—that is, iris or choroid coloboma, atresia of choanae, and hypoplastic semicircular canals—were analyzed. Of the 32 individuals, 10 displayed all three of Verloes’s major signs, and all were found to have a CHD7 mutation (P=.005) (table 6). Interestingly, these 10 individuals also displayed Blake’s four major criteria and had associated cranial nerve dysfunction and characteristic ear abnormalities. Eight of these individuals also had a cardiovascular malformation. We suggest that individuals with combined coloboma, choanal atresia, and hypoplastic semicircular canals are likely to have mutations in the CHD7 gene. Many individuals with all four of Blake's major criteria also have mutations in CHD7 (table 7). The minor criteria of Verloes were difficult to score accurately for each individual, because growth hormone and gonadotropin deficiencies were not documented for most of the individuals in the study group. Also, formal assessment of mental retardation, another minor sign, was not performed for these individuals. Similarly, distinctive CHARGE facies, one of Blake’s minor criteria, was difficult to score for many individuals enrolled through the Baylor College of Medicine Cardiovascular Genetics Web site by various referring physicians.
Table 6.
Characterization of Verloes’s Major Signs in 32 Individuals with CHARGE Syndrome[Note]
|
No. of Verloes’sMajor Signsa |
|||||
| Individuals | 1 | 2 | 3 | Total | P |
| CHD7 mutation negative | 4 | 5 | 0 | 9 | |
| CHD7 mutation positive | 1 | 12 | 10 | 23 | .005 |
| All | 5 | 17 | 10 | 32 | |
Note.— Significant results are in bold italics.
Major signs include coloboma (iris or choroid, with or without microphthalmia), atresia of choanae, and hypoplastic semicircular canals (Verloes 2005).
Table 7.
Characterization of Blake’s Major Criteria in 98 Individuals with CHARGE Syndrome
|
No. of Blake’sMajor Criteriaa |
||||||
| Individuals | 1 | 2 | 3 | 4 | Total | P |
| CHD7 mutation negative | 1 | 10 | 16 | 12 | 39 | |
| CHD7 mutation positive | 0 | 3 | 29 | 27 | 59 | .014 |
| All | 1 | 13 | 45 | 39 | 98 | |
Major criteria include ocular coloboma (iris, retina, choroid, or disc; microphthalmia), choanal atresia (unilateral/bilateral; stenosis/atresia), cranial nerve anomalies (facial palsy-unilateral/bilateral, sensorineural deafness, and/or swallowing problems), and characteristic ear anomalies (external ear-lop or cup-shaped ear, middle ear ossicular malformations, mixed deafness, and cochlear defects) (Blake et al. 1998).
In the initial study, CHD7 mutations and/or deletions were seen in ∼63% (12/19) of individuals with CHARGE syndrome (Vissers et al. 2004). These data are similar to those of our present study and suggest that CHARGE syndrome is likely a genetically heterogeneous condition. This is also implied by various chromosomal aberrations described in rare individuals with CHARGE. Additionally, the microarray analysis of gene expression with the use of lymphoblastoid cell lines from CHD7 mutation-positive and mutation-negative individuals with CHARGE syndrome shows significant gene expression differences between these two groups. The differential gene-expression pattern in the CHD7 mutation-negative group may further assist in the understanding of the molecular basis of the disorder in this group of affected children. Previously, we reported a de novo mutation in SEMA3E, a gene encoding a semaphorin protein involved in axon guidance and cell migration, in an individual with CHARGE syndrome (CHA6). SEMA3E became a candidate gene after a chromosomal breakpoint was narrowed in an unrelated affected individual, CHA193, with an apparently balanced chromosomal translocation, 46,XY(2;7)(p14;q21.11), within 200 kb of SEMA3E on chromosome 7 (Lalani et al. 2004). Both CHA6 and CHA193 were studied for mutations in CHD7. No mutation in the coding region or the exon-intron boundaries of CHD7 was detected in these individuals. Moreover, the lymphoblast cell line from CHA193 was also analyzed for gene-expression pattern and was consistent with a signature pattern seen in CHD7 mutation-negative individuals with CHARGE (fig. 2). It will be important to study the association between CHD7 and this semaphorin protein gene in a mutant animal model, to help further delineate the pathogenesis, particularly as it relates to the cranial nerve dysfunction seen so frequently in individuals with CHD7 mutation.
