Abstract
One function of lineage-restricted transcription factors may be to control the formation of tissue-specific chromatin domains. In erythroid cells, the β-globin gene cluster undergoes developmentally regulated hyperacetylation of histones at the active globin genes and the locus control region (LCR). However, it is unknown which transcription factor(s) governs the establishment of this erythroid-specific chromatin domain. We measured histone acetylation at the β-globin locus in the erythroid cell line G1E, which is deficient for the essential hematopoietic transcription factor GATA-1. Restoration of GATA-1 activity in G1E cells led to a substantial increase in acetylation of histones H3 and H4 at the β-globin promoter and the LCR. Time course experiments showed that histone acetylation occurred rapidly after GATA-1 activation and coincided with globin gene expression, indicating that the effects of GATA-1 are direct. Moreover, increases in histone acetylation correlated with occupancy of GATA-1 and the acetyltransferase CBP at the locus in vivo. Together, these results suggest that GATA-1 and its cofactor CBP are essential for the formation of an erythroid-specific acetylation pattern that is permissive for high levels of gene expression.
Formation of a permissive chromatin environment is a prerequisite for active transcription of eukaryotic genes. Genes that are expressed in a tissue-specific manner are thought to be embedded in tissue-specific chromatin regions that are characterized by elevated levels of histone acetylation. For example, high levels of histone acetylation are observed in mammalian and avian erythroid cells at the α- and β-globin genes (1, 8, 12, 22, 26, 28; for a review, see reference 5). Histone acetylation at the globin gene clusters is regulated throughout ontogeny, with the highest levels of acetylation found at the actively transcribed globin genes at a given stage of development. Additional peaks of histone acetylation are observed at the DNase I-hypersensitive sites of the β-globin locus control region (LCR) (8, 22, 28). The LCR, which in the human β-globin locus spans more than 17 kb and resides ∼6 kb upstream of the closest embryonic β-like globin gene, is required for high-level expression of all β-like globin genes (for reviews, see references 7, 14, 21, and 24). In transgenic mice, the LCR confers high-level expression of linked transgenes in a position-independent, copy number-dependent manner, suggesting a role in regulating chromatin structure. However, deletion of the murine and human LCR indicated that, while required for globin gene expression, it is dispensable for the formation of an open chromatin domain and for generalized histone hyperacetylation throughout the globin gene locus (2, 27-29). These results suggest that additional, perhaps redundant, sequences outside the LCR might contribute to the regulation of chromatin structure at the β-globin locus.
Erythroid-specific acetylation at the β-globin gene cluster is believed to be mediated at least in part by erythroid transcription factors that bind to the LCR and the globin gene promoters. Candidate factors include GATA-1, NF-E2, and EKLF. The observation that all of these factors associate with the histone acetyltransferases CBP/p300 (4, 6, 9, 17, 35) led to a model in which these proteins synergize in the recruitment of CBP/p300, thereby establishing a characteristic pattern of histone acetylation at the globin gene locus (for a review, see reference 3). In agreement with this model, interference with CBP/p300 function through forced expression of the adenoviral protein E1A inhibited erythroid cell differentiation and globin gene expression (4). The apparent simplicity of this model was challenged by a recent report showing that, in murine erythroid cells, NF-E2 is dispensable for histone acetylation at the LCR, which contains conserved and functional NF-E2 binding sites (18). In contrast, NF-E2 is required for high levels of histone acetylation at the β-globin gene promoter (18). These results raised the question as to which erythroid transcription factors mediate histone acetylation at the LCR and raised the possibility that histone acetylation is controlled by distinct mechanisms at the globin promoters and the LCR.
GATA-1 is a zinc finger transcription factor essential for normal development of erythroid cells, megakaryocytes, and eosinophils (references 15 and 34 and references therein). Functionally important GATA-1 binding sites are present at the globin gene promoters and the DNase I-hypersensitive sites (HS) 1 through 4 at the LCR, and GATA-1 binds to these regions in erythroid cells in vivo (16, 19). In this study we examined the role of GATA-1 in modulating histone acetylation at the globin gene locus. Using chromatin immunoprecipitation (ChIP) experiments, we measured histone acetylation at the LCR and the adult β-major globin gene promoter, comparing erythroid precursor cells that lack GATA-1 with those in which GATA-1 activity had been restored. We found that GATA-1 stimulates acetylation of both histones H3 and H4 at the LCR and the β-major globin gene promoter. Time course experiments further demonstrated a correlation between GATA-1 occupancy at GATA elements, recruitment of CBP, histone acetylation, and globin gene expression. Together, these results suggest that one essential function of GATA-1 is to establish an erythroid-specific pattern of histone hyperacetylation at the globin gene cluster and very likely other erythroid-expressed genes.
MATERIALS AND METHODS
Cell culture.
G1E cells were cultured as described previously (33). Cells were exposed to 1 μm estradiol (Sigma) where indicated.
ChIP assay.
