Abstract
A collection of Rhizobium etli promoters was isolated from a genomic DNA library constructed in the promoter-trap vector pBBMCS53, by their ability to drive the expression of a gusA reporter gene. Thirty-seven clones were selected, and their transcriptional start-sites were determined. The upstream sequence of these 37 start-sites, and the sequences of seven previously identified promoters were compared. On the basis of sequence conservation and mutational analysis, a consensus sequence CTTGACN16–23TATNNT was obtained. In this consensus sequence, nine on of twelve bases are identical to the canonical Escherichia coli σ70 promoter, however the R.etli promoters only contain 6.4 conserved bases on average. We show that the R.etli sigma factor SigA recognizes all R.etli promoters studied in this work, and that E.coli RpoD is incapable of recognizing them. The comparison of the predicted structure of SigA with the known structure of RpoD indicated that regions 2.4 and 4.2, responsible for promoter recognition, are different only by a single amino acid, whereas the region 1 of SigA contains 72 extra residues, suggesting that the differences contained in this region could be related to the lax promoter recognition of SigA.
INTRODUCTION
The Escherichia coli RNA polymerase holoenzyme consists of a core enzyme (α2ββ′), and one sigma factor (σ) subunit, which directs the core enzyme to specific sites on DNA to initiate transcription. In general, bacteria contain a main or housekeeping sigma factor, also known as sigma-70 (σ70) which controls the expression of most of the genes, in association with the RNApol core (1–4). Sequence alignments of several σ70 factors from different bacteria reveal four conserved regions (1 to 4), that have been further divided into subregions, involved in different aspects of transcription initiation, including promoter recognition, promoter melting, initiaton of RNA syntesis, abortive initiation and promoter escape (5).
The σ70 subunit of E.coli and Bacillus subtilis recognizes two types of promoter sequences: the first one, or ‘canonical promoter’, consisting of two hexamer sequences [TTGACA] and [TATAAT], centered respectively in the positions −10 and −35 relative to the transcription start-site (6,7). A helix–turn–helix motif located in the C-terminus of σ70 (subregion 4.2) recognizes the −35 box of the promoter, whereas a more centrally located α-helix (subregion 2.4) recognizes the −10 box (8). The second one, named ‘extended−10 promoter’ consists of an element [TATAAT] plus 2 nt [TG], located in the −15 and −14 positions. In this promoter type a −35 region or equivalent sequence does not exist. The N-terminal part of the subregion 3.0 (previously named 2.5) from σ70 is involved in the recognition of the conserved [TG] motif (9–11).
Rhizobium etli is a soil α-proteobacterium with the ability to colonize the roots of bean-plants to form nitrogen fixing nodules. The whole genomic sequence of R.etli CFN42 was recently concluded [(12); GeneBank accession no CP000133], and consists of a circular chromosome and six large plasmids, with an average G + C content of 61.5%. R.etli contains a large number of sigma factors (one housekeeping σ70 gene, two copies of rpoH, two copies of rpoN and 18 genes of the extracitoplasmic factor group), a feature shared with other nitrogen fixing organisms, like Bradyrhizobium japonicum, Mesorhizobium loti and Sinorhizobium meliloti (4). The physiological roles and the mechanisms of promoter recognition by these sigma factors remain unknown. Moreover, in these organisms only a few well-characterized promoters are available.
The R.etli housekeeping σ70 gene (sigA) encodes a protein of 685 amino acid residues with a molecular weight of 77.18 kDa. The primary structure of SigA is very similar to RpoD (the σ70 factor of E.coli), especially in regions 2, 3 and 4, however SigA contains an extended region 1 (13).
S.meliloti, Caulobacter crescentus and Rhodobacter sphaeroides also have a larger SigA region 1, and most of their σ70 promoters that have been characterized are not transcribed by the E.coli RNApol in vivo or in vitro, but in contrast, the RNApol of these bacteria can initiate transcription of typical E.coli σ70 promoters (13–16). This observation suggests some differences between the transcriptional machineries of E.coli and the α-proteobacteria, perhaps at the level of promoter recognition by the σ70 factor.
The knowledge of the promoter structures and their regulation in a particular organism is the first step towards the molecular understanding of the regulatory networks, an essential prerequisite to develop tools for genetic engineering.
To analyze the molecular basis of R.etli gene expression in this study, we identified, characterized and sequenced active promoters in R.etli under exponential growth conditions. A consensus promoter sequence was deduced, and possesses some differences to the promoter consensus recognized by σ70 in E.coli and B.subtilis. We showed that the double helix stability and the stacking energy of the R.etli and E.coli promoter regions possess a minimal energy value around −10 box, suggesting that despite the differences in their promoter structures, the promoters of both bacteria preserve thermodynamic properties, which favor DNA denaturation and help the transcription initiation. We demonstrated that SigA specifically recognizes the R.etli promoters studied in this work, but in contrast, the E.coli RNApol holoenzyme is incapable of recognizing them. Finally, we propose that R.etli contains a less strict σ70 that recognizes a lax consensus promoter structure.
MATERIALS AND METHODS
Bacterial strains and growth conditions
The bacterial strains and plasmids used in this work are listed in Table 1. E.coli strains were grown at 37°C in Luria–Bertani (LB) medium. R.etli strains were grown at 30°C in PY medium (17) or minimal medium containing 1.2 mM K2HPO4, 0.8 mM MgSO4, 10 mM succinic acid, 10 mM NH4Cl, 1.5 mM CaCl2 and 0.0005% FeCl3, with the pH adjusted to 6.8 (18). Microaerobic conditions during growth were set up by putting the cultures in 150 ml bottles that were flushed with several volumes of a 1% oxygen-99% argon mixture and closed with an airtight stopper. Antibiotics were added at the following final concentrations (in µgml−1): gentamicin 30; chloramphenicol 25; ampicillin 100; nalidixic acid 20. When it was used, X-Gluc (5-Br-4-Cl-3-indolyl-β-D-glucuronic acid, Research Organics) was added at a final concentration of 20 µgml−1.
Table 1.
Bacterial strains and plasmids used
| Bacterial strains/plasmids | Relevant characteristics | Source/references |
|---|---|---|
| Strains | ||
| Rhizobium etli | ||
| CFN42 | Streptomycin resistant, nodulates Phaseolus vulgaris | (17) |
| Escherichia coli | ||
| DH5α | Host strain for plasmids | (47) |
| BW21038 | gusA derivarive | (48) |
| DH10B | lacZ derivative | GibcoBRL |
| BL21(DE3) | Host strain used for expression of target proteins | (22) |
| Plasmids | ||
| pBBR1MCS-5 | Gentamycin resistant, cloning vector replicable in R.etli | (49) |
| pBBMCS53 | pBBR1MCS-5 derivative carrying a promoter minus β-glucuronidase gene (gusA) | (20) |
| pRK415 | Tetracycline resistant, derivative of pRK290 | (50) |
| pLysS | Chloramphenicol resistant, production of lysozyme for repression of T7 polymerase | (22) |
| pET-28a(+) | Kanamycin resistant, expression vector | Novagen |
| pRKS70 | pRK415 derivative carrying a 1726 bp fragment containing the sigA gene of the CFN42 strain | This work |
| pGUSphemC | pBBMCS53 derivative carrying a 278 bp fragment containing the native promoter of hemC gene. The fragment was generated by PCR and carries 57 bp upstream of gene and 221 bp of hemC | This work |
| pGUSphemC-10 | A pGUSphemC derivative in which the promoter element −10 was changed from [TACGAG] to [GCGGAG] | This work |
| pGUSphemC-35 | A pGUSphemC derivative in which the promoter element −35 was changed from [ATTGCG] to [AAGACG] | This work |
| pGUSphemC-1035 | A pGUSphemC derivative, but with mutated −10 and −35 elements from the promoter. Both elements were changed to the sequences indicated before | This work |
DNA and RNA isolation and manipulation
Genomic DNA was isolated using components and instructions from GenomicPrep cells and tissue DNA isolation kit (Amersham). Plasmid DNA was isolated as described by Sambrook et al. (19). DNA was restricted and ligated under the conditions specified by the enzyme manufacturer (Invitrogen). Taq DNA polymerase (Altaenzymes) was used for PCR. The PCR products were cloned using a pMOSblue blunt-ended kit (Amersham). RNA was isolated using components and instructions from the High pure RNA isolation kit (Roche).