No significant distinct phenotype-genotype correlations were identified for different types of CHD7 mutations (missense, splice site, and truncating) in the mutation-positive cohort. Similarly, no correlation was observed between the clinical severity and the inferred length of the mutant CHD7 protein. Three unrelated individuals with CHARGE syndrome with the same mutation, R2653X, in the BRK domain of the CHD7 gene have different clinical phenotypes. We have also observed variable expression of a CHD7 mutation within individuals in the same family. The MZ twins with the same mutation, E1271X, have differences in their phenotype; twin A had atrioventricular canal defect, and twin B had a ventricular septal defect and patent ductus arteriosus. CHA166, with the R2319S mutation, has iris coloboma, ear malformation, mixed hearing loss, and facial palsy. Her mother, who has the same mutation, has iris coloboma, hearing loss, facial palsy, and a history of delayed puberty.
There was no correlation of severity of phenotype with mutation in specific domains of the protein. However, we have observed that five individuals with CHARGE syndrome with mutations in the SNF2 domain have variable cardiovascular malformations.
In this study, the CHD7 mutations in the coding region and exon-intron boundaries were analyzed, and these data were used for the statistical analyses. One limitation of the study is the lack of sequence information in the regulatory and promoter elements of the CHD7 gene. Heterozygous deletions within the CHD7 gene may not have been detected by sequencing. We addressed this concern by performing FISH analyses on 23 CHD7 mutation-negative individuals with CHARGE syndrome, using RP11-33I11 and RP11-174G clones encompassing CHD7, and excluded any larger heterozygous deletion in these individuals. However, microdeletions of ⩽100 kb could potentially be undetected by the FISH methodology used.
Previously, the expression profile by semiquantitative RT-PCR analysis of CHD7 showed ubiquitous expression (Vissers et al. 2004). Mouse embryo in situ hybridization, however, shows patterned expression of Chd7 in tissues that are frequently affected in CHARGE syndrome, such as optic stalk/optic vesicle, facio-acoustic preganglion complex, olfactory pit, forebrain, hindbrain, mandibular component of the first branchial arch, and outflow tract of the heart. Remodeling of eukaryotic chromatin is known to play a significant role in regulation of gene transcription. Tong et al. (1998) have shown that the ATP-dependent nucleosome-remodeling activity of the human histone deacetylase complex derives from the helicase/ATPase domains of CHD3 (Mi-2α) and CHD4 (Mi-2β) proteins. This functionally coupled complex is referred to as the “nucleosome remodeling and deacetylating (NuRD) complex.” The NuRD complex from HeLa cells contains seven major polypeptides, including Mi2, MTA2, MBD3, and the histone deacetylase core, HDAC1/2 and RbAp46/48. Current models suggest that NuRD complexes are involved in producing a repressing chromatin environment (Feng and Zhang 2001; Brehm et al. 2004). Through its interaction with MBD3, a protein related to the methyl-cytosine–binding protein, NuRD is involved in remodeling and deacetylating nucleosomes containing methylated DNA. It is also known that MeCP1 complex, which is composed of 10 major polypeptides, including MBD2, CHD3, and CHD4, and all of the other known NuRD components, preferentially binds, remodels, and deacetylates methylated nucleosomes to cause gene silencing (Zhang et al. 1999). Another chromatin-remodeling protein, Chd1, is known to be a component of yeast SAGA and SILK, two highly homologous and conserved multisubunit histone acetyltransferase complexes (Pray-Grant et al. 2005). Chd1 was shown to specifically interact with the methylated lysine 4 mark on histone H3, which is associated with transcriptional activity. The association of CHD7 with histone deacetylases has not been studied yet. It will be important to study the role of this chromodomain helicase DNA-binding protein in gene transcription and to understand its possible role in gene repression.
In conclusion, CHD7 mutations account for ∼60% of cases of CHARGE syndrome in our study. Cardiovascular malformations, coloboma, and facial asymmetry were seen more often in those with mutations in the CHD7 gene. We also suggest that most individuals with CHARGE syndrome with combined coloboma, choanal atresia, and hypoplastic semicircular canals will have mutations in the CHD7 gene. Functional studies are needed to address the question of how CHD7 mutation alters gene expression, as expected on the basis of its similarity to other chromodomain helicase DNA-binding proteins.