ChIP assays were performed as described elsewhere (8) with the following modifications: cells were fixed in 0.4% formaldehyde in phosphate-buffered saline (PBS) at room temperature, and chromatin was sonicated to sizes between 500 and 1,000 bp. Two additional high-salt washes were performed following immunoprecipitations, using 20 mM Tris (pH 8.1), 2 mM EDTA, 500 mM NaCl, 1% Triton X-100, 0.01% sodium dodecyl sulfate. Samples were eluted twice in 100 μl (instead of 150 μl) of elution buffer, and the final volume was brought to 400 μl in Tris-EDTA.
PCRs of input DNA and immunoprecipitated DNA were performed in parallel in the presence of [α-32P]dCTP. The products were separated by acrylamide gel electrophoresis and quantified using a PhosphorImager and the ImageQuant software (Molecular Dynamics). The band intensity of each PCR was plotted as a percentage fraction of the band intensity of the input PCR (unprecipitated DNA). Comparing the input with the precipitated chromatin normalizes for variations in the input signal that may arise from variations in chromatin preparations. All PCRs were calibrated to reflect a linear relationship between sample content and PCR product. PCR primers spanning the β-major globin gene promoter, HS1, -2, -3, and -4, the embryonic globin genes ɛy and βH1, and the necdin gene (Table 1) were exactly as described previously (8). All ChIP experiments were controlled by analyzing in parallel samples containing no chromatin, no antibody, nonimmune rabbit IgG as control for anti-acetyl H3, and rabbit nonimmune serum as control for anti-acetyl H4. While all controls were performed in each experiment, they are shown only in Fig. 1 due to space limitations.
TABLE 1.
Primer pairs used in this study
| Site | Sequence |
|---|---|
| β-Major promoter | GACAAACATTATTCAGAGGGAGTACCC, AGGTGCACCATGATGTCTGTTTCTGG |
| HS1 | GCCTTACATCCCTGGACAGATAAAC, GTGACATTGTGAGAAACTGAGACCC |
| HS2 | TTCCTACACATTAACGAGCCTCTGC, AACATCTGGCCACACACCCTAAGC |
| HS3 | TGTTTCCCTGATGAGGATTCAATGG, CCCACACATGGTCATCTATCTGAGC |
| HS4 | CCCTCTCTTACATCTAGCAGCAATG, GCATGAAGTTCCTGCCTCTAAATC |
| ɛy | CAAAGAGAGTTTTTGTTGAAGGAGGAG, AAAGTTCACCATGATGGCAAGTCTGG |
| βH1 | CAAGGTCCAGGGTGAAGAATAAAAGG, CAGGAGCCTAAGACATAGTGCATTAGC |
| Necdin | TTTACATAAGCCTAGTGGTACCCTTCC, ATCGCTGTCCTGCATCTCACAGTCG |
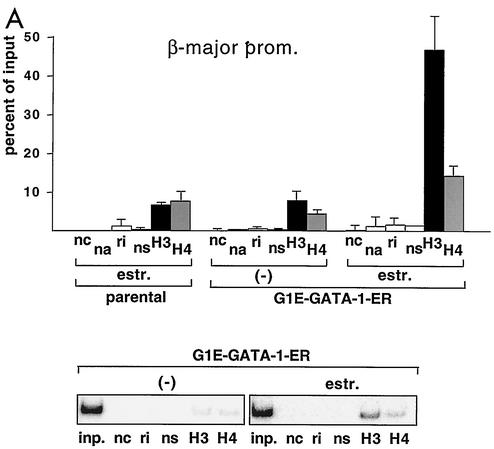
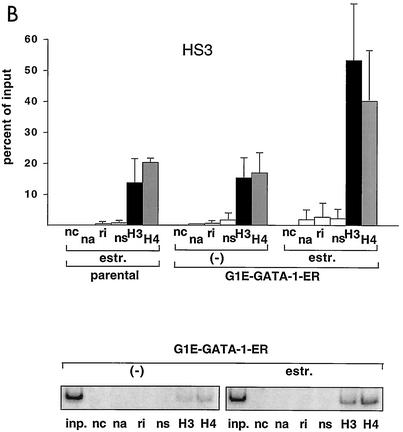
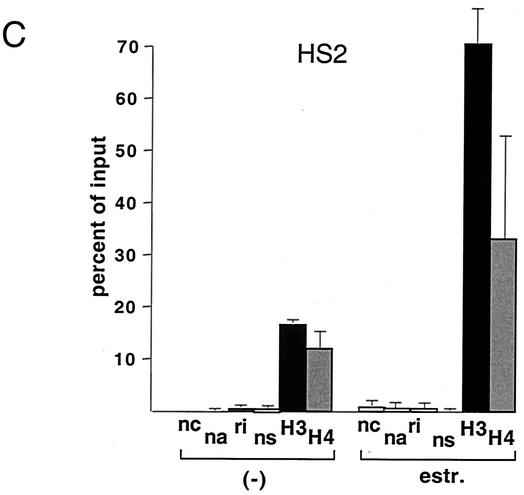
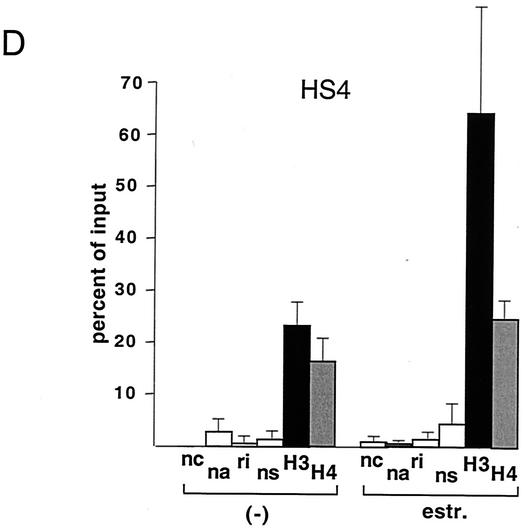
FIG. 1.