Construction of a R.etli promoter library
To isolate DNA fragments containing promoters, 10 mg chromosomal DNA of CFN42 strain was disrupted by nebulization during 2 min to 10 psi. Fragments ranging from 0.5 to 1 kb were purified from an agarose gel, blunted with Klenow and T4 pol enzymes, and ligated into the alkaline phosphatase treated SmaI sites of the promoter-trap vector pBBMCS53, which contains a promoterless gusA gene (20). The ligation mixture was electropored into DH10B using the instructions from ElectroMAX DH10B cell kit (GibcoBRL). Single colonies were selected in LB plates containing gentamicin.
Bacterial matings
pBBMCS53 derivatives were introduced by conjugation into R.etli CFN42 using pRK2013 as a helper plasmid. Strains were grown in PY liquid medium to a stationary phase, mixed in a proportion of donor–helper–receptor of 1:1:2 on PY plates and incubated at 30°C overnight. Cells were resuspended in fresh PY medium, and serial dilutions were plated in PY nalidixic acid 20, gentamicin 30 and X-Gluc. Clones containing fragments with a promoter were selected by blue/white screening of transconjugants.
β-Glucuronidase activity measurements
Overnight cultures of the strains carrying the desired constructs were grown in the presence of the appropriate selection. Cultures were diluted in fresh PY medium to an OD540 of 0.01 and grown to a final OD540 of 0.4. The culture (1 ml) was centrifuged and resuspended in a salt wash solution supplemented with chloramphenicol (100 mgml−1). Quantitative β-glucuronidase assays were performed with p-nitrophenyl glucuronide substrate as described previously (21). Data were normalized to total-cell protein concentration by the Lowry method (19). The results presented in the figures are the mean of three independent experiments.
Transcription start-sites determination
Transcription start-sites were mapped by means of 5′ RACE kit version 2.0 (Invitrogen). Briefly, DNA single chains (scDNA) were synthesized from 5 to 10 mg of total RNA using a gene-specific primer (GUS-LW5, CGATCCAGACTGAATGCCCAC) complementary to the region located in the position 96 to117 from gusA, and using a derivative of Moloney Murine Leukemia Virus Reverse Transcriptase (M-MLV RT). After first strand cDNA synthesis, the original mRNA template was removed by treatment with a RNase Mix (mixture of RNase H and RNase T1) and the scDNA was purified using a GlassMax DNA column. A homopolymeric tail was then added to the 3′ end of the scDNA using Terminal deoxynucleotidil transferase (TdT) and dCTP. In the next step, a PCR amplification was accomplished using Taq DNA polymerase, a second antisense primer (GUS-LW4, GTAACATAAGGGACTGACCTGC) that was complementary to gusA in the position 28 to 49, and a complementary homopolymer tail primer (AAP, GGCCACGCGTCGACTAGTACGGGIIGGGIIGGGIIG). When secondary bands were obtained, an additional PCR was done using an AAP primer and a third antisense primer (GUS-LW2, GCTTGGCGTAATCATGGTCAT), that was complementary to the DNA region located in the position 1 to 21 from gusA.
DNA sequencing
DNA sequencing reactions were performed using the GUS-LW primers and Big-Dye terminator kit version 3.1 in an automatic 310 DNA sequencer (Applied Biosystems).
Construction of promoter mutations
Mutant versions of the phemC and lac promoters were constructed by PCR using oligonucleotides carrying altered versions of the wild-type −35, −10 and both boxes simultaneously. Each PCR product was cloned in to pBBMCS53 vector and introduced into R.etli CFN42 by triparental conjugation.
Overexpression and purification of the R.etli SigA protein
A DNA fragment corresponding to the full-length sigA gene of R.etli was PCR amplified from total DNA, and cloned into pET-28a(+) (Novagen) to express sigA in BL21 (DE3)plysS strain (22). The induction in exponential growth phase (OD540 = 0.4) was obtained by the addition of isopropyl-β-d-thiogalactopyranoside (IPTG) up to the final concentration 1 mM. Cells were harvested after 4 h of cultivation and stored at −70°C. A total of 10 ml of culture usually yielded 2.5–3 mg of protein. Histidine tagged sigma protein was purified using HisTrap Kit (Amersham Biosciences). The cell pellet was suspended in 0.5 ml of binding buffer, containing 8 M urea and 0.25 mM phenylmethlysulfonyl fluoride (PMSF) [1× phosphate buffer (pH 7.4), 20 mM imidazole], and incubated 30 min at room temperature with gentle mixing. After sedimentation of cell debris, lyzate was loaded on to Ni-column and equilibrated with a binding buffer. The desired protein was eluted by phosphate buffer, containing 400 mM imidazol, collected in 0.5 ml fractions, and analyzed in 10% SDS–PAGE. The applied procedure provides protein of 95% purity. To avoid the influence of six-aminoacide histidine tagg, HisTrap purified protein was treated by thrombin (Thrombin CleanCleave Kit, Sigma) after buffer exchange by dialysis. After the thrombin treatment protein was dialyzed against TGED buffer [10 mM Tris–HCl (pH 7.6); 0.1 mM EDTA; 0.1 mM DTT; 0.1 M NaCl; 5% (v/v) glycerol] and storage buffer [10 mM Tris–HCl (pH 7.6); 0.1 mM EDTA; 0.1 mM DTT; 10 mM MgCl2; 0.2 M KCl; 50% (v/v) glycerol]. The final concentration of protein was evaluated by method of Bradford (Bio-Rad protein assay kit).
DNA binding shift assay
Linear DNA templates were randomly labeled with [α-P32]dCTP in a standard PCR, purified from 3.5% PAGE and used for DNA–protein complex formation. RNA polymerase was assembled from core and σ70 protein as it was described by Tang et al. (23). Binary complexes were allowed to preform for 20 min at 30°C, challenged by heparin (200 mg/ml) and immediately loaded on to 3.5% native PAGE, prewarmed up to the same temperature. After electrophoresis, gels were dried and radioautographed.
Software tools
Multiple DNA alignments were done using the programs CLUSTALW version 1.8.1 (24) and MultiAlin version 5.4.1 (25). Alignments were visually inspected and adapted with respect to the known transcriptional start-sites. Consensus matrices were generated and selected using WCONSENSUS version v5c (26). Briefly, to identify the −10 box, the first 20 bp upstream from the transcription start-site (+1) were used. To identify the −35 box, the 20 bp upstream to the −20 position were used as an input for WCONSENSUS. Algorithms available from server http://www.icgeb.trieste.it/dna (27) were used to evaluate double helix stability (28) and stacking energy (29). Calculations for each parameter were carried out with a window size of 10 and 20 nt, providing similar results. For each promoter position, the average value was calculated for the whole set of analyzed sequences.