Acknowledgments
We are deeply grateful to all the individuals with CHARGE syndrome and their families who continue to participate in this study. We also thank Benjamin B. Roa for assisting in this study. This work was funded by the Doris Duke Clinical Scientist Developmental Award (to S.R.L.) and National Institutes of Health (NIH) grant RO1HD39056 (to J.W.B.). J.M.G. was supported by SHARE’s Child Disability Center and the Steven Spielberg Pediatric Research Center. In addition, this work was supported by the University of California–Los Angeles Intercampus NIH/National Institute of General Medical Sciences Medical Genetics Training Program grant GM08243 and NIH/National Institute of Child Health and Human Development grant HD22657 from the U.S. Department of Health and Human Services, Public Health Service.
Appendix A
Table A1.
CHD7 Oligonucleotide Sequences
|
Oligonucleotide Sequence(5′→3′) |
|||
| Exon | Forward | Reverse | AnnealingTemperature(°C) |
| 2a | ttcaggtaccagaccagatacg | tacggactgattcattggagt | 60 |
| 2b | accagatacgagccccctac | acagcattggggtatcttgg | 60 |
| 2c | ggcagtattctcgatatccttacag | atcagtcgttcctggattgc | 60 |
| 2d | atgcagcagtctcgtccatt | acagggagattgatgcctga | 60 |
| 3 | gaaacatcagccactaactttca | cccctcatttcataggctgta | 60 |
| 4 | gccaatatgtatggatttatcagttg | caagataggggaggtcttgtg | 60 |
| 5 | gccactgtcttgggtttttg | ccaacattaggtggatgttcc | 60 |
| 6 | gtggtagcaaaggggaatga | tgggttaataaattagacaggattaga | 60 |
| 7 | tgtgctcatatgggagtaaatca | ccaaagtttcttaatttcccaac | 60 |
| 8 | tgttgctcagcagccttaat | atgcaagttgacagcaccaa | 60 |
| 9 | ttttatattgctgtgacccaaaa | gaccaggtctaggattctacca | 60 |
| 10 | gagcatgcttttccttaatgtg | ctccctggaactctccgatt | 60 |
| 11 | tggtaagaattggctgatgg | cacaaatgcatacccaaagg | 60 |
| 12 | agcctttgggtatgcatttg | ccttcccaagtcaccaagac | 60 |
| 13 | gagatctccaaagggataaatacg | gcatcaaattctgagcaacg | 60 |
| 14 | tgcctgattcctatactttgcat | ggttctgtgtactaggtgggaaa | 60 |
| 15 | cactgggctttgaaaaatgaa | caccatgaaatccccagtct | 60 |
| 16 | cagtttgcaatgggttttga | ttccacttttaggtggactgc | 60 |
| 17 | cgccaataaaccctatttgct | ctgagtgacgacgcaacatt | 60 |
| 18 | aggactccaccttcatgaccta | ttaaataaaggaaagtgccagaa | 60 |
| 19 | ggtttactctctttgagaaaaatgc | cccaatgcatcttgtaagca | 60 |
| 20, 21 | cggagcaaatacataaacaaaa | ggggtgtcacacaaattcaa | 60 |
| 22 | caacgctggtacctgacttaaa | acgaggcagggaaatatgg | 60 |
| 23 | gcctcgtgcattaagctctc | cctccaaatctgcgagttct | 60 |
| 24 | aggatgatggatgaacagca | gaaggttctgaaacacaatcca | 60 |
| 25 | cccaccatgctcagatgttt | gccaagagtcctttggaact | 60 |
| 26 | gttgtggcagtgctgtgatt | tgtgtactgcagggtaagaactg | 60 |
| 27, 28 | aaaagtagcactgggcagatt | gaaataaggtgttaagacactgctg | 60 |
| 29 | ccctttcccacactgtcatt | gagcctttctttggtggtca | 60 |
| 30 | ccacccccaaataactacca | tctgtaacacagaagggctca | 55 |
| 31a | gcaacaaagttctatacaaaaagacg | tgagcagatgagatgactgga | 60 |
| 31b | caaaacagaggggcaggtaa | ctcgtgaaaaagagcaggtg | 60 |
| 32 | tgcccaataccattctagcc | tctttgggctttttcagatca | 60 |
| 33 | tttgcatcttgatggatgtatt | gcaaggccagtgaaatataagc | 60 |
| 34 | aagccagcccatatagcagt | ggaggaagctggctttcata | 60 |
| 35 | ttgaaataggacattgtcagagg | attcaaggaaaaggcagagg | 60 |
| 36 | tctgacagttctctttggcatt | gaaggcaggataaaacacttcc | 60 |
| 37 | gaatgcgtgtgtgcgtgtat | ggccaacagaaaataagaaagac | 60 |
| 38a | atttggaatggcaggttcac | tcaggtccattccagcaaac | 60 |
| 38b | gaagaggaagaaggcccaaa | tcctggaggtagaaacatgg | 55 |
| 38c | tgactctgcgaatggatctg | tgaacattaactgtgagtgtaaacag | 60 |
Table A2.
