Abstract
Objective: To find new potential biomarkers and establish the patterns for the detection of ovarian cancer. Methods: Sixty one serum samples including 32 ovarian cancer patients and 29 healthy people were detected by surface-enhanced laser desorption/ionization mass spectrometry (SELDI-MS). The protein fingerprint data were analyzed by bioinformatics tools. Ten folds cross-validation support vector machine (SVM) was used to establish the diagnostic pattern. Results: Five potential biomarkers were found (2085 Da, 5881 Da, 7564 Da, 9422 Da, 6044 Da), combined with which the diagnostic pattern separated the ovarian cancer from the healthy samples with a sensitivity of 96.7%, a specificity of 96.7% and a positive predictive value of 96.7%. Conclusions: The combination of SELDI with bioinformatics tools could find new biomarkers and establish patterns with high sensitivity and specificity for the detection of ovarian cancer.
Keywords: Ovarian cancer, SVM, Diagnosis, SELDI-TOF, Proteomics
INTRODUCTION
Ovarian cancer is the most lethal gynecologic malignancy. Poor survival rates are mainly attributable to late diagnosis. Most patients at diagnosis have advanced stage disease. The 5-year survival rate for late clinical stage ovarian cancer is only 25%, but for early stage disease, the survival rate can be as high as 90%. CA125, the most widely used biomarker for ovarian cancer, does not have a satisfying positive predictive value. In early stage ovarian cancer, 40%–50% patients are CA125 negative, with high serum CA125 being seen in many benign gynecologic diseases and other types of cancer (Bast et al., 1998). Therefore, there is urgent need for new biomarkers for ovarian cancer.
A novel proteomic approach called surface enhanced laser desorption/ionization time-of-flight mass spectrometry (SELDI-TOF MS) and ProteinChip technology has been developed for the detection of early stage cancer (Petricoin and Liotta, 2004; Srinivas et al., 2002; Wiesner, 2004). SELDI-TOF MS combined with bioinformatics approach has successfully found some new biomarkers and achieved high sensitivity and specificity for the diagnosis of prostate (Adam et al., 2002), breast (Li et al., 2002; Hu et al., 2004), colorectal cancer (Chen et al., 2004; Yu et al., 2004), liver cancer (Poon et al., 2002) and so on.
This study project aimed at finding potential biomarkers in ovarian cancer and establishing the patterns for diagnosis of ovarian cancer.
MATERIALS AND METHODS
A total of 61 serum samples were obtained from the serum banks of the affiliated tumor hospital of Guangxi Medical University. The cancer group consisted of 31 serum samples from ovarian cancer patients at different clinical stages (the International Federation of Gynecology and Obstetrics (FIGO) staging system): FIGO I (n=3), FIGO II (n=5), FIGO III (n=13), FIGO IV (n=11). The median age of the ovarian cancer patients was 57 years (range, 14–68 years). The control group consisted of 29 serum samples from healthy women who were age and sex matched with the cancer group. Diagnoses were pathologically confirmed, and specimens were obtained before treatment. All samples were obtained in early morning and stored at −80 °C until use.
SELDI protein profiling
Serum samples in ice were thawed and centrifuged at 3000 rpm for 5 min at 4 °C, and supernatants were retained. Ninety μl of 5 g/L CHAPS (Sigma, USA) (pH 7.4) was added into PBS to make up 10 μl of each serum sample, and vortex-mixed. The diluted samples were added to 100 μl Cibacron Blue 3GA (Sigma, USA) (previously equilibrated thrice with 5 g/L CHAPS) in 96 well cell culture plate and agitated on a platform shaker at 4 °C for 60 min. After centrifugation at 1000 rpm, 50 μl supernatants were sampled and further diluted by 150 μl 20 mmol/L HEPES (pH 7.4) and applied to each well of a bioprocessor (Ciphergen Biosystems) containing hydrophobic surface (H4) chips previously activated with 20 mmol/L HEPES. The bioprocessor was then sealed and agitated on a platform shaker for 60 min at 4 °C. The excess serum mixtures were discarded, and the chips were washed thrice with 20 mmol/L HEPES and twice with deionized water. The chips were then removed from the bioprocessor and air-dried. Before SELDI analysis, 0.5 μl of a saturated solution of α-cyano-4-hydroxycinnamic acid (CHCA) in 0.5 L/L acetonitrile, and 5 ml/L trifluoroacetic acid was applied twice onto each chip, then air-dried.