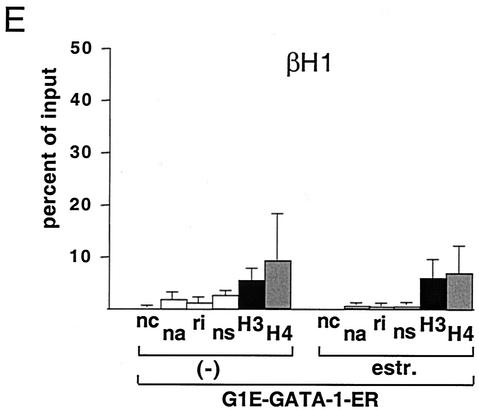
GATA-1-ER stimulates histone acetylation at the β-major globin gene promoter and the LCR. (A) ChIP assay using anti-diacetyl H3 and anti-tetraacetyl H4 antibodies. PCR primers span the β-major globin gene promoter region (see Table 1). Parental G1E cells lacking GATA-1-ER were treated with estradiol (estr.) for 72 h. G1E GATA-1-ER cells were left untreated (−) or were treated with estradiol for 72 h. Results are plotted as the percentage of nonprecipitated chromatin (input). Controls include no chromatin (nc), no antibody (na), rabbit nonimmune immunoglobulin G (ri), and rabbit nonimmune serum (ns). Results are averages of eight independent experiments. Error bars indicate standard deviations. The bottom panel is a representative polyacrylamide gel showing PCR products. (B) ChIP experiments using primers for HS3. Controls are as in panel A. Averages of six independent experiments are shown. Bottom panel: representative polyacrylamide gel showing PCR products. (C and D) Same experiment as shown in panel B, except that primers for HS2 and HS4 were used, respectively. Results are averages of three independent experiments. (E) Analysis of histone H3 and H4 acetylation at the embryonic βH1 globin gene. Results are averages of three independent experiments.
Antibodies.
Anti-diacetyl H3 and anti-tetraacetyl H4 antibodies were purchased from Upstate Biotechnology, and anti-GATA-1 (N6) and anti-CBP antibodies (A22) were obtained from Santa Cruz.
Northern blotting.
Total RNA was extracted using Trizol (Invitrogen). Equal amounts of RNA were loaded into each lane, separated on a 1.2% agarose gel, and transferred onto a nitrocellulose membrane. The membrane was probed with a full-length 32P-labeled β-major globin cDNA probe.
RESULTS AND DISCUSSION
Stimulation of histone acetylation by GATA-1 at the β-globin locus.
G1E cells are immature erythroid cells derived from in vitro-differentiated murine embryonic stem cells that have been rendered null for GATA-1 through gene targeting (33). Restoration of GATA-1 activity in G1E cells leads to their differentiation, as defined by expression of globins and other erythroid-restricted gene products, cell cycle arrest, and mature cell morphology (33). G1E cells thus provide a useful system to study GATA-1 function in its natural environment. To restore GATA-1 activity in all erythroid cells in a synchronous fashion, G1E cells were engineered to stably express GATA-1 fused to the ligand-binding domain (LBD) of the human estrogen receptor (ER) (G1E GATA-1-ER). G1E GATA-1-ER clones were selected that express GATA-1-ER protein at levels comparable to GATA-1 in normal erythroid cells (data not shown). Addition of estradiol to G1E GATA-1-ER cells leads to rapid, synchronous differentiation, thus mimicking the function of wild-type GATA-1 (10, 31). Using the ChIP assay with antibodies against diacetyl histone H3 and tetraacetyl histone H4, we measured acetylation of histones H3 and H4 at the globin gene locus in G1E GATA-1-ER cells. Activation of GATA-1 by treatment with estradiol for 72 h stimulated acetylation of histones H3 and H4 by ∼6-fold and ∼3-fold, respectively, at the β-major gene promoter (Fig. 1A). To determine whether GATA-1 also regulates histone acetylation at the LCR, PCR primer pairs were used to amplify sequences corresponding to HS3 of the LCR. The results showed that GATA-1-ER also stimulated acetylation of histones H3 (3.5-fold) and H4 (2.4-fold) at this site (Fig. 1B). At HS2, GATA-1-ER activation led to a 4.3-fold and 2.8-fold stimulation in acetylation of histones H3 and H4, respectively, while at HS4 the increases were 3-fold and 1.5-fold in H3 and H4 acetylation, respectively (Fig. 1C and D). Finally, preliminary data suggest that GATA-1 also augments acetylation of histones H3 and H4 at HS1 (data not shown). Thus, GATA-1 appears to be limiting for histone acetylation at the β-globin gene promoter and throughout the LCR, consistent with the presence of GATA-1 protein at these sites in vivo (19). In general, GATA-1-ER-induced changes in histone acetylation were more pronounced at the β-major gene promoter than the LCR. Given the abundance of transcription factor binding sites at the LCR, this might reflect partial compensation of acetyltransferase-recruiting activity by other transcriptional regulators. Consistent with this notion is the observation that histone acetylation levels in the absence of GATA-1 are higher at the LCR than at the β-major promoter. Of note, the levels of histone acetylation in the absence of estradiol were similar to those observed in estradiol-treated parental cells lacking GATA-1. Thus, in the absence of estradiol, GATA-1-ER appears to be inactive despite the presence of GATA-1-ER protein (see Fig. 5A).