RESULTS
Isolation of R.etli promoter sequences
To determine the general structure of the R.etli promoters, a random genomic library with an average insert size of 0.5 kb was constructed in the promoter-test vector pBBMCS53 and introduced by conjugation into R.etli CFN42. Direct screening on PY plates containing X-Gluc allowed the selection of transconjugants with an active reporter gene (gusA), suggesting that the constructs contained R.etli promoter regions. The β-glucuronidase activity of 100 transconjugans was determined in PY liquid cultures in exponential and stationary phases of growth. Sixty-eight of these clones showed β-glucuronidase activity under both conditions, while the remaining 32 transconjugants expressed the reporter gene only in PY plates, and for this reason were not used in further experiments. The ability to express the reporter gene of the 68 strains described above was tested in different conditions: low oxygen (1%), minimal medium (MM) and at different temperatures (30, 42 or −10°C). All but three expressed the reporter gene in all conditions, but at different levels, suggesting that the transconjugants carry constructs with constitutive promoters. The DNA sequences of 36 inserts of these constructs were determined. These promoter sequences, plus seven R.etli promoters experimentally identified in other studies were selected for subsequent analyses (30–35).
Thirty-three promoters were mapped within intergenic regions; eight within the coding regions of the immediately preceeding gene, and three in DNA sections that contained overlapped coding regions.
The genetic compartments of the 44 promoters of our collection were next: thirty-nine were located on the chromosome, two on plasmid p42d and one in each of these plasmids: p42b, p42e and p42f.
Identification of transcriptional start-sites
The transcription start-sites are good reference points to identify the upstream conserved sequence characteristics of the promoters. Thus, the 5′ ends of the mRNAs synthesized from each of the selected 36 promoter regions selected, were successfully identified by 5′ RACE using the total RNA isolated from PY cultures in exponential growth conditions of the transconjugants. In general, a single RT–PCR product was obtained, but in one clone, two conspicuous amplification products were observed. In this case the bands were independently isolated, cloned and sequenced, showing that these products are transcripts associated to the same gene.
Thirty-eight of the transcription start-sites (86%) fell within the first 250 bp upstream from coding regions, and four transcription start-sites corresponded to the nucleotide A of the start codon ATG from their respective ORF (Table 2).
Table 2.
Distances between the experimentally determined transcriptional start-sites and the ATG for each gene
| Gene | Distance between +1 to ATG (bp) | Gene product | References |
|---|---|---|---|
| ctRNA | NA | Antisense RNA regulator | (30) |
| repA | 45 | Plasmid partitioning protein RepAd | (31) |
| thiC | 209 | Thiamine biosynthesis | (32) |
| recA | 37 | Recombinase A | (33) |
| lipA | 63 | Lipoic acid synthetase | (34) |
| nifR | 71 | Nitrogen regulation | (35) |
| nifR | 28 | Nitrogen regulation | (35) |
| sigA | 230 | RNA polymerase sigma factor protein (sigma-70) | This work |
| ypf00079 | 18 | Probable methyl-accepting chemotaxis protein | This work |
| glyQ | 27 | Glycyl-tRNA synthetase alpha subunit | This work |
| yhch00468 | 37 | Hypothetical conserved protein | This work |
| yhch00334 | 54 | Hypothetical conserved protein | This work |
| yhch01154 | 229 | Hypothetical conserved protein | This work |
| Eda | 35 | KHG-KDPG aldolase | This work |
| yhch00528 | 19 | Hypothetical conserved protein | This work |
| engB | 29 | GTP-binding protein | This work |
| 16S rRNA | NA | 16S ribosomal RNA | This work |
| yhch01141 | 67 | Hypothetical conserved protein | This work |
| yhch00226 | 23 | Hypothetical conserved protein | This work |
| yhch00739* | A | Hypothetical conserved protein | This work |
| hemC | 18 | Porphobilinogen deaminase | This work |
| clpP | 55 | ATP-dependent Clp protease proteolytic subunit 2 | This work |
| rpoH1 | 101 | RNA polymerase sigma factor protein (sigma-32) | This work |
| aroG | 164 | Phospho-2-dehydro-3- deoxyheptonate aldolase | This work |
| ypch00196 | 41 | Putative transcriptional regulator | This work |
| ureA | 34 | Urease (urea amidohydrolase) gamma subunit | This work |
| ypch00131 | 268 | Probable transcriptional regulator | This work |
| ndvB | 24 | Beta-1,2-glucan production associated transmembrane protein | This work |
| ype00047* | 29 | Putative tail-specific protease | This work |
| yhch00197 | 21 | Hypothetical conserved protein | This work |
| pfs | 272 | mta/sah nucleosidase (P46) bifunctional protein | This work |
| cycH | A | cytochrome c-type biogenesis protein | This work |
| ypch00552 | A | Probable cytochrome P450 hydroxylase | This work |
| yhch00096 | 7 | Hypothetical conserved protein | This work |
| ypch00132 | 155 | Putative glutamine amidotransferase | This work |
| pepN | 23 | Aminopeptidase N | This work |
| pepN | 14 | Aminopeptidase N | This work |
| yhch00326 | 65 | Hypothetical conserved protein | This work |
| rpsP | 92 | 30S ribosomal protein S16 | This work |
| yhch01115 | 123 | Hypothetical conserved | This work |
| yhch00280 | 375 | Hypothetical conserved | This work |
| ispB | A | Octaprenyl-diphosphate synthase | This work |
| yhch00076 | 381 | Hypothetical conserved protein | This work |
| exoR2 | 242 | Exopolysaccharide production negative regulator | This work |
*pseudogene; A, adenine from ATG; NA, non apply.
Determination of a promoter consensus sequence
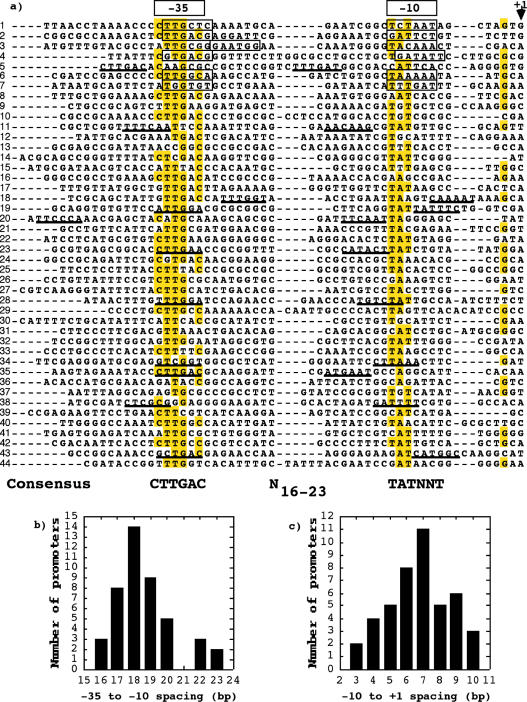
The promoter consensus sequence was derived by aligning 50 bp upstream from the transcription start-site of each promoter region. The identification of −10 and −35 boxes was corroborated with the WCONSENSUS program as described in Materials and Methods. As shown in Figure 1a, over-represented nucleotides clearly appeared around the −35 region and, in a lesser degree, in the −10 region. Then, the putative −10 consensus box [TATNNT] showed four identical positions to the −10 box of the E.coli consensus promoter [TATAAT] recognized by σ70 factor. The 3′ end of the R.etli −10 consensus boxes was positioned from 3 to 10 bp upstream from the transcriptional start-site (Figure 1c). Six promoter regions (14%) showed the dinucleotide [TG] in positions −15 and −14, which resembled the structure of the E.coli −10 extended promoter. The −35 box consensus sequence [CTTGAC], is identical in five positions to the equivalent box of the E.coli consensus promoter [TTGACA] recognized by σ70. The spacing between the −35 and −10 regions ranges from 16 to 23 bp (Figure 1b). The promoter sequences contain 6.4 conserved bases on average, of the 12 conforming −10 and −35.