Chips were detected on the Protein Biological System II (PBS-II) plus mass spectrometer reader (Ciphergen Biosystems). Data were collected by averaging 65 laser shots with an intensity of 135, a detector sensitivity of 7, a highest mass of 30000 Da and an optimized range of 2000–20000 Da. Mass accuracy was calibrated to less than 0.1% using the All-in-1 peptide molecular mass standard (Ciphergen Biosystems).
Bioinformatics analysis
The spectra intensities of all samples were normalized to the total ion current of mass to charge ratios (m/z) between 2000 and 30000 Da. Noise was filtered from the spectra and peaks were detected with an automatic peak detection pass. Peak clusters were completed to cluster the peaks in different samples with similar masses (defined by a mass window of 0.3% mass error). All these were performed using ProteinChip Software 3.1 (Ciphergen). The peak intensities were preprocessed by scaling all the data to the range [−1, 1].
SVM classifier
SVM is a new machine learning approach originally proposed and developed by Vapnik (1995). SVM applications are being actively pursued in various areas recently, from face recognition to genomics. It is a powerful tool for analyzing complex data derived from SELDI-MS. We constructed a non-linear SVM classifier with a radial based function (RBF) kernel, and with the parameter Gamma 0.6, being the cost of the constrain violation 19 to discriminate the different groups. Ten folds cross-validation approach was applied to estimate the accuracy of the classifier. This approach randomly selected the 9/10 of all the samples to be the blinded training set, and the remaining 1/10 samples to be the test set and repeated the procedure 10 times. SVM classifier is based on the shareware program OSU_SVM v.3.00 Toolbox of Junshui Ma and Yi Zhao.
Feature selection and model establishment
The power of each peak in discriminating different groups was estimated by receive option curve (ROC). The greater area under the curve value of the peak shows the higher relative importance value of the ability to accurately distinguish the different groups. The peaks with lower area under the curve values are excluded. To further select the set of candidate biomarkers, a stepwise approach was used for training many SVMs. The top 1 peak which had the highest ability to predict the two groups (having the highest area under curve values) was selected as single input to build the SVM. The discriminating ability of this SVM was estimated by the accuracy of blind test set. Then, the top 2 peaks were inputted to the SVM and the accuracy was calculated. The following peaks were added in input stepwise fashion to train the SVM and the accuracy was calculated. In this way, many models with different peaks were built. The peaks inputted to the model with highest accuracy were selected as the set of potential biomarkers. And the SVMs with the highest accuracy were selected for detecting ovarian cancer.
RESULTS
After filtrating noise by Ciphergen ProteinChip Software 3.1, 220 peaks were detected. The peaks were 2 kDa to 30 kDa. Peaks with m/z<2 kDa were mainly ion noise from the matrix and therefore excluded.
The 220 qualified peaks detected from the two groups were ranked by ROC. The top 10 peaks with higher area under curve values were selected for further analysis. The top 5 peaks were finally selected as potential biomarkers by using the stepwise approach. SVMs combined with different peaks achieved highest accuracy of 96.7%. The accuracies of these 10 models are plotted in Fig.1.
Fig. 1.