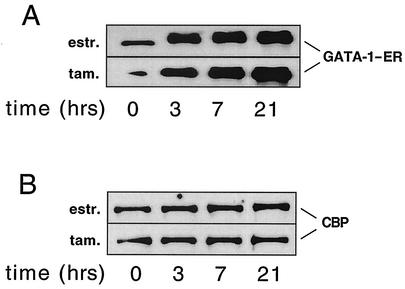
FIG. 5.
Nuclear extracts from cells treated with estradiol for the indicated amounts of time were analyzed by Western blotting with anti-GATA-1 (A) and anti-CBP (B) antibodies.
G1E cells represent definitive erythroid precursor cells, as reflected in their expression of adult-type globin genes (33). To determine whether the effects of GATA-1-ER on histone acetylation were specific for adult globin genes, we measured histone acetylation at the silent embryonic βH1 globin gene. The results show that GATA-1-ER activation did not appreciably augment acetylation of histones H3 and H4 (Fig. 1E). This indicates that the GATA-1-induced increases in histone acetylation at the β-major promoter and the LCR are specific and are not simply the consequence of maturation-induced global changes in histone acetylation.
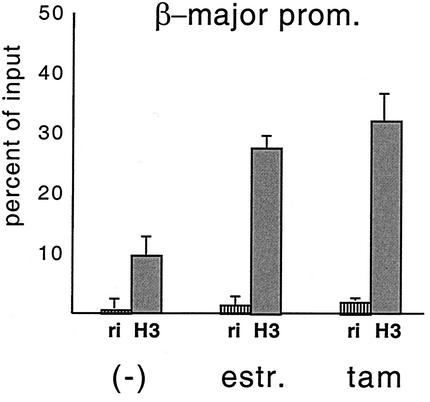
Fusion of transcription factors to the LBD of the ER might affect their function, since the LBD forms ligand-dependent interactions with transcription coactivators, some of which possess histone acetyltransferase activity (25). Thus, we determined whether the biological function of GATA-1 is altered due to the presence of the ER LBD. In contrast to the effects of estradiol, tamoxifen treatment not only fails to induce coactivator recruitment by the ER LBD but also triggers recruitment of corepressor complexes (11, 30). Hence, properties conferred to GATA-1 by the ER portion would be revealed by phenotypic and molecular differences upon treatment with tamoxifen instead of estradiol. When G1E GATA-1-ER cells were treated with tamoxifen, cells underwent differentiation, cell cycle arrest, and globin gene induction in a manner similar to that observed upon estradiol treatment (see Fig. 4 and data not shown), consistent with previous results (10, 31, 33). ChIP experiments performed with tamoxifen-treated G1E GATA-1-ER cells showed a comparable increase in histone H3 and H4 acetylation at the β-major globin gene promoter (Fig. 2) and the HS3 of the LCR (data not shown) compared to that with estradiol-treated cells. Thus, the effects of GATA-1-ER on cellular differentiation and histone acetylation are the result of the action of the GATA-1 moiety and not of the ER portion. This interpretation is further supported by the observation that GATA-1 and GATA-1-ER have the same biological activities in G1E cells (33) and primary differentiating erythroid cells (13). Moreover, GATA-1 and GATA-1-ER show the same DNA binding characteristics at the β-globin locus in vivo (19). Taken together, these results are consistent with GATA-1 being a critical regulator of histone acetylation at the LCR and β-globin promoter. However, it is possible that GATA-1 provides essential transcriptional activation functions in addition to its ability to recruit acetyltransferase activity to the locus.
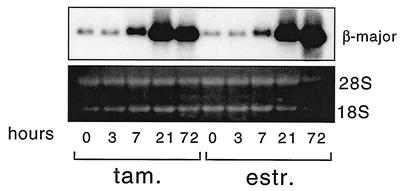
FIG. 4.