Figure 1.
(a) Alignment of the promoter regions identified in R.etli. The position of the −10 and −35 elements are indicated at the top of the alignment. The −10 and −35 elements identified by other authors are indicated within boxes. The nucleotides that appear in ≥51% of the cases in any given position are marked in yellow. The transcription start-sites are marked with +1. Alternative promoters with suboptimal properties are underline. The consensus sequence derived from the alignment are indicated at the bottom of the alignment. The promoters identified were: 1, prepA; 2, pctRNA; 3, precA; 4, plipA; 5, pthiC; 6, pnifR promoter 1; 7, pnifR promoter 2; 8, psigA; 9, pypf00079; 10, pglyQ; 11, pyhch00468; 12, pyhch00334; 13, pyhch01154; 14, peda; 15, pyhch00528; 16, pengB; 17, p16S rRNA; 18, pyhch01141; 19, pyhch00226; 20, pyhch00739 pseudogene; 21, phemC; 22, pclpP; 23, prpoH1; 24, paroG; 25, pypch00196; 26, pureA; 27, pypch00131; 28, pndvB; 29, pype00047 pseudogene; 30, pyhch00197; 31, ppfs; 32, pcycH; 33, pypch00552; 34, pyhch00096; 35, pypch00132; 36, ppepN promoter 1; 37, ppepN promoter 2; 38, pyhch00326; 39, prpsP; 40, pyhch01115; 41, pyhch00280; 42, pispB; 43, pyhch00076; 44, pexoR2. (b) Distribution of spacer lengths for the 44 promoters of the (a) (c) Distribution of the distance between the 3′ of the −10 element (TATNNT) and the +1
Suboptimal promoters were identified in 11 of the 44 promoter regions as shown in Figure 1a, but they were not taken in account to generate the consensus promoter sequence for the following reasons: none of them contained both hexameric boxes recognized by the best statistic parameters of WCONSENSUS. Nevertheless, in six of these promoters the −35 box overlapped with one proposed by us. In one case the −10 box overlapped the region proposed here. Moreover, some of the suboptimal promoters contained the spacer region two long (up to 27 bp) or to short (11 or 12 bp). In two cases the transcription start-sites were located far of the −10 boxes (up to 20 bp).
The results presented here showed that the R.etli promoter consensus possesses some similarities with the E.coli promoter consensus recognized by the σ70 factor, but the variability in the nucleotide frequency in each position of the consensus was greater than that observed in E.coli promoters (7).
Thermodynamic properties of the R.etli promoter regions
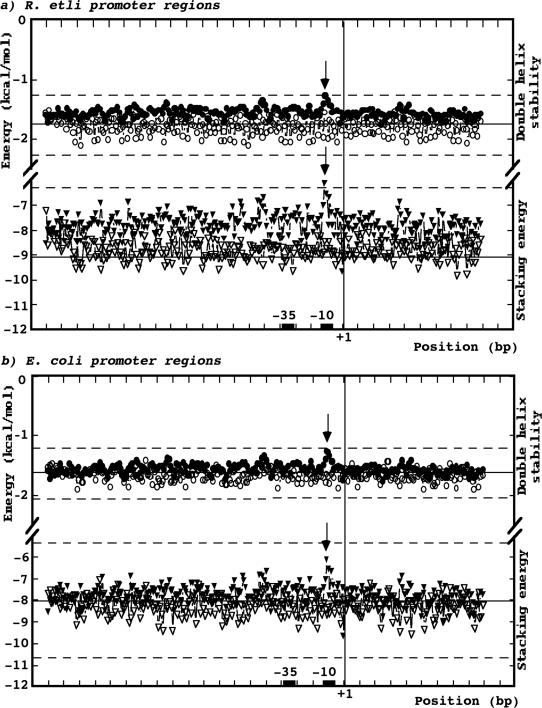
It was recently demonstrated in several bacteria genomes, that the intergenic regions containing promoters are generally less stable and less flexible than coding regions (36). To determine if these thermodynamic properties are also present in the promoter regions of R.etli, the stacking energy and the double helix stability were evaluated in our collection of 44 promoters. For this analysis the sequence of 200 bp upstream and 100 bp downstream from transcription start-site of all members of the promoter collection were taken in account. As an internal control, a set similar in size and length was constructed with the immediately downstream coding region of each member of the promoter collection. The same properties were predicted and contrasted with two E.coli equivalent sets: one containing 44 randomly chosen promoter sequences experimentally characterized and putatively recognized by σ70, and the other of 44 coding region sequences that corresponded to the genes controlled by the selected promoters. As shown in Figure 2a, the distribution profiles of double helix stability for coding and no coding regions of R.etli were similar, however, a significative local minimum around the −10 element was detected in our collection of promoter regions. The average stability of the E.coli coding regions was higher than the average stability of their promoter regions, and like R.etli, a prominent feature of the promoter regions was a minimum stability around the −10 box (Figure 2b).
Figure 2.
Structural properties of R.etli end E.coli promoters (a) Double helix stability and stacking energy of R.etli promoter regions (b) Double helix stability and stacking energy of E.coli promoter regions. Average representation for extended promoter regions from −200 to +100 bp with respect to +1. The x-axis represents the position in the promoter region (in bp), and the y-axis the energy (in kcal/mol). A vertical line marked with +1 indicates the position of the transcription start-sites. The position of −10 and −35 elements are marked with black boxes. The white symbols correspond to average values of a control set formed by 301 bp of coding regions. The black symbols correspond to average values of extended promoter regions. The circles represent the values of double helix stability, and the triangles show the values of stacking energy. Local minimum values for double helix stability and stacking energy are marked with arrows. The calculations for each parameter were carried out with a window size of 10 nt. Continuous and discontinuous lines indicate the average value of the collection and the average deviation, respectively.
The stacking energy profiles of R.etli and E.coli promoter regions were variable, but with a tendency to low negative values (low stability), nevertheless local minimum values were located around the −10 box. In contrast, the stacking energy profiles of R.etli and E.coli coding regions were similar: both showed more negative values that corresponded to great stability (Figure 2a and b). These results suggest that despite the variability in the nucleotide composition of R.etli promoters, these regions possess thermodynamic and structural properties similar to the E.coli promoter regions.
Promoter sequence mutagenesis
To demonstrate that the promoter consensus proposed here is correct, the DNA sequence of three R.etli mutant promoters (from repA, the antisense RNA gene present in the p42d repABC operon and recA) with the reduced transcriptional activity previously characterized was located in our alignment. As shown in Figure 3, these mutations fall in the regions centred in −10 and −35 of the consensus, suggesting that these elements were correctly identified.
Figure 3.
Genetic analysis of R.etli promoter regions All R.etli mutant promoters with reduced transcriptional activity previously characterized, plus mutant derivatives of the phemC promoter constructed in this work were aligned with the R.etli promoter consensus. The changes that produced a reduced transcriptional activity are marked with lower-case letters.