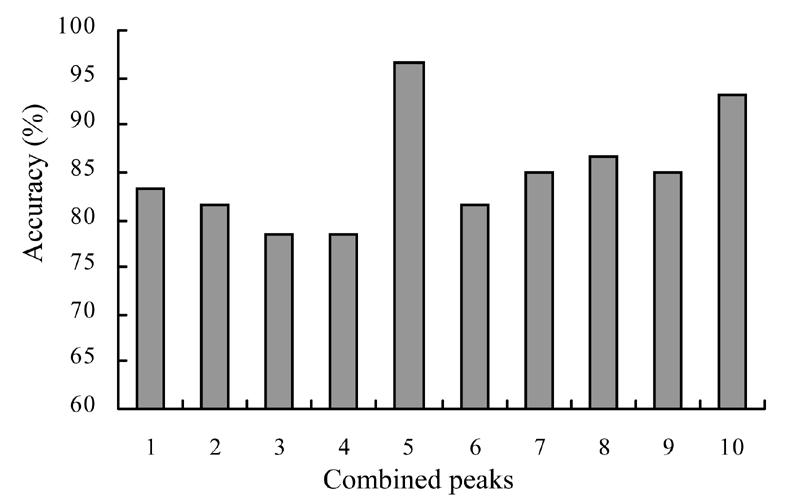
The accuracies of SVMs combined with different peaks
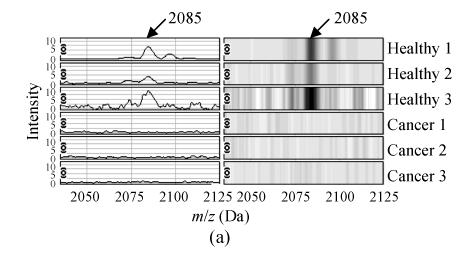
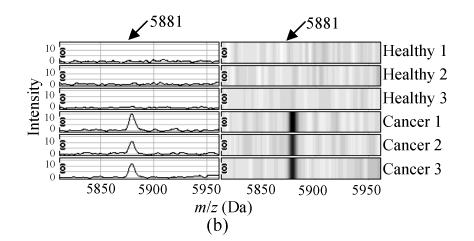
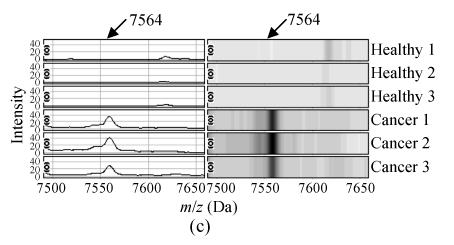
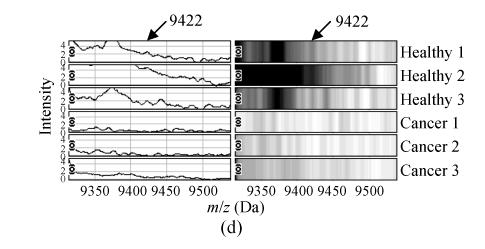
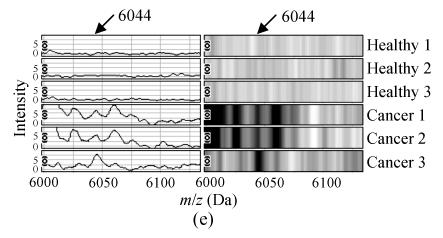
The m/z of the 5 candidate biomarkers were 2085 Da, 5881 Da, 7564 Da, 9422 Da, 6044 Da. The peaks with m/z of 5881 Da, 7564 Da, 6044 Da were highly expressed in ovarian cancer but weakly expressed in healthy people, as shown in Figs.2b, 2c and 2e, but the peaks with m/z of 2085 Da, 9422 Da appeared to be expressed in a contrary way, as shown in Figs.2a and 2d. In the two groups, the P values of t-tests and the area under the ROC curve showed the statistical significance of all the 5 peaks. Table 1 gives the descriptive statistics of the 5 peaks.
Fig. 2.
The spectra and gel maps of potential biomarkers. (a) The spectrum and the gel map of 2085 Da; (b) The spectrum and the gel map of 5881 Da; (c) The spectrum and the gel map of 7564 Da; (d) The spectrum and the gel map of 9422 Da; (e) The spectrum and the gel map of 6044 Da
Table 1.
The statistics of the candidate biomarkers
| m/z | AUC | P value (×10−5) | Healthy | Cancer | Mean S/N* of healthy | Mean S/N* of cancer |
| 2085 | 0.89 | 0.03 | 5.64±3.20 | 1.41±2.91 | 4.61 | 1.00 |
| 5881 | 0.87 | 0.06 | 1.73±1.28 | 6.55±4.41 | 1.67 | 6.13 |
| 7564 | 0.86 | 0.11 | 0.52±0.90 | 13.32±17.27 | 0.61 | 20.7 |
| 9422 | 0.80 | 5.26 | 2.20±1.00 | 1.21±0.75 | 4.02 | 2.08 |
| 6044 | 0.79 | 13.00 | 0.38±0.57 | 1.88±2.06 | 0.43 | 1.52 |
S/N: Signal/noise
The 5 peaks were combined and evaluated by 10 folds cross-validation SVM. The SVM was trained with 55 samples and tested with the remaining 6 samples. This procedure was repeated 10 times. For the 10 folds cross-validation SVM, the estimated specificity of the test sets was 96.7%, the estimated sensitivity was 96.7%, the estimated positive predictive value was 96.7%. Table 2 shows the results for this classifier.
Table 2.