Northern analysis of RNA derived from G1E GATA-1-ER cells treated with estradiol (estr.) and tamoxifen (tam.) for the indicated number of hours. Blots were probed with β-major globin cDNA. 28S and 18S rRNAs are shown to indicate equal amounts of RNA loading.
FIG. 2.
Tamoxifen and estradiol treatment of G1E GATA-1-ER cells for 72 h elicits comparable increases in histone H3 acetylation at the β-major promoter. The mean values of two independent experiments are shown.
Histone acetylation occurs rapidly following GATA-1 activation.
GATA-1 controls the expression of numerous erythroid genes, including the erythroid transcription factors EKLF and NF-E2. Therefore, it is possible that changes in histone acetylation are an indirect consequence of GATA-1 activity rather than a direct effect resulting from GATA-1-mediated recruitment of acetyltransferase activity. To address this issue, we performed time course ChIP experiments using the anti-acetyl histone antibodies described above. G1E GATA-1-ER cells were harvested for ChIP analysis following 3, 7, 21, and 72 h of estradiol treatment. The results in Fig. 3 show that histone acetylation at both the β-major promoter and HS3 was substantially increased as early as 7 h following GATA-1-ER activation (Fig. 3).
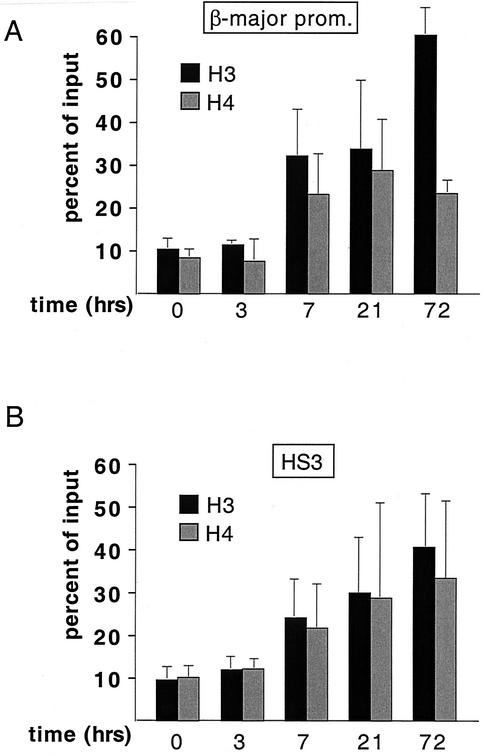
FIG. 3.
Stimulation of histone H3 and H4 acetylation rapidly follows activation of GATA-1-ER at the β-major promoter (A) and HS3 (B). Cells were treated with estradiol for the indicated amounts of time. The results are averages of three independent experiments.
Histone acetylation precedes transcriptional activation at the globin gene locus during development and can even occur in the absence of active globin gene transcription (8, 12, 28, 29). This suggests that histone acetylation is a prerequisite rather than a consequence of active transcription. To examine the relationship between histone acetylation and GATA-1-controlled globin gene activation, we measured β-major globin mRNA levels at various time points following GATA-1-ER activation. The Northern analysis in Fig. 4 shows that β-major globin mRNA accumulation begins at 7 h following GATA-1 activation, peaking at 21 h. Thus, maximal β-major globin mRNA accumulation coincides or slightly lags behind the peak in histone acetylation, supporting the notion that histone acetylation is a prerequisite for globin gene transcription. These results further suggest that GATA-1-mediated histone acetylation might be limiting for globin gene expression.
Estradiol treatment not only facilitates DNA binding of GATA-1-ER but also appears to stabilize the fusion protein (Fig. 5A). Accumulation of GATA-1-ER protein over time slightly preceded the increase in histone acetylation (compare with Fig. 3), suggesting that GATA-1 is directly responsible for the increases of histone acetylation at the β-globin gene locus.
GATA-1 occupancy at the β-globin locus in vivo correlates with histone acetylation.
The above studies suggest that GATA-1 functions by recruiting acetyltransferase activity to the globin locus. To test whether GATA-1-ER binding to the β-globin locus correlates with increased levels of histone acetylation, we determined the kinetics of GATA-1-ER binding to the LCR and the β-globin gene promoter in vivo. ChIP experiments with anti-GATA-1 antibodies showed that GATA-1-ER binding to HS3 can be detected as early as 3 h following estradiol treatment (Fig. 6A), suggesting that GATA-1-ER occupancy precedes the rise in histone acetylation. Very similar results were observed at the β-major globin promoter (data not shown). These results strongly suggest that GATA-1 directly recruits histone acetyltransferase activity to the β-globin locus.