Additionally the promoter consensus was verified in the following way: the constitutive promoter (phemC) of hemC, a gene encoding a porphobilinogen deaminase protein, was selected for a mutagenic analysis. This promoter possessed five conserved positions from consensus and showed moderate activity. Three mutant derivatives were constructed that deviated the promoter sequence from the consensus. First, the putative −35 element was altered, changing sequence [ATTGCG] to [AAGACG] to generate the construct pGUSphemC-35. Second, the putative −10 element was changed from [TACGAG] to [GCGGAG] to produce construct pGUSphemC-10. Third, the putative wild-type −35 and −10 elements were changed at the same time to the sequences indicated before, to generate plasmid pGUSphemC-1035. These constructs were introduced into R.etli CFN42, and the β-glucuronidase activities of the transconjugants were determined. As shown in Figure 3, all mutant constructs showed a low level of β-glucuronidase activity, indicating that the −35 and −10 elements of the R.etli promoter consensus were correctly identified.
Activity of the R.etli promoters in E.coli
As described in the previous section, the R.etli σ70 promoter consensus sequence is more variable than that described for E.coli. This observation suggests that the E.coli σ70 is unable to recognize the R.etli promoters or recognizes them less efficiently. To prove this possibility, the 36 transcriptional fusions of the collection were introduced into the BW21038 (gusA) strain, and their β-glucuronidase activities were evaluated in plates supplemented with X-Gluc. In this experiment only three promoters were functional, but showed low β-glucuronidase activity. These results suggest that the 33 transcriptional fusions are poorly or unrecognized by the E.coli sigma factors. It is possible than the E.coli σ70 is incapable of binding the R.etli promoters because it is more stringent in its recognition pattern, and imposes a strict functional barrier. If this hypothesis is true, then the R.etli σ70 (SigA) activates the transcriptional fusions described above in E.coli. To demonstrate this, all strains containing the gusA fusions were transformed with a plasmid expressing the R.etli σ70 gene under the control of the lac promoter and the β-glucuronidase activity of the transformants were evaluated in LB plates containing X-Gluc. Representative results are shown in Figure 4. The expression of sigA allowed the gusA fusions to express their reporter gene in the 36 R.etli promoter regions studied here, however, the strains containing sigA constructs were unstable, and a mixure of blue and white colonies was observed in the plates (Figure 4). The results suggest that R.etli σ70 is responsible for the transcriptional activity of these promoters, and that the structural variability of R.etli promoters cannot be recognized by E.coli σ70.
Figure 4.
β-glucuronidase activities of the R.etli gusA fusions in E.coli Qualitative evaluation of the β-glucuronidase activities of E.coli BW21038 transformants containing different R.etli promoter regions fused to the gusA gene of the promoter-trap vector pBBMCS53, with or without the expression of sigA present in the plasmid pRKS70. Each plate included positive (pHER; + symbol) and negative (pBBMCS53; − symbol) controls. Each pair of strains, marked with the same number, contain the same transcriptional fusion. Strains marked with an asterisk harbor the pRKS70, which expresses the sigA gene of R.etli. The strains contain the following promoter regions: 1, pglyQ; 2, pyhch1154; 3, prpoH1; 4, pyhch0528; 5, pypch00326; 6, pyhch00096; 7, pyhch00468; 8, pyhch00334 and 9, peda.
The E.coli σ70 is incapable of binding in vitro R.etli promoters
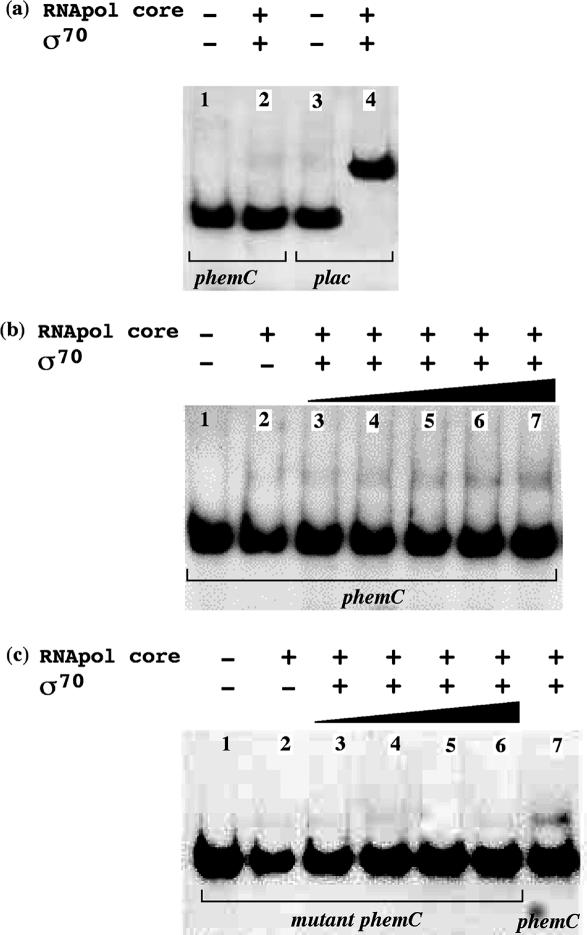
To verify if the R.etli promoters are recognized by the E.coli σ70 factor, electrophoretic mobility shift assay (EMSA) experiments were done using E.coli RNApol containing E.coli σ70 and a PCR product containing the promoter of hemC gene (phemC), plus the PCR products of four promoter regions selected at random: ureA, cycH, yhch00197 and yhch00468. As a negative control, a PCR product of a mutant derivative of promoter phemC containing the changes in the −10 and −35 elements described previously was used, and as a positive control, a PCR product containing the lac promoter was used (lac promoter is functional in R.etli). Representative results are shown in Figure 5a. In all experiments the E.coli RNApol holoenzyme was unable to bind the R.etli promoters tested, indicating that E.coli σ70 cannot recognize R.etli promoters.
Figure 5.
Gel mobility shift assays of the E.coli lac promoter, the R.etli phemC promoter or its mutant derivative by E.coli RNApol holoenzyme or by an E.coli RNApol core -SigA chimera. (a) The E.coli RNApol holoenzyme is capable of binding the E.coli lac promoter but not the R.etli phemC promoter. The DNA fragments used in the binding reactions, as targets were 5 fmol of 32P-labeled PCR products containing the phemC promoter (164 bp) or containing the lac promoter (167 bp). Each product was incubated with 12 pmol of RNApol core enzyme and 12 pmol of RpoD. The volume of the reaction was 15 µl. Lane 1 shows the PCR containing the lac promoter, and lane 2 shows the same product but with the addition of the protein mix. Lane 3 shows the PCR containing the phemC promoter, and lane 4 shows the same product but with the addition of the protein mix. (b) Gel mobility shift assays demonstrating the ability of an E.coli RNApol core -SigA chimera to bind the R.etli phemC promoter. Lane 1 shows 5 fmol of 32P-labeled PCR product containing the phemC promoter (164 bp), used as target. Lane 2 shows the same PCR product mixed only with 12 pmol of E.coli RNApol core enzyme. Lanes 3–7, show the target DNA with 12 pmol of E.coli RNApol core enzyme and SigA at the following concentrations: lane 3, 2 pmol; lane 4, 4 pmol; lane 5, 8 pmol; lane 6, 10 pmol, and lane 7, 12 pmol. The volume of each reaction was 15 µl. (c) Gel mobility shift assays showing the incapacity of an E.coli RNApol core -SigA chimera to bind to the R.etli phemC 1035 promoter, a derivative carrying mutations in the −10 and −35 elements. Lane 1 shows 5 fmol of 32P-labeled PCR product containing the phemC 1035 promoter (164 bp), used as target, and lane 2, the same target DNA mixed with 12 pmol of E.coli RNApol core enzyme. Lanes 3–6, show the target DNA mixed with 12 pmol of E.coli RNApol core enzyme and SigA at the following concentrations: lane 3, 4 pmol; lane 4, 8 pmol; lane 5, 10 pmol, and lane 6, 12 pmol. Lane 7 contains a 32P-labeled PCR product of the wild-type phemC promoter (164 bp) mixed with 12 pmol of RNApol core enzyme and 12 pmol of SigA. The volume of each reaction was 15 µl.