The predicted results of 10 folds cross-validation SVM (the sample size was the sum of 10 times training and test)
| Test set (6 cases×10) |
Training set (55 cases×10) |
|||||
| Healthy | Cancer | Sum | Healthy | Cancer | Sum | |
| Healthy | 29 | 1 | 30 | 231 | 29 | 260 |
| Cancer | 1 | 29 | 30 | 20 | 270 | 290 |
| Sum | 30 | 30 | 60 | 251 | 299 | 550 |
| Specificity | 96.7% (29/30) | 88.8% (231/260) | ||||
| Sensitivity | 96.7% (29/30) | 93.1% (270/290) | ||||
| Positive predictive value | 96.7% (29/30) | 90.3% (270/299) | ||||
DISCUSSION
The most commonly used biomarker for clinical screening and prognosis in patients with ovarian cancer is CA125. Serum CA125 levels are elevated in 80% of patients with advanced-stage epithelial ovarian cancer but are increased in only 50%–60% of patients with early-stage disease. With a cutoff of 30–35 units/ml, serum CA125 has been shown to have a sensitivity of only 50%–60%. Mok et al.(2001) reported that CA125 had a sensitivity of 64.9% and specificity of 94%. Rai et al.(2002) reported that the recommended cutoff of 35 units/ml for CA125 resulted in 65.6% sensitivity and 97.2% specificity.
Because of the multi-factorial nature of ovarian cancer, it is very clear that the combination of several markers is necessary to effectively detect and diagnose ovarian cancer. SELDI-MS and the ProteinChip technology coupled to sophisticated bioinformatics tools for complex data analysis will find the “fingerprints” of ovarian cancer and build the diagnosis model. Petricoin et al.(2002) first reported that SELDI profiling coupled to a learning algorithm that compared combination of five protein peaks led to a sensitivity of 100% and specificity of 95% in differentiating ovarian cancer. Ye et al.(2003) found a serum biomarker at 1700 Da up-regulated in cancer by SELDI which was identified to be haptoglobin-subunit., Zhang et al.(2004) reported that the SELDI pattern of three biomarkers was able to diagnose early stage ovarian cancer with 74% sensitivity at 97% specificity.
We developed the integrated approach of bioinformatics and biostatistics tools to analyze the large data of spectra. The ROC curve was applied to rank and select the peaks according to their contribution to the separation of two groups. For further estimating the effect of the multi-peak combination, the stepwise method built many models with varied peaks combination. The peaks combination having the highest accuracy would be selected as the potential biomarkers. To accurately estimate the sensitivity and specificity of the classifiers, the test sets were randomly selected many times, and separated from training sets each time.
In all the test sets, only two samples were inaccurately predicted. One was a healthy woman and the other was a patient of ovarian cancer (IV stage). All the patients of the early stage were accurately diagnosed. It showed the pattern is effective in early screening.
Sixty-one ovarian cancer patients and healthy women were detected by SELDI-MS and the complex data were analyzed by an SVM classifier. Five potential biomarkers were found, the diagnostic pattern was established, and specificity of 96.7%, sensitivity of 96.7%, positive predictive value of 96.7% were achieved.
The 15 ovarian cancer samples, which were collected in one week postoperatively, were also detected. The results showed no remarkable difference between the preoperative samples and postoperative ones, and demonstrated that the proteomic fingerprint had few changes in one week postoperatively.
More samples should be collected to validate the model and identify the selected biomarkers. The biomarkers were purified by fractionation and SDS-PAGE, and identified by tryptic digestion and online database searching.
In conclusion, the approach applying SELDI-MS and ProteinChip technology in combination with sophisticated bioinformatics tools can facilitate the discovery of new biomarkers and establish patterns with high sensitivity and specificity for the detection of ovarian cancer.