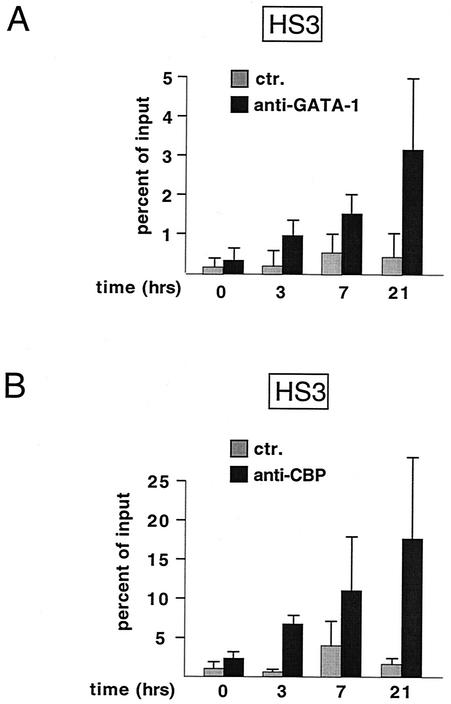
FIG. 6.
GATA-1 occupancy correlates with CBP occupancy at HS3. ChIP analysis of G1E GATA-1-ER cells at indicated time points following estradiol treatment using anti-GATA-1 (A) and anti-CBP (B) antibodies. Note that the presence of GATA-1 and CBP at HS3 precedes increases in histone acetylation (compare with Fig. 3). The results at each time point are the mean of at least three independent experiments.
GATA-1 occupancy at the β-globin locus in vivo correlates with CBP recruitment.
Our laboratory reported previously that GATA-1 binds to the acetyltransferase CBP in vitro and in vivo (4). CBP acetylates both histones H3 and H4, while most other acetyltransferases show a preference for either H3 or H4 (for a review, see reference 23), supporting a model in which CBP is the enzyme responsible for GATA-1-induced histone H3 and H4 acetylation. To test this model directly, a time course ChIP experiment was performed using anti-CBP antibodies. Association of CBP with HS3 was detectable as early as 3 h following estradiol treatment and peaked at 21 h (Fig. 6B). Thus, the presence of CBP at the locus correlated well with that of GATA-1. Together with our previous studies showing a physical interaction between GATA-1 and CBP (4), these results strongly suggest that GATA-1 directly recruits CBP to the globin gene locus, leading to hyperacetylation of histones. CBP protein levels did not change significantly throughout G1E cell differentiation (Fig. 5B), supporting the model that GATA-1 recruits CBP to select sites rather than globally enhancing histone acetylation through stimulation of CBP expression. In agreement with this interpretation, GATA-1-ER activation did not affect the levels of histone acetylation at the active housekeeping gene glyceraldehyde-3-phosphate dehydrogenase (data not shown).
Low levels of globin gene expression are observed in primary GATA-1-deficient erythroblasts (32) and in G1E cells without GATA-1 (Fig. 4). In agreement with these findings, we observed that histone acetylation at the globin locus in nonrescued G1E cells was elevated when compared to the brain-specific necdin gene that is silent in the erythroid lineage (data not shown). This suggests that additional erythroid transcription factors mediate GATA-1-independent histone acetylation at the β-globin locus. Likely candidates include GATA-2, which is expressed in normal hematopoietic progenitors and is elevated more than 20-fold above normal in GATA-1-deficient erythroblasts. Similar to GATA-1, GATA-2 also interacts with CBP (4). Other factors include EKLF and NF-E2, both of which bind cognate elements within the globin locus and interact with CBP/p300 (6, 9, 17, 35). It appears that the basal level of acetylation observed in the absence of GATA-1 is insufficient to support high-level globin gene expression, consistent with the model that acetylation is limiting for gene transcription. This interpretation is in agreement with the presence of elevated levels of histone acetylation in erythroid cells at silent globin genes and the peaks of acetylation found at the active gene promoters.
While the time course experiments strongly suggest that the effects of GATA-1 and CBP are direct, they do not rule out that the products of certain GATA-1 target genes, such as EKLF and NF-E2, contribute to the recruitment of acetyltransferases to the β-globin locus. However, an essential role for GATA-1 in this process is further supported by the observation that, in the absence of GATA-1, a substantial amount of NF-E2 is already present at HS2 (19), yet hyperacetylation at HS2 is still GATA-1 dependent. Furthermore, both NF-E2 and EKLF proteins are present in G1E cells lacking GATA-1 and do not undergo substantial increases upon GATA-1 activation (20). Thus, GATA-1 appears to be limiting for acetyltransferase recruitment.
One interesting finding of this study is the observation that GATA-1 appears to mediate acetylation of both histones H3 and H4. Since CBP/p300 acetylate both of these histones in vitro, this supports a model in which CBP/p300 are the acetyltransferases recruited by GATA-1 to the β-globin locus. While the anti-CBP ChIP experiments confirmed this model, it remains possible that additional acetyltransferases participate in establishing specific patterns of histone acetylation at the globin gene locus. In order to detect any changes in histone acetylation, our ChIP experiments were designed to cast a broad net by using antibodies that recognize most acetylated forms of histones H3 and H4. These antibodies fail to distinguish between the individual acetylated residues within histones H3 and H4. Therefore, a more detailed characterization using specific antibodies against each acetylated lysine residue might reveal whether the acetylated sites exactly match those acetylated by CBP/p300 in vitro. If GATA-1 mediates acetylation of additional lysine residues not targeted by CBP, this might implicate the presence of additional acetyltransferases at the globin locus. Experiments are under way to address these questions.