SigA binds in vitro R.etli promoters
To determine if SigA is capable of recognizing the R.etli promoters, binding shift experiments were done using a chimeric RNApol holoenzyme composed of E.coli RNApol core and R.etli σ70 factor assembled in vitro. As a target DNA, the PCR products containing the promoters mentioned above (phemC and the promoters of ureA, cycH, yhch00197 and yhch00468 genes) were used in these experiments. Representative results are shown in Figure 5. The chimeric RNApol holoenzyme bound the five PCR products, but at low efficiency (Figure 5b). In contrast, no DNA binding shifts were observed using a PCR control with the mutant promoter phemC (Figure 5c). These results showed that the chimeric RNApol holoenzyme specifically recognizes the R.etli promoters.
Comparative analysis of the σ70 consensus sequences from different bacteria
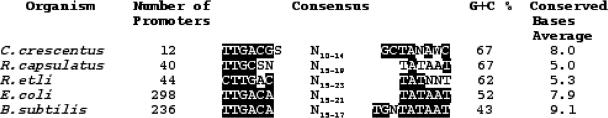
The R.etli σ70 promoter consensus was compared with those of the: (i) α-proteobacteria C.crescentus (G + C% of 67) (37) and Rhodobacter capsulatus (G + C% of 67) (38); (ii) the γ-proteobacteria E.coli (G + C% of 52) (6); and (iii) B.subtilis (G + C% of 43) (39). In the promoter consensus sequences of C.crescentus, R.capsulatus and R.etli, the −35 boxes are more conserved than the −10 boxes (Figure 6). Furthermore, the inspection of the all sequences showed that each bacterial species has a consensus sequence, with a different degree of variability. The R.etli and R.capsulatus promoters contain 6.4 and 5.0 conserved bases on average, respectively, less conserved in comparison with the promoter consensus of C.crescentus, that contains 8.0, E.coli 7.9 and B.subtilis 9.1 conserved nucleotides (7,39,40). Finally, the number of conserved nucleotides, especially in the −10 box, tends to diminish when genomic G + C content is above 60%. It is worth noting that the E.coli and B.subtilis −10 and −35 boxes are A + T rich and conserve 10 of 12 positions.
Figure 6.
Comparison of R.etli consensus promoter with other σ70 bacterial consensus promoters Black boxes correspond to nucleotides that appear in >51% of promoter consensus sequences analyzed. S (C or G), W (A or T), N (any nucleotide).
DISCUSSION
R.etli is a bacterium that inhabits diverse environments and has genome rich in genes related to transcriptional regulation, suggesting that it contains a complex genetic regulation system. To characterize the molecular basis of the transcription, and particulary, what kind of sequences act as promoters in R.etli, we identified and characterized a promoter consensus sequence recognized by the R.etli σ70 factor (SigA) in this paper.
The R.etli σ70 promoter consensus sequence [CTTGAC] N16–23 [TATNNT] was deduced from 44 promoter regions. Using site-directed mutagenesis of one representative R.etli promoter isolated in this work, and a collection of mutants of three promoter regions described previously, we found that all changes fell within the −10 or −35 boxes identified in this work, suggesting that the consensus sequence is correct. Moreover, two different lines of evidence indicate that this promoter consensus sequence is recognized by the R.etli σ70. The first one is that gusA fusions containing R.etli promoters are not functional in E.coli, but all regain their activity when complemented with SigA. Second, SigA is able to bind to the R.etli promoters but not to their mutant derivatives that partially eliminate the −35 or −10 boxes.
The location of the 44 promoter regions in the R.etli genome showed that 33 are located within intergenic regions, and that 8 (18%), fall within coding regions. This situation has also been observed in other bacteria, i.e. in E.coli 18% of the promoters are within genes (41), and around of 20% in Prochlorococcus marinus (42).
Of the promoters 86% analyzed here were located less than 250 bp from the translation start codons of their genes. This property has been widely studied in E.coli, where it is known that 90% of the promoters are located less than 250 bp upstream from the gene that they regulate (41).
Although the R.etli, B.subtilis and E.coli promoter consensus display a common structure, the R.etli consensus sequence contains only 6.4 conserved bases on average, revealing a more variable composition than in the E.coli and B.subtilis consensus, where 7.9 and 9.1 nt are conserved on average, respectively (7,39,40). Additionally, the R.etli promoter consensus shows that the sequence length between the −10 and −35 boxes varies from 16 to 23 nt, nevertheless, 25 of the promoters (56.81%) present a distance of 17 ± 1 nt. This variation range is similar to that found in E.coli (15 to 21 nt) and in B.subtilis (14 to 20 nt). The percentage of promoters with 17 ± 1 nt in length between the hexameric boxes, is quite low compared to E.coli (92%) and B.subtilis (84.5%) (6,39). The distance between +1 nt and −10 box 3′ end fluctuates from 3 to 10 bases, as found in E.coli and B.subtilis (6,39). Six promoters (14%) were found with a putative −10 extended sequence, and their frequency is lower than those reported for this kind of promoter in E.coli (20%) and B.subtilis (45%) (39,43,44).
A computer analysis of DNA double helix stability and stacking energy showed that the promoter regions presented a minimum value for both parameters around −10 box, as found in E.coli promoters. Having these results in mind, it is reasonable to conclude that in spite of their sequence heterogeneity, the promoter regions preserve thermodynamic properties which have been selected to favor DNA denaturation and help to start the transcription. Similar conclusions have been drawn for other bacterial promoter regions (36,45).
The R.etli promoters tested are not recognized by E.coli σ70, however, the E.coli lac promoter is recognized by R.etli σ70, suggesting that R.etli σ70 is more promiscuous than E.coli σ70.
The deduced primary structure of SigA is 47% similar to the E coli σ70, especially in regions 2.4 and 4.2, which are responsible for promoter recognition. The most notable difference between SigA and RpoD is the larger size of region 1 of SigA, which contains 72 extra amino acids. This region could be related to the lax promoter recognition of SigA. Thus, E. coli σ70 being the strictest factor for promoter recognition, it allows less structural variation, which is reflected in a robust consensus sequence. The opposite case is observed in α-proteobacteria, where the promiscuous σ70 allows a larger variation in its promoter structure.
A promiscuous σ70 factor could be an adaptive advantage for bacteria like the α-proteobacteria that inhabit environments with high biological and enviromental diversity, because they can take the benefits of the genetic information acquired by horizontal transfer in a fast and efficient way, avoiding a barrier that could limit the acquisition of new functions required for the adaptation to new environmental conditions.
Our comparison of the □□□□□□□□ consensus of several bacteria suggests that differences in the genomic G + C content are related to promoter structure: the bacteria with high G + C content tend to be more variable, and to have −10 boxes different in sequence to those from bacteria with low G + C content, and vice versa. Previous evidence in other organisms supports this conclusion (46).