Annotated Transcript List from Microarray Cluster Analysis
| Transcript | Nucleotide Accession Number | Affy | Entrez Gene ID | Description |
| 1 | U17714 | 208667_s_at | 6767 | Suppression of tumorigenicity 13 (colon carcinoma) (Hsp70 interacting protein) |
| 2 | AI742901 | 213889_at | 9487 | Phosphatidylinositol glycan, class L |
| 3 | AA675892 | 202704_at | 10140 | Transducer of ERBB2, 1 |
| 4 | BC004954 | 208929_x_at | 6137 | Ribosomal protein L13 |
| 5 | NM_002607 | 205463_s_at | 5154 | Platelet-derived growth factor alpha polypeptide |
| 6 | NM_001975 | 201313_at | 2026 | Enolase 2 (gamma, neuronal) |
| 7 | AV704017 | 222376_at | … | … |
| 8 | NM_016548 | 217771_at | 51280 | Golgi phosphoprotein 2 |
| 9 | NM_002012 | 206492_at | 2272 | Fragile histidine triad gene |
| 10 | BC004399 | 210652_s_at | 22996 | Chromosome 1 ORF 34 |
| 11 | NM_024843 | 217889_s_at | 79901 | Cytochrome b reductase 1 |
| 12 | NM_005908 | 203778_at | 4126 | Mannosidase, beta A, lysosomal |
| 13 | NM_003272 | 204137_at | 7107 | Transmembrane 7 superfamily member 1 (upregulated in kidney) |
| 14 | BF513474 | 226918_at | … | … |
| 15 | NM_003275 | 203662_s_at | 7111 | Tropomodulin 1 |
| 16 | BC002660 | 203661_s_at | 7111 | Tropomodulin 1 |
| 17 | NM_003794 | 205329_s_at | 8723 | Sorting nexin 4 |
| 18 | AK000208 | 225775_at | 340348 | Hypothetical protein MGC50844 |
| 19 | NM_003254 | 201666_at | 7076 | Tissue inhibitor of metalloproteinase 1 |
| 20 | AV728526 | 212611_at | 23220 | Deltex 4 homolog (Drosophila) |
| 21 | NM_018090 | 220731_s_at | 55707 | FLJ10420 |
| 22 | AI377685 | 235537_at | 54940 | OCIA domain–containing 1 |
| 23 | NM_002371 | 204777_s_at | 4118 | Mal; T-cell differentiation protein |
| 24 | NM_004182 | 218495_at | 8409 | … |
| 25 | NM_004199 | 202733_at | 8974 | Procollagen-proline; 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase) |
| 26 | NM_002513 | 204862_s_at | 4832 | Nonmetastatic cells 3 (protein expressed in) |
| 27 | NM_001349 | 201624_at | 1615 | Aspartyl-tRNA synthetase |
| 28 | CA418310 | 1568611_at | 8974 | Procollagen-proline; 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase) |
| 29 | BG034328 | 203588_s_at | 7029 | Transcription factor Dp-2 (E2F dimerization partner 2) |
| 30 | AB055977 | 223376_s_at | 25798 | Brain protein I3 |
| 31 | NM_024728 | 219655_at | 79783 | Chromosome 7 ORF 10 |
| 32 | AI801879 | 238377_s_at | … | … |
| 33 | BF224279 | 242823_at | … | … |
| 34 | NM_024006 | 217949_s_at | 79001 | Vitamin K epoxide reductase complex, subunit 1 |
| 35 | NM_006992 | 206076_at | 10233 | B7 gene |
| 36 | BG485163 | 224572_s_at | 359948 | Interferon regulatory factor 2–binding protein 2 |
| 37 | NM_002624 | 207132_x_at | 5204 | Prefoldin 5 |
| 38 | AB055804 | 210908_s_at | 5204 | Prefoldin 5 |
| 39 | AF083068 | 209940_at | 10039 | Poly (ADP-ribose) polymerase family, member 3 |
| 40 | NM_006893 | 218253_s_at | 1939 | Ligatin |
| 41 | AW157311 | 226608_at | 388272 | … |
| 42 | NM_006276 | 201129_at | 6432 | Splicing factor, arginine/serine-rich 7 (35 kDa) |
| 43 | AW574664 | 212191_x_at | 6137 | Ribosomal protein L13 |
| 44 | AF116707 | 223162_s_at | 57189 | LCHN protein |
| 45 | AF083033 | 209139_s_at | 8575 | Protein kinase; interferon-inducible double-stranded RNA-dependent activator |