Footnotes
Project (No. G1998051200) supported by the National Basic Research Program (973) of China
References
- 1.Adam B, Qu Y, Davis JW, Ward MD, Clements MA, Cazares LH, Semmens OJ, Schellhammer PF, Yasui Y, Ziding F, et al. Serum protein fingerprinting coupled with a pattern-matching algorithm distinguishes prostate cancer from benign prostate hyperplasia and healthy men. Cancer Res. 2002;62(13):3609–3614. [PubMed] [Google Scholar]
- 2.Bast RCJr, Xu FJ, Yu YH, Barnhill S, Zhang Z, Mills GB. CA125: the past and the future. Int J Biol Markers. 1998;13(4):179–187. doi: 10.1177/172460089801300402. [DOI] [PubMed] [Google Scholar]
- 3.Chen YD, Zheng S, Yu JK, Hu X. Application of serum protein pattern model in diagnosis of colorectal cancer. Zhonghua Zhong Liu Za Zhi. 2004;26(7):417–420. (in Chinese) [PubMed] [Google Scholar]
- 4.Hu Y, Zhang SZ, Yu JK, Liu J, Zheng S. Detection and evaluation of serum proteomic patterns by SELDI-TOF-MS in breast cancer. Zhong Hua Jian Yan Zha Zhi. 2004;27(10):646–648. (in Chinese) [Google Scholar]
- 5.Li J, Zhang Z, Rosenzweig J, Wang YY, Chan DW. Proteomics and bioinformatics approached for identification of serum biomarkers to detect breast cancer. Clin Chem. 2002;48(8):1296–1304. [PubMed] [Google Scholar]
- 6.Mok SC, Chao J, Skates S, Wong K, Yiu GK, Muto MG, Berkowitz RS, Cramer DW. Prostasin, a potential serum marker for ovarian cancer: identification through microarray technology. J Natl Cancer Inst. 2001;93(19):1458–1464. doi: 10.1093/jnci/93.19.1458. [DOI] [PubMed] [Google Scholar]
- 7.Petricoin EF, Liotta LA. SELDI-TOF-based serum proteomic pattern diagnostics for early detection of cancer. Curr Opin Biotechnol. 2004;15(1):24–30. doi: 10.1016/j.copbio.2004.01.005. [DOI] [PubMed] [Google Scholar]
- 8.Petricoin EF, Ardekani AM, Hitt BA, Levine PJ, Fusaro VA, Steinberg SM, Mills GB, Simone C, Fishman DA, Kohn EC, et al. Use of proteomic patterns in serum to identify ovarian cancer. Lancet. 2002;359(9306):572–577. doi: 10.1016/S0140-6736(02)07746-2. [DOI] [PubMed] [Google Scholar]
- 9.Poon TCW, Yip TT, Chan ATC, Yip C, Yip V, Mok TSK, Lee CCY, Leung TWT, Ho SKW, Johnson PJ. Comprehensive proteomic profiling identifies serum proteomic signatures for detection of hepatocellular carcinoma and its subtypes. Clin Chem. 2002;49(5):752–760. doi: 10.1373/49.5.752. [DOI] [PubMed] [Google Scholar]
- 10.Rai AJ, Zhang Z, Rosenzweig J, Shih IeM, Pham T, Fung ET, Sokoll LJ, Chan DD. Proteomic approaches to tumor marker discovery. Arch Pathol Lab Med. 2002;126(12):1518–1526. doi: 10.5858/2002-126-1518-PATTMD. [DOI] [PubMed] [Google Scholar]
- 11.Srinivas PR, Verma M, Zhao Y, Srivastava S, Clark RA, Tockman MS. Proteomics for cancer biomarker discovery. Clin Chem. 2002;48(8):1160–1169. [PubMed] [Google Scholar]
- 12.Vapnik V. The Nature of Statistical Learning Theory. New York: Springer Verlag; 1995. [Google Scholar]
- 13.Wiesner A. Detection of tumor markers with ProteinChip technology. Curr Pharm Biotechnol. 2004;51:45–67. doi: 10.2174/1389201043489675. [DOI] [PubMed] [Google Scholar]
- 14.Ye B, Cramer DW, Skates SJ, Gygi SP, Pratomo V, Fu L, Horick NK, Licklider LJ, Schorge JO, Berkowitz RS, et al. Haptoglobin-alpha subunit as potential serum biomarker in ovarian cancer: identification and characterization using proteomic profiling and mass spectrometry. Clin Cancer Res. 2003;9(8):2904–2915. [PubMed] [Google Scholar]
- 15.Yu JK, Chen YD, Zheng S. An integrated approach to the detection of colorectal cancer utilizing proteomics and bioinformatics. World J Gastroenterol. 2004;10(21):3127–3131. doi: 10.3748/wjg.v10.i21.3127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zhang Z, Bast RCJr, Yu Y, Li J, Sokoll LJ, Rai AJ, Rosenzweig JM, Cameron B, Wang YY, Meng XY, et al. Three biomarkers identified from serum proteomic analysis for the detection of early stage ovarian cancer. Cancer Res. 2004;64(16):5882–5890. doi: 10.1158/0008-5472.CAN-04-0746. [DOI] [PubMed] [Google Scholar]