Acknowledgments
The authors wish to thank Marco Zúñiga, Rosa Elena Gómez Barreto, Ismael Hernández, Patricia Bustos, Rosa Isela Santamaría and José Espíritu for their skillful technical support. The authors also thank Paul Gaytán and Eugenio López for the synthesis of oligonucleotides. This work was funded by CONACyT grants N-028, N-U46333-Q for emergent areas, and the Universidad Nacional Autónoma de México. Funding to pay the Open Access publication charges for this article was provided by Universidad Nacional Autónoma de México.
Conflict of interest statement. None declared.
REFERENCES
- 1.Lonetto M., Gribskov M., Gross C.A. The σ70 family: sequence conservation and evolutionary relationships. J. Bacteriol. 1992;174:3843–3849. doi: 10.1128/jb.174.12.3843-3849.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Wosten M.M.S.M. Eubacterial sigma-factors. FEMS Microbiol. Rev. 1998;22:127–150. doi: 10.1111/j.1574-6976.1998.tb00364.x. [DOI] [PubMed] [Google Scholar]
- 3.Maeda H., Fujita N., Ishihama A. Competition among seven Escherichia coli sigma subunits: relative binding affinities to the core RNA polymerase. Nucleic Acids Res. 2000;28:3497–3503. doi: 10.1093/nar/28.18.3497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Mittenhuber G. An inventory of genes encoding RNA polymerase sigma factors in 31 completely sequenced eubacterial genomes. J. Mol. Microbiol. Biotechnol. 2002;4:77–91. [PubMed] [Google Scholar]
- 5.Paget M.S., Helmann J.D. The sigma70 family of sigma factors. Genome Biol. 2003;4:203.1–203.6. doi: 10.1186/gb-2003-4-1-203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Harley C.B., Reynolds R.P. Analysis of E.coli promoter sequences. Nucleic Acids Res. 1987;15:2343–2361. doi: 10.1093/nar/15.5.2343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lisser S., Margalit H. Compilation of E.coli mRNA promoter sequences. Nucleic Acids Res. 1993;21:1507–1516. doi: 10.1093/nar/21.7.1507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Murakami K.S., Masuda S., Darst S.A. Structural basis of transcription initiation: RNA polymerase holoenzyme at 4 A resolution. Science. 2002;296:1280–1284. doi: 10.1126/science.1069594. [DOI] [PubMed] [Google Scholar]
- 9.Kumar A., Malloch R.A., Fujita N., Smillie D.A., Ishihama A., Hayward R.S. The minus 35-recognition region of Escherichia coli sigma 70 is inessential for initiation of transcription at an ‘Extended minus 10’ promoter. J. Mol. Biol. 1993;232:406–418. doi: 10.1006/jmbi.1993.1400. [DOI] [PubMed] [Google Scholar]
- 10.Barne K.A., Bown J., Busby S.J., Minchin S.D. Region 2.5 of the Escherichia coli RNA polymerase sigma70 subunit is responsible for the recognition of the ‘extended-10’ motif at promoters. EMBO J. 1997;16:4034–4040. doi: 10.1093/emboj/16.13.4034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Campbell E.A., Muzzin O., Chlenov M., Sun J.L., Olson C.A., Weinman O., Trester-Zedlitz M.L., Darst S.A. Structure of the bacterial RNA polymerase promoter especificity sigma subunit. Mol. Cell. 2002;9:527–539. doi: 10.1016/s1097-2765(02)00470-7. [DOI] [PubMed] [Google Scholar]
- 12.González, et al. The partitioned Rhizobium etli genome: Genetic and metabolic redundancy in seven interacting replicons. Proc. Natl Acad. Sci. USA. 2005;103:3834–3839. doi: 10.1073/pnas.0508502103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Luka S., Patriarca E.J., Riccio A., Iaccarino M., Defez R. Cloning of the rpoD analog from Rhizobium etli: sigA of R.etli is growth phase regulated. J. Bacteriol. 1996;178:7138–7143. doi: 10.1128/jb.178.24.7138-7143.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Peck M.C., Gaal T., Fisher R.F., Gourse R.L., Long S.R. The RNA polymerase alpha subunit from Shinorhizobium meliloti can assemble with RNA polymerase subunits from Escherichia coli and function in basal and activated transcription both in vivo and in vitro. J. Bacteriol. 2002;184:3808–3814. doi: 10.1128/JB.184.14.3808-3814.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Cullen P.J., Kaufman C.K., Bowman W.C., Kranz R.G. Characterization of the Rhodobacter capsulatus housekeeping RNA polymerase. In vitro transcripcion of photosynthesis and other genes. J. Biol. Chem. 1997;272:27266–27273. doi: 10.1074/jbc.272.43.27266. [DOI] [PubMed] [Google Scholar]
- 16.Wu J., Ohta N., Benson A.K., Ninfa A.J., Newton A. Purification, characterization, and reconstitution of DNA-dependent RNA polymerases from Caulobacter crescentus. J. Biol. Chem. 1997;272:21558–21564. doi: 10.1074/jbc.272.34.21558. [DOI] [PubMed] [Google Scholar]
- 17.Noel K.D., Sánchez A., Fernández L., Leemans J., Cevallos M.A. Rhizobium phaseoli symbiotic mutants with transposon Tn5 insertions. J. Bacteriol. 1984;158:148–155. doi: 10.1128/jb.158.1.148-155.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Bravo A., Mora J. Ammonium assimilation in Rhizobium phaseoli by the glutamine synthetase-glutamate synthase pathway. J. Bacteriol. 1988;170:980–984. doi: 10.1128/jb.170.2.980-984.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Sambrook J., Fritsch E.F., Maniatis T. Molecular Cloning. A Laboratory Manual. 2nd edn. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1989. [Google Scholar]
- 20.Corvera A., Promé D., Promé J.C., Martínez-Romero E., Romero D. The nolL gene from Rhizobium etli determines nodulation efficiency by mediating the acetylation of the fucosyl residue in the nodulation factor. Mol. Plant–Microbe Interact. 1999;12:236–246. doi: 10.1094/MPMI.1999.12.3.236. [DOI] [PubMed] [Google Scholar]
- 21.Wilson K.J., Huges S.G., Jefferson R.A. The Escherichia coli gus operon, induction and expression of the gus operon in E.coli and the occurrence and use of GUS in other bacteria. In: Gallagher S.R., editor. Gus Protocols, Using the Gus Gene as a Reporter of Gene Expression. Vol. 1. San Diego, CA: Academic Press; 1992. pp. 7–23. [Google Scholar]
- 22.Studier F.W., Rosenberg A.H., Dunn J.J., Dubendorff J.W. Use of T7 RNA polymerase to direct expression of cloned genes. Meth. Enzymol. 1990;185:60–89. doi: 10.1016/0076-6879(90)85008-c. [DOI] [PubMed] [Google Scholar]
- 23.Tang H., Severinov K., Goldfarb A., Ebright H. Rapid RNA polymerase genetics: one-day, no column preparation of reconstituted recombinant Escherichia coli RNA polymerase. Proc. Natl Acad. Sci. USA. 1995;92:4902–4906. doi: 10.1073/pnas.92.11.4902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Higgins D., Thompson J., Gibson,Thompson J.D., Higgins D.G., Gibson T.J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22:4673–4680. doi: 10.1093/nar/22.22.4673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Corpet F. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res. 1988;16:10881–10890. doi: 10.1093/nar/16.22.10881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Hertz G.Z., Stormo G.D. Identifying DNA and protein patterns with statistically significant alignments of multiple sequences. Bioinformatics. 1999;15:563–577. doi: 10.1093/bioinformatics/15.7.563. [DOI] [PubMed] [Google Scholar]