| 46 | BC005871 | 224435_at | 84293 | Chromosome 10 ORF 58 |
| 47 | AF237771 | 222455_s_at | 55742 | Parvin, alpha |
| 48 | AI364950 | 215880_at | 4669 | N-acetylglucosaminidase, alpha (Sanfilippo disease IIIB) |
| 49 | AB030653 | 220228_at | … | … |
| 50 | NM_015974 | 220753_s_at | 51084 | Crystallin, lambda 1 |
| 51 | BE348448 | 241567_at | 8715 | Nucleolar protein 4 |
| 52 | BE220308 | 237826_at | … | … |
| 53 | AW297742 | 243890_at | 5362 | Plexin A2 |
| 54 | BG029530 | 203297_s_at | 3720 | Jumonji; AT-rich interactive domain 2 |
| 55 | AW206458 | 230102_at | … | … |
| 56 | AF279372 | 210740_s_at | 3705 | Inositol 1,3,4-triphosphate 5/6 kinase |
| 57 | AI765312 | 242392_at | 148581 | … |
| 58 | AW016812 | 231644_at | … | … |
| 59 | AF043584 | 216495_x_at | … | … |
| 60 | T79953 | 203143_s_at | 9674 | KIAA0040 |
| 61 | AL135735 | 212506_at | 8301 | Phosphatidylinositol-binding clathrin assembly protein |
| 62 | BF510728 | 229750_at | 5452 | POU domain, class 2, transcription factor 2 |
| 63 | NM_016281 | 220761_s_at | 51347 | TAO kinase 3 |
| 64 | AW070877 | 229147_at | … | … |
| 65 | NM_000118 | 201809_s_at | 2022 | Endoglin (Osler-Rendu-Weber syndrome 1) |
| 66 | BF968097 | 229072_at | … | … |
| 67 | BC001312 | 208639_x_at | 10130 | Thioredoxin domain–containing 7 (protein disulfide isomerase) |
| 68 | NM_001766 | 205789_at | 912 | CD1D antigen, d polypeptide |
| 69 | NM_025159 | 220252_x_at | 80231 | … |
| 70 | NM_016096 | 218059_at | 51123 | HSPC038 protein |
| 71 | NM_014314 | 218943_s_at | 23586 | DEAD (Asp-Glu-Ala-Asp) box polypeptide 58 |
| 72 | BF701166 | 222651_s_at | 7227 | Trichorhinophalangeal syndrome I |
| 73 | AA741307 | 228531_at | 54809 | … |
| 74 | D31765 | 213449_at | 10940 | Processing of precursor 1; ribonuclease P/MRP subunit (Saccharomyces cerevisiae) |
| 75 | AI703496 | 227749_at | … | … |
| 76 | AK026716 | 234111_at | … | … |
| 77 | BC014496 | 1558186_s_at | … | … |
| 78 | NM_002296 | 201795_at | 3930 | Lamin B receptor |
| 79 | AK001166 | 226980_at | 55789 | DEP domain–containing 1B |
| 80 | AW664953 | 227335_at | 85362 | … |
| 81 | BG571343 | 1552334_at | … | … |
| 82 | NM_017654 | 219691_at | 54809 | Sterile alpha motif domain–containing 9 |
| 83 | L21934 | 221561_at | 6646 | Sterol O-acyltransferase (acyl-coenzyme A: cholesterol acyltransferase) 1 |
| 84 | NM_002758 | 205698_s_at | 5608 | Mitogen-activated protein kinase kinase 6 |
| 85 | AK090449 | 1566433_at | 122618 | … |
| 86 | AF124491 | 209876_at | 9815 | G protein-coupled receptor kinase interactor 2 |
| 87 | AW303300 | 225600_at | 286144 | Hypothetical protein LOC286144 |
| 88 | NM_015474 | 204502_at | 25939 | SAM domain and HD domain 1 |
| 89 | AF264784 | 224218_s_at | 7227 | Trichorhinophalangeal syndrome I |
| 90 | AL534702 | 213017_at | 171586 | Abhydrolase domain–containing 3 |
| 91 | AI670947 | 229116_at | 22866 | Connector enhancer of kinase suppressor of Ras 2 |
| 92 | AI348094 | 212956_at | 23158 | KIAA0882 protein |
| 93 | NM_006281 | 204068_at | 6788 | Serine/threonine kinase 3 (STE20 homolog; yeast) |
| 94 | BC006436 | 224518_s_at | 84527 | … |
| 95 | BF446390 | 227208_at | 338657 | Similar to DLNB14 |
| 96 | NM_005931 | 206247_at | 4277 | MHC class I polypeptide-related sequence B |
| 97 | BF222929 | 242346_x_at | … | … |