Acknowledgments
We thank E. Bresnick, M. Crossley, and S. Liebhaber for critical comments on the manuscript and M. Groudine for insightful discussions.
G.A.B. is supported by NIH grant DK54937, and D.L.L. is supported by NIH training grant T32GM008216.
REFERENCES
- 1.Anguita, E., C. A. Johnson, W. G. Wood, B. M. Turner, and D. R. Higgs. 2001. Identification of a conserved erythroid specific domain of histone acetylation across the alpha-globin gene cluster. Proc. Natl. Acad. Sci. USA 98:12114-12119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bender, M. A., M. Bulger, J. Close, and M. Groudine. 2000. Beta-globin gene switching and DNase I sensitivity of the endogenous beta-globin locus in mice do not require the locus control region. Mol. Cell 5:387-393. [DOI] [PubMed] [Google Scholar]
- 3.Blobel, G. A. 2000. CBP/p300: molecular integrators of hematopoietic transcription. Blood 95:745-755. [PubMed] [Google Scholar]
- 4.Blobel, G. A., T. Nakajima, R. Eckner, M. Montminy, and S. H. Orkin. 1998. CREB-binding protein (CBP) cooperates with transcription factor GATA-1 and is required for erythroid differentiation. Proc. Natl. Acad. Sci. USA 95:2061-2066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bulger, M., T. Sawado, D. Schubeler, and M. Groudine. 2002. ChIPs of the beta-globin locus: unraveling gene regulation within an active domain. Curr. Opin. Genet. Dev. 12:170-177. [DOI] [PubMed] [Google Scholar]
- 6.Cheng, X., M. J. Reginato, N. C. Andrews, and M. A. Lazar. 1997. The transcriptional integrator CREB-binding protein mediates positive cross talk between nuclear hormone receptors and the hematopoietic bZip protein p45/NF-E2. Mol. Cell. Biol. 17:1407-1416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Engel, J. D., and K. Tanimoto. 2000. Looping, linking, and chromatin activity: new insights into beta-globin locus regulation. Cell 100:499-502. [DOI] [PubMed] [Google Scholar]
- 8.Forsberg, E. C., K. M. Downs, H. M. Christensen, H. Im, P. A. Nuzzi, and E. H. Bresnick. 2000. Developmentally dynamic histone acetylation pattern of a tissue-specific chromatin domain. Proc. Natl. Acad. Sci. USA 97:14494-14499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Forsberg, E. C., K. Johnson, T. N. Zaboikina, E. A. Mosser, and E. H. Bresnick. 1999. Requirement of an E1A-sensitive coactivator for long-range transactivation by the β-globin locus control region. J. Biol. Chem. 274:26850-26859. [DOI] [PubMed] [Google Scholar]
- 10.Gregory, T., C. Yu, A. Ma, S. H. Orkin, G. A. Blobel, and M. J. Weiss. 1999. GATA-1 and erythropoietin cooperate to promote erythroid cell survival by regulating bcl-xL expression. Blood 94:87-96. [PubMed] [Google Scholar]
- 11.Halachmi, S., E. Marden, G. Martin, H. MacKay, C. Abbondanza, and M. Brown. 1994. Estrogen receptor-associated proteins: possible mediators of hormone-induced transcription. Science 264:1455-1458. [DOI] [PubMed] [Google Scholar]
- 12.Hebbes, T. R., A. L. Clayton, A. W. Thorne, and C. Crane-Robinson. 1994. Core histone hyperacetylation co-maps with generalized DNase I sensitivity in the chicken β-globin chromosomal domain. EMBO J. 13:1823-1830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Heyworth, C., S. Pearson, G. May, and T. Enver. 2002. Transcription factor-mediated lineage switching reveals plasticity in primary committed progenitor cells. EMBO J. 21:3770-3781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Higgs, D. R. 1998. Do LCRs open chromatin domains? Cell 95:299-302. [DOI] [PubMed] [Google Scholar]
- 15.Hirasawa, R., R. Shimizu, S. Takahashi, M. Osawa, S. Takayanagi, Y. Kato, M. Onodera, N. Minegishi, M. Yamamoto, K. Fukao, H. Taniguchi, H. Nakauchi, and A. Iwama. 2002. Essential and instructive roles of GATA factors in eosinophil development. J. Exp. Med. 195:1379-1386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Horak, C. E., M. C. Mahajan, N. M. Luscombe, M. Gerstein, S. M. Weissman, and M. Snyder. 2002. GATA-1 binding sites mapped in the beta-globin locus by using mammalian chIp-chip analysis. Proc. Natl. Acad. Sci. USA 99:2924-2929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hung, H. L., A. Y. Kim, W. Hong, C. Rakowski, and G. A. Blobel. 2001. Stimulation of NF-E2 DNA binding by CREB-binding protein (CBP)-mediated acetylation. J. Biol. Chem. 276:10715-10721. [DOI] [PubMed] [Google Scholar]