- 27.Vlahovicek K., Kaján L., Pongor S. DNA analysis servers: plot.it, bend.it, model.it and IS. Nucleic Acids Res. 2003;31:3686–3687. doi: 10.1093/nar/gkg559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Sugimoto N., Nakano S., Yoneyama M., Honda K. Improved thermodynamic parameters and helix initiation factors to predict stability of DNA duplexes. Nucleic Acids Res. 1996;24:4501–4505. doi: 10.1093/nar/24.22.4501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Ornstein R.L., Rein R., Breen D.L., MacElroy R.D. An optimized potential function for the calculation of nucleic acid interaction energies I. Base stacking. Biopolymers. 1978;17:2341–2360. doi: 10.1002/bip.1978.360171005. [DOI] [PubMed] [Google Scholar]
- 30.Venkova-Canova T., Soberón N.E., Ramírez-Romero M.A., Cevallos Miguel A. Two discrete elements are required for the replication of repABC plasmid: an antisense RNA and a stem–loop structure. Mol. Microbiol. 2004;54:1431–1444. doi: 10.1111/j.1365-2958.2004.04366.x. [DOI] [PubMed] [Google Scholar]
- 31.Ramírez-Romero M.A., Téllez-Sosa J., Barrios H., Pérez-Oseguera A., Rosas V., Cevallos M.A. RepA negatively autoregulates the transcription of the repABC operon of the Rhizobium etli symbiotic plasmid basic replicon. Mol. Microbiol. 2001;42:195–204. doi: 10.1046/j.1365-2958.2001.02621.x. [DOI] [PubMed] [Google Scholar]
- 32.Miranda-Rios J., Morera C., Taboada H., Dávalos A., Encarnación S., Mora J., Soberón M. Expressión of thiamin biosynthetic genes (thiCOGE) and production of symbiotic terminal oxidase cbb3 in Rhizobium etli. J. Bacteriol. 1997;179:6887–6893. doi: 10.1128/jb.179.22.6887-6893.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Tapias A., Fernández de Henestrosa A.R., Barbé J. Characterization of the promoter of the Rhizobium etli recA gene. J. Bacteriol. 1997;179:1573–1579. doi: 10.1128/jb.179.5.1573-1579.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Taté R., Riccio A., Iaccarino M., Patriarca E.J. Cloning and transcriptional análisis of the lipA (lipoic acid synthetase) gene from Rhizobium etli. FEMS Microbiol. Lett. 1997;149:165–172. doi: 10.1111/j.1574-6968.1997.tb10324.x. [DOI] [PubMed] [Google Scholar]
- 35.Patriarca E.J., Riccio A., Taté R., Colonna-Romano S., Iaccarino M., Defez R. The ntrBC genes of Rhizobium leguminosarum are part of a complex operon subject to negative regulation. Mol Microbiol. 1993;9:569–577. doi: 10.1111/j.1365-2958.1993.tb01717.x. [DOI] [PubMed] [Google Scholar]
- 36.Pedersen A.G., Jensen L.J., Brunak S., Staerfeldt H.H., Ussery D.W. A DNA structural atlas for Escherichia coli. J. Mol. Biol. 2000;299:907–930. doi: 10.1006/jmbi.2000.3787. [DOI] [PubMed] [Google Scholar]
- 37.Malakooti J., Wang S.P., Ely B. A consensus promoter sequence for Caulobacter crescentus genes involved in biosynthetic and housekeeping functions. J. Bacteriol. 1995;177:4372–4376. doi: 10.1128/jb.177.15.4372-4376.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Richard C.L., Tandon A., Kranz R.G. Rhodobacter capsulatus nifA1 promoter: High-GC−10 regions in high-GC bacteria and basis for their transcription. J. Bacteriol. 2004;186:740–749. doi: 10.1128/JB.186.3.740-749.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Helmann J.D. Compilation and analysis of Bacillus subtilis σA-dependent promoter sequences: evidence for extended contact between RNA polymerase and upstream promoter DNA. Nucleic Acids Res. 1995;23:2351–2360. doi: 10.1093/nar/23.13.2351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Ozoline O.N., Deev A.A., Arkhipova M.V. Non-canonical sequence elements in the promoter structure. Cluster analysis of promoters recognized by Escherichia coli RNA polymerase. Nucleic Acids Res. 1997;25:4703–4709. doi: 10.1093/nar/25.23.4703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Huerta M.A., Collado-Vides J. Sigma70 promoters in Escherichia coli: specific transcription in dense regions of overlapping promoter-like signals. J. Mol. Biol. 2003;333:261–278. doi: 10.1016/j.jmb.2003.07.017. [DOI] [PubMed] [Google Scholar]
- 42.Vogel J., Axmann I.M., Herzel H., Hess W.R. Experimental and computer analysis of transcriptional start sites in the cyanobacterium Prochlorococcus MED4. Nucleic Acids Res. 2003;31:2890–2899. doi: 10.1093/nar/gkg398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Burr T., Mitchell J., Kolb A., Minchin S., Busby S. DNA sequence elements located immediately upstream of the −10 hexamer in Escherichia coli promoters: a systematic study. Nucleic Acids Res. 2000;28:1864–1870. doi: 10.1093/nar/28.9.1864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Mitchell J.E., Zheng D., Busby S.J., Minchin S.D. Identification and analysis of ‘extended –10’ promoters in Escherichia coli. Nucleic Acids Res. 2003;31:4689–4695. doi: 10.1093/nar/gkg694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Lisser S., Margalit H. Determination of common structural features in Escherichia coli promoters by computer analysis. Eur. J. Biochem. 1994;223:823–830. doi: 10.1111/j.1432-1033.1994.tb19058.x. [DOI] [PubMed] [Google Scholar]
- 46.Morrison D.A., Jaurin B. Sttreptococcus pneumoniae possesses canonical Escherichia coli (sigma 70) promoters. Mol. Microbiol. 1990;4:1143–1152. doi: 10.1111/j.1365-2958.1990.tb00689.x. [DOI] [PubMed] [Google Scholar]
- 47.Hanahan D. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. 1983;166:557–580. doi: 10.1016/s0022-2836(83)80284-8. [DOI] [PubMed] [Google Scholar]
- 48.Metcalf W.W., Wanner B.L. Construction of new β-glucuronidase cassettes for making transcriptional fusions and their use with new methods for allele replacement. Gene. 1993;129:17–25. doi: 10.1016/0378-1119(93)90691-u. [DOI] [PubMed] [Google Scholar]
- 49.Kovach M.E., Elzer P.H., Hill D.S., Robertson G.T., Farris M.A., Roop R.M., Peterson K.M. Four new derivatives of the broad-host range cloning vector pBBR1MCS, carrying different antibiotic resistance cassettes. Gene. 1995;166:175–176. doi: 10.1016/0378-1119(95)00584-1. [DOI] [PubMed] [Google Scholar]
- 50.Keen N.T., Tamaki S., Kobayashi D., Trollinger D. Improved broad-host range plasmids for DNA cloning in gram-negative bacteria. Gene. 1988;70:191–197. doi: 10.1016/0378-1119(88)90117-5. [DOI] [PubMed] [Google Scholar]