| 98 | AI306487 | 222849_s_at | 79634 | Secernin 3 |
| 99 | AW470003 | 212149_at | 23167 | KIAA0143 protein |
| 100 | NM_016397 | 220607_x_at | 51497 | TH1-like (Drosophila) |
| 101 | AA551075 | 212188_at | 115207 | Potassium channel tetramerisation domain–containing 12 |
| 102 | NM_014736 | 202503_s_at | 9768 | KIAA0101 |
| 103 | NM_002128 | 200680_x_at | 3146 | High-mobility group box 1 |
| 104 | AB014555 | 38340_at | 9026 | Huntingtin interacting protein-1 related |
| 105 | NM_006565 | 202521_at | 10664 | CCCTC-binding factor (zinc-finger protein) |
| 106 | BF797555 | 212718_at | 10914 | Poly(A) polymerase alpha |
| 107 | NM_006457 | 203243_s_at | 10611 | PDZ and LIM domain 5 |
| 108 | BC005357 | 223608_at | 84288 | … |
| 109 | AB028628 | 203428_s_at | 25842 | ASF1 antisilencing function 1 homolog A (S. cerevisiae) |
| 110 | AA114243 | 244743_x_at | 7697 | Zinc-finger protein 138 (clone pHZ-32) |
| 111 | D86983 | 212013_at | 7837 | Melanoma-associated gene |
| 112 | BF342851 | 212012_at | 7837 | Melanoma-associated gene |
| 113 | BE618656 | 224626_at | 113829 | Solute carrier family 35, member A4 |
| 114 | AK096683 | 1558586_at | 7558 | Zinc-finger protein 11b (KOX 2) |
| 115 | AF043586 | 216430_x_at | 3538 | Immunoglobulin lambda constant 2 (Kern-Oz marker) |
| 116 | AB013384 | 209559_at | 9026 | Huntingtin interacting protein-1 related |
Web Resources
Accession numbers and URLs for data presented herein are as follows:
- Baylor College of Medicine Cardiovascular Genetics, http://www.cardiogene.org/
- ChipST2C, http://chipst2c.org/ChipST2C.html
- GenBank, http://www.ncbi.nlm.nih.gov/Genbank/ (for gene sequences [accession number NM_017780], clones RP11-33I11 [accession number AC113143] and RP11-174G1 [accession number AC022182], and cDNA fragment of KIAA1416 [accession number BC034239])
- Online Mendelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/Omim/ (for CHARGE syndrome)
References
- Amiel J, Attiee-Bitach T, Marianowski R, Cormier-Daire V, Abadie V, Bonnet D, Gonzales M, Chemouny S, Brunelle F, Munnich A, Manach Y, Lyonnet S (2001) Temporal bone anomaly proposed as a major criteria for diagnosis of CHARGE syndrome. Am J Med Genet 99:124–127 [DOI] [PubMed] [Google Scholar]
- Blake KD, Davenport SL, Hall BD, Hefner MA, Pagon RA, Williams MS, Lin AE, Graham JM Jr (1998) CHARGE association: an update and review for the primary pediatrician. Clin Pediatr (Phila) 37:159–173 [DOI] [PubMed] [Google Scholar]
- Blake KD, Ratcliffe JM, Wyse RK (1989) CHARGE association in two monozygous triplets. Int J Cardiol 25:339–341 10.1016/0167-5273(89)90225-8 [DOI] [PubMed] [Google Scholar]
- Brehm A, Tufteland KR, Aasland R, Becker PB (2004) The many colours of chromodomains. Bioessays 26:133–140 10.1002/bies.10392 [DOI] [PubMed] [Google Scholar]
- Clementi M, Tenconi R, Turolla L, Silvan C, Bortotto L, Artifoni L (1991) Apparent CHARGE association and chromosome anomaly: chance or contiguous gene syndrome. Am J Med Genet 41:246–250 10.1002/ajmg.1320410223 [DOI] [PubMed] [Google Scholar]
- De Krijger RR, Mooy CM, Van Hemel JO, Sulkers EJ, Kros JM, Bartelings MM, Govaerts LC (1999) CHARGE association-related ocular pathology in a newborn with partial trisomy 19q and partial monosomy 21q, from a maternal translocation (19;21) (q13.1;q22.3). Pediatr Dev Pathol 2:577–581 10.1007/s100249900165 [DOI] [PubMed] [Google Scholar]