- 18.Johnson, K. D., H. M. Christensen, B. Zhao, and E. H. Bresnick. 2001. Distinct mechanisms control RNA polymerase II recruitment to a tissue-specific locus control region and a downstream promoter. Mol. Cell 8:465-471. [DOI] [PubMed] [Google Scholar]
- 19.Johnson, K. D., J. A. Grass, M. E. Boyer, C. M. Kiekhaefer, G. A. Blobel, M. J. Weiss, and E. H. Bresnick. 2002. Cooperative activities of hematopoietic regulators recruit RNA polymerase II to a tissue-specific chromatin domain. Proc. Natl. Acad. Sci. USA 99:11760-11765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kiekhaefer, C. M., J. A. Grass, K. D. Johnson, M. E. Boyer, and E. H. Bresnick. 2002. Hematopoietic-specific activators establish an overlapping pattern of histone acetylation and methylation within a mammalian chromatin domain. Proc. Natl. Acad. Sci. USA 99:14309-14314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Levings, P. P., and J. Bungert. 2002. The human beta-globin locus control region. Eur. J. Biochem. 269:1589-1599. [DOI] [PubMed] [Google Scholar]
- 22.Litt, M. D., M. Simpson, F. Recillas-Targa, M. N. Prioleau, and G. Felsenfeld. 2001. Transitions in histone acetylation reveal boundaries of three separately regulated neighboring loci. EMBO J. 20:2224-2235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Marmorstein, R. 2001. Structure of histone acetyltransferases. J. Mol. Biol. 311:433-444. [DOI] [PubMed] [Google Scholar]
- 24.Martin, D. I., S. Fiering, and M. Groudine. 1996. Regulation of beta-globin gene expression: straightening out the locus. Curr. Opin. Genet. Dev. 6:488-495. [DOI] [PubMed] [Google Scholar]
- 25.McDonnell, D. P., and J. D. Norris. 2002. Connections and regulation of the human estrogen receptor. Science 296:1642-1644. [DOI] [PubMed] [Google Scholar]
- 26.Myers, F. A., D. R. Evans, A. L. Clayton, A. W. Thorne, and C. Crane-Robinson. 2001. Targeted and extended acetylation of histones H4 and H3 at active and inactive genes in chicken embryo erythrocytes. J. Biol. Chem. 276:20197-20205. [DOI] [PubMed] [Google Scholar]
- 27.Reik, A., A. Telling, G. Zitnik, D. Cimbora, E. Epner, and M. Groudine. 1998. The locus control region is necessary for gene expression in the human beta-globin locus but not for the maintenance of an open chromatin structure in erythroid cells. Mol. Cell. Biol. 18:5992-6000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Schubeler, D., C. Francastel, D. M. Cimbora, A. Reik, D. I. Martin, and M. Groudine. 2000. Nuclear localization and histone acetylation: a pathway for chromatin opening and transcriptional activation of the human beta-globin locus. Genes Dev. 14:940-950. [PMC free article] [PubMed] [Google Scholar]
- 29.Schubeler, D., M. Groudine, and M. A. Bender. 2001. The murine beta-globin locus control region regulates the rate of transcription but not the hyperacetylation of histones at the active genes. Proc. Natl. Acad. Sci. USA 98:11432-11437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Shang, Y., X. Hu, J. DiRenzo, M. A. Lazar, and M. Brown. 2000. Cofactor dynamics and sufficiency in estrogen receptor-regulated transcription. Cell 103:843-852. [DOI] [PubMed] [Google Scholar]
- 31.Tsang, A. P., J. E. Visvader, C. A. Turner, Y. Fujiwara, C. Yu, M. J. Weiss, M. Crossley, and S. H. Orkin. 1997. FOG, a multitype zinc finger protein, acts as a cofactor for transcription factor GATA-1 in erythroid and megakaryocytic differentiation. Cell 90:109-119. [DOI] [PubMed] [Google Scholar]
- 32.Weiss, M. J., G. Keller, and S. H. Orkin. 1994. Novel insights into erythroid development revealed through in vitro differentiation of GATA-1 embryonic stem cells. Genes Dev. 8:1184-1197. [DOI] [PubMed] [Google Scholar]
- 33.Weiss, M. J., C. Yu, and S. H. Orkin. 1997. Erythroid-cell-specific properties of transcription factor GATA-1 revealed by phenotypic rescue of a gene-targeted cell line. Mol. Cell. Biol. 17:1642-1651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Yu, C., A. B. Cantor, H. Yang, C. Browne, R. A. Wells, Y. Fujiwara, and S. H. Orkin. 2002. Targeted deletion of a high-affinity GATA-binding site in the GATA-1 promoter leads to selective loss of the eosinophil lineage in vivo. J. Exp. Med. 195:1387-1395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Zhang, W., and J. J. Bieker. 1998. Acetylation and modulation of erythroid Kruppel-like factor (EKLF) activity by interaction with histone acetyltransferases. Proc. Natl. Acad. Sci. USA 95:9855-9860. [DOI] [PMC free article] [PubMed] [Google Scholar]