- Devriendt K, Swillen A, Fryns JP (1998) Deletion in chromosome region 22q11 in a child with CHARGE association. Clin Genet 53:408–410 [DOI] [PubMed] [Google Scholar]
- Feng Q, Zhang Y (2001) The MeCP1 complex represses transcription through preferential binding, remodeling, and deacetylating methylated nucleosomes. Genes Dev 15:827–832 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graham JM Jr (2001) A recognizable syndrome within CHARGE association: Hall-Hittner syndrome. Am J Med Genet 99:120–123 [DOI] [PubMed] [Google Scholar]
- Hall BD (1979) Choanal atresia and associated multiple anomalies. J Pediatr 95:395–398 [DOI] [PubMed] [Google Scholar]
- Hittner HM, Hirsch NJ, Kreh GM, Rudolph AJ (1979) Colobomatous microphthalmia, heart disease, hearing loss, and mental retardation—a syndrome. J Pediatr Ophthalmol Strabismus 16:122–128 [DOI] [PubMed] [Google Scholar]
- Irizarry RA, Bolstad BM, Collin F, Cope LM, Hobbs B, Speed TP (2003a) Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res 31:e15 10.1093/nar/gng015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP (2003b) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4:249–264 10.1093/biostatistics/4.2.249 [DOI] [PubMed] [Google Scholar]
- Issekutz KA, Graham JM Jr, Prasad C, Smith IM, Blake KD (2005) An epidemiological analysis of CHARGE syndrome: preliminary results from a Canadian study. Am J Med Genet A 133:309–317 [DOI] [PubMed] [Google Scholar]
- Kaufman MH (1992) The Atlas of mouse development. University of Edinburgh, Edinburgh, United Kingdom [Google Scholar]
- Lalani SR, Safiullah AM, Fernbach SD, Phillips M, Bacino CA, Molinari LM, Glass NL, Towbin JA, Craigen WJ, Belmont JW (2005) SNP genotyping to screen for a common deletion in CHARGE syndrome. BMC Med Genet 6:8 10.1186/1471-2350-6-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lalani SR, Safiullah AM, Molinari LM, Fernbach SD, Martin DM, Belmont JW (2004) SEMA3E mutation in a patient with CHARGE syndrome. J Med Genet 41:e94 10.1136/jmg.2003.017640 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lalani SR, Stockton DW, Bacino C, Molinari LM, Glass NL, Fernbach SD, Towbin JA, Craigen WJ, Graham JM Jr, Hefner MA, Lin AE, McBride KL, Davenport SL, Belmont JW (2003) Toward a genetic etiology of CHARGE syndrome. I. A systematic scan for submicroscopic deletions. Am J Med Genet A 118:260–266 10.1002/ajmg.a.20002 [DOI] [PubMed] [Google Scholar]
- Lee WT, Hou JW, Yau KI, Wang TR (1995) Trisomy 18 in a patient with CHARGE association. J Formosan Med Assoc 94:60–62 [PubMed] [Google Scholar]
- Lev D, Nakar O, Bar-Am I, Zudik A, Watemberg N, Finkelstien S, Katzin N, Lerman-Sagie T (2000) CHARGE association in a child with de novo chromosomal aberration 46, X,der(X)t(X;2)(p22.1;q33) detected by spectral karyotyping. J Med Genet 37:E47 10.1136/jmg.37.12.e47 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lubinsky MS (1994) Properties of associations: identity, nature, and clinical criteria, with a commentary on why CHARGE and Goldenhar are not associations. Am J Med Genet 49:21–25 10.1002/ajmg.1320490106 [DOI] [PubMed] [Google Scholar]
- Mitchell JA, Giangiacomo J, Hefner MA, Thelin JW, Pickens JM (1985) Dominant CHARGE association. Ophthalmic Paediatr Genet 6:271–276 [PubMed] [Google Scholar]
- North KN, Wu BL, Cao BN, Whiteman DA, Korf BR (1995) CHARGE association in a child with de novo inverted duplication (14)(q22→q24.3). Am J Med Genet 57:610–614 10.1002/ajmg.1320570419 [DOI] [PubMed] [Google Scholar]
- Pagon RA, Graham JM Jr, Zonana J, Yong SL (1981) Coloboma, congenital heart disease, and choanal atresia with multiple anomalies: CHARGE association. J Pediatr 99:223–227 [DOI] [PubMed] [Google Scholar]
- Pray-Grant MG, Daniel JA, Schieltz D, Yates JR 3rd, Grant PA (2005) Chd1 chromodomain links histone H3 methylation with SAGA- and SLIK-dependent acetylation. Nature 433:434–438 10.1038/nature03242 [DOI] [PubMed] [Google Scholar]
- Sanlaville D, Romana SP, Lapierre JM, Amiel J, Genevieve D, Ozilou C, Le Lorch M, Brisset S, Gosset P, Baumann C, Turleau C, Lyonnet S, Vekemans M (2002) A CGH study of 27 patients with CHARGE association. Clin Genet 61:135–138 10.1034/j.1399-0004.2002.610208.x [DOI] [PubMed] [Google Scholar]
- Storey JD, Tibshirani R (2003) Statistical significance for genomewide studies. Proc Natl Acad Sci USA 100:9440–9445 10.1073/pnas.1530509100 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Studer M (2001) Initiation of facial motoneurone migration is dependent on rhombomeres 5 and 6. Development 128:3707–3716 [DOI] [PubMed] [Google Scholar]
- Tellier AL, Cormier-Daire V, Abadie V, Amiel J, Sigaudy S, Bonnet D, de Lonlay-Debeney P, Morrisseau-Durand MP, Hubert P, Michel JL, Jan D, Dollfus H, Baumann C, Labrune P, Lacombe D, Philip N, LeMerrer M, Briard ML, Munnich A, Lyonnet S (1998) CHARGE syndrome: report of 47 cases and review. Am J Med Genet 76:402–409 [DOI] [PubMed] [Google Scholar]
- Tellier AL, Lyonnet S, Cormier-Daire V, de Lonlay P, Abadie V, Baumann C, Bonneau D, Labrune P, Lacombe D, Le Merrer M, Nivelon A, Philip N, Briard ML, Munnich A (1996) Increased paternal age in CHARGE association. Clin Genet 50:548–550 [DOI] [PubMed] [Google Scholar]
- Tong JK, Hassig CA, Schnitzler GR, Kingston RE, Schreiber SL (1998) Chromatin deacetylation by an ATP-dependent nucleosome remodelling complex. Nature 395:917–921 10.1038/27699 [DOI] [PubMed] [Google Scholar]
- Verloes A (2005) Updated diagnostic criteria for CHARGE syndrome: a proposal. Am J Med Genet A 133:306–308 [DOI] [PubMed] [Google Scholar]
- Vissers LE, van Ravenswaaij CM, Admiraal R, Hurst JA, de Vries BB, Janssen IM, van der Vliet WA, Huys EH, de Jong PJ, Hamel BC, Schoenmakers EF, Brunner HG, Veltman JA, van Kessel AG (2004) Mutations in a new member of the chromodomain gene family cause CHARGE syndrome. Nat Genet 36:955–957 10.1038/ng1407 [DOI] [PubMed] [Google Scholar]
- Wilkinson DG (1992) In situ hybridization: a practical approach. Oxford University Press, London [Google Scholar]
- Woodage T, Basrai MA, Baxevanis AD, Hieter P, Collins FS (1997) Characterization of the CHD family of proteins. Proc Natl Acad Sci USA 94:11472–11477 10.1073/pnas.94.21.11472 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y, Ng HH, Erdjument-Bromage H, Tempst P, Bird A, Reinberg D (1999) Analysis of the NuRD subunits reveals a histone deacetylase core complex and a connection with DNA methylation. Genes Dev 13:1924–1935 [DOI] [PMC free article] [PubMed] [Google Scholar]